Abstract
The ruminal bacterium Butyrivibrio fibrisolvens is being engineered by the introduction of heterologous xylanase genes in an attempt to improve the utilization of plant material in ruminants. However, relatively little is known about the diversity and distribution of the native xylanase genes in strains of B. fibrisolvens. In order to identify the most appropriate hosts for such modifications, the xylanase genotypes of 28 strains from the three 16S ribosomal DNA (rDNA) subgroups of Butyrivibrio fibrisolvens have been investigated. Only 4 of the 20 strains from 16S rDNA group 2 contained homologues of the strain Bu49 xynA gene. However, these four xynA-containing strains, and two other group 2 strains, contained members of a second xylanase gene family clearly related to xynA (subfamily I). Homologues of xynB, a second previously described xylanase gene from B. fibrisolvens, were identified only in three of the seven group 1 strains and not in the group 2 and 3 strains. However, six of the group 1 strains contained one or more members of the two subfamilies of homologues of xynA. The distribution of genes and the nucleotide sequence relationships between the members of the two xynA subfamilies are consistent with the progenitor of all strains of B. fibrisolvens having contained a xynA subfamily I gene. Since many xylanolytic strains of B. fibrisolvens did not contain members of either of the xynA subfamilies or of the xynB family, at least one additional xylanase gene family remains to be identified in B. fibrisolvens.
Strains of Butyrivibrio fibrisolvens and Butyrivibrio-like organisms may frequently constitute a significant proportion of the organisms in the rumens of many species of ruminants (3, 23). Most strains of B. fibrisolvens are xylanolytic (4), while a smaller number have significant fibrolytic capabilities. For these reasons strains of B. fibrisolvens have been targeted for transformation with genes to improve their ability to degrade plant fiber prior to reintroduction into the rumen (5). Although two recently described approaches involved the introduction of a bacterial xylanase (15) and a modified fungal xylanase (8, 35), little work has been undertaken to characterize the repertoire of endogenous xylanase-encoding genes in diverse strains of B. fibrisolvens. An understanding of the evolution of the native xylan-degrading systems and the natures of the genes contained in the target strains for modification is required for the most effective construction of recombinant xylanolytic strains of B. fibrisolvens.
The xylanase and associated activities of B. fibrisolvens have been characterized in greatest detail from the type strain, D1 (12, 13), and from strains Nor37 (32–34) and H17c (11–13, 18). However, xylanase-encoding genes have been characterized only from two B. fibrisolvens strains, H17c (19) and Bu49 (21). The genes were designated xynA from Bu49 and xynB from H17c. In hybridization studies using a range of strains of B. fibrisolvens, homologues of the Bu49 gene, xynA, were detected in H17c and CF3 but not in Nor37 and D1 (21). However, a weakly hybridizing band was detected under low-stringency conditions with D1 and a number of other strains and also as an additional band in Bu49, H17c, and CF3. It was suggested that this band might represent an additional homologue of xynA (21).
Analysis of the xylanases expressed by B. fibrisolvens Bu49 and H17c suggested that the two strains produce quite different complements of enzymes. Two enzymes with xylanase activities were detected in Bu49, XynA (40 to 45 kDa with a neutral-to-alkaline pI) and a second, smaller, 12- to 16-kDa enzyme with a more acid pI (12, 21). In contrast, H17c exhibited a much more complex pattern and appeared to synthesize as many as 11 different enzymes with xylanase activities (18). The difference in xylanase patterns is somewhat surprising, as Bu49 and H17c are thought to be very similar strains of B. fibrisolvens (20).
Here we present an analysis of the distribution and evolution of homologues of xynA and xynB in strains in the three different groups of B. fibrisolvens. The results of the analysis highlight the diversity of xylanase genotypes in B. fibrisolvens and also identify a case of mistaken identity with respect to H17c.
MATERIALS AND METHODS
Bacterial strains and culture conditions.
Isolates and strains of B. fibrisolvens are listed in Table 1. Origins of the strains are as previously described (8, 14, 22). Additionally, Nor37 was obtained from C. Orpin and H17c(SA) was obtained from J. A. Thomson. B. fibrisolvens was maintained in rumen fluid medium containing 0.5% (wt/vol) cellobiose (3). The cellobiose was replaced with 0.5% (wt/vol) birch wood xylan (BWX) to determine the organism’s xylanolytic phenotype. Stocks were characterized by randomly amplified polymorphic DNA (RAPD) PCR and 16S ribosomal DNA (rDNA) analysis (see below) and repurified if necessary by streaking for single colonies on rumen fluid agar plates. For PCR, aliquots of overnight cultures were stored at −20°C and thawed as required to provide samples of DNA.
TABLE 1.
Xylanase genotypes and phenotypes of strains of B. fibrisolvens
| 16S rDNAa
|
Strain | Gene of xynA subfamilyf
|
Gene of xynB family | Xylanolytic | Reference(s) for xylanase phenotype | ||
|---|---|---|---|---|---|---|---|
| Group | Type | A | I | ||||
| 1 | 1 | D1b | —e (NSB) | xynK (WB) | xynY | + | 13 |
| 1 | 2 | A38c | xynC | xynD + xynH | xynZ | + | 4, 13 |
| LP210B | — | xynG1 | —e | + | This paper | ||
| LP461A | — | xynG2 | — | + | This paper | ||
| AR11 | — | — | + | This paper | |||
| 1 | New | AR10 | — | xynL | — | + | This paper |
| 1 | 6 | H17cSA | xynF | xynE | xynB | + | 18 |
| 2 | 8 | H17c | xynA1 (SB) | xynI1 (WB) | — | + | 13 |
| Bu49 | xynA1 (SB) | xynI1 (WB) | — | + | 13 | ||
| 2 | 11 | CF3 | xynA2 (SB) | xynI2 (WB) | — | + | 13 |
| 2 | P. ruminis | A46 | xynA3 | xynI3 | — | NDg | |
| E14 | — | xynJ1 | — | + | This paper | ||
| VV1 | — | xynJ2 | — | ND | |||
| LP445C | — | — | — | + | This paper | ||
| LP1309 | — | — | — | −h | This paper | ||
| LP1273 | — | — | — | + | This paper | ||
| LP1265 | — | — | — | ND | |||
| 2 | 9 | Nor37d | — (NSB) | — (NWB) | — | + | 32 |
| GS113 | — | — | — | + | 27 | ||
| AR9 | — | — | ND | ||||
| AR11a | — | — | — | + | This paper | ||
| AR16 | — | — | — | + | This paper | ||
| AR73 | — | — | — | + | This paper | ||
| LP1028 | — | — | — | − | This paper | ||
| 2 | New | AR27 | — | — | — | + | This paper |
| 2 | New | LP1266 | — | — | — | + | This paper |
| LP91.4.1 | — | — | — | + | This paper | ||
| 3 | OB156 | — | — | — | −i | 35 | |
ATCC 19171, type strain.
ATCC 27208.
Also known as NCFB2249 and NCDO2249.
No bona fide PCR product detected even under very-low-stringency conditions.
NSB, no strongly hybridizing band with xynA probe (21); SB, strongly hybridizing band with xynA probe (21); WB, weakly hybridizing band with xynA probe (21); NWB, no weakly hybridizing band with xynA probe (21).
ND, not done.
No significant growth on BWX.
We have not observed significant growth on BWX, but xylanase activity has been reported (15).
PCR amplification.
Purified DNA or 0.5-μl samples of stationary-phase cultures of strains of B. fibrisolvens were used as the sources of template DNAs for all PCRs. For RAPD analysis of strains, amplification was by the method of Ruma et al. (24) with the primer pairs 1-08 and 1-20 and 1-05 and 1-13 (Table 2). Samples were analyzed on high-resolution agarose gels, 2% Metaphor agarose (FMC), or 2% SFR agarose (Amresco), and for detailed comparisons, relevant samples were run in adjacent lanes. To amplify 16S rDNA, the PCR method described by Stackebrandt and Liesack (28) with primers 27f and 1492r was used (Table 2). PCR products were digested with restriction enzymes and run on high-resolution agarose gels (see above) alongside standard strains. Xylanase gene homologues were amplified with the PCR primers described in Table 2 under the following conditions: 95°C for 30 s, 46°C for 30 s, and 72°C for 30 s for 35 cycles. The annealing temperature was increased or reduced as required to alter the stringencies of the amplifications.
TABLE 2.
Design of oligonucleotide primers for RAPD and PCR amplification and sequencing of 16S rDNAs and xylanase gene fragments from B. fibrisolvens
| Primer | Sequencec | Location (nucleotides) | Use | Design |
|---|---|---|---|---|
| 1-05 | ACTTGGCGGCCT | RAPD PCR | 24 | |
| 1-08 | CAGGCGAAGGTT | RAPD PCR | 24 | |
| 1-13 | ACGGCTGGTTCC | RAPD PCR | 24 | |
| 1-20 | CAACAGCCCCCA | RAPD PCR | 24 | |
| 27f | GAGTTTGATCCTGGCTCAG | 16S rDNA PCR amplification and sequencing | 16 | |
| 1492r | ACGGCTACCTTGTTACGACTT | 16S rDNA PCR amplification | 16 | |
| 500r | GWATTACCGCGGCKGCTG | 16S rDNA sequencing | 16 | |
| Xyn10f Xyn10r | ATGAGRGGNCAYACNCTNGTNTGG AGRTGNGAYTGCATNCCYTYNCC | 697–720a 499–522b 1114–1136a 898–920b | Generic primers for PCR amplification of genes encoding ghf10 enzymes | Designed with the DNA sequence encoding two regions of highly conserved amino acid sequence in ghf10 (glycosylhydrolase family) enzymes, one of the major families of xylanases (10); sequence biased towards B. fibrisolvens xynA and xynB genes; produces a 440-bp product from xynA subfamily A and I genes and a 422-bp product from xynB family genes |
| XynAf XynAr | ACTATGGACGAACGTCTTGA CCTTTTTACCCTGGTTGAT | 790–809a 1087–1104a | PCR amplification of both subfamilies of xynA | Designed with two regions of DNA sequence conserved in both subfamilies of the xynA gene family but not in genes of the xynB family; sequence designed to have a similar number of mismatches to each of the known xynA subfamily A and I sequences used to derive the primer sequence (Fig. 1); produces a 360-bp product. |
| XynIJf | CGTCTTGAAATGTATGTA | 802–819a | PCR amplification of xynA subfamily I genes | Designed to specifically amplify xynA subfamily I genes in the presence of a xynA subfamily A gene |
| XynBf XynBr | AAGTGGTTTTTCTGCGA TAATGAAGTCTCTTTTCC | 538–554b 836–853b | PCR amplification of xynB family genes | Designed with two regions of DNA sequence conserved in xynB family genes but not in xynA subfamily A and I genes |
| Xyn38f1 | TATGCATGGGATGTATGT | See Fig. 2 | Resolution of the number of xynA subfamily I genes in A38 | Designed to be specific for the two different DNA sequences determined in the original PCR products from B. fibrisolvens A38 |
| Xyn38f2 | TATGCATGGGACGTTTGC | See Fig. 2 | ||
| Xyn38r1 | AGCTTGATTACATCGTTT | See Fig. 2 | ||
| Xyn38r2 | AGTGTGATTACGTTGTTA | See Fig. 2 |
Cloning of PCR products and sequence analysis.
PCR products were purified with Wizard PCR Preps (Promega) and cloned into pGEM-T. Ligated plasmid DNA was electroporated into Escherichia coli XL1-Blue and plated on Luria broth plates containing X-Gal (5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside; 20 μg ml−1), IPTG (isopropyl-β-d-thiogalactopyranoside; 1 mM), and 200 μg of ampicillin ml−1. Plasmid DNA was prepared from 2-ml overnight cultures grown in Luria broth containing 200 μg of ampicillin ml−1 with plasmid miniprep kits (Bio-Rad or Bresatec). Plasmid DNAs were sequenced with standard SP6 and T7 primers (Promega) with dye terminators (Applied Biosystems) and run on model 373 or 377 sequencers (Applied Biosystems). For each gene (except where stated below) at least one cloned PCR product was completely sequenced on both strands and additional cloned PCR products were entirely sequenced on at least one strand to identify errors that may have been introduced during PCR amplification. The sequence of xynF was from a single cloned PCR product. The sequence of xynZ was derived from four PCR products between the sites of primers xynBf and Br and from one PCR product outside this region. The sequences of xynD and xynH were determined by direct sequencing of PCR products between the XynA38f1 plus XynA38r1 and XynA38f2 plus XynA38r2 primer binding sites and from two cloned PCR products outside of these sites. Some other PCR products, derived under conditions in which mixed PCR products were not produced, were also sequenced directly with the PCR primers used for initial amplification. In this case, the sequences used were derived from double-stranded sequencing of the products of a single PCR amplification.
Sequence data were processed with Sequencer (Gene Codes Corporation), MacVector (Oxford Molecular Group), SeqPup, and CLUSTAL X (29). Dendrograms were drawn from the output of CLUSTAL X with njplot. Searches of databases were carried out by using BLAST at the National Center for Biotechnology Information web site.
Nucleotide sequence accession numbers.
The nucleotide sequences have been deposited in the GenBank database under accession no. AF125880 through AF125902.
RESULTS
Grouping of strains of B. fibrisolvens.
Amplified 16S rDNA PCR products were analyzed by restriction enzyme digestion with HpaII. Six different patterns where identified. Strains were then allocated to the appropriate group and, within group 1, to the appropriate type (Table 1). Strains in group 2 were allocated to types with the four restriction enzymes HhaI, Eco72I, Hsp92II, and MfeI (Table 1). The presence or absence of particular sites for these four enzymes allowed us to discriminate among all of the types in group 2 described by Willems et al. (31) and also Pseudobutyrivibrio ruminis (30). All strains were also typed by RAPD analyses to identify closely related strains. Only strains with clearly different RAPD patterns with at least one set of primers were included in further analyses (Table 1). In medium containing BWX, strains either reached an optical density at 600 nm of greater than 1 within 24 h or, with LP1309, LP1028, and OB156, did not show any detectable increase in their optical density at 600 nm in 24 h (Table 1). There was no clustering of these strains into particular 16S rDNA groups or types (Table 1).
Distribution of xynA and xynB in group 2 strains of B. fibrisolvens.
The pair of degenerate primers Xyn10f and Xyn10r amplified one major product of 440 bp in 4 of the 20 group 2 strains of B. fibrisolvens (Table 1). Representative cloned PCR products from H17c, Bu49, and CF3 were sequenced. Clones of the A46 product could not be isolated. The H17c sequence (xynA1) was identical to that of xynA from Bu49, and the CF3 sequence (xynA2) contained two silent base changes (Fig. 1).
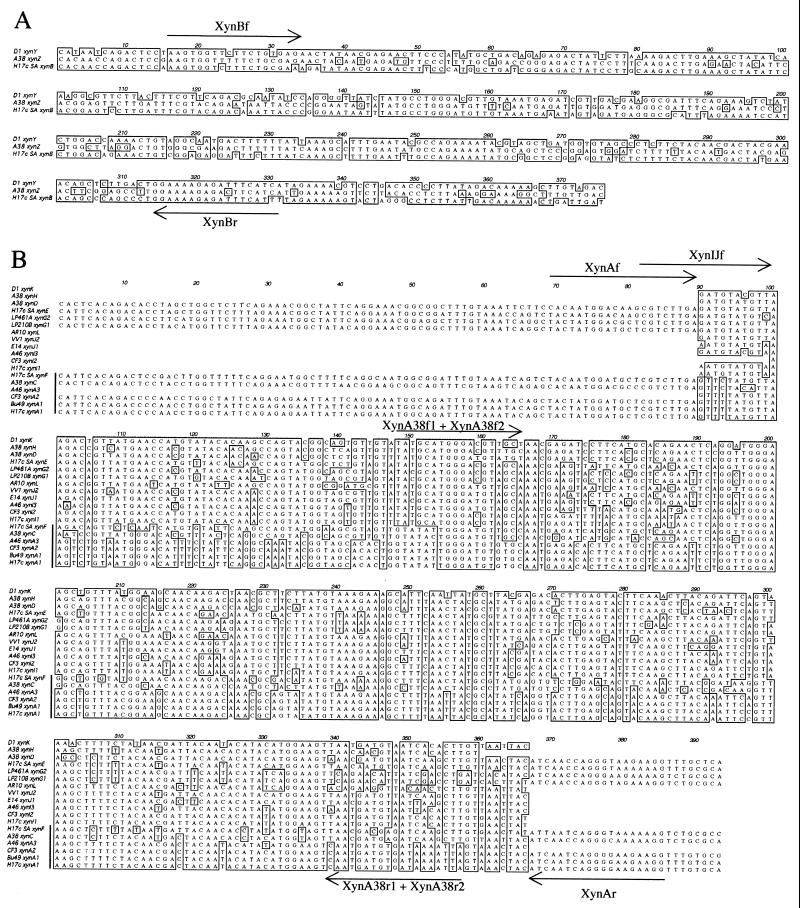
FIG. 1.
Alignment of nucleotide sequences of PCR products derived from the xylanase genes of B. fibrisolvens. (A) xynB family genes. Sequences shown are internal to the binding sites of the Xyn10f and Xyn10r primers. The locations of internal primers are indicated. Nucleotides conserved in two or more of the sequences are boxed. (B) xynA subfamily A and I genes. Sequences shown are internal to the binding sites of the Xyn10f and Xyn10r primers. The locations of internal primers are indicated. Nucleotides conserved in more than 50% of the sequences and internal to the binding sites of the XynAf and XynAr primers are boxed. A vertical line indicates the members of xynA subfamily A.
There was no evidence for a PCR product derived from xynB, reported to have been cloned from H17c (19), in any of the group 2 strains, including H17c itself. In order to identify the reason for the apparent absence of xynB from H17c, a culture of the strain used as the source of DNA for the cloning of xynB was obtained. This strain is henceforth referred to as H17c(SA). Unlike with H17c, PCR products of both 422 and 440 bp were amplified from strain H17c(SA) with primers Xyn10f and Xyn10r. The PCR products were cloned and sequenced. Three different xylanase genes were identified: xynB, identical to the published sequence (19), and two genes, xynE and xynF, each exhibiting approximately 70% DNA sequence identity with xynA1 (Fig. 1).
H17c(SA) and H17c are completely different strains of B. fibrisolvens.
The results described above suggested that H17c and H17c(SA) were different strains of B. fibrisolvens. RAPD analysis of H17c(SA) and H17c with two different sets of primers did not identify any common bands (Fig. 2). However, H17c had many bands in common with Bu49, as was expected from previous observations (20). Thus, H17c appears to be the same as the H17c used by Hespell and coworkers (11–13). Furthermore, restriction fragment length polymorphism (RFLP) analysis of the 16S rDNA PCR products amplified from H17c and H17c(SA) indicated that they were not closely related (Fig. 2). Sequence analysis of the region of the H17c(SA) rDNA gene between primers 27F and 500R (Table 2) placed the strain in group 1, type 6 (Fig. 3). In contrast, the equivalent region of the H17c 16S rDNA was identical to that previously reported from Bu49 (Fig. 3).
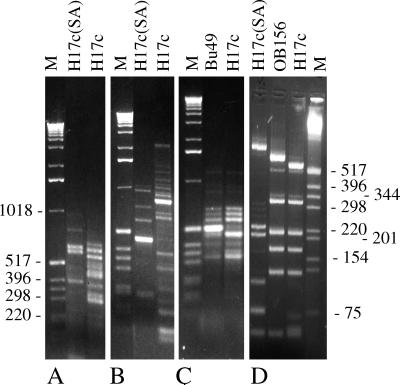
FIG. 2.
RAPD and 16S rDNA RFLP analysis of B. fibrisolvens Bu49, OB156, H17c, and H17c(SA). (A and C) RAPDs generated by primer pair 1-05 and 1-13. (B) RAPDs generated by primer pair 1-08 and 1-20. (D) HpaII-digested 16S rDNA PCR amplification products. Lengths of markers (M) are shown in base pairs.
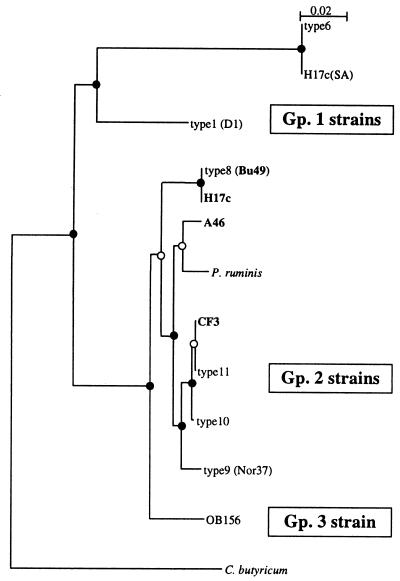
FIG. 3.
Dendrogram of 16S rDNA sequences from strains of B. fibrisolvens. Group 2 strains containing xynA subfamily A genes are indicated in bold. All sequences, except H17c(SA), H17c, A46, and CF3, were taken from GenBank. Clostridium butyricum was used as the outgroup. DNA sequences internal to the primers 27f and 500r were used to generate the dendrogram. The integrity of the topology was estimated by bootstrap resampling. Strongly supported branch points (bootstrap values of >75%) are indicated by filled circles, and marginally supported branches (bootstrap values of 50 to 74%) are indicated by open circles. Gp., group.
The data presented here concerning H17c and H17c(SA) indicate that it is probable that the other genes described from H17c(SA) (1, 2, 9, 17, 25, 26) are not applicable to the H17c originally isolated by Dehority (6).
Distribution of homologues of xynA and xynB in group 1 strains of B. fibrisolvens.
Primers Xyn10f and Xyn10r were used to amplify PCR products from several other members of group 1 B. fibrisolvens strains. The PCR products from A38, LP210B, LP461A, and D1 were cloned, and representatives of different groups of clones, identified by restriction enzyme analysis, were sequenced. No clones containing the PCR product from AR10 were isolated. One gene was identified in both LP210B (xynG1) and LP461A (xynG2) (Fig. 1), and no 422-bp PCR product characteristic of xynB homologues could be seen, even with the aid of restriction digestion (data not shown). In addition, only one gene, xynY, a homologue of xynB, was identified in D1 (Fig. 1). No 440-bp PCR product, characteristic of the xynA homologues, was present in the amplifications from D1, and the AluI patterns of the product were also consistent with the presence of only one xylanase gene (data not shown). The result was more complex for A38, from which a single cloned PCR product containing xynZ (a homologue of xynB) and multiple cloned PCR products containing xynC (an apparent homologue of xynA) were isolated and sequenced (Fig. 1). Three additional cloned PCR products, some of which appeared to represent chimeras generated during the PCR amplification from two other similar, but not identical, homologues of xynA, were also isolated. To determine the origin of the apparent chimeras, PCR primers specific for each of the two putative genes (Table 2) were used to obtain PCR products that confirmed the sequences of two closely related genes, xynD and xynH (Fig. 1).
To further investigate the apparent absence of homologues of xynB from many strains in group 1 and to obtain more clones of the A38 xynB homologue, xynZ, the primer pair XynBf and XynBr was used (Table 2). These primers amplified the known homologues of xynB from H17c(SA), A38, and D1, but no bona fide products were amplified from LP210B, LP461A, AR10, or AR11, even with annealing temperatures reduced to 38°C. The PCR products derived from A38 were cloned, and the sequence of xynB was confirmed (Fig. 1).
Reinvestigation of distribution of homologues of xynB in group 2 and group 3 strains of B. fibrisolvens.
The PCR primers XynBf and XynBr were used to screen the group 2 strains for the presence of homologues of xynB. Despite amplification being carried out at an annealing temperature of 40°C, no bona fide xynB homologues could be detected in any group 2 or 3 strains of B. fibrisolvens (Table 1).
Identification of further homologues of xynA with a second set of primers.
The original description of the cloning of xynA from Bu49 raised the possibility of the existence of an additional homologue of xynA in some strains of B. fibrisolvens (21). Our results with Xyn10f and Xyn10r did not clearly identify a second putative homologue of xynA, but minor PCR products were observed after restriction digestion (data not shown). Given the nature of PCR with highly degenerate primers and uncertainty concerning the level of conservation of the regions chosen for the original PCR primers, clearly some genes could have been missed. To address these issues, a further set of PCR primers, XynAf and XynAr, was designed (Table 2; Fig. 1) and used to screen all strains for the presence of additional homologues of xynA. Products of the expected size were amplified from a number of strains that had previously been negative with the primers Xyn10f and Xyn10r (Table 1). PCR products different from those xynA and xynB homologues already characterized from D1 (xynK), H17c (xynI1), and CF3 (xynI2) and products from the previously negative strains VV1 (xynJ2) and E14 (xynJ1) were cloned and sequenced (Fig. 1). In addition, a product from Bu49 with a restriction pattern identical to that of xynI1 from H17c was obtained. The products from two strains, A46 (xynA3) and AR10 (xynL), from which we had been unable to isolate clones of PCR products from the Xyn10f and Xyn10r amplifications were also cloned and sequenced (Fig. 1). To amplify a pure product of a second xynA homologue from A46, another internal primer, XynIJf (Table 2; Fig. 1), was used in conjunction with XynAr to clone and sequence xynI3 (Fig. 1).
The strains of B. fibrisolvens that were not positive with any of the sets of primers were screened with the primers XynAf and XynAr by using annealing temperatures down to 38°C. The Xyn10f-XynAr and XynAf-Xyn10r primer pairs were also used. In all cases, although a significant number of PCR products were amplified, no clear products of the expected sizes for bona fide xylanase genes were identified.
Analysis of the relationships between the sequenced xylanase gene fragments.
A dendrogram was constructed with all of the xylanase sequences obtained (Fig. 4). xynB, xynZ, and xynY (designated the xynB family) clearly form a separate cluster. The analysis supported the division of the genes homologous to xynA into two groups. The minor group contained xynA1 to -3, xynC, and xynF and has been designated subfamily A of the xynA gene family. The major group contained all of the other genes and has been designated subfamily I. Both subfamilies span the group 1 and group 2 B. fibrisolvens strains. Within subfamily I, the clustering of sequences was basically congruent with the 16S rDNA groups (Table 1), with the clear exception of the CF3 sequence (xynI2), which was almost identical to the H17c xynI1 sequence, while CF3 and H17c lay in quite distinct 16S rDNA types (Table 1). The members of subfamily A present in the group 2 strains, H17c, Bu49, CF3, and A46, were essentially identical (Fig. 1), but on the basis of 16S rDNA RFLPs, these strains lay in three different types (Table 1). In order to examine these inconsistencies in more detail, the sequences of the region of the 16S rDNAs between primers 27f and 500r were determined for A46 and CF3 and compared with this region in other group 2 strains of B. fibrisolvens (Fig. 3). Comparison of the two dendrograms confirms the lack of congruence between the subfamily A and I genes and the 16S rDNA of CF3 and between the subfamily A gene and the 16S rDNA of A46, when compared to the congruence of the subfamily A and I genes and 16S rDNAs of H17c and Bu49.
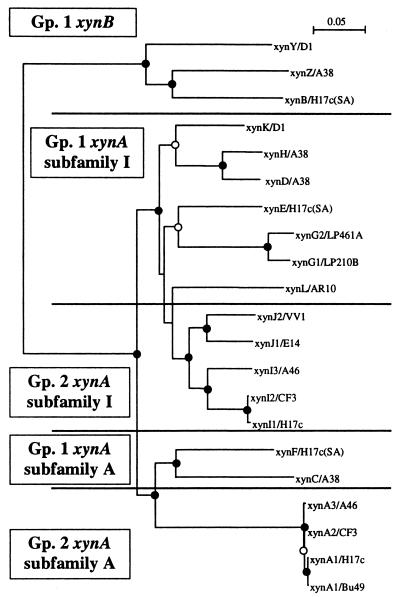
FIG. 4.
Dendrogram of xylanase gene sequences from strains of B. fibrisolvens. DNA sequences internal to the primers XynAf and XynAr for xynA subfamily A and I genes and the equivalent regions of xynB, xynX, and xynZ were used to generate the dendrogram. The xynB family genes were used as the outgroup for the analysis. The integrity of the topology was estimated by bootstrap resampling. Strongly supported branch points (bootstrap values of >75%) are indicated by filled circles, and marginally supported branches (bootstrap values of 50 to 74%) are indicated by open circles. Branch points without circles are not resolved. Gp., group.
DISCUSSION
We have identified two related, but distinct, subfamilies of xylanase genes in B. fibrisolvens that are similar to the previously characterized xynA gene from strain Bu49. The two subfamilies have been designated xynA subfamilies A and I. These genes are not present in the majority of B. fibrisolvens strains analyzed or even in the majority of xylanolytic strains. However, our findings suggest that the faint bands previously observed in a number of other strains after hybridization with xynA (21) probably also represent homologues of xynA. Indeed, since H17c, Bu49, CE-51, CE-52, and 12 form a cluster of closely related strains (group 2, type 8) (20, 31), it is likely that, in these strains, the genes are members of xynA subfamily I. Of this group, only H17c and Bu49 contain xynA. The sporadic distribution of members of xynA subfamily A and the high level of sequence similarity of subfamily A genes in the characterized members of the group 2 B. fibrisolvens strains suggest that xynA may have been horizontally transferred between a progenitor of H17c, CF3, or A46 and the progenitors of the other two strains. The alternative explanation of a very low rate of fixation of mutations in xynA subfamily A genes in the group 2 strains in conjunction with a high rate of gene loss appears less likely. The distribution of members of the xynA subfamily I across both group 1 and group 2 strains and the relatively good congruency between xylanase gene and 16S rDNA groupings suggest that the progenitor B. fibrisolvens contained at least a xynA subfamily I gene. Subsequent loss of the xynA subfamily I gene appears to have occurred a number of times during the evolution of B. fibrisolvens. The current data does not enable us to distinguish between the following two possible explanations for the current distribution of xynA subfamily A genes across both groups 1 and 2: first, duplication of a xynA subfamily I gene to generate a xynA subfamily A gene in a progenitor of only the group 1 B. fibrisolvens strains followed by horizontal transfer into the group 2 B. fibrisolvens strains and, second, the presence of a xynA subfamily A gene in the progenitors of both group 1 and 2 strains followed by extensive gene loss. For those strains with more than one xynA homologue, with the exception of xynD and xynH in A38, the data is certainly not consistent with the extra genes arising from recent gene duplication events. Interestingly, no strain containing only a xynA subfamily A gene has yet been identified.
Three group 1 strains, D1, A38, and H17c(SA), also contain a member of the second family of xylanases, xynB, identified in the B. fibrisolvens strains. The distribution of xynB family genes and their sequence relationships are consistent with the progenitor of the group 1 strains having contained a member of the xynB family. A number of losses of xynB family genes would be required to explain the current distribution. In particular, it would have been lost from a very early progenitor of the group 2 and 3 strains of B. fibrisolvens.
The predicted XynB product does not appear to have a bona fide signal sequence and therefore may be an intracellular enzyme (19). While it is likely that XynA and XynB play quite different roles in the degradation of plant fiber, it is not clear what differences might exist between the members of XynA subfamilies A and I. There is a fairly good correlation between the capacity of strains of B. fibrisolvens to degrade spear grass (22) and the presence of members of xynA subfamily I (xylanase-encoding genes). The correlation is better between the presence of xynA subfamily A members and degradation capacity. Five of the six best xylan-degrading strains of B. fibrisolvens included in our analysis contain a member of xynA subfamily A.
Bu49 has been reported to produce at least two xylanases, one of approximately 12 to 16 kDa and one of approximately 46 kDa (12, 21); the latter enzyme was assumed to be XynA. However, given the high level of predicted sequence similarity between XynA and XynI, it may well have been XynI or both XynA and XynI. The gene encoding the small acidic xylanase (21) has not yet been identified. Given the likely limitations of our primers and the impossibility of designing a single set of PCR primers that are universal for ghf10 xylanase-encoding genes, it cannot be ruled out that this as yet undiscovered gene may also encode an enzyme in ghf10. Experiments with a set of degenerate PCR primers designed against conserved regions of the ghf11 xylanases did not amplify any bona fide xylanase products from strains of B. fibrisolvens (unpublished). The same primers did amplify products from strains of Ruminococcus sp. and from a recombinant strain of B. fibrisolvens containing a gene encoding a ghf11 xylanase (unpublished). Homologues of the additional enzyme described from Bu49 may also be responsible for the xylanase activities observed in the majority of strains of B. fibrisolvens which, by our PCR-based screening, do not appear to contain members of either of the two subfamilies of xynA or the xynB family. The lack of a xynA homologue in one of these strains, Nor37, is consistent with the negative result obtained for the same strain by hybridization with xynA (20).
The work described here has increased our understanding of the evolution and distribution of xylanase genes in a diverse group of B. fibrisolvens strains. Many of the strains used in the analysis are candidates for transformation with genes encoding xylanases to increase their capacity to degrade plant material (8), and this work provides the basis for a more targeted approach to this manipulation. It also provides a framework for interpretation of the results of manipulation of xylanolytic activity by the addition of recombinant enzymes.
ACKNOWLEDGMENTS
We thank Jennifer A. Thomson for the gift of B. fibrisolvens H17c(SA); Keith Gregg for the gift of AR strains of B. fibrisolvens; and Jeff Cowley, Denis Krause, and Chris McSweeney for useful discussions.
This work was partly funded by the Australian Meat Research Corporation.
REFERENCES
- 1.Berger E, Jones W A, Jones D T, Woods D R. Cloning and sequencing of an endoglucanase (end1) gene from Butyrivibrio fibrisolvens H17c. Mol Gen Genet. 1989;219:193–198. doi: 10.1007/BF00261176. [DOI] [PubMed] [Google Scholar]
- 2.Berger E, Jones W A, Jones D T, Woods D R. Sequencing and expression of a cellodextrinase (ced1) gene from Butyrivibrio fibrisolvens H17c cloned in Escherichia coli. Mol Gen Genet. 1990;223:310–318. doi: 10.1007/BF00265068. [DOI] [PubMed] [Google Scholar]
- 3.Bryant M P, Burkey L A. Numbers and some predominant groups of bacteria in the rumen of cows fed different rations. J Dairy Sci. 1953;36:218–224. [Google Scholar]
- 4.Bryant M P, Small N. The anaerobic monotrichous butyric acid-producing curved rod-shaped bacteria of the rumen. J Bacteriol. 1955;72:16–21. doi: 10.1128/jb.72.1.16-21.1956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dalrymple B P, McSweeney C S. Manipulation of rumen function by the inoculation of novel microorganisms. Outlook Agric. 1998;27:107–113. [Google Scholar]
- 6.Dehority B A. Characterization of several bovine rumen bacteria isolated with a xylan medium. J Bacteriol. 1966;129:1724–1729. doi: 10.1128/jb.91.5.1724-1729.1966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Forster R J, Teather R M, Gong J, Deng S-J. 16S rDNA analysis of Butyrivibrio fibrisolvens: phylogenetic position and relation to butyrate-producing anaerobic bacteria from the rumen of white-tailed deer. Lett Appl Microbiol. 1996;23:218–222. doi: 10.1111/j.1472-765x.1996.tb00069.x. [DOI] [PubMed] [Google Scholar]
- 8.Gobius, K. S., G. P. Xue, J. H. Aylward, D. O. Krause, Y. J. Swadling, C. S. McSweeney, and B. P. Dalrymple. Transformation and expression of an anaerobic fungal xylanase in several strains of the rumen bacterium Butyrivibrio fibrisolvens. Submitted for publication. [DOI] [PubMed]
- 9.Goodman H J K, Woods D R. Cloning and nucleotide sequence of the Butyrivibrio fibrisolvens gene encoding a type III glutamine synthetase. J Gen Microbiol. 1993;139:1487–1493. doi: 10.1099/00221287-139-7-1487. [DOI] [PubMed] [Google Scholar]
- 10.Henrissat B. A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J. 1991;280:309–316. doi: 10.1042/bj2800309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hespell R B, Cotta M A. Degradation and utilization by Butyrivibrio fibrisolvens H17c of xylans with different chemical and physical properties. Appl Environ Microbiol. 1995;61:3042–3050. doi: 10.1128/aem.61.8.3042-3050.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hespell R B, Whitehead T R. Physiology and genetics of xylan degradation by gastrointestinal tract bacteria. J Dairy Sci. 1990;73:3013–3022. doi: 10.3168/jds.S0022-0302(90)78988-6. [DOI] [PubMed] [Google Scholar]
- 13.Hespell R B, Wolf R, Bothast R J. Fermentation of xylans by Butyrivibrio fibrisolvens and other ruminal bacteria. Appl Environ Microbiol. 1987;53:2849–2853. doi: 10.1128/aem.53.12.2849-2853.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hudman J F, Gregg K. Genetic diversity among strains of bacteria from the rumen. Curr Microbiol. 1989;19:313–318. [Google Scholar]
- 15.Kobayashi Y, Okuda N, Matsumoto M, Inoue K, Wakita M, Hoshino S. Constitutive expression of a heterologous Eubacterium ruminantium xylanase gene (xynA) in Butyrivibrio fibrisolvens. FEMS Microbiol Lett. 1998;163:11–17. doi: 10.1111/j.1574-6968.1998.tb13019.x. [DOI] [PubMed] [Google Scholar]
- 16.Lane D J. 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. Chichester, United Kingdom: John Wiley & Sons Ltd.; 1991. pp. 115–176. [Google Scholar]
- 17.Lin L L, Rumbak E, Zappe H, Thomson J A, Woods D R. Cloning, sequencing and analysis of expression of a Butyrivibrio fibrisolvens gene encoding a beta-glucosidase. J Gen Microbiol. 1990;136:1567–1576. doi: 10.1099/00221287-136-8-1567. [DOI] [PubMed] [Google Scholar]
- 18.Lin L L, Thomson J A. An analysis of the extracellular xylanases and cellulases of Butyrivibrio fibrisolvens H17c. FEMS Microbiol Lett. 1991;68:197–203. doi: 10.1016/0378-1097(91)90127-v. [DOI] [PubMed] [Google Scholar]
- 19.Lin L L, Thomson J A. Cloning, sequencing and expression of a gene encoding a 73 kDa xylanase enzyme from the rumen anaerobe Butyrivibrio fibrisolvens H17c. Mol Gen Genet. 1991;228:55–61. doi: 10.1007/BF00282447. [DOI] [PubMed] [Google Scholar]
- 20.Mannarelli B M. Deoxyribonucleic acid relatedness among strains of the species Butyrivibrio fibrisolvens. Int J Syst Bacteriol. 1988;38:340–347. [Google Scholar]
- 21.Mannarelli B M, Evans S, Lee D. Cloning, sequencing, and expression of a xylanase gene from the anaerobic ruminal bacterium Butyrivibrio fibrisolvens. J Bacteriol. 1990;172:4247–4254. doi: 10.1128/jb.172.8.4247-4254.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.McSweeney C S, Dulieu A, Bunch R. Butyrivibrio spp. and other xylanolytic microorganisms from the rumen have cinnamoyl esterase activity. Anaerobe. 1998;4:57–65. doi: 10.1006/anae.1997.0130. [DOI] [PubMed] [Google Scholar]
- 23.Orpin C G, Mathiesen S D, Greenwood Y, Blix A S. Seasonal changes in the ruminal microflora of the high-arctic Svalbard reindeer (Rangifer tarandus platyrhynchus) Appl Environ Microbiol. 1985;50:144–151. doi: 10.1128/aem.50.1.144-151.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ruma P, Chen S C, Sorrell T C, Brownlee A G. Characterization of Cryptococcus neoformans by random DNA amplification. Lett Appl Microbiol. 1996;23:312–316. doi: 10.1111/j.1472-765x.1996.tb00197.x. [DOI] [PubMed] [Google Scholar]
- 25.Rumbak E, Rawlings D E, Lindsey G G, Woods D R. Characterization of the Butyrivibrio fibrisolvens glgB gene, which encodes a glycogen-branching enzyme with starch-clearing activity. J Bacteriol. 1991;173:6732–6741. doi: 10.1128/jb.173.21.6732-6741.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rumbak E, Rawlings D E, Lindsey G G, Woods D R. Cloning, nucleotide sequence, and enzymatic characterization of an α-amylase from the ruminal bacterium Butyrivibrio fibrisolvens H17c. J Bacteriol. 1991;173:4203–4211. doi: 10.1128/jb.173.13.4203-4211.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sewell G W, Aldrich H C, Williams D, Mannarelli B, Wilkie A, Hespell R B, Smith P H, Ingram L O. Isolation and characterization of xylan-degrading strains of Butyrivibrio fibrisolvens from a Napier grass-fed anaerobic digester. Appl Environ Microbiol. 1988;54:1085–1090. doi: 10.1128/aem.54.5.1085-1090.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Stackebrandt E, Liesack W. The potential of rDNA in identification and diagnostics. In: Kessler C, editor. Nonradioactive labeling and detection of biomolecules. Berlin, Germany: Springer-Verlag; 1992. pp. 232–239. [Google Scholar]
- 29.Thompson J D, Gibson T J, Plewniak F, Jeanmougin F, Higgins D G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997;25:4876–4882. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.van Gylswyk N O, Hippe H, Rainey F A. Pseudobutyrivibrio ruminis gen. nov., sp. nov., a butyrate-producing bacterium from the rumen that closely resembles Butyrivibrio fibrisolvens in phenotype. Int J Syst Bacteriol. 1996;46:559–563. [Google Scholar]
- 31.Willems A, Amat-Marco M, Collins M D. Phylogenetic analysis of Butyrivibrio strains reveals three distinct groups of species within the Clostridium subphylum of the gram-positive bacteria. Int J Syst Bacteriol. 1996;46:195–199. doi: 10.1099/00207713-46-1-195. [DOI] [PubMed] [Google Scholar]
- 32.Williams A G, Withers S E. Factors affecting the formation of polysaccharide depolymerase and glycoside hydrolase enzymes by Butyrivibrio fibrisolvens NCDO 2249. J Appl Bacteriol. 1989;67:299–308. [Google Scholar]
- 33.Williams A G, Withers S E. The regulation of xylanolytic enzyme formation by Butyrivibrio fibrisolvens NCFB 2249. Lett Appl Microbiol. 1992;14:194–198. [Google Scholar]
- 34.Williams A G, Withers S E. Induction of xylan-degrading enzymes in Butyrivibrio fibrisolvens. Curr Microbiol. 1992;25:297–303. [Google Scholar]
- 35.Xue G P, Johnson J S, Bransgrove K L, Gregg K, Beard C E, Dalrymple B P, Gobius K S, Aylward J H. Improvement of expression and secretion of a fungal xylanase in the rumen bacterium Butyrivibrio fibrisolvens OB156 by manipulation of promoter and signal sequences. J Biotechnol. 1997;54:139–148. doi: 10.1016/s0168-1656(97)01671-4. [DOI] [PubMed] [Google Scholar]