Abstract
Synapses maintain their molecular composition, plasticity and function through the concerted action of protein synthesis and removal. The complex and polarized neuronal architecture poses specific challenges to the logistics of protein and organelle turnover since protein synthesis and degradation mainly happen in the cell soma. In addition, post-mitotic neurons accumulate damage over a lifetime, challenging neuronal degradative pathways and making them particularly susceptible to the effects of aging. This review will summarize the current knowledge on neuronal protein turnover mechanisms with a particular focus on the presynapse, including the proteasome, autophagy and the endolysosomal route and their roles in regulating presynaptic proteostasis and function. In addition, the author will discuss how physiological brain aging, which entails a progressive decline in cognitive functions, affects synapses and the degradative machinery.
Keywords: aging, autophagy, endolysosome, presynapse, proteostasis, ubiquitin proteasome system
Introduction
Protein homeostasis, or proteostasis, refers to all protein synthesis and degradation processes that preserve the essential balance of cellular protein levels. Synapses are especially dependent on the mechanisms for maintaining proteostasis as they contain an extremely dense network of proteins that needs to undergo constant remodeling to govern neurotransmission and synapse reshaping during learning and memory processes [1,2]. Most synapses are located far away from the soma, the main compartment for protein synthesis and degradation, creating extra challenges to the logistics of proteostasis pathways. In addition, because of their post-mitotic and long-lived nature, neurons and their synapses are at increased risk for age-related damage accumulation. During aging, cells encounter stressful situations, caused for instance by a buildup of unfolded proteins or an accumulation of reactive oxygen species (ROS) causing lipid peroxidation and protein oxidation [3]. At the synapse, high metabolic burdens and calcium insults during neuronal activity might even accelerate protein damages and turnover [4,5]. The burden on mature and aging synapses increases even more, as aging is associated with a progressive decline in the efficiency of degradation systems, such as the proteasome, endolysosomal and autophagy pathway. This review will first summarize current knowledge on the proteostasis processes that act to remove dysfunctional synaptic components (Figure 1) and how, for instance, these pathways are locally regulated to maintain proper functioning of the presynapse. Second, this review will address how physiological brain aging affects the synapse and its degradative capacity.
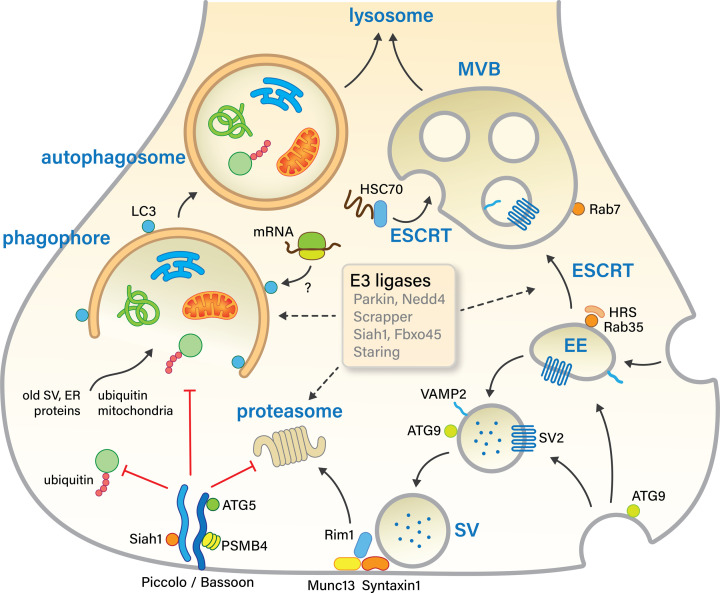
Figure 1. Degradative pathways and suggested presynaptic cargos.
Synaptic proteins and organelles, such as SVs and mitochondria, can be locally removed via alternative degradative routes. The proteasome removes predominantly cytosolic proteins, including active zone proteins RIM1 and Munc13. ESCRT-dependent endolysosomal sorting is thought to be required for the degradation of a subset of membrane proteins, for instance, SV proteins VAMP2 and SV2. Both proteins and organelles can be sequestered by a double-membrane structure called a phagophore, which then closes to form an autophagosome that eventually fuses with somatic lysosomes for degradation. Ubiquitination is a common signal for degradation in all of these major protein degradation pathways. Therefore, ubiquitin-ligases (E3) and deubiquitinating enzymes could play important roles in maintaining a healthy presynapse. The six E3 ligases indicated here either have known, mammalian, presynaptic targets (Nedd4, Scrapper, Fbxo45, Staring) or are locally regulated by presynaptic proteins (Siah1, Parkin); EE, early endosome; SV, synaptic vesicle; MVB, multivesicular body.
Ubiquitin–proteasome system (UPS)
A central feature of all major degradative pathways is the ability to selectivity target substrates by site-specific mono- and poly-ubiquitination. Ubiquitin is a 76-amino acid protein that covalently attaches to substrate proteins. This conjugation relies on the sequential activities of three classes of enzymes: E1 (ubiquitin-activating), E2 (ubiquitin-conjugating) and E3 (ubiquitin ligases). Ubiquitination is initiated by E1, an enzyme requiring ATP for the activation of ubiquitin. Ubiquitin is then handed over to an E2 to form an ubiquitin∼E2 conjugate via a thioester exchange reaction. The (>600 known) E3 ligases mediate the final step of ubiquitin transfer and provide the specificity for substrate recognition by interacting with both E2 and the target protein [6]. Ubiquitin can create different types of poly-ubiquitin chains via its lysine residues and each type of chain has the potential to act as a distinct intracellular signal. For instance, K48-linked chains are a prime signal for the ubiquitin–proteasome system (UPS) whereas K63-linked chains play a more prominent role in lysosomal degradation [7]. Deubiquitination enzymes (DUBs) serve as an important regulatory layer and contribute to the replenishment of ubiquitin pools.
The highly conserved UPS is responsible for the degradation of the majority of short-lived, cytosolic proteins. The proteasome consists of a barrel-shaped 20S catalytic core particle and 19S regulatory particles that select and unfold target proteins labeled with ubiquitin chains. During presynaptic assembly, a local decrease in UPS activity is proposed to aid in the formation of a stable cluster of synaptic vesicles (SVs) [8–10]. Research over the years also demonstrate critical functions for the UPS in mature synapses, with UPS involvement in the regulation of basal neurotransmission and synaptic plasticity [2,11,12]. Synaptic plasticity includes long-term potentiation (LTP) and long-term depression (LTD) that are thought to form the cellular basis of learning and memory, and their dysfunction may underlie memory loss and other cognitive disorders [13,14]. Incubation of hippocampal slices with proteasome inhibitors enhances early phase LTP induction [15] while impairing late phase LTP maintenance [11,16]. Bilateral infusion of lactacystin, a specific proteasome inhibitor, in the hippocampal CA1 area of rats is shown to impair memory formation and memory extinction [17,18]. The mechanisms that underlie proteasome-dependent plasticity have not been fully elucidated but probably depend on changes in the postsynaptic compartment [17,19]. In addition, proteasome inhibition is implicated in presynaptic neurotransmitter release. At the Drosophila neuromuscular synapse, acute pharmacological inhibition of the proteasome in the presynaptic neuron results in stabilization of active zone protein Unc-13/Munc13 and increased synaptic transmission due to a more efficient priming and release of SVs [20]. In cultured rat hippocampal neurons, proteasome inhibition causes an increase in the size of the recycling pool, although in this study no effect on SV release was observed [21]. Proteasome 20S complexes are mainly present in the cytosol, but a recent finding indicates that they can associate with the plasma membrane of neurons, where they are involved in the activity-dependent degradation of intracellular proteins, producing extracellular peptides that can modulate neuronal activity [22]. It is currently unclear how these surface-exposed proteasomes target to the plasma membrane and are embedded into the lipid bilayer.
Apart from changes in neurotransmission, proteasome dysregulation in mature synapses also causes structural presynaptic abnormalities. Inactivation of PI31, an adaptor for microtubule-based transport of proteasomes, results in the accumulation of ubiquitinated protein aggregates in mouse and fly axon terminals and is accompanied by distal axon swellings and presynapse loss [23,24], while leaving the postsynapse mostly unperturbed [23]. Whereas in these studies the identity of the accumulated proteins was not determined, other works have indicated that the levels of some presynaptic proteins, such as synaptophysin, liprin-alpha, RIM, Bassoon and above-mentioned Munc13 are controlled via UPS-mediated degradation [25–28]. At the mature mammalian presynapse, ubiquitin E3 ligases have been identified to induce proteasomal degradation of presynaptic proteins (Figure 1). The E3 enzyme Staring, for instance, ubiquitinates syntaxin 1, a member of the SNARE complex and an essential component of the SV fusion machinery [29]. E3 ligases SCRAPPER and Fbxo45 ubiquitinate active zone proteins RIM1 and Munc13, thereby contributing to the UPS-dependent regulation of SV release. Accordingly, loss of these E3 ligases results in a facilitation of neurotransmission [30,31]. In addition, the presynaptic active zone (AZ) proteins Bassoon and Piccolo are known to bind and inhibit the E3 ligase Siah1 [32]. Bassoon has also been found to interact with PSMB4, a structural subunit of the 20S core proteasome, thereby interfering with compartment-specific ubiquitination and proteasome biogenesis [33]. Accordingly, loss of Bassoon and Piccolo increases ubiquitination and degradation of SV proteins, eventually resulting in SV loss and synapse elimination. Thus, these AZ proteins seem to stabilize the AZ by inhibiting degradation. Interestingly, core AZ components show age-induced increases in Drosophila brains [34], but whether mammalian neurons show similar trends is unknown. Regarding the role of synaptic deubiquitylation very little is known. At the postsynapse, USP46 is identified as the DUB for AMPARs, stabilizing cell-surface expression levels [35]. Our knowledge regarding presynaptic ubiquitin regulation, including the identity, roles and substrates of DUBs is still very limited and thus warrants further investigation.
While the UPS locally and preferentially targets short-lived soluble proteins, the endolysosomal and autophagy pathways are responsible for the elimination of membrane proteins, long-lived proteins, insoluble protein aggregates and dysfunctional organelles [36]. In neuronal axons and synapses, both the endolysosomal and autophagy pathways are retrograde trafficking routes that deliver proteins to degradative lysosomes that are primarily present in neuronal somata [37].
The endolysosomal pathway
Transmembrane proteins and associated factors are typically degraded by the endolysosomal system and access this pathway by endocytosis. Pre-existing early endosomes then fuse with these endocytosed vesicles, permitting proteins to recycle back to the cell surface (regulated by small GTPases Rab4 and Rab11) or directing cargo to late endosomes or late endosomal multivesicular body (MVB) and finally to lysosomes for degradation (regulated by Rab7) [38]. During their transport towards the soma, axonal endosomes gradually maturate and acidify which may also control their degradative capacity [37,39]. In the pathway promoting degradation, Endosomal Sorting Complexes Required for Transport (ESCRT) is a core protein machinery, important for the recognition of ubiquitinated membrane cargo and the formation, trafficking, and fusion of MVBs. The ESCRT machinery is composed of multiprotein complexes known as ESCRT-0, I, II and III. The ESCRT-0 protein HRS initiates the process by binding to ubiquitinated cargos and tethering them to the surface of early endosomes by binding phosphatidylinositol 3-phosphate (PI[3]P). On endosomes, PI[3]P is generated by the Vps34 phosphatidylinositol 3-kinase complex II (Vps34/Vps15/Beclin1) that assembles at early endosomes via the association of Vps15 with Rab5 [40]. Recruitment of Rab5 effectors, such as Rabaptin-5 and Rabex-5, can regulate Rab5 activity and the formation of a local endosomal PI3P pool [41]. The sequential recruitment and activation of the other ESCRT complexes promotes inward vesicle budding and membrane sealing by the Vps4 ATPase [42]. Studies on ESCRT-dependent presynaptic protein degradation are scarce but evidence from Drosophila and cultured rat hippocampal neurons suggests that endosomal sorting is required for the degradation of a subset of ubiquitinated SV proteins (e.g. VAMP2 and SV2), a process involving active Rab35 dependent HRS recruitment to SV pools [43,44]. In agreement with this, mutations in CHMP2B, an ESCRT-III subunit involved in frontotemporal dementia, results in an increase in presynaptic endosomes and a selective retention of SV-associated proteins in aged mice [45]. In contrast, at the Drosophila photoreceptor axon terminals, sorting of SV proteins was found to be independent from Rab7 whereas degradative sorting of plasma membrane proteins was Rab7-dependent, indicating the involvement of multiple degradative sorting pathways [46]. Cytosolic proteins can also enter the endolysosomal system by “endosomal microautophagy”, a pathway dependent on ESCRTs, but not the core autophagy machinery (see below). This process relies on the cytosolic chaperone Hsc70 for substrate targeting and the subsequent invagination-mediated uptake by late endosomes/MVBs. In Drosophila neurons, endosomal microautophagy mediates the turnover of synaptic proteins harboring a KFERQ recognition motif, including for instance Unc-13/Munc13 and EndophilinA, thereby rejuvenating the protein pool and facilitating synaptic transmission [47]. Given the fact that in the presynapse most organelles and markers for the early stages of the endolysosomal route are present, including MVBs, Rabs and ESCRT components [48–50], this route might play an underappreciated but important role in synaptic proteostasis and function and therefore represent an interesting area for future research.
Autophagy at the presynapse
Although often used synonymously, there are at least three distinct types of autophagy; microautophagy, chaperone-mediated autophagy (CMA) and macroautophagy. Microautophagy (similar to endosomal microautophagy) is defined by the direct uptake of cargo by the lysosome via lysosomal membrane invagination [51]. In CMA, selective cytosolic substrate proteins bind the chaperone Hsc70 and are thought to be translocated across the lysosomal membrane via the receptor lysosome-associated membrane protein type 2A (LAMP2A) [52]. The most prevalent form of autophagy, however, is macroautophagy (hereafter called autophagy) that utilizes double membrane vesicles, called autophagosomes, that deliver their content to the lysosome. Under baseline conditions, neuronal autophagosomes are mostly generated in the axon tip and retrogradely transported to the cell soma for fusion with degradative lysosomes [53]. Autophagosome formation requires a “core machinery” that starts with initiation factor ULK1 kinase complex, which translocates to the endoplasmic reticulum (ER) membrane (or another membrane donor) to activate the VPS34/ATG14/Beclin1 complex, that in turn triggers the production of PI[3]P. The subsequent recruitment of the PI[3]P effector WIPI2B allows for further recruitment of autophagy-specific ubiquitin-like conjugation systems, including E1-like ATG7 and the E3-like ATG12-ATG5:ATG16L ligase complex, that lipidate LC3 and LC3-related proteins. The lipidated form of LC3 stably inserts itself into the growing inner and outer autophagosome membrane to promote membrane elongation, substrate recruitment and finally autophagosome closure [54]. Autophagy can be either a non-selective bulk process, or a selective process that utilizes so-called autophagy receptors that can recognize substrates and simultaneously interact with lipidated LC3, targeting specific substrates to the inner membrane of the growing autophagosome. The specific action of these receptors is regulated by protein modifications, such as ubiquitination, phosphorylation and oligomerization [55]. Autophagosomes are then transported to the soma in a dynein-dependent manner, a process that is linked to their maturation and gradual acidification, for example by fusion with late endosomes or lysosomes to form autolysosomes [56]. Interestingly, the canonical machinery involved in synaptic transmission can also act to locally regulate presynaptic autophagy. Presynaptic endocytic proteins Endophilin and Synaptojanin, for instance, promote autophagosome formation by aiding the lipidation of LC3 and autophagosome closure while the endocytic adaptor complex AP2 facilitates retrograde axonal transport of mature autophagosomes (also referred to as amphisomes) to promote TrkB/BDNF signaling [57]. Conversely, the AZ protein Bassoon inhibits autophagy by binding to autophagy protein ATG5 [58–61].
Autophagosome biogenesis and transport in axons have been studied in detail [62], and emerging studies now tackle the question what substrates are targeted by neuronal autophagy and how this process is involved in maintaining synapse function. At the post-synapse, recent studies show that dendritic autophagy is required for synaptic LTD and the removal of postsynaptic AMPA receptor subunits and scaffold proteins [63,64]. The main sites of autophagosome biogenesis are at distal axons where autophagosomes are proposed to contain cargoes such as damaged SV proteins [65,66] but also mitochondria [67,68], ER [69] and whole SVs [67,70]. Similar to proteins, organelles have finite lifetimes and accumulate damages over time that can, if not taken care of, lead to organelle dysfunction. In addition, during endo-exocytotic cycles, SVs are gradually contaminated with proteins that normally reside on the plasma membrane. According to one hypothesis the accumulation of a certain number of these proteins prevents aged SVs from fusion with the PM. Hence, such aged SVs need to be degraded [71]. In mouse dopaminergic neurons, induction of autophagy by treatment with the mTOR inhibitor rapamycin results in an ATG7-dependent reduction in the number of SVs and a consecutive decrease in evoked neurotransmitter release [72], suggesting that SVs as a whole can be targeted to the autophagy pathway. The synaptic vesicle-associated GTPase Rab26 is postulated to connect SV recycling and autophagy by binding to autophagy protein ATG16L and directing SVs as a whole to pre-autophagosomal structures [70,73]. Rab26 null mutant fruit flies suffer from defective stimulus-dependent neurotransmission, although this did not appear to be accompanied by changes in autophagosomal and SV markers, suggesting a possible autophagy-independent function of Rab26 [74]. Recent work indicates a role for ATG9, the only known transmembrane protein involved in the core autophagy pathway, by linking the activity-dependent SV cycle with autophagosome formation. ATG9 exhibits lipid scramblase activity and, together with the lipid transport protein ATG2, mediates the flow of lipids from a donor membrane to the growing autophagosome membrane [75]. ATG9 is enriched at presynaptic terminals where it plays a role in local autophagosome biogenesis [76]. Recent work in Caenorhabditis elegans and mouse primary neurons shows that ATG9-containing vesicles undergo exo-endocytosis at presynaptic sites in an activity-dependent manner, via shared components of the machinery for SV recycling. Mutants that disrupt synaptic endocytic traffic, including a Parkinson’s disease (PD) associated mutation in synaptojanin1/unc-26, result in presynaptic ATG9 accumulation and accelerate the age-dependent decay of C. elegans synapses and locomotor behavior [77]. How ATG9 cycling aides in autophagosome biogenesis and what autophagy substrates underlie the reported age-dependent changes remains to be explored.
Modulation of presynaptic neurotransmission by autophagy is not only proposed to occur via removal of SVs and their associated proteins. Knockout of the core autophagy protein ATG5 selectively in excitatory forebrain neurons results in increased excitatory neurotransmission at CA3-CA1 synapses in acute mouse hippocampal slices and in primary hippocampal cultures, an effect independent from SV number or SV protein levels [69]. Instead, the gain in neurotransmission results from elevated calcium release from endoplasmic reticulum (ER) tubules, which accumulates at presynaptic sites as a consequence of abolished autophagy. Another independent study also failed to detect major alterations in the levels of SV proteins in brains lacking ATG5 specifically in either excitatory or inhibitory neurons. Instead, they identified autophagy-dependent degradation of synaptic subunits of protein kinase A (PKA), a key regulatory enzyme involved in the phosphorylation of the synaptic proteome [78]. In contrast, a recent study using proteomic analysis of isolated autophagic vesicles from wild-type mouse brain tissue suggested synaptic proteins as basal autophagy cargoes [67]. This may reflect differences between the analysis of cargos under wild-type conditions or cargo accumulation in autophagy inhibited condition, a situation in which other mechanisms might be able to take over degradation of certain components. The same study revealed mitochondrial fragments as a major cargo within neuronal autophagosomes. Due to their role in sequestering cytosolic calcium or ATP production, presynaptic mitochondria can regulate presynaptic release, plasticity and increase synapse stability [79,80], suggesting that autophagy-mediated removal of axonal mitochondria modulates presynaptic processes. Indeed, in aged rodents and fruit flies, mitochondria lose their functionality possible as a consequence of malfunctional autophagy [81,82] (and see further below). Future studies will be needed to determine how synaptic cargo is recognized and what autophagy receptors and protein modifications might aid substrate recognition.
To allow neuronal synapses to modify their proteome during processes like plasticity, regulated protein degradation, but also protein synthesis must be implemented (for a nice overview on synaptic protein synthesis and delivery see [83]). Traditionally believed to be a purely somatic event, synthesis of proteins from messenger RNA (mRNA) can occur locally at the distal presynapse [84]. Interestingly, a recent study shows that the autophagy protein LC3B is an RNA-binding protein that can trigger rapid mRNA degradation upon autophagic activation in HEK293T cells, thereby mediating cross-talk between autophagy and local protein synthesis [85]. It would be interesting to see if LC3-dependent RNA degradation plays a role in regulating the local synaptic proteome.
Common signals and regulators of degradative pathways
Degradative pathways can respond to a multitude of stressors, including protein damage and nutrient starvation. These signaling pathways can regulate degradation pathways in different ways. For example, inhibition of the mechanistic target of rapamycin (mTOR) signaling is responsible for autophagy induction during nutrient starvation and neuronal activity [86–88] but is also involved in proteasomal inhibition [89]. Interestingly, mTOR is present at synapses. Here, its inactivation decreases the strength of evoked excitatory synaptic transmission via both post- and presynaptic mechanisms, including enhanced spontaneous SV vesicle fusion [90]. Another signal that can influence both UPS and autophagy is proteostatic imbalance in the ER (known as ER stress). ER stress can be a result of abnormalities in calcium homeostasis, oxidative stress and other conditions leading to protein folding defects, processes postulated to play an important role in the rate of aging. To restore balance, ER stress activates many different pathways, including autophagy and the unfolded protein response (UPR), a stress response mediated by ER transmembrane stress sensors (PERK, ATF6 and IRE1α) that induces transcriptional and translational programs to lower protein production and promote degradation. ER stress and the UPR are commonly associated with neurodegenerative diseases [91], and UPR manipulations can restore cognition and synapse function in aged mice [92,93]. Recent studies suggest that UPR can be locally initiated by activity in the distal dendrite and at postsynapses, thereby inducing dendritic growth [94], but the exact contributions of this broad process to synapse maintenance are presently unknown.
Although the above discussed degradation pathways use different molecular machineries, some of the key players are shared between them. A common denominator of all is the selection of substrates by ubiquitination. Ubiquitin ligases as well as DUBs can be shared between the degradation pathways. The E3 ubiquitin ligase PRKN/Parkin for instance, can catalyze a range of ubiquitination events, including mono-ubiquitination, K48- and K63-linked poly-ubiquitination. These events can mark substrates for the proteasome [95] but also play an important role in autophagic degradation of mitochondria [96] and, as more recently suggested, presynaptic vesicle autophagy [97]. Another E3 ligase, named Nedd4, induces rapid mGlu7 degradation via both proteasomal and lysosomal pathways [98]. Similarly, ubiquitin receptors can convey selectivity but are sometimes involved in multiple pathways, for instance, the autophagy receptor p62 brings ubiquitinated proteins to the autophagosome via LC3 binding but is also involved in proteasomal degradation [99]. As indicated above, the large AZ protein Bassoon can modulate the presynaptic proteome in several ways, by negatively regulating autophagy [60], proteasome activity [33] and protein ubiquitination [32]. Components of the degradative pathways are themselves often also substrates, proteasomes for example, can be degraded by bulk autophagy [100]. In addition, the MVB and autophagy pathways merge at the late endosome/lysosome [101], making both pathways sensitive to a block in lysosomal degradation by, for instance, proton pump or phosphoinositide 3-kinase (PI3K) inhibitors. Thus, when targeting specific protein degradation pathways or proteins, effects on downstream machineries or redundancy of degradative pathways need to be taken into consideration.
Global synapse changes during normal aging
Even in the absence of neurodegenerative diseases, the functional capabilities of our brain, such as memory and processing speed, progressively decline with age. Synapses form the requisite connections of the neuronal networks that underlie cognition and therefore represent an important target for the effects of aging. Human synapse formation starts prenatally and lasts until about 2 years after birth, after which half of all synapses are subsequently “pruned” during the following two decades [102]. After this developmental period, a proportion of synaptic connections will be stably maintained over life spans of months to years [103]. Other connections, however, are unstable with turnover rates in the range of a few days to weeks, likely depending on the brain region [104–106]. Alterations in synaptic turnover are correlated with the ability to learn and to retain information, indicating the importance for neuronal circuit plasticity [107,108]. A growing body of evidence supports the idea that disruptions in these synaptic connections drive brain changes and cognitive decline that occur during normal aging [109]. Decreases in spine density have been reported in aging rodents [92,110–112] (however also see [113,114]), non-human primates [115], and humans [116]. In vivo two-photon imaging of cortical spines and boutons in old mice (>20 months) revealed changes in basal synapse dynamics but not number [116,117] with for instance a reduction in learning-induced new spine formation [111]. The hippocampus, an area essential for learning and episodic memory, is particularly sensitive to aging [118] and a reduction in SV numbers is observed in aged hippocampal mossy fiber (MF)-CA3, and CA3-CA1 synapses [119]. These structural synaptic changes observed during aging also lead to functional and behavioral deteriorations. Electrophysiological studies in the context of aging consistently show changes in synaptic transmission, excitability and plasticity. In general, aging is often associated with a decreased synaptic transmission and a reduced ability to increase synaptic strength through LTP [120,121]. Not all synapses, however, are equally affected by aging. For instance, aged rodents display defective presynaptic LTP at MF-CA3 synapses [119] and reduced excitability in CA1 [122] but hyperactivation in CA3 pyramidal neurons [123]. A recent study combining electron microscopy and behavioral experiments, set out to link hippocampal CA1 and CA3 synaptic architecture with cognitive ability during aging. They demonstrated that synapse architecture did not change in old rats that were performing as well in learning-tasks as their young counterparts. In contrast, old but learning-impaired animals exhibited structural synaptic changes only in CA3 synapses. Notably, synapses within hippocampal region CA3 showed an increased number of perforated synapses, a synapse characterized by gaps in post- and presynaptic densities, and a different distribution of synaptic weight as measured by AMPAR expression [124]. Thus, although in some brain regions specific neuronal loss is observed during aging processes [125], it is clearly the more subtle changes in plastic synaptic connections that drive cognitive decline during normal aging. Other studies set out to address what molecular changes occur that could underlie these age-induced changes in synaptic connections. Studies have shown age-related decline of synaptic proteins such as synaptophysin [126] and NMDA receptor subunits [127]. Recently, multiple large datasets have become available that include age-related neuron and synapse changes in mice, monkeys and humans on multiple levels including the transcriptome [128–133], proteome [92,134–136], lipidome [137], epigenome [138] and metabolome [139]. These studies now provide an important basis for identifying signatures for cell changes during aging; however, we still lack detailed understanding on what underlies these changes and how they can be directly related to synaptic connectivity changes and cognitive decline during aging.
In agreement with previous studies in C. elegans [140,141], a recent study employing in vivo metabolic labeling and proteomic profiling of mouse brains, correlated normal aging of wild-type mice with decreases in global protein recycling [134]. Why global proteostasis declines during normal aging processes is not well understood but includes decreases in the efficiency and cooperation of protein quality control systems. The next paragraphs will focus on how physiological aging affects the major presynaptic protein degradation routes.
Age-related changes in presynaptic degradation pathways
During normal brain aging, malfunctioning of autophagy processes are described in many organisms such as D. melanogaster, rodents, and humans. This includes changes in mTOR activity, gene expression, down-regulation of autophagy proteins (including ATG5, ATG7 and Beclin), and decreases in the amount of autophagosomes or autophagic flux [142–146]. A number of studies have shown that autophagy contributes to memory formation [63,147,148] and emerging studies in mice demonstrate a link between age-induced impairment of hippocampus-dependent memories and the down-regulation of certain autophagy factors, highlighting the need for functional neuronal autophagy in learning and memory. Importantly, genetic manipulations restoring autophagic function are sufficient to rescue age-related memory decline [147]. Similarly, the autophagy inducer spermidine protects from age-related synaptic alterations at murine hippocampal MF-CA3 synapses and counteracts age-dependent memory impairment in mice and fruit flies [82,119,149]. Moreover, elevated dietary spermidine intake is associated with a lower risk for cognitive impairment and decline in humans [82]. How autophagy is differentially modulated at aging presynapses remains unclear. Age-induced increases in the levels of the active zone scaffolding protein Bassoon might inhibit autophagy and, thereby, reduce release probability and synaptic transmission. Studies performed in primary mouse dorsal root ganglion neurons have revealed an age-dependent decrease in autophagosome biogenesis, caused by a local loss of the PI[3]P effector WIPI2B. This leads to the stalling of autophagosomes and failure to recruit LC3 at axon terminals. Strikingly, overexpression of WIPI2 was sufficient to return axonal autophagosome biogenesis to levels observed in younger neurons [150]. Whether hippocampal synapses experience similar age-associated autophagy impairments remains to be determined.
The relatively newly discovered chaperone-mediated autophagy (CMA) also decreases with age, predominantly due to changes in lysosomal lipid composition leading to reduced stability of LAMP2A [151]. Recent work on rodent models with neuron-specific LAMP2A loss demonstrated changes reminiscent of brain aging, including accumulation of oxidized and ubiquitinated proteins, synapse loss and memory impairments [52,152]. Although presynaptic accumulation of α-synuclein is observed after CMA loss [153], there has been little research on CMA at synapses and its role in modulating synaptic content and function needs to be determined.
Defects in the neuronal endosome-lysosome and ESCRT pathway, including pH increases and enlargement of early endosomes, are one of the earliest observed changes in neurodegenerative diseases such as Alzheimer’s disease (AD) and PD [154]. However, few data are available on changes in this pathway during normal brain aging. Recent findings identified the BAG3-HSP70 complex as a mediator of endolysosomal degradation in rat cortical neurons, exerting its effects by disinhibiting Rab35 activity [155]. Interestingly, BAG3 expression is decreased in AD brain pathology [156] but is increased during normal aging [157]. Whereas ESCRT proteins control the pathway towards lysosomal degradation, another protein complex called Retromer (i.e. comprising the proteins Vps35, Vps26 and Vps29) mediates the recycling of membrane proteins to the Golgi complex or back to the cell surface. The association of the core Retromer complex with different sorting nexin (SNX) proteins provides Retromer diversity in terms of cargo recognition and trafficking pathways [158]. Retromer cargos include for instance postsynaptic neurotransmitter receptors [159,160] and the dopamine transporter DAT, a presynaptic protein required for dopamine clearance following release [161]. Although little is known about how aging influences Retromer efficiency, an age-dependent decrease of most Retromer components is observed in the mouse cortex [162]. When Retromer-dependent processes are disrupted, many protein cargoes are misdirected to the degradative pathway, leading to improper recycling of for instance lysosomal cathepsins and neurodegeneration [163]. Recent studies in Drosophila suggest that presynaptic Retromer regulates SV recycling [164,165], although Vps35 depletion in primary hippocampal mouse neurons did not alter synaptic vesicle release or retrieval [166] .
In the aging brain, the UPS is proposed to be modified in different ways, including alterations in the ubiquitin-conjugating and DUB enzyme landscape [167–170] and the expression or transport of proteasome subunits [171,172]. The accumulation of protein inclusions due to proteostasis failure can also clog and inactivate the proteasome [173]. This proteostasis feedback during aging makes it difficult to dissect primary causes, such as changes in UPS expression, from secondary consequences due to protein overload. In synaptosomal fractions from aged monkey brains, levels of ubiquitin-conjugating enzyme UBE2N increase with aging while UPS activity declines, potentially leading to an accumulation of ubiquitinated and misfolded proteins in neuronal terminals [170]. Similarly, proteasome activity is decreased during aging in Drosophila, and enhancement of neuronal proteasome function prolongs lifespan and delays age-related decline in memory and learning [174], although the background mechanisms are so far unknown. Little is known about age-related UPS changes and their impact on cognition in humans. A recent proteomic study on enriched synaptic fractions from human brain samples connected the levels of proteasome subunits to cognitive performance [135]. Here, proteasome subunit levels were decreased in subjects that, despite AD pathology, are cognitively unimpaired but elevated in aged individuals with low pathology but with impaired cognition. In conclusion, whereas most studies indicate age-related decline in proteasome function, the relationship between cognitive aging and impairment of the UPR are far from clear.
Conclusions and open questions
In recent years, we have gained fundamental insight into the degradative mechanisms involved in synaptic protein turnover and their relevance for neurotransmission. Both basal and stress-evoked pathways can damage synapse integrity and further research will be vital to uncover how synaptic components such as aged SVs are recognized and tagged for degradation. In addition, how do different regulators and signals such as neuronal activity, mTOR and ER-stress impinge on local degradative pathways in the synapse? How is the synaptic availability and activity of ubiquitin ligases and DUBs regulated, and how do all these different degradative pathways cooperate to sustain synapse integrity?
Much of what we know about aging comes from rodents, mostly from transgenic mouse model of age-related neurodegenerative diseases such as AD and PD. However, emerging studies show different, and often opposite, effects on the brain during neuropathology as compared with normal aging, indicating that neurodegenerative diseases are not a mere acceleration of aging. Whereas protein recycling in the brain decreases in aged, but otherwise healthy, mice, increased protein turnover is associated with aging in, for instance, mouse models of Alzheimer’s disease [134]. Here, differential effects on degradative pathways might account for the discrepancy. For instance during pathology an unusual high amount of misfolded proteins can clog the proteasome and other degradative pathways. It is also important to recognize that individual synapses show high degree of diversity arising from molecular and morphological differences [175]. We know that region-specific vulnerability underlies pathological brain aging, with for instance the hippocampus primarily affected in Alzheimer’s disease, the substantia nigra in Parkinson’s disease and the ventral spinal cord in amyotrophic lateral sclerosis [176]. During physiological aging, changes in synaptic architecture, composition, and function also vary depending on brain region and synapse type [110,124]. In addition, not all brain areas experience the same age-related changes in degradative pathways [177], indicating that physiological brain aging is as diverse as pathological brain aging. Therefore, we have to take into account that population-averaging measurements might not be able to detect all changes that happen with age. What is underlying this differential vulnerability of neuronal and synaptic populations is unclear but likely depends on intrinsic differences between synapses, such as their activity, their turnover rates or dependence on, and availability of, certain proteolytic systems. Recent studies also suggest cell extrinsic factors to play a role in age-dependent synapse decline, including systemic factors such as microbial metabolites (reviewed in [178]). How environmental factors shape protein homeostasis in humans is still an open debate but non-pharmacological methods such as caloric restriction and physical exercise increase neuronal plasticity and resistance of neurons to age-related decline in humans, possibly via common pathways such as mTOR and autophagy [179]. Finding the molecular pathways involved in synapse aging will be essential to design future interventions to halt or even reverse brain aging.
Acknowledgements
The author thanks Barth van Rossum (FMP, Berlin) for help with the artwork and Marta Maglione, Domenico Azarnia Tehran and Volker Haucke for critically reading the manuscript.
Abbreviations
- AD
Alzheimer’s disease
- CMA
chaperone-mediated autophagy
- mTOR
mechanistic target of rapamycin
- PD
Parkinson’s disease
- PI3K
phosphoinositide 3-kinase
- PKA
protein kinase A
- ROS
reactive oxygen species
- UPR
unfolded protein response
- UPS
ubiquitin–proteasome system
Data Availability
No data were generated for this review.
Competing Interests
The authors declare that there are no competing interests associated with the manuscript.
Funding
M.K. was supported by a Marie Skłodowska-Curie Postdoctoral Fellowship from the European Union [EU, Horizon 2020, 655604-SYNPT].
CRediT Author Contribution
Marijn Kuijpers: Conceptualization, Writing—original draft, Writing—review & editing.
References
- 1.Cajigas I.J., Will T. and Schuman E.M. (2010) Protein homeostasis and synaptic plasticity. EMBO J. 29, 2746–2752 10.1038/emboj.2010.173 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hegde A.N. (2017) Proteolysis, synaptic plasticity and memory. Neurobiol. Learn. Mem. 138, 98–110 10.1016/j.nlm.2016.09.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sonninen T.M., Goldsteins G., Laham-Karam N., Koistinaho J. and Lehtonen S. (2020) Proteostasis disturbances and inflammation in neurodegenerative diseases. Cells 9, 10.3390/cells9102183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Pulido C. and Ryan T.A. (2021) Synaptic vesicle pools are a major hidden resting metabolic burden of nerve terminals. Sci. Adv. 7, eabi9027 10.1126/sciadv.abi9027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Heo S., Diering G.H., Na C.H., Nirujogi R.S., Bachman J.L., Pandey A.et al. (2018) Identification of long-lived synaptic proteins by proteomic analysis of synaptosome protein turnover. Proc. Natl. Acad. Sci. U.S.A. 115, E3827–E3836 10.1073/pnas.1720956115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zheng N. and Shabek N. (2017) Ubiquitin ligases: structure, function, and regulation. Annu. Rev. Biochem. 86, 129–157 10.1146/annurev-biochem-060815-014922 [DOI] [PubMed] [Google Scholar]
- 7.Kocaturk N.M. and Gozuacik D. (2018) Crosstalk between mammalian autophagy and the ubiquitin-proteasome system. Front. Cell Dev. Biol. 6, 10.3389/fcell.2018.00128 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hamilton A.M., Oh W.C., Vega-Ramirez H., Stein I.S., Hell J.W., Patrick G.N.et al. (2012) Activity-dependent growth of new dendritic spines is regulated by the proteasome. Neuron 74, 1023–1030 10.1016/j.neuron.2012.04.031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Pinto M.J., Alves P.L., Martins L., Pedro J.R., Ryu H.R., Jeon N.L.et al. (2016) The proteasome controls presynaptic differentiation through modulation of an on-site pool of polyubiquitinated conjugates. J. Cell Biol. 212, 789–801 10.1083/jcb.201509039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Valnegri P., Huang J., Yamada T., Yang Y., Mejia L.A., Cho H.Y.et al. (2017) RNF8/UBC13 ubiquitin signaling suppresses synapse formation in the mammalian brain. Nat. Commun. 8, 1271 10.1038/s41467-017-01333-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Fonseca R., Vabulas R.M., Hartl F.U., Bonhoeffer T. and Nagerl U.V. (2006) A balance of protein synthesis and proteasome-dependent degradation determines the maintenance of LTP. Neuron 52, 239–245 10.1016/j.neuron.2006.08.015 [DOI] [PubMed] [Google Scholar]
- 12.Turker F., Cook E.K. and Margolis S.S. (2021) The proteasome and its role in the nervous system. Cell Chem Biol. 28, 903–917 10.1016/j.chembiol.2021.04.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Radulescu C.I., Cerar V., Haslehurst P., Kopanitsa M. and Barnes S.J. (2021) The aging mouse brain: cognition, connectivity and calcium. Cell Calcium 94, 102358 10.1016/j.ceca.2021.102358 [DOI] [PubMed] [Google Scholar]
- 14.Bliss T.V. and Collingridge G.L. (1993) A synaptic model of memory: long-term potentiation in the hippocampus. Nature 361, 31–39 10.1038/361031a0 [DOI] [PubMed] [Google Scholar]
- 15.Dong C., Upadhya S.C., Ding L., Smith T.K. and Hegde A.N. (2008) Proteasome inhibition enhances the induction and impairs the maintenance of late-phase long-term potentiation. Learn. Mem. 15, 335–347 10.1101/lm.984508 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Karpova A., Mikhaylova M., Thomas U., Knopfel T. and Behnisch T. (2006) Involvement of protein synthesis and degradation in long-term potentiation of Schaffer collateral CA1 synapses. J. Neurosci. 26, 4949–4955 10.1523/JNEUROSCI.4573-05.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lee S.H., Choi J.H., Lee N., Lee H.R., Kim J.I., Yu N.K.et al. (2008) Synaptic protein degradation underlies destabilization of retrieved fear memory. Science 319, 1253–1256 10.1126/science.1150541 [DOI] [PubMed] [Google Scholar]
- 18.Lopez-Salon M., Alonso M., Vianna M.R.M., Viola H., Souza T.M.E., Izquierdo I.et al. (2001) The ubiquitin-proteasome cascade is required for mammalian long-term memory formation. Eur. J. Neurosci. 14, 1820–1826 10.1046/j.0953-816x.2001.01806.x [DOI] [PubMed] [Google Scholar]
- 19.Ehlers M.D. (2003) Activity level controls postsynaptic composition and signaling via the ubiquitin-proteasome system. Nat. Neurosci. 6, 231–242 10.1038/nn1013 [DOI] [PubMed] [Google Scholar]
- 20.Speese S.D., Trotta N., Rodesch C.K., Aravamudan B. and Broadie K. (2003) The ubiquitin proteasome system acutely regulates presynaptic protein turnover and synaptic efficacy. Curr. Biol. 13, 899–910 10.1016/S0960-9822(03)00338-5 [DOI] [PubMed] [Google Scholar]
- 21.Willeumier K., Pulst S.M. and Schweizer F.E. (2006) Proteasome inhibition triggers activity-dependent increase in the size of the recycling vesicle pool in cultured hippocampal neurons. J. Neurosci. 26, 11333–11341 10.1523/JNEUROSCI.1684-06.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ramachandran K.V., Fu J.M., Schaffer T.B., Na C.H., Delannoy M. and Margolis S.S. (2018) Activity-dependent degradation of the nascentome by the neuronal membrane proteasome. Mol. Cell. 71, 169e6–177e6 10.1016/j.molcel.2018.06.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liu K., Jones S., Minis A., Rodriguez J., Molina H. and Steller H. (2019) PI31 is an adaptor protein for proteasome transport in axons and required for synaptic development. Dev. Cell. 50, 509e10–524e10 10.1016/j.devcel.2019.06.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Minis A., Rodriguez J.A., Levin A., Liu K., Govek E.E., Hatten M.E.et al. (2019) The proteasome regulator PI31 is required for protein homeostasis, synapse maintenance, and neuronal survival in mice. Proc. Natl. Acad. Sci. U. S. A 116, 24639–24650 10.1073/pnas.1911921116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jiang X.P., Litkowski P.E., Taylor A.A., Lin Y., Snider B.J. and Moulder K.L. (2010) A role for the ubiquitin-proteasome system in activity-dependent presynaptic silencing. J. Neurosci. 30, 1798–1809 10.1523/JNEUROSCI.4965-09.2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lazarevic V., Schone C., Heine M., Gundelfinger E.D. and Fejtova A. (2011) Extensive remodeling of the presynaptic cytomatrix upon homeostatic adaptation to network activity silencing. J. Neurosci. 31, 10189–10200 10.1523/JNEUROSCI.2088-11.2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Vitry S., Ausseil J., Hocquemiller M., Bigou S., Dos Santos Coura R. and Heard J.M. (2009) Enhanced degradation of synaptophysin by the proteasome in mucopolysaccharidosis type IIIB. Mol. Cell. Neurosci. 41, 8–18 10.1016/j.mcn.2009.01.001 [DOI] [PubMed] [Google Scholar]
- 28.Hakim V., Cohen L.D., Zuchman R., Ziv T. and Ziv N.E. (2016) The effects of proteasomal inhibition on synaptic proteostasis. EMBO J. 35, 2238–2262 10.15252/embj.201593594 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Chin L.S., Vavalle J.P. and Li L. (2002) Staring, a novel E3 ubiquitin-protein ligase that targets syntaxin 1for degradation. J. Biol. Chem. 277, 35071–35079 10.1074/jbc.M203300200 [DOI] [PubMed] [Google Scholar]
- 30.Tada H., Okano H.J., Takagi H., Shibata S., Yao I., Matsumoto M.et al. (2010) Fbxo45, a novel ubiquitin ligase, regulates synaptic activity. J. Biol. Chem. 285, 3840–3849 10.1074/jbc.M109.046284 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yao I., Takagi H., Ageta H., Kahyo T., Sato S., Hatanaka K.et al. (2007) SCRAPPER-dependent ubiquitination of active zone protein RIM1 regulates synaptic vesicle release. Cell 130, 943–957 10.1016/j.cell.2007.06.052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Waites C.L., Leal-Ortiz S.A., Okerlund N., Dalke H., Fejtova A., Altrock W.D.et al. (2013) Bassoon and Piccolo maintain synapse integrity by regulating protein ubiquitination and degradation. EMBO J. 32, 954–969 10.1038/emboj.2013.27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Montenegro-Venegas C., Fienko S., Anni D., Pina-Fernández E., Frischknecht R. and Fejtova A. (2021) Bassoon inhibits proteasome activity via interaction with PSMB4. Cell. Mol. Life Sci. 78, 1545–1563 10.1007/s00018-020-03590-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gupta V.K., Pech U., Bhukel A., Fulterer A., Ender A., Mauermann S.F.et al. (2016) Spermidine suppresses age-associated memory impairment by preventing adverse increase of presynaptic active zone size and release. PLoS Biol. 14, e1002563 10.1371/journal.pbio.1002563 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Huo Y., Khatri N., Hou Q., Gilbert J., Wang G. and Man H.Y. (2015) The deubiquitinating enzyme USP46 regulates AMPA receptor ubiquitination and trafficking. J. Neurochem. 134, 1067–1080 10.1111/jnc.13194 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Klionsky D.J. (2007) Autophagy: from phenomenology to molecular understanding in less than a decade. Nat. Rev. Mol. Cell Bio. 8, 931–937 10.1038/nrm2245 [DOI] [PubMed] [Google Scholar]
- 37.Kuijpers M., Azarnia Tehran D., Haucke V. and Soykan T. (2021) The axonal endolysosomal and autophagic systems. J. Neurochem. 158, 589–602 10.1111/jnc.15287 [DOI] [PubMed] [Google Scholar]
- 38.Huotari J. and Helenius A. (2011) Endosome maturation. Embo J. 30, 3481–3500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Overly C.C. and Hollenbeck P.J. (1996) Dynamic organization of endocytic pathways in axons of cultured sympathetic neurons. J. Neurosci. 16, 6056–6064 10.1523/JNEUROSCI.16-19-06056.1996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wallroth A. and Haucke V. (2018) Phosphoinositide conversion in endocytosis and the endolysosomal system. J. Biol. Chem. 293, 1526–1535 10.1074/jbc.R117.000629 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Borchers A.C., Langemeyer L. and Ungermann C. (2021) Who's in control? Principles of Rab GTPase activation in endolysosomal membrane trafficking and beyond J. Cell Biol. 220, 10.1083/jcb.202105120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Vietri M., Radulovic M. and Stenmark H. (2020) The many functions of ESCRTs. Nat. Rev. Mol. Cell Biol. 21, 25–42 10.1038/s41580-019-0177-4 [DOI] [PubMed] [Google Scholar]
- 43.Sheehan P., Zhu M., Beskow A., Vollmer C. and Waites C.L. (2016) Activity-dependent degradation of synaptic vesicle proteins requires Rab35 and the ESCRT pathway. J. Neurosci. 36, 8668–8686 10.1523/JNEUROSCI.0725-16.2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Uytterhoeven V., Kuenen S., Kasprowicz J., Miskiewicz K. and Verstreken P. (2011) Loss of skywalker reveals synaptic endosomes as sorting stations for synaptic vesicle proteins. Cell 145, 117–132 10.1016/j.cell.2011.02.039 [DOI] [PubMed] [Google Scholar]
- 45.Clayton E.L., Bonnycastle K., Isaacs A.M., Cousin M.A. and Schorge S. (2022) A novel synaptopathy-defective synaptic vesicle protein trafficking in the mutant CHMP2B mouse model of frontotemporal dementia. J. Neurochem. 160, 412–425 10.1111/jnc.15551 [DOI] [PubMed] [Google Scholar]
- 46.Jin E.J., Kiral F.R., Ozel M.N., Burchardt L.S., Osterland M., Epstein D.et al. (2018) Live observation of two parallel membrane degradation pathways at axon terminals. Curr. Biol. 28, 1027.e4–1038.e4 10.1016/j.cub.2018.02.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Uytterhoeven V., Lauwers E., Maes I., Miskiewicz K., Melo M.N., Swerts J.et al. (2015) Hsc70-Hsc74 deforms membranes to promote synaptic protein turnover by endosomal microautophagy. Neuron 88, 735–748 10.1016/j.neuron.2015.10.012 [DOI] [PubMed] [Google Scholar]
- 48.Birdsall V., Zhu M., Kirwan K., Imoto Y., Watanabe S. and Waites C.L. (2021) Axonal transport of Hrs is activity-dependent and rate limiting for synaptic vesicle protein degradation. bioRxiv 2020.04.16.044818 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ivanova D. and Cousin M.A. (2022) Synaptic vesicle recycling and the endolysosomal system: a reappraisal of form and function. Front. Synaptic Neurosci. 14, 826098 10.3389/fnsyn.2022.826098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Turegano-Lopez M., Santuy A., DeFelipe J. and Merchan-Perez A. (2020) Size, shape, and distribution of multivesicular bodies in the juvenile rat somatosensory cortex: a 3D electron microscopy study. Cereb. Cortex 30, 1887–1901 10.1093/cercor/bhz211 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Schuck S. (2020) Microautophagy - distinct molecular mechanisms handle cargoes of many sizes. J. Cell Sci. 133, jcs246322 10.1242/jcs.246322 [DOI] [PubMed] [Google Scholar]
- 52.Bourdenx M., Gavathiotis E. and Cuervo A.M. (2021) Chaperone-mediated autophagy: a gatekeeper of neuronal proteostasis. Autophagy 17, 2040–2042 10.1080/15548627.2021.1935007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Cheng X.T., Zhou B., Lin M.Y., Cai Q. and Sheng Z.H. (2015) Axonal autophagosomes use the ride-on service for retrograde transport toward the soma. Autophagy 11, 1434–1436 10.1080/15548627.2015.1062203 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Melia T.J., Lystad A.H. and Simonsen A. (2020) Autophagosome biogenesis: From membrane growth to closure. J. Cell Biol. 219, e202002085 10.1083/jcb.202002085 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Gubas A. and Dikic I. (2022) A guide to the regulation of selective autophagy receptors. FEBS J. 289, 75–89 10.1111/febs.15824 [DOI] [PubMed] [Google Scholar]
- 56.Cason S.E., Carman P.J., van Duyne C., Goldsmith J., Dominguez R. and Holzbaur E.L.F. (2021) Sequential dynein effectors regulate axonal autophagosome motility in a maturation-dependent pathway. J. Cell Biol. 220, e202010179 10.1083/jcb.202010179 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Kononenko N.L., Claßen G.A., Kuijpers M., Puchkov D., Maritzen T., Tempes A.et al. (2017) Retrograde transport of TrkB-containing autophagosomes via the adaptor AP-2 mediates neuronal complexity and prevents neurodegeneration. Nat. Commun. 8, 14819 10.1038/ncomms14819 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Soukup S.F., Kuenen S., Vanhauwaert R., Manetsberger J., Hernández-Díaz S., Swerts J.et al. (2016) A LRRK2-dependent EndophilinA phosphoswitch is critical for macroautophagy at presynaptic terminals. Neuron 92, 829–844 10.1016/j.neuron.2016.09.037 [DOI] [PubMed] [Google Scholar]
- 59.Hernandez-Diaz S., Ghimire S., Sanchez-Mirasierra I., Montecinos-Oliva C., Swerts J., Kuenen S.et al. (2022) Endophilin-B regulates autophagy during synapse development and neurodegeneration. Neurobiol. Dis. 163, 105595 10.1016/j.nbd.2021.105595 [DOI] [PubMed] [Google Scholar]
- 60.Okerlund N.D., Schneider K., Leal-Ortiz S., Montenegro-Venegas C., Kim S.A., Garner L.C.et al. (2017) Bassoon controls presynaptic autophagy through Atg5. Neuron 93, 897–+ 10.1016/j.neuron.2017.01.026 [DOI] [PubMed] [Google Scholar]
- 61.Vanhauwaert R., Kuenen S., Masius R., Bademosi A., Manetsberger J., Schoovaerts N.et al. (2017) The SAC1 domain in synaptojanin is required for autophagosome maturation at presynaptic terminals. EMBO J. 36, 1392–1411 10.15252/embj.201695773 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Stavoe A.K.H. and Holzbaur E.L.F. (2019) Axonal autophagy: Mini-review for autophagy in the CNS. Neurosci. Lett. 697, 17–23 10.1016/j.neulet.2018.03.025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Compans B., Camus C., Kallergi E., Sposini S., Martineau M., Butler C.et al. (2021) NMDAR-dependent long-term depression is associated with increased short term plasticity through autophagy mediated loss of PSD-95. Nat. Commun. 12, 2849 10.1038/s41467-021-23133-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kallergi E., Daskalaki A.D., Kolaxi A., Camus C., Ioannou E., Mercaldo V.et al. (2022) Dendritic autophagy degrades postsynaptic proteins and is required for long-term synaptic depression in mice. Nat. Commun. 13, 680 10.1038/s41467-022-28301-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Hill S.E., Kauffman K.J., Krout M., Richmond J.E., Melia T.J. and Colon-Ramos D.A. (2019) Maturation and clearance of autophagosomes in neurons depends on a specific cysteine protease isoform, ATG-4.2. Dev. Cell. 49, 251–266, e8 10.1016/j.devcel.2019.02.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Hoffmann S., Orlando M., Andrzejak E., Bruns C., Trimbuch T., Rosenmund C.et al. (2019) Light-Activated ROS Production Induces Synaptic Autophagy. J. Neurosci. 39, 2163–2183 10.1523/JNEUROSCI.1317-18.2019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Goldsmith J., Ordureau A., Harper J.W. and Holzbaur E.L.F. (2022) Brain-derived autophagosome profiling reveals the engulfment of nucleoid-enriched mitochondrial fragments by basal autophagy in neurons. Neuron 110, 967–976 10.1016/j.neuron.2021.12.029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Maday S. and Holzbaur E.L. (2012) Autophagosome assembly and cargo capture in the distal axon. Autophagy 8, 858–860 10.4161/auto.20055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Kuijpers M., Kochlamazashvili G., Stumpf A., Puchkov D., Swaminathan A., Lucht M.T.et al. (2022) Neuronal autophagy regulates presynaptic neurotransmission by controlling the axonal endoplasmic reticulum. Neuron 110, 734 10.1016/j.neuron.2022.01.029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Binotti B., Pavlos N.J., Riedel D., Wenzel D., Vorbrüggen G., Schalk A.M.et al. (2015) The GTPase Rab26 links synaptic vesicles to the autophagy pathway. Elife 4, e05597 10.7554/eLife.05597 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Truckenbrodt S., Viplav A., Jähne S., Vogts A., Denker A., Wildhagen H.et al. (2018) Newly produced synaptic vesicle proteins are preferentially used in synaptic transmission. EMBO J. 37, e98044 10.15252/embj.201798044 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Hernandez D., Torres C.A., Setlik W., Cebrián C., Mosharov E.V., Tang G.et al. (2012) Regulation of presynaptic neurotransmission by macroautophagy. Neuron 74, 277–284 10.1016/j.neuron.2012.02.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Lüningschrör P., Binotti B., Dombert B., Heimann P., Perez-Lara A., Slotta C.et al. (2017) Plekhg5-regulated autophagy of synaptic vesicles reveals a pathogenic mechanism in motoneuron disease. Nat. Commun. 8, 678 10.1038/s41467-017-00689-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kohrs F.E., Daumann I.M., Pavlovic B., Jin E.J., Kiral F.R., Lin S.C.et al. (2021) Systematic functional analysis of rab GTPases reveals limits of neuronal robustness to environmental challenges in flies. Elife 10, e59594 10.7554/eLife.59594 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Nakatogawa H. (2020) Mechanisms governing autophagosome biogenesis. Nat. Rev. Mol. Cell Biol. 21, 439–458 10.1038/s41580-020-0241-0 [DOI] [PubMed] [Google Scholar]
- 76.Stavoe A.K., Hill S.E., Hall D.H. and Colón-Ramos D.A. (2016) KIF1A/UNC-104 transports ATG-9 to regulate neurodevelopment and autophagy at synapses. Dev. Cell. 38, 171–185 10.1016/j.devcel.2016.06.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Yang S., Park D., Manning L., Hill S.E., Cao M., Xuan Z.et al. (2022) Presynaptic autophagy is coupled to the synaptic vesicle cycle via ATG-9. Neuron 110, 824.e10–840.e10 10.1016/j.neuron.2021.12.031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Overhoff M., Tellkamp F., Hess S., Tutas J., Tolve M., Faerfers M.et al. (2022) Autophagy regulates neuronal excitability by controlling cAMP/Protein Kinase A signaling. bioRxiv 2022.02.11.480034 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Lees R.M., Johnson J.D. and Ashby M.C. (2020) Presynaptic boutons that contain mitochondria are more stable. Front. Synaptic Neuro. 11, 37 10.3389/fnsyn.2019.00037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Todorova V. and Blokland A. (2017) Mitochondria and synaptic plasticity in the mature and aging nervous system. Curr. Neuropharmacol. 15, 166–173 10.2174/1570159X14666160414111821 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Liang Y., Piao C., Beuschel C.B., Toppe D., Kollipara L., Bogdanow B.et al. (2021) eIF5A hypusination, boosted by dietary spermidine, protects from premature brain aging and mitochondrial dysfunction. Cell Rep. 35, 108941 10.1016/j.celrep.2021.108941 [DOI] [PubMed] [Google Scholar]
- 82.Schroeder S., Hofer S.J., Zimmermann A., Pechlaner R., Dammbrueck C., Pendl T.et al. (2021) Dietary spermidine improves cognitive function. Cell Rep. 35, 108985 10.1016/j.celrep.2021.108985 [DOI] [PubMed] [Google Scholar]
- 83.Grochowska K.M., Andres-Alonso M., Karpova A. and Kreutz M.R. (2022) The needs of a synapse-How local organelles serve synaptic proteostasis. EMBO J. e110057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Hafner A.S., Donlin-Asp P.G., Leitch B., Herzog E. and Schuman E.M. (2019) Local protein synthesis is a ubiquitous feature of neuronal pre- and postsynaptic compartments. Science 364, 650–+ 10.1126/science.aau3644 [DOI] [PubMed] [Google Scholar]
- 85.Hwang H.J., Ha H., Lee B.S., Kim B.H., Song H.K. and Kim Y.K. (2022) LC3B is an RNA-binding protein to trigger rapid mRNA degradation during autophagy. Nat. Commun. 13, 1436 10.1038/s41467-022-29139-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Shehata M., Matsumura H., Okubo-Suzuki R., Ohkawa N. and Inokuchi K. (2012) Neuronal stimulation induces autophagy in hippocampal neurons that is involved in AMPA receptor degradation after chemical long-term depression. J. Neurosci. 32, 10413–10422 10.1523/JNEUROSCI.4533-11.2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Raab-Graham K.F., Haddick P.C., Jan Y.N. and Jan L.Y. (2006) Activity- and mTOR-dependent suppression of Kv1.1 channel mRNA translation in dendrites. Science 314, 144–148 10.1126/science.1131693 [DOI] [PubMed] [Google Scholar]
- 88.Takei N., Furukawa K., Hanyu O., Sone H. and Nawa H. (2014) A possible link between BDNF and mTOR in control of food intake. Front. Psychol. 5, 1093 10.3389/fpsyg.2014.01093 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Zhao J.H., Zhai B., Gygi S.P. and Goldberg A.L. (2015) mTOR inhibition activates overall protein degradation by the ubiquitin proteasome system as well as by autophagy. Proc. Natl. Acad. Sci. U. S. A. 112, 15790–15797 10.1073/pnas.1521919112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.McCabe M.P., Cullen E.R., Barrows C.M., Shore A.N., Tooke K.I., Laprade K.A.et al. (2020) Genetic inactivation of mTORC1 or mTORC2 in neurons reveals distinct functions in glutamatergic synaptic transmission. Elife 9, e51440 10.7554/eLife.51440 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Hetz C. and Saxena S. (2017) ER stress and the unfolded protein response in neurodegeneration. Nat. Rev. Neurol. 13, 477–491 10.1038/nrneurol.2017.99 [DOI] [PubMed] [Google Scholar]
- 92.Cabral-Miranda F., Tamburini G., Martinez G., Medinas D., Gerakis Y., Miedema T.et al. (2020) Control of mammalian brain aging by the unfolded protein response (UPR). bioRxiv, 10.1101/2020.04.13.039172 [DOI] [Google Scholar]
- 93.Krukowski K., Nolan A., Frias E.S., Boone M., Ureta G., Grue K.et al. (2020) Small molecule cognitive enhancer reverses age-related memory decline in mice. Elife 9, e62048 10.7554/eLife.62048 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Saito A., Cai L., Matsuhisa K., Ohtake Y., Kaneko M., Kanemoto S.et al. (2018) Neuronal activity-dependent local activation of dendritic unfolded protein response promotes expression of brain-derived neurotrophic factor in cell soma. J. Neurochem. 144, 35–49 10.1111/jnc.14221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Yoshii S.R., Kishi C., Ishihara N. and Mizushima N. (2011) Parkin mediates proteasome-dependent protein degradation and rupture of the outer mitochondrial membrane. J. Biol. Chem. 286, 19630–19640 10.1074/jbc.M110.209338 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Narendra D., Tanaka A., Suen D.F. and Youle R.J. (2008) Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 183, 795–803 10.1083/jcb.200809125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Hoffmann-Conaway S., Brockmann M.M., Schneider K., Annamneedi A., Rahman K.A., Bruns C.et al. (2020) Parkin contributes to synaptic vesicle autophagy in Bassoon-deficient mice. Elife 9, e56590 10.7554/eLife.56590 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Lee S., Park S., Lee H., Han S., Song J.M., Han D.et al. (2019) Nedd4 E3 ligase and beta-arrestins regulate ubiquitination, trafficking, and stability of the mGlu7 receptor. Elife 8, e44502 10.7554/eLife.44502 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Liu W.J., Ye L., Huang W.F., Guo L.J., Xu Z.G., Wu H.L.et al. (2016) p62 links the autophagy pathway and the ubiqutin-proteasome system upon ubiquitinated protein degradation. Cell. Mol. Biol. Lett. 21, 29 10.1186/s11658-016-0031-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Kristensen A.R., Schandorff S., Høyer-Hansen M., Nielsen M.O., Jäättelä M., Dengjel J.et al. (2008) Ordered organelle degradation during starvation-induced autophagy. Mol. Cell. Proteomics 7, 2419–2428 10.1074/mcp.M800184-MCP200 [DOI] [PubMed] [Google Scholar]
- 101.Andres-Alonso M., Kreutz M.R. and Karpova A. (2021) Autophagy and the endolysosomal system in presynaptic function. Cell. Mol. Life Sci. 78, 2621–2639 10.1007/s00018-020-03722-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Petanjek Z., Judaš M., Šimic G., Rasin M.R., Uylings H.B., Rakic P.et al. (2011) Extraordinary neoteny of synaptic spines in the human prefrontal cortex. Proc. Natl. Acad. Sci. U. S. A. 108, 13281–13286 10.1073/pnas.1105108108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Grutzendler J., Kasthuri N. and Gan W.B. (2002) Long-term dendritic spine stability in the adult cortex. Nature 420, 812–816 10.1038/nature01276 [DOI] [PubMed] [Google Scholar]
- 104.Attardo A., Fitzgerald J.E. and Schnitzer M.J. (2015) Impermanence of dendritic spines in live adult CA1 hippocampus. Nature 523, 592–596 10.1038/nature14467 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Pfeiffer T., Poll S., Bancelin S., Angibaud J., Inavalli V.K., Keppler K.et al. (2018) Chronic 2P-STED imaging reveals high turnover of dendritic spines in the hippocampus in vivo. Elife 7, e34700 10.7554/eLife.34700 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Quinn D.P., Kolar A., Harris S.A., Wigerius M., Fawcett J.P. and Krueger S.R. (2019) The stability of glutamatergic synapses is independent of activity level, but predicted by synapse size. Front. Cell Neurosci. 13, 291 10.3389/fncel.2019.00291 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Ash R.T., Fahey P.G., Park J., Zoghbi H.Y. and Smirnakis S.M. (2018) Increased axonal bouton stability during learning in the mouse model of MECP2 duplication syndrome. eNeuro 5, 10.1523/ENEURO.0056-17.2018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Holtmaat A. and Caroni P. (2016) Functional and structural underpinnings of neuronal assembly formation in learning. Nat. Neurosci. 19, 1553–1562 10.1038/nn.4418 [DOI] [PubMed] [Google Scholar]
- 109.Morrison J.H. and Baxter M.G. (2012) The ageing cortical synapse: hallmarks and implications for cognitive decline. Nat. Rev. Neurosci. 13, 240–250 10.1038/nrn3200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Cizeron M., Qiu Z., Koniaris B., Gokhale R., Komiyama N.H., Fransen E.et al. (2020) A brainwide atlas of synapses across the mouse life span. Science 369, 270–275 10.1126/science.aba3163 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Huang L.Y., Zhou H., Chen K., Chen X. and Yang G. (2020) Learning-dependent dendritic spine plasticity is reduced in the aged mouse cortex. Front. Neural Circuit 14, 581435 10.3389/fncir.2020.581435 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Wallace M., Frankfurt M., Arellanos A., Inagaki T. and Luine V. (2007) Impaired recognition memory and decreased prefrontal cortex spine density in aged female rats. Ann. N. Y. Acad. Sci. 1097, 54–57 10.1196/annals.1379.026 [DOI] [PubMed] [Google Scholar]
- 113.Davidson A.M., Mejia-Gomez H., Jacobowitz M. and Mostany R. (2020) Dendritic spine density and dynamics of layer 5 pyramidal neurons of the primary motor cortex are elevated with aging. Cereb. Cortex 30, 767–777 10.1093/cercor/bhz124 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Geinisman Y., Ganeshina O., Yoshida R., Berry R.W., Disterhoft J.F. and Gallagher M. (2004) Aging, spatial learning, and total synapse number in the rat CA1 stratum radiatum. Neurobiol. Aging 25, 407–416 10.1016/j.neurobiolaging.2003.12.001 [DOI] [PubMed] [Google Scholar]
- 115.Page T.L., Einstein M., Duan H., He Y., Flores T., Rolshud D.et al. (2002) Morphological alterations in neurons forming corticocortical projections in the neocortex of aged Patas monkeys. Neurosci. Lett. 317, 37–41 10.1016/S0304-3940(01)02428-4 [DOI] [PubMed] [Google Scholar]
- 116.Mostany R., Anstey J.E., Crump K.L., Maco B., Knott G. and Portera-Cailliau C. (2013) Altered synaptic dynamics during normal brain aging. J. Neurosci. 33, 4094–4104 10.1523/JNEUROSCI.4825-12.2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Grillo F.W., Song S., Teles-Grilo Ruivo L.M., Huang L., Gao G., Knott G.W.et al. (2013) Increased axonal bouton dynamics in the aging mouse cortex. Proc. Natl. Acad. Sci. U. S. A. 110, E1514–E1523 10.1073/pnas.1218731110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Burke S.N. and Barnes C.A. (2010) Senescent synapses and hippocampal circuit dynamics. Trends Neurosci. 33, 153–161 10.1016/j.tins.2009.12.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Maglione M., Kochlamazashvili G., Eisenberg T., Racz B., Michael E., Toppe D.et al. (2019) Spermidine protects from age-related synaptic alterations at hippocampal mossy fiber-CA3 synapses. Sci. Rep.-Uk 9, 19616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Rex C.S., Kramár E.A., Colgin L.L., Lin B., Gall C.M. and Lynch G. (2005) Long-term potentiation is impaired in middle-aged rats: regional specificity and reversal by adenosine receptor antagonists. J. Neurosci. 25, 5956–5966 10.1523/JNEUROSCI.0880-05.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Rizzo V., Richman J. and Puthanveettil S.V. (2014) Dissecting mechanisms of brain aging by studying the intrinsic excitability of neurons. Front. Aging Neurosci. 6, 337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Moyer J.R. Jr., Power J.M., Thompson L.T. and Disterhoft J.F. (2000) Increased excitability of aged rabbit CA1 neurons after trace eyeblink conditioning. J. Neurosci. 20, 5476–5482 10.1523/JNEUROSCI.20-14-05476.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Simkin D., Hattori S., Ybarra N., Musial T.F., Buss E.W., Richter H.et al. (2015) Aging-related hyperexcitability in CA3 pyramidal neurons is mediated by enhanced A-type K+ channel function and expression. J. Neurosci. 35, 13206–13218 10.1523/JNEUROSCI.0193-15.2015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Buss E.W., Corbett N.J., Roberts J.G., Ybarra N., Musial T.F., Simkin D.et al. (2021) Cognitive aging is associated with redistribution of synaptic weights in the hippocampus. Proc. Natl. Acad. Sci. U. S. A. 118, e1921481118 10.1073/pnas.1921481118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Xu B., Sun A., He Y., Qian F., Xi S., Long D.et al. (2018) Loss of thin spines and small synapses contributes to defective hippocampal function in aged mice. Neurobiol. Aging 71, 91–104 10.1016/j.neurobiolaging.2018.07.010 [DOI] [PubMed] [Google Scholar]
- 126.Smith T.D., Adams M.M., Gallagher M., Morrison J.H. and Rapp P.R. (2000) Circuit-specific alterations in hippocampal synaptophysin immunoreactivity predict spatial learning impairment in aged rats. J. Neurosci. 20, 6587–6593 10.1523/JNEUROSCI.20-17-06587.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Magnusson K.R., Nelson S.E. and Young A.B. (2002) Age-related changes in the protein expression of subunits of the NMDA receptor. Brain Res. Mol. Brain Res. 99, 40–45 10.1016/S0169-328X(01)00344-8 [DOI] [PubMed] [Google Scholar]
- 128.Kang H.J., Kawasawa Y.I., Cheng F., Zhu Y., Xu X., Li M.et al. (2011) Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 10.1038/nature10523 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Lu T., Pan Y., Kao S.Y., Li C., Kohane I., Chan J.et al. (2004) Gene regulation and DNA damage in the ageing human brain. Nature 429, 883–891 10.1038/nature02661 [DOI] [PubMed] [Google Scholar]
- 130.Song L., Pan S., Zhang Z., Jia L., Chen W.H. and Zhao X.M. (2021) STAB: a spatio-temporal cell atlas of the human brain. Nucleic Acids Res. 49, D1029–D1037 10.1093/nar/gkaa762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Tabula Muris C. (2020) A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 10.1038/s41586-020-2496-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Ximerakis M., Lipnick S.L., Innes B.T., Simmons S.K., Adiconis X., Dionne D.et al. (2019) Single-cell transcriptomic profiling of the aging mouse brain. Nat. Neurosci. 22, 1696–1708 10.1038/s41593-019-0491-3 [DOI] [PubMed] [Google Scholar]
- 133.Nowakowski T.J., Bhaduri A., Pollen A.A., Alvarado B., Mostajo-Radji M.A., Di Lullo E.et al. (2017) Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 10.1126/science.aap8809 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Andrews B., Murphy A.E., Stofella M., Maslen S., Almeida-Souza L., Skehel J.M.et al. (2021) Multidimensional dynamics of the proteome in the neurodegenerative and aging mammalian brain. Mol. Cell. Proteom. 21, 100192 10.1016/j.mcpro.2021.100192 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Carlyle B.C., Kandigian S.E., Kreuzer J., Das S., Trombetta B.A., Kuo Y.et al. (2021) Synaptic proteins associated with cognitive performance and neuropathology in older humans revealed by multiplexed fractionated proteomics. Neurobiol. Aging 105, 99–114 10.1016/j.neurobiolaging.2021.04.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Graham L.C., Naldrett M.J., Kohama S.G., Smith C., Lamont D.J., McColl B.W.et al. (2019) Regional molecular mapping of primate synapses during normal healthy aging. Cell Rep. 27, 1018–+ 10.1016/j.celrep.2019.03.096 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Fitzner D., Bader J.M., Penkert H., Bergner C.G., Su M., Weil M.T.et al. (2020) Cell-type- and brain-region-resolved mouse brain lipidome. Cell Rep. 32, 108132 10.1016/j.celrep.2020.108132 [DOI] [PubMed] [Google Scholar]
- 138.Zocher S., Overall R.W., Lesche M., Dahl A. and Kempermann G. (2021) Environmental enrichment preserves a young DNA methylation landscape in the aged mouse hippocampus. Nat. Commun. 12, 3892 10.1038/s41467-021-23993-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Ding J., Ji J., Rabow Z., Shen T., Folz J., Brydges C.R.et al. (2021) A metabolome atlas of the aging mouse brain. Nat. Commun. 12, 6021 10.1038/s41467-021-26310-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Ben-Zvi A., Miller E.A. and Morimoto R.I. (2009) Collapse of proteostasis represents an early molecular event in Caenorhabditis elegans aging. Proc. Natl. Acad. Sci. U. S. A. 106, 14914–14919 10.1073/pnas.0902882106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Walther D.M., Kasturi P., Zheng M., Pinkert S., Vecchi G., Ciryam P.et al. (2015) Widespread Proteome remodeling and aggregation in aging C. Elegans. Cell 161, 919–932 10.1016/j.cell.2015.03.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Yang F.Y., Chu X.L., Yin M.M., Liu X.L., Yuan H.R., Niu Y.M.et al. (2014) mTOR and autophagy in normal brain aging and caloric restriction ameliorating age-related cognition deficits. Behav. Brain Res. 264, 82–90 10.1016/j.bbr.2014.02.005 [DOI] [PubMed] [Google Scholar]
- 143.Feleciano D.R., Juenemann K., Iburg M., Bras I.C., Holmberg C.I. and Kirstein J. (2019) Crosstalk between chaperone-mediated protein disaggregation and proteolytic pathways in aging and disease. Front. Aging Neurosci. 11, 10.3389/fnagi.2019.00009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Lipinski M.M., Zheng B., Lu T., Yan Z.Y., Py B.F., Ng A.et al. (2010) Genome-wide analysis reveals mechanisms modulating autophagy in normal brain aging and in Alzheimer's disease. Proc. Natl. Acad. Sci. U. S. A. 107, 14164–14169 10.1073/pnas.1009485107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Fernández Á F., Sebti S., Wei Y., Zou Z., Shi M., McMillan K.L.et al. (2018) Disruption of the beclin 1-BCL2 autophagy regulatory complex promotes longevity in mice. Nature 558, 136–140 10.1038/s41586-018-0162-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Aman Y., Schmauck-Medina T., Hansen M., Morimoto R.I., Simon A.K., Bjedov I.et al. (2021) Autophagy in healthy aging and disease. Nat. Aging 1, 634–650 10.1038/s43587-021-00098-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Glatigny M., Moriceau S., Rivagorda M., Ramos-Brossier M., Nascimbeni A.C., Lante F.et al. (2019) Autophagy is required for memory formation and reverses age-related memory decline. Curr. Biol. 29, 435.e8–448.e8 10.1016/j.cub.2018.12.021 [DOI] [PubMed] [Google Scholar]
- 148.Pandey K., Yu X.W., Steinmetz A. and Alberini C.M. (2021) Autophagy coupled to translation is required for long-term memory. Autophagy 17, 1614–1635 10.1080/15548627.2020.1775393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Gupta V.K., Scheunemann L., Eisenberg T., Mertel S., Bhukel A., Koemans T.S.et al. (2013) Restoring polyamines protects from age-induced memory impairment in an autophagy-dependent manner. Nat. Neurosci. 16, 1453–+ 10.1038/nn.3512 [DOI] [PubMed] [Google Scholar]
- 150.Stavoe A.K., Gopal P.P., Gubas A., Tooze S.A. and Holzbaur E.L. (2019) Expression of WIPI2B counteracts age-related decline in autophagosome biogenesis in neurons. Elife 8, 10.7554/eLife.44219 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Kiffin R., Kaushik S., Zeng M., Bandyopadhyay U., Zhang C., Massey A.C.et al. (2007) Altered dynamics of the lysosomal receptor for chaperone-mediated autophagy with age. J. Cell Sci. 120, 782–791 10.1242/jcs.001073 [DOI] [PubMed] [Google Scholar]
- 152.Xilouri M., Brekk O.R., Polissidis A., Chrysanthou-Piterou M., Kloukina I. and Stefanis L. (2016) Impairment of chaperone-mediated autophagy induces dopaminergic neurodegeneration in rats. Autophagy 12, 2230–2247 10.1080/15548627.2016.1214777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Xilouri M., Brekk O.R., Landeck N., Pitychoutis P.M., Papasilekas T., Papadopoulou-Daifoti Z.et al. (2013) Boosting chaperone-mediated autophagy in vivo mitigates alpha-synuclein-induced neurodegeneration. Brain 136, 2130–2146 10.1093/brain/awt131 [DOI] [PubMed] [Google Scholar]
- 154.Hu Y.B., Dammer E.B., Ren R.J. and Wang G. (2015) The endosomal-lysosomal system: from acidification and cargo sorting to neurodegeneration. Transl. Neurodegener 4, 18 10.1186/s40035-015-0041-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Lin H., Tang M., Ji C., Girardi P., Cvetojevic G., Chen D.et al. (2021) BAG3 regulation of RAB35 mediates the endosomal sorting complexes required for transport/endolysosome pathway and tau clearance. Biol. Psychiatry 92, 10–24 10.1016/j.biopsych.2021.10.024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Zhou J., Chow H.M., Liu Y., Wu D., Shi M., Li J.et al. (2020) Cyclin-dependent kinase 5-dependent BAG3 degradation modulates synaptic protein turnover. Biol. Psychiatry 87, 756–769 10.1016/j.biopsych.2019.11.013 [DOI] [PubMed] [Google Scholar]
- 157.Tang M., Ji C., Pallo S., Rahman I. and Johnson G.V.W. (2018) Nrf2 mediates the expression of BAG3 and autophagy cargo adaptor proteins and tau clearance in an age-dependent manner. Neurobiol. Aging 63, 128–139 10.1016/j.neurobiolaging.2017.12.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Abubakar Y.S., Zheng W., Olsson S. and Zhou J. (2017) Updated insight into the physiological and pathological roles of the retromer complex. Int. J. Mol. Sci. 18, 10.3390/ijms18081601 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Choy R.W.Y., Park M., Temkin P., Herring B.E., Marley A., Nicoll R.A.et al. (2014) Retromer mediates a discrete route of local membrane delivery to dendrites. Neuron 82, 55–62 10.1016/j.neuron.2014.02.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Mikhaylova M., Bera S., Kobler O., Frischknecht R. and Kreutz M.R. (2016) A dendritic golgi satellite between ERGIC and retromer. Cell Reports 14, 189–199 10.1016/j.celrep.2015.12.024 [DOI] [PubMed] [Google Scholar]
- 161.Wu S.J., Fagan R.R., Uttamapinant C., Lifshitz L.M., Fogarty K.E., Ting A.Y.et al. (2017) The dopamine transporter recycles via a retromer-dependent postendocytic mechanism: tracking studies using a novel fluorophore-coupling approach. J. Neurosci. 37, 9438–9452 10.1523/JNEUROSCI.3885-16.2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Chu J. and Pratico D. (2017) The retromer complex system in a transgenic mouse model of AD: influence of age. Neurobiol. Aging 52, 32–38 10.1016/j.neurobiolaging.2016.12.025 [DOI] [PubMed] [Google Scholar]
- 163.Vagnozzi A.N. and Pratico D. (2019) Endosomal sorting and trafficking, the retromer complex and neurodegeneration. Mol. Psychiatr. 24, 857–868 10.1038/s41380-018-0221-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Inoshita T., Arano T., Hosaka Y., Meng H.R., Umezaki Y., Kosugi S.et al. (2017) Vps35 in cooperation with LRRK2 regulates synaptic vesicle endocytosis through the endosomal pathway in Drosophila. Hum. Mol. Genet. 26, 2933–2948 10.1093/hmg/ddx179 [DOI] [PubMed] [Google Scholar]
- 165.Ye H., Ojelade S.A., Li-Kroeger D., Zuo Z.Y., Wang L.P., Li Y.R.et al. (2020) Retromer subunit, VPS29, regulates synaptic transmission and is required for endolysosomal function in the aging brain. Elife 9, e51977 10.7554/eLife.51977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Vazquez-Sanchez S., Bobeldijk S., Dekker M.P., van Keimpema L. and van Weering J.R.T. (2018) VPS35 depletion does not impair presynaptic structure and function. Sci. Rep. 8, 2996 10.1038/s41598-018-20448-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Fletcher B.R., Hill G.S., Long J.M., Gallagher M., Shapiro M.L. and Rapp P.R. (2014) A fine balance: Regulation of hippocampal Arc/Arg3.1 transcription, translation and degradation in a rat model of normal cognitive aging. Neurobiol. Learn. Mem. 115, 58–67 10.1016/j.nlm.2014.08.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Koyuncu S., Loureiro R., Lee H.J., Wagle P., Krueger M. and Vilchez D. (2021) Rewiring of the ubiquitinated proteome determines ageing in C. elegans. Nature 596, 285–290 10.1038/s41586-021-03781-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Reddy Addi U., Jakhotia S., Reddy S.S. and Reddy G.B. (2022) Age-related neuronal damage by advanced glycation end products through altered proteostasis. Chem. Biol. Interact. 355, 109840 10.1016/j.cbi.2022.109840 [DOI] [PubMed] [Google Scholar]
- 170.Yin P., Tu Z., Yin A., Zhao T., Yan S., Guo X.et al. (2015) Aged monkey brains reveal the role of ubiquitin-conjugating enzyme UBE2N in the synaptosomal accumulation of mutant huntingtin. Hum. Mol. Genet. 24, 1350–1362 10.1093/hmg/ddu544 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Giannini C., Kloss A., Gohlke S., Mishto M., Nicholson T.P., Sheppard P.W.et al. (2013) Poly-Ub-substrate-degradative activity of 26S proteasome is not impaired in the aging rat brain. PloS ONE 8, e64042 10.1371/journal.pone.0064042 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Sacramento E.K., Kirkpatrick J.M., Mazzetto M., Baumgart M., Bartolome A., Di Sanzo S.et al. (2020) Reduced proteasome activity in the aging brain results in ribosome stoichiometry loss and aggregation. Mol. Syst. Biol. 16, e9596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Thibaudeau T.A., Anderson R.T. and Smith D.M. (2018) A common mechanism of proteasome impairment by neurodegenerative disease-associated oligomers. Nat. Commun. 9, 1097 10.1038/s41467-018-03509-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Munkácsy E., Chocron E.S., Quintanilla L., Gendron C.M., Pletcher S.D. and Pickering A.M. (2019) Neuronal-specific proteasome augmentation via Prosβ5 overexpression extends lifespan and reduces age-related cognitive decline. Aging Cell. 18, e13005 10.1111/acel.13005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Grant S.G.N. and Fransen E. (2020) The synapse diversity dilemma: molecular heterogeneity confounds studies of synapse function. Front. Synaptic Neurosci. 12, 590403 10.3389/fnsyn.2020.590403 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Mattson M.P. and Magnus T. (2006) Ageing and neuronal vulnerability. Nat. Rev. Neurosci. 7, 278–294 10.1038/nrn1886 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Keller J.N., Hanni K.B. and Markesbery W.R. (2000) Possible involvement of proteasome inhibition in aging: implications for oxidative stress. Mech. Ageing Dev. 113, 61–70 10.1016/S0047-6374(99)00101-3 [DOI] [PubMed] [Google Scholar]
- 178.Pluvinage J.V. and Wyss-Coray T. (2020) Systemic factors as mediators of brain homeostasis, ageing and neurodegeneration. Nat. Rev. Neurosci. 21, 93–102 10.1038/s41583-019-0255-9 [DOI] [PubMed] [Google Scholar]
- 179.Escobar K.A., Cole N.H., Mermier C.M. and VanDusseldorp T.A. (2019) Autophagy and aging: Maintaining the proteome through exercise and caloric restriction. Aging Cell. 18, 10.1111/acel.12876 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
No data were generated for this review.