Abstract
Current knowledge of genotypic and phenotypic diversity in the species Escherichia coli is based almost entirely on strains recovered from humans or zoo animals. In this study, we analyzed a collection of 202 strains obtained from 81 mammalian species representing 39 families and 14 orders in Australia and the Americas, as well as several reference strains; we also included a strain from a reptile and 10 from different families of birds collected in Mexico. The strains were characterized genotypically by multilocus enzyme electrophoresis (MLEE) and phenotypically by patterns of sugar utilization, antibiotic resistance, and plasmid profile. MLEE analysis yielded an estimated genetic diversity (H) of 0.682 for 11 loci. The observed genetic diversity in this sample is the greatest yet reported for E. coli. However, this genetic diversity is not randomly distributed; geographic effects and host taxonomic group accounted for most of the genetic differentiation. The genetic relationship among the strains showed that they are more associated by origin and host order than is expected by chance. In a dendrogram, the ancestral cluster includes primarily strains from Australia and ECOR strains from groups B and C. The most differentiated E. coli in our analysis are strains from Mexican carnivores and strains from humans, including those in the ECOR group A. The kinds and numbers of sugars utilized by the strains varied by host taxonomic group and country of origin. Strains isolated from bats were found to exploit the greatest range of sugars, while those from primates utilized the fewest. Toxins are more frequent in strains from rodents from both continents than in any other taxonomic group. Strains from Mexican wild mammals were, on average, as resistant to antibiotics as strains from humans in cities. On average, the Australian strains presented a lower antibiotic resistance than the Mexican strains. However, strains recovered from hosts in cities carried significantly more plasmids than did strains isolated from wild mammals. Previous studies have shown that natural populations of E. coli harbor an extensive genetic diversity that is organized in a limited number of clones. However, knowledge of this worldwide bacterium has been limited. Here, we suggest that the strains from a wide range of wild hosts from different regions of the world are organized in an ecotypic structure where adaptation to the host plays an important role in the population structure.
Escherichia coli is an abundant and usually harmless commensal of the human digestive flora (42). Nevertheless, pathogenic strains are an important cause of sickness and mortality throughout the world, particularly for children in underdeveloped countries (8). E. coli is also a common member of the microbial commensal community of mammals and birds. Surprisingly, little is known about the natural history and genetic structure of E. coli in populations of wild animals (42, 50). The standard reference collection of strains for this species, the ECOR collection (32), contains a number of strains isolated from five nonhuman mammalian orders, but most of the strains were recovered from domestic or zoo animals (32). The relevance of this limited sample has been questioned on the basis of possible cross-contamination from human host (42).
The first attempt to assess the genetic structure and diversity of E. coli was made by Milkman, who analyzed 829 isolates obtained mainly from humans (26). Using multilocus enzyme electrophoresis (MLEE) based on four loci, he determined that the average genetic diversity (H) of this species was 0.23. Subsequent studies that extended the work of Milkman were primarily concerned with the genotypic and phenotypic variation among strains isolated from the commensal fecal flora of humans and those responsible for neonatal septicemia, cystitis, pyelonephritis, and acute diarrhea (6, 38, 41, 47, 49, 50). Other studies have focused on various aspects of this species’ natural history, such as the turnover of strains in a single host (5), the sharing of clones among hosts (6), and the genetic structure and diversity of E. coli in its primary and secondary habitats (33, 46, 48). These and other studies have formed the basis of the clonal paradigm for the genetic structure of bacterial populations (10, 15, 16, 19, 25, 45) and have been reviewed by Selander et al. (42) and by Whittam (50).
Estimates of genetic diversity obtained by using strains isolated from human fecal samples range from 0.45 to 0.54 (42, 50). However, there is a caveat associated with this observation. The majority of strains from humans were isolated from people living in the developed countries of the West. Although some studies have included strains from Tonga, even these are suspect, since Tonga was used as a military base by both U.S. and New Zealand forces during World War II (4). It is therefore possible that the diversity of E. coli from humans is also underestimated. A high degree of genetic diversity (H = 0.61) has been found in E. coli isolated from sewage (33). These sewage samples undoubtedly represent two quite distinct sources of strains: those from the species’ primary habitat, the lower intestinal tract, and those from its secondary habitat, the sewage environment. Whittam and collaborators (46–48) have shown that sympatric E. coli populations inhabiting primary and secondary habitats are quite distinct from one another in terms of their clonal composition (50).
The objective of this study was to extend our understanding of the variation and genetic structure of E. coli by examining strains from a large variety of wild mammalian and avian hosts. We assessed genotypic diversity, phenotypic diversity, and genetic relatedness of 202 E. coli strains by using 11 loci in MLEE, plasmid profiles, resistance to six antibiotics, production of two toxins, and the utilization of 12 sugars. In this study, we analyzed how these genetic and phenotypic characteristics varied by host taxonomic group and geographic origin. E. coli was sampled mostly from wild mammals, and strains were taken from 81 species representing 39 families and 14 orders of mammals (31) in the Americas and Australia. We also included a sample from a reptile and 10 strains from birds from Mexico. As a reference, we also studied six strains from African baboons (35), 13 strains from the ECOR collection (32), and strain K-12 (2).
MATERIALS AND METHODS
Bacterial strains.
The complete list of hosts sampled by diet and geographic origin as well as their taxonomic classification by order and proximity to human environment is presented in Table A1 in the Appendix.
Strain isolation.
Strains collected in Mexico, Costa Rica, and Venezuela were recovered from captive or wild mammals and birds. Fecal samples were transported in swab-transport system containing Aimes media (Difco). Mexican samples were suspended in 1 ml of Luria broth and incubated; an aliquot was then streaked for single colonies on a minimal lactose plate. More than one isolate was sometimes taken from a single host individual. The Mexican samples were processed as follows. After incubation, the Lac+ colonies were tested for growth on minimal citrate plates. The Lac+ Cit− colonies were then tested to confirm that they matched the biochemical characteristics of E. coli: gas production positive, H2S negative, urea negative, methyl red positive, Voges-Proskauer negative (13). The Australian strains were collected and cultured by David Gordon. These strains were isolated from anal or cloacal samples or from fecal samples from wild mammals. A primary isolation of the strains was carried out by streaking the sample on a MacConkey plate to obtain single colonies. Only one isolate was taken from each individual host. Subsequently, single colonies from each plate were restreaked twice onto MacConkey plates. Colonies morphologically consistent with E. coli were then tested for growth on minimal lactose and minimal citrate plates. All Lac+ Cit− colonies were tested to confirm that they matched the biochemical characteristics of E. coli: phenylalanine negative, H2S negative, urea negative, indole positive, methyl red positive, Voges-Proskauer negative (13). In Australia and Mexico, all incubations were carried out overnight at 37°C. Following isolation, all strains were immediately stored at −80°C. All the strains classified by us as E. coli were confirmed as E. coli by serotyping in the Faculty of Medicine, Universidad Nacional Autónoma de Mexico.
In addition to the newly isolated strains, 13 strains from the E. coli reference collection ECOR (32), the sequenced K-12 strain MG1655 (2), and 6 strains from yellow baboons (Papio cynocephalus) were included in the analysis (35).
MLEE analysis.
MLEE using cellulose acetate membranes was carried out in Tris-glycine buffer (pH 8.5) (17). Ten enzymes were selected based on previous studies (40): ADH (alcohol dehydrogenase), ARK (arginine kinase), G6PDH (glucose-6-phosphate dehydrogenase), IDH (isocitrate dehydrogenase), MDH (malate dehydrogenase), ME (malic enzyme), MPI (mannose-6-phosphate isomerase), PEP (peptidase), PGM (phosphoglucomutase), and XDH (xanthine dehydrogenase). Eleven loci were resolved with those enzymes because ME exhibited two loci. All of the strains were examined at least twice to confirm their electrophoretic types (Table 1).
TABLE 1.
E. coli strains examined in this study and their electrophoretic profiles
| Host species (strain reference) or ECOR designationa | Allele number at locus:
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| IDH | PEP | PGM | ARK | ME1 | ME2 | MPI | ADH1 | MDH | 6G | XDH1 | |
| Gopherus flavomarginatus (57) | 0 | 2 | 4 | 1 | 4 | 9 | 7 | 1 | 3 | 3 | 2 |
| Tachyglosus aculeatus (TA309) | 2 | 3 | 3 | 3 | 2 | 6 | 4 | 3 | 3 | 0 | 2 |
| Tachyglosus aculeatus (TA310) | 2 | 3 | 4 | 3 | 2 | 6 | 3 | 3 | 3 | 3 | 3 |
| Tachyglosus aculeatus (TA311) | 3 | 3 | 4 | 3 | 2 | 6 | 3 | 3 | 3 | 3 | 4 |
| Ornithorhynchus anatinus (TA319) | 3 | 3 | 4 | 3 | 3 | 6 | 3 | 3 | 4 | 3 | 4 |
| Ornithorhynchus anatinus (TA32) | 2 | 3 | 3 | 3 | 2 | 6 | 4 | 3 | 3 | 3 | 3 |
| Marmosa camescens (64) | 4 | 3 | 5 | 3 | 3 | 7 | 3 | 3 | 2 | 3 | 4 |
| Didelphis virginiana (79) | 5 | 3 | 5 | 3 | 2 | 5 | 4 | 7 | 2 | 4 | 4 |
| Didelphis virginiana (2048) | 3 | 3 | 5 | 3 | 3 | 6 | 5 | 0 | 3 | 3 | 4 |
| Didelphis virginiana (2049) | 4 | 4 | 3 | 2 | 4 | 6 | 3 | 0 | 3 | 3 | 3 |
| Didelphis virginiana (2050) | 3 | 3 | 5 | 3 | 4 | 7 | 5 | 0 | 2 | 3 | 4 |
| Philander opossum (1698) | 3 | 3 | 3 | 2 | 2 | 7 | 4 | 4 | 3 | 3 | 5 |
| Cercatetus concinnus (TA002) | 2 | 4 | 3 | 2 | 3 | 7 | 3 | 3 | 5 | 3 | 4 |
| Antechinus bellus (TA148) | 2 | 4 | 3 | 2 | 3 | 7 | 3 | 3 | 3 | 3 | 4 |
| Antechinus flavipes (TA237) | 2 | 3 | 2 | 2 | 4 | 0 | 3 | 4 | 3 | 3 | 4 |
| Antechinus swainsonii (TA153) | 1 | 4 | 1 | 1 | 3 | 6 | 1 | 3 | 3 | 3 | 2 |
| Dasyurus geofroii (TA124) | 1 | 3 | 3 | 1 | 3 | 7 | 2 | 2 | 3 | 3 | 3 |
| Dasyurus hallucatus (TA260) | 2 | 4 | 2 | 2 | 1 | 6 | 3 | 0 | 1 | 3 | 4 |
| Dasyurus maculatus (TA036) | 3 | 3 | 2 | 3 | 3 | 6 | 4 | 3 | 3 | 3 | 3 |
| Sminthopsis macroura (TA244) | 2 | 4 | 2 | 2 | 4 | 7 | 5 | 4 | 3 | 3 | 4 |
| Sminthopsis murina (TA151) | 2 | 2 | 2 | 2 | 4 | 7 | 3 | 3 | 3 | 3 | 4 |
| Sminthopsis delichura (TA120) | 1 | 4 | 1 | 1 | 3 | 7 | 1 | 2 | 3 | 3 | 2 |
| Bettongia penicillata (TA115) | 3 | 4 | 4 | 3 | 2 | 6 | 5 | 4 | 3 | 3 | 0 |
| Lagorchestes hirsutus (TA263) | 2 | 3 | 2 | 2 | 3 | 7 | 3 | 0 | 3 | 3 | 4 |
| Macropus eugenii (TA052) | 3 | 3 | 2 | 3 | 3 | 6 | 4 | 3 | 3 | 3 | 3 |
| Macropus fulginosus (TA004) | 2 | 3 | 3 | 2 | 3 | 6 | 3 | 3 | 5 | 3 | 4 |
| Macropus giganteus (TA057) | 2 | 4 | 2 | 2 | 4 | 7 | 3 | 3 | 5 | 2 | 4 |
| Petrogale lateralis (TA135) | 3 | 3 | 0 | 2 | 3 | 6 | 3 | 3 | 3 | 3 | 3 |
| Perameles nasuta (TA020) | 2 | 3 | 2 | 2 | 2 | 7 | 3 | 3 | 4 | 2 | 4 |
| Petaurus breviceps (TA243) | 1 | 4 | 2 | 2 | 2 | 4 | 3 | 4 | 3 | 3 | 4 |
| Pseudocheirus peregrinus (TA022) | 2 | 4 | 2 | 2 | 3 | 7 | 3 | 3 | 5 | 3 | 4 |
| Trichosurus vulpecula (TA038) | 2 | 4 | 3 | 2 | 3 | 5 | 3 | 3 | 5 | 3 | 4 |
| Phascolarctos cinereus (TA003) | 1 | 3 | 1 | 1 | 2 | 6 | 2 | 2 | 3 | 3 | 2 |
| Sorex sp. (238) | 2 | 1 | 1 | 2 | 2 | 5 | 3 | 4 | 2 | 2 | 2 |
| Sorex sp. (240) | 6 | 2 | 4 | 2 | 3 | 8 | 7 | 0 | 4 | 3 | 4 |
| Tachyglosus aculeatus (TA051) | 2 | 4 | 3 | 2 | 4 | 7 | 4 | 3 | 5 | 2 | 4 |
| Tadarida brasiliensis (259) | 5 | 1 | 3 | 3 | 3 | 7 | 2 | 3 | 3 | 0 | 4 |
| Tadarida brasiliensis (261) | 5 | 1 | 3 | 3 | 3 | 7 | 2 | 3 | 4 | 3 | 4 |
| Tadarida brasiliensis (266) | 4 | 1 | 3 | 2 | 3 | 6 | 2 | 3 | 3 | 3 | 4 |
| Desmodus rotundus (25) | 4 | 2 | 0 | 4 | 3 | 5 | 1 | 3 | 2 | 3 | 4 |
| Desmodus rotundus (26) | 4 | 2 | 2 | 4 | 3 | 5 | 1 | 3 | 2 | 3 | 5 |
| Leptonycteris nivalis (31) | 3 | 2 | 2 | 2 | 3 | 5 | 1 | 2 | 2 | 2 | 3 |
| Leptonycteris nivalis (33) | 3 | 2 | 2 | 2 | 3 | 3 | 3 | 2 | 4 | 3 | 2 |
| Leptonycteris nivalis (36) | 3 | 1 | 3 | 2 | 3 | 5 | 5 | 2 | 6 | 4 | 2 |
| Carollia brevicauda (40) | 3 | 1 | 3 | 2 | 3 | 5 | 4 | 2 | 6 | 2 | 2 |
| Carollia brevicauda (41) | 0 | 1 | 3 | 2 | 3 | 5 | 4 | 2 | 6 | 3 | 2 |
| Carollia brevicauda (42) | 2 | 1 | 4 | 2 | 3 | 6 | 4 | 2 | 6 | 3 | 4 |
| Nyctophilus geoffroyi (TA225) | 2 | 2 | 2 | 2 | 4 | 0 | 2 | 4 | 3 | 3 | 4 |
| Alouatta palliata (1638) | 1 | 4 | 4 | 2 | 4 | 6 | 6 | 5 | 2 | 3 | 3 |
| Alouatta palliata (1639) | 2 | 4 | 4 | 2 | 4 | 6 | 6 | 5 | 2 | 3 | 4 |
| Alouatta palliata (1640) | 1 | 4 | 4 | 2 | 4 | 6 | 6 | 5 | 2 | 3 | 5 |
| Alouatta palliata (1731) | 3 | 5 | 4 | 4 | 3 | 3 | 4 | 0 | 2 | 3 | 3 |
| Alouatta pigra (1699) | 4 | 4 | 5 | 3 | 4 | 4 | 5 | 0 | 4 | 3 | 2 |
| Ateles geoffroyi (1669) | 0 | 2 | 2 | 2 | 2 | 6 | 3 | 5 | 4 | 4 | 2 |
| Papio cynocephalus (B184) | 3 | 3 | 2 | 2 | 3 | 7 | 3 | 3 | 3 | 4 | 4 |
| Papio cynocephalus (B197) | 2 | 3 | 0 | 2 | 3 | 7 | 3 | 3 | 3 | 4 | 4 |
| Papio cynocephalus (B307) | 2 | 3 | 1 | 2 | 3 | 6 | 3 | 3 | 2 | 3 | 4 |
| Papio cynocephalus (B349) | 2 | 3 | 2 | 2 | 3 | 7 | 4 | 3 | 3 | 4 | 4 |
| Papio cynocephalus (B427) | 3 | 3 | 2 | 2 | 1 | 4 | 3 | 3 | 1 | 3 | 4 |
| Papio cynocephalus (B430) | 3 | 3 | 2 | 3 | 3 | 7 | 3 | 3 | 3 | 3 | 4 |
| ECOR7 | 3 | 4 | 4 | 2 | 3 | 7 | 4 | 3 | 3 | 3 | 4 |
| Homo sapiens (45) | 3 | 2 | 2 | 3 | 3 | 5 | 4 | 0 | 3 | 3 | 3 |
| Homo sapiens (46) | 4 | 2 | 2 | 3 | 2 | 5 | 4 | 0 | 3 | 3 | 3 |
| Homo sapiens (47) | 1 | 2 | 3 | 3 | 2 | 5 | 4 | 0 | 3 | 3 | 3 |
| ECOR1 | 4 | 3 | 5 | 3 | 3 | 7 | 4 | 3 | 3 | 3 | 4 |
| ECOR5 | 5 | 3 | 2 | 5 | 4 | 8 | 6 | 3 | 2 | 3 | 3 |
| ECOR8 | 5 | 3 | 3 | 0 | 3 | 7 | 3 | 3 | 3 | 3 | 0 |
| ECOR10 | 3 | 3 | 5 | 5 | 4 | 7 | 4 | 3 | 3 | 3 | 4 |
| ECOR11 | 3 | 3 | 5 | 3 | 3 | 5 | 0 | 3 | 3 | 3 | 4 |
| ECOR12 | 4 | 3 | 5 | 3 | 3 | 6 | 5 | 4 | 3 | 3 | 4 |
| ECOR26 | 2 | 3 | 4 | 2 | 3 | 7 | 3 | 2 | 3 | 3 | 3 |
| ECOR41 | 2 | 3 | 3 | 2 | 1 | 6 | 3 | 2 | 1 | 3 | 3 |
| K-12 MG1655 | 3 | 4 | 5 | 3 | 3 | 5 | 0 | 3 | 3 | 3 | 4 |
| Sylvilagus sp. (80) | 3 | 3 | 3 | 3 | 4 | 6 | 4 | 7 | 4 | 0 | 4 |
| Oryctolagus cuniculus (2355) | 2 | 4 | 3 | 2 | 3 | 7 | 3 | 3 | 3 | 3 | 4 |
| Oryctolagus cuniculus (2356) | 2 | 0 | 3 | 2 | 2 | 6 | 3 | 3 | 2 | 3 | 4 |
| Oryctolagus cuniculus (2357) | 2 | 5 | 3 | 2 | 4 | 8 | 3 | 3 | 2 | 3 | 4 |
| Romerolagus diazi (2372) | 2 | 3 | 3 | 2 | 3 | 6 | 3 | 3 | 2 | 3 | 4 |
| Romerolagus diazi (2373) | 2 | 3 | 3 | 2 | 3 | 7 | 3 | 3 | 2 | 3 | 4 |
| Romerolagus diazi (2374) | 2 | 3 | 3 | 2 | 3 | 7 | 3 | 3 | 2 | 3 | 4 |
| Dasypus sp. (2395) | 3 | 3 | 1 | 1 | 2 | 6 | 3 | 2 | 3 | 3 | 2 |
| Dasypus sp. (2396) | 1 | 3 | 1 | 1 | 2 | 7 | 3 | 2 | 3 | 4 | 2 |
| Nyctomys sp. (69) | 1 | 3 | 3 | 2 | 3 | 3 | 1 | 3 | 4 | 3 | 4 |
| Baiomys musculus (75) | 3 | 2 | 3 | 2 | 3 | 5 | 5 | 6 | 4 | 4 | 4 |
| Habromys sp. (88) | 4 | 3 | 3 | 2 | 4 | 4 | 5 | 5 | 5 | 4 | 4 |
| Habromys sp. (89) | 3 | 3 | 3 | 2 | 3 | 5 | 5 | 8 | 5 | 3 | 4 |
| Peromyscus sp. (71) | 3 | 2 | 3 | 2 | 3 | 3 | 4 | 4 | 5 | 3 | 4 |
| Peromyscus sp. (72) | 3 | 2 | 3 | 4 | 3 | 3 | 4 | 4 | 5 | 3 | 4 |
| Peromyscus sp. (73) | 0 | 0 | 2 | 2 | 3 | 3 | 4 | 8 | 2 | 3 | 4 |
| Peromyscus megalops (95) | 2 | 3 | 3 | 3 | 2 | 5 | 6 | 3 | 5 | 3 | 4 |
| Peromyscus boylii (96) | 2 | 3 | 4 | 3 | 3 | 5 | 6 | 5 | 6 | 3 | 4 |
| Peromyscus boylii (97) | 1 | 3 | 4 | 3 | 3 | 5 | 6 | 6 | 6 | 3 | 4 |
| Oryzomys sp. (58) | 2 | 5 | 3 | 2 | 3 | 3 | 6 | 3 | 6 | 4 | 4 |
| Oryzomys sp. (59) | 2 | 2 | 3 | 3 | 2 | 5 | 4 | 5 | 2 | 4 | 4 |
| Oryzomys sp. (60) | 2 | 2 | 3 | 3 | 2 | 5 | 4 | 6 | 6 | 3 | 5 |
| Sigmodon mascotensis (67) | 1 | 3 | 4 | 2 | 3 | 5 | 5 | 3 | 5 | 3 | 2 |
| Sigmodon mascotensis (68) | 2 | 3 | 3 | 2 | 3 | 3 | 5 | 3 | 5 | 3 | 5 |
| Neotoma albigula (296) | 4 | 2 | 4 | 2 | 3 | 7 | 7 | 5 | 4 | 3 | 4 |
| Neotoma albigula (297) | 3 | 2 | 3 | 2 | 2 | 8 | 4 | 3 | 5 | 3 | 2 |
| Neotoma albigula (298) | 6 | 2 | 3 | 2 | 2 | 8 | 4 | 3 | 5 | 3 | 3 |
| Zyzomys argurus (TA140) | 2 | 3 | 3 | 2 | 3 | 7 | 3 | 3 | 2 | 3 | 4 |
| Zyzomys argurus (TA141) | 4 | 3 | 2 | 2 | 3 | 7 | 3 | 3 | 3 | 3 | 3 |
| Zyzomys argurus (TA150) | 4 | 4 | 1 | 2 | 3 | 7 | 2 | 3 | 2 | 2 | 4 |
| Rattus rattus (TA201) | 2 | 3 | 2 | 2 | 3 | 7 | 3 | 3 | 3 | 3 | 4 |
| Rattus rattus (TA063) | 3 | 4 | 3 | 2 | 3 | 7 | 2 | 3 | 2 | 2 | 3 |
| Rattus lutreolus (TA053) | 3 | 2 | 4 | 2 | 3 | 7 | 4 | 4 | 2 | 4 | 4 |
| Rattus lutreolus (TA107) | 3 | 4 | 4 | 2 | 3 | 6 | 4 | 3 | 3 | 3 | 4 |
| Rattus fuscipes (TA216) | 1 | 2 | 2 | 1 | 3 | 6 | 2 | 2 | 3 | 3 | 2 |
| Rattus fuscipes (TA001) | 4 | 4 | 3 | 2 | 3 | 7 | 4 | 3 | 3 | 4 | 4 |
| Pseudomys apodemoides (TA093) | 1 | 4 | 2 | 1 | 3 | 7 | 2 | 2 | 3 | 4 | 3 |
| Pseudomys apodemoides (TA096) | 4 | 3 | 1 | 2 | 3 | 7 | 4 | 3 | 3 | 3 | 4 |
| Mus musculus (TA072) | 2 | 3 | 3 | 2 | 3 | 7 | 3 | 3 | 2 | 3 | 4 |
| Mus musculus (TA074) | 2 | 4 | 2 | 2 | 3 | 7 | 3 | 3 | 3 | 3 | 4 |
| Mus musculus (TA105) | 3 | 4 | 2 | 3 | 3 | 7 | 4 | 3 | 2 | 2 | 3 |
| Mus musculus (3490) | 3 | 4 | 3 | 1 | 3 | 8 | 3 | 3 | 3 | 4 | 3 |
| Mus musculus (3491) | 2 | 4 | 2 | 1 | 3 | 8 | 3 | 2 | 3 | 4 | 3 |
| Mus musculus (3492) | 3 | 4 | 2 | 2 | 3 | 8 | 2 | 2 | 2 | 4 | 3 |
| Notomys mitchelli (TA098) | 2 | 4 | 3 | 2 | 3 | 7 | 3 | 3 | 3 | 3 | 4 |
| Notomys mitchelli (TA100) | 2 | 3 | 2 | 2 | 3 | 7 | 4 | 3 | 3 | 4 | 4 |
| Notomys mitchelli (TA102) | 2 | 3 | 2 | 1 | 4 | 7 | 2 | 2 | 3 | 3 | 2 |
| Hydrochaeris hydrochaeris (2079) | 3 | 2 | 3 | 3 | 2 | 6 | 4 | 3 | 3 | 3 | 0 |
| Hydrochaeris hydrochaeris (2080) | 3 | 3 | 3 | 3 | 2 | 6 | 4 | 3 | 3 | 3 | 0 |
| Liomys pictus (76) | 3 | 2 | 3 | 3 | 2 | 6 | 1 | 3 | 5 | 3 | 4 |
| Liomys pictus (78) | 3 | 2 | 3 | 4 | 2 | 6 | 1 | 3 | 5 | 4 | 4 |
| Liomys pictus (19) | 5 | 3 | 3 | 2 | 3 | 5 | 4 | 6 | 5 | 3 | 2 |
| Liomys pictus (20) | 5 | 3 | 3 | 4 | 3 | 3 | 4 | 6 | 5 | 3 | 2 |
| Liomys pictus (21) | 5 | 3 | 3 | 4 | 3 | 5 | 4 | 6 | 5 | 3 | 2 |
| Dipodomys merriami (286) | 3 | 2 | 3 | 3 | 3 | 6 | 3 | 5 | 5 | 2 | 3 |
| Dipodomys merriami (287) | 3 | 2 | 3 | 3 | 3 | 3 | 2 | 5 | 5 | 2 | 4 |
| Dipodomys merriami (288) | 3 | 2 | 3 | 3 | 3 | 3 | 3 | 5 | 5 | 2 | 4 |
| Perognathus penicillatus (820) | 2 | 2 | 3 | 2 | 3 | 3 | 7 | 5 | 4 | 3 | 4 |
| Perognathus penicillatus (821) | 2 | 2 | 3 | 2 | 3 | 3 | 7 | 4 | 4 | 3 | 0 |
| Perognathus penicillatus (822) | 2 | 2 | 3 | 2 | 3 | 3 | 7 | 5 | 4 | 3 | 0 |
| Tursiops truncatus (2025) | 2 | 3 | 3 | 3 | 3 | 6 | 4 | 5 | 3 | 2 | 4 |
| Tursiops truncatus (2026) | 2 | 3 | 3 | 4 | 3 | 6 | 4 | 5 | 3 | 2 | 4 |
| Tursiops truncatus (2027) | 2 | 3 | 2 | 4 | 3 | 6 | 4 | 5 | 3 | 2 | 4 |
| Felis concolor (268) | 3 | 3 | 2 | 3 | 2 | 6 | 4 | 3 | 1 | 3 | 2 |
| Felis concolor (269) | 3 | 3 | 3 | 2 | 1 | 6 | 5 | 3 | 3 | 3 | 0 |
| Felis catus (8) | 4 | 3 | 3 | 2 | 3 | 6 | 4 | 5 | 3 | 3 | 3 |
| Felis catus (9) | 4 | 3 | 3 | 2 | 3 | 3 | 3 | 5 | 3 | 3 | 4 |
| Felis catus (10) | 4 | 3 | 3 | 2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 |
| Panthera onca (65) | 3 | 3 | 4 | 2 | 2 | 7 | 5 | 3 | 6 | 4 | 3 |
| Panthera onca (66) | 3 | 3 | 4 | 2 | 2 | 7 | 5 | 3 | 6 | 4 | 3 |
| Panthera onca (270) | 3 | 3 | 4 | 2 | 3 | 3 | 3 | 4 | 6 | 3 | 3 |
| Lutra longicaudis (1930) | 2 | 2 | 3 | 0 | 3 | 6 | 3 | 3 | 2 | 3 | 3 |
| Lutra longicaudis (1931) | 2 | 3 | 3 | 3 | 3 | 6 | 3 | 4 | 3 | 3 | 4 |
| Lutra longicaudis (1932) | 2 | 3 | 2 | 2 | 3 | 6 | 2 | 3 | 2 | 3 | 4 |
| Bassariscus astutus (84) | 4 | 4 | 2 | 3 | 4 | 4 | 0 | 4 | 5 | 3 | 4 |
| Bassariscus astutus (815) | 2 | 4 | 3 | 2 | 3 | 5 | 4 | 3 | 4 | 4 | 2 |
| Bassariscus astutus (816) | 2 | 4 | 3 | 2 | 3 | 5 | 4 | 3 | 2 | 3 | 2 |
| Bassariscus astutus (817) | 2 | 4 | 2 | 2 | 3 | 3 | 4 | 2 | 2 | 3 | 2 |
| Nasua narica (2060) | 4 | 0 | 3 | 2 | 3 | 7 | 3 | 3 | 2 | 3 | 3 |
| Nasua narica (2062) | 2 | 3 | 3 | 2 | 3 | 7 | 2 | 2 | 3 | 3 | 3 |
| Procyon lotor (2064) | 5 | 2 | 5 | 3 | 3 | 7 | 3 | 4 | 2 | 4 | 0 |
| Procyon lotor (2065) | 5 | 2 | 5 | 3 | 3 | 7 | 3 | 4 | 2 | 4 | 0 |
| Procyon lotor (2066) | 5 | 2 | 3 | 2 | 3 | 5 | 2 | 2 | 2 | 0 | 0 |
| Canis latrans (90) | 3 | 4 | 4 | 3 | 4 | 4 | 4 | 3 | 2 | 4 | 1 |
| Canis latrans (91) | 3 | 4 | 4 | 3 | 3 | 5 | 4 | 4 | 3 | 4 | 1 |
| Canis latrans (92) | 3 | 4 | 4 | 3 | 3 | 5 | 4 | 4 | 3 | 4 | 1 |
| Canis latrans (824) | 3 | 3 | 3 | 2 | 3 | 6 | 4 | 3 | 1 | 4 | 4 |
| Canis latrans (825) | 3 | 3 | 3 | 3 | 3 | 6 | 4 | 3 | 3 | 3 | 4 |
| Canis latrans (830) | 0 | 3 | 3 | 4 | 3 | 3 | 4 | 5 | 3 | 3 | 3 |
| Canis familiaris (1) | 2 | 3 | 3 | 4 | 3 | 3 | 6 | 5 | 3 | 3 | 4 |
| Canis familiaris (2) | 2 | 3 | 4 | 4 | 3 | 5 | 6 | 4 | 2 | 3 | 4 |
| Canis familiaris (3) | 2 | 3 | 4 | 4 | 3 | 3 | 6 | 5 | 2 | 3 | 4 |
| Urocyon cinereoargenteus (272) | 4 | 4 | 4 | 3 | 1 | 1 | 0 | 3 | 3 | 3 | 1 |
| Urocyon cinereoargenteus (299) | 3 | 2 | 4 | 2 | 1 | 1 | 5 | 5 | 3 | 3 | 3 |
| Urocyon cinereoargenteus (300) | 4 | 4 | 4 | 2 | 3 | 5 | 5 | 5 | 3 | 3 | 4 |
| Urocyon cinereoargenteus (301) | 4 | 4 | 4 | 3 | 3 | 5 | 5 | 5 | 4 | 3 | 2 |
| ECOR23 | 3 | 4 | 4 | 2 | 3 | 7 | 4 | 3 | 3 | 3 | 4 |
| Trichechus manatus (1735) | 4 | 5 | 6 | 5 | 3 | 6 | 7 | 0 | 2 | 3 | 3 |
| Trichechus manatus (1736) | 4 | 5 | 6 | 3 | 3 | 6 | 7 | 0 | 2 | 3 | 3 |
| Tapirus bairdii (1666) | 4 | 3 | 4 | 2 | 3 | 6 | 5 | 0 | 5 | 3 | 4 |
| Tapirus bairdii (1667) | 0 | 3 | 4 | 2 | 3 | 8 | 5 | 5 | 5 | 3 | 4 |
| Tapirus bairdii (1668) | 0 | 3 | 4 | 2 | 3 | 8 | 4 | 4 | 5 | 4 | 3 |
| Tapirus bairdii (1937) | 2 | 1 | 3 | 0 | 3 | 6 | 3 | 4 | 2 | 3 | 4 |
| Tapirus terrestris (2381) | 4 | 3 | 4 | 3 | 2 | 7 | 4 | 4 | 3 | 4 | 4 |
| Tapirus terrestris (2382) | 4 | 3 | 4 | 3 | 3 | 5 | 5 | 3 | 3 | 4 | 3 |
| Equus caballus (807) | 3 | 2 | 4 | 2 | 3 | 5 | 4 | 0 | 5 | 3 | 4 |
| Equus caballus (808) | 4 | 2 | 3 | 2 | 3 | 5 | 3 | 0 | 6 | 3 | 4 |
| Equus caballus (809) | 3 | 2 | 3 | 2 | 3 | 6 | 4 | 0 | 5 | 3 | 4 |
| ECOR45 | 2 | 3 | 3 | 2 | 3 | 7 | 2 | 2 | 3 | 3 | 3 |
| Tayassu tajacu (2055) | 1 | 3 | 3 | 2 | 3 | 7 | 3 | 3 | 3 | 3 | 3 |
| Odocoeileus virginianus (63) | 2 | 2 | 3 | 2 | 2 | 7 | 3 | 3 | 2 | 3 | 1 |
| Odocoeileus virginianus (1984) | 5 | 4 | 4 | 2 | 3 | 6 | 3 | 3 | 3 | 3 | 3 |
| Odocoeileus virginianus (1985) | 3 | 3 | 4 | 2 | 2 | 6 | 4 | 0 | 3 | 3 | 4 |
| ECOR27 | 2 | 3 | 3 | 2 | 3 | 7 | 3 | 2 | 3 | 3 | 3 |
| Bos taurus (2376) | 5 | 3 | 4 | 2 | 3 | 7 | 4 | 3 | 2 | 3 | 4 |
| Bos taurus (2379) | 6 | 2 | 3 | 3 | 2 | 7 | 3 | 4 | 3 | 3 | 0 |
| ECOR33 | 2 | 3 | 3 | 2 | 3 | 6 | 3 | 2 | 3 | 3 | 3 |
| Ovis aries (1965) | 4 | 3 | 3 | 2 | 4 | 6 | 3 | 3 | 3 | 3 | 3 |
| Ovis aries (1966) | 4 | 3 | 3 | 2 | 4 | 6 | 3 | 4 | 3 | 3 | 3 |
| Ovis aries (1967) | 3 | 3 | 5 | 3 | 2 | 5 | 5 | 0 | 3 | 3 | 4 |
| Sula nebouxi (137) | 4 | 7 | 5 | 4 | 3 | 0 | 2 | 3 | 2 | 4 | 3 |
| Aquila chrysaetus (55) | 5 | 6 | 6 | 2 | 4 | 0 | 1 | 2 | 2 | 3 | 3 |
| Aratinga canicularis (48) | 5 | 6 | 4 | 3 | 3 | 0 | 1 | 3 | 1 | 3 | 4 |
| Calothorax lucifer (207) | 6 | 2 | 4 | 2 | 2 | 0 | 3 | 3 | 2 | 2 | 4 |
| Amazilia beryllina (225) | 3 | 4 | 4 | 2 | 1 | 0 | 1 | 3 | 1 | 1 | 4 |
| Henicorhina leucophrys (151) | 4 | 3 | 5 | 3 | 3 | 0 | 2 | 5 | 2 | 3 | 3 |
| Basileuterus belli (108) | 4 | 3 | 4 | 2 | 3 | 0 | 1 | 2 | 2 | 1 | 3 |
| Atlapetes brunneinucha (126) | 3 | 2 | 5 | 3 | 3 | 0 | 2 | 3 | 2 | 3 | 4 |
| Passer domesticus (345) | 6 | 7 | 4 | 4 | 2 | 0 | 3 | 3 | 2 | 4 | 4 |
| Empidonax sp. (161) | 4 | 3 | 2 | 3 | 4 | 0 | 1 | 4 | 2 | 1 | 2 |
For additional information regarding the ECOR isolates, see reference 32.
Biotype analysis.
All strains were taken from freezer cultures and grown on MacConkey plates. A single colony of each strain was tested for growth on minimal plates containing a 0.4% concentration (27) of one of the following sugars: adonitol, arabinose, dulcitol, inositol, maltose, mannitol, raffinose, rhamnose, salicin, sucrose, sorbitol, trehalose, and xylose.
Antibiotic resistance and toxin production.
Resistance to each of six antibiotics was tested by using Luria broth plates supplemented with one of the following antibiotics: ampicillin (50 μg/ml), chloramphenicol (12 μg/ml), kanamycin (50 μg/ml), neomycin (50 μg/ml), streptomycin (50 μg/ml), and tetracycline (25 μg/ml). The concentrations were based on previous studies of E. coli (27, 35).
Strains were also tested for hemolysin production with heart infusion agar supplemented with blood (5%) and were tested for verotoxin (VT) production with Rainbow Agar O157 (Biolog, Inc.). However, Rainbow Agar O157 is not a direct test for VT production; rather, it detects a trait that has been found to be highly correlated with VT production in clinically pathogenic strains such as O157:H7 (44). This pathogenic strain is β-glucuronidase negative (black colonies); other VT-producing strains typically overproduce β-galactosidase relative to β-glucuronidase (blue, purple, or violet colonies), and it is the production of these two compounds that Rainbow Agar has the ability to detect. We took dark and bluish colonies to represent VT-positive strains. To determine the average number of colonies per host group, we scored for dark colonies as follows: 2 for black colonies; 1 for dark blue, violet, and purple; and 0 for red and white colonies.
Plasmid analysis.
Isolated colonies were grown in Tris-borate medium and plasmids were extracted by the alkaline lysis procedure (12). Plasmid profiles were run in 0.7% agarose gel and stained with ethidium bromide. Megaplasmids were extracted directly in horizontal agarose gels by a modification of the procedure of Eckhardt (12). For purposes of statistical analysis, the observed plasmid bands were assigned to categories according to their size.
Statistical analysis.
The isolates were grouped on the basis of the taxonomic order of the host from which they were isolated, host diet, and host geographic origin. Some of these groups are naturally confounded (most carnivora eat meat and most rodents are granivores), and it is not possible to separate their contribution to phenotypic or genetic diversity. For example, Australia lacks native primates and Mexico lacks monotremes. To partially compensate for these confounding effects, various subsets of the data were used in the analyses.
Average genetic diversity per locus was estimated as H = Σhj/m, where m equals the number of loci scored and hj = [n/(n − 1)] (1 − Σpij2), where pij is the frequency of allele i at locus j, and n is the number of multilocus genotypes (30, 40). Standard error of H was obtained with the ETDIV program (46). We used modified Gst statistics to analyze the data. For example, the proportion of genetic variation attributable to geographic effects is (HT − HG)/HT, where HG is the arithmetic average of the H’s calculated separately for electrotypes (ETs) from each locality, and HT is the diversity of all strains regardless of locality (29, 30, 40). Standard error of Gst was also obtained with the ETDIV program (46). The statistical significance of Gst was analyzed with a χ2 test of independence, using the formula χ2 = nGst(a − 1), where n is the number of individuals and a is the total number of alleles; degrees of freedom are (k − 1)(a − 1), where k is the number of subdivisions (16).
The frequencies of different biotype traits were compared using χ2 tests or Fisher’s exact test when appropriate (37). To test for the association of phenotypic traits, a concordance analysis was done by using contingency coefficients (37). As contingency coefficients do not range from −1 to 1, as is the case for parametric measures of association, the absolute values of the coefficients are not presented.
A dendrogram was constructed using Nei’s genetic distances (30) and the neighbor-joining (NJ) method, using the Phylogeny Inference Package (PHYLIP, version 3.5c by Joseph Felsenstein, University of Washington).
RESULTS
Allozyme analysis.
The 202 strains examined yielded 187 genotypes (i.e., ETs). The number of alleles per locus averaged 6.8 (Table 2; range, 4 to 9). Null alleles were detected at all loci except MDH. The locus with the greatest allelic diversity was ME2 (0.80) and G6PD was the least variable (0.441). The average allelic diversity (H ± standard error [SE]) for the 11 loci was 0.682 ± 0.034 for the 187 ETs, and for the 202 isolates, we obtained an H of 0.673 ± 0.034 (Table 2).
TABLE 2.
Number of alleles and allelic diversity for 11 loci in 187 E. coli electrotypes
| Locus | No. of alleles | H |
|---|---|---|
| ADH1 | 9 | 0.736 |
| ARK | 6 | 0.617 |
| G6PD | 5 | 0.441 |
| IDH | 7 | 0.778 |
| MDH | 6 | 0.730 |
| ME1 | 4 | 0.531 |
| ME2 | 9 | 0.800 |
| MPI | 8 | 0.786 |
| PEP | 8 | 0.697 |
| PGM | 7 | 0.727 |
| XDH1 | 6 | 0.654 |
| Avg ± SE | 6.8 | 0.682 ± 0.034 |
In Table 3, we present genetic diversity and genetic differentiation results at different levels of analysis. At the geographic level, the diversity (H) ranges from 0.489 in the human-related strains of the ECOR collection to 0.705 in the strains isolated from Mexican mammals. The genetic differentiation ± SE among those data sets (Gst = 0.047 ± 0.014; χ2 = 630.9) is significantly different from zero (P < 0.0001).
TABLE 3.
Genetic diversity and genetic differentiation in E. coli from different geographical origins and with different host taxonomies and diets
| Level of analysis | Origin of the strain or description of host | No. of strains | No. of alleles | H | Gst (SE) | P |
|---|---|---|---|---|---|---|
| Geographic | Australia | 41 | 4.18 | 0.566 | ||
| Mexicoa | 131 | 6.82 | 0.705 | |||
| ECOR | 13 | 2.73 | 0.489 | 0.047 (0.014) | 0.00001 | |
| Host order | Carnivora | 34 | 4.55 | 0.653 | ||
| Rodentia | 51 | 4.82 | 0.657 | |||
| Marsupialia | 28 | 4.27 | 0.603 | |||
| Primates | 22 | 4.36 | 0.658 | |||
| Chiroptera | 14 | 4.18 | 0.665 | |||
| Artiodactyla | 11 | 3.09 | 0.511 | |||
| Perisodactyla | 10 | 3.18 | 0.608 | |||
| Aves | 10 | 3.36 | 0.630 | 0.075 (0.017) | 0.00001 | |
| Host diet | Omnivore | 66 | 5.45 | 0.646 | ||
| Granivore | 28 | 4.91 | 0.645 | |||
| Carnivore | 12 | 4.27 | 0.671 | |||
| Herbivore | 50 | 5.82 | 0.645 | |||
| Insectivore | 23 | 4.36 | 0.672 | 0.025 (0.007) | 0.126 | |
| Mexicob | 110 | 6.74 | 0.698 | 0.044 (0.012) | 0.0052 | |
| Australiab | 41 | 4.18 | 0.566 | 0.01 (0.01) | 1 | |
| Rodents | Australia | 17 | 2.91 | 0.515 | ||
| Mexico | 34 | 4.73 | 0.639 | 0.098 (0.03) | 0.00001 | |
| Australia | Rodents | 17 | 2.91 | 0.515 | ||
| Marsupials | 21 | 3.55 | 0.552 | 0.036 (0.011) | 0.184 |
The Mexican sample includes samples from 10 birds.
For space reasons, we included in these rows the average values for the different diets within each country.
If we take into account the host order, the genetic diversity ranges from 0.511 in artiodactyls to 0.665 in bats; the genetic differentiation among the strains from different hosts (Gst = 0.075 ± 0.017; χ2 = 898.56) is also significantly different from 0 (P < 0.0001; Table 3).
Considering host diet, the genetic diversity ranges from 0.672 in the strains associated with insectivores of different countries to 0.566 in the Australian strains with different diets. There is no genetic differentiation according to host diet (Gst = 0.025 ± 0.007; χ2 = 323.64, P = 0.126). If we analyze only the host diet in Australia (Gst = 0.01 ± 0.01; χ2 = 28.68, P = 0.999), we also do not find a significant difference. However, diet is significant in Mexico (Gst = 0.044 ± 0.012; χ2 = 296, P = 0.005).
The genetic differentiation between rodents from Australia and Mexico (Gst = 0.098 ± 0.03; χ2 = 234.85, P < 0.0001) was significantly different from zero (Table 3), with more diversity in the Mexican isolates (H = 0.639) than the isolates from Australian rodents (H = 0.515). However, partitioning the strains of Australia showed a nonsignificant differentiation (Gst = 0.036 = 0.011; χ2 = 48.95, P = 0.184), because the genetic diversity is very similar in marsupials (H = 0.552) and rodents (H = 0.515).
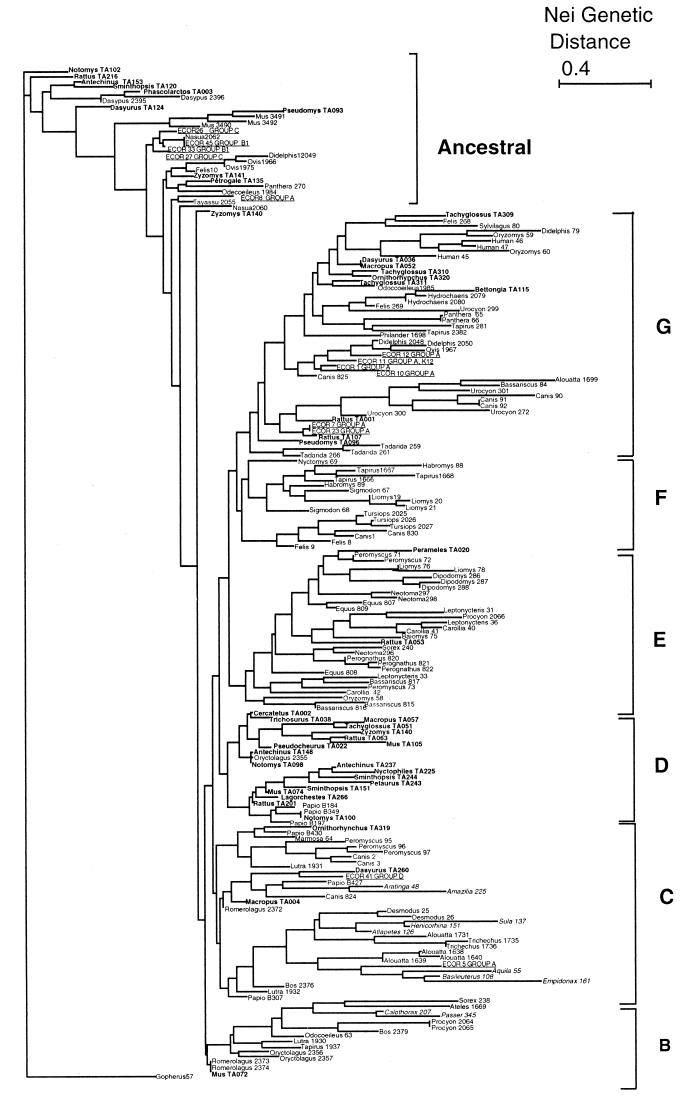
The genetic relationship among 202 strains was analyzed with Nei genetic distances, and we constructed an NJ dendrogram (Fig. 1). We also performed other phylogenetic analyses using UPGMA (using both the number of mismatches and Nei distances as genetic distances) distance method and constructed a PAUP tree using parsimony (figures not shown). All the analyzed trees showed results analogous to those of the NJ dendrogram. Based on biochemical tests and the UPGMA dendrograms, we decided to root the NJ dendrogram with the reptile strain, which shows a long branch. If this root is correct, in the dendrogram we can define an ancestral group from which all the other strains are derived. In this supposedly ancestral group, we found several strains (10) from marsupials and rodents from Australia as well as 11 strains from a diverse group of mammals from Mexico and three strains from the cosmopolitan house mouse Mus musculus. In this clade, we also observed the strains from groups B1 and C of the ECOR collection, as well as one strain from the group A (ECOR8). The majority of the human-associated ECOR group A isolates are present in the most differentiated cluster (G), along with strain K-12 (MG1655, identical in our analysis to ECOR11) and a group of strains from Mexican carnivores, several from rodents, and all the nonpathogenic strains from humans. The rest of the ECOR isolates (no. 5 and 41) are in cluster C along with most of the bird strains and isolates from the old world and new world monkeys. In cluster B, we observed a diverse group of strains mostly from Lagomorpha and carnivores. Cluster D has a tight group of Australian strains from marsupials and rodents. Strains from Mexican rodents and bats are mostly in clusters E and F. Cluster F also contains strains from domestic carnivores and wild dolphins. It is evident in this tree that most of the strains from wild animals are grouped by host order. It is also interesting that most of the Australian strains are in two clusters (A and G) while Mexican strains are dispersed along the tree.
FIG. 1.
Dendrogram depicting the strain similarities for the 202 strains of E. coli. The NJ tree was obtained from a Nei’s distance matrix derived from the multilocus electrophoresis data using the PHYLIP 3.5 program. For each strain, the name of the host along with the strain reference number is printed. Australian strains are in bold, bird strains are in italics, and ECOR strains are underlined.
Biotype analyses.
A minority of the strains could utilize adonitol, arabinose, dulcitol, inositol, or salicin, while most strains could ferment maltose, mannose, rhamnose, sorbose, trehalose, and xylose (Table 4). For 10 of the 12 substrates tested, the proportion of strains able to exploit a substrate varied significantly depending on the taxonomic group of the host from which they were isolated (Table 4). The ability of the strains to ferment maltose, rhamnose, and xylose was independent of host taxonomic group. Overall, strains isolated from hosts with diversified diets such as rodents, birds, and marsupials could ferment the greatest number of substrates, while strains from hosts with very specialized diets such as Monotremata, Cetacea, Xenarthra, and Sirenia could exploit fewer sugars.
TABLE 4.
Variation in sugar utilization among E. coli strains as a function of host taxonomic group
| Host groupa | No. of isolates | % Utilization of:
|
Total no. of sugars utilized | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ado | Ara | Dul | Ino | Mal | Man | Raf | Ram | Sal | Sor | Suc | Tre | Xyl | |||
| Aves | 10 | 20 | 10 | 50 | 40 | 80 | 80 | 40 | 90 | 50 | 100 | 80 | 100 | 90 | 13 |
| Artiodactyla | 12 | 0 | 0 | 8 | 0 | 100 | 25 | 50 | 75 | 0 | 58 | 33 | 91 | 75 | 9 |
| Carnivora | 34 | 6 | 0 | 39 | 0 | 100 | 76 | 67 | 91 | 0 | 91 | 70 | 100 | 100 | 10 |
| Cetaceab | 3 | 33 | 0 | 0 | 0 | 100 | 0 | 0 | 100 | 0 | 100 | 100 | 100 | 100 | 7 |
| Chiroptera | 15 | 29 | 0 | 79 | 29 | 93 | 100 | 79 | 93 | 50 | 86 | 100 | 100 | 93 | 12 |
| Insectivorab | 2 | 50 | 0 | 0 | 0 | 100 | 100 | 50 | 50 | 50 | 50 | 50 | 100 | 50 | 10 |
| Lagomorphab | 7 | 14 | 43 | 0 | 0 | 100 | 100 | 43 | 57 | 0 | 100 | 43 | 100 | 100 | 10 |
| Marsupialia | 28 | 4 | 4 | 11 | 11 | 100 | 86 | 39 | 86 | 4 | 89 | 71 | 100 | 100 | 13 |
| Montrematab | 5 | 0 | 0 | 0 | 0 | 100 | 0 | 0 | 80 | 0 | 100 | 60 | 100 | 100 | 6 |
| Perissodactylab | 9 | 33 | 0 | 0 | 0 | 100 | 56 | 33 | 67 | 0 | 89 | 22 | 100 | 78 | 9 |
| Primates | 24 | 5 | 5 | 14 | 0 | 100 | 54 | 50 | 68 | 0 | 55 | 64 | 91 | 91 | 11 |
| Rodentia | 51 | 10 | 4 | 10 | 2 | 90 | 64 | 50 | 78 | 10 | 84 | 70 | 90 | 90 | 13 |
| Statistical significancec | ∗ | ∗∗∗ | ∗∗∗ | ∗∗∗ | NS | ∗∗∗ | ∗ | NS | ∗∗∗ | ∗∗∗ | ∗∗ | ∗ | NS | ∗∗∗ | |
| ECOR | 13 | 32 | 39 | 81 | 0 | 39 | 11 | ||||||||
Does not include the single isolate from an elephant and a desert turtle.
Analysis excluded those host groups were the strain number is lower than 10.
NS, not significant; ∗, P < 0.05; ∗∗, P < 0.01; ∗∗∗, P < 0.001. Significance levels determined by χ2 tests.
Geographic effects on the frequency of sugar utilization could also be detected. A comparison of strains isolated from rodents in Australia and Mexico revealed that for mannitol only 32% (n = 16) of the strains from Australia could ferment this sugar, compared to 85% (n = 30) of the strains from Mexico (P < 0.001). Similarly, 32% of strains from Australian rodents can ferment raffinose compared to 64% of the strains from Mexican rodents (P < 0.02).
Antibiotic resistance, toxin production, and plasmid profiles.
Of the antibiotics tested, resistance to streptomycin was most frequent while resistance to chloramphenicol was rare (Table 5). Resistance to antibiotics was more frequent among strains from Mexico than among strains from Australia (9.6% of strains from Australia were resistant to one or more antibiotics compared to 41% of strains from Mexico). There was an insufficient number of strains with resistance to permit a detailed analysis of the distribution of antibiotic resistance as a function of host taxonomic group. However, considering the Mexican isolates alone, strains isolated from bats tended to have a much higher frequency of antibiotic resistance than other mammals. For example, 46% of strains from bats were ampicillin resistant, 100% were streptomycin resistant, and 15% presented neomycin resistance; none of those strains was resistant to the other antibiotics.
TABLE 5.
Overall frequency of antibiotic resistance in E. coli and comparison of frequencies between strains isolated from Australia and México
| Antibiotic for which resistance was tested | % of strains resistant to indicated antibiotic
|
Level of statistical significancea | ||
|---|---|---|---|---|
| All strains | Mexican strains (n = 135) | Australian strains (n = 46) | ||
| Ampicillin | 13.4 | 18.5 | 5.8 | ∗ |
| Chloramphenicol | 0.8 | 0.6 | 0 | NS |
| Streptomycin | 23.9 | 32.6 | 5.8 | ∗∗∗ |
| Kanomycin | 2.1 | 3.3 | 0 | NS |
| Neomycin | 3.4 | 2.6 | 0 | NS |
| Tetracycline | 5.9 | 7.8 | 0 | ∗ |
For explanation of significance levels, see footnote c, Table 4.
Overall, 44% of the strains produced more β-galactosidase than β-glucuronidase (i.e., were VT positive), and 2% of strains produced hemolysin. No difference in the overall frequency of hemolysins or VT production between strains from Australia and Mexico could be detected (Table 6; the average for Mexico was 0.64 ± 0.06 and the average for Australia was 0.57 ± 0.10 [not significant by Student’s t test]). The frequency of the marker for VT production varied with the host taxonomic group from which the strains were isolated (Table 6). Among the Mexican isolates, the frequency of positive strains ranged from 0.28 for strains isolated from Lagomorpha to 0.69 in strains isolated from rodents. Isolates from Australian rodents also showed a higher than average frequency of VT production compared to strains from marsupials. However, none of these differences are significant from zero.
TABLE 6.
Average and frequency of dark colonies in rainbow agar in E. coli strains isolated from various wild hosts
| Origin of host | Host order | No. of isolates | Avg no. of VT-positive strains ± SE | Frequency of VT-positive strains |
|---|---|---|---|---|
| Mexico and South America | All orders | 121a | 0.64 ± 0.06 | 0.60 |
| Ungulata | 18 | 0.39 ± 0.14 | 0.33 | |
| Carnivora | 34 | 0.53 ± 0.1 | 0.47 | |
| Chiroptera | 13 | 0.69 ± 0.17 | 0.61 | |
| Lagomorpha | 7 | 0.29 ± 0.17 | 0.28 | |
| Primates | 10 | 0.30 ± 0.14 | 0.33 | |
| Rodentia | 34 | 1.09 ± 0.14 | 0.69 | |
| Australia | All orders | 46a | 0.57 ± 0.1 | 0.45 |
| Marsupialia | 21 | 0.48 ± 0.16 | 0.33 | |
| Rodentia | 17 | 0.82 ± 0.15 | 0.66 |
The continental total includes small orders of mammalian hosts.
Concordance analysis (40) of the phenotypic traits using the strains isolated from Mexico revealed that the presence or absence of these traits is not independent (Table 7). Three major forms of association were detected: (i) the concurrence of the ability to utilize the less frequently fermented sugars, such as dulcitol and inositol; (ii) the concurrence of the inability to exploit sugars utilized by most strains, such as xylose and trehalose; and (iii) the negative association of two traits, such as where the ability to exploit one sugar (e.g., salicin) results in a lower-than-expected frequency of strains able to exploit a second sugar (e.g., sucrose). Resistance to various antibiotics was also found to be concurrent in a strain more often than would be expected by chance. Resistance to ampicillin was associated with the ability to utilize arabitol, adonitol, dulcitol, and salicin. Also the ability to exploit salicin was found to be more frequently associated with streptomycin resistance than expected. Similar kinds of association were seen in the strains from Australia (results not shown).
TABLE 7.
Associations of various phenotypic traits in E. coli isolated in Mexico
| Phenotype | Association between indicated phenotypesa
|
||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ara | Ado | Dul | Ino | Mal | Man | Raf | Rha | Sal | Sor | Suc | Tre | Xyl | Amp | Str | Kan | Neo | |
| Ado | 0 | ||||||||||||||||
| Dul | 0 | 0 | |||||||||||||||
| Ino | 0 | p | p | ||||||||||||||
| Mal | 0 | 0 | 0 | p | |||||||||||||
| Man | 0 | 0 | 0 | 0 | 0 | ||||||||||||
| Raf | 0 | 0 | p | 0 | 0 | 0 | |||||||||||
| Rha | 0 | 0 | p | 0 | 0 | 0 | 0 | ||||||||||
| Sal | 0 | p | p | p | 0 | 0 | − | 0 | |||||||||
| Sor | 0 | 0 | 0 | 0 | a | 0 | 0 | a | 0 | ||||||||
| Suc | 0 | 0 | p | 0 | 0 | 0 | p | 0 | − | 0 | |||||||
| Tre | 0 | 0 | 0 | 0 | a | a | 0 | 0 | 0 | 0 | 0 | ||||||
| Xyl | 0 | 0 | 0 | 0 | a | 0 | 0 | a | 0 | a | a | a | |||||
| Amp | p | p | p | 0 | 0 | − | 0 | 0 | p | 0 | 0 | 0 | 0 | ||||
| Str | 0 | 0 | 0 | 0 | 0 | − | 0 | 0 | p | 0 | 0 | 0 | 0 | p | |||
| Kan | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | p | 0 | ||
| Neo | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | p | 0 | p | |
| Tet | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | p | p | p |
Only those associations found to be significant at the P < 0.01 level are presented. p, joint presence of a rare trait; a, joint absence of a common trait; 0, no significant association; −, negative association of two traits.
We divided the samples by the distance of the host to the human environment (Table 8). We observed in the Mexican isolates and in the ECOR isolates a significantly higher amount of plasmids among isolates collected in cities and places close to humans than in those collected at isolated sites (χ2 = 28.98, P < 0.001) (Table 8). The effect of possible human contact on the number of plasmids was also observed on a larger geographical scale. In Mexican strains, we observed an average of 1.7 plasmids per strain, while in the less-populated Australia, the strains presented a significantly lower average (0.45 plasmids per strain; χ2 = 322.8, P < 0.001).
TABLE 8.
Antibiotic resistance and plasmid numbers in E. coli isolated from different sources in relation to their proximity to human environment
| Country of origin of isolate | Proximity to human environmenta | Source or type of isolate | No. of isolates | Avg no. of plasmids (SE) | Avg no. of antibiotic-resistant isolates (SE) |
|---|---|---|---|---|---|
| Australia | More than 100 miles from city | Wild mammals | 46 | 0.45 (0.09) | 0.19 (0.01) |
| Mexico | More than 100 miles from city | Wild mammals | 86 | 1.20 (0.12) | 0.58 (0.06) |
| Mexico | Less than 100 miles from city | Wild mammals | 14 | 1.55 (0.42) | 0.42 (0.09) |
| Mexico | Zoo | Wild mammals | 9 | 1.86 (0.45) | 0.33 (0.17) |
| Mexico | Cities | Domestic mammals | 23 | 2.09 (0.42) | 0.47 (0.08) |
| Mexico and United States | Cities | ECOR and nonpathogenic | 14 | 2.62 (0.65) | 0.66 (0.09) |
Ten strains were excluded from the analysis because field data were not complete.
On the other hand, E. coli associated with wild animals in isolated places in Mexico is, on average, resistant to as many antibiotics (average ± SE = 0.58 ± 0.06; Table 8) as E. coli in the other human-related sources from Mexico (weighted average ± SE of all the other sources = 0.43 ± 0.01). Our sample of the ECOR collection had the highest antibiotic resistance (average ± SE = 0.68 ± 0.02; data not shown in table), while Australian isolates had the lowest antibiotic resistance (average ± SE = 0.19 ± 0.01). In our sample, we did not observe that the antibiotic resistance is correlated with the mean number of plasmids. Most of the multidrug-resistant strains from wild mammals did not have any plasmids, suggesting that the genes for antibiotic resistance are chromosomal in some of the wild animal strains.
DISCUSSION
The estimated genetic diversity (H = 0.682) from our collection of E. coli is higher than the diversity reported for any collection of E. coli (5, 26, 33, 34, 38, 42, 46–48) and higher than most studied bacteria (7, 9, 10, 14, 28, 29, 39, 43, 45). This high diversity could be due in part to the fact that we intentionally selected a wide range of “good and informative characters” based on the analysis done by Selander and collaborators (40, 42). On the other hand, the observed genetic diversity would change if other loci were selected or if MLEE were performed with starch and a discontinuous buffer with a different pH (17). For example, based on 11 loci, the H was originally determined to be 0.42 for the ECOR collection (32). Subsequently, the number of loci examined was increased to 35, resulting in an H of 0.34 (41). However, the high genetic diversity we observed in our study could also be the result of the high diversity of hosts represented in our collection; the sample we studied includes isolates from all of the major mammalian orders (98.3% of the species are within those orders [31]). Even though isolates from some minor orders are lacking, when we add 10 isolates from 10 different birds the genetic diversity increased only 0.2%. The geographic scope of the sample is also patchy, with only a few strains from Africa or Asia and none from either polar region. The results suggest, however, that filling these gaps may not significantly increase estimates of the genetic diversity of E. coli. For example, in the Mexican mammalian samples alone, the genetic diversity (H) was estimated as 0.698 ± 0.048 (data not shown). But inclusion of the Australian samples, which represent an isolated continent with a unique mammalian fauna, lowered the estimate of the genetic diversity by 1%.
Previous studies of E. coli suggest that much of the observed allelic variation is selectively neutral and this, coupled with a large effective population size, can explain much of this species’ genetic diversity (42, 50). Population structure has been thought to account for little of the observed diversity (5, 11, 20, 42). However, genetic diversity in our collection of E. coli is probably ecologically (ecotypically) structured and as such, adaptation to the host plays an important role in population structure. We observed that host order and geographic origin are the most important factors to differentiate E. coli. Overall, 7.5% of the diversity can be explained by the order of the host, while nearly 10% of the genetic diversity of strains from rodents (Australia versus America) can be explained by geography alone. This contrasts with the observation that spatial structure accounts for only 2% of the genetic diversity among strains isolated from humans living in North America and Europe (27). That spatial structure accounts for so little of the genetic diversity of E. coli from humans probably reflects the fact that the samples are derived from a single, highly mobile host species. The larger geographic component of diversity observed in the strains from rodents can be attributed both to Australia’s long isolation and to the distinct rodent communities of the two continents. In Mexico, the diet of the host accounts for 4.4% of the diversity of E. coli. However, in Australia or in the total collection, the diet of the host was not an important factor differentiating populations.
The observed differentiation by host could be due in part to the presence of different ecological niches in the different orders. We observed that the type and range of sugars that can be exploited by E. coli strains are associated with the taxonomic group of the host from which the strains were isolated. Significant heterogeneity in sugar utilization also resulted when strains were grouped on the basis of host diet. Nevertheless, these results are somewhat ambiguous. This may reflect the joint problem of sample size for the diet grouping (for example, four dietary modes among the 14 bat hosts) and the inevitable interactions between host taxonomy and diet. Overall, E. coli from the less-represented hosts with specialized diets (monotremes, dolphins, Xenarthra, and Sirenia) exploited the fewest sugars. The strains from hoofed mammals also used few sugars, while bacteria from birds, marsupials, and rodents could utilize a larger number of sugars. The frequency with which strains can exploit some sugars was also found to vary with the geographic origin of the strains. For example, a greater proportion of strains from Mexican rodents can utilize raffinose and mannitol than strains from Australian rodents.
In previous studies, the majority of strains (69%) in the ECOR collection that were isolated from strictly herbivorous mammals could exploit raffinose, whereas in this study, less than half the strains recovered from the herbivorous groups could utilize raffinose (27, 32, 42). Only 30% of ECOR strains can ferment sorbose, compared to the 82% frequency of utilization observed in this study (27, 32, 42). No ECOR strains could use salicin, while in this study, 9% of the strains did so, in contrast with 41% in a population of strains from yellow baboons (35). Host digestive physiology and anatomy clearly vary among mammal species and reflect the interaction of phylogeny and diet. Host diet will in turn influence the kinds of growth substrates available to E. coli. As a result, E. coli strains will be confronted with very different biotic and abiotic environments in different host species. Our study suggests that these different environments result in E. coli that exhibit some degree of host specificity. Further work is required to determine the mechanisms responsible for this specificity and the role that such specificity plays in E. coli’s evolution.
The dendrogram depicted in Fig. 1 provides additional support for the observation that host and geographic effects contribute to the genetic structure of E. coli populations. For example, strains isolated from birds, rodents, and carnivores seem to cluster more often than other hosts, as do Australian strains. Strains from ungulates are in many clusters. In our tree, the ancestral cluster corresponds to some of the Australian strains along with the ECOR strains from groups B1 and C, while the most differentiated cluster presents strains from the ECOR group A along with carnivores from Mexico and other humans. Using the sequence data of 13 gene phylogenies, Lecointre et al. (23) observed that the ECOR strains had a different organization than previously reported. In their study the most pathogenic strains from group B2 were the ancestral group, while A and B1 were the most evolved sister groups. In their scenario, the capability of E. coli to invade other niches (extraintestinal virulence) is an ancestral trait that has evolved to the more benign intestinal strains in the other groups (23). We do not have strains from group B2 in our study, but preliminary data from our laboratory (data not shown) suggest a similar scenario for the evolution of diarrheic E. coli. We observed that four of the Mexican strains from our ancestral group present genes from the pathogenic island LEE (eae and espB) associated with an intestinal lesion in the EPEC and EHEC strains. We also observed in this ancestral clade six strains with ETEC serotypes. Preliminary analysis (data not shown) indicates that pathogenic chromosomal genes are common and diverse along the tree, suggesting an ancestral origin.
We also observed both that the VT marker is widespread in the tree and that VTs are more frequent in strains from rodents from both continents. The latter observation is consistent with the idea that rodents act as a reservoir of many diseases (1). On the other hand, artiodactyls have strains with the lower percentage of β-galactosidase production, contrary to the belief that domestic cattle naturally harbor β-galactosidase-producing E. coli (i.e., O157:H7) (36, 44). However, none of the levels of VT production per host are significantly different from the average for their country of origin. Another characteristic that is widespread in the collection is antibiotic resistance. With the exception of strains from Australian mammals, most of the E. coli strains present some antibiotic resistance, and multidrug resistance is frequent in Mexico. This could be in part the result of the widespread use of antibiotics and to the lack of real isolation from human environment in a country like Mexico. However, since antibiotic resistance is not randomly distributed in the wild E. coli strains (bats and rodents host strains with higher multidrug resistance), we believe that at least some of the antibiotic resistance may be related to additional nonhuman environmental pressures.
On the other hand, the number and size of plasmids are highly variable in E. coli, ranging from hundreds of base pairs to several hundred kilobase pairs. Their G+C contents can vary widely and are often different from that of their usual bacterial host, indicating a variety of sources from which plasmids (or part of them) are derived (3, 18, 21, 22, 24). No pattern has been observed in the number of plasmids in these bacteria, and this may be due in part to their possible heterogeneous origin (3, 21, 22). Nevertheless, we observed that with human proximity, the number of plasmids per strain increases. This could be a reflection of the acquisition of accessory elements due to increased densities of both hosts and bacteria in cities. Increased density could facilitate the movement of plasmids among strains of E. coli as well as other related bacteria. Such an exchange was observed by Boyd et al. (3), who analyzed the structure of F group-related plasmids in the ECOR collection. They found different phylogenetic relationships between plasmids and bacterial strains, suggesting that horizontal transfer of plasmids occurs at high rates within the ECOR collection.
ACKNOWLEDGMENTS
We thank the numerous colleagues who assisted us in obtaining samples. David Gordon collected all the Australian samples, helped with some statistical analysis, and reviewed many previous versions of the manuscript. Blanca Hernández, Jordan Goluvov, Meli Mandrujano, Rodrigo Medellin, Saul Aguilar, Osiris Gaona, Angeles Mendoza, Gabriel Pérez, Luis Medrano, Adolfo Navarro, Juan Castillo, Alejandro Zavala, Benjamin Morales, Ricardo Frias, Alejandro Velázques, Hector Arita, Carlos Alvarez, José Charles, Hilary Charles, Alvaro Miranda, David Valenzuela, Luisa Sandner, René Cerritos, and Jorge Ortega collected the Mexican samples. We also thank Antonio Cruz for valuable technical assistance and Armando Navarro, who performed the serotype analysis in the Faculty of Medicine, Universidad Nacional Autónoma de México. We thank Pilar Beltrán, Robert K. Selander, Brandon Gaut, Andrew Peek, and Juan Nuñez, who read the manuscript and gave useful comments. Richard Goldstein kindly provided the MG1655 strain of E. coli K-12, and Dan Dykhuizen sent us the yellow baboon strains.
This research was supported by DGAPA-UNAM IN208995 grant to V.S. and CONACyT 3675-N grant to L.E.E. and V.S., and M.R. was supported by a graduate student scholarship from CONACyT.
Appendix
TABLE A1.
Host characteristics and geographical origins of E. coli from wild mammals and birds
| Host species (strain number [s]) or ECOR designationa | Order | Diet | Habitat | Country or place of origin |
|---|---|---|---|---|
| Gopherus flavomarginatus (57) | Reptilia | Herbivore | Wild | Mexico |
| Tachyglosus aculeatus (TA309–311) | Monotremata | Insects | Wild | Australia |
| Ornithorhynchus anatinus (TA319, 320) | Monotremata | Insects | Wild | Australia |
| Marmosa canescens (64) | Marsupialia | Omnivore | Wild | Mexico |
| Didelphis virginiana (79, 2048–2050) | Marsupialia | Omnivore | Wild | Mexico |
| Philander opossum (1698) | Marsupialia | Omnivore | Wild | Mexico |
| Cercatetus concinnus (TA002) | Marsupialia | Omnivore | Wild | Australia |
| Antechinus bellus (TA148) | Marsupialia | Insects | Wild | Australia |
| Antechinus flavipes (TA237) | Marsupialia | Insects | Wild | Australia |
| Antechinus swainsonii (TA153) | Marsupialia | Insects | Wild | Australia |
| Dasyurus geofroii (TA124) | Marsupialia | Carnivore | Wild | Australia |
| Dasyurus hallucatus (TA260) | Marsupialia | Carnivore | Wild | Australia |
| Dasyurus maculatus (TA036) | Marsupialia | Carnivore | Wild | Australia |
| Sminthopsis macroura (TA244) | Marsupialia | Insects | Wild | Australia |
| Sminthopsis murina (TA151) | Marsupialia | Insects | Wild | Australia |
| Sminthopsis delichura (TA120) | Marsupialia | Insects | Wild | Australia |
| Bettongia penicillata (TA115) | Marsupialia | Herbivore | Wild | Australia |
| Lagorchestes hirsutus (TA263) | Marsupialia | Herbivore | Wild | Australia |
| Macropus eugenii (TA052) | Marsupialia | Herbivore | Wild | Australia |
| Macropus fulginosus (TA004) | Marsupialia | Herbivore | Wild | Australia |
| Macropus giganteus (TA057) | Marsupialia | Herbivore | Wild | Australia |
| Petrogale lateralis (TA135) | Marsupialia | Herbivore | Wild | Australia |
| Perameles nasuta (TA020) | Marsupialia | Insects | Wild | Australia |
| Petaurus breviceps (TA243) | Marsupialia | Omnivore | Wild | Australia |
| Pseudocheirus peregrinus (TA022) | Marsupialia | Herbivore | Wild | Australia |
| Dasypus sp. (2395, 2396) | Xerarntha | Omnivore | Wild | Mexico |
| Nyctomys sp. (69) | Rodentia | Granivore | Wild | Mexico |
| Baiomys musculus (75) | Rodentia | Granivore | Wild | Mexico |
| Habromys sp. (88–89) | Rodentia | Granivore | Close to city | Mexico |
| Peromyscus sp. (71–73) | Rodentia | Granivore | Wild | Mexico |
| Peromyscus megalops (95) | Rodentia | Granivore | Wild | Mexico |
| Peromyscus boylii (96, 97) | Rodentia | Granivore | Wild | Mexico |
| Oryzomys sp. (58–60) | Rodentia | Granivore | Wild | Mexico |
| Sigmodon mascotensis (67, 68) | Rodentia | Omnivore | Wild | Mexico |
| Neotoma albigula (296–298) | Rodentia | Omnivore | Wild | Mexico |
| Zyzomys argurus (TA140, -141, -150) | Rodentia | Granivore | Wild | Australia |
| Rattus rattus (TA201, -063) | Rodentia | Omnivore | Close to city | Australia |
| Rattus lutreolus (TA053, -107) | Rodentia | Omnivore | Wild | Australia |
| Rattus fuscipes (TA001, -216) | Rodentia | Omnivore | Wild | Australia |
| Pseudomys apodemoides (TA093, -096) | Rodentia | Herbivore | Wild | Australia |
| Mus musculus (TA072, -074, -105) | Rodentia | Omnivore | City | Australia |
| Mus musculus (3490–3492) | Rodentia | Omnivore | City | Mexico |
| Notomys mitchelli (TA098, -100, -102) | Rodentia | Granivore | Wild | Australia |
| Hydrochaeris hydrochaeris (2079, 2080) | Rodentia | Herbivore | Zoo | Venezuela |
| Liomys pictus (76, 78) | Rodentia | Granivore | Wild | Mexico |
| Liomys pictus (19–21) | Rodentia | Granivore | Zoo | Mexico |
| Dipodomys merriami (286–288) | Rodentia | Granivore | Wild | Mexico |
| Perognathus penicillatus (820–822) | Rodentia | Granivore | Wild | Mexico |
| Tursiops truncatus (2025–2027) | Cetacea | Carnivore | Wild | Mexico |
| Felis concolor (268, 269) | Carnivora | Carnivore | Wild | Mexico |
| Felis catus (8–10) | Carnivora | Omnivore | Domestic | Mexico |
| Panthera onca (65, 66, 270) | Carnivora | Carnivore | Wild | Mexico |
| Lutra longicaudis (1930–1932) | Carnivora | Carnivore | Zoo | Mexico |
| Trichosurus vulpecula (TA038) | Marsupialia | Omnivore | Wild | Australia |
| Phascolarctos cinereus (TA003) | Marsupialia | Herbivore | Wild | Australia |
| Sorex sp. (238, 240) | Insectivora | Insects | Wild | Mexico |
| Tachyglosus aculeatus (TA051) | Chiroptera | Insects | Wild | Australia |
| Tadarida brasiliensis (259, 261, 266) | Chiroptera | Insects | Wild | Mexico |
| Desmodus rotundus (25, 26) | Chiroptera | Blood | Close to city | Mexico |
| Leptonycteris nivalis (31, 33, 36) | Chiroptera | Nectar | Close to city | Mexico |
| Carollia brevicauda (40–42) | Chiroptera | Nectar | Wild | Mexico |
| Nyctophilus geoffroyi (TA225) | Chiroptera | Insects | Wild | Australia |
| Alouatta palliata (1638, 1639) | Primates | Herbivore | Wild | Mexico |
| Alouatta palliata (1731) | Primates | Herbivore | Zoo | Costa Rica |
| Alouatta pigra (1699) | Primates | Herbivore | Wild | Mexico |
| Ateles geoffroyi (1669) | Primates | Herbivore | Wild | Mexico |
| Papio cynocephalus (B184, -197, -307, -349, -427, -430) | Primates | Omnivore | Wild | Africa |
| ECOR7 orangutan | Primates | Omnivore | Zoo | United States |
| K-12 MG1655 | Primates | Omnivore | City | |
| Homo sapiens (45–47) | Primates | Omnivore | City | Mexico |
| ECOR1 human | Primates | Omnivore | City | United States |
| ECOR5 human | Primates | Omnivore | City | United States |
| ECOR8 human | Primates | Omnivore | City | United States |
| ECOR10 human | Primates | Omnivore | City | United States |
| ECOR11 human | Primates | Omnivore | City | Europe |
| ECOR12 human | Primates | Omnivore | City | Europe |
| ECOR26 human | Primates | Omnivore | City | United States |
| ECOR41 human | Primates | Omnivore | City | Australia |
| Sylvilagus sp. (80) | Lagomorpha | Herbivore | Wild | Mexico |
| Oryctolagus cuniculus (2355–2357) | Lagomorpha | Herbivore | Wild | Mexico |
| Romerolagus diazi (2372–2374) | Lagomorpha | Herbivore | Close to city | Mexico |
| Bassariscus astutus (84, 815–817) | Carnivora | Omnivore | Wild | Mexico |
| Nasua narica (2060, 2062) | Carnivora | Omnivore | Wild | Mexico |
| Procyon lotor (2064–2066) | Carnivora | Omnivore | Wild | Mexico |
| Canis latrans (90–92, 824, 825, 830) | Carnivora | Omnivore | Wild | Mexico |
| Canis familiaris (1–3) | Carnivora | Omnivore | Domestic | Mexico |
| Urocyon cinereoargenteus (272, 299–301) | Carnivora | Omnivore | Wild | Mexico |
| ECOR23 elephant | Pachydermata | Herbivore | Zoo | United States |
| Trichechus manatus (1735, 1736) | Sirenia | Herbivore | Wild | Mexico |
| Tapirus bairdii (1666–1668, 1937) | Perissodactyla | Herbivore | Wild | Mexico |
| Tapirus terrestris (2381, 2382) | Perissodactyla | Herbivore | Wild | Venezuela |
| Equus caballus (807, 808) | Perissodactyla | Herbivore | Domestic | Mexico |
| ECOR45 pig | Perissodactyla | Herbivore | Domestic | Asia |
| Tayassu tajacu (2055) | Artiodactyla | Herbivore | Wild | Mexico |
| Odocoeileus virginianus (63, 1984, 1985) | Artiodactyla | Herbivore | Wild | Mexico |
| ECOR27 giraffe | Artiodactyla | Herbivore | Zoo | United States |
| Bos taurus (2376, 2379) | Artiodactyla | Herbivore | Domestic | Mexico |
| ECOR33 sheep | Artiodactyla | Herbivore | Domestic | United States |
| Ovis aries (1965–1967) | Artiodactyla | Herbivore | Close to city | Mexico |
| Sula nebouxi (137) | Aves | Carnivore | Wild | Mexico |
| Aquila chrysaetus (55) | Aves | Carnivore | Wild | Mexico |
| Aratinga canicularis (48) | Aves | Frugivore | Wild | Mexico |
| Calothorax lucifer (207) | Aves | Nectar | Wild | Mexico |
| Amazilia beryllina (225) | Aves | Nectar | Wild | Mexico |
| Henicorhina leucophrys (151) | Aves | Insects | Wild | Mexico |
| Basileuterus belli (108) | Aves | Insects | Wild | Mexico |
| Atlapetes brunneinucha (126) | Aves | Insects | Wild | Mexico |
| Passer domesticus (345) | Aves | Granivore | Wild | Mexico |
| Empidonax sp. (161) | Aves | Insects | Wild | Mexico |
For additional information regarding the ECOR isolates, see reference 32.
REFERENCES
- 1.Baquero F, Blazquez J. Evolution of antibiotic resistance. Trends Ecol Evol. 1998;12:482–488. doi: 10.1016/s0169-5347(97)01223-8. [DOI] [PubMed] [Google Scholar]
- 2.Blattner F R, Plunkett I G, Bloch C A, Perna N T, Burland V, Riley M, Collado-Vides J, Glasner J D, Rode C K, Mayhew G F, Gregor J, Davis N W, Kirkpatrick H A, Goeden M A, Rose D J, Mau B, Shao Y. The complete genome sequence of Escherichia coli K-12. Science. 1997;277:1453–1462. doi: 10.1126/science.277.5331.1453. [DOI] [PubMed] [Google Scholar]
- 3.Boyd E F, Hill C W, Rich S M, Hartl D L. Mosaic structure of plasmids from natural populations of Escherichia coli. Genetics. 1996;143:1091–1100. doi: 10.1093/genetics/143.3.1091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Campbell I C. Island kingdom: Tonga ancient and modern. Christchurch, New Zealand: Canterbury University Press; 1992. [Google Scholar]
- 5.Caugant D A, Levin B R, Selander R K. Genetic diversity and temporal variation in the E. coli population of a human host. Genetics. 1981;98:467–490. doi: 10.1093/genetics/98.3.467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Caugant D A, Levin B R, Lidin-Janson G, Whittam T S, Svanborg Eden C, Selander R K. Genetic diversity and relationships among strains of Escherichia coli in the intestine and those causing urinary tract infections. Prog Allergy. 1983;33:203–227. doi: 10.1159/000318331. [DOI] [PubMed] [Google Scholar]
- 7.Caugant D A, Mocca L F, Frasch C E, Frøholm L O, Zollinger W D, Selander R K. Genetic structure of Neisseria meningitidis populations in relation to serogroup, serotype, and outer membrane protein pattern. J Bacteriol. 1987;169:2781–2792. doi: 10.1128/jb.169.6.2781-2792.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cravioto A, Tello A, Navarro A, Ruiz J, Villafán H, Uribe F, Eslava C. Association of Escherichia coli Hep-2 adherence patterns with type and duration of diarrhoea. Lancet. 1991;337:262–264. doi: 10.1016/0140-6736(91)90868-p. [DOI] [PubMed] [Google Scholar]
- 9.Denny T P, Gilmour M N, Selander R K. Genetic diversity and relationships of two pathovars of Pseudomonas syringae. J Gen Microbiol. 1988;134:1949–1960. doi: 10.1099/00221287-134-7-1949. [DOI] [PubMed] [Google Scholar]
- 10.Duncan K E, Ferguson N, Kimura K, Zhou X, Istock C A. Fine-scale genetic and phenotypic structure in natural populations of Bacillus subtilis and Bacillus licheniformis: implications for bacterial evolution and speciation. Evolution. 1994;48:2002–2025. doi: 10.1111/j.1558-5646.1994.tb02229.x. [DOI] [PubMed] [Google Scholar]
- 11.Dykhuizen D E, Green L. Recombination in Escherichia coli and the definition of biological species. J Bacteriol. 1991;173:7257–7268. doi: 10.1128/jb.173.22.7257-7268.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Eckhardt T. A rapid method for the identification of plasmid deoxyribonucleic acid in bacteria. Plasmid. 1978;1:584–588. doi: 10.1016/0147-619x(78)90016-1. [DOI] [PubMed] [Google Scholar]
- 13.Ewing W H. Edwards and Ewing’s identification of Enterobacteriaceae; 4th ed. New York, N.Y: Elsevier Science Publishing Co.; 1986. [Google Scholar]
- 14.Gordon D M, Wexler M, Reardon T B, Murphy P J. The genetic structure of Rhizobium populations. Soil Biol Biochem. 1995;27:491–499. [Google Scholar]
- 15.Guttman D S, Dykhuizen D E. Clonal divergence in Escherichia coli as a result of recombination, not mutation. Science. 1994;266:1380–1383. doi: 10.1126/science.7973728. [DOI] [PubMed] [Google Scholar]
- 16.Hagen M J, Hamrick J L. Population level processes in Rhizobium leguminosarum bv. trifolii: the role of founder effects. Mol Ecol. 1996;5:707–714. doi: 10.1111/j.1365-294x.1996.tb00305.x. [DOI] [PubMed] [Google Scholar]
- 17.Hebert P D N, Beaton M J. Methodologies for allozyme analysis using cellulose acetate electrophoresis. Beaumont, Tex: Helena Laboratories; 1993. [Google Scholar]
- 18.Hughes V M, Datta N. Conjugative plasmids in bacteria of the “pre-antibiotic era”. Nature (London) 1983;302:725–726. doi: 10.1038/302725a0. [DOI] [PubMed] [Google Scholar]
- 19.Istock C A, Duncan K E, Ferguson N, Zhou X. Sexuality in a natural population of bacteria: Bacillus subtilis challenges the clonal paradigm. Mol Ecol. 1992;1:95–103. doi: 10.1111/j.1365-294x.1992.tb00161.x. [DOI] [PubMed] [Google Scholar]
- 20.Jensen R A. Biochemical pathways can be traced backward through evolutionary time. Mol Biol Evol. 1985;2:92–108. doi: 10.1093/oxfordjournals.molbev.a040338. [DOI] [PubMed] [Google Scholar]
- 21.Jones C, Stanley J. Salmonella plasmids of the pre-antibiotic era. J Gen Microbiol. 1992;138:189–197. doi: 10.1099/00221287-138-1-189. [DOI] [PubMed] [Google Scholar]
- 22.Joset F, Guespin-Michel J. Procaryotic genetics. Cambridge, Mass: Blackwell Science; 1993. [Google Scholar]
- 23.Lecointre G, Rachdi L, Darlu P, Denamur E. Escherichia coli molecular phylogeny using the incongruence length difference test. Mol Biol Evol. 1998;15:1685–1695. doi: 10.1093/oxfordjournals.molbev.a025895. [DOI] [PubMed] [Google Scholar]
- 24.Maloy S R, Cronan J E, Freifelder D. Microbial genetics. 2nd ed. Boston, Mass: Jones and Bartlett Publishers; 1994. [Google Scholar]
- 25.Maynard Smith J, Smith N H, O’Rourke M, Spratt B G. How clonal are bacteria? Proc Natl Acad Sci USA. 1993;90:4384–4388. doi: 10.1073/pnas.90.10.4384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Milkman R. Electrophoretic variation in Escherichia coli from natural sources. Science. 1973;182:1024–1026. doi: 10.1126/science.182.4116.1024. [DOI] [PubMed] [Google Scholar]
- 27.Miller R D, Hartl D L. Biotyping confirms a nearly clonal population structure in Escherichia coli. Evolution. 1986;40:1–12. doi: 10.1111/j.1558-5646.1986.tb05712.x. [DOI] [PubMed] [Google Scholar]
- 28.Musser J M, Barenkamp S J, Granoff D M, Selander R K. Genetic relationships of serologically nontypable and serotype b strains of Haemophilus influenzae. Infect Immun. 1986;52:183–191. doi: 10.1128/iai.52.1.183-191.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nei M. F-statistics and analysis of gene diversity in subdivided populations. Ann Hum Genet Lond. 1977;41:225–233. doi: 10.1111/j.1469-1809.1977.tb01918.x. [DOI] [PubMed] [Google Scholar]
- 30.Nei M. Genetic distance between populations. Amer Nat. 1972;106:283–292. [Google Scholar]
- 31.Nowak R M. Walker’s mammals of the world. 5th ed. Baltimore, Md: Johns Hopkins University Press; 1991. [Google Scholar]
- 32.Ochman H, Selander R K. Standard reference strains of Escherichia coli from natural populations. J Bacteriol. 1984;157:690–693. doi: 10.1128/jb.157.2.690-693.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Pupo G M, Richardson B J. Biochemical genetics of a natural population of Escherichia coli: seasonal changes in alleles and haplotypes. Microbiology. 1995;141:1037–1044. doi: 10.1099/13500872-141-4-1037. [DOI] [PubMed] [Google Scholar]
- 34.Rocha M. Undergraduate dissertation. México D.F.: Universidad Nacional Autónoma de México; 1996. [Google Scholar]
- 35.Routman E, Miller R D, Philips-Conroy J, Hartl D L. Antibiotic resistance and population structure in Escherichia coli from free-ranging African yellow baboons. Appl Environ Microbiol. 1985;50:749–754. doi: 10.1128/aem.50.4.749-754.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Schaechter M, Curtis R, Ingraham J, Lin E, Brooks K, Magasanik B, Neidhardt D, Reznikoff W, Riley M, Umbarger E. The view from here. In: Neidhardt F C, Ingraham J L, Low K B, Magasanik B, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella typhimurium: cellular and molecular biology. Washington, D.C: American Society for Microbiology; 1996. pp. 1625–1648. [Google Scholar]
- 37.Seigel S. Nonparametric statistics for the behavioral sciences. London, England: McGraw-Hill, Inc.; 1956. [Google Scholar]
- 38.Selander R K, Levin B R. Genetic diversity and structure in Escherichia coli. Science. 1980;210:545–547. doi: 10.1126/science.6999623. [DOI] [PubMed] [Google Scholar]
- 39.Selander R K, McKinney R M, Whittam T S, Bibb W F, Brenner D J, Nolte F S, Pattison P E. Genetic structure of populations of Legionella pneumophilia. J Bacteriol. 1985;163:1021–1037. doi: 10.1128/jb.163.3.1021-1037.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Selander R K, Caugant D A, Ochman H, Musser J M, Whittam T S. Methods of multilocus enzyme electrophoresis for bacterial population genetics and systematics. Appl Environ Microbiol. 1986;51:873–884. doi: 10.1128/aem.51.5.873-884.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Selander R K, Korhonen T K, Väisänen-Rhen V, Williams P H, Pattison P E, Caugant D A. Genetic relationships and clonal structure of strains of Escherichia coli causing neonatal septicemia and meningitis. Infect Immun. 1986;52:213–222. doi: 10.1128/iai.52.1.213-222.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Selander R K, Caugant D A, Whittam T S. Genetic structure and variation in natural populations of Escherichia coli. In: Neidhardt F C, Ingraham J L, Low K B, Magasanik B, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella typhimurium: cellular and molecular biology. Washington, D.C: American Society for Microbiology; 1987. pp. 1625–1648. [Google Scholar]
- 43.Selander R K, Beltran P, Smith N H, Helmuth R, Rubin F A, Kopecko D J, Ferris K, Tall B D, Cravioto A, Musser J M. Evolutionary genetic relationships of clones of Salmonella serovars that cause human typhoid and other enteric fevers. Infect Immun. 1990;58:2262–2275. doi: 10.1128/iai.58.7.2262-2275.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Smith H R, Scotland S M. Vero cytotoxin-producing strains of Escherichia coli. J Med Microbiol. 1988;26:77–85. doi: 10.1099/00222615-26-2-77. [DOI] [PubMed] [Google Scholar]
- 45.Souza V, Eguiarte L, Avila G, Cappello R, Gallardo C, Montoya J, Piñero D. Genetic structure of Rhizobium etli biovar phaseoli associated with wild and cultivated bean plants (Phaseolus vulgaris and Phaseolus coccineus) in Morelos, Mexico. Appl Environ Microbiol. 1994;60:1260–1268. doi: 10.1128/aem.60.4.1260-1268.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Whittam T S, Ochman H, Selander R K. Geographic components of linkage disequilibrium in natural populations of Escherichia coli. Mol Biol Evol. 1983;1:67–83. doi: 10.1093/oxfordjournals.molbev.a040302. [DOI] [PubMed] [Google Scholar]
- 47.Whittam T S, Ochman H, Selander R K. Multilocus genetic structure in natural populations of Escherichia coli. Proc Natl Acad Sci USA. 1983;80:1751–1755. doi: 10.1073/pnas.80.6.1751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Whittam T S. Clonal dynamics of Escherichia coli in its natural habitat. Antonie Leeuwenhoek. 1989;55:23–32. doi: 10.1007/BF02309616. [DOI] [PubMed] [Google Scholar]
- 49.Whittam T S, Wolfe M L, Wachsmuth I K, Ørskov F, Ørskov I, Wilson R A. Clonal relationships among Escherichia coli strains that cause hemorrhagic colitis and infantile diarrhea. Infect Immun. 1993;61:1619–1629. doi: 10.1128/iai.61.5.1619-1629.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Whittam T S. Genetic variation and evolutionary processes in natural populations of Escherichia coli. In: Neidhardt F C, Ingraham J L, Low K B, Magasanik B, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella typhimurium: cellular and molecular biology. Washington, D.C: American Society for Microbiology; 1996. pp. 2708–2728. [Google Scholar]