Abstract
In two published reports using monoclonal antibodies (MAbs) generated against whole cells, Olsen et al. showed that strain-specific antigens on the surface of cultured cells of Sinorhizobium meliloti were diminished or absent in the endophytic cells (bacteroids) recovered from alfalfa nodules, whereas two common antigens were not affected by bacterial differentiation (P. Olsen, M. Collins, and W. Rice, Can. J. Microbiol. 38:506–509, 1992; P. Olsen, S. Wright, M. Collins, and W. Rice, Appl. Environ. Microbiol. 60:654–661, 1994). The nature of the antigens (i.e., the MAb epitopes), however, were not determined in those studies. For this report, the epitopes for five of the anti-S. meliloti MAbs were identified by polyacrylamide gel electrophoresis-immunoblot analyses of the polysaccharides extracted from S. meliloti and Sinorhizobium fredii. This showed that the strain-specific MAbs recognized K antigens, whereas the strain-cross-reactive MAbs recognized the lipopolysaccharide (LPS) core. The MAbs were then used in the analysis of the LPS and K antigens extracted from S. meliloti bacteroids, which had been recovered from the root nodules of alfalfa, and the results supported the findings of Olsen et al. The size range of the K antigens from bacteroids of S. meliloti NRG247 on polyacrylamide gels was altered, and the epitope was greatly diminished in abundance compared to those from the cultured cells, and no K antigens were detected in the S. meliloti NRG185 bacteroid extract. In contrast to the K antigens, the LPS core appeared to be similar in both cultured cells and bacteroids, although a higher proportion of the LPS fractionated into the organic phase during the phenol-water extraction of the bacteroid polysaccharides. Importantly, immunoblot analysis with an anti-LPS MAb showed that smooth LPS production was modified in the bacteroids.
Gram-negative bacteria of the family Rhizobiaceae participate in a mutualistic symbiosis with legumes. The infection process is initiated by an exchange of signal molecules in the form of plant-derived flavonoids and bacterial Nod factors (5). In the course of infection, the bacteria undergo morphological changes, which result in the inclusion of highly differentiated cells, termed bacteroids, in the root nodules of the host plant. Although there is little information available on specific changes in the cell surface chemistry of Sinorhizobium spp. during infection and differentiation, Olsen et al. (10, 11) used monoclonal antibodies (MAbs) in enzyme-linked immunosorbent assays (ELISAs) and immunofluorescence studies of whole cells to show that unidentified strain-specific antigens on the surface of cultured cells of Sinorhizobium meliloti were diminished or absent in bacteroids recovered from alfalfa nodules. In contrast, certain common antigens were not affected by bacterial differentiation. In this study, we determined the nature of the antigens and used the MAbs in analysis of bacteroid extracts.
A recent report showed that capsular polysaccharide (K antigens) and lipopolysaccharide (LPS) are important surface antigens of Sinorhizobium spp. (16). S. meliloti and Sinorhizobium fredii typically produce two forms of LPS: rough LPS (R-LPS), which consists of a lipid A membrane anchor and conserved core oligosaccharides, and smooth LPS (S-LPS), which includes the O antigen (or O polysaccharide), and past studies have shown that the core oligosaccharides are structurally similar in both the R-LPS and the S-LPS of Sinorhizobium spp. (17). There is limited variation in O-polysaccharide structure among Sinorhizobium strains, and when present, the S-LPS migrate as two or three distinct bands in polyacrylamide electrophoresis (PAGE) analyses. Characterization of two forms of S-LPS from S. fredii USDA205 showed that the primary O antigen is a glucan and a secondary O antigen is a xylomannan (17). In this regard, Sinorhizobium spp. are unusual, as the O antigens of most gram-negative bacteria are highly variable, strain-specific surface antigens (19); in this genus, that role is fulfilled by the K antigens.
The K antigens of Sinorhizobium spp. are major strain-specific antigens, which commonly consist of small repeating units of a hexose and 1-carboxy-2-keto-3-deoxy sugars, such as sialic acid or 3-deoxy-d-manno-2-octulosonic acid (Kdo), although they vary in glycosyl residue composition, substitution patterns, linkage points, anomeric configuration, and size range (6, 14–16). The K antigens of rhizobia do not possess a lipid anchor and can be separated from the LPS by preparative PAGE, based on the presence of the lipid A moiety on the LPS (8). The LPS and K antigens can also be identified in PAGE analyses by differential staining and by the use of nondetergent gels (8, 9, 15). Past studies have shown that the K antigens, as produced by cultured cells, may be involved in an early recognition step in S. meliloti-alfalfa interactions (1) and that K-antigen production is affected by plant-derived signals (17, 18). This suggests that K-antigen expression is tightly regulated by the bacterium and that it may be modified in the endophytic cell.
We employed PAGE-immunoblot analyses of extracted polysaccharides to identify the epitopes for a panel of monoclonal antibodies (MAbs) that were generated against whole cells of S. meliloti by Olsen et al. (10, 11). We found that three strain-specific MAbs recognized the K antigens of the homologous strains and that two strain-cross-reactive MAbs recognized the LPS core. Three of the MAbs were then used in the analysis of the polysaccharides extracted from bacteroids of S. meliloti NRG247 and S. meliloti NRG185, which were recovered from alfalfa nodules. The results showed that the K antigens produced by the S. meliloti NRG247 bacteroids were greatly diminished in abundance and had altered mobilities on polyacrylamide gels, and no K antigens were detected in the polysaccharide preparation from S. meliloti NRG185 bacteroid. In contrast, the LPS core production did not appear to be significantly modified in the endophytic bacteria, although the S. meliloti NRG185 bacteroids were shown to produce distinct forms of S-LPS.
Note that the terms “strain-specific” and “strain-cross-reactive” were used in the previous reports (10, 11), so they are used in this report. However, these are relative descriptions, as the strain-specific MAbs recognize a limited number of other strains, and the strain-cross-reactive MAbs recognize most but not all S. meliloti strains.
Epitope identification for the anti-S. meliloti MAbs.
The strains used in this study are described in Table 1. Cells were stored at −70°C in 7.5% glycerol and cultured in tryptone-yeast extract broth (S. meliloti) or yeast extract-mannitol (S. fredii) at 24°C, as previously described (15, 17). Prior to immunoblot analysis, the polysaccharides extracted from the cultured cells of each strain were analyzed by deoxycholic acid-PAGE, using 18% acrylamide gels, as previously described in detail (16), to determine the relative mobilities of the LPS and K antigens (data not shown). The cell-associated polysaccharides from cell pellets of 3-ml cultures were extracted by a mini-phenol-water method (16); the cultures were grown to the same point (optical density at 600 nm [OD600], 0.8 to 1.0), and the polysaccharide preparations were dissolved in identical volumes of PAGE sample buffer. The identity of each bacterial product (R-LPS, S-LPS, and K antigens) on polyacrylamide gels has been unequivocally established in past studies (6, 9, 12, 14–18). The R-LPS migrates as a disperse banding region, consisting of multiple bands (due to heterogeneity in the core), and the S-LPS migrates as two or three low-mobility bands (16). In contrast, the K antigens, which are polymers of conserved repeating units, migrate in distinct ladder patterns (6, 15, 16). The PAGE migration patterns of each component were also verified for this study by differential staining. Only the LPS appeared on silver-stained gels, and a specific Alcian blue-silver stain procedure that omits the oxidation step visualized only the K antigens (3, 15). Furthermore, only the K antigens, which lack a lipid moiety, exhibited normal migration in nondetergent polyacrylamide gels, whereas the LPS appeared as a smear at the top of the lane (data not shown).
TABLE 1.
Bacterial strains used in this study
| Straina | Characteristics and origin | Source |
|---|---|---|
| Sinorhizobium meliloti | ||
| Rm41 | Wild type from Medicago sativa; Europe | Bánfalvi et al. |
| Rm1021 | Str-resistant derivative of wild-type strain RCR2011 | Walker et al. |
| NRG23 | Wild type from M. sativa; North America | Olsen et al. |
| NRG34 | Wild type from M. sativa; North America | Philom Bios |
| NRG43 | Wild type from M. sativa; North America | Olsen et al. |
| NRG53 | Wild type from M. sativa; North America | Olsen et al. |
| NRG133 | Wild type from M. sativa; North America | Olsen et al. |
| NRG185 | Wild type from M. sativa; North America | Olsen et al. |
| NRG247 (Balsac)b | Wild type from M. sativa; North America | Olsen et al. |
| NRG282 | Wild type from M. sativa; North America | Olsen et al. |
| NRG286 | Wild type from M. sativa; North America | Olsen et al. |
| Sinorhizobium sp. strain NGR234 | Wild type from Lablab purpureus; New Guinea | Trinick |
| Sinorhizobium fredii | ||
| HH103 | Wild type from soil; Hubei province, China | Dowdle and Bohlool |
| HH303 | Wild type from soil; Hubei province, China | Dowdle and Bohlool |
| USDA191 | Wild type from soil; Shanghai province, China | Keyser et al. |
| USDA192 | Wild type from Glycine soja; China | Keyser et al. |
| USDA196 | Wild type from soil; Hubei province, China | Dowdle and Bohlool |
| USDA197 | Wild type from soil; Hubei province, China | Dowdle and Bohlool |
| USDA201 | Wild type from Glycine max cv. Peking; China | Keyser et al. |
| USDA205 | Wild type from G. max cv. Peking; China | Keyser et al. |
| USDA208 | Wild type from G. max cv. Peking; China | Keyser et al. |
| USDA257 | Wild type from G. max cv. Peking; China | Keyser et al. |
S. meliloti NRG34, S. meliloti NRG185, and S. meliloti NRG247 (Balsac) were included in this study because the MAbs were raised against whole cells of these strains (10) (see Table 2). S. meliloti NRG23, NRG43, NRG53, NRG133, NRG286, and NRG289 were chosen because they were negative for MAb 10 in the previous study (11), which was the exception. S. meliloti Rm1021, S. meliloti Rm41, and S. fredii USDA257 were used because they are the most commonly studied microsymbionts in serogroup A. S. fredii USDA205 and Sinorhizobium sp. strain NGR234 were analyzed because they are in serogroup B.
S. meliloti NRG247 is referred to as Balsac in the report of Olsen et al. (11).
Initial immunoblot analyses were performed on the extracts from a subgroup of S. meliloti strains used in the original work of Olsen et al. (11) (Table 1). The LPS and K antigens were separated on polyacrylamide gels and blotted to Nytran+ membranes (Schleicher and Schuell, Keene, N.H.) with a Trans-Blot SD apparatus (Bio-Rad), as previously described (16). Individual strips (lanes) were probed with each of five MAbs (Table 2), which were provided by Perry Olsen (Agriculture Canada, Beaverlodge, Alberta, Canada). Examples of the immunoblots are shown in Fig. 1 (not all negative results are shown). The MAbs were delineated into two classes: (i) the strain-specific MAbs that recognized the K antigens (MAbs 7 to 9) and (ii) the strain-cross-reactive MAbs that recognized LPS (MAbs 10 and 11). We were unable to determine the epitope for a third strain-cross-reactive MAb (MAb 6) described in the report of Olsen et al. (11).
TABLE 2.
MAbs used in this study
| MAba | Descriptionb | Specificityc |
|---|---|---|
| MAb 6 | Anti-S. meliloti NRG247 (Balsac) IgG3 (F2P18E) | Strain-cross-reactive |
| MAb 7 | Anti-S. meliloti NRG34 IgM (F12P92F) | Strain-specific |
| MAb 8 | Anti-S. meliloti NRG185 IgM (F4P26B) | Strain-specific |
| MAb 9 | Anti-S. meliloti NRG247 (Balsac) IgM (F12P92F) | Strain-specific |
| MAb 10 | Anti-S. meliloti NRG34 IgG3 (F12P64C) | Strain-cross-reactive |
| MAb 11 | Anti-S. meliloti NRG185 IgM (F3P49C) | Strain-cross-reactive |
All MAbs used in this study were obtained from P. Olsen and are described in detail elsewhere (11). The numerical designations are those used by Olsen et al. (11).
Each MAb was raised against whole cells of the designated strains. IgG3, immunoglobulin G3.
The terms “strain-cross-reactive” and “strain-specific” were used in the previous report (11).
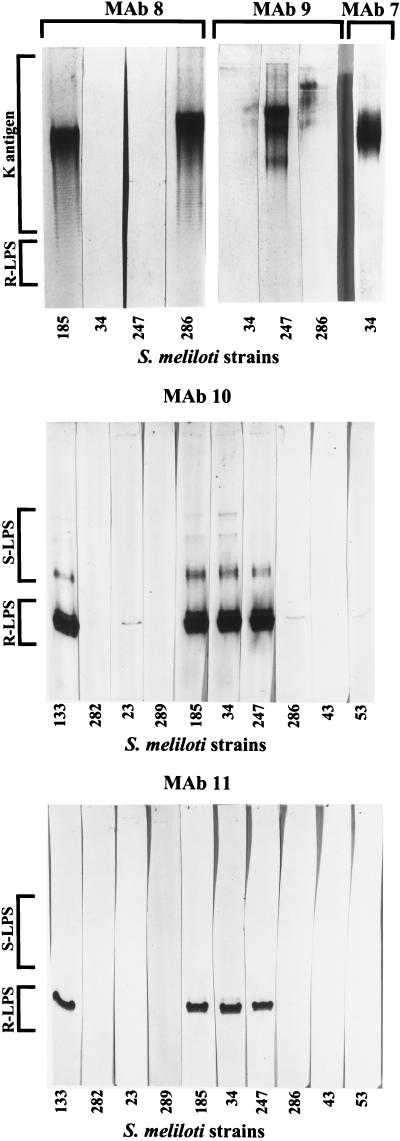
FIG. 1.
Immunoblot analysis of the polysaccharide extracts from cultured cells of S. meliloti. The immunoblots were probed with the MAbs shown in Table 2. The MAb designations are given above the lanes. The strains are identified in abbreviated form beneath the lane as follows: 23, S. meliloti NRG23; 34, S. meliloti NRG34; 43, S. meliloti NRG43; 53, S. meliloti NRG53; 133, S. meliloti NRG133; 185, S. meliloti NRG185; 247, S. meliloti NRG247; 282, S. meliloti NRG282; 286, S. meliloti NRG286; 289, S. meliloti NRG289. The R-LPS, S-LPS, and K-antigen labels identify the antigens bound by the MAbs (the S-LPS and K antigens comigrate on polyacrylamide gels, so those regions overlap). Only a few examples of negative results are shown.
Anti-K antigen MAbs.
MAbs 7, 8, and 9 each recognized a ladder pattern on the immunoblots, which is due to the K antigens (6, 14–18). Of the strains tested in this study, MAbs 7 and 9 were positive only with the K antigens from the homologous strains (i.e., the strains used to generate the MAbs), S. meliloti NRG34 and S. meliloti NRG247, respectively (Fig. 1). Although there was a marginal reaction of MAb 9 with S. meliloti NRG286, this was probably not due to the primary K antigen, as the lane was stained above the K antigen (compare MAb 9 with MAb 8 [Fig. 1]). MAb 8 recognized the K antigens from both the homologous strain, S. meliloti NRG185, and S. meliloti NRG286, which is in agreement with the results of Olsen et al. (11).
In the ELISA of untreated cells performed by Olsen et al. (11), MAb 7 was also positive only with the homologous strain of the 60 strains tested, whereas MAbs 8 and 9 each recognized 19 of 60 strains; however, this included four strains that were recognized by both MAbs 8 and 9 (11). This observation is interesting, as the structures of the primary K antigens from S. meliloti NRG185 and S. meliloti NRG247 have been determined, and they are quite different (16). The K antigen from S. meliloti NRG247 consists of glucose and sialic acid [-α-Glc→α-NeuNAc-]n and that of S. meliloti NRG185 contains N-acetylglucosamine and Kdo [-β-GlcNAc→β-Kdo-]n. Therefore, the cross-reactivity of MAbs 8 and 9 with some strains must be due to common structural features or similar secondary K antigens, not identical primary K antigens.
Anti-LPS MAbs.
Strain-cross-reactive MAb 10 appeared to recognize several forms of R-LPS, as shown by the disperse signal in the R-LPS banding region, as well as the S-LPS from the same strains (Fig. 1), showing that the epitope does not include the O-antigen linkage point. Interestingly, the presence and abundance of the highest-mobility S-LPS band (near the lower end of the region labeled S-LPS in Fig. 1) vary from preparation to preparation, and it often does not appear at all in Alcian blue-silver stained polyacrylamide gels (see reference 16). In this study, the band was much fainter than the lower-mobility S-LPS bands on polyacrylamide gels (data not shown) but yielded a much stronger signal in the immunoblot (Fig. 1).
In contrast to MAb 10, MAb 11 recognized only one major form (band) of R-LPS from the serogroup A strains, and it did not bind the S-LPS from any strain. The latter phenomenon was not due to a lower titer or weaker signal, as overloaded blots, more-concentrated antibody, and longer blot development yielded much more signal for the R-LPS and still no binding of the S-LPS (data not shown). In addition, MAb 11 had negative reactions with S. meliloti NRG23, S. meliloti NRG53, and S. meliloti NRG286, whereas MAb 10 yielded a marginal reaction with the R-LPS from all three strains.
A second series of analyses was performed with MAbs 10 and 11, using S. meliloti strains from other sources (listed in Table 1) as well as several S. fredii strains and Sinorhizobium sp. strain NGR234 (data not shown). The results of all immunoblot analyses of LPS are summarized in Table 3. We found that MAbs 10 and 11 were positive with the R-LPS from all strains previously shown to be positive with anti-Rm41 polyclonal antiserum (16), including many strains of S. fredii, and negative with all LPS recognized by anti-Rf205 antiserum (16). Thus, at least four LPS core serogroups have been identified in the genus (11, 16; this report). Although Table 3 includes only those strains that have been tested with all MAbs and antisera, if the results of the ELISAs from Olsen et al. (11) are included, 61 of 73 (84%) strains tested fall into serogroup A, 4 strains are in serogroup B (5%), 3 strains are in serogroup C (4%), and the remainder are untyped.
TABLE 3.
Antibody specificity for LPS core serogroups of Sinorhizobium spp.a
| LPS core serogroup | Immunoblot response of LPSb | MAb(s) or antiserumc | Sinorhizobium strains |
|---|---|---|---|
| Serogroup Ad | + | MAbs 10 and 11 | S. meliloti strains (Rm41, Rm1021, NRG34, NRG133, NRG185, and NRG247) and S. fredii strains (USDA192, USDA196, USDA197, USDA201, USDA257, and HH103) |
| + | Anti-Rm41 | ||
| − | Anti-Rf205 | ||
| Serogroup B | − | MAbs 10 and 11 | S. fredii USDA191, S. fredii USDA205, S. fredii USDA208, and Sinorhizobium sp. strain NGR234 |
| − | Anti-Rm41 | ||
| + | Anti-Rf205 | ||
| Serogroup C | +/− | MAb 10 | S. meliloti NRG23, S. meliloti NRG53, and S. meliloti NRG286 |
| − | MAb 11 | ||
| +/− | Anti-Rm41 | ||
| − | Anti-Rf205 | ||
| Untyped | − | All MAbs and antisera | S. fredii HH303, S. meliloti NRG43, S. meliloti NRG282, and S. meliloti NRG289 |
The LPS serogroup designations are a result of this study and have not been used previously.
Based on degree of reaction with the R-LPS (Fig. 1). Symbols: +, positive; +/−, marginal; −, negative.
Specificity was determined by immunoblot analyses performed with MAb 10 and MAb 11 (Table 2) in this study or with polyclonal antisera in a earlier report (16). Anti-Rm41, polyclonal antiserum raised against whole cells of S. meliloti Rm41; Anti-Rf205, polyclonal antiserum raised against whole cells of S. fredii USDA205.
Analysis of the K antigens and LPS from bacteroids.
Three of the MAbs were then used in an analysis of polysaccharide preparations from bacteroids of S. meliloti NRG247 and S. meliloti NRG185. For this analysis, alfalfa (Medicago sativa cv. Apollo) seeds were sprouted in sterile 15-cm-diameter plastic pots on 5-cm-deep coarse horticultural vermiculite (Schundler, Metuchen, N.J.) and 2-cm-deep Fafard Mix No. 3-B (Conrad Fafard, Agawam, Mass.), and the seeds were covered with 1-cm-deep vermiculite. The plants were grown in a controlled growth chamber with 14 h of light (23°C) and 10 h of dark (18°C) and 70% humidity. The seedlings were inoculated 72 h after germination by applying 0.5 ml of bacterial culture (OD600, 0.8 to 1.0) along the base (root pole) of the hypocotyl with a Pasteur pipette. Healthy, pink root nodules of similar size were then removed at 8 weeks postinoculation, and the bacteroids were recovered by crushing the nodules in a mortar and pestle in 50 mM Tris-HCl, followed by filtration of the suspension through glass wool (1 cm thick) in a 10-ml disposable syringe. The eluted bacteroids were then pelleted, and the polysaccharides were extracted with hot phenol-water, as described above (although convenient, collecting and freezing the nodules prior to crushing was found to yield inconsistent results, so bacteroid recovery and polysaccharide extraction were performed immediately after removing the nodules from the plants). The polysaccharide preparations were then dissolved in PAGE sample buffer to the same concentration as for the preparations from cultured cells, and separate immunoblots were probed with MAb 8, 9, or 10.
The results showed a significantly reduced signal with MAb 9 (anti-K antigen) in the immunoblot analysis of the polysaccharide preparation from S. meliloti NRG247 bacteroids compared to that from cultured cells, and the bacteroid K antigen migrated as a low-mobility smear, near the top of the lane (Fig. 2). In contrast, there was no detectable binding in the MAb 8 immunoblot analysis of the bacteroid polysaccharides from S. meliloti NRG185, suggesting that there was no K antigen produced by the bacteroids or that it was structurally modified. These results fully corroborate those of previous studies (10, 11), which showed a marginal reaction of MAb 9 with S. meliloti NRG247 bacteroids (whole cells) and no reaction of MAb 8 with S. meliloti NRG185 bacteroids.
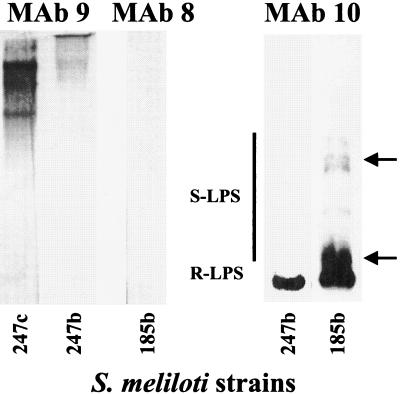
FIG. 2.
Immunoblot analysis of the polysaccharide extracts from alfalfa bacteroids of S. meliloti NRG185 (185b) and S. meliloti NRG247 (247b). The leftmost lane contains the extract from cultured cells of S. meliloti NRG247 (247c) for comparison. The immunoblots were probed with the MAbs shown in Table 2. The two arrows indicate the migration positions of two bacteroid-specific S-LPS banding regions not found in the analysis of cultured cells (the fact that these bands were recognized by MAb 10 showed that it was LPS).
The MAb 10 immunoblot of the bacteroid extracts from both S. meliloti NRG247 and S. meliloti NRG185 showed the presence of R-LPS with a mobility on polyacrylamide gels similar to that of R-LPS from the cultured cells (Fig. 2). This shows that the MAb 10 epitope is present in the R-LPS of bacteroids, indicating that the LPS core structure is similar in both cultured cells and bacteroids. However, this did not preclude the possibility of changes in the R-LPS that do not affect the MAb 10 epitope; in fact, approximately 50% of the R-LPS from the bacteroids fractionated into the phenol phase during the extraction compared to <5% in the extraction of the cultured cells, and the phenol-soluble bacteroid LPS was also recognized by MAb 10 (data not shown). This explained the relatively low response in the immunoblot (compare Fig. 2 and 1) and suggested that there is some difference in the fatty acid content of the lipid A from the bacteroid.
In contrast to the R-LPS, clear differences in the bacteroid-derived S-LPS from S. meliloti NRG185 were apparent in the immunoblot analysis. Interestingly, no S-LPS was detected in the polysaccharide preparation from the S. meliloti NRG247 bacteroids (Fig. 2), showing that specific bacteroid structures may vary significantly from strain to strain in the same host plant.
In their analyses of endophytic sinorhizobia, Olsen et al. (11) found that of the 51 strains (untreated cells) that were positive for MAb 10 in ELISA, 50 gave a positive response with bacteroids (98%), showing that the MAb 10 epitope in the LPS core is essentially unchanged in the bacteroids. Similar results were obtained in immunofluorescence studies (10), in which MAb 10 showed an equivalent affinity for cultured cells or bacteroids. We have shown that MAb 10 did, in fact, bind the extracted R-LPS from bacteroids of S. meliloti NRG185 and S. meliloti NRG247. These results are in agreement with a study of “induced” cells of S. fredii (17), which showed that the addition of apigenin or soybean root extract elicited changes in O-antigen and K-antigen production, but no changes were found in LPS core structure. In addition, the fact that much more of the R-LPS from the S. meliloti bacteroids fractionated into the organic phase during extraction suggested that lipid A structure (fatty acid content) may also be modified during morphogenesis to the bacteroid state, without affecting the MAb 10 epitope.
In contrast to the conserved production of the LPS epitopes in the bacteroids, the immunofluorescence studies of Olsen et al. (10) showed that the strain-specific epitopes (i.e., the K antigens) were diminished (S. meliloti NRG247) or absent (S. meliloti NRG185) in bacteroids, and in ELISAs (11), the strain-specific MAbs recognized a total of 35 strains (untreated cells), but only 2 of the 35 (6%) were positive with the bacteroids (including S. meliloti NRG247). We also showed that the S. meliloti NRG247 bacteroids produced K antigens that were recognized by MAb 9, although they were less abundant and larger, whereas no K antigens were detected with MAb 8 in immunoblot analyses of the polysaccharide extracts from S. meliloti NRG185 bacteroids. Unfortunately, it was not possible with PAGE analyses of these preparations to determine whether structurally modified K antigen, which does not bind the respective MAbs, was produced by the bacteroids of either strain because the polyacrylamide gels showed an abundance of Alcian blue-stained material of unknown nature (data not shown). The material may have been modified K antigen, unrelated bacteroid-specific polysaccharides, contaminating plant-derived material, or some combination of the three. We are currently working towards improved methods of bacteroid recovery to overcome this problem.
Other studies in progress have already shown that the analyses of bacteroid structure will not be simple (13). For example, we have found that changes in abiotic factors, such as temperature, in both the bacterial culture and plant growth conditions, also result in changes to the cell surface of the bacteria and bacteroids. The use of MAbs to study specific cell surface components from the bacteroids of Rhizobium leguminosarum (2, 4, 7) have yielded interesting information about morphogenesis in the pea symbiont, including the temporal expression of specific epitopes during nodule development. We hope that our future studies of bacteroids will provide insight into the differences and similarities in bacterial morphogenesis in narrow-host-range symbiosis (S. meliloti) versus broad-host-range systems (S. fredii), the latter of which includes both determinate and indeterminate host plants.
Acknowledgments
This work was supported in part by grant MCB-9728564 from the National Science Foundation (to B. L. Reuhs) and by the U.S. Department of Energy-funded Center for Plant and Microbial Complex Carbohydrate Research (grant DE-FG02-93ER-20097). T. Ojanen-Reuhs was supported by the Academy of Finland.
REFERENCES
- 1.Becquart-de Kozak I, Reuhs B L, Buffard D, Breda C, Kim J S, Esnault R, Kondorosi A. Role of the K-antigen subgroup of capsular polysaccharides in the early recognition process between Rhizobium meliloti and alfalfa leaves. Mol Plant-Microbe Interact. 1997;10:114–123. [Google Scholar]
- 2.Carlson R W, Reuhs B L, Forsberg L S, Kannenberg E L. Rhizobial cell surface carbohydrates. In: Goldberg J B, editor. Genetics of bacterial polysaccharides. Boca Raton, Fla: CRC Press; 1999. pp. 53–90. [Google Scholar]
- 3.Corzo J, Pérez-Galdona R, León-Barrios M, Gutiérrez-Navarro A M. Alcian Blue fixation allows silver staining of the isolated polysaccharide component of bacterial lipopolysaccharides in polyacrylamide gels. Electrophoresis. 1991;12:439–441. doi: 10.1002/elps.1150120611. [DOI] [PubMed] [Google Scholar]
- 4.de Maagd R A, de Rijk R, Mulders I H M, Lugtenberg B J J. Immunological characterization of Rhizobium leguminosarum outer membrane antigens by use of polyclonal and monoclonal antibodies. J Bacteriol. 1989;171:1136–1142. doi: 10.1128/jb.171.2.1136-1142.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Denarie J, Roche P. Rhizobium nodulation signals. In: Verma D P S, editor. Molecular signals in plant-microbe communications. Boca Raton, Fla: CRC Press; 1992. pp. 295–324. [Google Scholar]
- 6.Forsberg L S, Reuhs B L. Structural characterization of the K antigens from Rhizobium fredii USDA257: evidence for a common structural motif, with strain-specific variation, in the capsular polysaccharides of Rhizobium spp. J Bacteriol. 1997;179:5366–5371. doi: 10.1128/jb.179.17.5366-5371.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kannenberg E L, Perotto S, Bianciotto V, Rathbun E A, Brewin N J. Lipopolysaccharide epitope expression of Rhizobium bacteroids as revealed by in situ immunolabelling of pea root nodule sections. J Bacteriol. 1994;176:2021–2032. doi: 10.1128/jb.176.7.2021-2032.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kim J S, Reuhs B L, Rahman M M, Ridley B, Carlson R W. Separation of bacterial capsular and lipopolysaccharides by preparative electrophoresis. Glycobiology. 1996;6:433–437. doi: 10.1093/glycob/6.4.433. [DOI] [PubMed] [Google Scholar]
- 9.Kiss E, Reuhs B L, Kim J S, Kereszt A, Petrovis G, Putnoky P, Dusha I, Carlson R W, Kondorosi A. The rkpGHI and -J genes are involved in capsular polysaccharide production by Rhizobium meliloti. J Bacteriol. 1997;179:2132–2140. doi: 10.1128/jb.179.7.2132-2140.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Olsen P, Collins M, Rice W. Surface antigens present on vegetative Rhizobium meliloti cells may be diminished or absent when cells are in the bacteroid form. Can J Microbiol. 1992;38:506–509. [Google Scholar]
- 11.Olsen P, Wright S, Collins M, Rice W. Patterns of reactivity between a panel of monoclonal antibodies and forage Rhizobium strains. Appl Environ Microbiol. 1994;60:654–661. doi: 10.1128/aem.60.2.654-661.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Petrovics G, Putnoky P, Reuhs B L, Kim J S, Thorp T A, Noel D, Carlson R W, Kondorosi A. The presence of a novel type of surface polysaccharide in Rhizobium meliloti requires a new fatty acid synthase-like gene cluster involved in symbiotic nodule development. Mol Microbiol. 1993;8:1093–1094. doi: 10.1111/j.1365-2958.1993.tb01653.x. [DOI] [PubMed] [Google Scholar]
- 13.Reuhs, B. L. Unpublished data.
- 14.Reuhs B L. Acidic capsular polysaccharides (K antigens) of Rhizobium. In: Stacey G, Mullin B, Gresshoff P M, editors. Biology of plant-microbe interactions. St. Paul, Minn: International Society for Molecular Plant-Microbe Interactions; 1996. pp. 331–336. [Google Scholar]
- 15.Reuhs B L, Carlson R W, Kim J S. Rhizobium fredii and Rhizobium meliloti produce 3-deoxy-d-manno-2-octulosonic acid-containing polysaccharides that are structurally analogous to group II K antigens (capsular polysaccharides) found in Escherichia coli. J Bacteriol. 1993;175:3570–3580. doi: 10.1128/jb.175.11.3570-3580.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Reuhs B L, Geller D P, Kim J S, Fox J E, Kumar Kolli V S, Pueppke S G. Sinorhizobium fredii and Sinorhizobium meliloti produce structurally conserved lipopolysaccharides and strain-specific K antigens. Appl Environ Microbiol. 1998;64:4930–4938. doi: 10.1128/aem.64.12.4930-4938.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Reuhs B L, Kim J S, Badgett A, Carlson R W. Production of the cell-associated polysaccharides of Rhizobium fredii USDA205 is modulated by apigenin and host root extract. Mol Plant-Microbe Interact. 1994;7:240–247. doi: 10.1094/mpmi-7-0240. [DOI] [PubMed] [Google Scholar]
- 18.Reuhs B L, Williams M N V, Kim J S, Carlson R W, Cote F. Suppression of the Fix− phenotype of Rhizobium meliloti exoB mutants by lpsZ is correlated to a modified expression of the K polysaccharide. J Bacteriol. 1995;177:4289–4296. doi: 10.1128/jb.177.15.4289-4296.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Whitfield C, Valvano M A. Biosynthesis and expression of cell-surface polysaccharides in Gram-negative bacteria. Adv Microb Physiol. 1993;35:135–246. doi: 10.1016/s0065-2911(08)60099-5. [DOI] [PubMed] [Google Scholar]