Abstract
Motivation
DNA methylation is a key epigenetic factor regulating gene expression. While promoter methylation has been well studied, recent publications have revealed that functionally important methylation also occurs in intergenic and distal regions, and varies across genes and tissue types. Given the growing importance of inter-platform integrative genomic analyses, there is an urgent need to develop methods to discover and characterize gene-level relationships between methylation and expression.
Results
We introduce a novel sequential penalized regression approach to identify methylation-expression quantitative trait loci (methyl-eQTLs), a term that we have coined to represent, for each gene and tissue type, a sparse set of CpG loci best explaining gene expression and accompanying weights indicating direction and strength of association. Using TCGA and MD Anderson colorectal cohorts to build and validate our models, we demonstrate our strategy better explains expression variability than current commonly used gene-level methylation summaries. The methyl-eQTLs identified by our approach can be used to construct gene-level methylation summaries that are maximally correlated with gene expression for use in integrative models, and produce a tissue-specific summary of which genes appear to be strongly regulated by methylation. Our results introduce an important resource to the biomedical community for integrative genomics analyses involving DNA methylation.
Availability and implementation
We produce an R Shiny app (https://rstudio-prd-c1.pmacs.upenn.edu/methyl-eQTL/) that interactively presents methyl-eQTL results for colorectal, breast and pancreatic cancer. The source R code for this work is provided in the Supplementary Material.
Supplementary information
Supplementary data are available at Bioinformatics online.
1 Introduction
DNA methylation involves the addition of a methyl group to cytosine residues, predominantly in the context of CpG dinucleotides in a DNA sequence. It is among the best studied epigenetic modifications, plays an important role in the regulation of gene transcription, and is associated with numerous key biological processes and diseases.
The increasing availability of multi-platform genomics and epigenomics datasets in cancer and other diseases has motivated researchers to develop unified modeling frameworks to perform integrative analyses relating the various platforms to each other, such as iBAG (integrative Bayesian Analysis of Genomics Data) (Wang et al., 2013; Jennings et al., 2013) and iCluster (integrative clustering) (Shen et al., 2009), which decompose gene expression into components explained by upstream genetic and epigenetic platforms (e.g. copy number, methylation and miRNA) and then incorporate these components as predictors in a clinical model. These integrative models require calculation of gene-level summaries for each genomic platform. For a platform such as copy number, it is relatively easy to come up with a reasonable strategy for computing gene level summaries (e.g. average copy number in coding region of gene), but a simple strategy like this may not work well for platforms like methylation that affect expression in more complex and subtle ways. In existing literature, we have encountered two strategies for constructing gene-level methylation summaries: (i) computing the average methylation level across all probes located in the gene’s promoter region, or (ii) using the methylation level for the single probe that appears to be most negatively correlated with gene expression. While reasonable, both of these strategies appear to be simplistic and could miss the most important epigenetic effects for a given gene.
This can be seen in a study of methylation and expression of two clinically relevant genes in colorectal cancer (CRC), AREG and EREG, that motivated this work (Lee et al., 2016). Initially, their integrative analysis of methylation and expression focused on the single CpG site within the promoter region of each gene that was identified to be most negatively correlated with gene expression based on a pan-cancer analysis spanning all cancers in The Cancer Genome Atlas (TCGA). Fitting the iBAG model to this data, of expression variability was explained by methylation for EREG, while for AREG, less than 10% of gene expression was found to be explained by methylation in the CRC samples. Upon further examination, cg03244277 was the CpG site whose methylation was most correlated with AREG expression when looking across all cancers, but for CRC the methylation at this site had very little association with AREG expression. However, the methylation for two other sites, cg02334660 and cg26611070, had strong association with AREG expression (34% and 35% of expression variability explained, respectively) in CRC, and one of these markers was located in the gene body of AREG, not the promoter region. This demonstrated that (i) functionally important methylation can vary across cancer types, (ii) not all important methylation occurs in the promoter region of the gene and (iii) the choice of any single CpG to represent methylation level for a given gene has the potential to miss important signals. If the original CpG derived from the pan-cancer analysis was utilized, it would have resulted in an incorrect conclusion that AREG is not strongly regulated by methylation. This motivated the development of a more comprehensive strategy for studying associations of methylation and expression in a tissue-specific manner, not restricting focus to promoter regions, and allowing for the potential of multiple important CpG sites per gene, to ensure that important discoveries are not missed.
Initial studies of methylation focused primarily on promoter-associated CpG islands, which are CpG-rich short regions found within the promoter regions of 70% human genes (Laurent et al., 2010) that are frequently linked with gene silencing. However, with the advent of high-throughput DNA methylation profiling methods that survey a more extensive set of CpG sites, it has become clear that the relationship between DNA methylation and gene transcription is much more complicated than previously expected (Teschendorff and Relton, 2018). For instance, the comprehensive high-throughput array-based relative methylation analysis of human colon cancer methylome showed that methylation in CpG shores, flanking regions up to 2 kb away from CpG islands, is strongly related to gene expression (Irizarry et al., 2009). Extensive positive correlations between gene body methylation and gene expression have been observed and reported in multiple genome-wide studies of epigenomic and gene expression data (Kulis et al., 2013). Moreover, recent studies have suggested regulatory roles for CpGs in distal elements such as enhancers (Aran et al., 2013; Kulis et al., 2013; Wagner et al., 2014; Yao et al., 2015; Rhie et al., 2016). These recent findings indicate that regulatory methylation sites do not occur exclusively in CpG islands or gene promoters, and the effect of DNA methylation on gene expression can depend strongly on the genomic location of the CpG with respect to the gene body (Thingholm et al., 2016).
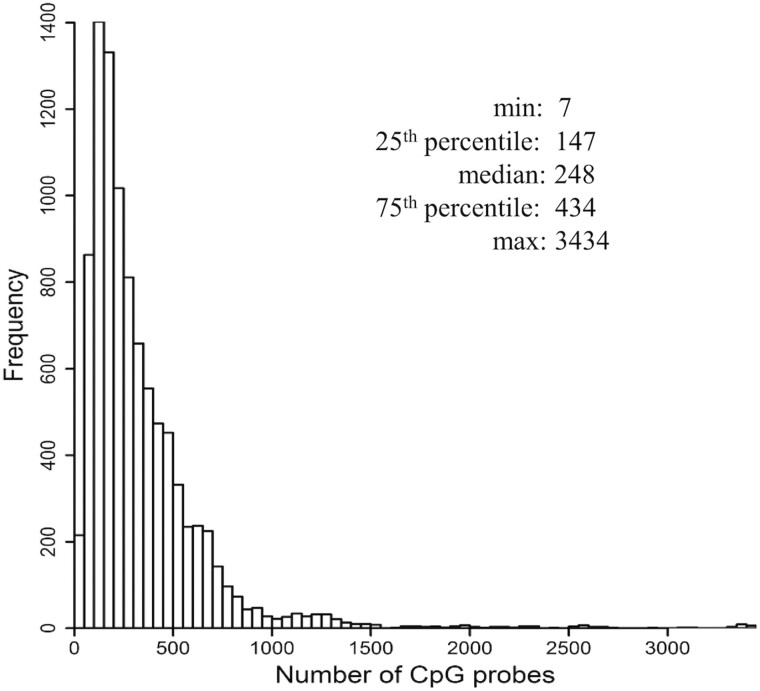
One simple approach to study the association of methylation and gene expression is to look at CpG sites one at a time, e.g. by calculating marginal correlations between gene expression and methylation of each CpG site in some neighborhood of the gene, ideally expanding to the flanking regions outside the gene body by at least several hundred kilobases to capture potential distal regulatory elements (Aran et al., 2013). As shown in Figure 1, for the Illumina 450k array, there are often hundreds to thousands of CpGs inside or within ±500 kb flanking a gene. While this simple approach can reveal important insights, it might fail to detect some of the true associations because the surveying of such a large number of CpGs per gene requires adjustment for multiple testing, and one-at-a-time analyses ignore the fact that multiple methylation sites can work together to regulate gene transcription in a coordinated manner rather than independently (Denis and Tadesse, 2016). In particular, Zhong et al. (2019) investigated different models to predict gene expression from DNA methylation, and found that compared to simple linear regression models that use any given CpG site alone, simultaneously utilizing all CpG sites in the gene region as predictors substantially improved the model’s predictive performance.
Fig. 1.
Distribution of the number of CpG probes located within ±500 kb of the gene region. For the 9569 genes satisfying our selection criteria, the distribution of the number of CpG probes located within the gene body or the flanking region of ±500 kb on either end from the Illumina 450 K methylation array per gene is shown
Under the assumption that functional importance of a particular CpG is often related to its relative location within or around the gene (Thingholm et al., 2016), several studies (Brenet et al., 2011; Lou et al., 2014; Rhee et al., 2013; VanderKraats et al., 2013; Schlosberg et al., 2017) have investigated the combinatorial effect of methylation of different components of a transcription unit on gene expression, and constructed quantitative models to predict gene expression based on methylation of different genomic regions. However, the quantitative relationships between methylation of different genomic regions and gene expression could greatly differ across genes in the genome. Moreover, these studies limit their analyses to CpGs in the gene body and very close to the gene body (1–2 kb), whereas more distal CpGs, e.g. in enhancer regions, can also have an important effect on expression. It is also worth noting that methylation plays a fundamental role in regulating the expression levels for some genes, while it is a much less important factor for others. These general modeling approaches do not provide a summary of which genes are strongly regulated by CpG methylation.
To address these issues, we have developed a penalized regression approach to identify gene-specific methylation-expression quantitative trait loci (methyl-eQTLs). As eQTL analysis has been extensively performed in human genetic studies to identify genetic variants that explain gene expression, in our context, we coin the term ‘methyl-eQTLs’ to represent the methylation loci that are most strongly associated with downstream gene expression. Methyl-eQTLs consist of a tissue-specific, gene-specific set of CpG loci whose methylation status best predicts expression for that gene, and corresponding weights that indicate the relative importance and direction of their associations. We focus on cis methyl-eQTLs, which are CpG sites located inside the gene body or within 500 kb of the gene region, but utilize a sequential modeling strategy that incorporates biological information about which CpGs are a priori most likely to be associated with expression based on their characteristics. Methyl-eQTLs can be used to compute gene-specific methylation scores that are maximally correlated with gene expression, and provide a gene-specific measure of percentage of expression variability explained by methylation status. We stress that the contribution of this work is not merely the introduction of a technically novel sequential penalized regression approach that makes effective use of prior biological information. More importantly, we highlight the limitations of simplistic approaches commonly used in the literature for methylation-expression integration, and strongly advocate performing variable selection to find a sparse set of CpGs from hundreds to thousands of potentially important ones located in a broader region around the gene, so as to better understand the cis-regulatory effects of methylation on gene expression and to provide a gene-specific measure of proportion of expression explainable by methylation.
In simulations that are constructed to mimic real data, our proposed sequential approach is shown to outperform alternative methods for integrating gene expression and DNA methylation in terms of finding a balance between predictive ability and model sparsity. We apply this approach to study methyl-eQTLs in the setting of CRC, using TCGA tumor samples to build the models, and assessing the performance in terms of sparsity and ability to explain expression variability using cross validation as well as an independent validation dataset consisting of tumor samples from The University of Texas M.D. Anderson Cancer Center (MDACC). We also apply this approach to identify methyl-eQTLs for breast cancer (BRCA) and pancreatic cancer (PAAD). Methyl-eQTL analysis is performed separately for each cancer since methylation patterns vary strongly across tissue types (Jones, 2012), and as illustrated by our AREG/EREG example, global modeling across all tissue types may miss out on relationships specific to a particular tissue type. We develop a freely available Shiny App containing the methyl-eQTLs of all genes in the genome for each of the 3 tumor types.
2 Material and methods
2.1 Data
Colorectal cancer data: To fit our model and find the methyl-eQTLs, we used TCGA primary solid tumor samples in colon (COAD) or rectal (READ) whose methylation data and matching gene expression data are both available, pooling them as a CRC cohort. There are 369 samples in total. To validate the methyl-eQTLs, we used a cohort of 163 tumor samples from primary resection specimens from colon or rectal cancer patients at MDACC.
DNA methylation data for both cohorts were generated from the Illumina Infinium HumanMethylation 450K BeadChip and quantified by beta values, which range from 0 to 1, with 0 indicating no methylation of a CpG site at either allele, 1 indicating methylation at both alleles. For gene expression data, level 3 RNASeqV2 gene-level expression data which were generated from the Illumina HiSeq platform were used for the TCGA cohort; gene expression data for the MDACC cohort were generated from Agilent microarrays.
The epigenome data generated from 17 primary solid tumor samples at MDACC were used to annotate the chromatin states of CpGs (Orouji et al., 2021). Based on enrichment patterns of six histone modification marks, we trained a ChromHMM model (Ernst and Kellis, 2010) that consists of 10 chromatin states. These data were used to assess whether the CpGs selected by our algorithm were enriched for any particular chromatin state.
Data of other cancer types: We also applied our approach to find methyl-eQTLs using TCGA primary solid tumor samples in pancreas or breast whose methylation data and matching gene expression data are both available. There are 782 breast cancer samples and 178 pancreatic cancer samples. For chromatin state annotation, we downloaded reference epigenome data from the matched tissue type generated by NIH Roadmap Epigenomics Program, and used an 18-state model learned by ChromHMM (Kundaje et al., 2015).
More detailed descriptions about these data and their preprocessing procedures can be found in Supplementary Section S1 in Supplementary Material.
2.2 Identification of methyl-eQTLs
2.2.1. Overview
Because methylation in distal regulatory elements may also be functionally related, we considered all CpGs located up to 500 kb upstream of TSS through 500 kb downstream of TES as potentially important. Previous studies show that distal regulatory elements are generally located within this neighborhood (Wagner et al., 2014; Aran et al., 2013), so we decided not to consider more distant sites to reduce false positives.
The selection of important CpGs for each gene can be cast as a linear regression variable selection problem. The lasso is a commonly used variable selection technique that utilizes a L1-norm penalty on the regression coefficients to provide a sparse solution (Tibshirani, 1996). The lasso considers all predictors equally in variable selection, while it is known that certain CpGs sites are a priori more likely to be functionally important than others depending on the CpG type and relative location to the gene body, as described in Section 1. To incorporate such prior knowledge, we propose a sequential, two-step variable selection approach. Our sequential lasso procedure (referred to as Seq-Lasso) implements lasso regression in two steps, first considering CpG sites that are most likely to be associated with gene expression a priori, which are determined by a preliminary genome-wide study, and then checking the rest of the CpG sites to see if any of those can further explain the variability in gene expression.
2.2.2. Notation and model
Let yik denote the gene expression data for individual i () at gene k (). Suppose that for gene k, there are Jk CpG probes located within the gene body or in the flanking ±500 kb on either end. Let xijk denote the methylation beta value for individual i at CpG site j () of gene k. A standard linear regression model is assumed to relate gene expression with methylation, i.e.
| (1) |
where is an vector of expression data, and Xk is an matrix of methylation data. For gene k, Seq-Lasso performs variable selection among the Jk associated CpGs to identify a sparse subset whose methylation can best predict expression. For these selected CpGs, the regression coefficients βjk can be interpreted as weights that indicate the direction (positive or negative) and strength (absolute magnitude) of the association between methylation at CpG site j and expression of gene k.
Let denote the epigenomic characteristics of the CpG site j with respect to the target gene k, including (i) its relative location to the gene k, (ii) the type of the CpG site j relative to the CpG island (CpG island, CpG shore, CpG shelf or open sea), (iii) the size of the gene k and (iv) the direction of marginal association ρjk between methylation at the CpG site j and expression of the gene k. Formal definitions of are given in Supplementary Section S2 in Supplementary Material.
2.2.3. Procedures
Step 1 : Conduct a preliminary whole-genome analysis to build a model to estimate the probability that a given CpG site is found to be associated with expression based on its epigenomic characteristics . We accomplish this by first applying simple lasso regression of the expression data on the methylation data Xk for each gene k to get an estimate , then modeling the probability of selection by lasso as a function of (Wood, 2006), i.e. . Calculation details of are given in Supplementary Section S2 in Supplementary Material.
Step 2 : For each gene k, apply lasso regression to the expression data and the methylation data of a subset of CpG sites that are considered a priori likely to be important based on our model in Step 1. We denote this subset by , which includes the CpG sites such that for some pre-chosen threshold π that could depend on the size of the gene k and the direction of marginal association ρjk between methylation at the CpG site j and expression of the gene k. Guidance on selection of π is provided in Supplementary Section S2 in Supplementary Material.
| (2) |
The regularization parameter λ1 is chosen by 5-fold cross validation. Denote the indices of the CpG sites with non-zero estimated coefficients among by . Regress on the methylation data in and get the residual vector .
Step 3 : For each gene k, apply lasso regression to the residual vector from Step 2 as the new response, and the methylation data of the CpG sites in , i.e.
| (3) |
The regularization parameter λ2 is once again chosen by 5-fold cross validation. Denote the indices of the CpG sites with non-zero estimated coefficients among by .
Step 4 : For each gene k, the regression model (1) is refitted by ridge regression only for the CpGs selected in Step 2 and 3.
Apply ridge regression to expression data and the methylation data of CpG sites in , i.e.
| (4) |
The regularization parameter λ3 is again chosen by 5-fold cross validation. Refitting the coefficients of CpG sites selected in Step 2 and 3 by ridge regression helps reduce the bias in estimation of large coefficients that plagues lasso regression (Zou, 2006).
Our Seq-Lasso gives priority to the CpG sites with characteristics that make them more likely to be selected as important based on information learned from the whole-genome analysis by letting them be selected first, and then other CpGs with characteristics that make them less likely to be important are only considered in the second step. This has several key benefits. First, it tends to lead to a parsimonious model that uses as few CpGs as possible in explaining the expression variability. Second, if multiple CpGs are correlated with each other and with expression, our approach will tend to select the one that is a priori more likely to be important (in ) based on its characteristics rather than just arbitrarily selecting one, as a single-step lasso would do, which may make them more likely to be functionally important. Together, these contribute to a model that is more likely to discover biologically interpretable and reproducible results.
3 Results
3.1 Selection probability functions
Figure 2 shows the results of our genome-wide analysis of CpG selection probability in terms of importance for predicting gene expression as a function of its epigenomic characteristics in CRC, which reveal very interesting patterns. First, selection probabilities are always much higher for methylation sites within or very close to the gene body compared to more distant regions, where the selection probability decreases monotonically and then reaches a plateau. This suggests that functionally related CpG sites tend to concentrate around the gene body, as expected. Second, a sharp peak occurs near the TSS in the probability curve for CpG sites whose methylation levels are negatively associated with gene expression for all CpG types and gene sizes. Again, this is not surprising, as one expects that methylation in or near the promoter region of a gene should be associated with gene silencing. In contrast, the selection probability curves for positively associated CpG sites do not show this pattern, but instead show higher probabilities across the gene body, in most cases highest near the TES. Third, when controlling for direction of association and gene size, differences between the selection probability curves across CpG types are evident. Methylation sites in CpG islands are least likely to be selected in all cases. These selection probability functions are shared together with the source code in Supplementary Material.
Fig. 2.

The probability of selection as a function of a CpG’s epigenomic characteristics in CRC. Each subfigure shows the probability that a CpG probe is selected (indicated by y-axis) as a function of its relative distance to the TSS and TES of the associated gene (indicated by x-axis), CpG type (CpG island, CpG shore, CpG shelf, open sea), gene size (small, medium, large), separately for CpGs negatively (top row) or positively (bottom row) associated with gene expression
We formally tested if CpG sites are significantly more likely to be chosen for certain CpG types and locations. Preliminary analyses suggested that the relationships among the CpG types vary across positively and negatively correlated CpG sites and inside/outside the gene body, but not by more specific location or gene size, so for simplicity we reported analyses aggregating over these factors. The odds ratios of selection for each pair of CpG types are shown in Figure 3. In all cases, CpG sites in CpG islands are least likely to be selected as important for predicting expression. Open seas are most likely to be selected for positively correlated CpG sites and all CpG sites outside the gene body, while CpG shores are most likely to be selected for negatively correlated CpG sites within the gene body. These formal testing results confirm the previous finding that functionally important methylation occurs more often in CpG shores than CpG islands in colon cancer (Irizarry et al., 2009), and also add to the growing body of literature on which CpG types are more likely to be related to gene expression.
Fig. 3.
The odds ratios of selection for each pair of CpG types. For every pair of CpG types, the odds ratios (OR) of selection are shown for CpGs (A) inside the gene body and (B) outside the gene body, with P-value testing whether OR = 1 (corrected for multiple testing) given in parentheses. In each table, the odds ratio of every pair of CpG types (CpG type in the column header as numerator; CpG type in the row header as denominator) for marginally positively correlated CpGs are shown in the upper half; the odds ratio of every pair of CpG types (CpG type in the row header as numerator; CpG type in the column header as denominator) for marginally negatively correlated CpGs are shown in the lower half. For example, for a marginally positively correlated CpG in the gene body, a CpG located in a shelf is 40% more likely to be selected than one from a CpG shore (OR = 1.40, P-value <1e-4); for marginally negatively correlated CpG sites in the gene body, a CpG located in the shelf is 18% less likely to be selected than one from a CpG shore (OR = 0.82, P-value =1e-4)
3.2 Performance comparison of Seq-Lasso with other methods
Next, we assessed the performance of Seq-Lasso relative to other methods for identifying methyl-eQTLs in terms of model sparsity and predictive accuracy. These alternative methods include Lasso which performs lasso regression on CpGs located in the gene body or 500 kb on either end, E-NET which performs elastic net regression (Zou and Hastie, 2005) on CpGs located in the gene body or 500 kb on either end, E-NET-local which performs elastic net regression on CpGs located in the gene body or up to 2 kb upstream of TSS, which performs iterative adaptive lasso (Sun et al., 2010) on CpGs located in the gene body or 500 kb on either end, which selects the single CpG that is most negatively correlated with gene expression among methylation sites in the gene body or up to 2 kb upstream of TSS, NegCor-all which selects the single CpG that is most negatively correlated with gene expression among methylation sites in the gene body or 500 kb on either end, and which uses the average methylation level of CpGs located in a 2 kb neighborhood centered at TSS to correlate with expression. Among these alternatives, Lasso, E-NET and E-NET-local are natural statistical learning strategies that one might try, and is a variable selection approach originally developed for genome-wide simultaneous multiple-loci mapping, all of which have not been used in existing literature for methylation-expression integration to our knowledge. Like Seq-Lasso, the models selected by Lasso, E-NET and E-NET-local are refitted using ridge regression to minimize attenuation of large coefficients. and are widely adopted in the literature to integrate methylation and gene expression, and NegCor-all can be viewed as an extended version of that allows us to select the most negatively correlated CpG from the same pool of methylation sites as Seq-Lasso, Lasso and E-NET.
3.2.1. Simulation study
We first conducted simulations to evaluate the performance of these methods.
Simulation setup: We randomly selected 500 genes with at least moderate correlations between gene expression and methylation in CRC data from the 9527 genes satisfying our selection criteria (which are described in Supplementary Section S1.1.5 in Supplementary Material). To construct simulations that mimic real data, we directly used the CRC methylation data of CpGs located in the gene body or 500 kb on either end from the TCGA cohort as the design matrix Xk for gene k, and generated gene expression data following the model (1), where the true regression coefficient and the error variance are obtained from fitting Lasso to the real expression and methylation data in the same cohort. Such simulation design preserves the complex patterns of correlations among CpGs observed in real methylation data, and closely mimics the underlying dependence structure of ‘signals’ (i.e. functionally relevant methylation) on various epigenomic characteristics, and does not give our proposed Seq-Lasso any unfair advantage. We simulated 100 replicate datasets per gene.
Simulation results: For each simulated dataset, we used two-thirds of the entire cohort () as training data to obtain an estimate of the regression coefficient, and then predicted gene expressions on the remaining test data (). For each integrative approach, we assessed model sparsity on the training data, which is defined as the number of non-zero coefficient estimates, and the predictive performance on the test data, which is measured by (i) the Spearman correlation between actual and predicted expressions and (ii) the relative prediction error (RPE) defined by (5).
| (5) |
The simulation results are shown in Figure 4. The methods that only consider CpGs in or very close to the gene body (E-NET-local, and ) are clearly outperformed by the other methods that also look at CpGs in the 500 kb neighborhood in terms of prediction accuracy, suggesting the usefulness of distal CpGs in explaining expression in many genes. Among the latter class of methods, Seq-Lasso, Lasso and E-NET have substantially better predictive performance than and NegCor-all. Compared to Lasso and E-NET, Seq-Lasso tends to produce a much sparser model at the cost of only slightly compromised prediction accuracy.
Fig. 4.
Performance comparison of Seq-Lasso with other methods in the simulation study. For the 500 randomly selected genes, the subfigures compare the performance of Seq-Lasso with alternatives for identifying methyl-eQTLs in terms of model sparsity as measured by the number of selected CpGs on the training data in (A), and predictive accuracy on the test data as measured by the Spearman correlation between actual and predicted gene expressions in (B) and relative prediction error in (C). We take the median across 100 replicates to compute a gene-specific average for each performance measure
3.2.2. Application to CRC data
We then applied all the integrative methods to the 9527 genes satisfying our selection criteria in CRC data (which are described in Supplementary Section S1.1.5 in Supplementary Material), and evaluated the performance of each method using 3-fold cross validation on the TCGA cohort and independent validation on the MDACC cohort. For cross validation, we calculated the Spearman correlation between the predicted and actual gene expressions across 369 samples in the TCGA cohort. For independent validation, we first obtained the methyl-eQTLs from the TCGA cohort and used it to predict the gene expressions for the MDACC cohort, and then calculated the Spearman correlation between the predicted and actual gene expressions. One caveat of independent validation data is that it uses a different platform (Agilent) to measure gene expression from TCGA (RNAseq), leading to measurements on different scales. In spite of this limitation, we still think it instructive to look at Spearman correlations between predicted and real expressions in this dataset. To focus on genes with a reasonably high percentage of expression variability explainable by methylation, these comparisons were restricted to the 5586 genes with a Spearman correlation of at least 0.40 between the actual and predicted expressions based on 3-fold cross validation of the TCGA data using at least one of the methods above.
The comparisons across all methods are presented in Figure 5 and lead to conclusions highly consistent with those made in the simulation study. Comparing the standard methods commonly used in the literature, tends to produce significantly higher correlations than , but there is a clear gap in predictive performance between these and the penalized regression approaches (Lasso, Seq-Lasso, E–NET, IAL) that consider CpGs in the 500 kb neighborhood, suggesting that useful information is lost by collapsing all methylation information into one single site per gene. This difference is seen in both the TCGA data and the independent validation data (Fig. 5B and C). Note that across all methods the Spearman correlations are lower in the independent validation dataset, which is at least partially due to the different gene expression platforms.
Fig. 5.
Performance comparison of with other methods in the CRC data. For the 5586 genes with a Spearman correlation of at least 0.40 between actual and predicted gene expressions in the TCGA cohort using at least one method, the subfigures compare the performance of with alternatives for identifying methyl-eQTLs in terms of model sparsity as measured by the number of selected CpGs in (A), and the ability to explain expression variability as measured by the Spearman correlation between actual and predicted gene expressions in the TCGA cohort in (B) and in the MDACC cohort in (C)
For cross validation of the TCGA data, Lasso, Seq-Lasso, E–NET all produce much higher correlations than (median = 0.42). Lasso and E-NET have slightly higher correlations (median = 0.53 for Lasso; median = 0.54 for E-NET) than Seq-Lasso (median = 0.51) but also incorporate many more CpGs (Fig. 5A). The median number of CpGs selected by Seq-Lasso is 8, fewer than two-thirds of the median for Lasso (13) and one-third of the median for E-NET (25). For independent validation involving the Agilent gene expression data, the Spearman correlation distributions for Seq-Lasso (median = 0.35), Lasso (median = 0.36), E-NET (median = 0.37) and (median = 0.34) are very similar.
Taken together, the comparisons in Sections 3.2.1 and 3.2.2 demonstrate the benefit of Seq-Lasso in balancing predictive performance and model sparsity. We provide more results about CRC methyl-eQTLs identified by Seq-Lasso using the TCGA data in Supplementary Section S4 in Supplementary Material, as well as the R code to implement each integrative method.
3.3 Enrichment of chromatin states in selected CpG sites
We computed the fold enrichment of various chromatin states in the CpG sites selected by Seq-Lasso. The chromatin state annotations are based on a 10-state model applied to 17 CRC primary tumor samples (see Section 2.1). For our enrichment analysis, two chromatin states that have different patterns of combinatorial histone marks but are both annotated as active enhancers are combined into one state. For a given CpG site, we assigned the most frequently predicted chromatin state among the 17 CRC tumor samples. Fold enrichment is defined as the ratio of odds of being assigned to a particular chromatin state for selected CpGs to the odds of being assigned to this chromatin state for unselected CpGs.
Figure 6 shows the fold enrichment along with 95% confidence interval of different chromatin states in the CpG sites identified by Seq-Lasso from the 5586 genes with a moderate or strong association between methylation and expression. As shown in Figure 6, most active states are enriched in the identified CpG sites, including active enhancer state, actively transcribed state and transcribed state with enhancer signature, while the inactive states (Polycomb repressed and heterochromatin repressed) are underrepresented. Interestingly, the active promoter/enhancer state is underrepresented in the identified CpG sites. Fisher’s exact tests show that the enrichment of each chromatin state among identified CpG sites is significantly different from 1.00 (P-value 1e-4). The chromatin state enrichment patterns for CpGs identified by the alternative methods are qualitatively similar and presented in Supplementary Section S5 in Supplementary Material.
Fig. 6.

Fold enrichment of annotated chromatin states in CpG sites identified by in CRC. The plot shows the estimated odds ratio of each annotated chromatin state for CpG sites identified by , relative to the unselected CpG sites. The open circles represent the point estimates of odds ratio, and the horizontal bars denote 95% confidence intervals. The chromatin state annotations are based on 17 CRC primary tumor samples with epigenomes generated at MDACC
3.4 Methyl-eQTLs
3.4.1. Motivating example: AREG and EREG
For AREG and EREG from the motivating example of this study, the methyl-eQTLs identified by Seq-Lasso and other methods are shown in Figures 7A–C and 8A–C, with selected CpGs and their weights highlighted in the figure. The gene body is indicated by the dark gray bar, with an arrow demonstrating the direction of transcription, and the nearby flanking genes are indicated by light gray bars with gene names listed at the top.
Fig. 7.

Methyl-eQTLs of AREG in CRC, and heatmap of absolute correlations across CpGs. (A)–(C) show the methyl-eQTLs identified by different methods, with the marks at the top indicating all CpG sites in the region (CpG island-red; CpG shore-pink; CpG shelf-green; open sea-black), and the marks at the bottom indicating active chromatin states predicted using ChromHMM (active TSS-red; flanking TSS-orange red; active enhancer-orange; transcribed enhancer-green yellow). The heatmaps (D)–(F) show the pairwise absolute Pearson correlation coefficients between CpGs selected by (arranged in columns) and all CpGs within ±500 kb of AREG (arranged in rows). The red marks on the left of each heatmap denote the CpGs identified by , and the black marks denote the CpGs that are selected by but not
Fig. 8.

Methyl-eQTLs of EREG in CRC, and heatmap of absolute correlations across CpGs. (A)–(C) show the methyl-eQTLs identified by different methods, with the marks at the top indicating all CpG sites in the region (CpG island-red; CpG shore-pink; CpG shelf-green; open sea-black), and the marks at the bottom indicating active chromatin states predicted using ChromHMM (active TSS-red; flanking TSS-orange red; active enhancer-orange; transcribed enhancer-green yellow). The heatmaps (D)–(F) show the pairwise absolute Pearson correlation coefficients between CpGs selected by (arranged in columns) and all CpGs within ±500 kb of EREG (arranged in rows). The red marks on the left of each heatmap denote the CpGs identified by , and the black marks denote the CpGs that are selected by but not
Figure 7A shows the methyl-eQTLs of AREG in colorectal cancer. Both Seq-Lasso and Lasso identify a CpG probe in the middle of the gene body whose methylation exhibits strong positive correlation with gene expression, and capture two CpG sites located at a distance from the gene body of AREG, for which the methylation levels are strongly inversely related to gene expression. Interestingly, one of them is located exactly at the TSS of EREG, which is situated roughly 50 kb upstream of AREG. The predicted expression by Seq-Lasso has a Spearman correlation of 0.664 and 0.666 with actual AREG expression respectively for 3-fold cross-validation in the TCGA cohort and independent validation in the MDACC cohort, respectively meaning that of the variation in AREG expression ranks is explained by methylation in the TCGA (MDACC) cohort. Both Seq-Lasso and Lasso yield much higher Spearman correlations than (0.320 for 3-fold cross validation and 0.298 for independent validation) and (0.333 for 3-fold cross validation and 0.298 for independent validation, not shown), indicating that these naïve methods miss functionally related CpGs when integrating methylation and gene expression.
Figure 7B and C show the methyl-eQTLs of AREG in breast cancer and pancreatic cancer respectively. A comparison of Figure 7A–C reveals that the methyl-eQTLs of AREG are very different across tissue types. Focusing on Seq-Lasso, the identified CpG sites have modest overlaps among three tumor types considered. In addition, the Spearman correlations between the actual and predicted expressions greatly differ across tissue types. The association between AREG expression and methylation is very strong in colorectal cancer, moderate in breast cancer and weak in pancreatic cancer.
Figure 8A shows the methyl-eQTLs of EREG in colorectal cancer. Seq-Lasso identifies 3 neighboring CpG sites at the TSS of EREG as functionally important. picks out one very strong negative signal at the TSS, and yields similar Spearman correlation between the actual and predicted EREG expressions as Seq-Lasso, suggesting that methylation at this single CpG site can explain as much variation in EREG expression. Lasso finds many more CpGs with only marginally higher correlation achieved. Figure 8B and C show the methyl-eQTLs of EREG in breast cancer and pancreatic cancer respectively, indicating that the methyl-eQTLs of EREG also depend on the tissue type.
Figure 8D–F shows the pairwise absolute Pearson correlation coefficients between the CpGs identified by Seq-Lasso and all the CpGs located within ±500 kb of EREG, respectively for the three tumors considered. In each heatmap, the rows labeled by black marks often include at least one light colored cell, indicating that a CpG identified by Lasso but not Seq-Lasso is often highly correlated (or anti-correlated) with at least one CpG identified by Seq-Lasso. This phenomenon results from the sequential nature of our approach. After Seq-Lasso identifies important CpGs in the first step, the methylation sites that are correlated with expression because of their strong correlation with those identified CpGs would not be selected in the second step, leading to a sparser set of methyl-eQTLs than Lasso. This phenomenon can also be observed for AREG (Fig. 7D–F).
3.4.2. Shiny App for methyl-eQTLs
To facilitate the search and visualization of methyl-eQTLs, we have developed a freely available R Shiny app that can be accessed at https://rstudio-prd-c1.pmacs.upenn.edu/methyl-eQTL/. This Shiny app can interactively display the methyl-eQTLs of any gene in the genome for colorectal, breast and pancreatic cancer, which includes a tissue-specific, gene-specific list of important CpGs and their corresponding weights, along with the Spearman correlation between actual gene expression and predicted expression based on 3-fold cross validation in the TCGA cohort for each gene. Note that a measure of percent of expression variability explained by the methyl-eQTLs can be computed by squaring the Spearman correlations. Given methyl-eQTLs, gene-specific methylation scores (GSMSs) can be computed for each subject that can serve as gene-level summary of methylation and be used in graphical displays of methylation data (which are described in Supplementary Section S6 in Supplementary Material). Our Shiny app can also interactively display the GSMS heatmaps of tumor samples for gene sets from common biological pathways. In Supplementary Section S7 in Supplementary Material, we show how to use the Shiny app and provide additional examples of interesting methyl-eQTLs; we also compute GSMSs of the TCGA and MDACC colorectal cohorts and compare them across four consensus molecular subtypes of colorectal cancer recently discovered and validated by Guinney et al. (2015).
4 Discussion
We propose Seq-Lasso, a novel penalized regression approach with an iterative nature to integrate DNA methylation and gene expression data. We consider CpG sites in a 500 kb neighborhood of the gene, where distal regulatory elements such as enhancers are usually located. By applying a sparsity-inducing penalty and using a sequential modeling strategy that incorporates information regarding which CpGs are a priori most likely to be associated with expression, Seq-Lasso is able to produce sparse tissue-specific, gene-specific lists of potentially functionally important CpGs and corresponding weights, which we refer to as methyl-eQTLs. Methyl-eQTLs can be used to construct gene-level methylation summaries that are maximally correlated with gene expressions, and produce tissue-specific, gene-specific measures of proportion of expression variation explainable by methylation.
Another key contribution of our work is to bring to attention the limitations of standard approaches currently used in the literature to integrate methylation and gene expression, which summarize gene-level methylation by taking the most negatively correlated probe or the average methylation level around the TSS. We find that methyl-eQTLs often include many distal CpG sites and intergenic regions far from the promoter region, and the inclusion of these sites leads to significantly better ability to explain expression. Therefore, it is clearly necessary to look at CpGs far beyond the TSS or the gene body to better understand the cis-effects of methylation, and we advise taking a variable selection approach to this purpose. In this work we built on Lasso to develop Seq-Lasso. By incorporating prior knowledge learned from the whole genome, Seq-Lasso is able to essentially accomplish the same predictive ability as Lasso while using a significantly smaller set of CpGs that may also tend to be potentially more functionally relevant and biologically interpretable. More generally, this two-step procedure can be combined with any given variable selection approach, such as E-NET and , to make better use of the biological information in this context and potentially improve the performance of the original approach.
The analysis of methyl-eQTLs for individual genes also reveals that the CpG sites associated with gene expression vary greatly across genes and tissue types, demonstrating the importance of adopting a tissue-specific and gene-specific approach to integrating methylation and gene expression.
In multi-platform integration, it is frequently useful to compute gene-level summaries of various platforms. Our approach yields gene-level summary of methylation which is sparse and maximally correlated with expression. These gene-level methylation summaries can be used in integrative models, and can also be associated directly with demographical and prognostic factors, as illustrated by our ANOVA analysis of GSMSs and consensus molecular subtypes in CRC in Supplementary Section S6 in Supplementary Material. A similar strategy could be used to construct gene-level summaries for other genomic platforms, such as DNA mutation and miRNA.
To identify the CpG sites which are most likely to be associated with gene expression a priori, our model combined information across the genome to estimate this prior probability as a function of epigenomic characteristics of a CpG site. Apart from guiding the sequential selection, these selection probability functions constitute important biological findings by themselves, which may contribute to the unraveling of complicated regulatory roles taken by methylation. They also corroborate some previous findings regarding the methylation-expression relationship, including the frequently observed association between promoter hyper-methylation and gene silencing, and the finding that CpG shore methylation is strongly associated with gene expression (Irizarry et al., 2009).
The CpGs selected by our model are further studied by cross-reference with predicted chromatin states from ChromHMM. As found in our analyses, CpGs at genomic locations with certain predicted chromatin states tend to have higher or lower propensity for selection as being functionally important for predicting expression.
It should be pointed out that our modeling approach assumed a linear regression setting for interpretability, implying the effect of methylation sites on gene expression is linear and additive (Wang et al., 2013). It would be possible to extend this approach to consider more general and flexible models with non-parametric non-linear effects and interactions allowed, but these are outside the scope of this study. In addition, the conclusions drawn from our integrative approach should not be interpreted as strict causal relationships, but instead represent strongly associated CpG sites that are potential key methylation switches and need further functional validation to confirm any causative relationships, such as modulation using CRISPR/dCas9 fusion proteins to methylate/demethylate the region and then measuring the changes in gene expression levels. While we do not have wet lab to validate these results, we freely provide these methyl-eQTL results across the whole genome for selected cancer types so interested biomedical researchers could biologically validate them. Also, based on our analysis, it is clear that the use of the 27K methylation array, which contains roughly one or two CpGs per gene mostly found in the promoter region (Jeschke et al., 2015), would miss many important CpGs relative to the 450K methylation array. Recently, whole-genome bisulfite sequencing has been used to interrogate CpGs at single-base resolution (Lou et al., 2014). Given sufficient samples, we could rerun our methyl-eQTL analysis on the whole methylome data to obtain more accurate gene-level methylation summaries.
In our current Shiny R app, we interactively visualize the methyl-eQTLs for colon, rectal, breast and pancreatic cancer types. Our future plans include expanding results to include methyl-eQTLs for all TCGA cancers, which will be included in future updates of our Shiny app, and given healthy tissue for various tissue types, to expand this to identify methyl-eQTLs for normally functioning tissues.
Currently, we focus on cis methyl-eQTL analysis in this article, and it may be interesting to extend the scope of potential CpGs to study trans methyl-eQTLs in the future, which could regulate the expression of distant target genes through pathways or transcription factors. Finally, Seq-Lasso is developed to identify methyl-eQTLs independently for each gene, which can be cast as a variable selection problem with a univariate response. An interesting extension of this work is to generalize Seq-Lasso to simultaneously identify methyl-eQTLs of multiple genes, which should presumably improve upon Seq-Lasso by borrowing strength across genes, since we expect that genes with similar expression profiles are likely to be co-regulated by a common set of CpG loci. This extension requires the use of a variable selection approach that models multivariate responses and encourages correlated responses to have similar regression coefficients. Numerous multivariate penalized regression approaches in the literature (Cheng et al., 2014, 2016; Deshpande et al., 2019; Chun and Keles, 2009) are well suited to this purpose, and could be combined with our sequential modeling strategy to accomplish simultaneous identification of methyl-eQTLs for multiple genes. This is outside the scope of the current work and we leave it to future work.
Supplementary Material
Acknowledgements
The authors thank the editor, the associate editor and two referees for their valuable comments, which have helped us substantially improve this article.
Funding
This work was supported by R01-CA178744, UH2-CA207101, P30-CA016672, R01-CA244845 and P50-CA221707 from the National Cancer Institute, 1550088 and UL1-TR001878 from the National Science Foundation, the University of Texas M.D. Anderson Cancer Center Colorectal Cancer Moonshot, and the Del and Dennis McCarthy Distinguished Professorship Endowment in Gastrointestinal Cancer Research.
Conflict of Interest: none declared.
Contributor Information
Yusha Liu, Department of Human Genetics, The University of Chicago, Chicago, IL 60637, USA.
Keith A Baggerly, Department of Bioinformatics and Computational Biology, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Elias Orouji, Department of Genomic Medicine, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Ganiraju Manyam, Department of Bioinformatics and Computational Biology, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Huiqin Chen, Department of Bioinformatics and Computational Biology, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Michael Lam, Department of Gastrointestinal Medical Oncology, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Jennifer S Davis, Department of Epidemiology, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Michael S Lee, Department of Medicine, The University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA.
Bradley M Broom, Department of Bioinformatics and Computational Biology, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
David G Menter, Department of Gastrointestinal Medical Oncology, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Kunal Rai, Department of Genomic Medicine, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Scott Kopetz, Department of Gastrointestinal Medical Oncology, M.D. Anderson Cancer Center, Houston, TX 77030, USA.
Jeffrey S Morris, Department of Biostatistics, Epidemiology and Informatics, The University of Pennsylvania, Philadelphia, PA 19104-6021, USA.
References
- Aran D. et al. (2013) DNA methylation of distal regulatory sites characterizes dysregulation of cancer genes. Genome Biol., 14, R21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brenet F. et al. (2011) DNA methylation of the first exon is tightly linked to transcriptional silencing. PLoS One, 6, e14524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng W. et al. (2014) Graph-regularized dual lasso for robust eQTL mapping. Bioinformatics, 30, i139–i148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng W. et al. (2016) Sparse regression models for unraveling group and individual associations in eQTL mapping. BMC Bioinformatics, 17, 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chun H., Keles S. (2009) Expression quantitative trait loci mapping with multivariate sparse partial least squares regression. Genetics, 182, 79–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denis M., Tadesse M.G. (2016) Evaluation of hierarchical models for integrative genomic analyses. Bioinformatics, 32, 738–746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deshpande S.K. et al. (2019) Simultaneous variable and covariance selection with the multivariate spike-and-slab lasso. J. Comput. Graph. Stat., 28, 921–931. [Google Scholar]
- Ernst J., Kellis M. (2010) Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat. Biotechnol., 28, 817–825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guinney J. et al. (2015) The consensus molecular subtypes of colorectal cancer. Nat. Med., 21, 1350–1356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Irizarry R.A. et al. (2009) The human colon cancer methylome shows similar hypo-and hypermethylation at conserved tissue-specific CpG island shores. Nat. Genet., 41, 178–186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jennings E.M. et al. (2013) Bayesian methods for expression-based integration of various types of genomics data. EURASIP J. Bioinf. Syst. Biol., 2013, 13–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeschke J. et al. (2015) DNA methylome profiling beyond promoters–taking an epigenetic snapshot of the breast tumor microenvironment. FEBS J., 282, 1801–1814. [DOI] [PubMed] [Google Scholar]
- Jones P.A. (2012) Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet., 13, 484–492. [DOI] [PubMed] [Google Scholar]
- Kulis M. et al. (2013) Intragenic DNA methylation in transcriptional regulation, normal differentiation and cancer. Biochim. Biophys. Acta Gene Regul. Mech., 1829, 1161–1174. [DOI] [PubMed] [Google Scholar]
- Kundaje A. et al. ; Roadmap Epigenomics Consortium. (2015) Integrative analysis of 111 reference human epigenomes. Nature, 518, 317–330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laurent L. et al. (2010) Dynamic changes in the human methylome during differentiation. Genome Res., 20, 320–331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee M.S. et al. (2016) Association of CpG island methylator phenotype and EREG/AREG methylation and expression in colorectal cancer. Br. J. Cancer, 114, 1352–1361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lou S. et al. (2014) Whole-genome bisulfite sequencing of multiple individuals reveals complementary roles of promoter and gene body methylation in transcriptional regulation. Genome Biol., 15, 408–421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orouji E. et al. (2021) Chromatin state dynamics confers specific therapeutic strategies in enhancer subtypes of colorectal cancer. bioRxiv, doi: 10.1136/gutjnl-2020-322835. [DOI] [PMC free article] [PubMed]
- Rhee J.-K. et al. (2013) Integrated analysis of genome-wide DNA methylation and gene expression profiles in molecular subtypes of breast cancer. Nucleic Acids Res., 41, 8464–8474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rhie S.K. et al. (2016) Identification of activated enhancers and linked transcription factors in breast, prostate and kidney tumors by tracing enhancer networks using epigenetic traits. Epigenet. Chromatin, 9, 50–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schlosberg C.E. et al. (2017) Modeling complex patterns of differential DNA methylation that associate with gene expression changes. Nucleic Acids Res., 45, 5100–5111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen R. et al. (2009) Integrative clustering of multiple genomic data types using a joint latent variable model with application to breast and lung cancer subtype analysis. Bioinformatics, 25, 2906–2912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun W. et al. (2010) Genomewide multiple-loci mapping in experimental crosses by iterative adaptive penalized regression. Genetics, 185, 349–359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teschendorff A.E., Relton C.L. (2018) Statistical and integrative system-level analysis of DNA methylation data. Nat. Rev. Genet., 19, 129–147. [DOI] [PubMed] [Google Scholar]
- Thingholm L.B. et al. (2016) Strategies for integrated analysis of genetic, epigenetic, and gene expression variation in cancer: addressing the challenges. Front. Genet., 7, 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tibshirani R. (1996) Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Ser. B (Methodological), 58, 267–288. [Google Scholar]
- VanderKraats N.D. et al. (2013) Discovering high-resolution patterns of differential DNA methylation that correlate with gene expression changes. Nucleic Acids Res., 41, 6816–6827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wagner J.R. et al. (2014) The relationship between DNA methylation, genetic and expression inter-individual variation in untransformed human fibroblasts. Genome Biol., 15, R37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang W. et al. (2013) iBAG: integrative Bayesian analysis of high-dimensional multiplatform genomics data. Bioinformatics, 29, 149–159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wood S.N. (2006). Generalized Additive Models: An Introduction with R. Chapman and Hall/CRC, Boca Raton. [Google Scholar]
- Yao L. et al. (2015) Inferring regulatory element landscapes and transcription factor networks from cancer methylomes. Genome Biol., 16, 105–121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhong H. et al. (2019) Predicting gene expression using DNA methylation in three human populations. PeerJ, 7, e6757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zou H. (2006) The adaptive lasso and its oracle properties. J. Am. Stat. Assoc., 101, 1418–1429. [Google Scholar]
- Zou H., Hastie T. (2005) Regularization and variable selection via the elastic net. J. R. Stat. Soc. Ser. B (Stat. Methodol.), 67, 301–320. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.