Abstract
The enzymatic transformation of 2,4,6-trinitrotoluene (TNT) by purified XenB, an NADPH-dependent flavoprotein oxidoreductase from Pseudomonas fluorescens I-C, was evaluated by using natural abundance and [U-14C]TNT preparations. XenB catalyzed the reduction of TNT either by hydride addition to the aromatic ring or by nitro group reduction, with the accumulation of various tautomers of the protonated dihydride-Meisenheimer complex of TNT, 2-hydroxylamino-4,6-dinitrotoluene, and 4-hydroxylamino-2,6-dinitrotoluene. Subsequent reactions of these metabolites were nonenzymatic and resulted in predominant formation of at least three dimers with an anionic m/z of 376 as determined by negative-mode electrospray ionization mass spectrometry and the release of ∼0.5 mol of nitrite per mol of TNT consumed. The extents of the initial enzymatic reactions were similar in the presence and in the absence of O2, but the dimerization reaction and the release of nitrite were favored under aerobic conditions or under anaerobic conditions in the presence of NADP+. Reactions of chemically and enzymatically synthesized and high-pressure liquid chromatography-purified TNT metabolites showed that both a hydroxylamino-dinitrotoluene isomer and a tautomer of the protonated dihydride-Meisenheimer complex of TNT were required precursors for the dimerization and nitrite release reactions. The m/z 376 dimers also reacted with either dansyl chloride or N-1-naphthylethylenediamine HCl, providing evidence for an aryl amine functional group. In combination, the experimental results are consistent with assigning the chemical structures of the m/z 376 species to various isomers of amino-dimethyl-tetranitrobiphenyl. A mechanism for the formation of these proposed TNT metabolites is presented, and the potential enzymatic and environmental significance of their formation is discussed.
Bacterial transformation of the explosive 2,4,6-trinitrotoluene (TNT) (Table 1, compound 1) has been shown to proceed via two different biochemical routes: direct 2e− reduction of the aromatic ring via hydride addition and successive 2e− reduction of the nitro groups. The extents of these reactions and the predominant products depend on the type of microorganism involved and the environmental conditions. Here we consider the action of an enzyme purified from an aerobic bacterium, Pseudomonas fluorescens I-C.
TABLE 1.
TNT metabolites produced by XenB as revealed by ESI-MS and HPLC analyses
| Compound(s) | Name | Chemical compositiona | Molecular mass (Da)b | HPLC retention time(s) (min)c |
|---|---|---|---|---|
| 1 | TNT | C7H5O6N3 | 226 | 18.9 |
| 2 | H-TNT | C7H6O6N3− | 228 | 23.6 |
| 3–6 | 2H-TNT · H+ | C7H8O6N3− | 230 | 6.4, 10.1, 13.4, 17.7 |
| 7 | 2-Hydroxylamino-4,6-dinitrotoluene | C7H7O5N3 | 212 | 11.6 |
| 8 | 4-Hydroxylamino-2,6-dinitrotoluene | C7H7O5N3 | 212 | 12.1 |
| 9 | 2-Amino-4,6-dinitrotoluene | C7H7O4N3 | 196 | 15.0 |
| 10 | 4-Amino-2,6-dinitrotoluene | C7H7O4N3 | 196 | 15.5 |
| 11 | Azoxynitrotoluene isomers | C14H10O9N6 | 405 | 24.5–25.5 |
| 12 | Amino-dimethyl-tetranitrobiphenyl isomers | C14H11O8N5 | 376 | 24.5–25.5 |
| 13 | Metabolite from anaerobic incubation | C7H7O4N3 | 196 | 1.5–4.0 |
| 14 | Metabolite from anaerobic incubation | C7H9O4N3 | 198 | 1.5–4.0 |
Chemical composition of the compound.
Molecular mass of the anion as determined by negative-mode ESI-MS.
Retention time(s) as determined by the ion pair HPLC separations described in Materials and Methods.
Direct reduction of the aromatic ring has been reported to be catalyzed by aerobic bacteria (8, 31, 32), and the products include a hydride-Meisenheimer complex (H-TNT), a dihydride-Meisenheimer complex (2H-TNT), and several protonated tautomers of 2H-TNT (2H-TNT · H+). Reduction of the nitro group in aerobic environments results in accumulation of 2-hydroxylamino-4,6-dinitrotoluene, 4-hydroxylamino-2,6-dinitrotoluene, 2-amino-4,6-dinitrotoluene, 4-amino-2,6-dinitrotoluene, 2,4-diamino-6-nitrotoluene, isomers with other combinations of partially or fully reduced nitro groups, and azoxy dimers (11, 21, 32, 33). Cometabolic oxidation of TNT to 2,4,6-trinitrobenzyl alcohol and 2,4,6-trinitrobenzoate has also been reported for propane-grown Mycobacterium vaccae (30), but putative enzymatic steps or intermediates in these reactions have not been described.
Unlike TNT, dinitrotoluenes are known to be growth substrates for aerobic bacteria (24). In the initial steps of the metabolic pathways, nitrite is apparently released from intermediates of the hydroxylation or dioxygenation reactions by α,β elimination. Thus, for aerobic metabolism of TNT, release of nitrite from the aromatic ring appears to be a critical requirement for further metabolism. Mineralization of TNT by TOL transformants of Pseudomonas sp. clone A has been reported (6, 11), suggesting that cometabolic hydroxylation reactions may indeed facilitate nitrite release. 2,4-Dinitrotoluene has been suggested as a TNT by-product, but mass balance confirmation of nitrite release was not provided (11). Likewise, a Mycobacterium strain reduced TNT to H-TNT and apparently released nitrite (31), but subsequent studies revealed false-positive reactions of the Griess reagent used for colorimetric nitrite quantitation (32), making the assessment of the nitrite release mechanism equivocal. Pentaerythritol tetranitrate (PETN) reductase purified from Enterobacter cloacae has also been reported to catalyze TNT transformation by aromatic ring reduction and to release nitrite, again on the basis of the use of the Griess reagent (8, 9). In these previous studies, the carbon-containing metabolites associated with nitrite release were not identified, leaving the mechanism of nitrite release unresolved.
Recently, we have worked with the flavoprotein xenobiotic reductases XenA from Pseudomonas putida II-B and XenB from P. fluorescens I-C (1, 2). Along with PETN reductase and other enzymes, XenA and XenB are members of an evolutionarily related family of NAD(P)H-dependent flavoprotein oxidoreductases. Although XenA and XenB have high primary sequence identity, suggesting similar three-dimensional structures, they exhibit different selectivities for removal of the nitro groups from nitroglycerin. These differences in catalytic selectivity extend to reactions with TNT. While XenA catalyzes only nitro group reductions, XenB transforms TNT via nitro group reduction and direct reduction of the aromatic ring. Using ion pair high-pressure liquid chromatography (HPLC), ion chromatography, liquid scintillation counting, optical, and mass spectral techniques, we have detected 13 TNT products resulting from XenB-initiated TNT transformations. Among the reactions, a nonenzymatic reaction of the hydroxylamino-dinitrotoluene (HADNT) isomers and 2H-TNT · H+ results in nitrite release and the formation of various isomers with proposed chemical structures corresponding to that of amino-dimethyl-tetranitrobiphenyl. Since the same products are observed after transformation of TNT by P. fluorescens I-C, potential accumulation of these novel biphenyl dimers in the environment is indicated.
MATERIALS AND METHODS
Chemicals.
Nitroglycerin was provided by Olin Corporation (Baraboo, Wis.). TNT was obtained from Chem Service (West Chester, Pa.). 2,4-Dinitrotoluene, 2,6-dinitrotoluene, 1,3-dinitrobenzene, nitrobenzene, and the three nitrotoluene isomers were purchased from Aldrich Chemical Company (Milwaukee, Wis.). [U-14C]TNT (21.58 mCi/mmol; purity, greater than 95%, as determined by thin-layer chromatography) was purchased from ChemSyn Science Labs (Lenexa, Kans.). Dansyl chloride was purchased from Molecular Probes (Eugene, Oreg.). Isotopically labeled 18OH2 (97% 18O content) was obtained from ICON (Summit, N.J.). All other chemicals were analytical grade or better.
Chemical syntheses.
The HADNT isomers were chemically synthesized by using zinc dust and ammonium chloride (19) and were purified by HPLC as described below. H-TNT was chemically synthesized by using sodium borohydride (11, 16) and was purified by HPLC as described below. Isomers of 2H-TNT · H+ were also produced by reduction of TNT with sodium borohydride and were purified by HPLC.
Dansyl chloride was prepared as a 100 mM solution in acetonitrile. Dansylation reactions were performed by mixing 15 μl of dansyl solution, 100 μl of NaHCO3 (50 mM, pH 8.3), and 50 μl of HPLC-purified compounds resuspended in water. Reactions were carried out in the dark at room temperature. After 30 min of incubation, the dansylation reaction was stopped by addition of 25 μl of 0.1 N NaOH (10).
For thin-layer chromatography, HPLC-purified compounds were resuspended in acetonitrile:water (1:1, vol/vol) and spotted onto silica gel thin-layer chromatography plates. Each silica plate was sprayed with NaNO2 (0.1% [wt/vol] solution in 1 N HCl), followed by N-1-naphthylethylenediamine HCl (0.04% [wt/vol] solution prepared in methanol). The plate was air dried and then heated at 100°C for 1 h and exposed to UV light (27).
Bacterial strains, culturing methods, and whole-cell assays.
The bacterium used in this study, P. fluorescens I-C, was isolated (2) from nitroglycerin-contaminated soil obtained from the Badger Army Ammunition Plant (Baraboo, Wis.) on the basis of its ability to tolerate high concentrations of nitroglycerin in the growth medium. For whole-cell assays, P. fluorescens I-C was grown aerobically in Luria-Bertani medium at 30°C. After overnight incubation, the cells from 1 ml of the medium were isolated by centrifugation at 14,000 × g, washed twice with 100 mM phosphate buffer (pH 7), and resuspended in 1 ml of the same buffer. A 500-μl aliquot of 0.56 mM TNT dissolved in deionized water was added to the culture, and TNT transformation products were determined by HPLC.
Enzyme reactions.
XenB was purified by standard chromatographic methods and was characterized as previously reported (2). One unit of XenB activity was defined as the amount of enzyme that catalyzed consumption of 1 μmol of NADPH per min. Reactions were carried out in 1 ml of 100 mM phosphate (pH 7.0) in the presence of 300 μM nitro-containing substrate and 130 μM NADPH. TNT transformation reactions were performed in 1.5-ml polyethylene microcentrifuge tubes containing 900 μl of 100 mM phosphate (pH 7.0), 100 nmol of TNT, and 300 nmol of NADPH. The reactions were initiated by the addition of purified XenB to give 5.4 μg of enzyme per reaction mixture.
HPLC methods.
TNT metabolites were separated by ion pair chromatography by using a Beckman (Fullerton, Calif.) HPLC equipped with a Nova-Pak radial compression C18 column (bead size, 4 μm; 5 by 100 mm; Waters Chromatography, Milford, Mass.). The ion pair reagent tetrabutylammonium dihydrogen phosphate (pH 7.5) was added to both deionized water (aqueous buffer) and acetonitrile (acetonitrile buffer) to a final concentration of 5 mM. The initial composition of the mobile phase was 75% aqueous buffer and 25% acetonitrile buffer. This composition was maintained for 1 min. The percentage of acetonitrile buffer was then linearly increased to 32% over 12.2 min, linearly increased to 64% over 10.5 min, and increased stepwise to 90%. The 90% acetonitrile buffer composition was maintained for 2.3 min. At the end of the separation protocol, the column was reequilibrated to 25% acetonitrile buffer over 5 min. TNT and metabolites were detected by UV-visible absorption spectrometry by using a diode array detector. Metabolites derived from aromatic ring reduction were detected at 477 nm, while metabolites derived from nitro group reduction were detected at 230 nm. The retention times observed for TNT and other metabolites by the ion pair HPLC method and elution protocol described above are summarized in Table 1.
TNT metabolites resulting from dimerization reactions (Table 1, compounds 11 and 12) were not completely separated by the ion pair HPLC method and were analyzed by performing a different gradient elution with 20 mM ammonium acetate buffer (pH 7.5) and acetonitrile. During separation, an initial mobile phase composition of 100% ammonium acetate buffer was maintained for 1 min. The percentage of acetonitrile was linearly increased to 20% over 2 min and maintained for 5 min, linearly increased to 25% over 2 min and maintained for 2 min, increased to 35% over 0.5 min and maintained for 7.5 min, and increased to 90% over 0.5 min and maintained for 14.5 min.
Ion chromatography.
Nitrite was detected and quantified by using an LC20 ion chromatograph (Dionex, Sunnyvale, Calif.) equipped with a GP40 gradient pump, an ED40 electrochemical detector, an IonPac AG11 guard column (4 by 50 mm), an IonPac ATC-1 anion trap column, an ASRS-Ultra 4-mm suppression system, and an IonPac AS11 anion-exchange column (4 by 250 mm). NaNO2 was used as a standard for calibration curves.
ESI-MS analysis.
Electrospray ionization mass spectral (ESI-MS) data were obtained with a Perkin-Elmer Sciex API 365 triple quadrupole spectrometer at the Mass Spectrometry Facility of the University of Wisconsin Biotechnology Center. When the TNT metabolites were recovered after ion pairing separation, the fractions collected were dried with a speed vacuum, resuspended in acetonitrile, and then separated from the pairing reagent by HPLC by the ammonium acetate method described above. Metabolites recovered after HPLC separation by the ammonium acetate method were dried and resuspended on acetonitrile-water (1:1, vol/vol) for injection into the spectrometer.
Liquid scintillation counting.
[U-14C]TNT was mixed with TNT in acetonitrile to produce a 10 mM solution with a specific activity of 8,770 dpm/nmol. Ten-microliter portions of this solution were added to enzyme reaction mixtures, giving a total of 100 nmol of TNT per reaction mixture. The distribution of 14C in TNT and the various metabolites was evaluated by injecting 100 μl of a reaction mixture into the HPLC column and collecting 0.5-ml aliquots of the HPLC eluent throughout the entire HPLC run. A scintillation cocktail (4.5 ml; ScintiSafe Econo 2; Fisher) was added to the vials, and the counts per minute were determined with a Packard Tri-Carb 2100TR liquid scintillation analyzer. Individual scintillation vials were counted either for 1 min or to achieve an E2ς value of more than 2%. The 14C elution profile was correlated with the retention times of TNT and the known metabolites to determine the relative percentage of each compound in the enzyme reaction mixtures. In addition, the 14C elution profile was used to define the retention times and quantities of unknown metabolites.
RESULTS
Reaction of purified XenB with TNT.
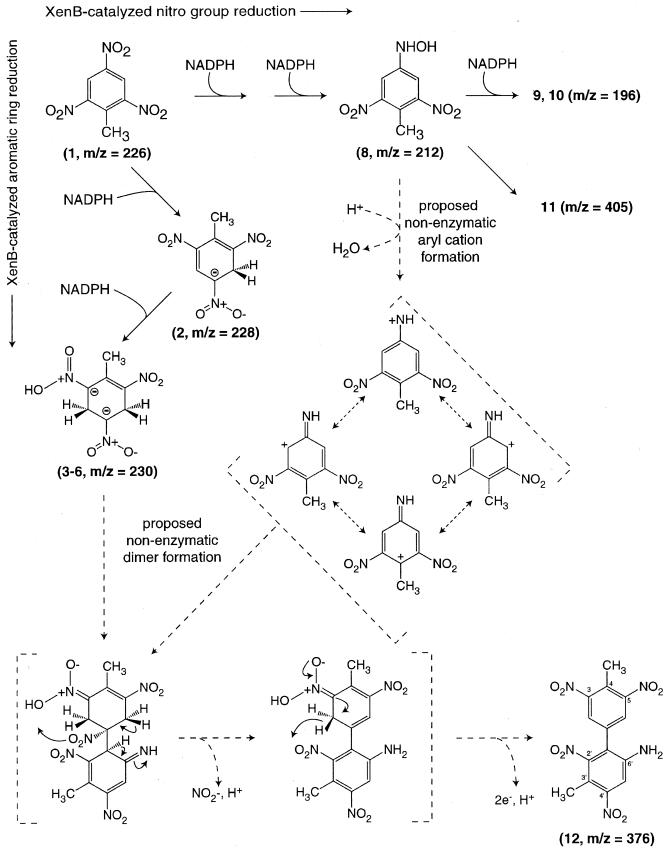
In this work, we focused on characterization of metabolites produced by the reaction of purified XenB with TNT under either aerobic or anaerobic conditions. In addition, chemically synthesized compounds corresponding to the metabolites isolated from the enzyme reaction mixtures were used to investigate the potential contributions of successive enzymatic reductions and abiotic reactions. A summary of the TNT metabolites detected is presented in Table 1, while Fig. 1 shows the products of XenB-catalyzed reduction of TNT. Figure 1 also shows a proposed mechanism for abiotic reactions leading to the release of nitrite and the formation of dimeric compounds that are proposed here to be various isomers of amino-dimethyl-tetranitrobiphenyl.
FIG. 1.
XenB-catalyzed transformation of TNT and proposed subsequent nonenzymatic reactions leading to nitrite release and formation of amino-dimethyl-tetranitrobiphenyl. The m/z values indicated were determined by ESI-MS for the anion of each compound detected during the reactions. Compounds in brackets are proposed intermediates not detected in the experiments.
The ability of XenB to reduce TNT and other nitroaromatic compounds was initially evaluated by measuring the stimulation of NADPH consumption in the presence of a nitro-containing compound. As shown in Table 2, the rate of TNT-stimulated NADPH consumption was about 40% of that observed for the comparable reaction with nitroglycerin. Purified XenB also reacted with 2,4-dinitrotoluene and 1,3-dinitrobenzene (Table 2) but did not react with 2,6-dinitrotoluene or any of the mononitrated aromatic compounds tested. Compared with XenB evaluations, evaluations performed with the related flavoprotein xenobiotic reductase XenA (1) revealed ∼20% of the reactivity with TNT and no substantial reactivity with any of the other nitroaromatic compounds shown in Table 2. Thus, XenA reactions were not characterized further in this work.
TABLE 2.
Specific activities of XenB with different substrates
| Substrate | Sp act (U/mg)a |
|---|---|
| Nitroglycerin | 6.8 |
| TNT | 2.6 |
| 2,4-Dinitrotoluene | 0.1 |
| 1,3-Dinitrobenzene | 0.1 |
| 2,6-Dinitrotoluene | 0 |
| Nitrobenzene | 0 |
| 2-Nitrotoluene | 0 |
| 3-Nitrotoluene | 0 |
| 4-Nitrotoluene | 0 |
One unit of activity is defined as 1 μmol of NADPH oxidized per min. The reported activities are averages based on three independent measurements. The standard errors for measurable activities were less than 10%.
Release of nitrite from TNT.
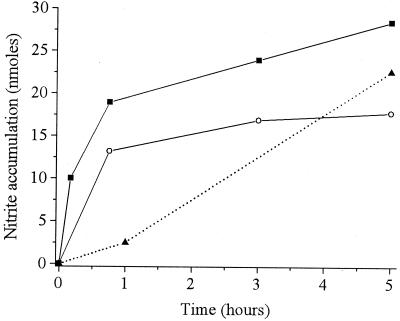
Ion chromatography was used to demonstrate unambiguously and to quantify the release of nitrite from TNT during XenB-catalyzed reactions (Fig. 2). From the 100 nmol of TNT originally present in the aerobic enzyme reaction mixtures, ∼28 nmol of nitrite was recovered after 5 h and ∼50 nmol was recovered after 24 h. With anaerobic enzyme reaction mixtures, ∼18 nmol of nitrite was recovered after 5 h and ∼48 nmol was recovered after 24 h. Nitrite was also observed during aerobic incubation of TNT with P. fluorescens I-C cells (Fig. 2), with ∼23 nmol of nitrite recovered after 5 h and ∼37 nmol recovered after 24 h. In all the experiments, at least 50% of the TNT had been transformed after only 10 min of incubation and about 90% had been transformed after 30 min. Thus, a primary goal of this work was to characterize the TNT metabolites that were produced by the nitrite release reaction.
FIG. 2.
Nitrite accumulation during transformation of TNT by XenB and whole cells of P. fluorescens I-C. The initial conditions were 100 nmol of TNT and 300 nmol of NADPH in 100 mM phosphate buffer (pH 7). Symbols: ■, aerobic transformation by XenB; ○, anaerobic transformation by XenB; ▴, aerobic transformation by P. fluorescens I-C.
Products observed from aromatic ring reduction.
Upon addition of TNT, reaction mixtures containing XenB changed from colorless to red and then to yellow. According to previous descriptions (31, 32), these changes suggested successive formation of H-TNT, 2H-TNT, and various 2H-TNT · H+. HPLC analyses of the XenB reaction mixtures demonstrated transient accumulation of compound 2, which had an HPLC retention time of 23.6 min (Table 1) and an optical spectrum (λmax, ∼480 nm [31, 32]) indistinguishable from that of chemically synthesized H-TNT. After resuspension of compound 2, negative-mode ESI-MS revealed an m/z value of 228, which corresponded to the chemical formula of the H-TNT anion (Table 1). Control experiments performed with TNT, NADPH, and no enzyme showed no conversion of TNT, demonstrating that XenB was required to catalyze the formation of compound 2. Thus, compound 2 was identified as H-TNT, a transient product of the NADPH-dependent reaction of XenB and TNT.
The subsequent appearance of a yellow color in the reaction mixture was associated with accumulation of four distinct metabolites (compound 3, with a retention time of 6.4 min, a UV λmax of 236 nm, and a visible λmax of 449 nm; compound 4, with a retention time of 10.1 min, a UV λmax of 262 nm, and a visible λmax of 492 nm; compound 5, with a retention time of 13.4 min, a UV λmax of 262 nm, and a visible λmax of 466 nm; and compound 6, with a retention time of 17.7 min, a UV λmax of 262 nm, and a visible λmax of 487 nm) (Table 1; Fig. 1). These products had UV-visible spectra similar to those reported for metabolites arising from reduction of H-TNT (8, 32) and subsequently identified (32) as various tautomers of 2H-TNT · H+. Negative-mode ESI-MS analyses of the HPLC-purified metabolites revealed an m/z value of 230 for each of the four compounds, which corresponded to the molecular mass of 2H-TNT · H+ (Table 1).
Enzyme reactions in which chemically synthesized compound 2 was used as the substrate resulted in rapid accumulation of compounds 3 through 6. In contrast, control reaction mixtures that contained compound 2, NADPH, and no enzyme yielded the previously demonstrated (11, 32) reverse transformation of H-TNT to TNT (∼98% conversion after 12 h) and no accumulation of compounds 3 through 6. These results demonstrate that XenB also catalyzes reduction of H-TNT to 2H-TNT.
Further studies of compounds 3 through 6 revealed that their interconversion was not enzymatic and that compound 5 was the most stable of these four metabolites. Thus, when HPLC-purified compound 3 (i.e., compound 3 chemically synthesized by reduction of TNT with sodium borohydride, collected from an HPLC eluent, and dried with a speed vacuum) was resuspended in phosphate buffer (100 mM, pH 7.0) and reinjected into the HPLC after 3 h, compound 3 (6% ± 1% [average ± standard deviation based on at least two experiments]), compound 5 (79% ± 3%), and compound 6 (15% ± 2%) were detected. Similarly, resuspension and reinjection of compound 4 into the HPLC resulted in the appearance of compounds 3 (6% ± 2%), 4 (20% ± 1%), 5 (62% ± 2%), and 6 (12% ± 3%). Resuspension and reinjection of compound 5 resulted in mainly compound 5 (81% ± 4%) and smaller amounts of compounds 3 (5% ± 1%) and 6 (14% ± 2%). Finally, reinjection of compound 6 revealed mainly compound 5 (81% ± 2%) and smaller fractions compounds 3 (5% ± 1%) and 6 (14% ± 1%). When HPLC-purified compounds 3 through 6 were individually used as substrates for XenB reactions, no further metabolites were detected and the distribution of HPLC peak areas among the resultant products (compounds 3 through 6) was nearly indistinguishable from that obtained in the absence of XenB.
Taken together, these experiments demonstrated that XenB can catalyze the reduction of TNT (compound 1) to H-TNT (compound 2) and the reduction of H-TNT to 2H-TNT. Further rearrangements among the various 2H-TNT tautomers (compounds 3 through 6) was nonenzymatic. These results are summarized in Fig. 1. Our data were not sufficient to determine the exact chemical structures of compounds 3 through 6. Nevertheless, the results show that compounds 3 through 6 are related to each other by reversible reactions and that their relative concentrations in the reaction mixtures reach chemical equilibrium in either the presence or the absence of XenB and NADPH.
Products observed from nitro group reduction.
In addition to the products derived from aromatic ring reduction, the XenB-catalyzed reaction with TNT yielded substantial amounts of 2HADNT (compound 7, with a retention time of 11.6 min) and 4HADNT (compound 8, with a retention time of 12.1 min) (Table 1; Fig. 1), and compound 8 was the major product. Identification of these metabolites was based on the following experimental evidence: (i) the HPLC retention times of the enzyme products closely matched those of chemically synthesized compounds 7 and 8; (ii) the UV-visible spectra of the enzyme products matched those of the chemically synthesized compounds; and (iii) negative-mode ESI-MS revealed an m/z value of 212 for both compound 7 and compound 8, which corresponded to the predicted mass of the anion of the two possible HADNT isomers (Table 1). The individual isomers were assigned to the corresponding HPLC peaks on the basis of the higher yield of compound 8 obtained from the chemical synthesis (19) and on the basis of differences in the UV-visible spectra reported for the two isomers (15).
For longer incubation periods, small amounts of 2-amino-4,6-dinitrotoluene (compound 9, with a retention time of 15.0 min) and 4-amino-2,6-dinitrotoluene (compound 10, with a retention time of 15.5 min) were also identified on the basis of the similarity of the HPLC retention times with those of available standards and on the basis of the m/z value of 196 determined by ESI-MS analyses (Table 1).
Use of [U-14C]TNT.
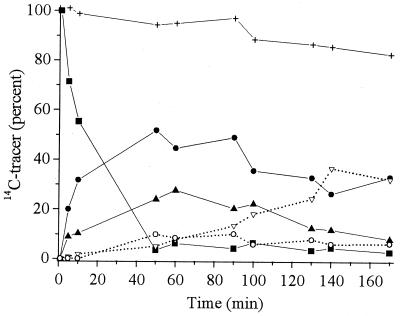
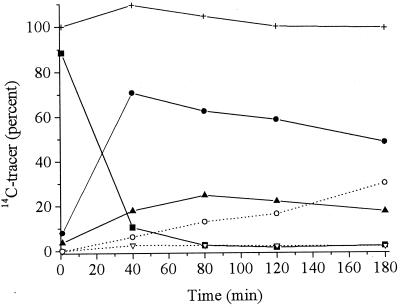
[U-14C]TNT was used to ascertain the mass balance for the metabolites produced by XenB. Figure 3 shows the time course for distribution of 14C during a typical aerobic enzyme reaction. For simplicity, the 14C counts were treated as counts from five groups of metabolites, reflecting unreacted TNT (compound 1), metabolites from aromatic ring reduction (compounds 2 through 6), metabolites from nitro group reduction (compounds 7 through 10), dimerization products (compounds 11 and 12), and two unidentified metabolites having retention times of 1.5 to 4.0 min (compounds 13 and 14). About 6 to 10% of 14C was associated with compounds 13 and 14, and this proportion did not change substantially during the course of the aerobic reactions. Recovery of 14C during HPLC separations was greater than 80% throughout the entire reaction period.
FIG. 3.
Mass balance for TNT metabolites during aerobic transformation with XenB based on the recovery of 14C from [U-14C]TNT. Symbols: ■, TNT (compound 1); ●, metabolites from aromatic ring reduction (compounds 2 through 6); ▴, metabolites from nitro group reduction (compounds 7 through 10); ▿, dimerized products (compounds 11 and 12); ○, unidentified metabolites; +, total 14C recovery.
After 50 min, ∼3% of the 14C was in unreacted TNT, 52% was in products derived from aromatic ring reduction (compounds 3 through 6), 24% was in products derived from nitro group reduction (compounds 7 and 8, 21%; compounds 9 and 10, 3%), and 5% was in HPLC peaks with retention times of 24.5 to 25.5 min (unresolved compounds 11 and 12) (Table 1). For reaction times greater than 50 min, the 14C contents of the products derived from both aromatic ring reduction (compounds 3 through 6) and nitro group reduction (compounds 7 through 10) decreased, while the 14C contents of the HPLC peaks of compounds 11 and 12 increased noticeably. After 170 min, comparable amounts of 14C were present in the HPLC peaks of compounds 11 and 12 and the products of aromatic ring reduction (33%, mostly compound 5) (see above), while the amount of 14C in the products of nitro group reduction (particularly compounds 7 and 8) had decreased to ∼8%. Thus, accumulation of compounds 11 and 12 occurred along with a decrease in the amounts of the products derived from both aromatic ring reduction and nitro group reduction.
At the end of the reaction, ∼18% of the original 14C was not recovered. However, after repeated extraction of the reaction tubes with acetonitrile, an additional 9% of the original 14C was recovered, indicating that adsorption was the most likely cause of 14C loss. When this acetonitrile-extracted material was subjected to HPLC analysis, compound 12 was identified as the predominant adsorbed metabolite. Thus, compound 12 was the major end product in aerobic reaction mixtures and contained ∼42% of the original TNT.
ESI-MS of compounds 11 and 12 produced by XenB.
For better separation of metabolites with retention times of 24.5 to 25.5 min (Table 1), a different HPLC method with ammonium acetate and acetonitrile was used, as described in Materials and Methods. With this method, three TNT metabolites with an m/z of 376 (Table 1, compound 12) eluted with retention times of 23.5, 23.8, and 24.1 min. The metabolite exhibiting the largest absorption peak eluted at 23.8 min. In addition, a metabolite with an m/z of 405 (compound 11) eluted at 24.3 min. The molecular mass of compound 11 corresponded to that of previously well-characterized azoxydinitrotoluene dimers (11, 21, 33). In contrast, the molecular mass of compound 12 suggested an unknown compound whose mass was consistent with dimerization of TNT metabolites in which nitrite was also released (Table 1; Fig. 1).
An m/z of 376 is consistent with the molecular anion of a compound with an odd number of nitrogen atoms (18). In principle, the mass of compound 12 could be accounted for by an azoxy dimer having one nitrite group replaced by a hydroxyl group. However, when compound 12 was formed in buffer prepared by using 18O-enriched water, an m/z of 376 was still obtained, showing that replacement of a nitrite group by a hydroxyl group was not required to form compound 12. Furthermore, the 18O experiment suggested that compound 12 did not contain a nitroso functional group whose O atom would likely exchange with water.
Combined chemical and enzymatic synthesis of compounds 11 and 12.
A mixture of compounds 7 and 8 was chemically synthesized and used as the substrate for an aerobic XenB reaction. After 24 h, compounds 7 and 8 were completely consumed, and only compound 11 was observed by combined HPLC and ESI-MS analyses. A control experiment conducted with compounds 7 and 8 and without XenB also gave only compound 11, showing that formation of compound 11 was not enzyme catalyzed. Since azoxy dimers are formed from HADNT isomers (33), compound 11 was assigned to represent all isomeric azoxy dimers having an m/z of 405 that were observed in this work (Table 1). When similar aerobic XenB reactions were undertaken by using the HPLC-purified, individual, aromatic ring reduction products (compounds 3 through 6) as the substrates, no further enzymatic transformation was observed (see above), and notably, neither compound 11 nor 12 was produced.
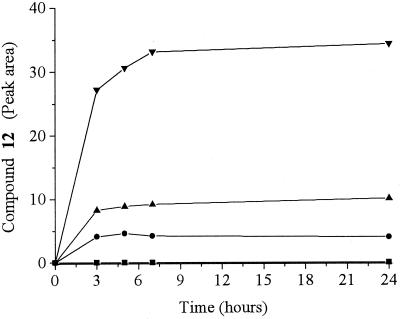
The results in Fig. 3 suggested that compound 12 may have arisen from combination of the products of aromatic ring reduction and nitro group reduction. Thus, by combination of chemically synthesized and HPLC-purified compounds 7 and 8 with enzymatically synthesized and HPLC-purified compound 5 (the major species of compounds 3 through 6), compounds 11 and 12 were produced, and compound 12 was the major product. In aerobic reactions in which the synthesized precursors were used, the rates of formation of compound 12 were similar regardless of whether XenB or NADPH was present. Moreover, in experiments performed with an excess of chemically synthesized compounds 7 and 8, the amount of compound 12 formed was proportional to the amount of compound 5 present (Fig. 4). Direct quantification of the nitrite released during the reaction of compound 5, with compounds 7 and 8 was unfortunately not possible because at high concentrations of compounds 7 and 8, a peak resulting from these compounds was not fully resolved from the nitrite peak. Nevertheless, the results of these experiments show that compound 12 was formed by the nonenzymatic reaction of an HADNT (compound 7 or 8) and a metabolite derived from aromatic ring reduction (compound 3, 4, 5, or 6), and the molecular weight of compound 12 is consistent with the release of a nitro group during the reaction. Since both compounds 7 and 8 were consumed in the reaction, at least two isomers of compound 12 are likely. Furthermore, the observed solution equilibrium involving compounds 3 through 6 did not permit identification of which compound reacted to give compound 12, but if more than one compound did react, additional isomers of compound 12 would be likely.
FIG. 4.
Nonenzymatic formation of compound 12 by reaction of excess chemically synthesized compounds 7 and 8 (780 nmol) with enzymatically synthesized compound 5. Symbols: ■, no compound 5 added; ●, 10 nmol of compound 5 added; ▴, 27 nmol of compound 5 added; ▾, 76 nmol of compound 5 added.
In order to investigate the possible functional groups present in compound 12, colorimetric reactions diagnostic for the presence of amine groups were performed (13, 27). A control reaction of TNT with dansyl chloride at pH 8.3 gave no modification of TNT as detected by HPLC, showing that nitro groups were not reactive. In contrast, reaction with compound 10 gave a complete reaction, showing that the amine group was reactive. A comparable reaction of compound 12 with dansyl chloride also gave a complete reaction, as judged from the disappearance of compound 12 from the HPLC chromatogram. Since the 18O labeling experiments indicated that a hydroxyl group was not present, the dansylation reaction indicated that a reactive amino group was present in compound 12. Further evidence for the presence of an aromatic amine was obtained by use of NaNO2 and N-1-naphthylethylenediamine HCl, which react to give a characteristic UV-active derivative of the amine functional group. While TNT did not give a colored product from this reaction, both compounds 10 and 12 were purple under UV illumination, providing further evidence for the existence of an aryl amine functional group.
Anaerobic transformation of TNT by XenB.
Figure 5 shows the time course for distribution of 14C during a typical anaerobic enzyme reaction. As in the aerobic mass balance study (Fig. 3), the 14C counts were treated as counts for five groups of metabolites. In this experiment, the percentage of total 14C recovered from HPLC separation was greater than 99% during the reaction period. After 80 min of incubation, 62% of the 14C was in products derived from aromatic ring reduction (mostly compounds 3, 5, and 6), 25% was in products derived from nitro group reduction (compounds 7 and 8, 21%; compounds 9 and 10, 4%), and ∼13% was in two unidentified compounds (compounds 13 and 14; retention times, 1.5 to 4.0 min). Notably, compounds 11 and 12 were not produced under anaerobic conditions and accounted for only 2% of the 14C after 80 min.
FIG. 5.
Mass balance of TNT metabolites during anaerobic transformation with XenB based on recovery of 14C from [U-14C]TNT. Symbols: ■, TNT (compound 1); ●, metabolites from aromatic ring reduction (compounds 3 through 6); ▴, metabolites from nitro group reduction (compounds 7 through 10); ▿, dimerized products (compounds 11 and 12); ○, unidentified metabolites; +, total 14C recovery.
In the anaerobic reactions, accumulation of compounds 13 and 14 was more evident with longer incubation times, and after 180 min 30% of the 14C was in compounds 13 and 14, 48% was in compounds 3 through 6, and 17% was in compounds 7 through 10. Interestingly, compounds 13 and 14 were not detected in the HPLC elution profiles by optical detection at either 230 or 477 nm. However, when fractions of the HPLC eluent were collected at 1.5 to 4.0 min and subjected to negative-mode ESI-MS, compounds 13 and 14 were determined to have m/z values of 196 and 198, respectively. Although these masses are possibly consistent with further reduction of compounds 9 and 10 to nitroso-hydroxylamino-nitrotoluene and dihydroxylamino-nitrotoluene, the absence of an optical spectrum characteristic of an aromatic ring suggests that other structures are possible, and thus these products have not been conclusively identified.
After 180 min, the anaerobic reaction vial was opened to the atmosphere and incubated aerobically for an additional 90 min. After this treatment, 17% of the 14C was converted to compounds 11 and 12, metabolites that did not accumulate under anaerobic conditions. In addition, the amount in compounds 3 through 6 decreased to 29% and the amount in compounds 7 through 10 decreased to 7%, supporting the proposed reaction of compounds 3 through 6 with compounds 7 and 8 to give compound 12. Furthermore, the amount in compounds 13 and 14 decreased to 20% upon exposure to air. The total recovery of 14C after 180 min was 76%, a value which probably reflected losses due to adsorption.
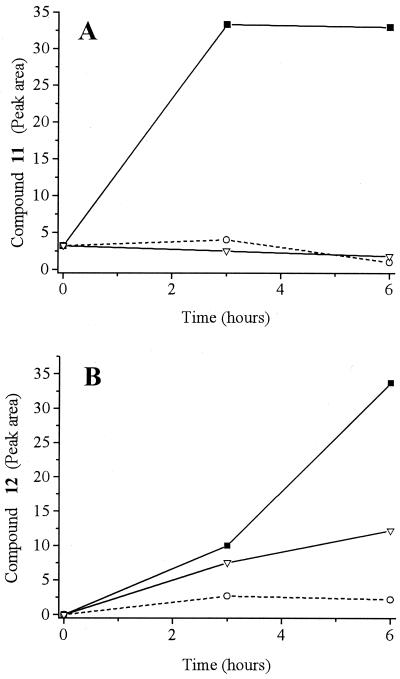
Figure 6 shows that formation of compound 11, which occurs in the presence of oxygen, did not occur under anaerobic conditions. In contrast, formation of compound 12 can occur under both aerobic and anaerobic conditions, but in the absence of oxygen an oxidizing agent, such as NADP+, is required for the reaction to proceed (Fig. 6B).
FIG. 6.
Nonenzymatic formation of compound 11 (A) and compound 12 (B) by reaction of compound 5 with compounds 7 and 8. Symbols: ■, aerobic conditions; ○, anaerobic conditions; ▿, anaerobic conditions with excess (0.5 mM) NADP+.
TNT transformation by whole cells.
When P. fluorescens I-C cells grown on Luria-Bertani medium and resuspended in 100 mM phosphate buffer (pH 7) were amended with TNT, products derived from both aromatic ring and nitro group reduction were detected. As observed with the purified enzyme, the culture medium immediately changed from colorless to red and then to yellow upon the addition of TNT. Figure 1 shows that nitrite was released in the whole-cell reactions, albeit in lower amounts than those observed in the purified enzyme reactions. After 45 min of incubation, nitrite accumulations in whole-cell and enzyme reaction mixtures were 2.5 and 18 nmol, respectively, while after 24 h of incubation, nitrite accumulations in whole-cell and enzyme reaction mixtures were 37 and 50 nmol, respectively. HPLC separation with detection at 477 nm revealed transient formation of compound 2 and accumulation of compounds 3 through 6, while detection at 230 nm revealed accumulation of compounds 7 and 8 and their subsequent conversion to compounds 9 and 10. Accumulation of the latter two metabolites was substantially greater in the whole-cell reaction mixtures than in the enzyme reaction mixtures. Furthermore, both compound 11 and compound 12 were detected in the whole-cell experiments, although accumulation of these compounds was less than that observed in the enzymatic reactions. In whole-cell reaction mixtures, accumulation of compound 12 after 24 h of incubation was 33% less than that in enzymatic reaction mixtures, a difference that correlates well with the observed difference in nitrite accumulation (e.g., there was 26% less nitrite in whole-cell reaction mixtures). The decrease in the accumulation of nitrite and compound 12 was likely due to more extensive conversion of compounds 7 and 8 to compounds 9 and 10, resulting in decreased availability of intermediates required for synthesis of compounds 11 and 12.
DISCUSSION
Our results show that XenB has the ability to catalyze transformation of TNT by aromatic ring reduction and by nitro group reduction. Furthermore, our results show that nonenzymatic reaction of the products leads to release of nitrite under oxidizing conditions (in the presence of either O2 or NADP+) and to accumulation of dimers thought to be isomers of amino-dimethyl-tetranitrobiphenyl. Since P. fluorescens I-C containing XenB produced the same products as those observed with the purified enzyme, it is possible that amino-dimethyl-tetranitrobiphenyls may be produced in natural environments. This suggested pathway provides a new focus for enzymatic enhancement of TNT bioremediation.
Nitrite release from TNT initiated by XenB-catalyzed reduction.
Like XenB, PETN reductase from E. cloacae PB2 has been reported to transform TNT by both aromatic ring reduction and nitro group reduction (8). As determined by colorimetric assays, nitrite release was reported in PETN reductase studies and in other studies (6–8, 31). However, no major carbon-containing products of the denitration reaction were identified in these previous studies, which ultimately limited both the insight into the mechanism of nitrite release and the prospects for improving the putative biotransformations. By using purified XenB, ion chromatography, 14C tracer methods, and reactions of chemically synthesized precursors, we found that up to 0.50 nmol of nitrite can be released per nmol of TNT consumed (Fig. 1). On the basis of enzymatic and chemical characterizations, it now appears that compounds 3 through 6 and compound 7 or 8 are required for the nitrite release reaction described here. The half-integral stoichiometry for the amount of nitrite released relative to the amount of TNT consumed supports the proposed nonenzymatic dimerization mechanism for nitrite release (Fig. 1). On the basis of these results, other transformations would be required to increase the stoichiometry of nitrite release during TNT metabolism.
Proposed mechanism for nitrite release and dimerization.
Our results show that XenB catalyzes reduction of TNT to compounds 3 through 6 and compounds 7 through 10. In contrast, the predominant end product isolated from the reaction mixtures has been proposed to be compound 12, an amino-dimethyl-tetranitrobiphenyl. A mechanism accounting for the formation of compound 12 is shown in Fig. 1. In this proposal, dimerization is initiated by the interaction of resonance-stabilized aryl nitrenium cation and aryl carbanion species. Thus, ion pairing and hydrophobic stacking interactions may be implicit features of this reaction. Acid-catalyzed decomposition of an aromatic hydroxylamine, which is the first step of the Bamberger rearrangement (17), provides a precedent for the proposed formation of a resonance-stabilized aryl nitrenium cation and is consistent with the requirement for an HADNT isomer during the formation of compound 12. Furthermore, compound 2 and compounds 3 through 6 are resonance-stabilized carbanionic intermediates due to the presence of the multiple nitro group substituents, which permits extensive delocalization of electron density from the added hydride equivalents. In this work, the dihydride level intermediates compounds 3 through 6 were shown to also participate in the formation of compound 12.
As noted above, compound 12 likely represents a variety of possible isomers. Figure 1 shows a representative dimerization reaction that gives the sterically least-hindered product; analogous reactions can be drawn to potentially give five other isomers. Formation of the C—C bond between C—4 of 2H-TNT · H+ and C—3 of the aryl nitrenium cation would place a nitro group originally from 2H-TNT · H+ in a position suitable to be eliminated as nitrite, while rearrangement of the imine group would lead to rearomatization of the aromatic ring originally from HADNT. Elimination of nitrite as shown in Fig. 1 would be favorable, as nitrite is an excellent leaving group at neutral pH. Indeed, α,β elimination of nitrite has been observed in a number of other relevant enzyme reactions, notably nitrobenzene dioxygenase (20), 1,3-dinitrobenzene dioxygenase (5), 2-nitrotoluene dioxygenase (12), 2,4-dinitrotoluene dioxygenase (23, 26, 28), and nitrophenol oxygenase (25) reactions. Furthermore, rearrangement of the imine with subsequent rearomatization should be a highly favorable reaction leading to the formation of an aryl amine functional group. Due to a reduced pKa value caused by the ring nitro substituents, the aryl amine should readily react with an acylating agent, such as dansyl chloride, at neutral pH or with NaNO2 and N-1-naphthylethylenediamine HCl under more extreme conditions. Thus, the quantitative conversion of amino-dinitrotoluenes and compound 12 and the lack of reactivity of TNT with these diagnostic reagents provide chemical evidence for the presence of an aryl amine group.
The final rearomatization reaction and the formation of compound 12 would require 2e− oxidation and H+ release. This reaction would be analogous to the well-documented conversion of H-TNT to TNT (31), a process that requires 2e− oxidation and H+ release. As shown in Fig. 6B, this reaction can be mediated by either O2 or NADP+. In principle, a variety of other oxidants could also function in this final oxidation step. The difference between the rate of nitrite release and the rate of formation of compound 12 under either aerobic or anaerobic conditions could in part be due to a faster rearomatization reaction in the presence of O2 than in the presence of other oxidants or due to the effects of the different concentrations of the oxidants on the bimolecular reaction.
Enzymology of TNT metabolism.
XenB is a member of a family of NAD(P)H-dependent flavoprotein xenobiotic reductases (1). Primary sequence analysis of two members of this family, XenA and XenB, revealed 34% primary sequence identity and 51% similarity, suggesting that these enzymes may have related three-dimensional structures. However, catalytic characterizations revealed substantial differences in the regioselectivity for nitrite removal from nitroglycerin and the reduction of cyclohexenone (1, 2). In this work, initial characterizations revealed that XenA had ∼20% of the reactivity of XenB with TNT when preparations were indirectly assayed by determining NADPH consumption and that XenA gave only products derived from nitro group reduction, while XenB could transform TNT into compounds 2 through 6 and compounds 7 through 10.
The ability of other xenobiotic reductases to release nitrite via the dimerization reaction described here may depend on the ability to match the products given by XenB, which include products of both aromatic ring reduction and nitro group reduction. Alternatively, xenobiotic reductases capable of producing primarily hydroxylamino metabolites may primarily yield azoxy dimers in aerobic environments, while xenobiotic reductases capable of producing primarily aromatic ring reduction metabolites offer alternative, although currently poorly understood, possibilities for TNT bioremediation.
Studies with P. fluorescens I-C revealed increased accumulation of amino-dinitrotoluenes compared to that observed in the purified enzyme reactions, suggesting the presence of another enzyme that enhances production of these metabolites. The formation of amino-dinitrotoluenes would also contribute to the observed decreases in the amount of compound 12 and nitrite release, since a required substrate would be consumed by an alternative reaction.
Environmental implications.
The denitration mechanism proposed here provides insight into a potential constraint on microbial utilization of TNT as a nitrogen source, as nitrite release would depend on the availability of both compounds 3 through 6 and compound 7 or 8. Rapid conversion of TNT to amino-dinitrotoluenes, as observed in the whole-cell experiments, would also effectively minimize the nitrite release reaction, which could explain why Rhodococcus erythropolis HL PM-1 failed to utilize TNT as a nitrogen source even though the two required TNT reduction pathways were present (32). Furthermore, production of the precursors compounds 3 through 6 and compound 7 or 8 would require at a minimum consumption of seven equivalents of NAD(P)H in order to produce one equivalent of utilizable nitrogen (e.g., four equivalents consumed by XenB to yield compound 12 and three equivalents consumed by nitrite reductase [EC 1.6.6.4] to convert nitrite to ammonia). This substantial energy demand and the limited nitrite release may effectively prevent utilization of TNT as a sole source of nitrogen by the mechanism proposed in Fig. 1. However, regardless of whether TNT can be used as a source of nitrogen, the results also suggest that polynitrated biphenyl compounds may be formed in the environment from TNT as a consequence of reductive cometabolism. While these compounds have not been reported in previous nitroaromatic transformation studies, the similar chemical reactivities of nitrite and chlorine suggest that various isomers of compound 12 may have properties similar to those of polychlorinated biphenyls, including insolubility, toxicity (4, 14), and recalcitrance to further bioremediation (3, 22, 29). Whether these compounds can be further transformed in either the laboratory or the natural environment remains to be determined.
ACKNOWLEDGMENTS
This work was partially supported by grant MCB 9733734 from the National Science Foundation (to B.G.F.) and contract DAA21-93-C-1034 from the U.S. Army ARDEC (to G.H.C.). Additional support was provided by the College ot Agricultural and Life Sciences (to G.H.C.) and the College of Engineering (to D.R.N.), University of Wisconsin-Madison.
We thank A. Harms (University of Wisconsin Biotechnology Center) for expert help with the ESI-MS analysis.
REFERENCES
- 1.Blehert D S, Fox B G, Chambliss G H. Cloning and sequence analysis of two Pseudomonas flavoprotein xenobiotic reductases. J Bacteriol. 1999;181:6254–6263. doi: 10.1128/jb.181.20.6254-6263.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Blehert D S, Knoke K L, Fox B G, Chambliss G H. Regioselectivity of nitroglycerin denitration by flavoprotein nitroester reductases purified from two Pseudomonas species. J Bacteriol. 1997;179:6912–6920. doi: 10.1128/jb.179.22.6912-6920.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brenner V, Arensdorf J J, Focht D D. Genetic construction of PCB degraders. Biodegradation. 1994;5:359–377. doi: 10.1007/BF00696470. [DOI] [PubMed] [Google Scholar]
- 4.Culp S J, Roberts D W, Talaska G, Lang N P, Fu P P, Lay J O, Jr, Teitel C H, Snawder J E, Von Tunglen L S, Kadlubar F F. Immunochemical, 32P-postlabeling, and GC/MS detection of 4-aminobiphenyl-DNA adducts in human peripheral lung in relation to metabolic activation pathways involving pulmonary N-oxidation, conjugation, and peroxidation. Mutat Res. 1997;378:97–112. doi: 10.1016/s0027-5107(97)00101-2. [DOI] [PubMed] [Google Scholar]
- 5.Dickel O, Knackmuss H-J. Catabolism of 1,3-dinitrobenzene by Rhodococcus sp. QT-1. Arch Microbiol. 1991;157:76–79. doi: 10.1007/BF00245339. [DOI] [PubMed] [Google Scholar]
- 6.Duque E, Haidour A, Godoy F, Ramos J L. Construction of a Pseudomonas hybrid strain that mineralizes 2,4,6-trinitrotoluene. J Bacteriol. 1993;175:2278–2283. doi: 10.1128/jb.175.8.2278-2283.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Fiorella P D, Spain J C. Transformation of 2,4,6-trinitrotoluene by Pseudomonas pseudoalcaligenes JS52. Appl Environ Microbiol. 1997;63:2007–2015. doi: 10.1128/aem.63.5.2007-2015.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.French C E, Nicklin S, Bruce N C. Aerobic degradation of 2,4,6-trinitrotoluene by Enterobacter cloacae PB2 and by pentaerythritol tetranitrate reductase. Appl Environ Microbiol. 1998;64:2864–2868. doi: 10.1128/aem.64.8.2864-2868.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.French C E, Nicklin S, Bruce N C. Sequence and properties of pentraerythritol tetranitrate reductase from Enterobacter cloacae PB2. J Bacteriol. 1996;178:6623–6627. doi: 10.1128/jb.178.22.6623-6627.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gros C, Labouesse B. Study of the dansylation reaction of amino acids, peptides, and proteins. Eur J Biochem. 1969;7:463–470. doi: 10.1111/j.1432-1033.1969.tb19632.x. [DOI] [PubMed] [Google Scholar]
- 11.Haidour A, Ramos J L. Identification of products resulting from the biological reduction of 2,4,6-trinitrotoluene, 2,4-dinitrotoluene, and 2,6-dinitrotoluene by Pseudomonas sp. Environ Sci Technol. 1996;30:2365–2370. [Google Scholar]
- 12.Haigler B E, Wallace W H, Spain J C. Biodegradation of 2-nitrotoluene by Pseudomonas sp. strain JS42. Appl Environ Microbiol. 1994;60:3466–3469. doi: 10.1128/aem.60.9.3466-3469.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Haugland R P. Handbook of fluorescent probes and research chemicals. 6th ed. Eugene, Oreg: Molecular Probes, Inc.; 1996. [Google Scholar]
- 14.Hirayama T, Iguchi K, Yoshida S, Yamanaka Y, Watanabe T. Structural determination of a directly mutagenic amino-nitrobiphenyl as the S9 metabolite of 2,4,2′,4′-tetranitrobiphenyl in Salmonella typhimurium TA98. Mutat Res. 1991;262:203–207. doi: 10.1016/0165-7992(91)90023-w. [DOI] [PubMed] [Google Scholar]
- 15.Huang S, Lindahl P A, Wang C, Bennett G N, Rudolph F B, Hughes J B. 2,4,6-Trinitrotoluene reduction by carbon monoxide dehydrogenase from Clostridium thermoaceticum. Appl Environ Microbiol. 2000;66:1474–1478. doi: 10.1128/aem.66.4.1474-1478.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kaplan L A, Siedle A R. Studies on boron hydrides. IV. Stable hydride Meisenheimer adducts. J Org Chem. 1971;36:937–939. [Google Scholar]
- 17.March J. Advanced organic chemistry. Reactions, mechanisms, and structure. 4th ed. New York, N.Y: John Wiley & Sons; 1992. [Google Scholar]
- 18.McLafferty F W, Tureček F. Interpretation of mass spectra. 4th ed. Mill Valley, Calif: University Science Books; 1993. [Google Scholar]
- 19.Mori M, Kudo Y, Nunozawa T, Miyahara T, Kozuka H. Intestinal metabolism of 2,4-dinitrotoluene in rats. Chem Pharm Bull (Tokyo) 1985;33:327–332. doi: 10.1248/cpb.33.327. [DOI] [PubMed] [Google Scholar]
- 20.Nishino S F, Spain J C. Oxidative pathway for the biodegradation of nitrobenzene by Comamonas sp. strain JS765. Appl Environ Microbiol. 1995;61:2308–2313. doi: 10.1128/aem.61.6.2308-2313.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rieger P-G, Knackmuss H-J. Basic knowledge and perspectives on biodegradation of 2,4,6-trinitrotoluene and related nitroaromatic compounds in contaminated soil. In: Spain J C, editor. Biodegradation of nitroaromatic compounds. New York, N.Y: Plenum Press; 1995. pp. 1–18. [Google Scholar]
- 22.Shannon M J, Unterman R. Evaluating bioremediation: distinguishing fact from fiction. Annu Rev Microbiol. 1994;47:715–738. doi: 10.1146/annurev.mi.47.100193.003435. [DOI] [PubMed] [Google Scholar]
- 23.Spain J C. Bacterial degradation of nitroaromatic compounds under aerobic conditions. In: Spain J C, editor. Biodegradation of nitroaromatic compounds. New York, N.Y: Plenum Press; 1995. pp. 19–35. [Google Scholar]
- 24.Spain J C. Biodegradation of nitroaromatic compounds. Annu Rev Microbiol. 1995;49:523–555. doi: 10.1146/annurev.mi.49.100195.002515. [DOI] [PubMed] [Google Scholar]
- 25.Spain J C, Gibson D T. Pathway for biodegradation of p-nitrophenol in a Moraxella sp. Appl Environ Microbiol. 1991;57:812–819. doi: 10.1128/aem.57.3.812-819.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Spanggord R J, Spain J C, Nishino S F, Mortelmans K E. Biodegradation of 2,4-dinitrotoluene by a Pseudomonas sp. Appl Environ Microbiol. 1991;57:3200–3205. doi: 10.1128/aem.57.11.3200-3205.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Stahl E, editor. Thin layer chromatography. Berlin, Germany: Springer Verlag; 1965. [Google Scholar]
- 28.Suen W-C, Spain J C. Cloning and characterization of Pseudomonas sp. strain DNT genes for 2,4-dinitrotoluene degradation. J Bacteriol. 1993;175:1831–1837. doi: 10.1128/jb.175.6.1831-1837.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Timmis K N, Steffan R S, Unterman R. Designing microorganisms for the treatment of toxic wastes. Annu Rev Microbiol. 1994;48:525–557. doi: 10.1146/annurev.mi.48.100194.002521. [DOI] [PubMed] [Google Scholar]
- 30.Vanderberg L A, Perry J J, Unkefer P J. Catabolism of 2,4,6-trinitrotoluene by Mycobacterium vaccae. Appl Microbiol Biotechnol. 1995;43:937–945. doi: 10.1007/BF02431931. [DOI] [PubMed] [Google Scholar]
- 31.Vorbeck C, Lenke H, Fischer P, Knackmuss H-J. Identification of a hydride-Meisenheimer complex as a metabolite of 2,4,6-trinitrotoluene by a Mycobacterium strain. J Bacteriol. 1994;176:932–934. doi: 10.1128/jb.176.3.932-934.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Vorbeck C, Lenke H, Fischer P, Spain J C, Knackmuss H-J. Initial reductive reactions in aerobic microbial metabolism of 2,4,6-trinitrotoluene. Appl Environ Microbiol. 1998;64:246–252. doi: 10.1128/aem.64.1.246-252.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Won W D, Heckly R J, Glover D J, Hoffsommer J C. Metabolic disposition of 2,4,6-trinitrotoluene. Appl Microbiol. 1974;27:513–516. doi: 10.1128/am.27.3.513-516.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]