Abstract
Cryptosporidium parvum oocysts were stained with the fluorogenic dyes SYTO-9 and SYTO-59 and sorted by flow cytometry in order to determine whether the fluorescent staining intensity correlated with the ability of oocysts to infect neonatal CD-1 mice. Oocysts that did not fluoresce or that displayed weak fluorescent intensity when stained with SYTO-9 or SYTO-59 readily established infections in mice, whereas those oocysts that fluoresced brightly did not. Although fluorescent staining profiles varied among different batches of oocysts, a relative cutoff in fluorescent staining intensity that correlated with animal infectivity was observed for all batches.
Cryptosporidium parvum is now recognized as a frequent cause of waterborne disease in humans (1, 10, 16, 17, 20). A primary means of parasite transmission is via drinking water, through the use of untreated surface water, contaminated distribution systems, or water treatment facilities employing only chlorine disinfection protocols. Significant morbidity and mortality have been associated with outbreaks of this parasite, particularly in immunocompromised individuals and in children (10).
An ongoing challenge of detection and disinfection of Cryptosporidium spp. is the difficulty in determining whether a parasite is viable. The presence of dead parasites in finished water is of little concern for disease transmission. Animals have been used as surrogates for determining the infectious potential of C. parvum oocysts (13). However, the animal infectivity method is tedious, difficult, and expensive and is not readily amenable to normal laboratory analysis in the water industry. Several methods have been used to estimate the viability of parasites, including in vitro excystation (1, 6, 7, 25, 26), infection of cell lines (12, 24, 27), parasite morphology by light microscopy, the uptake or exclusion of fluorogenic dyes (4, 5, 8, 9, 23), and animal infectivity (2–5, 15, 18, 19, 21, 22, 29, 30). Other assays that allow determination of viability of C. parvum oocysts include immunomagnetic capture PCR (28) and fluorescence in situ hybridization (FISH) techniques (11, 32, 33). Of all of these methods, only animal infectivity provides direct information about the ability of the parasite to cause disease.
In previous studies, we examined the potential use of fluorescent nucleic acid binding dyes as indicators of C. parvum oocyst viability under different experimental conditions. We reported that the staining of C. parvum oocysts with the nucleic acid binding dyes SYTO-9 and SYTO-59 correlated with the viability of these organisms, with heat-killed oocyst preparations used as a positive control (4, 5). In the present study, we demonstrate that fluorescence intensity of SYTO-9- and SYTO-59-stained C. parvum oocysts directly correlates with animal infectivity.
Source of C. parvum oocysts.
The strain of C. parvum used in this study was originally isolated by Harley Moon (National Animal Disease Center, Ames, Iowa) and is referred to as the Iowa strain. C. parvum oocysts were isolated from the feces of infected neonatal Holstein calves by methods described elsewhere (19). Oocysts were used within 90 days of isolation in all experiments. In our studies, C. parvum oocysts are never exposed to 2.5% potassium dichromate or sodium hypochlorite, a common procedure, in order to minimize oxidative damage incurred on the oocysts by this treatment.
Determining C. parvum infections in neonatal CD-1 mice.
A neonatal mouse model was used to evaluate infectivity of C. parvum oocysts (14, 15). Breeding pairs of outbred CD-1 mice were obtained from Charles River Breeding Laboratories (St. Constant, Quebec, Canada). The animals were given food and water ad libitum and were housed in cages with covers fitted with a 0.22-μm-pore-diameter filter in a specific-pathogen-free (P-2 level) animal facility.
Oocyst doses were prepared from the stock or flow cytometer-sorted suspensions by serial dilution to obtain the required dose. The actual dose given to the mice was determined from quadruplicate hemocytometer counts of the stock suspension. Five-day-old neonatal mice were inoculated intragastrically with feeding needles containing a known number of oocysts suspended in 50 μl of deionized water. Two hours prior to infection, the neonatal mice (5 days old) were taken away from the mothers to ensure that their stomachs were empty and ready to receive the intragastric inoculum of C. parvum. In addition, neonates from multiple litters were pooled and randomly selected for infection, in order to minimize variability introduced by inherent resistance or susceptibility of neonatal littermates to infection with C. parvum. The infectivity of the oocysts was determined 7 days after infection.
Two methods of detection of parasites in exposed mice were employed in the experiments. The first method was microscopic examination of mouse intestinal homogenates, by procedures described elsewhere (19). In the second method, C. parvum infections were evaluated by staining mouse intestinal homogenates with fluorescein-labeled anti-C. parvum monoclonal antibody (Immucell, Portland, Maine) and by flow cytometry to detect the presence of fluorescent oocysts (FACSCalibur; Becton Dickinson). Briefly, mice were killed by cervical dislocation, and the lower half of the small intestine was removed and placed in 10 ml of deionized water. The intestines were homogenized for 45 to 60 s in a Sorvall Omni-Mixer and washed by centrifugation at 2,000 × g for 15 min. The pellet was then washed once with deionized water containing 0.01% Tween 20 at 2,000 × g for 15 min. The supernatant was discarded, and the cell pellet was disrupted by vigorous mixing. Twenty microliters of the viscous pellet was pipetted into a 35-μm-pore-diameter sieve fitted onto a 6-ml flow cytometer polystyrene tube (Becton Dickinson), and the sieve was flushed with 450 μl of 1% bovine serum albumin (BSA; fraction V; Boehringer Mannheim) in phosphate-buffered saline (PBS). The strained suspension was incubated for 15 min at room temperature in order to block nonspecific adsorption of monoclonal antibodies to intestinal contents. Homogenates were stained with a 1:2,000 final dilution of fluorescein-labeled anti-C. parvum monoclonal antibody (Immucell) at 37°C for 30 min. Detection of C. parvum oocysts was done with a FACSCalibur flow cytometer programmed under the following settings: (i) forward side scatter photodiode setting = E00 and amp gain = 4.00; (ii) side scatter photomultiplier setting = 402, and (iii) FL1 photomultiplier setting = 470 (Fig. 1). Fifty thousand events were collected for each sample. A stock oocyst suspension was used to define a region based on size (i.e., forward light scatter) and internal complexity (i.e., side scatter) of C. parvum oocysts. This defined region (region 1) was subsequently used to discriminate potential oocysts in mouse intestinal homogenates. An additional criterion (i.e., gate) within this region was defined based on the fluorescent staining intensity (i.e., FL1) of particles within this region.
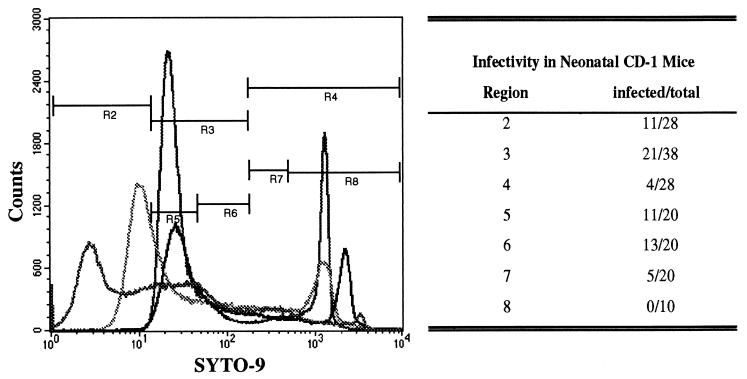
FIG. 1.
Comparison between SYTO-9 staining and infectivity of C. parvum oocysts. Oocysts were stained with SYTO-9 and sorted with the flow cytometer based on their fluorescence intensity (regions 2 to 8 [R2 to R8, respectively]). Sorted oocysts were used to infect neonatal CD-1 mice (100 oocysts/mouse). The ability of the sorted oocysts to establish infections in the neonatal mice is shown in the table next to the fluorogram. The results are from four independent sorting trials with four different batches of oocysts done on different days. The results demonstrate that although staining profiles may vary from batch to batch, the ability of stained oocysts to cause infections in neonatal mice is stable.
Evaluation of nucleic acid staining of C. parvum oocysts.
Staining of C. parvum oocysts (107) was done in a total volume of 500 μl of deionized water containing 10 μM SYTO-9 or 60 μM SYTO-59. All staining procedures were done with microcentrifuge tubes at 37°C for 30 min (SYTO-9) or 1 h (SYTO-59). Unbound dye was washed out by centrifuging the parasite suspension (10,000 × g for 10 min), removing the supernatant, and resuspending the parasite pellet in 1 ml of deionized water. Parasites were washed twice to remove unbound dye before oocysts were analyzed by flow cytometry. Analysis of C. parvum oocysts was done based on three parameters: (i) forward light scatter = voltage E00, amp gain of 4.00; (ii) side light scatter = voltage 402, amp gain of 4.00; and (iii) fluorescent intensity (FL1 for SYTO-9 = voltage of 480, amp gain of 1.00; or FL3 for SYTO-59 = voltage of 675, Amp gain of 1.00).
To ensure that daily fluctuations in the flow cytometer were controlled, calibrations on the flow cytometer were done daily with Calibrite beads (Becton Dickinson). “Channel targeting” was used as a method to ensure that samples obtained on different days were analyzed with identical instrument performances. Channel targeting was accomplished with fluorescein isothiocyanate (FITC) or PerCP-labeled Calibrite beads (Becton Dickinson). Beads were run on the flow cytometer and targeted to a mean channel setting in the appropriate fluorescence spectrum (e.g., FL1 = 98.93 or FL3 = 1,267.55). These mean channel values were used as target settings for the flow cytometer. Daily fluctuations in flow cytometer performance were corrected by adjusting instrument settings (i.e., photodiode and photomultiplier tube settings), so that similar target settings were consistently achieved. Lot-to-lot variation in Calibrite beads was also accounted for by comparing channel target settings between bead lots. After channel targeting was done, samples containing oocysts were subsequently run on the flow cytometer. Deionized water was used as the sheath fluid for experiments involving SYTO-9 or SYTO-59 dye staining. All samples were analyzed at a high flow rate through the flow cytometer (approximately 60 μl/min).
Oocysts stained with SYTO-9 or SYTO-59 displayed heterogeneous staining patterns, with a spectrum of fluorescent intensities observed among individual oocysts in the population (Fig. 1 and 2). The majority of oocysts within a given preparation remained unstained or weakly stained (regions 2 and 3 in Fig. 1 and 2); oocysts were thought to be viable based on our previous reports characterizing SYTO dye fluorescent staining intensities by using epifluorescence microscopy (4, 5). The proportion of unstained or weakly stained oocysts ranged from 75 to 95%, depending on the batch of C. parvum oocysts. Although fluorescent profiles were consistent when a single batch of oocysts was stained (data not shown), staining profiles varied when different batches of oocysts were stained with SYTO dyes (Fig. 1 and 2).
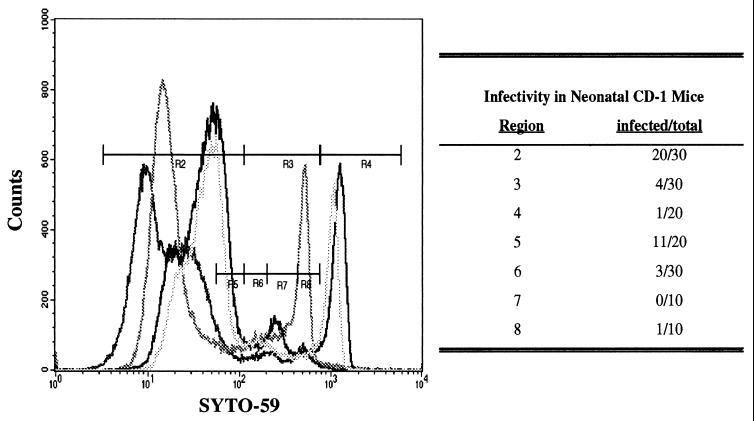
FIG. 2.
Relationship between SYTO-59 staining and infectivity of C. parvum oocysts. Oocysts were stained with SYTO-59 and sorted on the flow cytometer on the basis of their fluorescence intensity (regions 2 to 8 [R2 to R8, respectively]). Sorted oocysts were used to infect neonatal CD-1 mice (100 oocysts/mouse). The ability of the sorted oocysts to establish infections in the neonatal mice is shown in the table next to the fluorogram. The results are from four independent sorting trials with four different batches of oocysts done on different days. The results demonstrate that although staining profiles may vary from batch to batch, the ability of stained oocysts to cause infections in neonatal mice is stable.
Oocysts of various staining intensities were sorted into a cell concentrator module (Becton Dickinson) containing a 25-mm-diameter tissue culture insert (pore size, 1 μm; Becton Dickinson). The tissue culture insert had been previously incubated with 1% BSA–PBS for 30 min at room temperature to block adsorption of the parasites to the membrane and ensure efficient recovery of the parasites. Oocysts were sorted at a low flow rate (∼12 μl/min) by using an exclusion sort mode. Because sorting accuracy by flow cytometry approximates 95%, a reduced infectivity approach was used to identify a relative cutoff in fluorescence staining that could classify a subpopulation of oocysts as noninfectious or infectious. Theoretically, a sort of noninfectious oocysts may contain as many as 5% contaminating infectious oocysts, resulting in the occasional mouse becoming infected. For this reason, the experimental approach required that regions in flow cytometry be defined on the basis of a reduced infective potential and not absolute infectivity.
A portion of the stained and flow cytometer-sorted oocysts was used to inoculate neonatal CD-1 mice for infectivity analysis. Unstained oocysts were highly infectious to neonatal CD-1 mice (region 2, Fig. 1 and 2). As few as 10 weakly stained sorted oocysts caused infections in neonatal CD-1 mice (data not shown). Brightly stained oocysts (region 4, Fig. 1 and 2) were considerably less infectious than unstained or weakly stained oocysts.
Although oocysts that stained brightly with SYTO dyes were not infectious to neonatal mice, it was possible that binding of the dyes to sporozoite DNA (or RNA) impaired the ability of these parasites to replicate within host cells. Thus, the relationship we observed between infectivity and SYTO dye staining may be an experimental artifact due to the mutagenic properties of the dyes. However, we consistently observed that highly fluorescent oocysts were smaller and had greater internal complexity than unstained oocysts (Fig. 3). We subsequently sorted unstained oocysts, based on their size and internal complexity only, and infected neonatal mice. The smaller and more compact oocysts (oocysts that fluoresce brightly when stained with SYTO dyes) did not infect neonatal CD-1 mice (Fig. 3), whereas the larger and less compact oocysts (oocysts that do not fluoresce when stained with SYTO dyes) readily established infections in these mice (Fig. 3). These results indicate that the inability of intensely stained oocysts to infect neonatal mice was not a result of SYTO dyes binding to nucleic acids and impairing cellular function or parasite replication within host cells. Furthermore, it would appear that death of an oocyst is accompanied by physiological changes mediating compaction of the oocyst.
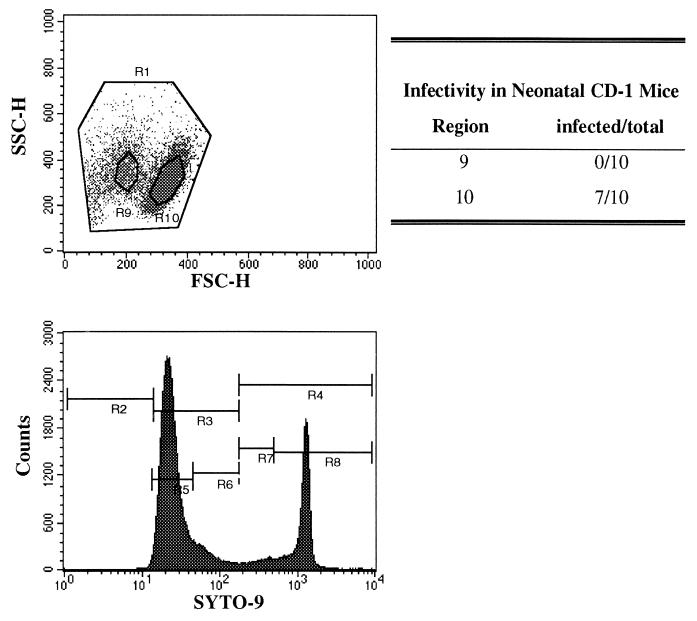
FIG. 3.
Results showing that the observed relationship between SYTO-9 staining and infectivity of C. parvum oocysts is not an indirect result of the binding of SYTO-9 to cellular nucleic acids. The events in region 9 (R9) shown on the left dot plot, represent oocysts that fluoresced brightly when stained with SYTO-9 (oocysts obtained from region 4 [R4], lower fluorescence histogram). The events in region 10 (R10) represent oocysts that stained weakly with SYTO-9 (oocysts obtained from region 3 [R3], lower fluorescence histogram). Regions 9 and 10 were drawn around these foci and subsequently used as sorting gates for an unstained preparation of C. parvum oocysts. Oocysts falling into these regions were sorted, and 100 oocysts were administered to neonatal CD-1 mice. The infectivity results of these two regions are shown to the right of the dot plot. FSC-H, forward light scatter (size); SSC-H, side light scatter (internal complexity).
These results were consistently observed between different batches of oocysts (four different batches used in these experiments [Fig. 1 and 2]). Although flow cytometric profiles of SYTO dye-stained oocysts varied from batch to batch, a relative cutoff in fluorescence intensity of stained parasites could be used as an objective criterion for classifying an oocyst as being infectious or noninfectious.
Evaluation of nucleic acid staining by using confocal microscopy.
Confocal microscopy was done with a Molecular Dynamics 2001 confocal microscope (Sunnyvale, Calif.). The instrument settings for all confocal analyses were as follows: ×100 objective lens, excitation wavelength of 488, 565 beamsplitter, 530 DF 30 filter for FITC detection, photomultiplier gain of 700 V, and pinhole 100 μm in diameter. The fluorescence intensity was measured with Image Space 3.10 software. The mean fluorescence intensity of oocysts within the sorted population was determined by overlaying a circle (diameter of 6 μm) on individual oocysts in confocal images, and their fluorescent intensity per image pixel was measured. Ten oocysts were randomly chosen from these images to approximate the mean fluorescent intensity of oocysts in sorted populations. Confocal microscopy verified the observation that more intensely stained oocysts were less infectious than unstained or weakly stained oocysts (Fig. 4). Noninfectious oocysts had bright internal staining of the sporozoites within the oocysts.
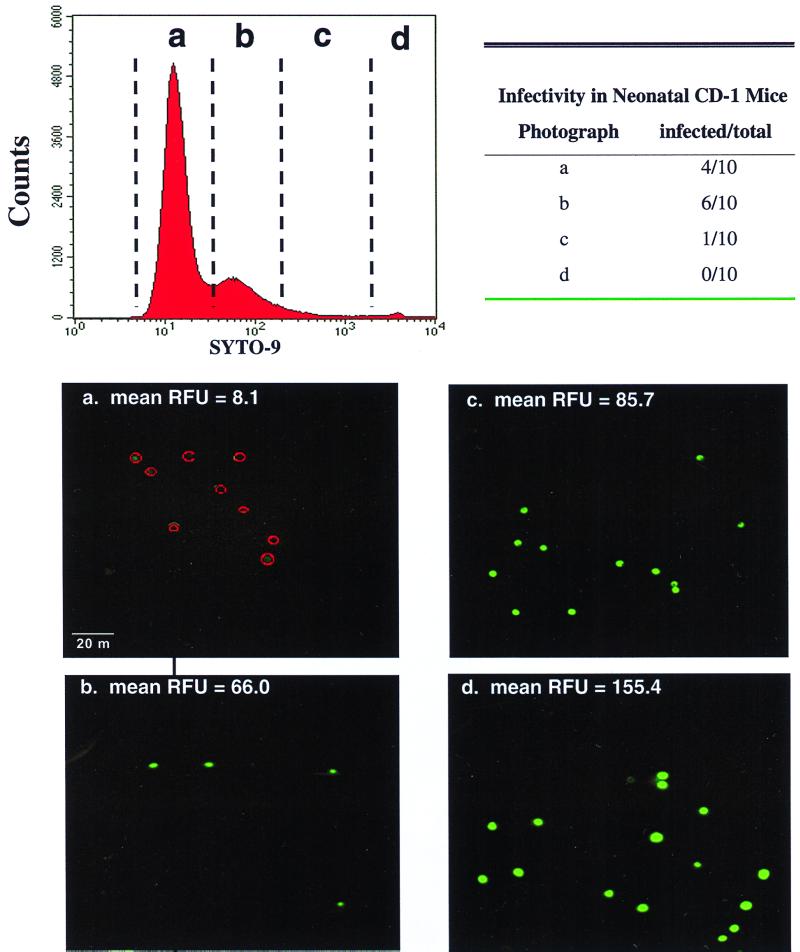
FIG. 4.
Confocal image of sorted oocysts. SYTO-9-stained C. parvum oocysts were sorted by flow cytometry, and split samples were used to infect neonatal CD-1 mice or to visualize fluorescent staining profiles by confocal microscopy. Panels a to d represent the different oocyst subpopulations sorted by flow cytometry. Red circles in panel a indicate where unstained oocysts were located. The level of staining (relative fluorescence units [RFU]) was quantified by confocal microscopy. The infective potentials of the oocysts depicted in the confocal micrographs are listed in the table.
Application to the water industry.
The direct correlation between animal infectivity and SYTO dye staining may be useful as a technique for identifying infectious C. parvum oocysts in source drinking waters. SYTO-59 may be particularly useful for this application, since its fluorescence spectrum does not overlap with that of FITC, and therefore it may be used in conjunction with commercially available FITC-labeled anti-C. parvum monoclonal antibodies to detect and determine the viability or infectivity of oocysts in environmental samples. At present, we are evaluating the staining characteristics of different strains and genotypes of C. parvum.
Recently, several other methods have been developed to estimate the viability and/or infectivity of C. parvum oocysts. In vitro excystation is commonly used as an indicator of C. parvum viability (1, 7, 23). However, its applicability for detection of viable oocysts in natural waters is limited, due to the abundance and diversity of microorganisms within natural water samples and, conversely, the small numbers of C. parvum oocysts found in these samples. Nevertheless, in vitro excystation is still used to estimate the viable numbers of parasites in stock oocyst preparations and to determine the efficacy of inactivation of oocysts after chemical inactivation (7, 15, 23, 25). Although in vitro excystation is used as a measure of oocyst viability, viability does not necessarily equate to infectivity. In fact, in vitro excystation has been shown to repeatedly overestimate the infectivity of C. parvum oocysts (7, 15). Moreover, it is assumed that intact oocysts that remain after excystation are nonviable and therefore are not infectious. Conversely, those oocysts that do excyst are viable and therefore capable of establishing infections. We have recently demonstrated that intact oocysts isolated by flow cytometry, after in vitro excystation, are capable of establishing infections in neonatal CD-1 mice (data not shown).
Because in vitro excystation is primarily used as a measure of viability and not infectivity, several researchers have used excystation in conjunction with in vitro infection of immortalized mammalian cell lines to assess the infectivity of C. parvum oocysts. In these studies, oocysts are excysted and the suspension containing infective sporozoites is inoculated into cell culture. Infection and replication within cells in vitro can be measured with different assays. Slifko et al. (27) have recently developed a focus detection method for determination of the infectivity of excysted parasites. In this assay, infected cell cultures are stained with an anti-C. parvum sporozoite-merozoite rabbit polyclonal antibody, and the number of infectious foci in cell cultures was determined by using an indirect fluorescent antibody assay. Rochelle et al. (24) use reverse transcriptase-PCR to detect the C. parvum heat shock protein 70 (hsp70) in infected cell cultures. Di Giovanni et al. (12) used PCR to detect the hsp70 gene of C. parvum in infected cell lines. Although all of these assays measure the establishment and infectivity of the excysted parasites, there are several potential problems with the in vitro cell culture infectivity assays. Since excystation is used as a method for obtaining infectious sporozoites, limitations in the excystation procedure are added to the evaluation sensitivity of the in vitro cell culture infectivity assays. Thus, oocysts that do not excyst, but are infectious, cannot be detected by these assays. Moreover, oocyst preparations require some degree of activation and sterilization (i.e., exposure to bleach to prevent bacterial contamination of the cell cultures) prior to the addition of the parasites to the cell cultures. Arguably, sterilization procedures may adversely affect oocyst viability and/or infectivity. In addition, it has been demonstrated that various cell lines display different degrees of susceptibility to infection with C. parvum (31), and the correlation between in vitro infectivity and animal infectivity has yet to be established.
Dye permeability assays using fluorogenic dyes such as 4′6-diamidino-2-phenylindole (DAPI) and propidium iodide (PI) have also been used as indicators of C. parvum oocyst viability (8, 9). Although DAPI-PI staining of C. parvum oocysts correlates with in vitro excystation (8), a direct correlation between animal infectivity and DAPI-PI is yet to be established.
Recently, a FISH technique has also been developed that shows considerable promise as an indicator of C. parvum oocyst viability (11, 32, 33). In these assays, a fluorescent DNA probe is targeted to the 18S rRNA of C. parvum. The basis for this assay is the premise that 18S rRNA is usually present in viable organisms and is degraded by cellular RNases in dead or dying cells. A distinct advantage of the FISH technique over SYTO-9 or SYTO-59 staining is the apparent high degree of species specificity: the ability to distinguish viable C. parvum from other Cryptosporidium species. However, unlike the SYTO dyes, existing FISH techniques are limited to measuring the viability of C. parvum oocysts and not their infectivity.
At present, we are evaluating whether SYTO dyes can be used in conjunction with flow cytometry to determine the levels of inactivation of C. parvum oocysts after exposure to various chemical disinfectants. Since the precise mechanisms of inactivation of protozoan cysts or oocysts are not known, it is premature to assume that nucleic acid dyes may be useful as indicators of oocyst viability or infectivity following chemical disinfection. Chemical disinfectants may potentially alter cell wall integrity, mediating changes in permeability of the dyes into oocysts (i.e., increasing or decreasing fluorescence intensity). The use of the nucleic acid dyes as indicators of viability or infectivity for chemically inactivated parasites will require more research to determine the effects of each disinfectant on oocyst staining intensity.
Acknowledgments
We thank Cezary Kucharsky and Shannon Lefevbre for technical assistance.
This work was supported by the American Water Works Association Research Foundation (AWWARF) and by the Natural Sciences and Engineering Council of Canada (NSERC) to M.B. and G.R.F. L.L.G. was supported by an NSERC postdoctoral fellowship, and N.F.N. was supported by a Province of Alberta doctoral fellowship.
REFERENCES
- 1.Barer M R, Wright A E. A review: Cryptosporidium in water. Lett Appl Microbiol. 1990;11:271–277. [Google Scholar]
- 2.Belosevic M, Faubert G M. Giardia muris: correlation between oral dosage, course of infection and trophozoite distribution in the mouse small intestine. Exp Parasitol. 1983;56:93–100. doi: 10.1016/0014-4894(83)90100-5. [DOI] [PubMed] [Google Scholar]
- 3.Belosevic M, Faubert G M, MacLean J D, Law C, Croll N A. Giardia lamblia infections in Mongolian gerbils: an animal model. J Infect Dis. 1983;147:222–226. doi: 10.1093/infdis/147.2.222. [DOI] [PubMed] [Google Scholar]
- 4.Belosevic M, Guy R A, Taghi-Kilani R, Neumann N F, Gyürék L L, Liyanage L R J, Millard P J, Finch G R. Nucleic acid stains as indicators of Cryptosporidium parvum oocyst viability. Int J Parasitol. 1997;27:787–798. doi: 10.1016/s0020-7519(97)00033-7. [DOI] [PubMed] [Google Scholar]
- 5.Belosevic M, Taghi-Kilani R, Guy R A, Neumann N F, Finch G R, Gyürék L L, Liyanage L R J. Vital dye staining of Giardia and Cryptosporidium. Denver, Colo: American Water Works Association; 1997. [Google Scholar]
- 6.Bingham A K, Meyer E A. Giardia excystation can be induced in vitro in acidic solutions. Nature. 1979;277:301–302. doi: 10.1038/277301a0. [DOI] [PubMed] [Google Scholar]
- 7.Black E K, Finch G R, Taghi-Kilani R, Belosevic M. Comparison of assays for Cryptosporidium parvum oocysts viability after chemical disinfection. FEMS Microbiol Lett. 1996;135:187–189. doi: 10.1111/j.1574-6968.1996.tb07987.x. [DOI] [PubMed] [Google Scholar]
- 8.Campbell A T, Robertson L J, Smith H V. Viability of Cryptosporidium parvum oocysts: correlation of in vitro excystation with inclusion or exclusion of fluorogenic vital dyes. Appl Environ Microbiol. 1992;58:3488–3493. doi: 10.1128/aem.58.11.3488-3493.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Campbell A T, Robertson L J, Smith H V. Effects of preservatives on viability of Cryptosporidium parvum oocysts. Appl Environ Microbiol. 1993;59:4361–4362. doi: 10.1128/aem.59.12.4361-4362.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Casemore D P, Wright S E, Coop R L. Cryptosporidiosis: human and animal epidemiology. In: Fayer R, editor. Cryptosporidium and cryptosporidiosis. Boca Raton, Fla: CRC Press; 1997. pp. 65–92. [Google Scholar]
- 11.Deere D, Vesey G, Milner M, Williams K, Ashbolt N, Veal D. Rapid method for fluorescent in situ ribosomal RNA labelling of Cryptosporidium parvum. J Appl Microbiol. 1998;85:807–818. doi: 10.1046/j.1365-2672.1998.00589.x. [DOI] [PubMed] [Google Scholar]
- 12.Di Giovani G, LeChevallier M, Battigelli E, Campbell A, Abbaszagedan M. Proceedings of the AWWA Water Quality and Technology Conference. Denver, Colo: American Water Works Association; 1998. Detection of Cryptosporidium parvum oocysts recovered from environmental water samples using immunomagnetic separation (IMS) and integrated cell culture-PCR (CC-PCR) [Google Scholar]
- 13.Enriquez F J, Sterling C A. Cryptosporidium infections in inbred strains of mice. J Protozool. 1991;38:100S–102S. [PubMed] [Google Scholar]
- 14.Ernest J A, Blagburn B L, Lindsay D S, Current W L. Infection dynamics of Cryptosporidium parvum (Apicomplexa: Cryptosporiidae) in neonatal mice (Mus musculus) J Parasitol. 1986;72:796–798. [PubMed] [Google Scholar]
- 15.Finch G R, Black E K, Gyürék L L, Belosevic M. Ozone inactivation of Cryptosporidium parvum in demand-free phosphate buffer determined by in vitro excystation and animal infectivity. Appl Environ Microbiol. 1993;59:4203–4210. doi: 10.1128/aem.59.12.4203-4210.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gallaher M M, Herndon J L, Nims L J, Sterling C R, Grabowski D J, Hull H F. Cryptosporidiosis and surface water. Am J Public Health. 1989;79:39–42. doi: 10.2105/ajph.79.1.39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Grimason A M, Smith H V, Smith P G, Jackson M E, Girdwood R W A. Waterborne cryptosporidiosis and environmental health. Environ Health. 1990;98:228. [Google Scholar]
- 18.Gyürék L L, Finch G R, Belosevic M. Modeling chlorine inactivation requirements of Cryptosporidium parvum oocysts. J Environ Eng. 1997;123:865–875. [Google Scholar]
- 19.Gyürék, L. L., H. Li, M. Belosevic, and G. R. Finch. Ozone inactivation kinetics of Cryptosporidium parvum in oxidant demand-free phosphate buffer. J. Environ. Eng., in press. [DOI] [PMC free article] [PubMed]
- 20.Hayes E B, Matte T D, O'Brien T R, McKinley T W, Logsdon G S, Rose J B, Ungar B L P, Word D M, Pinsky P F, Cummings M L, Wilson M A, Long E G, Hurwitz E S, Juranek D D. Large community outbreak of cryptosporidiosis due to contamination of a filtered public supply. N Engl J Med. 1989;320:1372–1376. doi: 10.1056/NEJM198905253202103. [DOI] [PubMed] [Google Scholar]
- 21.Labatiuk C W, Finch G R, Belosevic M. Proceedings of the Water Quality Technology Conference. Denver, Colo: American Water Works Association; 1991. Comparison of a fluorogenic dye and animal infectivity for viability determination of Giardia following ozone disinfection; pp. 789–803. [Google Scholar]
- 22.Labatiuk C W, Schaefer III F W, Finch G R, Belosevic M. Comparison of animal infectivity, excystation, and fluorogenic dye as measures of Giardia muris cyst inactivation by ozone. Appl Environ Microbiol. 1991;57:3187–3192. doi: 10.1128/aem.57.11.3187-3192.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Robertson L J, Campbell A T, Smith H V. Survival of Cryptosporidium parvum oocysts under various environmental pressures. Appl Environ Microbiol. 1992;58:3494–3500. doi: 10.1128/aem.58.11.3494-3500.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rochelle P A, De Leon R, Stewart M H, Wolfe R L. Comparison of primers and optimization of PCR conditions for detection of Cryptosporidium parvum and Giardia lamblia in water. Appl Environ Microbiol. 1997;63:106–114. doi: 10.1128/aem.63.1.106-114.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rose J B. Occurrence and control of Cryptosporidium in drinking water. In: McFeters G A, editor. Drinking water microbiology. New York, N.Y: Springer-Verlag; 1990. pp. 294–321. [Google Scholar]
- 26.Schaefer F W., III . Methods for excystation of Giardia. In: Meyer E A, editor. Giardiasis. Vol. 3. Amsterdam, The Netherlands: Elsevier; 1990. pp. 111–136. [Google Scholar]
- 27.Slifko T R, Friedman D, Rose J B, Jakubowski W. An in vitro method for detecting infectious Cryptosporidium oocysts with cell culture. Appl Environ Microbiol. 1997;63:3669–3674. doi: 10.1128/aem.63.9.3669-3675.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Stinear T, Matusan A, Hines K, Sandery M. Detection of a single viable Cryptosporidium parvum oocyst in environmental water concentrates by reverse transcription-PCR. Appl Environ Microbiol. 1996;62:3385–3390. doi: 10.1128/aem.62.9.3385-3390.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Taghi-Kilani R, Gyürék L L, Liyanage L, Guy R A, Finch G R, Belosevic M. Vital dye staining of Giardia and Cryptosporidium. In Chlorine dioxide: drinking water, process water, and wastewater issues. 1995. Proceedings of the Third International Symposium. American Water Works Association, Denver, Colo. [Google Scholar]
- 30.Taghi-Kilani R, Gyürék L L, Millard P J, Finch G R, Belosevic M. Vital dyes as indicators of Giardia muris viability following cyst inactivation. Int J Parasitol. 1996;26:437–446. doi: 10.1016/0020-7519(96)00033-1. [DOI] [PubMed] [Google Scholar]
- 31.Upton S J. In vitro cultivation of Cryptosporidium. In: Fayer R, editor. Cryptosporidium and cryptosporidiosis. Boca Raton, Fla: CRC Press; 1997. pp. 181–208. [Google Scholar]
- 32.Vesey G, Deere D, Dorsch M, Veal D, Williams K, Ashbolt N. Proceedings of the International Symposium on Waterborne Cryptosporidium. Denver, Colo: American Water Works Association; 1997. Fluorescent in-situ labeling of viable Cryptosporidium parvum in water samples; pp. 21–29. [Google Scholar]
- 33.Vesey G, Ashbolt N, Fricker E J, Deere D, Williams K L, Veal D A, Dorsch M. The use of a ribosomal RNA targeted oligonucleotide probe for fluorescent labeling of viable Cryptosporidium parvum oocysts. J Appl Microbiol. 1998;85:429–440. doi: 10.1046/j.1365-2672.1998.853496.x. [DOI] [PubMed] [Google Scholar]