Abstract
We describe a rapid, reproducible, and sensitive method for detection and quantification of archaea in naturally occurring microbial communities. A domain-specific PCR primer set and a domain-specific fluorogenic probe having strong and weak selectivity, respectively, for archaeal rRNA genes (rDNAs) were designed. A universal PCR primer set and a universal fluorogenic probe for both bacterial and archaeal rDNAs were also designed. Using these primers and probes, we demonstrated that detection and quantification of archaeal rDNAs in controlled microbial rDNA assemblages can be successfully achieved. The system which we designed was also able to detect and quantify archaeal rDNAs in DNA samples obtained not only from environments in which thermophilic archaea are abundant but also from environments in which methanogenic archaea are abundant. Our findings indicate that this method is applicable to culture-independent molecular analysis of microbial communities in various environments.
Recent molecular phylogenetic analyses based on small-subunit (SSU) rRNA gene (rDNA) sequencing have revealed the remarkable phylogenetic diversity of archaea and their potential ecological significance not only in extreme environments but also in nonextreme environments (3, 4, 7, 10, 12–14, 16, 18, 19, 25, 34, 38, 40, 42, 44). It has been accepted that the abundant occurrence of archaea is not a local event, as these organisms are widely observed in various microbial habitats on the earth. Hence, it is likely that assessment of archaeal community diversity is important for elucidating the structure, function, and interactions of naturally occurring microbial communities. For detection and quantification of archaea in an entire microbial community, quantitative nucleic acid hybridization (11, 12, 17) and whole-cell in situ fluorescent hybridization (2, 13, 34) with archaeal rRNA-targeted nucleotide probes and competitive quantitative PCR (9, 33) have been used.
A fluorogenic PCR method using the TaqMan probe designed to hybridize with a region within the targeted amplicon allows detection and quantification of the initial template concentration (5, 23, 24). Using the TaqMan fluorogenic PCR system, a rapid and reproducible detection and quantification technique has been developed for several infectious and harmful microorganisms (5, 15, 20, 26, 28, 30, 31, 37). This system is probably quite sensitive for detection and quantification of small amounts of the targeted amplicon and can be used for large numbers of samples and specimens due to the advantages of PCR. These features permit possible application for rapid detection and quantification of archaeal rRNA or rDNA in nucleic acid extracts prepared from low-biomass environmental samples and a great number of samples, such as samples taken from long drilled cores and water columns at short intervals. The proportion of archaeal rRNA or rDNA in the whole microbial rRNA or rDNA provides an important clue for estimating the abundance of archaea in naturally occurring microbial communities. Here, we describe a rapid, reproducible, and sensitive method for detection and quantification of archaeal rDNAs in DNA extracts from various environments. This method is based on the TaqMan fluorogenic PCR system and will be integrated into a comprehensive, culture-independent, molecular analysis system for environmental microbiology.
Establishment of quantitative fluorogenic PCR system.
A multiple alignment was obtained from Ribosomal Database Project II (22), and an updated multiple alignment of archaeal SSU rDNA sequences was constructed by including various nearly complete or partial rDNA sequences from cultivated members and environmental clones reported previously (38, 42, 44). Based on the alignments, appropriate sites for SSU rDNA-targeted universal and domain-specific oligonucleotide primers and probes for the TaqMan fluorogenic PCR system were chosen. Following the manufacturer's manual provided with the TaqMan fluorogenic PCR system (PE Applied Biosystems, Foster City, Calif.), we designed the universal and domain-specific primers to have appropriate melting temperatures between 55 and 60°C and to produce rDNA amplicons of appropriate lengths (<500 bp). The universal and domain-specific TaqMan probes were designed to have melting temperatures greater than 65°C, which are higher than those of the primers which were designed, and appropriate binding positions as recommended by the manufacturer. The primers were Arch349F (5′-GYGCASCAGKCGMGAAW-3′) and Arch806R (5′-GGACTACVSGGGTATCTAAT-3′) for archaeal rDNA, Bac349F (5′-AGGCAGCAGTDRGGAAT-3′) and Bac806R (5′-GGACTACYVGGGTATCTAAT-3′) for bacterial rDNA, and Uni340F (5′-CCTACGGGRBGCASCAG-3′) and Uni806R (5′-GGACTACNNGGGTATCTAAT-3′) for prokaryotic universal rDNA (the number in each primer or probe designation indicates the position of the 5′ end of the primer or probe in Escherichia coli 16S rRNA). The TaqMan probes were Arch516F (5′-TGYCAGCCGCCGCGGTAAHACCVGC-3′) for archaeal rDNA, Bac516F (5′-TGCCAGCAGCCGCGGTAATACRDAG-3′) for bacterial rDNA, and Uni516F (5′-TGYCAGCMGCCGCGGTAAHACVNRS-3′) for prokaryotic universal rDNA. These probes each had a fluorescent reporter dye (6-carboxyfluorescein) covalently attached to the 5′ end and a fluorescent quencher dye (6-carboxytetramethylrhodamine) attached six or more bases downstream from the reporter dye. All primers and probes were analyzed by using the PROBE_CHECK program from the Ribosomal Database Project (22) and the gapped-BLAST search algorithm (1, 6) to examine selectivity for the targeted domain or universality for both prokaryotic domains. The computer analyses indicated that the domain-specific primers and probes bound specifically to the appropriate sites in the targeted domains of the rDNA. The TaqMan probes were obtained from PE Applied Biosystems.
In order to check the selectivity of the domain-specific primers for the targeted domains of the rDNA during the PCR, a nonfluorogenic PCR in the absence of the TaqMan probes was performed by using LA Taq polymerase with GC buffer (TaKaRa, Kyoto, Japan) and representative bacterial and archaeal rDNAs (almost the full length between primer Bac27F [21] or Arch21F [12] and primer 1492R [21]) directly amplified and purified from genomic DNAs and environmental clones as templates. The genomic DNAs of Rhodothermus obamensis JCM9785 (35), Thermaerobacter marianensis JCM10246 (39), Hydrogenobacter sp. strain JCM10560, Pseudomonas aeruginosa JCM5962, Clostridium perfringens JCM1290, Cytophaga marinoflava JCM8517, Shewanella violacea JCM10179 (29), Bacillus subtilis JCM1465, Thermosipho japonicus JCM10495 (41), E. coli INVα (Invitrogen, Carlsbad, Calif.), Haloarcula japonica JCM7785, and Palaeococcus ferrophilus JCM10417 (43) were preserved at our institute, and samples of genomic DNAs of Pyrobaculum sp. strain JCM10595 and Sulfurisphaera sp. were kindly provided by Yoshihiko Sako and Takuro Nunoura, Kyoto University, Kyoto, Japan. The genomic DNA samples were extracted by a standard method (45) and frozen at −20°C. Environmental archaeal rDNA clones of deep-sea hydrothermal vent euryarchaeotic groups 2 (pISA42), 6 (pISA48), and 7 (pISA16) and marine group I (pMC1A11) were obtained from various deep-sea hydrothermal vent environments (38) and were preserved at our institute.
Reaction mixtures were prepared in which the concentration of each oligonucleotide primer was 0.4 μM and the concentration of the rDNA template was 10 pg/μl. Two-step thermal cycling was performed by using the GeneAmp 9600 PCR system (Perkin-Elmer, Foster City, Calif.), and the conditions were as follows: denaturation at 96°C for 25 s and annealing and extension at every 1°C interval between 55 and 60°C for 360 s for a total of 25 cycles. The rDNA templates that were prepared contained equal proportions of each type of bacterial rDNA or equal proportions of each type of archaeal rDNA. When the prokaryotic universal PCR primer set was used, almost identical amplification efficiencies were obtained with either the bacterial or archaeal rDNA templates at all temperatures between 55 and 60°C. The archaeon-specific primer set provided the proper product from only archaeal rDNA templates at temperatures above 55°C, whereas the bacterium-specific primer set provided products from both bacterial and archaeal rDNA templates even at 60°C. These results indicated that the prokaryotic universal primer set that was designed effectively retrieved phylogenetically diverse prokaryotic rDNA and that the archaeon-specific primer set effectively detected only archaeal rDNA.
Using the prokaryotic universal and archaeon-specific primer sets and probes, the conditions for fluorogenic PCR were established with purified, representative bacterial and archaeal rDNAs as the templates. PCR and fluorescence signal monitoring were performed with the GeneAmp 5700 sequence detection system (PE Applied Biosystems). Reaction mixtures for the fluorogenic PCR were prepared in which the concentrations of each primer and the TaqMan probe were optimized (0.8 and 0.2 μM, respectively) and the concentration of the rDNA template was varied between 1 fg/μl and 20 pg/μl. The TaqMan Universal PCR Master Mix (PE Applied Biosystems), the LA Taq polymerase with GC buffer (TaKaRa), or the Ex Taq polymerase (TaKaRa) was used. All cycles began with 2 min at 50°C and then 10 min at 96°C in the case of the TaqMan Universal PCR Master Mix or 1 min at 96°C in the case of the LA Taq or Ex Taq polymerase system. Following these initial steps, two-step cycles of 25 s at 96°C and 6 min at 57 or 59°C were used for amplification of the prokaryotic rDNA or the archaeal rDNA. All assays were performed at least in triplicate. Post-PCR analysis was performed by using the GeneAmp 5700 SDS software (PE Applied Biosystems) and basically the analysis procedure described previously (5). Of the various Taq polymerase-buffer systems tested, the LA and Ex Taq polymerase systems provided more stable exponential fluorogenic amplification and more efficient amplification (lower threshold cycles at a given initial template concentration) than the TaqMan Universal PCR Master Mix. In addition, the minimum detection limit for the initial template concentration was lower with the LA Taq polymerase (approximately 1 fg of rDNA/μl) than with the Ex Taq polymerase or the TaqMan Universal PCR Master Mix (approximately 5 fg of rDNA/μl). Based on these results, the LA Taq polymerase system was selected for use in further experiments.
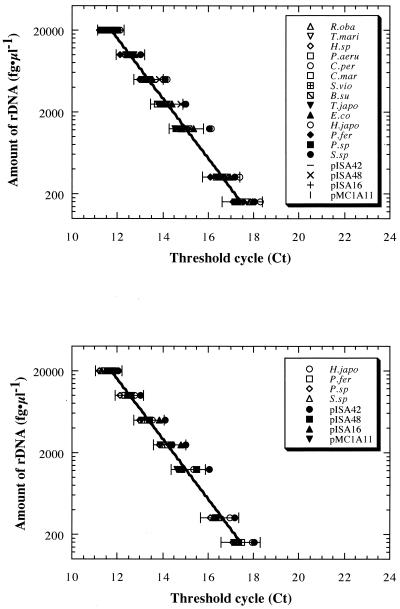
Fluorogenic amplification of SSU rDNA was tested by using different concentrations of rDNAs from representative bacterial and archaeal species as the templates (Fig. 1). All of the templates (10 bacterial and eight archaeal templates) gave reliable exponential fluorogenic amplification patterns (correlation coefficient, >0.999) dependent on the initial template concentration in the range from 100 fg of rDNA/μl to 20 pg of rDNA/μl when the universal primer set and probe were used (Fig. 1A). The archaeal primer set and probe also provided reliable fluorogenic amplification (correlation coefficient, >0.999) dependent on the initial template concentration in the range from 100 fg of rDNA/μl to 20 pg of rDNA/μl when any of the representative archaeal rDNA templates was used (Fig. 1B). The exponential fluorogenic amplification pattern obtained with each of the representative rDNAs when either the universal or archaeal primer set and probe were used was within the 95% confidence range of an average exponential fluorogenic amplification pattern, and the data were strongly correlated with each other (Fig. 1). With both the universal and archaeal systems, it was possible to detect fluorogenic amplification at the lowest template concentration (1 fg of rDNA/μl), but the reliability of quantification (correlation coefficient, <0.992) obtained with initial template concentrations below 100 fg of rDNA/μl was low (data not shown).
FIG. 1.
Fluorogenic amplification patterns dependent on homogeneous rDNA template concentrations, obtained by using a prokaryotic universal primer set and probe (A) or an archaeon-specific primer set and probe (B). Concentration-dependent changes in the threshold cycle were obtained in assays performed with rDNAs from representative bacterial and archaeal strains and environmental clones as the templates. All assays were performed at least in triplicate. The lines are exponential regression lines based on an average threshold cycle value obtained from all of the prokaryotic rDNA (A) and all of the archaeal rDNA (B) templates. The bars indicate the 95% confidence ranges of average regressions. Representative rDNAs were obtained from Rhodothermus obamensis JCM9785 (R.oba), Thermaerobacter marianensis JCM10246 (T.mari), Hydrogenobacter sp. strain JCM10560 (H.sp), Pseudomonas aeruginosa JCM5962 (P.aeru), Clostridium perfringens JCM1290 (C.per), Cytophaga marinoflava JCM8517 (C.mar), Shewanella violacea JCM10179 (S.vio), Bacillus subtilis JCM1465 (B.su), Thermosipho japonicus JCM10495 (T.japo), Escherichia coli INVα (E.co), Haloarcula japonica JCM7785 (H.japo), Palaeococcus ferrophilus JCM10417 (P.fer), Pyrobaculum sp. strain JCM10595 (P.sp), Sulfurisphaera sp. (S.sp), and environmental archaeal rDNA clones pISA42, pISA48, pISA16, and pMC1A11.
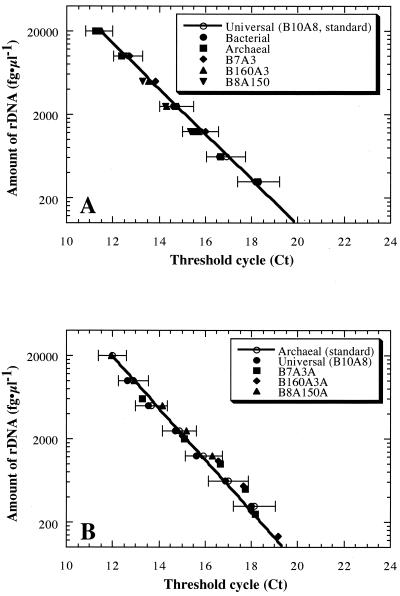
The detection and quantification of different proportions of archaeal rDNA in heterogeneous rDNA samples were examined (Fig. 2). Standard curves dependent on the initial template concentration were constructed by using a prokaryotic universal rDNA mixture (B10A8 mixture) containing equal amounts of 10 bacterial rDNAs and eight archaeal rDNAs for quantification of the prokaryotic rDNA and an archaeal rDNA mixture containing equal amounts of eight archaeal rDNAs for quantification of the archaeal rDNA. During preparation of the standard curves, samples with various initial template concentrations were always assayed in the same assay plate. The heterogeneous rDNA samples (bacterial rDNA mixture containing equal amounts of 10 bacterial rDNAs, archaeal rDNA mixture containing equal amounts of eight archaeal rDNAs, B7A3 mixture containing 30% archaeal rDNA, B160A3 mixture containing 1.8% archaeal rDNA, and B8A150 mixture containing 95% archaeal rDNA) were used as the templates, and the initial template concentrations were between 100 fg of rDNA/μl and 20 pg of rDNA/μl, a range which was within the range for reliable quantification. The universal primer set and probe resulted in stable fluorogenic amplification patterns that were almost identical for all of the rDNA mixtures (Fig. 2A). The archaeal primer set and probe provided archaeal rDNA-dependent fluorogenic amplification from heterogeneous rDNA samples, indicating that the threshold cycle was dependent on the archaeal rDNA concentrations in the rDNA mixtures (Fig. 2B). No fluorogenic amplification was observed with rDNA samples lacking archaeal rDNA. In any case, when either the universal or archaeal primer set and probe were used, the exponential fluorogenic amplification pattern was strongly correlated with the standard amplification pattern as it was within the 95% confidence range (Fig. 2). These results suggest that the fluorogenic PCR system using the archaeon-specific primers and probe which we designed can be used to detect and quantify archaeal rDNA in DNA samples extracted from heterogeneous, often unbalanced, naturally occurring microbial communities.
FIG. 2.
Fluorogenic amplification patterns dependent on heterogeneous rDNA template concentrations, obtained by using a prokaryotic universal primer set and probe (A) or an archaeon-specific primer set and probe (B). Concentration-dependent changes in the threshold cycle were obtained in assays by using heterogeneous rDNA mixtures as the templates. All assays were conducted at least in triplicate. The lines are standard curves obtained with the B10A8 mixture (A) and the archaeal rDNA mixture (B). The bars indicate the 95% confidence ranges of the standard curves. The heterogeneous rDNA samples (B10A8 mixture containing equal amounts of 10 bacterial and eight archaeal rDNAs, bacterial rDNA mixture containing equal amounts of 10 bacterial rDNAs, archaeal rDNA mixture containing equal amounts of eight archaeal rDNAs, B7A3 mixture containing 30% archaeal rDNA, B160A3 mixture containing 1.8% archaeal rDNA, and B8A150 mixture containing 95% archaeal rDNA) were used as the templates; the initial template concentrations were between 100 fg of rDNA/μl and 20 pg of rDNA/μl, a range which was within the reliable quantification range.
Comparison with dot blot hybridization analysis.
Dot hybridization analysis was performed with the heterogeneous rDNA samples described above in order to compare the stabilities and sensitivities of detection and quantification of the fluorogenic quantitative PCR method and the conventional dot blot hybridization analysis method. Dilution series were prepared for the universal rDNA (B10A8) mixture, bacterial rDNA mixture, archaeal rDNA mixture, B7A3 mixture, B160A3 mixture, and B8A150 mixture, and the diluted samples were denatured at 100°C for 10 min and cooled on ice. The denatured rDNA samples were dotted onto positively charged nylon membranes (Roche Diagnostics) and cross-linked to the membranes by exposure to 120 mJ of UV light energy by using a UV Stratalinker 1800 (Stratagene, Torrey Pines, Calif.). The oligonucleotide probes used were Uni1392R (5′-ACGGGCGGTGTGTRC-3′) (32) and Arch915R (5′-GTGCTCCCCCGCCAATTCCT-3′) (36), and these probes were conjugated at their 5′ ends to digoxigenin by the supplier (Amersham Pharmacia). The hybridization and wash conditions were empirically optimized for each probe. The optimal conditions were defined as the conditions that gave the strongest signal for the intended targets while minimizing cross-reactivity. Hybridization was conducted overnight in hybridization buffer (pH 7.0) containing 750 mM NaCl, 75 mM sodium citrate, 0.02% (wt/vol) sodium dodecyl sulfate (SDS), 0.1% (wt/vol) sodium lauroylsarcosine, and 2% (wt/vol) blocking reagent (Roche Diagnostics) at the optimized temperature (42°C for the universal probe and 54°C for the archaeal probe). After hybridization, the filters were washed twice with buffer (pH 7.0) containing 300 mM NaCl, 30 mM sodium citrate, and 0.1% SDS at room temperature for 5 min and then twice with buffer (pH 7.0) containing 15 mM NaCl, 1.5 mM sodium citrate, and 0.1% SDS at same temperatures used for hybridization for 30 min. Probe-target hybrids were detected with a DIG luminescent detection kit (Roche Diagnostics), and the chemical luminescence signal was developed and quantified by using NIH Image, version 1.62.
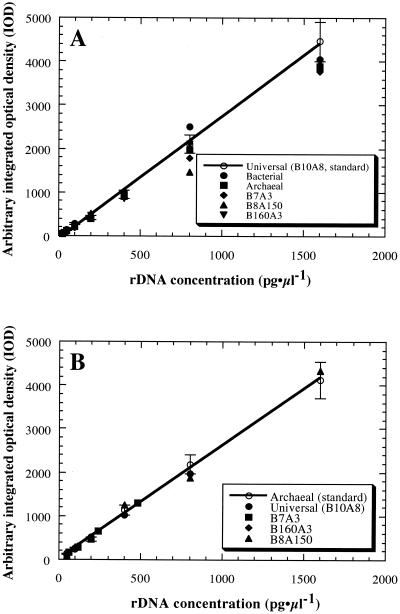
The hybridization pattern obtained with prokaryotic or archaeal rDNA when the universal or archaeal probes were used was stable at rDNA concentrations ranging from 50 pg of rDNA/μl to 1.5 ng of rDNA/μl, while several patterns varied from the 95% confidence range of the standard pattern (Fig. 3). At concentrations below 50 pg of rDNA/μl, no stable hybridization signal was obtained with either the universal or archaeal probe. When we compared the invariability of quantification and the detection limit obtained with the fluorogenic quantitative PCR and dot blot hybridization analysis methods, the fluorogenic PCR method performed with the TaqMan probes was more effective for detection and quantification of rDNA than the dot blot hybridization analysis method under the conditions examined in this study.
FIG. 3.
Hybridization patterns dependent on rDNA concentrations, obtained by using a prokaryotic universal oligonucleotide probe (A) or an archaeon-specific oligonucleotide probe (B). Concentration-dependent changes in hybridization signal strength were obtained in assays in which heterogeneous rDNA mixtures were used. All assays were conducted at least in triplicate. The lines are standard curves obtained from the B10A8 mixture (A) and the archaeal rDNA mixture (B). The bars indicate the 95% confidence ranges of the standard curves. The heterogeneous rDNA samples (B10A8 mixture, bacterial rDNA mixture, archaeal rDNA mixture, B7A3 mixture, B160A3 mixture, and B8A150 mixture) were used, and the rDNA concentrations were between 50 pg of rDNA/μl and 1.5 ng of rDNA/μl, a range which was within the reliable quantification range.
Application to naturally occurring microbial communities.
The fluorogenic PCR method using the prokaryotic universal and archaeon-specific primer sets and probes was used in an analysis of DNA samples extracted from various naturally occurring microbial communities. The samples used for DNA extraction were hot spring water and sediment samples obtained from Goshogake Hot Spring (GHS) (85°C, pH 5.0) and Fukeyu Hot Spring (FHS) (85°C, pH 4.1) in Akita prefecture, Japan (K. Takai, A. Inoue, and K. Horikoshi, Shinkai Symp. 1998, p. 52, 1998), effluent vent water from a deep-sea hydrothermal vent in Iheya Basin (IVW) (38), freshwater sediment from a depth of 15 cm in Takatori Creek (TC), and sediment from a depth of 15 cm in Nojima Estuary (NE) near the Japan Marine Science & Technology Center. The microbial community structures and the archaeal rDNA compositions of the samples from GHS, FHS, and IVW were examined previously by a PCR-mediated sequencing analysis of rDNA clone libraries constructed with the Uni515F and Uni1408R (42) primers and the Arch21F (12) and 1492R (21) primers (38; Takai et al., Shinkai Symp. 1998). The universal PCR primer libraries obtained from GHS and FHS consisted of only archaeal rDNA clones, and no bacterial rDNA was obtained (44 of 44 clones and 56 of 56 clones, respectively, were hyperthermophilic crenarchaeotic rDNA sequences) (Takai et al., Shinkai Symp. 1998). The archaeal PCR primer libraries from both hot spring samples contained rDNA clones phylogenetically associated with the orders Thermoproteales, Igneococcales, and Sulfolobales. Likewise, partial sequencing of rDNA clones in the universal PCR primer library obtained from IVW revealed that the archaeal rDNA clones comprised about 30% of the total rDNA community (38). For the sediment samples from TC and NE, the total cell density and the autofluorescent cell density (including F420-dependent autofluorescent methanogens) were determined by epifluorescence microscopy with and without DAPI (4′,6′-diamidino-2-phenylindole) staining, respectively. Most of the autofluorescent cells were found to form sarcinalike structures that were typically observed in methanogenic archaeal members of the family Methanosarcinaeae (8), and some were long straight rods. The proportions of autofluorescent cells were about 1.4 and 9.6% of the total cells in the sediments from TC and NE, respectively.
Nucleic acids were extracted and purified from the GHS, FHS, and IVW samples as described previously (38; Takai et al., Shinkai Symp. 1998). Nucleic acids were extracted from the TC and NE sediments by using an Ultraclean Soil DNA MegaPrep kit (MO BIO Laboratories, Inc., Solana Beach, Calif.). Approximately 5 g of wet sediment was used for DNA extraction, performed by using the manufacturer's suggested protocol. Although the DNA samples obtained with the kit contained little contaminating RNA, 0.02% (wt/vol) RNase A (Sigma) was added to the samples, and the contaminating RNA was degraded at 37°C for 3 h. The mixtures were extracted with an equal volume of phenol saturated with 100 mM Tris-HCl (pH 8.0), followed by extraction with phenol-chloroform-isoamyl alcohol (24:24:1, vol/vol/vol) and chloroform-isoamyl alcohol (24:1, vol/vol). DNA was precipitated from the solutions by adding a 3 volumes of ethanol and was recovered by centrifugation.
Using the purified DNA samples as templates, detection and quantification of the archaeal rDNA were conducted under the optimized conditions described above. A dilution series for each of the DNA samples and the samples were assayed by using the universal rDNA (B10A8) mixture and the archaeal rDNA mixture as the standards for quantification of whole microbial rDNA and archaeal rDNA, respectively (Table 1). The initial rDNA template concentrations in the environmental DNA samples were between 20 fg of rDNA/μl and 20 pg of rDNA/μl, a range chosen based on calculation of the reliable quantification range. As expected from the results of previous studies (38; Takai et al., Shinkai Symp. 1998), the fluorogenic PCR quantification results indicated that the amounts of archaeal rDNA were equivalent to the amounts of total prokaryotic rDNA in the samples from the hot springs and that about 30% of the prokaryotic rDNA in the deep-sea hydrothermal vent water was derived from members of the archaeal group (Table 1). These results indicate that the fluorogenic PCR method is effective for detecting and quantifying the proportions of archaeal rDNA in whole microbial rDNA assemblages obtained from naturally occurring microbial communities. Not only in the extreme environments where archaea were relatively abundant but also in nonextreme environments, the fluorogenic PCR method successfully detected archaeal rDNA (Table 1). In freshwater and estuarial anoxic sediments, the most abundant members of the archaeal group are considered to be methanogens. In fact, observation by epifluorescence microscopy indicated the presence of cells displaying autofluorescence and a typical pseudosarcina structure, which were likely distinctive morphological characteristics of members of the Methanosarcinaceae (8), at levels proportional to the number of cells stained by DAPI (ca. 1% of the total cells in the freshwater sediment and ca. 10% of the total cells in the estuarial sediment). The proportion of archaeal rDNA in the total prokaryotic rDNA as determined by fluorogenic PCR matched the proportion of autofluorescent cells presumably consisting of methanogenic archaea in the sediment samples (Table 1). These results suggested that the fluorogenic PCR could be used to detect and quantify archaeal rDNA in DNA samples obtained not only from environments where thermophilic archaea are abundant but also from environments where methanogenic archaea are abundant.
TABLE 1.
Quantification of archaeal rDNA in DNA samples obtained from various naturally occurring microbial communities
| Sample | Temp (°C) | pH | Depth | Amt used for extraction | Cell densitya | DNA yieldb | Concn of prokaryotic universal rDNA | Concn of archaeal rDNA | Proportion of archaeal rDNA (%)c | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| Water and sediments from GHS | 85 | 5.0 | 800 g (wet wt) | 106 cells/g (wet wt) | 3.8 ng/g (wet wt) | 69.4 ± 10.0 fg/g (wet wt) | 73.4 ± 10.3 fg/g (wet wt) | 106 ± 35 | Takai et al.d | |
| Water and sediments from FHS | 85 | 4.1 | 900 g (wet wt) | 106 cells/g (wet wt) | 3.9 ng/g (wet wt) | 44.8 ± 0.7 fg/g (wet wt) | 43.8 ± 0.8 fg/g (wet wt) | 97.8 ± 3.3 | Takai et al.d | |
| Vent water from IVW | ∼300 | 972 m | 12 liters | 107 cells/liter | 12.5 ng/liter | 12.5 ± 3.5 fg/liter | 4.2 ± 0.7 fg/liter | 33.6 ± 15.8 | 38 | |
| Sediments from TC | 15–20 | 15 cm | 5 g (wet wt) | 109 cells/g (wet wt) | 6.5 μg/g (wet wt) | 4,460 ± 840 fg/g (wet wt) | 42.0 ± 6.0 fg/g (wet wt) | 0.94 ± 0.18 | This study | |
| Sediments from NE | 15–20 | 15 cm | 5 g (wet wt) | 109 cells/g (wet wt) | 3.5 μg/g (wet wt) | 1,160 ± 140 fg/g (wet wt) | 152 ± 24 fg/g (wet wt) | 13.1 ± 4.2 | This study |
Direct cell count as determined with DAPI.
DNA concentrations were measured with a spectrophotometer.
Proportion of archaeal rDNA in whole prokaryotic universal rDNA.
Takai et al., Shinkai Symp. 1998.
In conclusion, the quantitative PCR method established in the present study by using prokaryotic universal and archaeon-specific primers and probes was probably more sensitive (able to detect a lower concentration of rDNA) and more invariable (stable and able to quantify the archaeal rDNA) than the conventional dot blot hybridization analysis method targeting rRNA genes, and it can be readily applied to large numbers of samples. In this study, we sought to design domain-specific primers and a domain-specific probe for bacterial rDNA at the same site as the archaeon-specific primers and probe. Although the bacterium-specific primers which we designed were not fully selective for bacterial rDNA, strongly selective bacterium-specific primers could be constructed with different binding sites within the SSU rDNA. By using a combination of universal, bacterial, and archaeal primer and probe sets, quantification of members of microbial communities based on rDNA structures should be further improved. Fragmentation of the microbial chromosomal DNA, contamination by organic or inorganic substances, and contamination by eukaryotic DNA might influence fluorogenic amplification. However, it seems likely that improvements in methods of extraction of DNA from environmental samples, such as elimination of eukaryotic cells by size by prefiltration and use of an optimized technique for extraction of DNA (27), might reduce the effects of such negative factors on reliable quantification by the fluorogenic PCR method. Using the fluorogenic PCR method in combination with other culture-independent molecular techniques should facilitate elucidation of the structures, functions, and interactions of natural microbial communities.
ACKNOWLEDGMENTS
We thank Yoshihiko Sako, Norimichi Nomura, and Takuro Nunoura for kindly providing samples of purified genomic DNA of several archaeal strains and Wayne R. Bellamy for editing English usage in the manuscript.
This work was supported in part by a domestic research fellowship provided by the Japan Science and Technology Corporation.
REFERENCES
- 1.Altschul S F, Madden T L, Schäffer A A, Zhang J, Zhang Z, Miller W, Lipman D J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Amann R I, Springer N, Ludwig W, Gortz H D, Schleifer K H. Identification in situ and phylogeny of uncultured bacterial endosymbionts. Nature. 1991;351:161–164. doi: 10.1038/351161a0. [DOI] [PubMed] [Google Scholar]
- 3.Barns S M, Fundyga R E, Jeffries M W, Pace N R. Remarkable archaeal diversity detected in a Yellowstone National Park hot spring environment. Proc Natl Acad Sci USA. 1994;91:1609–1613. doi: 10.1073/pnas.91.5.1609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Barns S M, Delwiche C F, Palmer J D, Pace N R. Perspectives on archaeal diversity, thermophily and monophyly from environmental rRNA sequences. Proc Natl Acad Sci USA. 1996;93:9188–9193. doi: 10.1073/pnas.93.17.9188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bassler H A, Flood S J A, Livak K J, Marmaro J, Knorr R, Batt C A. Use of a fluorogenic probe in a PCR-based assay for detection of Listeria monocytogenes. Appl Environ Microbiol. 1995;61:3724–3728. doi: 10.1128/aem.61.10.3724-3728.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Benson D A, Boguski M S, Lipman D J, Ostell J, Ouellette B F F. GenBank. Nucleic Acids Res. 1998;26:1–7. doi: 10.1093/nar/26.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bintrim S B, Donohue T J, Handelsman J, Robert G P, Goodman R M. Molecular phylogeny of Archaea from soil. Proc Natl Acad Sci USA. 1997;94:277–282. doi: 10.1073/pnas.94.1.277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Boone D R, Mah R A. Group I. Methanogenic archaeaobacteria. In: Staley J T, Bryant M P, Pfennig N, Holt J G, editors. Bergey's manual of systematic bacteriology. Vol. 3. Baltimore, Md: The Williams & Wilkins Co.; 1989. pp. 2173–2216. [Google Scholar]
- 9.Brunk C F, Eis N. Quantitative measure of small-subunit rRNA gene sequences of the kingdom Korarchaeota. Appl Environ Microbiol. 1998;64:5064–5066. doi: 10.1128/aem.64.12.5064-5066.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Buckley D H, Graber J R, Schmidt T M. Phylogenetic analysis of nonthermophilic members of the kingdom Crenarchaeota and their diversity and abundance in soils. Appl Environ Microbiol. 1998;64:4333–4339. doi: 10.1128/aem.64.11.4333-4339.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chin K J, Lukow T, Conrad R. Effect of temperature on structure and function of the methanogenic archaeal community in an anoxic rice field soil. Appl Environ Microbiol. 1999;65:2341–2349. doi: 10.1128/aem.65.6.2341-2349.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Delong E F. Archaea in coastal marine environments. Proc Natl Acad Sci USA. 1992;89:5685–5689. doi: 10.1073/pnas.89.12.5685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Delong E F, Taylor L T, Marsh T L, Preston C M. Visualization and enumeration of marine planktonic archaea and bacteria by using polyribonucleotide probes and fluorescent in situ hybridization. Appl Environ Microbiol. 1999;65:5554–5563. doi: 10.1128/aem.65.12.5554-5563.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Delong E F, Wu K Y, Prezelin B B, Jovine R V M. High abundance of Archaea in Antarctic marine picoplancton. Nature. 1994;371:695–697. doi: 10.1038/371695a0. [DOI] [PubMed] [Google Scholar]
- 15.Desjardin L E, Chen Y, Perkins M D, Teixeira L, Cave M D, Eisenach K D. Comparison of the ABI 7700 system (TaqMan) and competitive PCR for quantification of IS6110 DNA in sputum during treatment of tuberculosis. J Clin Microbiol. 1998;36:1964–1968. doi: 10.1128/jcm.36.7.1964-1968.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fuhrman J A, McCallum K, Davis A A. Novel major archaebacterial group from marine plankton. Nature. 1992;356:148–149. doi: 10.1038/356148a0. [DOI] [PubMed] [Google Scholar]
- 17.Giovannoni S J, Britschgi T B, Moyer C L, Field F G. Genetic diversity of Sargasso Sea bacterioplankton. Nature. 1990;345:60–65. doi: 10.1038/345060a0. [DOI] [PubMed] [Google Scholar]
- 18.Hershberger K L, Barns S M, Reysenbach A L, Dawson S C, Pace N R. Wide diversity of Crenarchaeota. Nature. 1996;384:420. doi: 10.1038/384420a0. [DOI] [PubMed] [Google Scholar]
- 19.Hinrichs K U, Hayes J M, Sylva S P, Brewer P G, Delong E F. Methane-consuming archaebacteria in marine sediments. Nature. 1999;398:802–805. doi: 10.1038/19751. [DOI] [PubMed] [Google Scholar]
- 20.Kim Y R, Czajka J, Batt C A. Development of a fluorogenic probe-based PCR assay for detection of Bacillus cereus in nonfat dry milk. Appl Environ Microbiol. 2000;66:1453–1459. doi: 10.1128/aem.66.4.1453-1459.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lane D J. 16S/23S sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. New York, N.Y: John Wiley & Sons Ltd.; 1985. pp. 115–176. [Google Scholar]
- 22.Larsen N, Olsen G J, Maidak B L, McCaughey M J, Overbeek R, Macke T J, Marsh T L, Woese C R. The Ribosomal Database Project. Nucleic Acids Res. 1993;21:3021–3023. doi: 10.1093/nar/21.13.3021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lie S Y, Petropoulos C J. Advances in quantitative PCR technology: 5′ nuclease assays. Curr Opin Biotechnol. 1998;9:43–48. doi: 10.1016/s0958-1669(98)80082-7. [DOI] [PubMed] [Google Scholar]
- 24.Livak K J, Flood S J A, Marmaro J, Giusti W, Deetz K. Oligonucleotides with fluorescent dyes at opposite ends provide a quenched probe system useful for detecting PCR product and nucleic acid hybridization. PCR Methods Appl. 1995;4:357–362. doi: 10.1101/gr.4.6.357. [DOI] [PubMed] [Google Scholar]
- 25.MacGregor B J, Moser D P, Alm E W, Nealson K H, Stahl D A. Crenarchaeota in Lake Michigan. Appl Environ Microbiol. 1997;63:1178–1181. doi: 10.1128/aem.63.3.1178-1181.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Martell M, Gomez J, Esteban J I, Sauleda S, Quer J, Cabot B, Esteban R, Guardia J. High-throughput real-time reverse transcription-PCR quantitation of hepatitis C virus RNA. J Clin Microbiol. 1999;37:327–332. doi: 10.1128/jcm.37.2.327-332.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Miller D N, Bryant J E, Madsen E L, Ghiorse W C. Evaluation and optimization of DNA extraction and purification procedures for soil and sediment samples. Appl Environ Microbiol. 1999;65:4715–4724. doi: 10.1128/aem.65.11.4715-4724.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Morris T, Robertson B, Gallagher M. Rapid reverse transcription-PCR detection of hepatitis C virus RNA in serum by using the TaqMan fluorogenic detection system. J Clin Microbiol. 1996;34:2933–2936. doi: 10.1128/jcm.34.12.2933-2936.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nogi Y, Kato C, Horikoshi K. Taxonomic studies of deep-sea barophilic Shewanella strains and description of Shewanella violacea sp. nov. Arch Microbiol. 1998;170:331–338. doi: 10.1007/s002030050650. [DOI] [PubMed] [Google Scholar]
- 30.Norton D M, Batt C A. Detection of viable Listeria monocytogenes with a 5′ nuclease PCR assay. Appl Environ Microbiol. 1999;65:2122–2127. doi: 10.1128/aem.65.5.2122-2127.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Oberst R D, Hays M P, Bohra L K, Phebus R K, Yamashiro C T, Paszko-Kolva C, Flood S J A, Sargeant J M, Gillespie J R. PCR-based DNA amplification and presumptive detection of Escherichia coli O157:H7 with an internal fluorogenic probe and the 5′ nuclease (TaqMan) assay. Appl Environ Microbiol. 1998;64:3389–3396. doi: 10.1128/aem.64.9.3389-3396.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Olsen G J, Lane D J, Giovannoni S J, Pace N R, Stahl D A. Microbial ecology and evolution: a ribosomal RNA approach. Annu Rev Microbiol. 1986;40:337–365. doi: 10.1146/annurev.mi.40.100186.002005. [DOI] [PubMed] [Google Scholar]
- 33.Piatak M, Jr, Luk K C, Williams B, Lifson J D. Quantitative competitive polymerase chain reaction for accurate quantification of HIV DNA and RNA species. BioTechniques. 1993;14:70–80. [PubMed] [Google Scholar]
- 34.Preston C M, Wu K Y, Molinski T F, Delong E F. A psychrophilic crenarchaeon inhabits a marine sponge: Cenarchaeum symbiosum gen. nov., sp. nov. Proc Natl Acad Sci USA. 1996;93:6241–6246. doi: 10.1073/pnas.93.13.6241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Sako Y, Takai K, Uchida A, Ishida Y, Katayama Y. Rhodothermus obamensis sp. nov., a modern lineage of extremely thermophilic marine bacterium. Int J Syst Bacteriol. 1996;46:1099–1104. doi: 10.1099/00207713-46-4-1099. [DOI] [PubMed] [Google Scholar]
- 36.Stahl D A, Amann R. Development and application of nucleic acid probes in bacterial systematics. In: Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. New York, N.Y: John Wiley & Sons Ltd.; 1991. pp. 205–248. [Google Scholar]
- 37.Swan D C, Tucker R A, Holloway B P, Icenogle J P. A sensitive, type-specific, fluorogenic probe assay for detection of human papillomavirus DNA. J Clin Microbiol. 1997;35:886–891. doi: 10.1128/jcm.35.4.886-891.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Takai K, Horikoshi K. Genetic diversity of archaea in deep-sea hydrothermal vent environments. Genetics. 1999;152:1285–1297. doi: 10.1093/genetics/152.4.1285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Takai K, Inoue A, Horikoshi K. Thermaerobacter marianensis gen. nov., sp. nov., an aerobic extremely thermophilic marine bacterium from the 11,000 m deep Mariana Trench. Int J Syst Bacteriol. 1999;49:619–628. doi: 10.1099/00207713-49-2-619. [DOI] [PubMed] [Google Scholar]
- 40.Takai K, Horikoshi K. Molecular phylogenetic analysis of archaeal intron-containing genes coding for rRNA obtained from a deep-subsurface geothermal water pool. Appl Environ Microbiol. 1999;65:5586–5589. doi: 10.1128/aem.65.12.5586-5589.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Takai K, Horikoshi K. Thermoshipho japonicus sp. nov., an extremely thermophilic bacterium isolated from a deep-sea hydrothermal vent in Japan. Extremophiles. 2000;4:9–17. doi: 10.1007/s007920050002. [DOI] [PubMed] [Google Scholar]
- 42.Takai K, Sako Y. A molecular view of archaeal diversity in marine and terrestrial hot water environments. FEMS Microbiol Ecol. 1999;28:177–188. [Google Scholar]
- 43.Takai K, Sugai A, Itoh T, Horikoshi K. Palaeococcus ferrophilus gen. nov., sp. nov., a barophilic hyperthermophilic archaeon from a deep-sea hydrothermal vent chimney. Int J Syst Evol Microbiol. 2000;50:489–500. doi: 10.1099/00207713-50-2-489. [DOI] [PubMed] [Google Scholar]
- 44.Vetriani C, Jannasch H W, MacGregor B J, Stahl D A, Reysenbach A-L. Population structure and phylogenetic characterization of marine benthic Archaea in deep-sea sediments. Appl Environ Microbiol. 1999;65:4375–4384. doi: 10.1128/aem.65.10.4375-4384.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wilson K. Preparation of genomic DNA from bacteria. In: Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Short protocols in molecular biology. New York, N.Y: John Wiley & Sons; 1992. pp. 2.10–2.12. [Google Scholar]