Abstract
In spite of the variety of initial reactions, the aerobic biodegradation of aromatic compounds generally yields dihydroxy intermediates for ring cleavage. Recent investigation of the degradation of nitroaromatic compounds revealed that some nitroaromatic compounds are initially converted to 2-aminophenol rather than dihydroxy intermediates by a number of microorganisms. The complete pathway for the metabolism of 2-aminophenol during the degradation of nitrobenzene by Pseudomonas pseudoalcaligenes JS45 has been elucidated previously. The pathway is parallel to the catechol extradiol ring cleavage pathway, except that 2-aminophenol is the ring cleavage substrate. Here we report the elucidation of the pathway of 2-amino-4-methylphenol (6-amino-m-cresol) metabolism during the degradation of 4-nitrotoluene by Mycobacterium strain HL 4-NT-1 and the comparison of the substrate specificities of the relevant enzymes in strains JS45 and HL 4-NT-1. The results indicate that the 2-aminophenol ring cleavage pathway in strain JS45 is not unique but is representative of the pathways of metabolism of other o-aminophenolic compounds.
Nitroaromatic compounds are important industrial feedstocks due to the versatile chemistry of the nitro group (22). Many nitroaromatic compounds are harmful, and their release into the environment has caused concern. The microbial degradation of nitroaromatic compounds is usually initiated by an enzymatic attack on the nitro group (22). One of the strategies involves partial reduction of the nitro group to a hydroxylamino group. The hydroxylamino intermediates can be transformed to catechols which enter the ring cleavage pathways of aerobic degradation of aromatic compounds, as observed during degradation of 4-nitrobenzoate (6, 33), 3-nitrophenol (19), and 4-nitrotoluene (7, 26). Alternatively, the hydroxylamino intermediates can be rearranged, by an intramolecular transfer of the hydroxyl group (10; L. J. Nadeau, Z. He, and J. C. Spain, unpublished data), to ortho-aminophenols, as observed during biodegradation of nitrobenzene (21, 24), 3- or 4-chloronitrobenzene (2, 16, 24), 3-nitrophenol (28), 2-chloro-5-nitrophenol (29), 4-nitrotoluene (30), and 2,4,6-trinitrotoluene (15) by various microorganisms. The initial steps of each of the degradation pathways have been elucidated; however, the lower pathway has been determined only for the metabolism of 2-aminophenol during the degradation of nitrobenzene by Pseudomonas pseudoalcaligenes JS45 (11, 14). The pathway is parallel to the catechol extradiol ring cleavage pathway, except that 2-aminophenol is the ring cleavage substrate.
Mycobacterium strain HL 4-NT-1 is able to grow on 4-nitrotoluene as the sole source of nitrogen, carbon, and energy (30). 4-Nitrotoluene is converted to 2-amino-4-methylphenol (6-amino-m-cresol) via 4-hydroxylaminotoluene in reactions catalyzed by a nitroreductase and an aminohydroxymutase. In reactions analogous to those used by strain JS45, extracts from induced cells of strain HL 4-NT-1 catalyzed the conversion of 2-amino-4-methylphenol to a transient yellow intermediate (A385) which was subsequently converted into another compound (A270) identified as 5-methylpicolinic acid (30). When NAD+ was included in the reaction mixture, 5-methylpicolinic acid accumulated to a lesser extent and ammonia was released (30). Based on the preliminary observations and by analogy with the system in strain JS45, the authors tentatively proposed that 2-amino-5-methylmuconic semialdehyde and 2-amino-5-methylmuconic acid were produced from 2-amino-4-methylphenol (30). The goal of the present work was to rigorously determine the steps in the lower pathway including the mechanism of release of ammonia in Mycobacterium strain HL 4-NT-1 grown on 4-nitrotoluene. We also compared the substrate specificities of the relevant enzymes in strains JS45 and HL 4-NT-1 to provide a preliminary evaluation of the relationship between the two pathways.
MATERIALS AND METHODS
Bacterial strains and growth conditions.
Mycobacterium strain HL 4-NT-1 was grown on nitrogen-free mineral medium with succinate (10 mM) and 4-nitrotoluene (0.5 mM) as described previously (30). Fully induced cells of strain HL 4-NT-1 were obtained by incubation of the harvested cells in fresh medium with 4-nitrotoluene as the sole carbon source overnight. Uninduced cells were obtained by substitution of NH4Cl for 4-nitrotoluene during growth. Cells were harvested by centrifugation and then broken using a French press as described previously (30).
Enzyme assays.
The activity of the ring cleavage dioxygenase (0.1 mg of protein/ml) in crude extracts was determined spectrophotometrically by monitoring the rate of the disappearance of the substrates (0.05 mM) in potassium phosphate buffer (50 mM, pH 8.0). The semialdehyde dehydrogenase activity (0.084 mg of protein/ml) was assayed by measurement of the decrease of A375 (ɛ = 44,000 M−1) using 2-hydroxymuconic semialdehyde (0.038 to 0.041 mM) as the substrate in the presence of 0.2 mM NAD+ in potassium phosphate buffer (50 mM, pH 8.0) (11). Dehydrogenation of amino-substituted semialdehydes was assayed as reported previously by estimation of the increase in the A326 which was contributed by the products of dehydrogenation (9). The deaminase activity was determined by monitoring the decrease of A326 of a reaction mixture containing enzyme preparations (0.016 to 0.063 mg of protein/ml), substrate (2-aminomuconate, 0.088 mM, ɛ = 17,800 M−1 [13], or 2-amino-5-methylmuconate, 0.098 mM, ɛ = 15,400 M−1 [this work]), and 50 mM Tris-HCl buffer, pH 7.5, as described previously (12, 13). 4-Oxalocrotonate decarboxylase activity was determined by the initial rate of decrease in A236 caused by the disappearance of 2-oxo-3-hexene-1,6-dioate (4-oxalocrotonate, 0.06 to 0.07 mM, ɛ = 7,200 M−1) in the presence of 1 mM MgSO4 and enzyme (0.035 mg of protein/ml) (8). Protein was measured by the Coomassie Plus protein assay reagent from Pierce (Rockford, Ill.) by using bovine serum albumin as a standard.
Ion-exchange chromatography.
A Hitrap Q column (5 ml; Pharmacia) was equilibrated with potassium phosphate buffer (10 mM, pH 7.0), loaded with 12.5 ml of crude extract from strain HL 4-NT-1, and washed with 25 ml of the buffer. Proteins were eluted with a 48-ml linear gradient from 0 to 0.5 M NaCl in buffer at a flow rate of 0.8 ml · min−1. Fractions (2 ml) were collected and assayed for enzyme activities and protein concentration.
Preparation of 2-amino-5-methylmuconate.
The muconate was prepared from 4-methyl-2-aminophenol by the combined action of 2-aminophenol 1,6-dioxygenase of strain JS45 (5) and the partially purified 2-amino-5-methylmuconic semialdehyde dehydrogenase obtained by the ion-exchange chromatography described above. The molar extinction coefficient of 2-amino-5-methylmuconate at 326 nm was determined after the reaction went to completion in the presence of excess dehydrogenase as indicated by the absence of accumulation of ring cleavage intermediates (A382). The accumulation of NADH was eliminated by the addition of pyruvate (0.5 mM) and lactate dehydrogenase (5 U per ml). The product was purified by ion-exchange chromatography as described previously for 2-aminomuconate (13).
Biochemical reagents and chemicals.
Purified 2-aminomuconate semialdehyde dehydrogenase and 2-aminomuconate deaminase from strain JS45 were obtained as described previously (9, 12). 2-Aminophenol 1,6-dioxygenase was obtained from an Escherichia coli clone containing the structural genes encoding the enzyme from strain JS45 (5). The 4-oxalocrotonate decarboxylase from strain JS45 was partially purified as described previously (11). 2-Hydroxy-5-methylmuconic semialdehyde was prepared from 4-methylcatechol by the catechol 2,3-dioxygenase in an E. coli clone containing the appropriate gene from Comamonas sp. strain JS765 (23). 2-Aminomuconate was enzymatically synthesized as previously described (13). 2-Hydroxymuconic semialdehyde and 2-oxo-3-hexene-1,6-dioate (4-oxalocrotonate) were prepared as reported previously (9, 14). All other chemicals were from Sigma (St. Louis, Mo.) or Aldrich (Milwaukee, Wis.).
RESULTS
Ring cleavage dioxygenase and semialdehyde dehydrogenase activities in crude extracts.
As observed previously (30), the crude extracts of strain HL 4-NT-1 catalyzed the oxidation of 2-amino-4-methylphenol and 2-aminophenol, but not 4-methylcatechol (Table 1). In contrast to the previous report (30), we found that catechol was also a substrate of the dioxygenase, but the activity ceased after a half-minute, indicating that catechol was a suicide substrate of the 2-amino-4-methylphenol 1,6-dioxygenase, as it is for the 2-aminophenol 1,6-dioxygenase from strain JS45 (5).
TABLE 1.
Comparison of the relevant enzyme activities in Mycobacterium strain HL 4-NT-1 and P. pseudoalcaligenes JS45
| Enzyme and substrate | Relative activity (sp act [μmol min−1 mg of protein−1])
|
|
|---|---|---|
| HL 4-NT-1a | JS45b | |
| Dioxygenase | ||
| 2-Aminophenol | 17 | 100 (21.6)c |
| 2-Amino-4-methylphenol | 100 (0.13) | 25c |
| Catechold | 23 | 13c |
| 4-Methylcatechol | NDe | NDc |
| Dehydrogenase | ||
| 2-Aminomuconic semialdehyde | 34 | 311 |
| 2-Amino-5-methylmuconic semialdehyde | 112 | 9 |
| 2-Hydroxymuconic semialdehyde | 100 (0.17) | 100 (4.3) |
| 2-Hydroxy-5-methylmuconic semialdehyde | 111 | 32 |
| Deaminase | ||
| 2-Aminomuconate | 100 (1.5) | 100 (37.8) |
| 2-Amino-5-methylmuconate | 10 | 1.2 |
| Decarboxylase | ||
| 2-Oxo-3-hexene-1,6-dioate | 100 (0.41) | 100 (1.7) |
| 5-Methyl-2-oxo-3-hexene-1,6-dioate | 2.5 | 9.5 |
Data were obtained with partially purified enzymes, except those for the dioxygenase activity, which were obtained with crude extracts.
Data were obtained with purified dioxygenase, dehydrogenase, and deaminase. Data for the decarboxylase activity were obtained with partially purified enzyme.
Data are from reference 17.
Catechol was a suicide substrate.
ND, not detected.
The A375 of 2-hydroxymuconic semialdehyde decreased when the crude extracts of strain HL 4-NT-1 were incubated with the substrate in the presence of NAD+, but no decrease in the absorbance was observed in the absence of NAD+. The observation indicated that strain HL 4-NT-1 contains a semialdehyde dehydrogenase but not a semialdehyde hydrolase. The result suggested that 2-amino-5-methylmuconic semialdehyde, the ring fission product from 2-amino-4-methylphenol, is dehydrogenated to 2-amino-5-methylmuconate, as proposed previously (30), and by the same mechanism as in the metabolism of 2-aminophenol in strain JS45 (14). Incubation of 2-amino-4-methylphenol with the crude extracts in the presence of NAD+, however, did not lead to a spectral change characteristic of the proposed 2-amino-5-methylmuconate, probably due to a high activity of the subsequent deaminase enzyme.
Partial purification of 2-amino-5-methylmuconic semialdehyde dehydrogenase and production of 2-amino-5-methylmuconate.
To investigate the individual enzymes, the cell extracts of strain HL 4-NT-1 were loaded on a Hitrap-Q anion-exchange column and partially purified enzymes were obtained as described in Materials and Methods. The 2-amino-4-methylphenol 1,6-dioxygenase was not detected spectrophotometrically in any fractions, probably due to the lability of the dioxygenase. 2-Hydroxymuconate semialdehyde was used for routine assay of the activity of the semialdehyde dehydrogenase because 2-amino-5-methylmuconic semialdehyde is unstable and spontaneously converts to 5-methylpicolinate (17, 30). The dehydrogenase activity eluted at 0.35 M NaCl (Fig. 1).
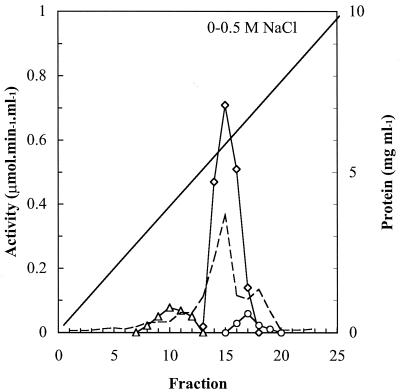
FIG. 1.
Hitrap-Q chromatography of crude extracts of strain HL 4-NT-1. Diagonal line, NaCl elution gradient from 0 to 0.4 M; dotted line, protein concentration; ○, semialdehyde dehydrogenase; ▵, 2-amino-5-methylmuconate deaminase; ◊, 4-oxalocrotonate decarboxylase.
The activity of the semialdehyde dehydrogenase from strain HL 4-NT-1 was optimal at pH 8, whereas 2-aminomuconic semialdehyde dehydrogenase from strain JS45 had an optimal activity at pH 7.3 (9). Both enzymes were able to act on the hydroxy analogs (Table 1). The 2-aminophenol 1,6-dioxygenase (5) was used to produce amino-substituted semialdehydes. The incubation of 2-amino-4-methylphenol with the combination of the partially purified semialdehyde dehydrogenase and the 2-aminophenol 1,6-dioxygenase produced a previously unreported metabolite with an absorbance maximum at 326 nm (Fig. 2). When the purified 2-aminomuconic semialdehyde dehydrogenase (9) from strain JS45 was substituted for the dehydrogenase fraction from strain HL 4-NT-1, identical results were obtained. When the purified 2-aminomuconate deaminase (12) from strain JS45 acted on the product, stoichiometric release (90%) of ammonia was observed. Based on the catalytic mechanisms of the dioxygenase and dehydrogenase in strain JS45 (9, 14) and the stoichiometry of ammonia release, the product seems to be 2-amino-5-methylmuconate. The spectrum of 2-amino-5-methylmuconate was similar to that of 2-aminomuconate, but the molar extinction coefficient of the former was determined to be 15.4 ± 0.2 mM−1 cm−1 (average of three determinations), whereas that of the latter is 17.8 mM−1 cm−1 (13).
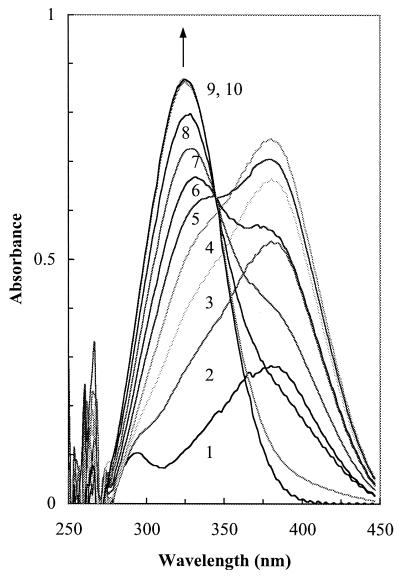
FIG. 2.
Enzymatic formation of 2-amino-5-methylmuconate by the action of partially purified 2-amino-5-methylmuconic semialdehyde dehydrogenase (fraction 17). 2-Amino-5-methylmuconic semialdehyde (A382) was instantly produced from 2-amino-4-methylphenol (0.05 mM) by the 2-aminophenol 1,6-dioxygenase of strain JS45 (0.31 mg of protein per ml). The reaction mixture also contained potassium phosphate (50 mM, pH 8.0), fraction 17 (0.084 mg of protein per ml), and 0.2 mM NAD+. Recordings were taken at 0.5-min intervals. The A326 increased during the course of the reaction.
Deamination of 2-amino-5-methylmuconate.
When 2-amino-5-methylmuconate was incubated with crude extracts of HL 4-NT-1, the A326 decreased with time. The enzyme activity eluted from the Hitrap-Q anion-exchange column at 0.21 M NaCl (Fig. 1). When 2-amino-5-methylmuconate (113 μmol in 1 ml) was incubated with the active fraction, stoichiometric release of ammonia (112 μmol) was observed. This result indicated that the enzyme was 5-methyl-2-aminomuconate deaminase. During the deamination of 2-amino-5-methylmuconate, a new absorbance maximum appeared at 297 nm along with an increase of absorbance at about 237 nm (Fig. 3A). The enzyme also catalyzed the deamination of 2-aminomuconate with a higher relative activity (Table 1). During deamination of 2-aminomuconate by the HL 4-NT-1 deaminase, the absorbance changed directly from 326 to 236 nm (the putative oxo isomer, 2-oxo-3-hexene-1,6-dioate) without a significant A297 (the putative enol isomer, 2-hydroxymuconate) (Fig. 3B). The spectral change was similar to that catalyzed by the purified 2-aminomuconate deaminase from strain JS45 (12). Control experiments indicated that the product from 5-methyl-2-aminomuconate in reactions catalyzed by purified JS45 deaminase also had an absorbance maximum at 297 nm (data not shown). The relative contributions of the enol and oxo isomers of 5-methyl-2-oxo-3-hexene-1,6-dioate to the absorbance spectra have not been determined rigorously. Based on analogy with previous work, the reaction catalyzed by the HL 4-NT-1 deaminase seems to be the same as that catalyzed by the JS45 deaminase. That is, the product of deamination of 5-methyl-2-aminomuconate seems to be the oxo isomer 5-methyl-2-oxo-3-hexene-1,6-dioate, not the enol isomer 5-methyl-2-hydroxymuconate.
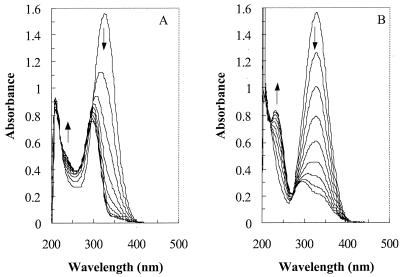
FIG. 3.
Spectral changes during metabolism of 2-amino-5-methylmuconate (A) and 2-aminomuconate (B). Buffer was Tris-HCl (50 mM, pH 7.5). The amount of partially purified deaminase (fraction 10) from strain HL 4-NT-1 in the reaction mixture (1 ml) for panels A and B was 0.063 and 0.016 mg of protein, respectively. The intervals between scans are 4 min for panel A and 2 min for panel B.
The decarboxylase-hydratase fraction.
The protein fraction that eluted at 0.31 M NaCl from the Hitrap-Q anion-exchange column catalyzed the decarboxylation of 2-oxo-3-hexene-1,6-dioate in the presence of MgSO4, as monitored by the decrease of the A237 (8). There was no activity in the absence of MgSO4. During the decarboxylation, a transient absorbance peak at 265 nm was observed (Fig. 4), indicating that the hydratase coeluted with the decarboxylase, a common observation for the two enzymes (8, 11, 18). It was not possible to determine accurately the specific activity of decarboxylation of 5-methyl-2-oxohexene-1,6-dioate because no authentic substrate was available. The decarboxylases from both bacteria showed less than 10% of the relative activity on the 5-methyl analog (Table 1). The existence of the decarboxylase and hydratase in strain HL 4-NT-1 suggested that the later steps in the pathway of the metabolism of 4-methyl-2-aminophenol were similar to the corresponding steps in catechol extradiol pathways.
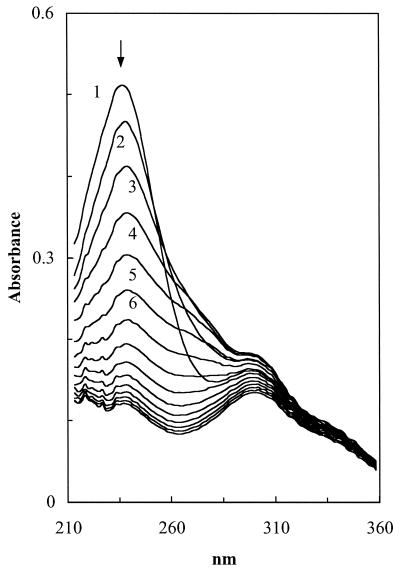
FIG. 4.
Spectral changes during metabolism of 2-oxo-3-hexene-1,6-dioate. The substrate (A236) was preequilibrated with its enol form (2-hydroxymuconate, A296) in Tris-HCl buffer (50 mM, pH 7.0). Fraction 14 was used for the source of the decarboxylase (0.035 mg of protein per ml). Reactions were initiated by the addition of MgSO4 (1 mM final concentration). Recordings were taken at 0.5-min intervals. The transient increase of absorbance around 265 nm indicates that the 2-oxo-4-pentenoate hydratase coexisted in the fraction.
DISCUSSION
Microorganisms exhibit extensive metabolic diversity that enables them to degrade a variety of aromatic compounds. In general, pathways for the aerobic degradation of aromatic compounds can be considered to consist of two parts: an upper pathway and a lower pathway (25, 32). Since the catechol extradiol pathway (Fig. 5) was reported for the first time in the 1960s (3, 4, 27), it has been found to be one of the most common lower pathways involved in degradation of both naturally occurring and man-made aromatic compounds (1, 22, 25, 32). The ortho-aminophenol intermediates produced during degradation of some nitroaromatic compounds do not have the two adjacent hydroxyl groups on the aromatic ring that were previously thought to be necessary for metabolism through the extradiol ring cleavage pathway. The pathway for the metabolism of 2-aminophenol during the degradation of nitrobenzene by strain JS45 is parallel to the catechol extradiol pathway (Fig. 5) (11, 14). Results presented here suggest that the pathway is not confined to the degradation of nitrobenzene and might be common among bacteria that degrade nitroaromatic compounds or the corresponding aminophenols.
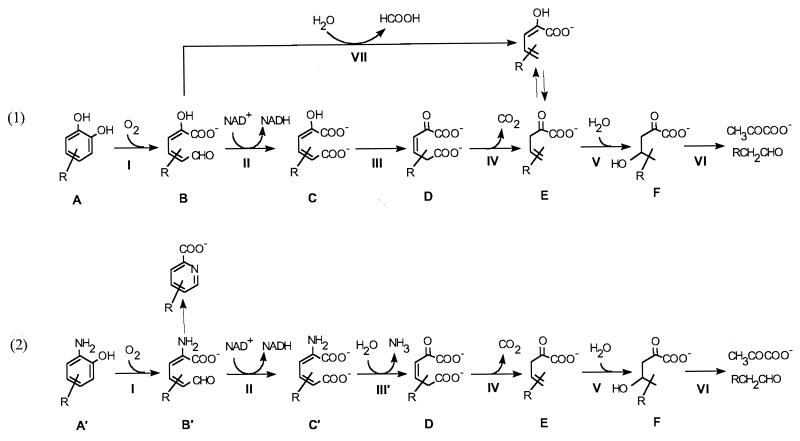
FIG. 5.
General catechol extradiol ring cleavage pathway (pathway 1) and general o-aminophenol extradiol-like ring cleavage pathway (pathway 2). In pathway 1, R can be hydrogen or a methyl, hydroxyl, carboxyl, chloro, or other group; in pathway 2, R has been shown to be a hydrogen (11) and methyl group (this work). I, ring cleavage dioxygenase; II, aldehyde dehydrogenase; III, tautomerase; III′, deaminase; IV, decarboxylase; V, hydratase; VI, aldolase; VII, hydrolase. Vertical arrows represent spontaneous conversion. When A is (4-methyl)catechol, B is 2-hydroxy-(5-methyl)muconic semialdehyde, C is 2-hydroxy-(5-methyl)muconate, D is (5-methyl)-2-oxo-3-hexene-1,6-dioate, E is 2-oxo-4-pen(or hex)enoate, and F is 2-oxo-4-hydroxyvalerate(or hexanoate). When A′ is 2-amino-(4-methyl)phenol, B′ and C′ are the corresponding amino analogs of B and C.
Our results indicate that both 2-aminophenol and 2-amino-4-methylphenol are degraded through the o-aminophenol cleavage pathways in either strain JS45 or HL 4-NT-1 even though the specific activities of the enzymes were not the same in the two strains. Catechol 2,3-dioxygenase similarly catalyzes cleavage of 4-methylcatechol (1, 3). ortho-Aminophenolic compounds can be considered the analogs of catechols in which one of the hydroxyl groups is replaced by an amino group. The amino and hydroxyl groups are comparable in size and electronegativity, and both are electron donating by resonance effects. The amino group is basic, and the hydroxyl group shows similar but weaker basicity (20). However, 2-aminophenol is a very poor substrate for catechol 2,3-dioxygenase; it is oxidized at only about 0.01% of the rate of catechol (5). On the other hand, catechol was a suicide substrate of the two o-aminophenol 1,6-dioxygenases from strains JS45 and HL 4-NT-1. Therefore, considering the fact that there is only a single hydroxyl group in o-aminophenols, it seems more appropriate to categorize the o-aminophenol dioxygenases as extradiol-like ring cleavage dioxygenases. Because the two pathways are similar but not identical, it is better to name the o-aminophenol ring cleavage route an extradiol-like pathway.
In recent years, nitrobenzene, 3-chloronitrobenzene, 4-chloronitrobenzene, 3-nitrophenol, 2-chloro-5-nitrophenol, 4-nitrotoluene, and probably 2,4,6-trinitrotoluene have been reported to be transformed to the corresponding o-aminophenols via mutase-catalyzed rearrangements in various microorganisms (22). The proliferation of reports suggests that o-aminophenolic compounds may be central intermediates in the biodegradation of a variety of nitroaromatic compounds. It will be useful to elucidate the pathways used during the degradation of other nitroaromatic compounds via substituted o-aminophenol intermediates. As the catechol pathway is the archetype of extradiol cleavage pathways for degradation of various aromatic compounds, the 2-aminophenol pathway might serve as an archetypal extradiol-like ring cleavage pathway for degradation of relevant nitroaromatic compounds. Biochemical characterization and sequence analysis of 2-aminophenol 1,6-dioxygenases from strain JS45 and another bacterium, Pseudomonas sp. strain AP-3, show that the enzymes share some features with extradiol dioxygenases, but the amino acid sequences are only distantly related (5, 17, 31). 2-Aminomuconic semialdehyde dehydrogenase from P. pseudoalcaligenes JS45 is most similar to its counterparts in the catechol pathway in both biochemical properties and amino acid sequences (9). However, the substrates for 2-aminomuconate deaminase and 4-oxalocrotonate tautomerase are not interchangeable, even though the two reactions seem superficially similar. The N-terminal amino acid sequence of the deaminase from JS45 does not match that of the tautomerase (12). Further comparative investigation of the enzymology and molecular biology of the two pathways will be required to explain the biochemistry of the relevant reactions and the evolution of the extradiol-like pathway.
ACKNOWLEDGMENTS
The work was supported in part by the U.S. Air Force Office of Scientific Research and the Strategic Environmental Defense Research Program. This work was also supported in part by an appointment to the research Participation Program at the U.S. Air Force Research Laboratory, Tyndall Air Force Base, administered by the Oak Ridge Institute for Science and Education through an interagency agreement between the U.S. Department of Energy and Tyndall Air Force Base.
We thank Hiltrud Lenke for providing strain HL 4-NT-1, Becky Parales for providing the E. coli clone which contains the catechol 2,3-dioxygenase gene from JS765, and Shirley Nishino for reviewing the manuscript.
REFERENCES
- 1.Bayly R C, Barbour M G. The degradation of aromatic compounds by the meta and gentisate pathways. In: Gibson D T, editor. Microbial degradation of organic compounds. New York, N.Y: Marcel Dekker; 1984. pp. 252–294. [Google Scholar]
- 2.Corbett M D, Corbett B R. Metabolism of 4-chloronitrobenzene by the yeast Rhodosporidium sp. Appl Environ Microbiol. 1981;41:942–949. doi: 10.1128/aem.41.4.942-949.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dagley S, Chapman P J, Gibson D T, Wood J M. Degradation of the benzene nucleus by bacteria. Nature. 1964;202:775–778. doi: 10.1038/202775a0. [DOI] [PubMed] [Google Scholar]
- 4.Dagley S, Evans W C, Ribbons D W. New pathways in the oxidative metabolism of aromatic compounds by micro-organisms. Nature. 1960;188:560–566. doi: 10.1038/188560a0. [DOI] [PubMed] [Google Scholar]
- 5.Davis J K, He Z, Somerville C C, Spain J C. Genetic and biochemical comparison of 2-aminophenol 1,6-dioxygenase of Pseudomonas pseudoalcaligenes JS45 to meta cleavage dioxygenases: divergent evolution of 2-aminophenol meta cleavage pathway. Arch Microbiol. 1999;172:330–339. doi: 10.1007/s002030050787. [DOI] [PubMed] [Google Scholar]
- 6.Groenewegen P E J, Breeuwer P, van Helvoort J M L M, Langenhoff A A M, de Vries F P, de Bont J A M. Novel degradative pathway of 4-nitrobenzoate in Comamonas acidovorans NBA-10. J Gen Microbiol. 1992;138:1599–1605. doi: 10.1099/00221287-138-8-1599. [DOI] [PubMed] [Google Scholar]
- 7.Haigler B E, Spain J C. Biodegradation of 4-nitrotoluene by Pseudomonas sp. strain 4NT. Appl Environ Microbiol. 1993;59:2239–2243. doi: 10.1128/aem.59.7.2239-2243.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Harayama S, Rekik M, Ngai K-L, Ornston L N. Physically associated enzymes produce and metabolize 2-hydroxy-2,4-dienoate, a chemically unstable intermediate formed in catechol metabolism via meta cleavage in Pseudomonas putida. J Bacteriol. 1989;171:6251–6258. doi: 10.1128/jb.171.11.6251-6258.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.He Z, Davis J K, Spain J C. Purification, characterization, and sequence analysis of 2-aminomuconic 6-semialdehyde dehydrogenase from Pseudomonas pseudoalcaligenes JS45. J Bacteriol. 1998;180:4591–4595. doi: 10.1128/jb.180.17.4591-4595.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.He Z, Nadeau L J, Spain J C. Characterization of hydroxylaminobenzene mutase from pNBZ139 cloned from Pseudomonas pseudoalcaligenes JS45: a highly-associated sodium-dodecyl-sulfate-stable enzyme catalyzing an intramolecular transfer of hydroxyl group. Eur J Biochem. 2000;267:1110–1116. doi: 10.1046/j.1432-1327.2000.01107.x. [DOI] [PubMed] [Google Scholar]
- 11.He Z, Spain J C. Comparison of the downstream pathways for degradation of nitrobenzene by Pseudomonas pseudoalcaligenes JS45 (2-aminophenol pathway) and by Comamonas sp. JS765 (catechol pathway) Arch Microbiol. 1999;171:309–316. doi: 10.1007/s002030050715. [DOI] [PubMed] [Google Scholar]
- 12.He Z, Spain J C. A novel 2-aminomuconate deaminase in the nitrobenzene degradation pathway of Pseudomonas pseudoalcaligenes JS45. J Bacteriol. 1998;180:2502–2506. doi: 10.1128/jb.180.9.2502-2506.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.He Z, Spain J C. Preparation of 2-aminomuconate from 2-aminophenol by coupled enzymatic dioxygenation and dehydrogenation reactions. J Ind Microbiol Biotechnol. 1999;23:138–142. doi: 10.1038/sj.jim.2900705. [DOI] [PubMed] [Google Scholar]
- 14.He Z, Spain J C. Studies of the catabolic pathway of degradation of nitrobenzene by Pseudomonas pseudoalcaligenes JS45: removal of the amino group from 2-aminomuconic semialdehyde. Appl Environ Microbiol. 1997;63:4839–4843. doi: 10.1128/aem.63.12.4839-4843.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hughes J B, Wang C, Yesland K, Richardson A, Bhadra R, Bennett G, Ruldoph F. Bamberger rearrangement during TNT metabolism by Clostridium acetobutylicum. Environ Sci Technol. 1998;32:494–500. [Google Scholar]
- 16.Katsivela E, Wray V, Pieper D H, Wittich R. Initial reactions in the biodegradation of 1-chloro-4-nitrobenzene by a newly isolated bacterium, strain LW1. Appl Environ Microbiol. 1999;65:1405–1412. doi: 10.1128/aem.65.4.1405-1412.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lendenmann U, Spain J C. 2-Aminophenol 1,6-dioxygenase: a novel aromatic ring-cleavage enzyme purified from Pseudomonas pseudoalcaligenes JS45. J Bacteriol. 1996;178:6227–6232. doi: 10.1128/jb.178.21.6227-6232.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lian H, Whitman C P. Stereochemical and isotopic labelling studies of 4-oxalocrotonate decarboxylase and vinylpyruvate hydratase: analysis and mechanistic implications. J Am Chem Soc. 1994;116:10403–10411. [Google Scholar]
- 19.Meulenberg R, Pepi M, de Bont J A M. Degradation of 3-nitrophenol by Pseudomonas putida B2 occurs via 1,2,4-benzenetriol. Biodegradation. 1996;7:303–311. doi: 10.1007/BF00115744. [DOI] [PubMed] [Google Scholar]
- 20.Morrison R T, Boyd R T. Organic chemistry. 4th ed. Boston, Mass: Allyn and Bacon, Inc.; 1983. p. 616. [Google Scholar]
- 21.Nishino S F, Spain J C. Degradation of nitrobenzene by a Pseudomonas pseudoalcaligenes. Appl Environ Microbiol. 1993;59:2520–2525. doi: 10.1128/aem.59.8.2520-2525.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Nishino S F, Spain J C, He Z. Strategies for aerobic degradation of nitroaromatic compounds: process discovery to field application. In: Spain J C, Hughes J B, Knackmuss H-J, editors. Biodegradation of nitroaromatic compounds and explosives, in press. Boca Raton, Fla: CRC Press, Inc.; 2000. [Google Scholar]
- 23.Parales R E, Ontl T A, Gibson D T. Cloning and sequence analysis of a catechol 2,3-dioxygenase gene from the nitrobenzene-degrading strain Comamonas sp. JS765. J Ind Microbiol Biotechnol. 1997;19:385–391. doi: 10.1038/sj.jim.2900420. [DOI] [PubMed] [Google Scholar]
- 24.Park H, Lim S, Chang Y K, Livingston A G, Kim H. Degradation of chloronitrobenzenes by a coculture of Pseudomonas putida and a Rhodococcus sp. Appl Environ Microbiol. 1999;65:1083–1091. doi: 10.1128/aem.65.3.1083-1091.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Reineke W, Knackmuss H-J. Construction of haloaromatics utilising bacteria. Nature. 1979;277:385–386. doi: 10.1038/277385a0. [DOI] [PubMed] [Google Scholar]
- 26.Rhys-Williams W, Taylor S T, Williams P A. A novel pathway for the catabolism of 4-nitrotoluene by Pseudomonas. J Gen Microbiol. 1993;139:1967–1972. doi: 10.1099/00221287-139-9-1967. [DOI] [PubMed] [Google Scholar]
- 27.Sala-Trepat J M, Evans W C. The meta cleavage of catechol by Azotobacter species. Eur J Biochem. 1971;20:400–413. doi: 10.1111/j.1432-1033.1971.tb01406.x. [DOI] [PubMed] [Google Scholar]
- 28.Schenzle A, Lenke H, Fischer P, Williams P A, Knackmuss H-J. Catabolism of 3-nitrophenol by Ralstonia eutropha JMP 134. Appl Environ Microbiol. 1997;63:1421–1427. doi: 10.1128/aem.63.4.1421-1427.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Schenzle A, Lenke H, Spain J C, Knackmuss H-J. Chemoselective nitro group reduction and reductive dechlorination initiate degradation of 2-chloro-5-nitrophenol by Ralstonia eutropha JMP134. Appl Environ Microbiol. 1999;65:2317–2323. doi: 10.1128/aem.65.6.2317-2323.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Spiess T, Desiere F, Fischer P, Spain J C, Knackmuss H-J, Lenke H. A new 4-nitrotoluene degradation pathway in a Mycobacterium strain. Appl Environ Microbiol. 1998;64:446–452. doi: 10.1128/aem.64.2.446-452.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Takenaka S, Murakami S, Shinke R, Hatakeyama K, Yukawa H, Aoki K. Novel genes encoding 2-aminophenol 1,6-dioxygenase from Pseudomonas species AP-3 growing on 2-aminophenol and catalytic properties of the purified enzyme. J Biol Chem. 1997;272:14727–14732. doi: 10.1074/jbc.272.23.14727. [DOI] [PubMed] [Google Scholar]
- 32.Williams P A, Sayers J R. The evolution of pathways for aromatic hydrocarbon oxidation in Pseudomonas. Biodegradation. 1994;5:195–217. doi: 10.1007/BF00696460. [DOI] [PubMed] [Google Scholar]
- 33.Yabannavar A V, Zylstra G J. Cloning and characterization of the genes for p-nitrobenzoate degradation from Pseudomonas pickettii YH105. Appl Environ Microbiol. 1995;61:4282–4290. doi: 10.1128/aem.61.12.4284-4290.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]