Abstract
Introduction
With advances in HBV therapies, there is a need to identify serum biomarkers that assess the HBV cccDNA reservoir and predict functional cure in HIV/HBV co-infection.
Methods
In this retrospective study combining samples from HIV/HBV co-infected participants enrolled in two ACTG interventional trials, proportions achieving HBsAg <0.05 log10 IU/mL and HBV RNA <log10 1.65 U/mL or not detected (LLoQ/NEG) in response to DUAL [tenofovir TDF+emtricitabine (FTC)) vs. MONO (FTC or lamivudine (3TC)] HBV-active ART, were measured. Predictors of qHBsAg <0.05 log10 IU/mL were evaluated in logistic regression models.
Results
There were 88 participants [58% female, median age 34; 47 on DUAL vs. 41 on MONO HBV-active ART]. 21% achieved HBsAg <0.05 log10 IU/mL (30% DUAL vs. 10% MONO). Time to HBsAg <0.05 log10 IU/mL was lower (p=0.02) and the odds of achieving HBsAg <0.05 log10 IU/mL were higher (p=0.07) in DUAL participants. HBV RNA became <LLoQ/NEG in 47% (DUAL 60% vs. MONO 33%). qHBsAg <3 log10 IU/mL was the strongest predictor of HBsAg <0.05 log10 IU/mL.
Conclusion
This study supports current recommendations of TDF based DUAL-HBV active ART for initial use in HIV/HBV co-infection. HBV RNA could be a useful marker of treatment response in HIV/HBV co-infected patients on HBV-active ART.
Keywords: quantitative HBsAg, HBV RNA, HIV/HBV co-infection
Introduction
Chronic hepatitis B (HBV) is common among persons with HIV. In countries where both viruses are endemic, the prevalence of HIV/HBV co-infection is as high as 28% [1]. Current recommended therapies for the treatment of HIV/HBV co-infection include the combination of nucleoside analogs (NrtI), tenofovir (TDF) and lamivudine (3TC) or emtricitabine (FTC), and have been shown to effectively suppress HBV replication [2]. However, complete eradication of HBV virus is rarely achieved with NrtIs because they do not directly affect hepatocyte reservoirs of covalently closed circular DNA (cccDNA).
With recent progress in the development of novel anti-HBV drugs, and new strategies for treatment cessation with existing antivirals, there has been a renewed search for serum markers that more reliably measure the effects of therapy on viral replication, and the likelihood of achieving functional or complete cure. HBV DNA is a serum marker for cccDNA activity, however HBV DNA becomes rapidly undetectable in NrtI treated patients and therefore not an optimal marker of cccDNA activity in patients undergoing therapy. Since cccDNA itself is challenging to measure, requiring biopsy, surrogate markers that more effectively reflect the transcriptional activity of intrahepatic virus and the efficacy of treatment are needed [3].
Quantitative HBsAg (qHBsAg) and HBV RNA are serum markers, which have been shown to be positively correlated with intrahepatic transcription activity in chronic HBV and are being explored as prognostic markers [4–6]. HBsAg, which is a glycosylated envelope protein of HBV virions, is synthesized from translated messenger RNAs of transcriptionally active cccDNA as well as from integrated HBV DNA [7]. HBV RNA circulating in serum, recently confirmed as primarily pregenomic RNA (pgRNA), is derived from intrahepatic cccDNA and serves as the transcriptional intermediary in HBV replication [8,9]. Since HBsAg and HBV RNA have origins from cccDNA and integrated DNA (HBsAg), and are not directly affected by NrtI’s, they may be better surrogate markers of antiviral efficacy compared to HBV DNA [10]. In clinical studies, qHBsAg can predict spontaneous clearance of HBsAg, liver fibrosis and hepatocellular cancer (HCC) [11]. In HBV mono-infection, HBV RNA has been shown to have an important role in determining the potency of antivirals and predicting initial virologic response or earlier HBV suppression to antiviral therapies [8,12]. Both markers have been shown to predict HBV reactivation after discontinuation of NrtI therapy [13,14].
Measurement of qHBsAg and HBV RNA in HIV/HBV co-infected individuals specifically, may be useful to determine the efficacy of HBV active antiretroviral treatment (HBV-ART) and a more precise estimation of duration of therapy needed to achieve HBsAg clearance. These markers could also help to assess whether certain individuals could switch to more simplified ART regimens (containing only one or no HBV active drugs), which are increasingly available [15].
In this study we evaluated dynamic changes in qHBsAg and HBV RNA over 144 weeks in response to HBV-ART among HIV/HBV co-infected participants enrolled in two completed large multinational ART trials. Changes in HBV RNA, qHBsAg, and the proportion achieving HBsAg <0.05 log10 IU/mL and HBV RNA less than the lower limit of quantitation or negative (<LLOQ/NEG) respectively, were compared between participants on ART containing two (DUAL) vs. one (MONO) HBV active agent to evaluate differences in regimen efficacy. Predictors of HBsAg <0.05 log10 IU/mL and qHBsAg correlations with HBV RNA and HBV DNA were evaluated.
Methods
This retrospective observational study included HIV/HBV co-infected adult (>18y) participants who had enrolled in two multi-national randomized controlled trials (RCTs) for HIV-1 treatment; ACTG 5175 (PEARLS) and A5208 (OCTANE) [16,17]. The OCTANE study, comprised two concurrent, randomized, open-label trials of ART that included only women with and without prior exposure to single-dose nevirapine who were then randomly assigned at least six months post-partum to either ritonavir-boosted lopinavir plus tenofovir–emtricitabine (TDF-FTC) vs. nevirapine plus TDF-FTC, daily. ACTG 5175 (PEARLS) was a randomized, multinational clinical trial in HIV-1-infected, ART-naïve participants who were randomized to one of three ART regimens; i) zidovudine/lamivudine (ZDV-3TC) + efavirenz (EFV); ii) didanosine (DDI) + FTC + atazanavir (ATV), or iii) co-formulated TDF-FTC + EFV. HIV/HBV co-infected participants from these two studies were identified for inclusion in this study if they had a (1) positive qualitative HBsAg result from screening for the treatment study; (2) HBV DNA results from the completed molecular characterization study [18]; and (3) available specimens at baseline with ≥2ml plasma.
Informed consents for the ACTG 5175 (PEARLS) and 5208 (OCTANE) [16,17] parent RCTs were obtained from the study participants in their native language. This sub-study was approved by the local institutional review boards and Northwestern University IRB.
Laboratory testing
HBV RNA and qHBsAg levels were tested at Abbott Laboratories on serum specimens stored at −80°C and at 24-week intervals from baseline (week 0) to week 144. pgRNA was detected using the Abbott RealTime 0.2 mL HBV RNA Research Use Only (RUO) assay (Abbott Diagnostics, Abbott Park, IL), as previously described [19]. This dual-target assay is designed to quantify HBV RNA from highly conserved targets in the core and X genes located at the 5’ and 3’ ends of the full-length pgRNA, respectively. Primers amplify RNA from both pgRNA and HBV mRNA. Assay values are reported as log U/mL determined by calibration of the assay with a DNA secondary standard (Abbott Molecular, Des Plaines, IL) that is traceable to the World Health Organization HBV DNA standard where 1 U of HBV RNA is equal to 1 IU of HBV DNA. Levels detectable but less than the lower limit of quantification (<LLoQ) were calculated to be 1.81 log10 U/mL and 1.65 log10 U/L for core and X targets, respectively. HBV RNA below LLoQ that was not detected was classified as negative (NEG). HBsAg was quantitated using Abbott ARCHITECT HBsAg (Abbott Diagnostics, Sligo, Ireland), a quantitative chemiluminescent microparticle assay. Concentration values ≥0.05 log10 IU/ml are considered reactive and values <0.05 log10 IU/mL are considered nonreactive; however, it is possible that HBsAg is present below this limit.
Serologic testing for HBeAg, anti-HBe, HBV DNA (LLoQ 200 IU/mL) were performed in a characterization study [18] as previously described.
Statistical analysis
The qHBsAg results below LLoQ of 0.05 log10 IU/mL were assigned 0.025 (one-half of the LLoQ). Similarly, HBV RNA (U/mL) results below the LLoQ of 1.65 were assigned 0.825 and NEG results were assigned 0. Time to qHBsAg loss was calculated as the number of weeks between the ART study entry (week 0, baseline) and the sample date of the nonreactive qHBsAg. Time to first HBV RNA <LLoQ/NEG was computed as the number of weeks between the ART study entry (week 0) and the date of the earliest sample with HBV RNA <LLoQ or NEG. For participants who did not have the event, the follow-up time was censored at the last study visit with test result. Week 24 qHBsAg decline of at least 1 log10 from baseline was assessed for the participants whose baseline qHBsAg was at least 10 IU/mL (1 log10).
Fisher’s exact tests were conducted for comparisons of binary or categorical measures between groups, and Cochran-Mantel-Haenszel tests for stratified analyses. Wilcoxon rank-sum tests were conducted for comparisons of continuous measures between DUAL and MONO groups. DUAL participants were those who received TDF+ FTC in their ART regimen. MONO participants were those who received only FTC or 3TC in their ART regimen. Additional analyses stratified by sex at birth were also conducted, given the higher proportion of females in DUAL due to the OCTANE trial design. Comparisons of times-to-event were conducted by log-rank tests. Spearman rank-order correlation coefficient was used to evaluate associations between HBV markers. Fisher’s transformation with bias-adjustment was used to construct 95% confidence intervals for the correlation coefficients.
Logistic regression models were used to evaluate potential predictors for qHBsAg loss. Predictors included in the model were age, sex, baseline HBV DNA, HBV RNA, qHBsAg, ALT, HBeAg, CD4+ T-cell count, HIV RNA, MONO or DUAL for HBV. For variable selection, stepwise selection method was applied, using a 0.15 significance level for entry and 0.20 to remain in the model. Backward elimination method using a 0.20 significance level was also conducted to assess how sensitive the final model was to the selection method.
Results
Inclusion criteria were met in 99 participants. 11 were excluded from the final analysis: 9 participants had a non-reactive qHBsAg at baseline; one participant had a missing baseline qHBsAg result; and in one participant adefovir was the single HBV-active agent in their treatment regimen.
Of the 88 participants included in the analysis, 47 received DUAL (TDF+FTC) HBV-ART and 41 received MONO HBV-ART (18 on 3TC, 23 on FTC). Most (82%) participants were from non-US sites and were non-Hispanic Black [Table 1]. Overall, 59% were female, with more males (61%) in the MONO group and more females (77%) in the DUAL group. Hepatitis B genotype A was detected in most participants and 50% were HBeAg positive. HBV RNA was positive >1.65 log10 U/L (LLoQ) in 77% of participants. 89% (MONO 100%, DUAL 79%) had ≥1 log10 IU/mL HBsAg at baseline.
Table 1.
Baseline characteristics
| aAnti-HBV therapy | ||||
|---|---|---|---|---|
| Characteristic: Median (Q1,Q3) or Proportion (%) |
Total N=88 |
MONO (3TC or FTC) N=41 |
DUAL (TDF+FTC) N=47 | |
| Age (years) | 34 (30, 40) | 35 (32, 40) | 34 (29, 40) | |
| 20–29 | 17 (19%) | 5 (12%) | 12 (26%) | |
| 30–39 | 47 (53%) | 25 (61%) | 22 (47%) | |
| 40–49 | 20 (23%) | 10 (24%) | 10 (21%) | |
| 50–59 | 3 (3%) | 1 (2%) | 2 (4%) | |
| ≥60 | 1 (1%) | 0 (0%) | 1 (2%) | |
| Sex at birth | M | 36 (41%) | 25 (61%) | 11 (23%) |
| F | 52 (59%) | 16 (39%) | 36 (77%) | |
| Race/Ethnicity | White Non-Hispanic | 3 (3%) | 2 (5%) | 1 (2%) |
| Black Non-Hispanic | 72 (82%) | 31 (76%) | 41 (87%) | |
| Hispanic (Regardless of Race) | 6 (7%) | 4 (10%) | 2 (4%) | |
| Asian, Pacific Islander | 7 (8%) | 4 (10%) | 3 (6%) | |
| Country | Botswana | 3 (3%) | 0 (0%) | 3 (6%) |
| Haiti | 4 (5%) | 2 (5%) | 2 (4%) | |
| Kenya | 2 (2%) | 0 (0%) | 2 (4%) | |
| Malawi | 20 (23%) | 13 (32%) | 7 (15%) | |
| Peru | 6 (7%) | 4 (10%) | 2 (4%) | |
| South Africa | 13 (15%) | 3 (7%) | 10 (21%) | |
| Thailand | 7 (8%) | 4 (10%) | 3 (6%) | |
| United States | 10 (11%) | 8 (20%) | 2 (4%) | |
| Zimbabwe | 23 (26%) | 7 (17%) | 16 (34%) | |
| BMI * | 22.2 (19.9, 23.9) | 21.4 (19.8, 23.8) | 23.0 (20.7, 24.2) | |
| CD4 T cell (count/mm 3 ) | 136.5 (71.5, 218.5) | 161 (92, 241) | 119 (70, 194) | |
| Log10 HIV RNA (cp/mL) | 5.06 (4.54, 5.59) | 4.93 (4.41, 5.42) | 5.22 (4.69, 5.78) | |
| ALT (IU/L) | 25 (20, 38) | 28 (21, 38) | 24 (18, 34) | |
| ≥ULN | 20 (23%) | 10 (24%) | 10 (21%) | |
| AST (IU/L) | 35 (27, 44) | 39 (27, 52) | 33 (27, 40) | |
| ≥ULN | 42 (48%) | 19 (46%) | 23 (49%) | |
| HBeAg | Positive | 44 (50%) | 20 (49%) | 24 (51%) |
| Negative | 44 (50%) | 21 (51%) | 23 (49%) | |
| Anti-HDV | Positive | 13 (15%) | 1 (2%) | 12 (26%) |
| Negative | 75 (85%) | 40 (98%) | 35 (74%) | |
| HBV DNA log10 IU/mL | 5.17 (2.74, 8.13) | 5.75 (2.92, 8.20) | 4.39 (2.58, 7.52) | |
| HBV DNA ≥200 IU/mL | 77 (88%) | 36 (88%) | 41 (87%) | |
| HBV Genotype * | A | 65 (83%) | 29 (81%) | 36 (86%) |
| C | 4 (5%) | 3 (8%) | 1 (2%) | |
| D | 3 (4%) | 1 (3%) | 2 (5%) | |
| E | 3 (4%) | 1 (3%) | 2 (5%) | |
| F | 3 (4%) | 2 (6%) | 1 (2%) | |
| qHBsAg (log10 IU/mL) | 3.53 (2.73, 4.69) | 4.17 (2.99, 4.67) | 3.12 (2.19, 4.72) | |
| qHBsAg ≥1log10 IU/mL | 78 (89%) | 41 (100%) | 37 (79%) | |
| qHBsAg ≥1000 IU/mL | 56 (64%) | 30 (73%) | 26 (55%) | |
| HBV RNA (log10 U/mL) | 3.80 (1.78, 6.60) | 4.49 (2.00, 6.63) | 2.61 (0.83, 6.57) | |
|
HBV RNA (log10
U/mL) |
Negative | 3 (3%) | 0 (0%) | 3 (6%) |
| LLOQ (<1.65 log10)** | 17 (19%) | 8 (20%) | 9 (19%) | |
| Positive | 68 (77%) | 33 (80%) | 35 (74%) | |
DUAL regimens: tenofovir (TDF), emtricitabine (FTC) and lopinavir/ritonavir (LPV/r) 10 (21%); TDF, FTC and efavirenz (EFV) 20 (43%); TDF, FTC and nevirapine (NVP) 17 (36%); MONO regimens: FTC and atazanavir (ATV) 1 (2%); didanosine (DDI), FTC and ATV 22 (54%); combivir (zidovudine and lamivudine) and EFV 17 (41%), combivir 1 (1%).
Missing BMI in 27, all in DUAL; Missing Genotype in 10 (5 MONO, 5 DUAL)
Excludes Negatives
Changes in Quantitative HBsAg by ART regimen
Median baseline qHBsAg was 3.53 (Q1-Q3: 2.73 – 4.69) log10 IU/mL and higher among participants randomized to MONO vs. DUAL participants [4.17 vs 3.12 log10 IU/mL; p=0.040; stratified by sex, p=0.022]. Median qHBsAg at week 144 was 2.81 (Q1-Q3:1.83– 3.65) log10 IU/mL (n=46 due to loss of follow-up). Overall, 18/87 (21%) participants had reduced HBsAg below the assay’s 0.05 log10 IU/mL limit of detection [14/46 (30%) DUAL vs. 4/41(10%) MONO]. One participant was removed because final HBsAg status could not be confirmed due to a missing follow-up sample. The proportion of DUAL participants with HBsAg <0.05 log10 IU/mL was higher than the proportion of MONO participants with HBsAg <0.05 log10 IU/mL in the cross-sectional analyses throughout study follow-up [Figure 1a]. In a Kaplan-Meier plot, time to HBsAg <0.05 log10 IU/mL was lower in DUAL participants (p=0.02) with HBsAg losses occurring early within the first 24 weeks [Figure 1b].
Figure 1.
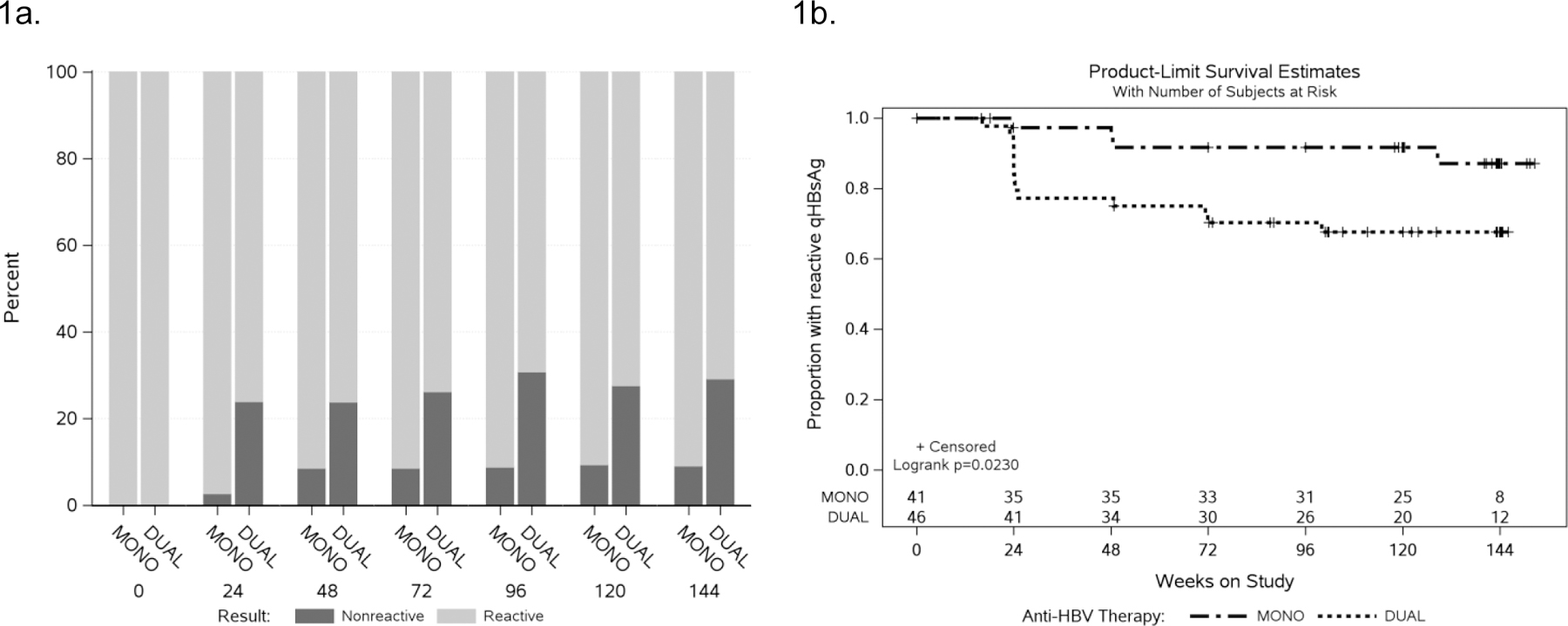
a. HBsAg status by anti-HBV ART regimen
b. Time to qHBsAg loss (<0.05 IU/mL) by anti-HBV ART regimen
Predictors of HBsAg loss
Factors associated with HBsAg <0.05 log10 IU/mL at the univariate level are shown in Table 2. The wide confidence intervals around the estimated odds ratio reflect the small number achieving HBsAg loss. In the final multivariate model using the stepwise selection method, baseline qHBsAg <3 log10 IU/mL, ALT≥ULN, positive HBeAg and HIV RNA ≥5 log10 were all associated with HBsAg<0.05 log10 IU/mL. Use of a DUAL regimen was also associated with higher odds of HBsAg loss, although this became less significant in the multivariate model (p=0.07). Age, sex and HBV RNA were not associated with HBsAg<0.05 log10 IU/mL. Additional analysis was conducted in the subset of participants with baseline qHBsAg ≥ 1 log (n=73). In this subset, Week 24 HBsAg drop from baseline ≥1log10 was significantly associated with subsequent HBsAg <0.05 log10 IU/mL (p= 0.01); other covariates were no longer significant.
Table 2.
Baseline risk factors associated with qHBsAg loss
| Covariate | Univariate OR (95% CI) |
P value | Multivariate OR (95% CI) |
P value |
|---|---|---|---|---|
| Age ≥35 vs.<35 years | 1.23 (0.43, 3.46) | 0.701 | - | - |
| Sex Female vs. Male | 1.54 (0.52, 4.57) | 0.438 | - | - |
| HBV DNA ≥10,000 vs. <10,000 IU/mL |
1.07 (0.37, 3.11) | 0.896 | - | - |
|
qHBsAg
≥3.0 vs <3.0 log10 IU/mL |
0.20 (0.07, 0.62) | 0.005 | 0.06 (<0.01, 0.38) | 0.003 |
| HBV therapy DUAL vs. MONO | 4.05 (1.21, 13.54) | 0.023 | 4.12 (0.87, 19.51) | 0.074 |
| ALT ≥ULN vs. <ULN | 2.74 (0.89, 8.43) | 0.079 | 5.78 (1.16, 28.66) | 0.032 |
| HBeAg positive vs. negative | 2.45 (0.83, 7.28) | 0.106 | 8.79 (1.35, 57.45) | 0.023 |
| CD4 ≥200 vs.<200 cells/mm3 |
0.43 (0.11, 1.63) | 0.213 | - | - |
| HIV RNA ≥5 log10 vs. <5 log10 copies/mL | 6.50 (1.72, 24.52) | 0.006 | 4.76 (1.01, 22.37) | 0.048 |
| HBV RNA ≥3 log10 vs. <3 log10 U/mL | 0.52 (0.18, 1.50) | 0.225 | - |
Based on 87 participants
Changes in HBV RNA by ART regimen
Median baseline HBV RNA was 3.80 (Q1-Q3: 1.78 – 6.60) log10 U/mL and higher in MONO vs. DUAL subjects (4.49 vs. 2.61; p=0.34; stratified by sex, p =0.146). Median HBV RNA at week 144 was 1.80 (Q1-Q3: 0.00 – 4.44, N=46) log10 U/mL. Among 88 participants, three participants (all DUAL) had no detectable HBV RNA (HBV RNA NEG) at baseline, and 17 (19%) had HBV RNA levels <LLoQ but detectable at baseline. Two of 3 participants with NEG HBV RNA at baseline, remained negative throughout the study. Eleven of 17 participants <LLoQ at baseline remained <LLoQ throughout the study or had subsequent missing results. Among the 68 participants with baseline HBV RNA >LLoQ, 32 (47%) [DUAL 21/35 (60%) vs. MONO 11/33 (33%)] achieved HBV RNA <LLoQ or NEG (LLOQ/NEG) on study. Figure 2a shows the proportion of participants achieving HBV RNA <LLoQ (black) and NEG (grey) by treatment at each time point. The proportion achieving HBV RNA <LLoQ/NEG remained higher in DUAL vs. MONO participants throughout follow-up. 23/32 participants who achieved HBV RNA <LLoQ/ NEG, remained <LLoQ/ NEG throughout the study or had missing subsequent HBV RNA results. The remaining 9 participants had HBV RNA fluctuate to above LLoQ during follow-up. In a Kaplan-Meier plot of the time to the first HBV RNA < LLoQ by HBV treatment regimen (DUAL vs. MONO) [Figure 2b], time to HBV <LLoQ was shorter in DUAL participants (p=0.08) with the greatest loss within the first 24 weeks.
Figure 2.

a. HBV RNA status by anti-HBV ART regimen
b. Time to first HBV RNA<LLOQ (<1.65 U/mL) by anti-HBV ART regimen
Changes in HBV DNA by ART regimen
Median baseline HBV DNA was 5.17 log10 IU/mL overall and higher among MONO vs. DUAL participants [5.75 vs. 4.39 log10 IU/mL]. By week 144, 46 (78%) participants achieved HBV DNA<200 IU/mL [26 (90%) DUAL vs. 20 (67%) MONO; p=0.06 unstratified, p=0.04 stratified by sex)]. The proportion of participants with HBV DNA<200 IU/mL all study time points was numerically greater in DUAL vs. MONO groups.
Correlations between qHBsAg, HBV RNA and HBV DNA
Estimated correlations between (i) qHBsAg and HBV RNA and (ii) qHBsAg and HBV DNA and their changes from baseline were all positive (p <0.01 at each study week, MONO and DUAL combined) [Figs 3a/3b]. Levels of qHBsAg correlated more closely with levels of HBV RNA over time than with HBV DNA especially in DUAL participants at weeks 24, 48 and 144. HBV RNA declines also correlated closely with qHBsAg declines in the 18 participants who lost HBsAg. Only 1/18 participant who lost HBsAg did not have an HBV RNA <LLoQ.
Figure 3.

a: Correlations between qHBsAg and HBV RNA changes from baseline
b: Correlations between qHBsAg and HBV DNA changes from baseline
Discussion
These data are some of the first to describe three-year quantitative changes in qHBsAg and HBV RNA after ART initiation in HIV/HBV co-infection. Declines in both markers were observed in response to ART, and higher proportions of participants achieved HBsAg <0.05 log10 IU/mL and HBV RNA <LLoQ with DUAL than MONO therapy.
There is a growing body of literature describing the utility of HBsAg quantification in predicting treatment responses to antivirals and disease progression in patients with chronic HBV [20]. Quantitative serum HBsAg levels have been shown to correlate well with both cccDNA and intrahepatic HBV DNA and are considered a better predictor of overall outcomes with NrtI therapy than serum DNA [21]. HBsAg loss, the most optimal endpoint in HBV, is associated with sustained suppression of HBV replication and viral protein expression even after treatment has been withdrawn. In this study, 21% participants achieved HBsAg <0.05 log10 IU/mL with a significantly higher proportion of loss (30%) observed in participants on DUAL HBV-ART. DUAL therapy also predicted HBsAg <0.05 log10 IU/mL in multivariate models, similar to other studies [2,23]. Several studies have reported similarly high HBsAg sero-clearance rates among HIV/HBV co-infected ranging from 6–19% over 1–5 years [22–28] vs.1–4% in those with HBV alone [29]. Potent antivirals (TDF and entecavir) suppress HBV DNA in HBV mono-infected patients but rarely lead to loss of HBsAg. Mechanisms proposed for the higher HBsAg clearance rates in HIV/HBV co-infection involve ART initiated immune reconstitution. Several observational studies of HIV/HBV co-infection have reported higher rates of HBsAg loss in patients with higher baseline CD4+ T cell counts at the end of follow-up and more robust increases in CD4+ T cell counts after initiation of ART [30,27,31,32]. In this study, we observed an inverse association between HBsAg <0.05 log10 IU/mL and baseline CD4+ T cell count, where participants with CD4 ≥200 were sixty times less likely to achieve HBsAg loss. Although this finding was not significant, we did observe a significant association between HBsAg <0.05 log10 IU/mL and higher baseline HIV viral loads, which in turn, have been shown previously to be a strong predictor of immune reconstitution in HIV studies [33]. Although we were unable to assess time updated CD4+ T cell count as a covariate in a cox regression models because of our small data set, we postulate that both the antiviral effect of dual HBV therapy and augmented immune response in persons with lower baseline CD4+ T cell counts, led to the high seroclearance of HBsAg. Other determinants of HBsAg loss reported in the literature include lower baseline HBsAg and/or early declines in HBsAg [2,34], HBeAg seropositive status [11], African origin [34], longer duration of HBV therapy and higher age of acquisition [35], female sex and lower HBV DNA levels [24], and the postpartum state [36]. We observed similar associations between low qHBsAg at baseline, 24 week ≥1 log drop, HBeAg sero-positive status and HBsAg <0.05 log10 IU/mL in our data.
We report some of the first data on changes in HBV RNA levels in HIV/HBV co-infected participants. HBV RNA is an emerging marker of interest in patients with chronic hepatitis B and thought to reflect viral replication and HBV activity more accurately. We observed baseline median HBV RNA levels that were lower than HBV DNA at study entry. Of note, lower levels of HBV RNA have been observed among HBeAg negative vs. positive patients [12] who accounted for half of all patients in our study cohort. The frequent measurements of HBV RNA allowed us to make important observations about the dynamics of HBV RNA after therapy. Notably, despite overall declines on treatment, more variation was seen in HBV RNA levels compared to HBV DNA. 9/23 (39%) participants who achieved <LLoQ/NEG on study did not remain undetectable during follow-up. Seven/20 participants who had either HBV RNA NEG or < LLoQ at baseline, also became detectable while on treatment. All but two of these participants (both with baseline HBV RNA LLoQ/NEG) achieved sustained HBV DNA virologic suppression during follow-up [data not shown]. The lack of direct activity of NrtIs on generation of HBV pgRNA virions from cccDNA may account for some of this RNA variability after treatment [8]. Although RNA has been shown to be of limited additional value to HBV DNA in untreated patients [37], it has been shown to be of value in patients on NrtIs, declining similarly to HBV DNA after treatment initiation and importantly predicting viral rebound when treatment is stopped [8,19, 38,39]. In addition, our data would suggest HBV RNA may have some value in determining ongoing viral activity where HBV DNA is negative.
Studies have also shown a close correlation between HBV RNA and HBsAg, and in one study, HBV RNA was a significant predictor of HBsAg loss [40,41]. In our study, qHBsAg correlated well with both HBV RNA (in all patients and those who lost HBsAg), with stronger correlations observed than those between qHBsAg and DNA especially in the first 48 weeks and among all DUAL participants. The weaker correlation with HBV DNA may be due to the underlying mechanisms of NrtIs which inhibit DNA polymerase but have no effect on other steps of viral replication, transcriptional activity of cccDNA and HBsAg production. HBV RNA was not a significant predictor of HBsAg <0.05 log10 IU/mL in multivariate analyses, however this could have been due to small numbers. In HIV/HBV co-infection, HBV RNA measurement may be of additional benefit in treatment monitoring in patients on HBV active-ART, more reliably assessing anti-HBV efficacy, and perhaps better predicting functional cure and helping to inform decision-making about ART switch or the addition of novel HBV therapeutic agents.
Strengths of the study included the frequent measurements of serum markers during follow-up, allowing us to assess dynamic changes early after ART initiation in this cohort. We also present the first direct comparison of HBsAg, HBV RNA, and HBV DNA and strength of association between these markers in HIV/HBV co-infected. An important limitation is that this study was not a randomized controlled trial and differences in baseline characteristics may have affected the results. Notably, qHBsAg levels were lower in DUAL participants (and a higher proportion of DUAL participants were women) which could have contributed to the finding of higher proportions of HBsAg loss within this group. In this study we did not measure intrahepatic cccDNA, so we were not able to determine how accurately qHBsAg and HBV RNA reflect hepatic cccDNA transcription activity in the presence of NrtIs. Although several studies have reported good correlations between qHBsAg and declines in intrahepatic cccDNA in HBV mono-infected after treatment, stronger correlations have been reported between HBcrAg and cccDNA which was also not measured in this study [42]. This may be in part due to expression of HBsAg from other sources such as integrated HBV DNA that is unaffected by suppression of HBV virus [43]. The data on HBV RNA is less consistent with some reports of stronger correlations between other markers (HBsAg, HBcrAg) and cccDNA or no correlation at all [8,42,44]; however, studies differ with respect to samples sizes, stage of infection and type of treatment. In this study, we did not examine associations between markers and HBeAg seroconversion. HBV RNA levels in particular have been shown to decline more rapidly among HBeAg positive patients with seroconversion compared to those without [43]. In this same study HBV RNA was more closely correlated with cccDNA in these individuals. Other limitations include the relatively small sample size and use of older ART regimens.
In conclusion, our study supports current recommendations of TDF/FTC containing HBV-ART for initial use in HIV/HBV co-infection. Our results also suggest HBV RNA could be a useful additional marker of treatment response in HIV/HBV co-infected participants. HBsAg and HBV RNA could be used to determine which co-infected patients could switch to more simplified ART on benefit from newer HBV regimens in the future, by better predicting functional cure. Larger clinical trials are needed to validate this however, as well as more studies assessing the role of these and other novel markers in predicting treatment outcomes and disease monitoring in HIV/HBV co-infection.
Acknowledgments
We are grateful to the staff and clients that participated in the ACTG 5175 (PEARLS) and A5208 (OCTANE) trials in Botswana, Haiti, Kenya, Malawi, Peru, South Africa, Thailand, United States and Zimbabwe and the contributions of Frederick Sawe (US Military HIV Research Program CTU, Kenya), Cynthia Riviere and Patrice Severe (Haiti CRS). The authors acknowledge the support of the ACTG network and the scientific committees for their contribution during protocol development and study conduct. The authors also acknowledge the following study team members for their contributions; laboratory data manager Kyle Whitson; Network Coordinating Center (NCC) Research Assistant Amy Dominguez and Triin Umbleja at the Statistical and Data Analysis Center (SDAC).
Sources of Funding:
Research reported in this publication was supported by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under Award Number UM1 AI068634, UM1 AI068636 and UM1 AI106701. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Footnotes
Conflicts of Interest:
GC, MK, VH, and MA are employees and shareholders of Abbott Laboratories.
MP is a Consultant to Antios and Aligos and spouse of an employee at Hoffman-La Roche
This study has been previously presented at the 2021 Virtual Annual ACTG Network Meeting June 16th, 2021, USA.
References
- 1.Matthews PC, Geretti AM, Goulder PJ, Klenerman P. Epidemiology and impact of HIV coinfection with hepatitis B and hepatitis C viruses in Sub-Saharan Africa. J Clin Virol 2014; 61:20–33. [DOI] [PubMed] [Google Scholar]
- 2.Singh KP, Crane M, Audsley J, Avihingsanon A, Sasadeusz J, Lewin SR. HIV-hepatitis B virus coinfection: epidemiology, pathogenesis, and treatment. AIDS 2017; 31:2035–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ou Q, Guo J, Zeng Y, Chen H. Insights for clinical diagnostic indicators of virus and host in chronic hepatitis B infection. J Viral Hepat 2020; 27:224–32. [DOI] [PubMed] [Google Scholar]
- 4.Thompson AJ, Nguyen T, Iser D, Ayres A, Jackson K, Littlejohn M et al. Serum hepatitis B surface antigen and hepatitis B e antigen titers: disease phase influences correlation with viral load and intrahepatic hepatitis B virus markers. Hepatology 2010; 51:1933–44. [DOI] [PubMed] [Google Scholar]
- 5.Giersch K, Allweiss L, Volz T, Dandri M, Lutgehetmann M. Serum HBV pgRNA as a clinical marker for cccDNA activity. J Hepatol 2017; 66:454–467. [DOI] [PubMed] [Google Scholar]
- 6.Chan HL, Wong VW, Wong GL, Tse CH, Chan HY, Sung JJ. A longitudinal study on the natural history of serum hepatitis B surface antigen changes in chronic hepatitis B. Hepatology 2010; 52:1232–41. [DOI] [PubMed] [Google Scholar]
- 7.Ungtrakul T, Sriprayoon T, Kusuman P, Chunnuan P, Soonklang K, Sornsamdang G, et al. Role of quantitative hepatitis B surface antigen in predicting inactive carriers and HBsAg seroclearance in HBeAg-negative chronic hepatitis B patients. Medicine (Baltimore) 2017; 96:e6554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Wang J, Shen T, Huang X, Kumar GR, Chen X, Zeng Z, et al. Serum hepatitis B virus RNA is encapsidated pregenome RNA that may be associated with persistence of viral infection and rebound. J Hepatol 2016; 65:700–10. [DOI] [PubMed] [Google Scholar]
- 9.Anderson M, Gersch J, Luk KC, Dawson G, Carey I, Agarwal K, et al. Circulating Pregenomic Hepatitis B Virus RNA Is Primarily Full-length in Chronic Hepatitis B Patients Undergoing Nucleos(t)ide Therapy. Clin Infectious Dis 2021; 72:2029–2031 [DOI] [PubMed] [Google Scholar]
- 10.Wang CC, Tseng KC, Hsieh TY, Tseng TC, Lin HH, Kao JH. Assessing the Durability of Entecavir-Treated Hepatitis B Using Quantitative HBsAg. Am J Gastroenterol 2016; 111:1286–94 [DOI] [PubMed] [Google Scholar]
- 11.Mak LY, Seto WK, Fung J, Yuen MF. Use of HBsAg quantification in the natural history and treatment of chronic hepatitis B. Hepatol Int 2020; 14:35–46 [DOI] [PubMed] [Google Scholar]
- 12.Huang YW, Takahashi S, Tsuge M, et al. On-treatment low serum HBV RNA level predicts initial virological response in chronic hepatitis B patients receiving nucleoside analogue therapy. Antivir Ther 2015; 20:369–75. [DOI] [PubMed] [Google Scholar]
- 13.Tsuge M, Murakami E, Imamura M, et al. Serum HBV RNA and HBeAg are useful markers for the safe discontinuation of nucleotide analogue treatments in chronic hepatitis B patients. J Gastroenterol 2013; 48:1188–204 [DOI] [PubMed] [Google Scholar]
- 14.Papatheodoridi M, Hadziyannis E, Berby F, Kalliopi Z, Testoni B, Rigopoulou E, et al. Predictors of HBsAg Loss, Relapse and Retreatment after Discontinuation of Effective Oral Antiviral Therapy in Non-Cirrhotic HBeAg-Negative Chronic Hepatitis B. J. Viral Hepat 2020; 27:118–126 [DOI] [PubMed] [Google Scholar]
- 15.Vitoria M, Rangaraj A, Ford N, Doherty M. Current and future priorities for the development of optimal HIV drugs. Curr Opin HIV AIDS 2019; 14:143–9. [DOI] [PubMed] [Google Scholar]
- 16.Campbell TB, Smeaton LM, Kumarasamy N, Flanigan T, Klingman K, Firnhaber C et al. Efficacy and safety of three antiretroviral regimens for initial treatment of HIV-1: a randomized clinical trial in diverse multinational settings. PLoS Med 2012; 9:e1001290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lockman S, Hughes MD, McIntyre J, Zheng Y, Chipato T, Conradie F, et al. Antiretroviral therapies in women after single-dose nevirapine exposure. N Engl J Med 2010; 363:1499–509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Thio CL, Smeaton L, Hollabaugh K, Saulnyas M, Hwang H, Saravanan S et al. Comparison of HBV-active HAART regimens in an HIV-HBV multinational cohort: outcomes through 144 weeks. AIDS 2015; 29:1173–82 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Butler EK, Gersch J, McNamara A, Luk KC, Holzmayer V, de Medina M, et al. Hepatitis B Virus Serum DNA and RNA Levels in Nucleos(t)ide Analog-Treated or Untreated Patients During Chronic and Acute Infection. Hepatology 2018; 68:2106–17. [DOI] [PubMed] [Google Scholar]
- 20.Tseng TC, Kao JH. Clinical utility of quantitative HBsAg in natural history and nucleos(t)ide analogue treatment of chronic hepatitis B: new trick of old dog. J Gastroenterol 2013; 48:13–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Chan HL, Wong VW, Tse AM, Chi-Hang Tse, Chim AM, Chan HY et al. Serum HBsAg levels correlate well with the cccDNA and intrahepatic HBV DNA. Low pretreatment HBsAg is better than HBV DNA to predict good response to peginterferon and lamivudine treatment. Clin Gastroenterol Hepatol 2007; 5:1462–8.18054753 [Google Scholar]
- 22.Boyd A, Gozlan J, Miailhes P, Lascoux-Combe C, Cam MS, Rougier H et al. Rates and determinants of hepatitis B ‘e’ antigen and hepatitis B surface antigen seroclearance during long-term follow-up of patients coinfected with HIV and hepatitis B virus. AIDS 2015; 29:1963–73. [DOI] [PubMed] [Google Scholar]
- 23.Psevdos G Jr., Kim JH, Suh JS, Sharp VL. Predictors of loss of hepatitis B surface antigen in HIV-infected patients. World J Gastroenterol 2010; 16:1093–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Vinikoor MJ, Sinkala E, Chilengi R, Mulenga LB, Chi BH, Zyambo Z et al. Impact of Antiretroviral Therapy on Liver Fibrosis Among Human Immunodeficiency Virus-Infected Adults With and Without HBV Coinfection in Zambia. Clin Infect Dis 2017; 64:1343–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Audsley J, Avihingsanon A, Littlejohn M, Bowden S, Matthews G, Fairley C et al. Long-Term TDF-Inclusive ART and Progressive Rates of HBsAg Loss in HIV-HBV Coinfection-Lessons for Functional HBV Cure? J Acquir Immune Defic Syndr 2020; 84:527–33. [DOI] [PubMed] [Google Scholar]
- 26.Martin-Carbonero L, Teixeira T, Poveda E, Plaza Z, Vispo E, Gonzalez-Lahoz J et al. Clinical and virological outcomes in HIV-infected patients with chronic hepatitis B on long-term nucleos(t)ide analogues. AIDS 2011; 25:73–9. [DOI] [PubMed] [Google Scholar]
- 27.Huang AJ, Nunez M. Outcomes in HIV/HBV-Coinfected Patients in the Tenofovir Era Are Greatly Affected by Immune Suppression. J Int Assoc Provid AIDS Care 2015; 14:360–8. [DOI] [PubMed] [Google Scholar]
- 28.Hamers RL, Zaaijer HL, Wallis CL, Siwale M, Ive P, Botes ME, et al. HIV-HBV coinfection in Southern Africa and the effect of lamivudine- versus tenofovir-containing cART on HBV outcomes. J Acquir Immune Defic Syndr 2013; 64:174–82 [DOI] [PubMed] [Google Scholar]
- 29.Yeo YH, Ho HJ, Yang HI, Tseng TC, Hosaka T, Trinh HN, et al. Factors Associated With Rates of HBsAg Seroclearance in Adults With Chronic HBV Infection: A Systematic Review and Meta-analysis. Gastroenterology 2019; 156:635–46 e9. [DOI] [PubMed] [Google Scholar]
- 30.Jain MK, Vigil KJ, Parisot P, Go G, Vu T, Li X, et al. Incidence and Predictors of Hepatitis B Surface Antigen Clearance in HIV Patients: A Retrospective Multisite Study. Open Forum Infect Dis 2021;8:ofab116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Arendt E, Jaroszewicz J, Rockstroh J, Olson DM, Zacher BJ, Mederack I, et al. Improved immune status corresponds with long-term decline of quantitative serum hepatitis B surface antigen in HBV/HIV co-infected patients. Viral Immunol 2012; 25:442–7. [DOI] [PubMed] [Google Scholar]
- 32.Jaroszewicz J, Reiberger T, Meyer-Olson D, Mauss S, Vogel M, Ingiliz P, et al. Hepatitis B surface antigen concentrations in patients with HIV/HBV co-infection. PLoS One 2012; 7:e43143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Novak RM, Richardson JT, Buchacz K, Chmiel JS, Durham MD, Palella FJ et al. Immune reconstitution inflammatory syndrome: incidence and implications for mortality. AIDS 2012; 26:721–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gantner P, Cotte L, Allavena C, Firouze BS, Huleux T, Duvivier C et al. Higher rates of HBsAg clearance with tenofovir-containing therapy in HBV/HIV co-infection. PLoS One 2019;14:e0215464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Chen CH, Lu SN, Hung CH, Wang JH, Hu TH, Changchien CS et al. The role of hepatitis B surface antigen quantification in predicting HBsAg loss and HBV relapse after discontinuation of lamivudine treatment. J Hepatol 2014; 61:515–22. [DOI] [PubMed] [Google Scholar]
- 36.Joshi SS, Coffin CS. Hepatitis B and Pregnancy: Virologic and immunologic characteristics. Hepatology Communications 2020; 4:157–171 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang J, Yu Y, Li G, Shen C, Li J, Chen S. Natural history of serum HBV-RNA in chronic HBV infection. J Viral Hepat 2018; 25:1038–47. [DOI] [PubMed] [Google Scholar]
- 38.Fan R, Zhou B, Xu M, Tan D, Niu J, Wang H et al. Association Between Negative Results From Tests for HBV DNA and RNA and Durability of Response After Discontinuation of Nucles(t)ide Analogue Therapy. Clin Gastroenterol Hepatol 2020; 18:719–27 e7. [DOI] [PubMed] [Google Scholar]
- 39.van Bommel F, Bartens A, Mysickova A, Hofmann J, Krüger DH, Berg T, et al. Serum hepatitis B virus RNA levels as an early predictor of hepatitis B envelope antigen seroconversion during treatment with polymerase inhibitors. Hepatology 2015; 61:66–76. [DOI] [PubMed] [Google Scholar]
- 40.Lim SG, Phyo WW, Ling JZJ, Cloherty G, Butler E, Kuhns M, et al. Comparative biomarkers for HBsAg loss with antiviral therapy shows dominant influence of quantitative HBsAg (qHBsAg). Aliment Pharmacol Ther 2021; 53:172–82. [DOI] [PubMed] [Google Scholar]
- 41.Farag MS, van Campenhout MJH, Pfefferkorn M, Fishcer J, Deichsel D, Boonstra A et al. Hepatitis B Virus RNA as Early Predictor for Response to Pegylated Interferon Alpha in HBeAg-Negative Chronic Hepatitis B. Clin Infect Dis 2021; 72:202–11. [DOI] [PubMed] [Google Scholar]
- 42.Gao Y, Li Y, Meng Q, Zhang Z, Zhao P, Shang Q et al. Serum Hepatitis B Virus DNA, RNA, and HBsAg: Which Correlated Better with Intrahepatic Covalently Closed Circular DNA before and after Nucleos(t)ide Analogue Treatment? J Clin Microbiol 2017; 55:2972–82 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kim JH, Ghosh A, Ayithan N, Romani S, Khanam A, Park JJ, et al. Circulating serum HBsAg level is a biomarker for HBV-specific T and B cell responses in chronic hepatitis B patients. Sci Rep 2020; 10:5947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Chen EQ, Wang ML, Tao YC, Wu DB, Liao J, He M, et al. Serum HBcrAg is better than HBV RNA and HBsAg in reflecting intrahepatic covalently closed circular DNA. J Viral Hepat 2019; 26:586–95 [DOI] [PubMed] [Google Scholar]


