SUMMARY
Trypanosoma cruzi I, a discrete typing unit (DTU) found in human infections in Venezuela and other countries of the northern region of South America and in Central America, has been recently classified into five intra-DTU genotypes (Ia, Ib, Ic, Id, Ie) based on sequence polymorphisms found in the spliced leader intergenic region. In this paper we report the genotype identification of T. cruzi human isolates from one outbreak of acute orally acquired Chagas disease that occurred in a non-endemic region of Venezuela and from T. cruzi triatomine and rat isolates captured at a guava juice preparation site which was identified as the presumptive source of infection. The genotyping of all these isolates as TcId supports the view of a common source of infection in this oral Chagas disease outbreak through the ingestion of guava juice. Implications for clinical manifestations and dynamics of transmission cycles are discussed.
Key words: Acute Chagas disease outbreak, DTU genotypes, oral transmission, Trypanosoma cruzi
INTRODUCTION
Chagas disease (CD) is a parasitic illness with an estimated prevalence of eight million infected people in Latin America, although the information currently available does not reflect the true extent of the problem [1]. Trypanosoma cruzi, its aetiological agent, is mainly transmitted to humans by different Reduviidae insect vectors or by blood transfusion. Congenital transmission from chronically infected women occurs in 2–8% of pregnancies depending on the endemic area [2, 3]. New epidemiological modes of infection have been detected such as oral transmission and domestic adaptation of wild triatomines. The oral route of transmission is well documented and constitutes a mode of infection that leads to microepidemics and outbreaks of a more severe clinical form of acute CD [4–8]. The latter via transmission is currently acquiring relevance due to its association with a high mortality rate. The severity of CD also depends on the genetic diversity of T. cruzi and the competence of the host immune system.
In Venezuela, a significant outbreak of acute CD occurred in December 2007 affecting 103 persons at a school in an urban neighbourhood of Caracas where vector transmission had not been reported previously [9]. The infection was associated with Panstrongylus geniculatus, a sylvatic triatomine vector that is in a progressive process of domiciliation [10–12] and which had been considered as an inefficient T. cruzi transmission vector. The outbreak was suspected of being linked to the ingestion of contaminated guava juice prepared under unsupervised hygienic conditions by a supplier in a distant slum on the western mountain slopes of the city, where triatomines were present. A significant positive correlation was found between ingestion of guava juice and risk of infection [9].
T. cruzi shows a genetic diversity that has been associated with the observed variability of clinical manifestations, geographical distribution and preferential parasite–vector interactions. To characterize the population structure, T. cruzi has been subdivided into six discrete typing units (DTUs), namely TcI, TcII, TcIII, TcIV, TcV and TcVI [13]. T. cruzi I (TcI) isolates are predominant in Venezuela and other countries of the northern region of South America and in Central America [14–17]. TcI has been mainly associated with low parasitaemia level in peripheral blood and very mild clinical symptoms. However, many symptomatic cases of severe acute CD patients infected with TcI in endemic regions of Venezuela and Colombia have also been described [14, 18, 19]. Five intra-DTU TcI genotypes (Ia, Ib, Ic, Id, Ie) have been described based on sequence polymorphisms found in the mini-exon gene's intergenic region [20–22]. So far, strains associated with documented cases of oral transmission have been characterized as TcII [23] and TcI [24, 25]. In this context we decided to determine the intra-DTU TcI genotypes of T. cruzi isolates from the orally transmitted outbreak described above which also allowed us to understand the current circulating infecting strains. Preliminary results revealed a marked genetic homogeneity, with all of the isolates belonging to the TcI lineage.
METHODS
T. cruzi isolates
The parasite isolates were obtained by haemoculture from peripheral blood of three patients orally infected in an urban outbreak at a school in Caracas, Venezuela, in December 2007 (including the index case); from the faeces of one infected P. geniculatus triatomine, and from the blood of a rat captured at the guava juice preparation site which was identified as the source of infection [9]. In addition, a cultured human stock from B. Alarcón de Noya's laboratory, isolate PM [26], as well as Munanta (DTU TcI), Ikiakarora (DTU TcIII) and Guateque (DTU TcVI) strains from M. C. López's laboratory [27] were included (Table 1).
Table 1.
Description of Trypanosoma cruzi isolates
| Isolate | Geographical origin | Host | Route of infection | DTU |
|---|---|---|---|---|
| MHOM/VE/2007/1593 | Chacao, Caracas | Human | Oral | Id |
| MHOM/VE/2007/1595 | Chacao, Caracas | Human | Oral | Id |
| MHOM/VE/2007/1598 | Chacao, Caracas | Human | Oral | Id |
| MHOM/VE/1955/PM | Valencia | Human | Vector | Ia/Ib |
| TGEN/VE/2008/AC1 | Altagracia, Caracas | Panstrongylus geniculatus | Id | |
| MRAT/VE/2008/RATA6 | Altagracia, Caracas | Rattus rattus | Id |
DTU, Discrete typing unit.
DNA extraction and PCR amplifications
Total DNA extraction from samples was performed by resuspension of parasite pellets in lysis solution (150 mm NaCl, 0·1 m EDTA, 0·1 mg Proteinase K, 0·4% SDS) and incubated for 2 h at 50°C, followed by phenol-chloroform extractions and RNAse treatment. DNA pellets were resuspended in distilled water and stored at −20°C. Amplification of spliced leader intergenic region (SL-IR) was performed in a multiplex reaction with primers TC (CCCCCCTCCCAGGCCACACTG), TC1 (GTGTCCGCCACCTCCTTCGGGCC) and TC2 (CCTGCAGGCACACGTGTGTGTG) [28, 29]. PCR was performed in a final volume of 25 μl containing 100 ng template DNA, 100 pmol of each primer, 2·5 mm of each dNTP, 2·5 U Taq polymerase, 2·5 μl of 10× Taq polymerase buffer, and 1·5 mm MgCl2. Amplification was performed in a MJR PTC-100 thermocycler (MJ Research, USA) as follows: one step of 1 min denaturation at 94°C, 27 cycles at 94 °C for 30 s, 55°C for 30 s, 72°C for 30 s and a final extension step at 72°C for 5 min. Ribosomal subunit amplification with primers D71 (AAGGTGCGTCGACAGTGTGG) and D72 (TTTTCAGAATGGCCAACAGT) were also performed [28] in a final volume of 25 μl containing 100 ng template DNA, 100 pmol of each primer, 2·5 mm of each dNTP, 2·5 U Taq polymerase, 2·5 μl of 10× Taq polymerase buffer and 1·5 mm MgCl2. The PCR products were analysed by 1·5% agarose gel electrophoresis for SL-IR amplification and by 12% acrylamide gel electrophoresis for 24Sα amplification. In both cases the DNA was detected by ethidium bromide staining and visualized under ultraviolet light. PCR products of 350 bp and 300 bp for SL-IR amplification were expected for TcI and for TcII, TcV and TcVI, respectively, while no bands were expected for TcIII and TcIV [30]. A product of 110 bp for 24Sα rDNA amplification was expected for TcI, TcIII and TcV, and products of 120 bp and 125 bp, for TcIV and for TcII and TcVI, respectively.
The amplification products for SL-IR were directly sequenced using the dideoxy chain termination method, and also purified from agarose gels using the Qualex kit (Qiagen, Germany) and cloned in p-GEM T vector (Promega, USA), using the blue/white selection with IPTG/X-gal for the identification of transformants. DNA from each one of the three clones was isolated with the SV Minipreps DNA Purification System (Promega) and the two chains sequenced using the dideoxy chain termination method.
DNA sequence and molecular phylogenetic analysis
Multiple sequence alignments were performed using ClustalW2 [31] and edited in BioEdit v. 7.0.9 (Ibis Biosciences, USA).
RESULTS
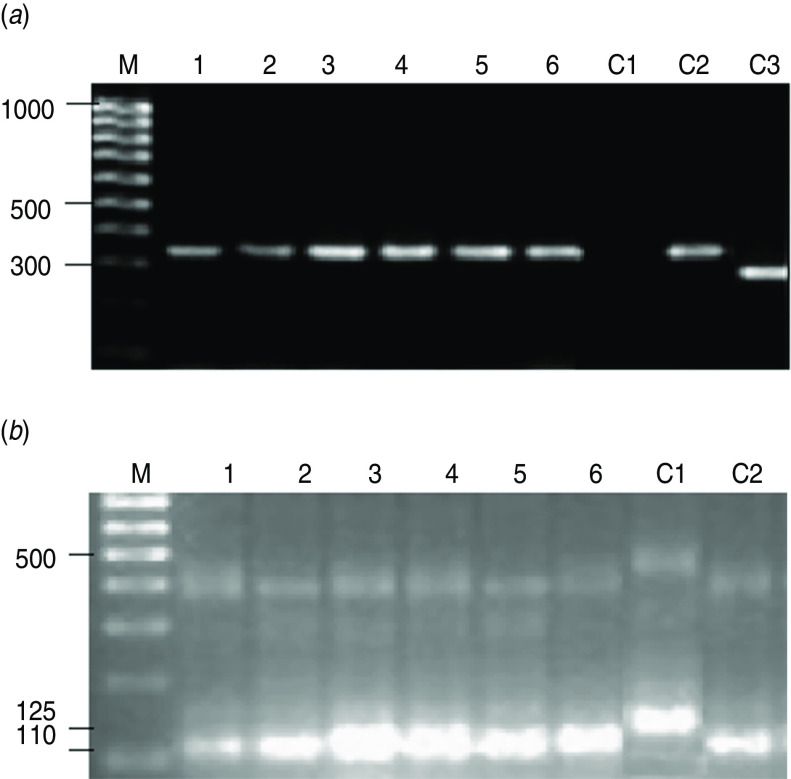
In order to characterize the infecting strain that caused the orally acquired CD acute infection, parasites were isolated from three patients from the Caracas outbreak and also from a triatomine and a rat captured at the guava juice preparation site (identified as the source of infection), grown and genotyped. Thus, genomic DNA (gDNA) from the different isolates was purified and PCR performed. As PCR controls, gDNA purified from known T. cruzi strains such as Munanta (DTU TcI), Ikiakarora (DTU TcIII) and Guateque (DTU TcVI) were employed as DNA template. As observed in Figure 1a, amplification of SL-IR in a multiplex reaction using the TC, TC1 and TC2 primers generated a 350 bp product when the T. cruzi gDNAs isolated from the three patients from Caracas (lanes 1–3), and those isolated from the triatomine (lane 5), the rat (lane 6) and control DTU TcI (lane C2) were employed. A single amplification band of 300 bp was generated when DTU TcVI gDNA (lane C3) was employed as DNA template and none when DTU TcIII gDNA (lane C1) was used. These data indicated that all the analysed parasite strains belonged to DTU TcI. This typing was corroborated by the PCR performed with specific primers for 24S amplification which produced amplified bands with the expected size for DTU TcI strains (Fig. 1b).
Fig. 1.
Gel electrophoresis of PCR products from T. cruzi isolates. (a) 1·5% Agarose gel electrophoresis of SL-IR amplicons. (b) 12% Acrylamide gel electrophoresis of 24Sα amplicons. Lane M, 100 bp DNA ladder (hyperladder IV, Bioline, UK); lanes 1–3, isolates 1593, 1595 and 1598 from patients infected in the Caracas outbreak; lane 4, laboratory strain PM; lane 5, isolate AC1 from an infected Panstrongylus geniculatus triatomine captured at the guava juice preparation site; lane 6, isolate RATA 6 from an infected rat captured at the guava juice preparation site; lanes C1, C2, C3, T. cruzi control strains Ikiakora (DTU TcIII), Munanta (DTU TcI) and Guateque (DTU TcVI).
The SL-IR amplified products were directly sequenced but also cloned into pGEM-T Easy Vector (Promega), in order to reveal the possible existence of mixed TcI SL-IR genotypes. The DNA from different clones was purified and subsequently sequenced. Sequence alignment of the different PCR products (whose electropherograms exhibited no nucleotide ambiguity) and clones corresponding to the isolated parasites in all the patients from Caracas were seen to be identical, indicating that they were all infected by a T. cruzi population belonging to a single TcI genotype. The sequences of the different PCR products and clones corresponding to the amplicons generated with the DNA of the parasites isolated from the triatomine and the rat were also identical to those obtained from the parasites isolated from the patients.
Following the intra-DTU TcI classification first proposed by Herrera et al. [20] and expanded by Cura et al. [22], when the first 60 bp of the SL-IR were aligned with reference sequences, it was observed that all the isolates from the Caracas outbreak presented a gap of 8 nt at position 20–27, as well as other single nucleotide polymorphisms (SNPs), considered as hallmarks of the DTU TcId (Fig. 2). These observations indicated that the triatomine and rat were also infected by a T. cruzi population belonging to the same TcI genotype.
Fig. 2.

Multiple alignment of 60-nt region within the SL-IR of the T. cruzi I sequences analysed. Partial reference sequences reported in [20, 22] are shown in bold: CGC (AM259467), JLC (AM259468), X380C (AM259472), JD1C (AM259478) and PALV2-2 Cl5 (GQ398812.1). Gaps (∼), conserved sites (.).
On the other hand, a SL-IR sequence divergence was observed in clones corresponding to the laboratory PM strain which was isolated from a patient with vector-acquired CD [26]. These sequences did not have the characteristic 8-nt gap of DTU TcId. Instead they bore the SNP characteristic of the SL-IR sequences from parasites belonging to DTUs TcIa and TcIb (Fig. 2) which was taken as an indication that the laboratory strain was a mixed population of TcIa and TcIb genotypes.
DISCUSSION
In this study we have demonstrated that three patients (including the index case) from an important outbreak of acute CD occurring at a school in an urban neighbourhood of Caracas, and a triatomine and a rat captured at the guava juice preparation site identified as the source of the parasite contamination, were all infected by a T. cruzi population belonging to the same TcI genotype. These observations strongly suggest that a single strain was implicated in this outbreak and reinforce the fact that the guava juice preparation site was the source of the infection, identifying the oral route of transmission as a common mechanism of infection [9]. On the other hand, our results indicate that the T. cruzi isolate PM, obtained from a patient with vector-acquired CD [26], was a mixed population of TcIa and TcIb genotypes. Recently, natural mixed populations of TcIa and TcId have been reported in faecal samples of Triatoma infestans from Paraguay [22] and in heart, blood and skin chagoma samples from an Argentinean patient [32].
According to current molecular evidence, TcI is one of the parental DTUs in the hybridization events that, separated by periods of clonal evolution, explain the genetic diversity of the extant T. cruzi populations. In Venezuela and Colombia, TcI isolates have been associated not only with low parasitaemia and very mild clinical symptoms, but also with severe acute symptomatic cases and death [14, 19, 20]. TcI has a notable intra-DTU diversity, recently classified into five genotypes (Ia, Ib, Ic, Id, Ie) [20–22], although it could not be definitely associated with the diversity of TcI biological characteristics observed, i.e. transmission cycles, host preferences, clinical manifestations or geographical distribution [33, 34]. In Colombia, an association was found for TcId with sylvatic populations of Rhodnius prolixus and wild reservoirs [21]. We here report its association with sylvatic populations of P. geniculatus. The close contact with infected sylvatic species might explain oral outbreaks in areas without a history of vector transmission. In fact, Caracas is a densely populated urban area currently considered non-endemic for CD, since active vector transmission had not been reported previously. However, a local sylvatic triatomine vector, P. geniculatus, is present in the surrounding mountain area where the nocturnal incursion of sylvatic infected triatomine bugs has allowed the opportunity of easy contamination of unprotected food and beverages with T. cruzi [9].
A multilocus microsatellite typing of T. cruzi DTU I [16] showed a sylvatic TcI genotype predominance in the Venezuelan adult intradomiciliary triatomine sampled, including Triatoma maculata, P. geniculatus, and R. prolixus. The TcI sylvatic genotype was different from the domestic one observed in TcI strains from infected humans sampled across the country. In Colombia, TcId has also been mainly associated with sylvatic triatomines and reservoir populations suggesting, in both countries, a possible and progressive introduction of a sylvatic sub-DTU I in the domestic environment, independent of the transmission route of the parasite or the species of the triatomine vector.
In a broader study TcId was also found in a variety of peridomestic, domestic and sylvatic triatomines, as well as in wild reservoirs and in human samples from South America [22]. These data are also consistent with the epidemiological changes that are taking place with regard to CD transmission since P. geniculatus had been considered as an inefficient T. cruzi transmission vector.
A preferential cardiac tropism for TcId has been suggested [32]. This tropism was consistent with the clinical manifestations of patients from the Caracas outbreak, where in 61 (59%) of the 103 confirmed cases, one or more abnormalities were observed on ECG recordings, 75% were symptomatic, 20·3% required hospitalization and a child aged 5 years died of acute chagasic myocarditis [9].
This report shows that T. cruzi isolates from sylvatic vectors and reservoirs, as well as isolates from humans infected in an orally acquired CD outbreak in a non-endemic region of Venezuela belong to a same T. cruzi I genotype, TcId. The similarity in the parasite isolates from three patients of the urban oral outbreak and the triatomine and the rat captured at the guava juice preparation site, allows us to suggest that the source of infection was common for these persons and the vector.
Although further analysis with more isolates is needed, this work contributes to the knowledge and possible association between the oral route of transmission, the severity of the disease and the genotypes of T. cruzi isolates, providing evidence that DTU TcId can successfully infect humans by the oral route.
ACKNOWLEDGEMENTS
This work received financial support from Sección de Inmunología, Instituto de Medicina Tropical, Universidad Central de Venezuela; Fondo Nacional de Ciencia y Tecnología (travelling expenses of Z.D. for a research cooperation with Spain, grant no. G-2005000387), (T.A., grant no. 2006000216); Plan Nacional I+D+i, MINECO, Spain (M.C.T. and M.C.L., grant nos. BFU2010-1670 and SAF2012-35777); FEDER (M.C.T. and M.C.L). M. C. Thomas, M. C. López, O. Noya and B. Alarcón de Noya are members of the NHEPACHA network (Latin American Network for Chagas disease) and PAICET network (Tri-Continental Altantic Platform for Research and Control of Tropical Diseases).
DECLARATION OF INTEREST
None.
REFERENCES
- 1.World Health Organization. Report of the scientific working group on Chagas disease. 2007. (http://whqlibdoc.who.int/hq/2007/TDR_SWG_09_spa.pdf). Accessed June 2012.
- 2.Carlier Y, Torrico F. Congenital infection with Trypanosoma cruzi from mechanism of transmisión to strategies for diagnosis and control. Revista da Sociedade Brasileria de Medicina Tropical 2003; 36: 767–771. [DOI] [PubMed] [Google Scholar]
- 3.World Health Organization. Control of Chagas disease. Second report of the WHO Expert Committee. World Health Organization, Technical Report Series 2002; 905: 59–90. [PubMed] [Google Scholar]
- 4.Coura JR, et al. Emerging Chagas' disease in Amazonian Brazil. Trends in Parasitology 2002; 18: 171–176. [DOI] [PubMed] [Google Scholar]
- 5.Nóbrega AA, et al. Oral transmission of Chagas disease by consumption of açaí palm fruit, Brazil. Emerging Infectious Diseases 2009; 15: 653–655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ríos JF, et al. Probable outbreak of oral transmission of Chagas disease in Turbo, Antioquia. Biomedica 2011; 31: 185–195. [DOI] [PubMed] [Google Scholar]
- 7.Santalla Vargas J, et al. First reported outbreak of Chagas disease in the Bolivian Amazonia: report of 14 acute cases by oral transmission of Trypanosoma cruzi in Guayaramerín, Beni-Bolivia. Biofarbo 2011; 19: 52–58. [Google Scholar]
- 8.Shikanai-Yasuda MA, Carvalho NB. Oral transmission of Chagas disease. Clinical Infectious Diseases 2012; 54: 845–852. [DOI] [PubMed] [Google Scholar]
- 9.Alarcón de Noya B, et al. Large urban outbreak of orally acquired acute Chagas disease at a school in Caracas, Venezuela. Journal of Infectious Diseases 2010; 201: 1308–1315. [DOI] [PubMed] [Google Scholar]
- 10.Reyes-Lugo M. Panstrongylus geniculatus Latreille 1811 (Hemiptera: Reduviidae: Triatominae), vector of Chagas disease in the home environment of North-Central Venezuela. Revista Biomédica 2009; 20: 180–205. [Google Scholar]
- 11.Feliciangeli MD, et al. Mixed domestic infestation by Rhodnius prolixus Stal, 1859 and Panstrongylus geniculatus Latreille, 1811, vector incrimination, and seroprevalence for Trypanosoma cruzi among inhabitants in El Guamito, Lara State, Venezuela. American Journal of Tropical Medicine and Hygiene 2004; 71: 501–505. [PubMed] [Google Scholar]
- 12.Carrasco HJ, et al. Risk of Trypanosoma cruzi I (Kinetoplastida: Trypanosomatidae) transmission by Panstrongylus geniculatus (Hemiptera: Reduviidae) in Caracas (Metropolitan District) and neighboring States, Venezuela. International Journal for Parasitology 2005; 35: 1379–1384. [DOI] [PubMed] [Google Scholar]
- 13.Zingales B, et al. A new consensus for Trypanosoma cruzi intraspecific nomenclature: second revision meeting recommends TcI to TcVI. Memorias do Instituto Oswaldo Cruz 2009; 104: 1051–1054. [DOI] [PubMed] [Google Scholar]
- 14.Añez N, et al. Predominance of lineage I among Trypanosoma cruzi isolates from Venezuelan patients with different clinical profiles of acute Chagas Disease. Tropical Medicine and International Health 2004; 9: 1319–1326. [DOI] [PubMed] [Google Scholar]
- 15.Samudio F, et al. Predominance of Trypanosoma cruzi I among Panamanian sylvatic isolates. Acta Tropica 2007; 101: 178–181. [DOI] [PubMed] [Google Scholar]
- 16.Llewellyn MS, et al. Genome-scale multilocus microsatellite typing of Trypanosoma cruzi discrete typing unit I reveals phylogeographic structure and specific genotypes linked to human infection. PLoS Pathogens 2009; 5: e1000410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Carrasco HJ, et al. Geographical distribution of Trypanosoma cruzi genotypes in Venezuela. PLoS Neglected Tropical Diseases 2012; 6: e1707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Triana O, et al. Trypanosoma cruzi: variability of stocks from Colombia determined by molecular karyotype and minicircle Southern blot analysis. Experimental Parasitology 2006; 113: 62–66. [DOI] [PubMed] [Google Scholar]
- 19.Salazar A, Schijman AG, Triana O. High variability of Colombian Trypanosoma cruzi lineage I stocks as revealed by low-stringency single primer-PCR minicircle signatures. Acta Tropica 2006; 100: 110–118. [DOI] [PubMed] [Google Scholar]
- 20.Herrera C, et al. Identifying four Trypanosoma cruzi I isolate haplotypes from different geographic regions in Colombia. Infection, Genetics and Evolution 2007; 7: 535–539. [DOI] [PubMed] [Google Scholar]
- 21.Falla A, Herrera C, et al. Haplotype identification within Trypanosoma cruzi I in Colombian isolates from several reservoirs, vectors and humans. Acta Tropica 2009; 110: 15–21. [DOI] [PubMed] [Google Scholar]
- 22.Cura CI, et al. Trypanosoma cruzi I genotypes in different geographical regions and transmission cycles based on a microsatellite motif of the intergenic spacer of spliced-leader genes. International Journal for Parasitology 2010; 40: 1599–1607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Steindel M, et al. Characterization of Trypanosoma cruzi isolated from humans, vectors, and animal reservoirs following an outbreak of acute human Chagas disease in Santa Catarina State, Brazil. Diagnostic Microbiology and Infectious Disease 2008; 60: 25–32. [DOI] [PubMed] [Google Scholar]
- 24.Marcili A, et al. Trypanosoma cruzi in Brazilian Amazonia: Lineages TCI and TCIIa in wild primates, Rhodnius spp. and in humans with Chagas disease associated with oral transmission. International Journal for Parasitology 2009; 39: 615–623. [DOI] [PubMed] [Google Scholar]
- 25.Andrade SG, et al. Biological, biochemical and molecular features of Trypanosoma cruzi strains isolated from patients infected through oral transmission during a 2005 outbreak in the state of Santa Catarina, Brazil: its correspondence with the new T. cruzi Taxonomy Consensus (2009). Memorias do Instituto Oswaldo Cruz 2011; 106: 948–956. [DOI] [PubMed] [Google Scholar]
- 26.Maekelt GA. The complement fixation test for Chagas disease. Zeitschrift Tropenmedizin und Parasitologie 1960; 11: 155–166. [PubMed] [Google Scholar]
- 27.Thomas MC, et al. Plasticity of the histone H2A genes in a Brazilian and six Colombian strains of Trypanosoma cruzi. Acta Tropica 2000; 25: 203–210. [DOI] [PubMed] [Google Scholar]
- 28.Souto RP, et al. DNA markers define two major phylogenetic lineages of Trypanosoma cruzi. Molecular and Biochemical Parasitology 1996; 83: 141–152. [DOI] [PubMed] [Google Scholar]
- 29.Fernandes O, et al. Brazilian isolates of Trypanosoma cruzi from humans and triatomines classified into two lineages using mini-exon and ribosomal RNA sequences. American Journal of Tropical Medicine and Hygiene 1998; 58: 807–811. [DOI] [PubMed] [Google Scholar]
- 30.Zingales B, et al. The revised Trypanosoma cruzi subspecific nomenclature: Rationale, epidemiological relevance and research applications. Infection, Genetics and Evolution 2012; 12: 240–253. [DOI] [PubMed] [Google Scholar]
- 31.Goujon M, et al. A new bioinformatics analysis tools framework at EMBL-EBI (2010). Nucleic Acids Research 2010; 38 (Suppl.): W695–699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Burgos JM, et al. Molecular identification of Trypanosoma cruzi discrete typing units in end-stage chronic Chagas heart disease and reactivation after heart transplantation. Clinical Infectious Diseases 2010; 51: 485–495. [DOI] [PubMed] [Google Scholar]
- 33.O'Connor O, Bosseno MF, et al. Genetic clustering of Trypanosoma cruzi I lineage evidenced by intergenic miniexon gene sequencing. Infection, Genetics and Evolution 2007; 7: 587–593. [DOI] [PubMed] [Google Scholar]
- 34.Mejía-Jaramillo AM, et al. Geographical clustering of Trypanosoma cruzi I groups from Colombia revealed by low-stringency single specific primer-PCR of the intergenic regions of spliced-leader gene. Parasitology Research 2009; 104: 399–410. [DOI] [PubMed] [Google Scholar]