SUMMARY
This study evaluates the usefulness of spatio-temporal statistical tools to detect outbreaks using routine surveillance data where limited epidemiological information is available. A dataset from 2002 to 2007 containing information regarding date, origin, source and serotype of 29 586 Salmonella isolates from Thailand was analysed. Data was grouped into human and non-human categories and the analysis was performed for the top five occurring serovars for each year of the study period. A total 91 human and 39 non-human significant spatio-temporal clusters were observed, accounting for 11% and 16% of the isolates, respectively. Serovar-specific associations between human and non-human clusters were also evaluated. Results show that these statistical tools can provide information for use in outbreak prevention and detection, in countries where only limited data is available. Moreover, it is suggested that monitoring non-human reservoirs can be relevant in predicting future Salmonella human cases.
Key words: Analysis of data, infectious disease control, outbreaks, Salmonella
INTRODUCTION
Salmonella is a major cause of human gastroenteritis and bacteraemia in both the industrialized and developing world, with most frequent cases being foodborne infections [1, 2]. Although Salmonella serovars Typhimurium and Enteritidis are the most common causes of human salmonellosis worldwide, other serovars have been reported to be more prevalent in some regions, e.g. S. Stanley and S. Weltevreden that are very common in South East Asia [3–5].
In a globalized world with constant movement of people, animals and goods, the importance of monitoring infectious diseases is growing. Alert systems, able to detect changes in disease trends and risk factors as soon as possible, are essential to allow quick control measures and prevent outbreaks and epidemics [6, 7]. One of the biggest challenges when developing these tools is how to handle data limitations. Surveillance datasets are often hampered by the lack of sufficient quality data and confounded by multiple human and environmental interactions.
Determining the sources of the infection of salmonellosis is often done retrospectively using clinical case information. To develop efficient methods for prospective surveillance it is fundamental not only to understand the mechanisms of infection development and spread but also survey the more probable sources of infection. However, monitoring of the frequent reservoirs for Salmonella is, in most countries, only based in sporadic sample collections rather than in focused structured surveillance programmes.
Statistical scans are a mathematical tool commonly used in outbreak detection and evaluation from cancer to infectious diseases including Salmonella [8–11]. These tools scan the data looking for unexpected cases, patterns or trends in spatial, temporal and also spatio-temporal dimensions.
The main objective of this study was to investigate spatio-temporal clusters using statistical scan methods on routine monitoring data of Salmonella infections. The dataset comprised isolate data from human and non-human sources in Thailand from 2002 to 2007.
A second objective was to evaluate the existence of serovar-specific associations between human and non-human clusters. This type of information could be useful in complementing incomplete datasets and helping the establishment of effective surveillance systems.
MATERIAL AND METHODS
Data
In Thailand, the monitoring of Salmonella infections is based on a passive surveillance scheme where the National Institute of Health (NIH) – Salmonella and Shigella Section, receives the clinical isolates suspected to be Salmonella from the diagnostic laboratories, hospitals and medical clinics across the country. For each confirmed case, the relevant clinical and epidemiological information is recorded [12].
Data on Salmonella from other sources (e.g. animals or food) is based on sporadic and ad-hoc sampling schemes implemented by the Thai authorities [12].
For this study, we used a dataset containing data on 29 586 Salmonella isolates collected during the period 2002–2007 from both human and other sources. Besides date, location and source of the sample, the isolate's serotype was also registered.
Database management, descriptive statistics and data arrangement were performed in SAS Enterprise Guide 3.0 (SAS Institute, USA).
Spatio-temporal scan statistics
The cluster analysis was performed using spatio-temporal statistical scan methods available at SaTScanTM v. 9.0.1 platform (Information Management Services Inc., USA) [13].
Scan statistics analyses are usually done by moving a scanning window (with different possibilities for shapes and sizes) through the space and time dimension of the data. For each location and window size, the number of observed and expected cases is compared and any not-expected excesses on the number of observed cases registered. The statistical significance of each potential cluster is then evaluated [8, 13]. Each cluster is determined independently so that not only the most likely cluster (MLC) is detected, but also all statistical significant clusters are as well [13, 14]. By doing a simultaneous spatial and temporal analysis it is possible to detect clusters that would not be apparent if looking only into one dimension. Spatio-temporal clusters can capture the mechanisms of infection spreading much more realistically [15].
For the human isolates, a retrospective space–time permutation scan statistic method was used. This method uses only case numbers, and does not require data on the background population at risk [16]. It only requires minimal assumptions about time and geographical location and has the advantage of adjusting automatically for natural purely spatial or temporal variation (e.g. seasonal variation) [8]. A cluster is detected in a region, if during a specific time interval there is a high proportion of excess cases or a smaller deficiency of cases than in the neighbouring regions [16]. The analyses were serovar-specific and performed by year (using a month as a unit of observation).
A Bernoulli retrospective scan statistic was used for the non-human isolates. This model is independent of the underlying population using the data grouped as cases vs. controls to determine if there is significant clustering of the distribution of cases compared to the distribution of controls [10, 17].
The analyses were per serovar, so the isolates from each specific serovar were defined as cases, while the isolates from the remaining serovars were defined as controls.
Regarding the human isolates, the analyses were performed by year (using a month as a unit of observation).
Other pre-defined settings common to both methods were: a scanning window with an elliptical shape, a maximum spatial cluster size of 50% of the population at risk, a maximum temporal cluster size of 50% of the study period (6 months), and no restrictions for reporting secondary clusters. A cluster was considered significant if the P value for its calculated likelihood was <0·05 and 999 Monte Carlo simulations were used for each assessment.
Besides the MLC, all significant secondary clusters were reported in the tool output. Secondary clusters with no overlap with the MLC were considered in the results. In contrast, secondary clusters that were just variations of the MLC were discarded, as the only information provided by those refers to some uncertainty on the exact boundaries (either in time or space) of the MLC.
The existence of a possible association between a human and a non-human cluster was also evaluated. We considered two clusters possibly associated if the clusters overlapped in time and space, if the clusters overlapped only in one dimension (either time or space) and were adjacent in the other or if the clusters were adjacent in both time and space.
The geographical information system ArcMap 9.0 (Environmental Systems Research Institute, USA) was used for providing spatial coordinates and for visualizing the clusters. The Google charts API tool (Google Inc., USA) was also used for cluster visualization.
RESULTS
Data description
The 29 586 isolates were collected from different sources. The human isolates accounted for 65% of the total, isolates from food sources represented 19% and animal sources 7%. Isolates from a non-descriptive ‘other’ source (e.g. environment) were 9% of the data.
Apart from the human cases, the data collected from other sources had too few isolates in each subcategory (e.g. food, animal, environmental or other) to allow a reliable analysis, so the data was aggregated into a generic non-human data category. A total of 194 and 177 different serovars were identified for the human and non-human categories, respectively (results not shown).
The data distribution throughout the study period was not uniform: 28% of the isolates were from 2002, 13% from 2003, 18% from 2004, 18% from 2005, 13% from 2006 and 11% from 2007. By performing the analyses for the complete study period as whole, the results would be driven by the year 2002, hiding possible clusters occurring later. The study period was then divided into years and the analyses run separately.
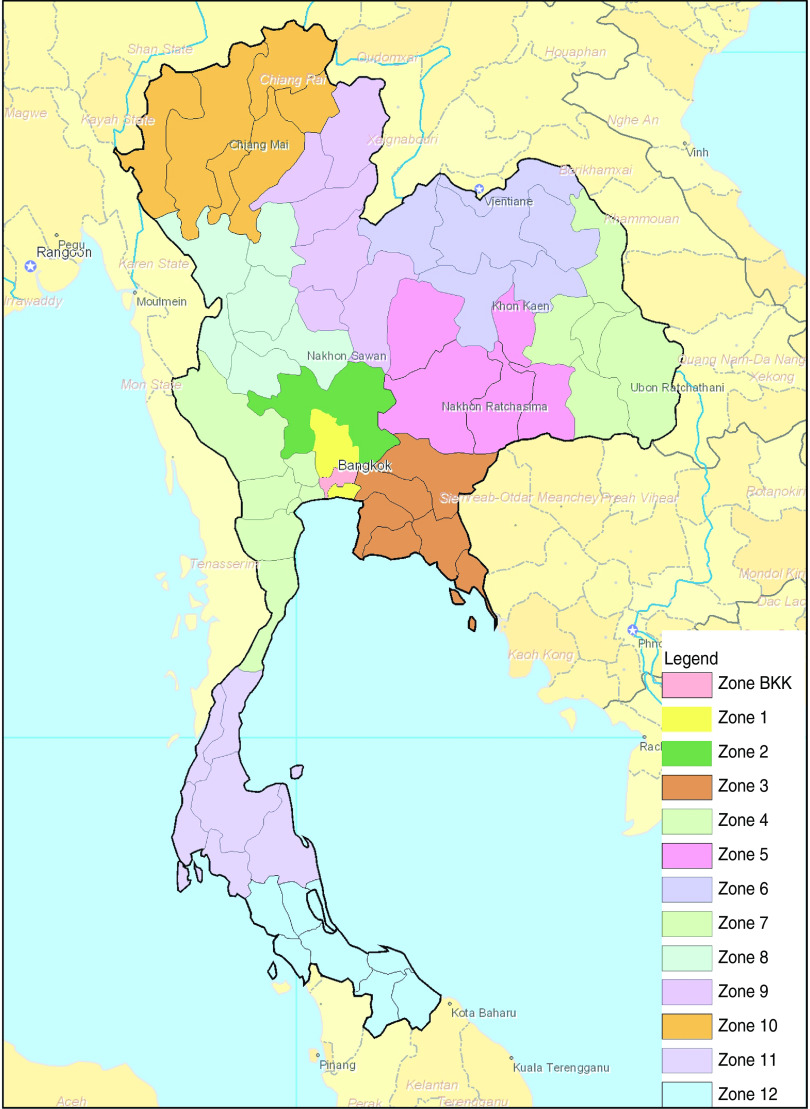
Thailand is organized into four main geographical regions: Central, Northeastern, Northern and Southern. These regions are further divided into 13 administrative zones (zones 1–12 and Bangkok), which are again further organized into 76 provinces [12, 18, 19]. The dataset contained data collected within 55 of the 76 provinces. Bangkok was the most represented province (34% of the isolates) followed by Ratchaburi (9%), Nonthaburi (8%), Khon Kaen (8%), Chiang Mai (6%), the remaining provinces each represented <5% of the isolates. The low number of isolates in some provinces was insufficient to allow an analysis at province level, so instead it was done at the zone level. Figure 1 shows a map of Thailand highlighting the 13 different administrative zones.
Fig. 1.
[colour online]. Map of Thailand representing the official 13 zones (map reproduced courtesy of the National Institute of Health, Thailand; modified with permission). BKK, Bangkok.
For each year, the top five most common serovars in the human category were selected for analysis (Table 1). S. Enteritidis, S. Stanley and S. Rissen were the most common serovars being present in the 6 years analysed, followed by S. Weltevreden (5 years), S. Anatum (3 years), S. Cholerasuis (2 years), S. Corvallis (1 year) and S. Typhimurium (1 year). However, in three instances, S. Cholerasuis (2006 and 2007) and S. Enteritidis (2007), there were not enough isolates in the non-human dataset to run the analysis, so the isolates of S. Corvallis (2006) and S. Weltevreden and S. Anatum (2007) were used instead.
Table 1.
Five most frequent serovars (and respective number of isolates) per year in the human dataset
| Rank | 2002 | 2003 | 2004 | 2005 | 2006 | 2007 |
|---|---|---|---|---|---|---|
| 1 | S. Enteritidis (515) | S. Enteritidis (304) | S. Stanley (447) | S. Stanley (456) | S. Enteritidis (541) | S. Enteritidis* (440) |
| 2 | S. Rissen (327) | S. Stanley (220) | S. Enteritidis (368) | S. Enteritidis (403) | S. Stanley (313) | S. Stanley (297) |
| 3 | S. Weltevreden (322) | S. Rissen (174) | S. Weltevreden (365) | S. Weltevreden (331) | S. Choleraesuis* (233) | S. Rissen (219) |
| 4 | S. Anatum (317) | S. Weltevreden (166) | S. Rissen (300) | S. Rissen (324) | S. Rissen (224) | S. Choleraesuis* (211) |
| 5 | S. Stanley (260) | S. Anatum (157) | S. Anatum (255) | S. Corvallis (208) | S. Weltevreden (202) | S. Typhimurium (202) |
| — | S. Corvallis (122) | S. Weltevreden (164) | ||||
| — | S. Anatum (122) |
Values in parentheses are number of isolates.
For three of the serovars (indicated by *) there were not enough isolates in the non-human dataset to run the analysis, therefore the analysis was run for the next serovar in the list that had enough available data.
Spatio-temporal scan statistics
A total of 91 human (involving 11% of the total human isolates) and 39 non-human (involving 16% of the total non-human isolates) significant spatio-temporal clusters were found distributed throughout the 6 years of data. The summarization of the results per year and serovar is shown on Table 2, while a complete description of the detected clusters for human and non-human isolates can be found in Tables 3 and 4, respectively. Figures 2 and 3 illustrate the clusters detected in 2003 for both categories.
Table 2.
Number of human and non-human clusters and respective associations detected per year and serovar
| No. of human clusters (n = 91) | No. of non-human clusters (n = 39) | No. of associations detected (%) (n = 23) | |
|---|---|---|---|
| Year | |||
| 2002 | 17 | 6 | 5 (29·4) |
| 2003 | 8 | 7 | 3 (37·5) |
| 2004 | 19 | 7 | 5 (26·3) |
| 2005 | 16 | 6 | 2 (12·5) |
| 2006 | 17 | 7 | 5 (29·4) |
| 2007 | 14 | 6 | 3 (21·4) |
| Serovar | |||
| S. Enteritidis | 16 | 5 | 2 (12·5) |
| S. Rissen | 21 | 9 | 8 (38·1) |
| S. Weltevreden | 16 | 8 | 5 (31·3) |
| S. Anatum | 14 | 7 | 4 (28·6) |
| S. Stanley | 16 | 8 | 4 (25) |
| S. Corvallis | 5 | 2 | 0 |
| S. Typhimurium | 3 | 0 | 0 |
Table 3.
Description of the clusters detected in the human dataset in the study period, 2002–2007
| Cluster ID | Serovar | Year | Zones | Duration | No. of isolates |
|---|---|---|---|---|---|
| 1 | S. Anatum | 2002 | 5 | Feb. | 12 |
| 2 | S. Anatum | 2002 | 3 + 11 | May | 7 |
| 3 | S. Anatum | 2002 | 2 | Aug. | 5 |
| 4 | S. Anatum | 2002 | 10 | Oct. | 7 |
| 5 | S. Anatum | 2002 | 1 + 2 | Nov. | 33 |
| 6 | S. Enteritidis | 2002 | Bangkok+5 | Feb. | 85 |
| 7 | S. Enteritidis | 2002 | 11 + 12 | June | 24 |
| 8 | S. Enteritidis | 2002 | Bangkok | July | 81 |
| 9 | S. Enteritidis | 2002 | 1 + 2+4 | Nov.–Dec. | 26 |
| 10 | S. Stanley | 2002 | 2 | Nov. | 13 |
| 11 | S. Rissen | 2002 | 3 + 11 + 12 | Jan. | 8 |
| 12 | S. Rissen | 2002 | 3 + 11 | May | 6 |
| 13 | S. Rissen | 2002 | Bangkok+4 | July | 26 |
| 14 | S. Rissen | 2002 | 8 | Dec. | 6 |
| 15 | S. Weltevreden | 2002 | 12 | Jan. | 45 |
| 16 | S. Weltevreden | 2002 | Bangkok+3 | May–Aug. | 40 |
| 17 | S. Weltevreden | 2002 | 8 + 9+2 + 10 + 1+4 | Oct.–Dec. | 32 |
| 18 | S. Anatum | 2003 | 4 | Jan. | 13 |
| 19 | S. Anatum | 2003 | 6 | June–July | 13 |
| 20 | S. Anatum | 2003 | 2 + 5+7 + 8+9 | Aug. | 9 |
| 21 | S. Enteritidis | 2003 | 6 + 9+10 | June | 13 |
| 22 | S. Enteritidis | 2003 | 1 | Nov.–Dec. | 19 |
| 23 | S. Stanley | 2003 | 6 + 8+9 | Nov. | 13 |
| 24 | S. Rissen | 2003 | 6 | July | 19 |
| 25 | S. Rissen | 2003 | 1 + 2+8 + 9 | Sep. | 7 |
| 26 | S. Anatum | 2004 | 4 | July | 34 |
| 27 | S. Anatum | 2004 | 12 | Aug | 7 |
| 28 | S. Anatum | 2004 | 3 + 5+6 | Sep. | 8 |
| 29 | S. Anatum | 2004 | 10 | Oct.–Dec. | 20 |
| 30 | S. Enteritidis | 2004 | Bangkok+1 + 3 | Mar.–Apr. | 39 |
| 31 | S. Enteritidis | 2004 | 12 | June–Aug. | 81 |
| 32 | S. Enteritidis | 2004 | 10 | Oct.–Dec. | 74 |
| 33 | S. Stanley | 2004 | 5 | Jan. | 27 |
| 34 | S. Stanley | 2004 | 4 + 8 | May | 26 |
| 35 | S. Stanley | 2004 | 1 + 2 | July | 10 |
| 36 | S. Stanley | 2004 | 10 | Oct.–Dec. | 33 |
| 37 | S. Rissen | 2004 | 1 | Mar.–May | 22 |
| 38 | S. Rissen | 2004 | 6 | June | 26 |
| 39 | S. Rissen | 2004 | 2 + 4 | July | 13 |
| 40 | S. Rissen | 2004 | 12 | Aug. | 12 |
| 41 | S. Rissen | 2004 | 10 | Oct.–Dec. | 20 |
| 42 | S. Weltevreden | 2004 | 1 + 3 | Mar.–Apr. | 23 |
| 43 | S. Weltevreden | 2004 | 12 | June | 62 |
| 44 | S. Weltevreden | 2004 | 10 | Oct.–Dec. | 30 |
| 45 | S. Enteritidis | 2005 | 5 + 9+10 | Feb. | 21 |
| 46 | S. Enteritidis | 2005 | 1 + 2+4 | Mar. | 21 |
| 47 | S. Enteritidis | 2005 | 12 | May–July | 47 |
| 48 | S. Enteritidis | 2005 | 3 + 5 | Aug. | 27 |
| 49 | S. Stanley | 2005 | 4 | Mar. | 22 |
| 50 | S. Stanley | 2005 | 12 | May–July | 54 |
| 51 | S. Stanley | 2005 | 2 + 8+9 + 10 | Nov.–Dec. | 45 |
| 52 | S. Stanley | 2005 | 6 | May | 24 |
| 53 | S. Rissen | 2005 | 1 | June | 24 |
| 54 | S. Rissen | 2005 | 12 | May–Sep. | 36 |
| 55 | S. Rissen | 2005 | 8 + 10 | Dec. | 20 |
| 56 | S. Weltevreden | 2005 | 5 + 7+9 | Feb. | 15 |
| 57 | S. Weltevreden | 2005 | 12 | May–July | 56 |
| 58 | S. Weltevreden | 2005 | 8 + 10 | Dec. | 24 |
| 59 | S. Corvallis | 2005 | 12 | May–July | 26 |
| 60 | S. Corvallis | 2005 | 1 + 5+7 | Aug. | 9 |
| 61 | S. Enteritidis | 2006 | 10 | Apr.–May | 42 |
| 62 | S. Enteritidis | 2006 | 6 + 9 | Aug. | 37 |
| 63 | S. Enteritidis | 2006 | 3 + 5 | Dec. | 46 |
| 64 | S. Stanley | 2006 | 5 + 9+10 | Apr. | 34 |
| 65 | S. Stanley | 2006 | 4 | July | 18 |
| 66 | S. Stanley | 2006 | Bangkok+1 + 2 | Aug.–Oct. | 41 |
| 67 | S. Rissen | 2006 | 6 | Feb. | 9 |
| 68 | S. Rissen | 2006 | 5 + 9+10 | Apr. | 16 |
| 69 | S. Rissen | 2006 | 4 | July | 16 |
| 70 | S. Rissen | 2006 | 2 + 5+7 | Dec. | 9 |
| 71 | S. Weltevreden | 2006 | 3 | Jan. | 10 |
| 72 | S. Weltevreden | 2006 | 5 + 9+10 | Apr. | 36 |
| 73 | S. Weltevreden | 2006 | 4 | July | 11 |
| 74 | S. Weltevreden | 2006 | 2 + 5+7 | Dec. | 16 |
| 75 | S. Corvallis | 2006 | 6 | Feb. | 3 |
| 76 | S. Corvallis | 2006 | 5 | Apr. | 3 |
| 77 | S. Corvallis | 2006 | 1 + 2+4 | July–Oct. | 26 |
| 78 | S. Anatum | 2007 | 8 + 9 | July | 14 |
| 79 | S. Anatum | 2007 | 3 + 5 | Aug. | 3 |
| 80 | S. Stanley | 2007 | 6 | May | 14 |
| 81 | S. Stanley | 2007 | 5 | Aug. | 12 |
| 82 | S. Stanley | 2007 | 2 + 8+9 + 10 | Dec. | 13 |
| 83 | S. Rissen | 2007 | 6 | May | 11 |
| 84 | S. Rissen | 2007 | 5 | Aug. | 8 |
| 85 | S. Rissen | 2007 | 1 | Sep.–Oct. | 16 |
| 86 | S. Weltevreden | 2007 | 6 | May | 13 |
| 87 | S. Weltevreden | 2007 | 5 | Aug. | 7 |
| 88 | S. Weltevreden | 2007 | 10 | Oct. | 10 |
| 89 | S. Typhimurium | 2007 | 6 + 9 | May | 11 |
| 90 | S. Typhimurium | 2007 | 3 + 5+7 + 11 + 12 | Aug. | 7 |
| 91 | S. Typhimurium | 2007 | 4 | Jan. | 9 |
For the number representing the geographical zones, refer to the key in Figure 1.
Table 4.
Description of the clusters detected in the non-human dataset in the study period, 2002–2007
| Cluster ID | Serovar | Year | Zones | Duration | No. of isolates |
|---|---|---|---|---|---|
| 1 | S. Anatum | 2002 | 6 | Apr. | 111 |
| 2 | S. Enteritidis | 2002 | 2 | Nov. | 35 |
| 3 | S. Stanley | 2002 | 1 | June–Nov. | 49 |
| 4 | S. Rissen | 2002 | 2 + 6+8 + 9+10 | Jan.–Apr. | 99 |
| 5 | S. Weltevreden | 2002 | 9 + 10 | Apr.–Aug. | 62 |
| 6 | S. Weltevreden | 2002 | 4 + 1 | Sep.–Dec. | 33 |
| 7 | S.Anatum | 2003 | Bangkok | Oct. | 18 |
| 8 | S.Anatum | 2003 | 6 + 8+9 + 10 | Nov.–Dec. | 78 |
| 9 | S. Enteritidis | 2003 | Bangkok+2 + 4+8 | Jan.–Apr. | 62 |
| 10 | S. Stanley | 2003 | 1 | Jan.–June | 27 |
| 11 | S. Rissen | 2003 | 1 | Feb. | 6 |
| 12 | S. Rissen | 2003 | 6 + 8+9 + 10 | July–Dec. | 90 |
| 13 | S. Weltevreden | 2003 | Bangkok | Dec. | 57 |
| 14 | S.Anatum | 2004 | 1 | Mar. | 23 |
| 15 | S.Anatum | 2004 | Bangkok+3 | May–June | 52 |
| 16 | S.Anatum | 2004 | 2 + 4+8 + 9+10 | Sep. | 10 |
| 17 | S. Enteritidis | 2004 | Bangkok | Sep. | 56 |
| 18 | S. Stanley | 2004 | Bangkok+4 | Aug. | 41 |
| 19 | S. Rissen | 2004 | Bangkok | Apr.–June | 58 |
| 20 | S. Weltevreden | 2004 | Bangkok | Oct.–Nov. | 31 |
| 21 | S. Enteritidis | 2005 | 3 | Dec. | 20 |
| 22 | S. Stanley | 2005 | Bangkok+4 + 11 + 12 | Feb. | 51 |
| 23 | S. Rissen | 2005 | Bangkok | Feb. | 74 |
| 24 | S. Rissen | 2005 | 5 + 6+7 + 8+9 | May–Oct. | 113 |
| 25 | S. Weltevreden | 2005 | Bangkok+4 + 8+10 | July–Dec. | 98 |
| 26 | S. Corvallis | 2005 | Bangkok | Oct. | 10 |
| 27 | S. Enteritidis | 2006 | 3 | Feb. | 29 |
| 28 | S. Stanley | 2006 | 1 + 2+6 + 8+9 + 10 | Feb.–May | 32 |
| 29 | S. Stanley | 2006 | 1 + 2+7 + 11 + 12 | Aug. | 6 |
| 30 | S. Rissen | 2006 | 1 + 2+4 + 6+8 + 9+10 | Jan.–Mar. | 29 |
| 31 | S. Weltevreden | 2006 | Bangkok+4 | May–Aug. | 48 |
| 32 | S. Weltevreden | 2006 | 3 | Aug.–Dec. | 46 |
| 33 | S. Corvallis | 2006 | Bangkok+4 | Jan.–Feb. | 39 |
| 34 | S. Anatum | 2007 | 3 | Jan.–Apr. | 12 |
| 35 | S. Stanley | 2007 | 4 | Mar. | 8 |
| 36 | S. Stanley | 2007 | 2 | Dec. | 7 |
| 37 | S. Rissen | 2007 | 2 + 4 | Apr.–Sep. | 23 |
| 38 | S. Rissen | 2007 | 1 + 2 | Feb.–Mar. | 13 |
| 39 | S. Weltevreden | 2007 | Bangkok+4 + 8 | Nov. | 6 |
For the number representing the geographical zones, refer to the key in Figure 1.
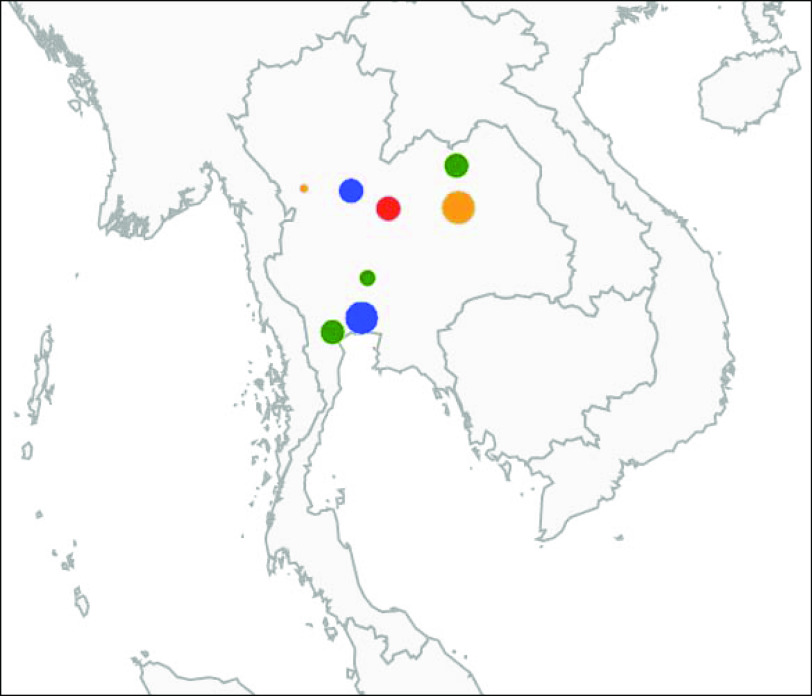
Fig. 2.
[colour online]. Representation of the human clusters detected in 2003. Each marker colour represents a serovar (blue: S. Anatum; green: S. Enteritidis; red: S. Stanley; orange: S. Rissen). The size of the markers is scaled according to the number of isolates involved in each cluster. The clusters represented are cluster numbers 18, 19, 20, 21, 22, 23, 24 and 25 from Table 3.
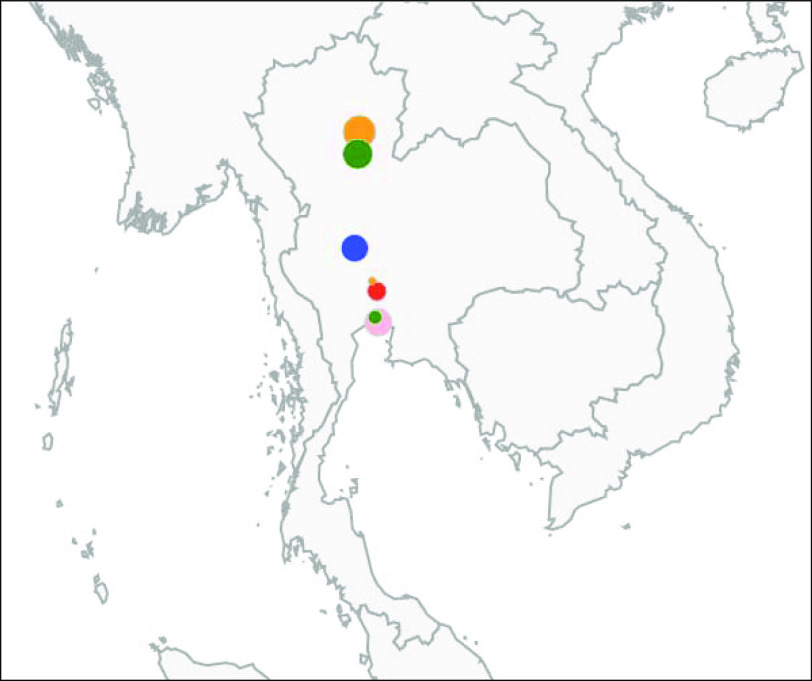
Fig. 3.
[colour online]. Representation of the non-human clusters detected in 2003. Each marker colour represents a serovar (blue: S. Anatum; green: S. Enteritidis; red: S. Stanley; orange: S. Rissen; pink: S. Weltevreden). The size of the markers is scaled according to the number of isolates involved in each cluster. The clusters represented are cluster numbers 7, 8, 9, 10, 11, 12 and 13 from Table 4.
In the results per year, for the human clusters, the number of detected clusters ranged from nine (2003) to 19 (2004). For the non-human clusters, the numbers were very similar between the years.
Looking at the results by serovar, it was for S. Rissen that more clusters were detected in both the human (21 clusters) and non-human (nine clusters) categories, followed by S. Weltevreden and S. Stanley each with 16 clusters in the human category and eight in the non-human group.
DISCUSSION
This study presents a retrospective spatio-temporal statistical scan analysis of Salmonella isolates in Thailand during the period 2002–2007. The objectives of the analysis were to evaluate the existence of significant clusters and possible relationships between the human and non-human clusters. We also discuss the usefulness of spatio-temporal statistical tools in analysing data with limited epidemiological information.
The need for efficient early-alert detection systems for infectious diseases is growing. Public health authorities have to be able to quickly assess a possible outbreak and take the appropriate actions to control it.
Mathematical methods that can quickly scan the data looking for unexpected patterns, in time and space, while adjusting for known covariates or risk factors (e.g. climate or for foodborne bacteria, consumption habits) are fundamental in prospective surveillance systems.
In this study, we look at not only clinical data but also data collected from common reservoirs for Salmonella.
For the analysis of human isolates, a space–time permutation model was used. The fact that the method does not require data on population at risk makes it a good choice when facing data limitations. Moreover, because it can adjust automatically for purely geographical variations (e.g. population density) and purely temporal variations (e.g. seasonal patterns) [20], it reduces the effect of possible data bias. The scan detected 91 significant clusters involving 11% of the total number of cases reported. The percentage of total reported salmonellosis cases associated with outbreaks has been estimated to be between 5 and 10% in New Zealand and in Europe [21, 22]. It was not possible to further investigate each cluster to evaluate if it corresponded to a true outbreak, but the results tend to agree with previous findings.
As an example, for S. Weltevreden, most of the clusters detected occurred in coastal regions (see cluster ID nos. 15, 16, 42, 43, 57 in Table 3). This is in agreement with Hendriksen et al. [23] who describe S. Weltevreden as being commonly associated with seafood. Moreover, for S. Rissen and S. Stanley, the results predict more clusters occurring in the central part of Thailand, which is a region associated with agriculture and more specifically intensive pig farming. Once again, in agreement with Hendriksen et al. [24], these serovars are frequently associated with pigs.
The data collection for the non-human category was mostly based on specific monitoring initiatives rather than in an established surveillance programme. Thus, there is both over- and under-representation of regions, periods of the year and reservoirs sampled in the dataset. The Bernoulli statistical method handles the data in two separate sets: cases and controls. By defining as cases the isolates from the serovar chosen for analysis, and as controls the rest of the isolates collected in the same year, the bias on the representativeness of the data is minimized.
The choice of serovars to be analysed was based on the most frequent serovars isolated in humans per year. For years 2002–2005 (results not shown) the most common serovars found in humans were also among those most common in the non-human category. However, in 2006 for S. Cholerasuis, and in 2007 for S. Cholerasuis and S. Enteritidis, there were not enough isolates to perform the scan, even though these serovars were among the most frequent cases in humans. This could be due to changes in the reservoirs sampled (e.g. not collecting samples from eggs in 2007 which would be the most common reservoir for S. Enteritidis), or by true decreases on prevalences conjugated for instance with increases in the consumption of imported foods that were not sampled.
Statistical scan methods, such as those presented here, have been frequently used to identify clusters of disease, both infectious and chronic [8, 10, 11, 17, 20, 25–27].
The results showed that the methods performed well in detecting significant clusters and handling data limitations. Both the space–time permutation model and the Bernoulli model could detect not only the most likely cluster but also detected two or three significant secondary clusters for many of the serovars. This suggests that the models are sensitive enough to detect all possible significant clusters according to the defined settings of likelihood. However, the scanning window used was limited to an elliptical cylinder shape (even if with various sizes), which could result in some clusters not being detected when scanning large geographical regions like Thailand [8].
Data on surveillance of the most common reservoirs for Salmonella can provide useful insight for the prevention or early detection of human outbreaks and should be handled together with clinical data. In this analysis, we evaluated cases where there was a geographical and temporal adjacency or overlap between human and non-human clusters for the same serovar.
In the cases where an overlap existed, the information from the non-human clusters could help in tracking the source of the outbreak. Furthermore, for cases when a non-human cluster occurred before the human cluster or in its bordering regions, the information could be used to prevent human outbreaks from occurring. This type of associations and information will be more reliable and significant as the data quality improves.
The associations detected were especially pronounced for S. Rissen. This serovar has often been linked with human infections through the consumption of pork products [23], which are very popular in Thai cuisine. More details on the source of the S. Rissen isolates could confirm the association detected.
Regarding other possible bias in the data collection, the decrease in the number of isolates throughout the study period could be explained by a real decrease in the number of cases due to surveillance changes and improvements, but it should also reflect changes in the data collection process. Demographic differences should also be taken into account, e.g. the fact that Bangkok and its bordering regions are much more populated than the northern part of Thailand, or that access to medical facilities is more difficult in rural areas than in the cities. Similarly, the main economic activity of each region influences the number of submitted isolates, for instance coastal areas have more tourism and during seasonal peaks population numbers can increase markedly.
The amount and distribution of available data forced the analysis to be run aggregating the isolates from non-human sources in one generic group. A more complete and systematic data collection would allow the analysis to be run accounting for specific source (food, animal or even more detailed as food type or animal species) and resulting in more definitive conclusions about possible associations detected. Nevertheless, these types of data restrictions reflect the reality for most countries, where a systematic and integrated surveillance system is not yet in place. It is important to develop methods that can work within these limitations and still have a reasonable predictive ability.
Further epidemiological investigations are needed to determine whether clusters represent real outbreaks or if they are a result of a temporary increase in the prevalence of endemic strains. Still, a significant cluster shows that some changes have occurred either in reservoirs or in the human population and this may require appropriate action by the relevant authorities. The observed temporal association between clusters appearing in non-human reservoirs prior to cases in humans could be used as warnings for the authorities and allow preventive actions to be taken before cases occur in humans.
The statistical analysis was done retrospectively using data from 2002 to 2007, but it still provides important indications on how the tools work and how limitations, such as the low amount of data collected and the fact that it is geographically unevenly dispersed over a large territory like Thailand, are handled. This can be useful for future adaptations of the methods to work in real-time and function as an outbreak detection tool.
In this study, spatio-temporal scan statistics proved to be an efficient and user-friendly platform for running a retrospective cluster analysis. The SatScan software comprises different statistical methods that can be adequate for different types of data and limitations as shown in this study. The use of approaches like the one presented here could provide methodological support to contribute to the implementation of efficient strategies to control and prevent Salmonella infections.
ACKNOWLEDGEMENTS
This study was supported by the Center for Genomic Epidemiology (09-067103/DSF) (www.genomicepidemiology.org).
DECLARATION OF INTEREST
None.
REFERENCES
- 1.Domingues AR, et al. Source attribution of human salmonellosis using a meta-analysis of case-control studies of sporadic infections. Epidemiology and Infection 2012; 140: 959–969. [DOI] [PubMed] [Google Scholar]
- 2.Pires SM, Hald T. Assessing the differences in public health impact of Salmonella subtypes using a bayesian microbial subtyping approach for source attribution. Foodborne Pathogens and Disease 2010; 7: 143–151. [DOI] [PubMed] [Google Scholar]
- 3.Aarestrup FM, et al. Antimicrobial susceptibility and occurrence of resistance genes among Salmonella enterica serovar Weltevreden from different countries. Journal of Antimicrobial Chemotherapy 2003; 52: 715–718. [DOI] [PubMed] [Google Scholar]
- 4.Hendriksen RS, et al. Antimicrobial resistance and molecular epidemiology of Salmonella Rissen from animals, food products, and patients in Thailand and Denmark. Foodborne Pathogens and Disease 2008; 5: 605–619. [DOI] [PubMed] [Google Scholar]
- 5.Herikstad H, Motarjemi Y, Tauxe RV. Salmonella surveillance: a global survey of public health serotyping. Epidemiology and Infection 2002; 129: 1–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Reintjes R, Baumeister HG, Coulombier D. Infectious disease surveillance in North Rhine-Westphalia: first steps in the development of an early warning system. International Journal of Hygiene and Environmental Health 2001; 203: 195–199. [DOI] [PubMed] [Google Scholar]
- 7.Zankari E, et al. Identification of acquired antimicrobial resistance genes. Journal of Antimicrobial Chemotherapy 2012; 67, 2640–2644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kulldorff M, et al. A space-time permutation scan statistic for disease outbreak detection. PLoS Medicine 2005; 2: e59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Vieira VM, et al. Spatial-temporal analysis of breast cancer in upper Cape Cod, Massachusetts. International Journal of Health Geographics 2008; 7: 46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hyder K, et al. Use of spatiotemporal analysis of laboratory submission data to identify potential outbreaks of new or emerging diseases in cattle in Great Britain. BMC Veterinary Research 2011; 7: 14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Alkhamis MA, et al. Temporospatial clustering of foot-and-mouth disease outbreaks in Israel and Palestine, 2006–2007. Transboundary and Emerging Diseases 2009; 56: 99–107. [DOI] [PubMed] [Google Scholar]
- 12.Bangtrakulnonth A, et al. Salmonella serovars from humans and other sources in Thailand, 1993–2002. Emerging Infectious Diseases 2004; 10: 131–136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kulldorff M. A spatial scan statistic. Communications in Statistics – Theory and Methods 1997; 26: 1481–1496. [Google Scholar]
- 14.Madin B. An evaluation of foot-and-mouth disease outbreak reporting in mainland South-East Asia from 2000 to 2010. Preventive Veterinary Medicine 2011; 102: 230–241. [DOI] [PubMed] [Google Scholar]
- 15.Bangtrakulnonth A, et al. Shigella from humans in Thailand during 1993 to 2006: spatial-time trends in species and serotype distribution. Foodborne Pathogens and Disease 2008; 5: 773–784. [DOI] [PubMed] [Google Scholar]
- 16.Kulldorff M. SaTScan User Guide v. 9.0, 2010.
- 17.Warden CR. Comparison of Poisson and Bernoulli spatial cluster analyses of pediatric injuries in a fire district. International Journal of Health Geographics 2008; 7: 51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Bangtrakulnonth A, Tishyadhigama P. Annual Report of Confirmed Salmonella and Shigella in Thailand 2006, 2007.
- 19.National Statistical Office, Thailand (http://web.nso.go.th/).
- 20.Stelling J, et al. Automated use of WHONET and SaTScan to detect outbreaks of Shigella spp. using antimicrobial resistance phenotypes. Epidemiology and Infection 2010; 138: 873–883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.European Food Safety Authority. The community summary report on trends and sources of zoonoses, zoonotic agents and foodborne outbreaks in the European union in 2008. EFSA Journal 2010; 8: 1496. [Google Scholar]
- 22.King N, Lake R, Campbell D. Source attribution of nontyphoid salmonellosis in new zealand using outbreak surveillance data. Journal of Food Protection 2011; 74: 438–445. [DOI] [PubMed] [Google Scholar]
- 23.Hendriksen RS, et al. Risk factors and epidemiology of the ten most common Salmonella serovars from patients in Thailand: 2002–2007. Foodborne Pathogens and Disease 2009; 6: 1009–1019. [DOI] [PubMed] [Google Scholar]
- 24.Hendriksen RS, et al. Characterization of Salmonella enterica serovar Enteritidis isolates recovered from blood and stool specimens in Thailand. BMC Microbiology 2012; 12: 92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Coleman M, et al. Using the SaTScan method to detect local malaria clusters for guiding malaria control programmes. Malaria Journal 2009; 8: 68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mostashari F, et al. Dead bird clusters as an early warning system for West Nile virus activity. Emerging Infectious Diseases 2003; 9: 641–646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Abe T, Martin IB, Roche LM. Clusters of census tracts with high proportions of men with distant-stage prostate cancer incidence in New Jersey, 1995 to 1999. American Journal of Preventive Medicine 2006; 30: S60–66. [DOI] [PubMed] [Google Scholar]