Abstract
The development of natural competence by bacteria in situ is considered one of the main factors limiting transformation-mediated gene exchanges in the environment. Ralstonia solanacearum is a plant pathogen that is also a naturally transformable bacterium that can develop the competence state during infection of its host. We have attempted to determine whether this bacterium could become the recipient of plant genes. We initially demonstrated that plant DNA was released close to the infecting bacteria. We constructed and tested various combinations of transgenic plants and recipient bacteria to show that the effectiveness of such transfers was directly related to the ratio of the complexity of the plant genome to the number of copies of the transgene.
Nucleotide sequences in databases indicate that some genes in genomes of soil bacteria actually came from plants (12, 37). The most likely mechanism by which such horizontal gene transfers could occur is natural transformation of bacteria by extracellular DNA released by plants (5). However, several physical and biological conditions must be fulfilled before a plant gene can be expressed by a bacterium, explaining why such events have remained rare and difficult to detect (20). These conditions include the release of DNA very close to those few bacteria that have the relevant molecular mechanisms and which actually develop the physiological state of competence, in which they are able to take up DNA (20). The naked DNA must also escape any rapid chemical or enzymatic degradation and avoid being irreversibly adsorbed onto soil components (16, 42, 43). Finally, transformation requires that the transforming DNA become integrated into the bacterial chromosome by homologous or more or less illegitimate recombination (21).
According to these various conditions, the soil is an unlikely environment for transformation-mediated gene transfers (5, 39), while much more favorable conditions could be encountered in plant tissues in which some symbiotic or pathogenic bacteria multiply actively. This is particularly the case with plants sensitive to the plant pathogen Ralstonia solanacearum, because a rare combination of the following positive factors should be examined for horizontal gene transfer (5). R. solanacearum is a naturally competent bacterium which was found to develop competence in vitro (3) but also in situ during the process of infection of the host plants (4). It can be hypothesized that the infection process could lead plant DNA released by decaying plant cells to be in close contact with these invading and metabolically active bacterial cells. Finally, even the traditional and genetic barriers to such interkingdom transfers due to the molecular mechanisms preventing recombination with foreign DNA in bacteria (3, 18) could be overcome with transgenic plant DNA because of the prokaryotic origin of the marker genes (11, 15).
Our objectives in this paper were to confirm experimentally these hypotheses and to determine the various physical or biological factors which control the transfer of prokaryotic sequences from plants to bacteria.
Evidence of direct contacts between plant DNA and infecting bacteria.
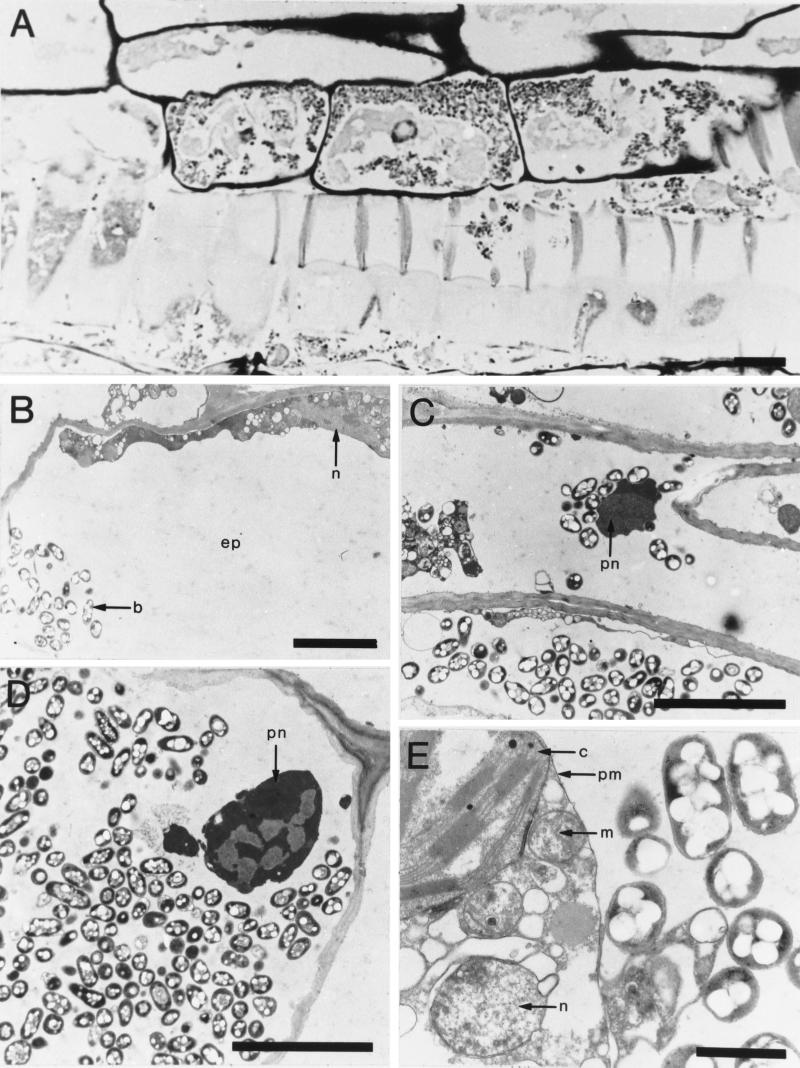
Our initial goal was to determine whether infection of sensitive plants by R. solanacearum resulted in close physical contact between the invading bacteria and the plant nuclear DNA. We used tomato plants infected with R. solanacearum strain GMI1000 (bv. 1), since this strain is virulent towards the tomato (28), with a multiplication rate in planta compatible with the development of the competence state (3). Seeds of the transgenic tomato pKHG3 (24) were sown in potting compost (Ets Grassot, Brignais, France), and the plantlets were grown for 2 weeks before being inoculated with a suspension of R. solanacearum bacteria according to the method of Bertolla et al. (4). Segments of leaves, including a portion of the main leaf vein, were then removed 5 days after inoculation at 30°C and prepared for microscopy by the protocol of Pépin and Boumendil (27). Semithin sections (0.7 to 1 μm thick) were cut for light microscopy, and ultrathin sections (0.07 μm thick) were cut for electron microscopy. Semithin sections stained with Richardson's mixture (30) showed numerous bacteria (Fig. 1A) in mature and differentiating xylem tissue both inside and outside the primary cell walls of the vessels under the light microscope. They were often in the sheath of cells associated with vessels, some of them having intact nuclei. Ultrathin sections were floated onto copper grids and contrasted with uranyl acetate and lead citrate (29). Bacteria were also seen inside cortical parenchyma cells, which contained abundant cytoplasmic and nuclear remnants (Fig. 1B, C, D, and E). Most of the infected cells were adjacent to intact collapsed cells that clearly contained nuclei and chloroplasts. Infection due to the intensive spread of bacteria involved bacterial lytic enzymes, such as endoglucanases, polygalacturonases, and pectin methylesterases (31, 34, 35, 38). Although these enzymes are not absolutely required for wilting, they permit bacteria to spread from cell to cell by degrading plant cell walls and tissues, thus releasing host DNA. Direct observations indicate that physical contact between the plant DNA and invading bacteria cells is possible. In view of results obtained by members of our group on the development of competence in planta (4), these data confirm the potential of an R. solanacearum-based model to fulfill most of the criteria required for natural gene transfer in the environment. These include the release of plant DNA in close contact with bacteria that are physiologically capable of being genetically transformed. We next provided the plants and bacteria with nucleotide sequences favoring the integration of the plant DNA into the recipient genome by homologous recombination and the expression of the marker genes.
FIG. 1.
(A) Light microscope observations of a longitudinal section of tomato xylem tissues infected by R. solanacearum. Bacteria were mainly inside the tracheids and associated parenchyma cells, some of which still contained a cytoplasm and a nucleus. (B, C, D and E) Transmission electron microscope views of tomato parenchyma cells infected by R. solanacearum. (B) R. solanacearum bacteria in a parenchyma cell, the contents of which were necrotic but which still had a recognizable nucleus, are shown. (C) R. solanacearum bacteria are shown in differentiating vascular tissues; an organelle could be identified as a degraded nucleus. (D) Heavy infection of a parenchyma cell, which still contained an organelle that could be a nucleus, is shown. (E) R. solanacearum bacteria are shown in the extraplasmatic space of a plasmolysed parenchyma cell containing living organelles, such as a chloroplast, mitochondria, and a nucleus. Abbreviations: b, bacteria; c, chloroplast; ep, extraplasmatic space; m, mitochondria; n, nucleus; pm, plasma membrane; pn, putative nucleus. Bars indicate lengths of 10 μm (A), 5 μm (B, C, and D), and 1 μm (E).
Construction of sequence-compatible transgenic plants and R. solanacearum strains.
The first strategy was to construct transgenic plants in which the transgene included sequences of the R. solanacearum strain flanking the marker plant genes, to provide homologous sequences between the bacterial and plant genomes. This required construction of the binary plasmid pZpop1, with spectinomycin and gentamycin resistance genes flanked by the popA gene sequences from R. solanacearum GMI1000 (Fig. 2). The plasmids used in the plant transformations were transferred into Agrobacterium tumefaciens EHA105 by electroporation (22). Disks of PBD6 tobacco leaves and tomato plants were transformed with A. tumefaciens according to the protocols from Rogers et al. (32) and Fillatti et al. (14), respectively. Calli were cultured on selective medium containing 100 mg of kanamycin/liter for tobacco or 50 mg of kanamycin/liter for tomato for about 5 weeks. The resulting resistant green plantlets were placed in soil, and their DNA was checked. Plant DNA was extracted from tobacco and tomato leaves by the cetyltrimethylammonium bromide method (13), and Southern hybridizations (33) indicated that one to four copies of transgenic DNA (T-DNA) had been integrated into the plant nuclear genomes (Table 1).
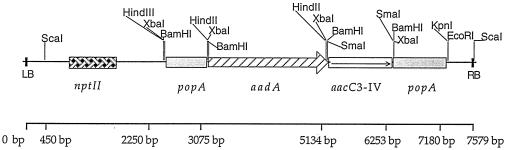
FIG. 2.
Physical map of the T-DNA of plasmid pZpop1 used to transform plants. The gentamycin resistance gene aacC3-IV that was extracted from pUC1813AM/Gm (9) after a SalI digest was cloned into the XhoI site of the pFB2 plasmid to give pFB21. A SacII cassette containing the antibiotic resistance genes flanked by 719 and 907 bp of the popA gene from the R. solanacearum GMI1000 strain was cloned into the SmaI site of the binary vector pPZP212 (17). LB and RB are the left and right borders of the T-DNA.
TABLE 1.
Bacterial strains, plasmids, and plants used in this study
| Strain, plasmid, or plant | Description and/or relevant genotype | Source or reference |
|---|---|---|
| Bacterial strains | ||
| Ralstonia solanacearum GMI1000 | Wild type | 26 |
| Ralstonia solanacearum GMI1000FAP2 | Recipient strain nptII | 4 |
| Ralstonia solanacearum GMI1000FB3 | Recipient strain ΔT-DNA::aacC3-IV | This work |
| Plasmids | ||
| pBluescript | Stratagene, La Jolla, Calif. | |
| pFAP2 | nptII gene from Tn5 | 4 |
| pFAP3 | nptII inactivated by a 324-bp deletion | This work |
| pFB1 | popA gene from R. solanacearum | This work |
| pFB2 | popA::aad | This work |
| pFB21 | popA::aad, aacC3-IV | This work |
| pFB3 | popA::ΔT-DNA::aacC3-IV | This work |
| pT-DNA | T-DNA from pBin19 | This work |
| pΔT-DNA | Deleted T-DNA (nptII inactivated by a 354-bp deletion) | This work |
| pPZP212 | Binary vector for plant transformation | 19 |
| pBin19 | Binary vector with T-DNA containing nptII | 6 |
| pKHG3 | Binary vector with T-DNA containing nptII | 2 |
| pZpop1 | Binary vector with popA::aad, aacC3-IV | This work |
| Plants | ||
| Nicotiana tabacum | ||
| cv. pBD6 | Wild type | |
| pKHG3-ER1 | Transgenic; 1 copy of T-DNA from pKHG3 | This work |
| pKHG3-ER2 | Transgenic; 3 copies of T-DNA from pKHG3 | This work |
| pBin19-ER1 | Transgenic; 1 copy of T-DNA from pKHG3 | This work |
| pBin19-ER2 | Transgenic; 3 copies of T-DNA from pKHG3 | This work |
| Zpop1-EP1 | Transgenic; 1 copy of T-DNA from pZpop1 | This work |
| Zpop1-EP2 | Transgenic; 4 copies of T-DNA from pZpop1 | This work |
| Lycopersicon esculentum | ||
| Ailsa Craig | Wild type | |
| pKHG3 | Transgenic; 1 copy of T-DNA from pKHG3 | 27 |
| Zpop1 | Transgenic; 2 copies of T-DNA from pZpop1 | This work |
We also modified the R. solanacearum strain with sequences from the transgenic plant. Because some of the transgenic plants contained the nptII marker gene, conferring resistance to kanamycin, we cloned a part of this nptII gene in R. solanacearum to provide the recipient bacterial strain with sequences in which homologous recombination could occur. We constructed the plasmid pFB3 containing the deleted nptII gene from the plasmid pBin19 flanked with sequences of the popA gene from the R. solanacearum strain GMI1000 (Fig. 3). pFB3 was transferred to strain GMI1000 by natural transformation according to the method of Bertolla et al. (3). Other transformations of R. solanacearum were also performed by this method. The popA sequences permitted the integration of the deleted nptII gene into the R. solanacearum genome by homologous recombination. The recipient strain GMI1000FB3 was checked by PCR using primers FGPnptII1544 and FGPnptII2347′ (4). Moreover, the sensitivity of the gentamycin-resistant clones to ampicillin confirmed the absence of replicative plasmids in the strain GMI1000FB3.
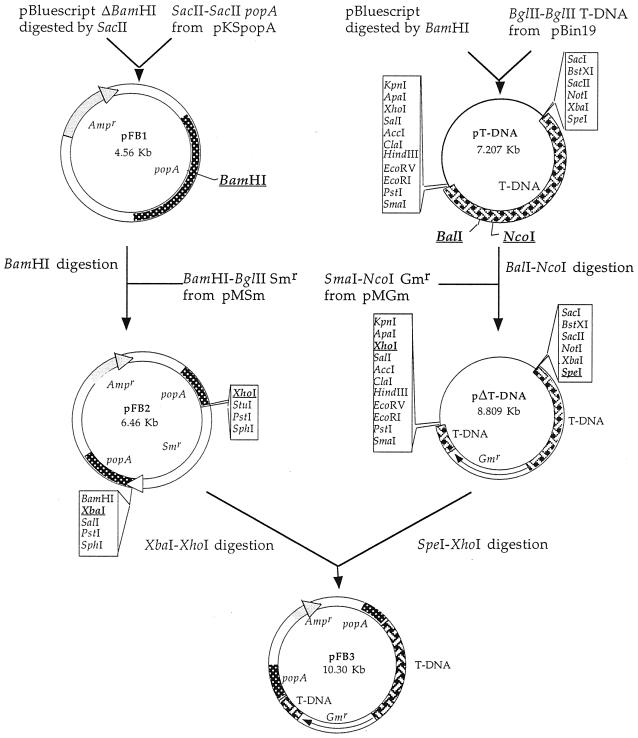
FIG. 3.
Construction of the pFB3 plasmid containing a defective T-DNA. The BglII-BglII T-DNA fragment from pBin19 (6) was ligated into the BamHI linearized pBluescript vector to create plasmid pT-DNA. This plasmid was digested with NcoI and BalI to generate a deletion of 354 bp within the nptII gene before being ligated to the gentamycin resistance cassette resulting from the digestion of pMGm with NcoI-SmaI to give plasmid pΔT-DNA. The SacII-SacII popA gene that was recovered from pKSpopA (1) was ligated into a SacII linearized pBluescript whose BamHI restriction site was deleted previously to give plasmid pFB1. pFB1 was modified by the ligation of the BamHI-BglII aad gene (40) into the unique BamHI site of the popA gene, creating pFB2 in order to provide suitable conditions for the integration of this defective T-DNA. Plasmid pFB3 was constructed by cloning the 5.5-kb SpeI-XhoI fragment of defective T-DNA from pΔT-DNA into the SpeI-XhoI sites of the vector pFB2.
Plasmids pKHG3 and pBin19, which contained the nptII marker gene in the T-DNA region, were used to transform tobacco and tomato plants and so to provide the corresponding transgenic plants (Table 1).
Virulence of the recipient R. solanacearum strains.
The symptoms of infection of the transgenic tomato plants Lycopersicon esculentum pKHG3 and ZPop1, inoculated by the strain GMI1000FB3 and the wild-type strain GMI1000, respectively, were similar, including severe necrosis around the infection sites and the pith of wilted plants, which appeared to be water soaked, brown and hollow. Infected plants wilted within 6 days after the first appearance of symptoms (at 2 days). The infected stems were crushed and homogenized in sterile distilled water with an Ultra-Turrax T25 homogenizer at 25,000 rpm (Janke and Kunkel, Staufen, Germany) to determine the bacterial population dynamics. The plant tissue suspensions were then plated on Boucher gelose (BG) media (8), supplemented with 12 μg of gentamycin/ml for strain GMI1000FB3. The population kinetics in planta of the GMI1000 and GMI1000FB3 strains were similar and reached 4.5 × 109 ± 1.7 × 109 and 6.6 × 109 ± 1 × 109 CFU g of fresh material−1 (n = 3) after 5 days, respectively.
In vitro transformation of the recipient R. solanacearum strains with plasmid and transgenic plant DNA.
Two conditions were tested to validate the various donor plant DNA-recipient bacterial strain combinations. We first transformed the recipient R. solanacearum strains in vitro with the various plasmids used to transform the plants. R. solanacearum GMI1000 was transformed with 0.1 μg of the binary recombinant plasmid pZpop1. Clones resistant to gentamycin (12 μg ml−1) and spectinomycin (40 μg ml−1) were detected at frequencies reaching 3.76 × 10−7 ± 2.14 × 10−8 (n = 3). Similar results were obtained with strain GMI1000FB3 transformed by plasmids pKHG3 and pBin19 on BG media supplemented with 25 μg of kanamycin/ml (Table 2). Natural transformation of this recipient strain by plasmids restored a functional copy of the nptII kanamycin resistance gene. The presence of the marker genes in transformants was checked by PCR with primers complementary to part of the aacC3-IV gene (FGPaac1, 5′-TCCTTCTGAAGGCTCTTCTC-3′, and FGPaac2, 5′-GCAATACGAATGGCGAAAAG-3′) and with the set of primers targeting the nptII gene (4). The expected PCR products were detected, including 601- and 803-bp-long DNA fragments for the aacC3-IV gene in strain GMI1000 and the restored nptII marker gene in GMI1000FB3, respectively (data not shown). Neither the back trap method (41) nor the plasmid extraction kit from Qiagen (Courtaboeuf, France) detected any plasmid, indicating that the marker gene was integrated into the recipient genome (results not shown). This set of experiments indicated that the targeted sequences could be integrated and expressed in R. solanacearum, whatever the endogenous (popA gene) or exogenous (nptII gene) origin of the sequences involved in the homologous recombination mechanism.
TABLE 2.
Transformation frequencies of R. solanacearum recipient strains with transgenic plant DNA and plasmids containing T-DNA
| Donor DNA | T-DNA/plant genomea | Transformation frequency for R. solanacearum strainb:
|
|
|---|---|---|---|
| GMI1000 | GMI1000FB3 | ||
| Plasmid pZpop1 | 3.76 × 10−7 ± 2.1 × 10−8 | ||
| Lycopersicon esculentum Zpop1 | 2 | <1.53 × 10−9 | |
| Nicotiana tabacum Zpop1-EP1 | 1 | <1.6 × 10−9 | |
| Nicotiana tabacum Zpop1-EP2 | 4 | <1.36 × 10−9 | |
| Plasmid pKHG3 | 1.43 × 10−7 ± 5.3 × 10−8 | ||
| Lycopersicon esculentum pKHG3 | 1 | <4.38 × 10−9 | |
| Nicotiana tabacum pKHG3-ER1 | 1 | <3.2 × 10−9 | |
| Nicotiana tabacum pKHG3-ER2 | 3 | <2.91 × 10−9 | |
| Plasmid pBin19 | 8.9 × 10−8 ± 2.3 × 10−8 | ||
| Nicotiana tabacum pBin19-ER1 | 1 | <1.95 × 10−9 | |
| Nicotiana tabacum pBin19-ER2 | 3 | <2.35 × 10−9 | |
The genome sizes of Lycopersicon esculentum and Nicotiana tabacum are 1 × 109 and 4.6 × 109 bp, respectively.
The length of sequence similarity of each side of the marker mutation between the donor DNA and the recipient strain is 719–907 bp for GMI1000 and 700–600 bp for GMI1000FB3.
As a second step, recipient strains GMI1000 and GMI1000FB3 were transformed in vitro with DNA extracted from the transgenic tobacco and tomato plants Zpop1, pKHG3, and pBin19 (Table 1). Concentrations of DNA from 0.1 to 1,000 μg ml−1 were tested. We also used plant DNA that had been treated with restriction endonucleases that did not affect the transgene region to mimic the partial degradation of DNA that could occur in the plant. Transformants remained undetectable at all the DNA concentrations tested and with long and restricted DNA fragments, indicating that transformation, if any, occurred at frequencies below 1.36 × 10−9 (Table 2). The transformation efficiency was not increased when the R. solanacearum recipient strains were transformed by electroporation (10), indicating that the lack of transformation was not due to the natural process of DNA uptake.
In planta tests to track plant-bacterium gene transfers.
Transgenic tomato plants (pKHG3) were inoculated with R. solanacearum strain GMI1000FB3 containing the deleted nptII gene under conditions that permitted the bacterium to become competent. Infected stems were removed and crushed 5 days later and were plated on BG media supplemented with 25 μg of kanamycin/ml. Kanamycin-resistant bacteria were detected at about 102 CFU g of fresh plant material−1. However, hybridization of the genomic DNA of these clones with a nptII probe remained negative, indicating that kanamycin resistance was not encoded by nptII (data not shown). Moreover, their amplified and restricted 16S ribosomal gene, with the FGPS5 and FGP1509′ primers (25), showed patterns that did not correspond to those for R. solanacearum (data not shown). These clones did not belong to R. solanacearum but were part of the indigenous, epiphytic, potentially opportunistic but unidentified microflora that naturally contained the gene for kanamycin resistance but which did not demonstrate that horizontal gene transfer had occurred.
We also tested strain GMI1000 as a potential recipient of the R. solanacearum-indigenous popA sequences, specifically cloned in the transgenic tomato plant Zpop1 (Fig. 2). The crushed infected plant tissues did not produce any spectinomycin- and gentamycin-resistant colonies when plated on BG medium supplemented with 50 μg of spectinomycin/ml and 12 μg of gentamycin/ml, indicating that no gene transfer was detected.
It can be argued that a transfer event would not have any positive effect on the fitness of the strain which would have permitted the specific multiplication of the transformants and favored their detection. We therefore tried to provide the natural medium with selection pressure by inoculating the transgenic plant tissues on days 2 and 5 after infection with 0.1 ml of antibiotic solution (50 μg of kanamycin/ml for plants pKHG3; 24 μg of gentamycin/ml and 100 μg of spectinomycin/ml for plants Zpop1), according to the protocol described by Bertolla et al. (4). The plants were then crushed on day 7. In spite of these numerous assays to boost the growth of potential recombinant clones, transformants remained undetectable. These negative results confirmed those in which R. solanacearum strains were transformed in vitro with plant DNA and indicated that the frequency of horizontal gene transfer in planta must be below about 4.27 × 10−9 ± 8.8 × 10−10 transformants per recipient cell (n = 20).
Factors limiting transformation.
Some of the numerous factors which may limit gene exchanges between plants and bacteria could not be used to explain these negative results. The successful transformation of R. solanacearum with plasmid DNA issued from Escherichia coli (3) and A. tumefaciens (19) could exclude any influence of restriction and modification mechanisms (7). This could mean that R. solanacearum belongs to the group of competent bacteria in which DNA is translocated into the cytoplasm as single-stranded molecules, so escaping the degradation mechanisms specific to double-stranded DNA.
The successful transformation of another naturally competent bacterium, such as Acinetobacter calcoaceticus, by DNA from transgenic plants (11, 15) confirms that plant DNA can be internalized by bacteria. In our study we hypothesized that specific proteins or histones bound to plant DNA which did not inhibit transformation in A. calcoaceticus would not affect specifically R. solanacearum. The fact that these two bacteria differ in their efficiency of transformation, the frequency of which is routinely 10−2 in A. calcoaceticus (26) and less than 10−6 in R. solanacearum, could explain the different rates of transformation of these two bacteria by plant DNA.
Another key factor to be considered is the characteristics of the donor DNA: its number of copies of target sequences, the size of the plant genome, and the ratio between the transgene and the whole plant genome. An increase of the complexity of the donor DNA should reduce the frequency of transformation for a given gene due to increased competition with nontarget sequences.
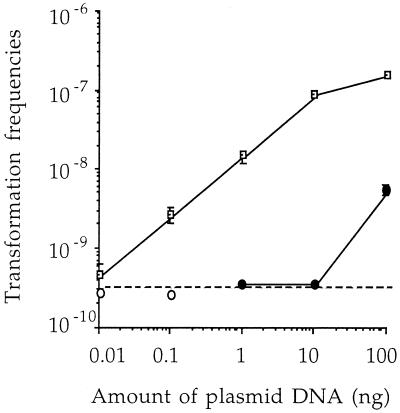
In order to check the influence of donor DNA complexity on the frequency of transformation, R. solanacearum was transformed in vitro with various amounts of pZpop1 plasmid DNA (100, 10, 1, 0.1 and 0.01 ng), alone or diluted in 5 μg of nontransgenic plant DNA (from L. esculentum, var. Ailsa Craig). The minimal amount of pZpop1 plasmid providing detectable transformants was 0.01 ng when the pure plasmid solution was used (Fig. 4), corresponding to 6.7 × 105 copies of transforming sequences. This value was actually lower than the actual number of transforming sequences in 5 μg of transgenic tomato or tobacco plant DNA, indicating that the absolute copy number of selectable sequences in transgenic plant DNA would be high enough to provide transformants at detectable frequencies. Moreover, plasmid DNA mixed with plant DNA background always provided a 10-fold-lower transformation frequency than an identical concentration of pure plasmid DNA solution (Fig. 4). The minimal amount of plasmid pZpop1 (1 ng) in mixed DNA that provided detectable transformants corresponded to 6.7 × 107 transforming molecules, 15 times higher than the actual copy number in the genome of transgenic tomato plants (genome, about 1 × 109 bp) and 67 times more than that in tobacco plants (genome, about 4.6 × 109 bp). These data confirm that the efficiency of transformation of R. solanacearum is directly related to the complexity of the donor DNA. The efficiency of transformation of a marker gene present as only a few copies in transgenic plant DNA must be at least 100-fold less than that for plasmid solutions. Transfers could thus occur at frequencies not lower than 10−11, which is 6 orders of magnitude higher than those expected for Erwinia chrysanthemi, another bacterial plant pathogen (36). Moreover, the fact that R. solanacearum develops competence in planta and acts directly to release plant DNA indicates that such transfer events could occur in the environment at quite relevant frequencies. These data confirm the usefulness of models based on direct symbiotic but mainly pathogenic plant-bacterium relationships for investigating horizontal gene transfers in situ.
FIG. 4.
Influence of the concentration of pZpop1 on the transformation frequencies of R. solanacearum GMI1000. R. solanacearum GMI1000 cells were transformed with 100, 10, 1, 0.1, and 0.01 ng of a pure pZpop1 solution (open squares). Transformations were also conducted with the same amounts of plasmid diluted in 5 μg of wild-type tomato DNA (solid circles). The open circles indicate transformation frequencies below the detection limit (dotted line). Error bars show standard deviations of triplicate experiments. The mean value symbols from three replicate experiments occasionally obscure the smaller standard error bars.
Acknowledgments
We thank Stephane Peyrard and the Electron Microscopy Center (CMEABG) of the Claude Bernard University in Lyon for technical assistance.
This work was supported by the Biotechnology program of the Ministère Français de l'Enseignement Supérieur et de la Recherche (MENRT). F.B. and A.S. were funded by a grant from the MENESER and by the European Agriculture and Fisheries Program (contract number FAIR-98-5002), respectively.
REFERENCES
- 1.Arlat M, Van Gijsegem F, Huet J C, Pernollet J C, Boucher C A. PopA1, a protein which induces a hypersensitivity-like response on specific Petunia genotypes, is secreted via the Hrp pathway of Pseudomonas solanacearum. EMBO J. 1994;13:543–553. doi: 10.1002/j.1460-2075.1994.tb06292.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Béclin C, Charlot F, Botton E, Jouanin L, Doré C. Potential use of the aux2 gene from Agrobacterium rhizogenes as a conditional negative marker in transgenic cabbage. Transgenic Res. 1993;2:48–55. [Google Scholar]
- 3.Bertolla F, Van Gijsegem F, Nesme X, Simonet P. Conditions for natural transformation of Ralstonia solanacearum. Appl Environ Microbiol. 1997;63:4965–4968. doi: 10.1128/aem.63.12.4965-4968.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bertolla F, Brito B, Frostegard A, Nesme X, Simonet P. During infection of its host, the plant pathogen Ralstonia solanacearum naturally develops a state of competence and exchanges genetic material. Mol Plant-Microbe Interact. 1999;12:467–472. [Google Scholar]
- 5.Bertolla F, Simonet P. Horizontal gene transfers in the environment: natural transformation as a putative process for gene transfers between transgenic plants and soil micro-organisms. Res Microbiol. 1999;150:1–10. doi: 10.1016/s0923-2508(99)80072-2. [DOI] [PubMed] [Google Scholar]
- 6.Bevan M. Binary Agrobacterium vectors for plant transformation. Nucleic Acids Res. 1984;12:8711–8721. doi: 10.1093/nar/12.22.8711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bickle T A, Krüger D H. Biology of DNA restriction. Microbiol Rev. 1993;57:434–450. doi: 10.1128/mr.57.2.434-450.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Boucher C A, Barberis P A, Trigalet A P, Demery D A. Transposon mutagenesis of Pseudomonas solanacearum: isolation of Tn5 induced avirulent mutants. J Gen Microbiol. 1985;131:2449–2457. [Google Scholar]
- 9.Braü B, Pilz U, Piepersberg W. Genes for gentamicin-(3)-N-acetyltransferases III and IV: nucleotide sequence of the AAC(3)-IV gene and possible involvement of an IS40 element in its expression. Mol Gen Genet. 1984;193:179–187. doi: 10.1007/BF00327434. [DOI] [PubMed] [Google Scholar]
- 10.Burns J L, Hedin L A. Genetic transformation of Pseudomonas cepacia using electroporation. J Microbiol Methods. 1991;13:215–221. [Google Scholar]
- 11.De Vries J, Wackernagel W. Detection of nptII (kanamycin resistance) gene in genomes of transgenic plants by marker-rescue transformation. Mol Gen Genet. 1998;257:606–613. doi: 10.1007/s004380050688. [DOI] [PubMed] [Google Scholar]
- 12.Doolittle R F. The case of gene transfer between very distantly related organisms. In: Syvanen M, Kado C I, editors. Horizontal gene transfer. London, United Kingdom: Chapman and Hall; 1998. pp. 311–320. [Google Scholar]
- 13.Doyle J, Doyle J L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull. 1987;1:11–15. [Google Scholar]
- 14.Fillatti J J, Kiser J, Rose R, Comai L. Efficient transfer of a glyphosate tolerance gene into tomato using a binary Agrobacterium tumefaciens vector. Biotechnology. 1987;5:726–730. [Google Scholar]
- 15.Gebhard F, Smalla K. Transformation of Acinetobacter sp. BD413 by transgenic sugar beet DNA. Appl Environ Microbiol. 1998;64:1550–1554. doi: 10.1128/aem.64.4.1550-1554.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gebhard F, Smalla K. Monitoring field releases of genetically modified sugar beets for persistence of transgenic plant DNA and horizontal gene transfer. FEMS Microbiol Ecol. 1999;28:261–272. [Google Scholar]
- 17.Hajdukiewicz P, Svab Z, Maliga P. The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Mol Biol. 1994;25:989–994. doi: 10.1007/BF00014672. [DOI] [PubMed] [Google Scholar]
- 18.Khasanov F K, Zvingila D J, Zainullin A A, Prozorov A A, Bashkirov V I. Homologous recombination between plasmid and chromosomal DNA in Bacillus subtilis requires approximately 70 bp of homology. Mol Gen Genet. 1992;234:494–497. doi: 10.1007/BF00538711. [DOI] [PubMed] [Google Scholar]
- 19.Le T, Leccas D, Boucher C. Proceedings of the 4th International Conference on Pathogenic Bacteria. Angers, France: Institut National de la Recherche Agronomique; 1978. Transformation of Pseudomonas solanacearum K60; pp. 819–822. [Google Scholar]
- 20.Lorenz M G, Wackernagel W. Bacterial gene transfer by genetic transformation in the environment. Microbiol Rev. 1994;58:563–602. doi: 10.1128/mr.58.3.563-602.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Matic I, Rayssiguier C, Radman M. Interspecies gene exchange in bacteria: the role of SOS and mismatch repair systems in evolution of species. Cell. 1995;80:507–515. doi: 10.1016/0092-8674(95)90501-4. [DOI] [PubMed] [Google Scholar]
- 22.Mersereau M, Pazour G J, Das A. Efficient transformation of Agrobacterium tumefaciens by electroporation. Gene. 1990;90:149–151. doi: 10.1016/0378-1119(90)90452-w. [DOI] [PubMed] [Google Scholar]
- 23.Message B, Boistard P, Pitrat M, Smith J, Boucher C. Proceedings of the 4th International Conference on Pathogenic Bacteria. Angers, France: Institut National de la Recherche Agronomique; 1978. A new class of fluidal avirulent mutant of Pseudomonas solanacearum unable to induce a hyper sensitive reaction; pp. 823–833. [Google Scholar]
- 24.Moreau-Mhiri C, Morel J B, Audeon C, Ferault M, Grandbastien M A, Lucas H. Regulation of expression of the tobacco Tnt1 retrotransposon in heterologous species following pathogen-related stresses. Plant J. 1996;9:409–419. [Google Scholar]
- 25.Normand P, Orso S, Cournoyer B, Jeannin P, Chapelon C, Dawson J, Evtushenko L, Misra A. Molecular phylogeny of the genus Frankia and related genera and emendation of the family Frankiaceae. Int J Syst Bacteriol. 1996;46:1–9. doi: 10.1099/00207713-46-1-1. [DOI] [PubMed] [Google Scholar]
- 26.Palmen R, Buijsman P, Hellingwerf K J. Physiological regulation of competence induction for natural transformation in Acinetobacter calcoaceticus. Arch Microbiol. 1994;162:344–361. [Google Scholar]
- 27.Pépin R, Boumendil J. Préservation de l'ultrastructure du sclérote de Sclerotinia tuberosa (Hedw.) Fuckel (champignon discomycète). Un modèle pour la préparation des échantillons imperméables et hétérogènes. Cytologia. 1982;47:359–377. [Google Scholar]
- 28.Prior P, Steva H. Characteristics of strains of Pseudomonas solanacearum from French West Indies. Plant Dis. 1990;74:13–17. [Google Scholar]
- 29.Reymolds E S. The use of lead citrate as an electron opaque stain in electron microscopy. J Cell Biol. 1963;17:208–212. doi: 10.1083/jcb.17.1.208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Richardson K C, Jarret L, Finke E H. Embedding in epoxy resin for ultrathin sectioning in electron microscopy. Stain Technol. 1961;35:313. doi: 10.3109/10520296009114754. [DOI] [PubMed] [Google Scholar]
- 31.Roberts D P, Denny T P, Schell M. Cloning of the egl gene of Pseudomonas solanacearum and analysis of its role in phytopathogenicity. J Bacteriol. 1988;170:1445–1451. doi: 10.1128/jb.170.4.1445-1451.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Rogers S G, Horsch R B, Fraley R T. Gene transfer in plants: production of transformed plants using Ti plasmid vectors. Methods Enzymol. 1986;118:627–640. [Google Scholar]
- 33.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 34.Schell M A, Roberts D P, Denny T P. Cloning of the pglA gene of Pseudomonas solanacearum and its involvement in phytopathogenicity. J Bacteriol. 1988;170:4501–4508. doi: 10.1128/jb.170.10.4501-4508.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Schell M A, Denny T P, Huang J. Extracellular virulence factors of Pseudomonas solanacearum: role in disease and their regulation. In: Kado C I, Crosa J, editors. Molecular mechanisms of bacterial virulence. Dordrecht, The Netherlands: Kluwer Academic Publishers; 1993. pp. 311–324. [Google Scholar]
- 36.Schlüter K, Fütterer J, Potrykus I. Horizontal gene transfer from transgenic potato line to a bacterial pathogen (Erwinia chrysanthemi) occurs—if at all—at an extremely low frequency. Biotechnology. 1995;13:1094–1098. doi: 10.1038/nbt1095-1094. [DOI] [PubMed] [Google Scholar]
- 37.Smith M W, Feng D F, Doolittle R F. Evolution by acquisition, the case of horizontal gene transfers. Trends Biochem Sci. 1992;17:489–493. doi: 10.1016/0968-0004(92)90335-7. [DOI] [PubMed] [Google Scholar]
- 38.Spök A, Stubenrauch G, Schorgendorfer K, Schwab H. Molecular cloning of a pectin esterase gene from Pseudomonas solanacearum. J Gen Microbiol. 1991;137:131–140. doi: 10.1099/00221287-137-1-131. [DOI] [PubMed] [Google Scholar]
- 39.Stotzky G. Gene transfer among bacteria in soil. In: Levy S B, Miller R V, editors. Gene transfer in the environment. New York, N.Y: McGraw-Hill Book Co.; 1989. pp. 165–222. [Google Scholar]
- 40.Ubben D, Schmitt R. A transposable promoter and transposable promoter probes derived from Tn1721. Gene. 1987;53:127–134. doi: 10.1016/0378-1119(87)90100-4. [DOI] [PubMed] [Google Scholar]
- 41.Wheatcroft R, McRae D G, Miller W. Changes in the Rhizobium meliloti genome and the ability to detect supercoiled plasmids during bacteroid development. Mol Plant-Microbe Interact. 1990;3:9–17. [Google Scholar]
- 42.Widmer F, Seidler R J, Wartud L S. Sensitive detection of transgenic plant marker gene in persistence in soil microcosms. Mol Ecol. 1996;5:603–613. [Google Scholar]
- 43.Widmer F, Seidler R J, Donegan K K, Reed G L. Quantification of transgenic plant marker gene persistence in the field. Mol Ecol. 1997;6:1–7. [Google Scholar]