Abstract
A mutant (XT906) of Xanthomonas campestris pv. citri, the causal agent of citrus canker, was induced by insertion of the transposon Tn5tac1 and isolated. This mutant did not grow or elicit canker disease in citrus leaves but was still able to induce a hypersensitive response in a nonhost plant (the common bean). The mutant was also unable to grow on minimal medium containing fructose or glycerol as the sole carbon source. A 2.5-kb fragment of wild-type DNA that complemented the mutant phenotype of XT906 was isolated. Sequence analysis revealed that this DNA fragment encoded a protein of 562 amino acids that shows homology to phosphoglucose isomerase (PGI). Enzyme activity assay confirmed that the encoded protein possesses PGI activity. Analysis of the activity of the promoter of the pgi gene revealed that it was inhibited by growth in complex medium but induced by culture in plant extract. These results demonstrate that PGI is required for pathogenicity of X. campestris pv. citri.
Xanthomonas campestris pv. citri is the agent responsible for bacterial canker in citrus (10), which is a problem worldwide in the production of citrus fruits (28). The pathogen induces the formation of cankers and water-soaked lesions surrounded by chlorotic haloes on leaves as well as surface-penetrating necrotic lesions on fruit. It enters its host through natural openings (such as stomata) or wounds on the plant surface (14, 28), where the environment is less harsh and rich in nutrients that include glucose, fructose, xylose, sucrose, and galactose (21).
The interaction between plant and pathogen is a dynamic process that involves the exchange of signals. An important subset of pathogenicity genes performs regulatory functions that are likely required for the bacteria to adapt to the environmental and physiological changes encountered during infection. The expression of pathogenicity or pathogenicity-related genes is regulated by environmental factors (such as pH and osmotic strength), plant signals (such as carbon sources, organic nitrogen, and phosphate), and catabolite repression (23, 26).
In bacterial phytopathogens, the expression of hrp genes, which determine basic pathogenicity to the host and the ability to cause a hypersensitive reaction (HR) in nonhost plants, is under the control of plant signals or environmental factors (23, 26). A cluster of genes is responsible for the production and regulation in Xanthomonas and Pseudomonas that controls the synthesis of virulence factors in response to physiological or environmental changes (26). A small diffusible signal molecule is required for a regulatory system for pathogenicity of X. campestris (3). In Agrobacterium- or Rhizobium-plant interactions, specific plant phenolic compounds act as inducers of bacterial genes important for crown gall pathogenesis (35) and nodulation (20), respectively. The products of such pathogenicity or pathogenicity-related genes may function as signals indicating that the environment is suitable for pathogen growth and initial pathogenesis (23, 26). However, the molecular mechanisms in Xanthomonas that underlie plant-pathogen signal exchange and the corresponding signaling pathways remain unclear.
We have now characterized a Tn5tac1 insertional mutant, XT906, of X. campestris pv. citri that exhibits defects in pathogenicity and virulence. The transposon was shown to disrupt a phosphoglucose isomerase (PGI) gene in this mutant. Our data also indicate that PGI is required for the pathogenicity of Xanthomonas. These observations suggest that PGI performs dual functions and have implications for our understanding of the metabolism and pathogenicity of Xanthomonas.
MATERIALS AND METHODS
Bacterial strains, plasmids, and culture media.
The bacterial strains and plasmids used in this study are listed in Table 1. Xanthomonas campestris pv. citri strains were routinely grown at 28°C in TSG nutrient medium (10 g of Bacto Tryptone, 5 g of Bacto Soytone, 5 g of NaCl, and 2 g of glucose [each] per liter), XVM2 medium [20 mM NaCl, 10 mM (NH4)2SO4, 5 mM MgSO4, 1 mM CaCl2, 0.16 mM KH2PO4, 0.32 mM K2HPO4, 0.01 mM FeSO4, 0.03% Casamino Acids (pH 6.7), 10 mM fructose, 10 mM sucrose] (32), or modified synthetic M9 medium (25) supplemented with various carbon sources as indicated. Escherichia coli strains were grown in Luria-Bertani medium at 37°C. Media were supplemented with kanamycin (50 μg/ml), ampicillin (50 μg/ml), or gentamicin (5 μg/ml) as required.
TABLE 1.
Bacterial strains and plasmids used in the present study
| Strain or plasmid | Relevant characteristic(s)a | Reference or source |
|---|---|---|
| Strains | ||
| E. coli | ||
| HB101 | supE44 hsdS20(rK− mK+) recA13 ara-14 proA2 lacY1 galK2 rpsL20 xyl-5 mtl-1 | 4a |
| DH5αMCR | F−recA j80 ΔlacZDM15 mcrA mcrB mcrC | BRL |
| X. campestris pv. citri | ||
| XW47 | Natural isolate from citrus from Taiwan | 28 |
| XT10 | Nonpathogenic Tn5tac1 insertional mutant of XW47 | 28 |
| XT906 | Nonpathogenic Tn5tac1 insertional mutant of XW47 | This study |
| Plasmids | ||
| pBR322::Tn5tac1 | pBR322 containing Tn5tac1 (lacIq Ptac); Kmr Ampr Tetr | 6 |
| pHC79 | pBR322 derivative containing one cos site | 16 |
| pHT906 | pHC79 clone containing a 30- to 40-kb insert of XT906 DNA and Tn5tac1 | This study |
| pHW906 | pHC79 clone containing a 30- to 40-kb insert of XW47 DNA | This study |
| pBluescript KS(+) | Ampr cloning vector | Stratagene |
| pUFR047 | incW Gmr Amprmob+ Pmob lacZA+ par+ | 11 |
| pUW906XH | pUFR047 clone containing a 5-kb XhoI-HindIII fragment of pgi | This study |
| pUW906XAp | pUFR047 clone containing a 2.5-kb XhoI-ApaI fragment of pgi | This study |
| pUW906P | pUFR047 clone containing an internal PstI fragment of pgi | This study |
| pUGUS | pUFR047 containing a promoterless gus gene | This study |
| pUW906PGUS | pUGUS containing an internal PstI fragment of pgi | This study |
| pUW10PGUS | pUGUS containing an internal PstI fragment of hrpX | 28 |
Kmr, Ampr, Tetr, and Gmr, kanamycin, ampicillin, tetracycline, and gentamicin resistance, respectively.
Transposition mutagenesis.
The suicide plasmid pBR322::Tn5tac1 (6) (0.5 to 1 μg) was introduced directly into X. campestris pv. citri XW47 by electroporation (36). Given that pBR322 is not able to replicate in Xanthomonas and contains a kanamycin resistance gene and that X. campestris pv. citri XW47 is sensitive to kanamycin, we selected kanamycin-resistant colonies to examine further for evidence of Tn5tac1 insertion.
Plant material and plant inoculations.
Citrus lemon, tobacco (Nicotiana tabacum cultivar W38), and common bean (Phaseolus vulgaris L. var. humilis Alef) were grown in a greenhouse. For disease symptom assays, 20 to 30 puncture wounds were introduced per 1-cm2 area of citrus leaf with a standard 26-gauge needle. Bacteria that had been grown overnight in TSG broth, harvested by centrifugation, and resuspended in 0.85% (wt/vol) NaCl at a density of ∼108 CFU/ml were then applied in 10-μl droplets directly onto the puncture wound sites on the citrus leaves. The plant reaction was assessed 7 to 14 days after inoculation. For HR tests, X. campestris pv. citri XW47 and its mutant were grown as for the symptom assays, adjusted to a density of 106 CFU/ml, and injected with a fine hypodermic needle into the intercellular spaces of a common bean leaf. The injection was administered near the lateral veins of the leaf, where the tissue is thicker (17). After infection, plants were maintained in the greenhouse.
For in planta growth assays, periodic sampling was performed by excising two of the inoculated leaves and harvesting bacteria from leaf disks. These punches (1 cm in diameter) were removed and shattered at various times after inoculation. Bacteria were eluted from the pieces in 1 ml of sterile tap water, and appropriate dilutions were plated to obtain single colonies on medium containing appropriate antibiotics. Colonies were counted after 48 h; the results were calculated as CFU per square centimeter of leaf tissue.
For the preparation of leaf extracts, fully expanded leaves from tobacco, common bean, and lemon plants were harvested and frozen in liquid nitrogen. After removal of the main veins, the tissue was homogenized in 1 mM MgCl2 (1 g [fresh weight] per ml) in the presence of polyvinylpyrrolidone (1 g [fresh weight] per 0.1 ml). The homogenate was centrifuged for 30 min at 50,000 × g, and the resulting supernatant was tested at a dilution of 1/20 for induction assays.
DNA manipulation and Southern hybridization.
Standard microbiological and molecular genetic techniques were used (25) unless otherwise stated. Total genomic DNA was isolated by the cetyltrimethylammonium bromide method (2) from bacterial cells grown in liquid culture. The integration of Tn5tac1 was analyzed by Southern blot hybridization: DNA fragments produced by digestion with restriction endonucleases were fractionated by electrophoresis and then transferred to membranes (Genescreen Plus; DuPont Biotechnology Systems, Boston, Mass.). Hybridization and detection protocols were performed with the use of nonradioactive DNA labeling and detection kits (Boehringer Mannheim, Mannheim, Germany).
DNA sequence analyses.
Sequence data were analyzed with the University of Wisconsin Genetic Computer Group software package (version 9.0). The SwissProt 23.0 CDS translation from the GenBank Release 73.1 database was searched with the predicted amino acid sequences of the pgi gene and the Blast program, run at the National Center for Biotechnology Information network service in Bethesda, Md.
PGI enzyme assay.
The assay of PGI activity by electrophoresis and staining was performed according to a method similar to that described by Gottlieb and Higgins (13). Cultured cells (5 × 108) were collected by centrifugation, washed once with 100 mM NaCl, and resuspended in 100 μl of grinding buffer (11). The cells were disrupted by ultrasonic treatment with a heat systems ultrasonicator (New Highway, Farmingdale, N.Y.) and then centrifuged at 12,000 × g for 10 min. The supernatant was subjected to electrophoresis, and the gel was stained with a solution containing 0.1 M Tris-HCl (pH 8.0), 5 mM fructose-6-phosphate dehydrogenase, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (80 ng/ml), and phenazine methosulfate (20 ng/ml) and then developed at 37°C (29).
Assay of pgi promoter activity.
A DNA fragment containing the putative promoter sequence of X. campestris pv. citri pgi was cloned into the PstI site of pUGUS, which contains a promoterless β-glucuronidase (GUS) gene. For the assay of GUS activity, bacteria transformed with the resulting construct were grown overnight in TSG medium, collected by centrifugation, and washed twice in 1 mM MgCl2 before inoculation into complex medium or minimal XVM2 medium supplemented with various carbon sources. After incubation for 16 h, cells were harvested by centrifugation and resuspended in assay buffer (16). Portions of the suspension were then assayed for GUS activity. The number of bacteria (CFU) per assay was determined by plating appropriate dilutions on selective medium. GUS activity was measured with a fluorometer and 4-methylumbelliferyl glucuronide as the substrate, as previously described (16). One unit of GUS activity is defined as 1 nmol of 4-methyl-umbelliferone produced per min.
Nucleotide sequence accession number.
The GenBank accession number of the reported sequence of the 2.5-kb DNA fragment complementing the mutant phenotype of XT906 is AF054807.
RESULTS
Isolation and characterization of the nonpathogenic mutant XT906.
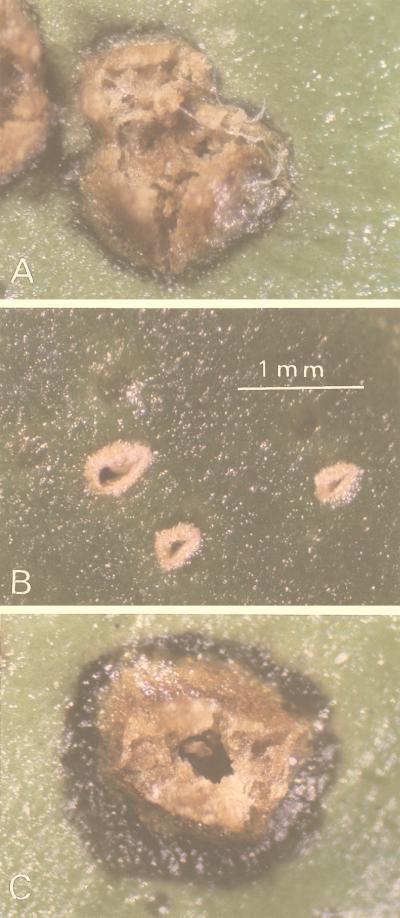
To identify pathogenicity-related genes and investigate the mechanism of citrus canker disease, we have used the transposon Tn5tac1 to induce mutations that affect the pathogenicity of X. campestris pv. citri. The suicide plasmid pBR322::Tn5tac1 was introduced directly into X. campestris pv. citri XW47 by electroporation. About 1,000 kanamycin-resistant clones were selected and tested for their pathogenicity by inoculation into puncture wounds of citrus leaves. Inoculated leaves were examined over the 2-week period normally required for disease development. Typical symptoms of citrus canker caused by wild-type bacteria include raised eruptions surrounded by water-soaked lesions (Fig. 1A). The mutant XT906 failed to induce disease symptoms in the host plant for up to 1 month after inoculation (Fig. 1B).
FIG. 1.
Symptoms expressed on citrus leaves 2 weeks after inoculation with X. campestris pv. citri XW47 (A), XT906 (B), or XT906(pUW906XAp) (C).
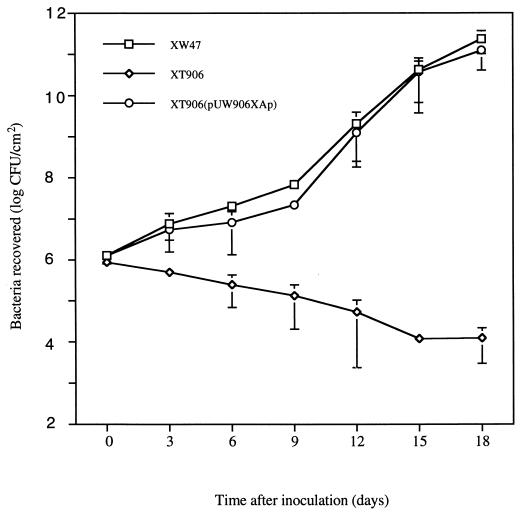
The ability of this mutant to elicit an HR in a nonhost plant was investigated by the injection of bacterial cells into the intercellular space of common bean leaves. The mutant retained the ability to induce an HR in this plant species (data not shown). Thus, the XT906 mutant showed a Path− HR+ phenotype. Examination of bacterial growth in planta 18 days after inoculation revealed that whereas the population of the wild-type strain XW47 had increased by a factor of 105, that of the mutant XT906 had decreased by a factor of 102 (Fig. 2).
FIG. 2.
Growth in planta of X. campestris pv. citri XW47, XT906, and XT906(pUW906XAp). Data are means ± standard errors of the means (SEM) of values from three independent experiments, each performed in duplicate.
Cloning of a DNA fragment required for growth in citrus and pathogenicity.
Southern blot analysis confirmed the presence of a single copy of Tn5tac1 in the mutant XT906 (data not shown). To clone the transposon-tagged gene, we screened a cosmid pHC79 library prepared with XT906 DNA for clones containing Tn5tac1. From one of the resulting positive clones, pHT906, a 3-kb EcoRI-HindIII fragment containing a portion of Tn5tac1 and X. campestris pv. citri flanking DNA was isolated and subcloned into the pBluescript KS(+) vector, yielding pKT906IEH. The DNA flanking Tn5tac1 was used as a probe to isolate clones from a pHC79 library prepared with genomic DNA from XW47. One of the resulting positive clones, pHW906, was selected for subcloning into a low-copy-number, broad-host-range plasmid, pUFR047, and for further complementation studies.
Subclones containing a 5-kb XhoI-HindIII fragment (pUW906XH) or a 2.5-kb XhoI-ApaI fragment (pUW906XAp) were introduced separately into the mutant XT906 by electroporation. Kanamycin- and gentamicin-resistant transformants were then tested for pathogenicity on citrus leaves. The Tn5tac1-induced mutation in XT906 was complemented by both plasmid constructs, as evidenced by the induction of disease symptoms (Fig. 1C and 3A), but not by pUFR047. The growth of the complemented strain XT906(pUW906XAp) on citrus leaves also resembled that of the wild-type strain (Fig. 2). The 2.5-kb DNA fragment contained in pUW906XAp thus appeared to complement fully the pathogenicity defect of the mutant XT906.
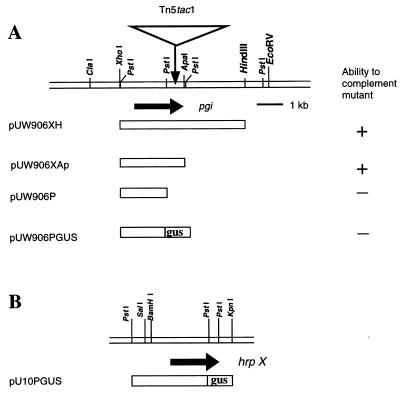
FIG. 3.
Structural organization of the pgi (A) and hrpX (B) loci of X. campestris pv. citri and of plasmids derived therefrom. (A) Restriction map of the genomic region containing pgi showing the orientation of the gene (horizontal arrow) and the site of transposon insertion in the mutant XT906. The plasmids pUW906XH, pUW906XAp, and pUW906P were used for complementation studies (the results of which are summarized), and the plasmid pUW906PGUS (containing a pgi::gus fusion gene) was used for the analysis of regulation of pgi expression. (B) Restriction map of the genomic region containing hrpX showing the orientation of the gene (horizontal arrow) and the structure of the plasmid pUW10PGUS, which contains an hrpX::gus fusion gene.
Identification of the mutated gene in XT906.
The 2.5-kb DNA fragment that complemented the mutant phenotype of XT906 was sequenced and found to contain an open reading frame (ORF) that encodes a polypeptide of 562 amino acids. The Tn5tac1 transposon was shown to be inserted in this ORF in XT906 at nucleotide position 1937. The predicted amino acid sequence of the ORF most closely resembled that of PGI from pgi, with which it shared 44% identity and 61% similarity.
To confirm that the ORF encoded a PGI enzyme, we assayed extracts of bacterial strains for PGI activity. PGI activity was detected in the wild-type strain XW47 and the complemented strain XT906(pUW906XAp) but not in XT906 or XT906(pUFR047) (data not shown). The PGI activity of strain XT10, an HR− mutant of XW47 in which the hrpX gene is disrupted (30), was similar to that of the wild-type strain. Thus, the defects in both pathogenicity and PGI activity of XT906 were complemented by the same 2.5-kb fragment of wild-type genomic DNA.
To examine the effect of the PGI mutation on bacterial growth in culture, we tested the ability of XT906 to utilize glucose, sucrose, fructose, glycerol, or mannitol. Whereas the wild-type strain grew on medium containing any of these carbon sources except mannitol, XT906 was not able to utilize mannitol, fructose, or glycerol; the growth of XT906 on medium containing glucose or sucrose was similar to that of XW47 (Table 2). The growth defect of XT906 in culture was corrected by transformation with pUW906XAp but not with pUFR047.
TABLE 2.
Growth of X. campestris pv. citri strains on minimal medium supplemented with various carbon sourcesa
| Carbon source | Strain
|
|||
|---|---|---|---|---|
| XW47 | XT906 | XT906(pUFR047) | XT906(pUW906XAp) | |
| None | − | − | − | − |
| Glucose | ++ | ++ | ++ | ++ |
| Mannitol | − | − | − | − |
| Fructose | + | − | − | + |
| Glycerol | + | − | − | + |
| Sucrose | + | + | + | + |
Bacterial suspensions (5,000 CFU per 10 μl) were plated in M9 medium supplemented with test carbon sources at a concentration of 0.01 M. After incubation at 28°C for 10 days, the extent of bacterial growth was assessed as ++ (substantial growth), + (less marked growth), or − (no growth).
Regulation of the expression of pgi.
The mutation in pgi resulted in the inability of X. campestris pv. citri to utilize fructose or glycerol as carbon sources, to grow in plant tissue, and to cause typical canker symptoms. These observations prompted us to investigate the expression of pgi under various conditions, including complex medium, minimal medium containing different carbon sources, and plant extracts. The synthetic medium XVM2 was shown to induce expression of hrp loci (hrpA, hrpX, and hrpG) of X. campestris pv. vesicatoria to an extent similar to that apparent during growth in plant extracts (32). Expression of the hrpX gene of X. campestris pv. vesicatoria is induced in XVM2 medium but not in NYG medium (31). We examined the expression of X. campestris pv. citri pgi with a similar approach and compared the results with those obtained for hrpX.
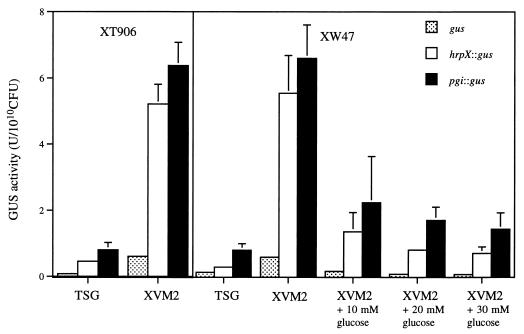
For analysis of the expression of pgi, we measured the activity of a fusion construct containing a promoterless GUS gene in both the wild-type and XT906 mutant strains under various growth conditions. We thus constructed plasmids containing pgi::gus (pUW906PGUS) and hrpX::gus (pUW10PGUS) fusion genes (Fig. 3) and introduced them into X. campestris pv. citri. When bacteria were grown in the complex medium TSG, the amount of GUS activity in extracts of XW47 harboring the pgi::gus or hrpX::gus constructs was similar to that in extracts of XW47 transformed with the control plasmid pUGUS; similar results were obtained with strain XT906 (Fig. 4). However, when XW47 was grown in XVM2 (which does not contain glucose), the amount of GUS activity in extracts of cells harboring pgi::gus or hrpX::gus was markedly greater than that in extracts of cells harboring pUGUS as well as that in extracts of bacteria grown in TSG medium. Addition of glucose to XVM2 resulted in a concentration-dependent decrease in the GUS activity of XW47 cells harboring either fusion construct. To analyze whether pgi is subjected to feedback regulation, we compared the GUS activities of the wild-type and XT906 strains transformed with the pgi::gus construct and grown in XVM2 medium. In the presence of feedback regulation mediated by the pgi promoter, the activity of the pgi::gus construct would be expected to be altered by the end product of PGI action in the wild type but not in the pgi mutant. No difference in GUS activity was apparent between XW47 and XT906 strains, indicating the absence of feedback regulation.
FIG. 4.
Expression of pgi and hrpX genes in the wild-type XW47 and mutant XT906 strains of X. campestris pv. citri grown in various culture media. Strains XW47 or XT906 were transformed with the plasmid pUW906PGUS (pgi::gus), the plasmid pUW10PGUS (hrpX::gus), or the promoterless control pUGUS (gus) and were grown for 16 h in either TSG or XVM2 medium in the absence or presence of 10, 20, or 30 mM glucose, as indicated. Cell extracts were then prepared and assayed for GUS activity. Data are means ± SEM of values from three experiments.
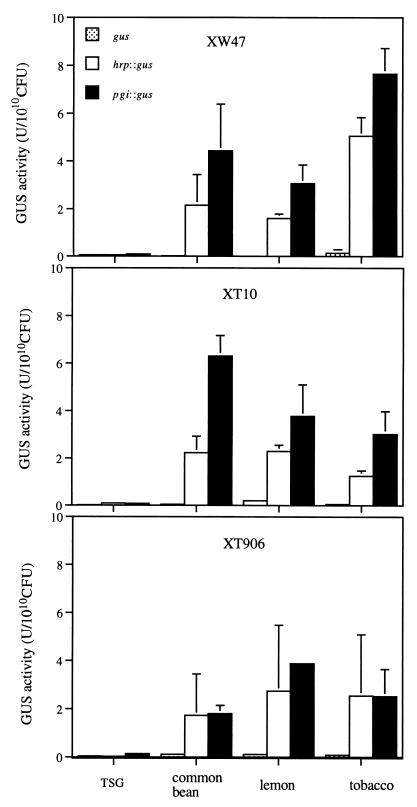
Given that hrpX (Path− HR−) and pgi (Path− HR+) mutants both exhibit a pathogenicity defect but that the pgi mutant is still able to induce an HR in the common bean, we examined the expression of hrpX::gus and pgi::gus fusion genes in bacteria grown in leaf extracts of lemon, common bean, and tobacco (a nonhost of X. campestris pv. citri that does not exhibit an HR response). When the bacteria were grown in the presence of any of the three plant extracts, the amount of GUS activity in strains XW47, XT906, and XT10 harboring either fusion gene was markedly greater than that apparent when they were grown in TSG medium (Fig. 5); the activities of the fusion genes were markedly greater than that of the promoterless gus gene for all strains grown in the presence of plant extract. Thus, extracts of both host and nonhost plants induced the expression of hrpX and pgi genes of X. campestris pv. citri.
FIG. 5.
Expression of pgi and hrpX genes in the wild-type XW47 (top) and mutant XT10 (middle) and XT906 (bottom) strains of X. campestris pv. citri grown in the presence of extracts of lemon, common bean, or tobacco plants. Bacteria were transformed with pUW906PGUS (pgi::gus), pUW10PGUS (hrpX::gus), or pUGUS (gus) and grown for 16 h in either TSG medium or plant extract. Bacterial extracts were then prepared and assayed for GUS activity. Data are means ± SEM of values from three experiments.
DISCUSSION
With the use of transposon mutagenesis, we have isolated a mutant, XT906, of X. campestris pv. citri that is nonpathogenic to the host plant. A 2.5-kb fragment of wild-type genomic DNA complemented the pathogenicity defect of this mutant and was shown to contain an ORF that encodes PGI. Our results demonstrate that a single mutation abolished both PGI activity and pathogenicity, suggesting that PGI contributes to pathogenicity, either directly or indirectly.
PGI is a dimeric enzyme that catalyzes the reversible isomerization of glucose 6-phosphate and fructose 6-phosphate and is essential for fructose catabolism by Pseudomonas (1). Although PGI-deficient xanthomonads have not previously been described, Xanthomonas is closely related evolutionarily to Pseudomonas (1, 9). Mutants of Pseudomonas aeruginosa that lack PGI activity grow slowly on glycerol and fructose media but normally on glucose medium (22), characteristics similar to those described here for the pgi mutant of X. campestris pv. citri. Xanthomonas utilizes glucose and sucrose as primary carbon sources for its growth on prepared medium (21); most glucose is catabolized by the Entner-Doudoroff pathway (37), thereby generating pyruvate and glyceraldehyde 3-phosphate. PGI does not participate in the Entner-Doudoroff pathway, which supplies most energy for cell growth (33), suggesting that the PGI enzyme is not essential for metabolism based on glucose as the primary carbon source.
It is possible that the pathogenicity defect of the Xanthomonas mutant XT906 is the result of insufficient PGI activity required for growth in planta. However, various carbon sources present in plant tissue, such as glucose and sucrose (21), should be able to support the growth of XT906. Thus, it is unlikely that the lack of growth in planta and the pathogenicity of XT906 is due to an inability to utilize fructose as a carbon source because of PGI deficiency. Thus, the XT906 mutant did not multiply in plant tissue, even in the presence of carbon sources that should be able to support its growth.
Various mutations have been shown to exert effects on catabolism and pathogenicity, with the affected catabolites contributing to regulation of the expression of pathogenicity-related genes (12, 19, 24). Isolation of a phosphoglyceromutase mutant of Pseudomonas syringae revealed that this glycolytic enzyme is required for both bacterial growth in the host and the utilization of carboxylic acids (15, 19). The pgm mutant is nonpathogenic in the host, but it still induces an HR in nonhost plants (7), characteristics similar to those of the pgi mutant of X. campestris pv. citri. Furthermore, the pgm mutant grows well on complete medium as well as on media containing citrate plus either glucose or fructose but grows to a limited extent on media containing fructose, glucose, or glycerol as the only carbon source. It is possible that metabolites generated by the pathways that include PGI or phosphoglyceromutase regulate genes essential for cell propagation in planta.
Genes that encode bacterial virulence factors are often subject to coordinate regulation, and these regulatory systems are able to respond to various environmental signals that may be encountered during the cycle of infection (18). Thus, regulation of the expression of pgi and other pathogenicity-related genes such as hrpX, which is also required for pathogenesis and for an HR in nonhost plants (4, 31), warrants further investigation. In Agrobacterium tumefaciens, sugars and phenolic compounds synergistically induce the expression of vir genes through a distinct regulatory pathway that includes VirA and the chromosomally encoded virulence protein ChvE (5, 34). Similar regulation of virulence genes may occur in other pathogens, including Erwinia, Pseudomonas, and Xanthomonas. Expression of the hrpX gene of X. campestris pv. vesicatoria is regulated by various factors; it is inhibited by growth in complex medium and is inducible in the host plant (31). We have now shown that the expression of both hrpX and pgi genes of X. campestris pv. citri cultured in plant extracts or the synthetic medium XVM2 is markedly greater than that apparent for cells cultured in the complex medium TSG, suggesting that a factor (or factors) present in plants and XVM2 regulates the promoter activities of these genes. Our results thus indicate that both hrpX and pgi are coordinately regulated by similar catabolic systems. These catabolic systems may therefore play an important role in pathogenicity and may regulate the expression of other pathogenicity-related genes.
The bacterial pathogenicity genes that determine the nature of the interaction between the bacterium and the plant can be divided into two classes: disease-specific (dsp) genes, which are associated with disease development in host plants but not with the induction of an HR in nonhost plants (8, 27), and hrp genes, which are required for both the pathogenic interaction with host plants and the induction of an HR in nonhost plants (26). The hrp genes of X. campestris pv. campestris are induced in both host and nonhost plants (26), suggesting that the conditions required for the induction of hrp genes in a certain pathovar are not plant species specific and that the host range of the pathogen is controlled by some other mechanism. The properties of the pgi gene characterized in the present study resemble those of dsp genes more closely than they do those of hrp genes. Our experiments with hrpX::gus and pgi::gus fusion genes revealed that the expression of both was induced by the culture of bacteria in leaf extracts of citrus, common bean, or tobacco, suggesting that both hrp and pgi genes are induced in host and nonhost plants, even in a nonhost plant (such as tobacco) that does not exhibit an HR to X. campestris pv. citri. Thus, although hrp and pgi mutants show different phenotypes with regard to their effects on nonhost plants, the corresponding genes may not play a major role in determining plant species specificity.
Many genes whose products participate in bacterial metabolism may be regulated in response to changes in the nutrient environment and thereby contribute to phytopathogenicity. The pgi gene of X. campestris pv. citri, which has now been shown to contribute to plant-pathogen interaction, appears to fall into this category.
ACKNOWLEDGMENTS
This research was supported by the Institute of Molecular Biology, Academia Sinica, and the National Science Council (NSC grant 88-2311-B001-085), Taiwan, Republic of China.
We thank Dean W. Gabriel for providing plasmid pUFR047 and Jychian Chen for comments on the manuscript.
REFERENCES
- 1.Allenza P, Lee Y N, Lessie T G. Enzymes related to fructose utilization in Pseudomonas cepacia. J Bacteriol. 1982;150:1348–1356. doi: 10.1128/jb.150.3.1348-1356.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ausubel F M, Brent R, Kingston R E, Moore S S, Seidman J G, Smith J A, Struhl K. Current protocols in molecular biology. Vol. 1. New York, N.Y: Wiley; 1987. [Google Scholar]
- 3.Barber C E, Tang J L, Feng J X, Pan M Q, Wilson T J G, Slater H, Dow J M, Williams P, Daniels M J. A novel regulatory system required for pathogenicity of Xanthomonas campestris is mediated by a small diffusible signal molecule. Mol Microbiol. 1997;24:555–566. doi: 10.1046/j.1365-2958.1997.3721736.x. [DOI] [PubMed] [Google Scholar]
- 4.Bonas U, Fenselau S, Horns T, Maric C, Moussian B, Pierre M, Wengelnik K, den Ackerveken V. hrpX and avirulence genes of Xanthomonas campestris pv. vesicatoria controlling the interaction with pepper and tomato. In: Daniels M J, et al., editors. Advances in molecular genetics of plant-microbe interactions. Vol. 3. Amsterdam, The Netherlands: Kluwer; 1994. pp. 57–64. [Google Scholar]
- 4a.Boyer H W, Roulland-Dussoix D. A complementation analysis of the restriction and modification of DNA in Escherichia coli. J Mol Biol. 1969;14:459–472. doi: 10.1016/0022-2836(69)90288-5. [DOI] [PubMed] [Google Scholar]
- 5.Cangelosi G A, Ankenbauer R G, Nester E W. Sugars induce the Agrobacterium virulence genes through a periplasmic binding protein and a transmembrane signal protein. Proc Natl Acad Sci USA. 1990;87:6708–6712. doi: 10.1073/pnas.87.17.6708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chow W Y, Berg D E. Tn5tac1, a derivative of transposon Tn5 that generates conditional mutations. Proc Natl Acad Sci USA. 1988;85:6468–6472. doi: 10.1073/pnas.85.17.6468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cuppels D A. Generation and characterization of Tn5 insertion mutations in Pseudomonas syringae pv. tomato. Appl Environ Microbiol. 1986;51:323–327. doi: 10.1128/aem.51.2.323-327.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Daniels M J, Barber C E, Turner P C, Cleary W G, Sawczyc M K. Isolation of mutants of Xanthomonas campestris pv. campestris showing altered pathogenicity. J Gen Microbiol. 1984;130:2447–2455. [Google Scholar]
- 9.De Crécy-Lagard O M, Bouvet M, Lejeune P, Danchin A. Fructose catabolism in Xanthomonas campestris pv. campestris. J Biol Chem. 1991;266:8154–8161. [PubMed] [Google Scholar]
- 10.Dye D W, Bradbury J F, Goto M, Hayward A C, Lelliott R A, Schroth M N. International standards for naming pathovars of phytopathogenic bacteria and a list of pathovar names and pathotype strains. Rev Plant Pathol. 1980;59:153–167. [Google Scholar]
- 11.Fraenkel D G, Levisohn S R. Glucose and gluconate metabolism in an Escherichia coli mutant lacking phosphoglucose isomerase. J Bacteriol. 1967;93:1571–1578. doi: 10.1128/jb.93.5.1571-1578.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gold S, Nishio S, Tsuyumu S, Keen N T. Analysis of the pelE promoter in Erwinia chrysanthemi EC16. Mol Plant-Microbe Interact. 1992;5:170–178. [PubMed] [Google Scholar]
- 13.Gottlieb L D, Higgins R C. Phosphoglucose isomerase expression in species of clarkia with and without a duplication of the coding gene. Genetics. 1984;107:131–140. doi: 10.1093/genetics/107.1.131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Graham J H, Gottwald T R, Riley T D, Bruce M A. Susceptibility of citrus fruit to bacterial spot and citrus canker. Phytopathology. 1992;82:452–457. [Google Scholar]
- 15.Jackson D P, Gray D A, Morris V L. Identification of a DNA region required for growth of Pseudomonas syringae pv. tomato on tomato plants. Can J Microbiol. 1992;38:883–890. [Google Scholar]
- 16.Jefferson R A, Burgess S M, Hirsh D. β-Glucuronidase from Escherichia coli as a gene-fusion marker. Proc Natl Acad Sci USA. 1986;83:8447–8451. doi: 10.1073/pnas.83.22.8447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Klement Z. Rapid detection of the pathogenicity of phytopathogenic pseudomonads. Nature. 1963;199:299–300. doi: 10.1038/199299b0. [DOI] [PubMed] [Google Scholar]
- 18.Miller J F, Mekalanos J J, Falkow S. Coordinate regulation and sensory transduction in the control of bacterial virulence. Science. 1989;243:916–922. doi: 10.1126/science.2537530. [DOI] [PubMed] [Google Scholar]
- 19.Morris V L, Jackson D P, Grattan M, Ainsworth T, Cuppels D A. Isolation and sequence analysis of the Pseudomonas syringae pv. tomato gene encoding a 2,3-diphosphoglycerate-independent phosphoglyceromutase. J Bacteriol. 1995;177:1727–1733. doi: 10.1128/jb.177.7.1727-1733.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Mulligan J T, Long S R. Induction of Rhizobium nodC expression by plant exudate requires nodD. Proc Natl Acad Sci USA. 1985;82:6609–6613. doi: 10.1073/pnas.82.19.6609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Padmanbhan D, Vidhyasekaran P, Rajagopalan C K S. Changes in photosynthesis and carbohydrate content in canker and halo regions in Xanthomonas citri infected citrus leaves. Indian J Phytopathol. 1974;26:215–217. [Google Scholar]
- 22.Phibbs P V, McCowen S M, Feary T M, Blevins W T. Mannitol and fructose catabolic pathway of Pseudomonas aeruginosa carbohydrate-negative mutant and pleiotropic effects of certain enzyme deficiencies. J Bacteriol. 1978;133:717–728. doi: 10.1128/jb.133.2.717-728.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Rahm L G, Mindrinos M N, Panopoulos N J. Plant and environmental sensor signals control the expression of hrpX genes in Pseudomonas syringae pv. phaseolicola. J Bacteriol. 1992;174:3499–3507. doi: 10.1128/jb.174.11.3499-3507.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Reverchon S, Expert D, Robert-Bandouy J. The cyclic AMP receptor protein is the main activator of pectinolysis genes in Erwinia chrysanthemi. J Bacteriol. 1997;179:3500–3508. doi: 10.1128/jb.179.11.3500-3508.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 26.Schulte R, Bonas U. Expression of the Xanthomonas campestris pv. vesicatoria hrp gene cluster, which determines pathogenicity and hypersensitivity on pepper and tomato, is plant inducible. J Bacteriol. 1992;174:815–823. doi: 10.1128/jb.174.3.815-823.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Seal S E, Cooper R M, Clarkson J M. Identification of a pathogenicity locus in Xanthomonas campestris pv. vesicatoria. Mol Gen Genet. 1990;222:452–456. doi: 10.1007/BF00633855. [DOI] [PubMed] [Google Scholar]
- 28.Stall R E, Seymour C P. Canker, a threat to citrus in the Gulf-Coast states. Plant Dis. 1983;67:581–585. [Google Scholar]
- 29.Tait R C, Froman B E, Laudencia-Chingcuanco D L, Gottlieb L D. Plant phosphoglucose isomerase genes lack introns and are expressed in Escherichia coli. Plant Mol Biol. 1988;11:381–388. doi: 10.1007/BF00039018. [DOI] [PubMed] [Google Scholar]
- 30.Tung, S. Y., and T. T. Kuo. Isolation and characterization of mutants of the citrus canker pathogen Xanthomonas campestris pv. citri that induce distinct patterns of disease. Submitted for publication.
- 31.Wengelnik K, Bonas U. HrpXv, an AraC-type regulator, activates expression of five of the six loci in the hrp cluster of Xanthomonas campestris pv. vesicatoria. J Bacteriol. 1996;178:3462–3469. doi: 10.1128/jb.178.12.3462-3469.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wengelnik K, Marie C, Russel M, Bonas U. Expression and localization of HrpA1, a protein of Xanthomonas campestris pv. vesicatoria essential for pathogenicity and induction of the hypersensitive reaction. J Bacteriol. 1996;178:1061–1069. doi: 10.1128/jb.178.4.1061-1069.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Whitfield C I, Sutherland W, Cripps R E. Glucose metabolism in Xanthomonas campestris. J Gen Microbiol. 1982;128:981–985. [Google Scholar]
- 34.Winans S C, Kerstetter R A, Nester E W. Transcriptional regulation of the virA and virG genes of Agrobacterium tumefaciens. J Bacteriol. 1988;170:4047–4054. doi: 10.1128/jb.170.9.4047-4054.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Winans S C. Transcriptional induction of an Agrobaterium regulatory gene at tandem promoters by plant-released phenolic compounds, phosphate starvation, and acidic growth media. J Bacteriol. 1990;172:2433–2438. doi: 10.1128/jb.172.5.2433-2438.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yang M K, Su W C, Kuo T T. Highly effecient transfection of Xanthomonas campestris by electroporation. Bot Bull Acad Sin (Taipei) 1991;32:197–203. [Google Scholar]
- 37.Zagallo A C, Wang C H. Comparative glucose catabolism of Xanthomonas species. J Bacteriol. 1967;93:970–975. doi: 10.1128/jb.93.3.970-975.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]