Abstract
Despite the strong prognostic stratification of circulating tumor cells (CTCs) enumeration in metastatic breast cancer (MBC), current clinical trials usually do not include a baseline CTCs in their design. This study aimed to generate a classifier for CTCs prognostic simulation in existing datasets for hypothesis generation in patients with MBC. A K-nearest neighbor machine learning algorithm was trained on a pooled dataset comprising 2436 individual MBC patients from the European Pooled Analysis Consortium and the MD Anderson Cancer Center to identify patients likely to have CTCs ≥ 5/7 mL blood (StageIVaggressive vs StageIVindolent). The model had a 65.1% accuracy and its prognostic impact resulted in a hazard ratio (HR) of 1.89 (Simulatedaggressive vs SimulatedindolentP < .001), similar to patients with actual CTCs enumeration (HR 2.76; P < .001). The classifier’s performance was then tested on an independent retrospective database comprising 446 consecutive hormone receptor (HR)-positive HER2-negative MBC patients. The model further stratified clinical subgroups usually considered prognostically homogeneous such as patients with bone-only or liver metastases. Bone-only disease classified as Simulatedaggressive had a significantly worse overall survival (OS; P < .0001), while patients with liver metastases classified as Simulatedindolent had a significantly better prognosis (P < .0001). Consistent results were observed for patients who had undergone CTCs enumeration in the pooled population. The differential prognostic impact of endocrine- (ET) and chemotherapy (CT) was explored across the simulated subgroups. No significant differences were observed between ET and CT in the overall population, both in terms of progression-free survival (PFS) and OS. In contrast, a statistically significant difference, favoring CT over ET was observed among Simulatedaggressive patients (HR: 0.62; P = .030 and HR: 0.60; P = .037, respectively, for PFS and OS).
Keywords: clinical trial model, machine learning, liquid biopsy, biomarker, K-nearest neighbor
Current clinical trials usually do not include a baseline circulating tumor cells (CTCs) enumeration in the trial design. This article reports a classifier to simulate the prognostic stratification of CTCs in existing datasets for hypothesis generation in metastatic breast cancer scenarios. This classifier has the potential to inform and potentially drive future clinical trial design.
Implications for Practice.
Circulating tumor cells (CTC)-based risk stratification may have a role for future treatment strategies, as it enables the selection of subgroups with differential response potential. Machine learning can simulate CTCs-based staging in scenarios of particular interest to identify subpopulations for hypothesis generation which may benefit from higher intensity treatments due to a more aggressive outcome and inform future clinical trials designs.
Background
Metastatic disease occurs in approximately 20%-50% of patients with early breast cancer (BC) and in 6%-10% of newly diagnosed BC cases. Different disease subtypes account, at least partially, for the variability in overall survival (OS) which can range from months to several years.1,2 As increasing knowledge is generated regarding new therapeutic agents, it is important to identify new predictive factors that help treatment selection. To date, the most established predictive markers in metastatic breast cancer (MBC) are the expression of HR and HER2. Patients with HR-positive/HER2-negative MBC often respond to endocrine therapy (ET) alone or in combination with targeted agents with generally fewer side effects and toxicities than chemotherapy (CT). ET-based therapies in combination with CDK4/6 inhibitors are therefore the preferred treatment approach in most cases of HR-positive MBC, reserving CT for patients with extensive symptomatic visceral disease and/or defined endocrine resistance. To date, there are no predictive biomarkers driving treatment choice regarding targeted therapies such as inhibitors of cyclin-dependent kinase (CDK) 4/6.
While circulating tumor cells (CTCs) are recognized as an independent prognostic marker for OS, their role in supporting clinical management of MBC is still not well defined.3-5 A previous effort to prospectively evaluate the clinical utility of CTCs enumeration in MBC was performed in the SWOG 0500 trial. In this study, clinicians were guided to maintain or switch chemotherapy regimen based on an early CTCs evaluation after 21 days of therapy.6 Although the study showed no OS differences in patients with persistently elevated CTCs that changed CT regimen, the prognostic potential of CTCs was further confirmed. The study’s sampling timeframe, however, was not strictly biology driven, since CTCs dynamics is not just treatment-induced but also likely the result of tumor biology evolution.7 Moreover, the CT selection was driven by the clinician’s choice and not by biology-defined targets.
The strong prognostic stratification achieved by CTCs enumeration can have potential applications in identifying patients that will likely benefit from intensive treatments, while reserving less toxic treatments for those with an inherently indolent disease.8
Current clinical trials usually do not include a baseline CTCs enumeration in their design, primarily due its perceived costs and technical complexity, notwithstanding software solutions that minimize inter-operator variability. Therefore, there is an unmet need to evaluate biomarker-assisted decisional algorithms.9
The aim of this study was to generate a classifier to simulate the prognostic stratification of CTCs in existing datasets for hypothesis generation in precise MBC scenarios. This classifier has the potential to inform and potentially drive future clinical trials design.
Methods
Study Population and Ethical approval
The model was trained on a pooled dataset based on data from 2436 individual MBC patients from 17 European Centers participating in the European Pooled Analysis Consortium (EPAC) and the MD Anderson Cancer Center (MDACC).5 The database characteristics were previously published.4
The anonymized data were transferred to the Robert H. Lurie Comprehensive Cancer Center-Bioinformatics Core Facility. A retrospective Institutional Review Board-approved protocol was used to access and analyze the data. CTCs enumeration was performed through the CellSearch platform (Menarini Silicon Biosystems, PA) and the patients were classified accordingly based on a 5 CTCs per 7.5 mL cut off in Stage IVaggressive (5 ≥ CTCs) and Stage IVindolent (5 < CTCs).3,4
Features Selection and Training of the Machine Learning Classifier
K-nearest neighbor (KNN) is a supervised machine learning algorithm that can be used to solve both classification and regression problems. After a training phase during which labeled data are analyzed, the model abstracts a function that can be used to infer an appropriate output when new unlabeled data are evaluated. The KNN algorithm classifies objects based on their proximity in the feature space through a majority vote of its neighbors. The object is therefore assigned to the class that is most common among its KNN, where K is a positive integer that is typically small. If K = 1, then the object is assigned to the class of its nearest neighbor.
Baseline features linked to tumor biology were tested through Chi square test for association with respect to Stage IVaggressive and consistently selected based on both clinical and statistical significance (Table 1).
Table 1.
Patients’ characteristics across the CTCs enumeration subgroups.
| Variable | Stage IVindolent | Stage IVaggressive | P-value |
|---|---|---|---|
| ER | .001 | ||
| Negative | 372 (3.22%) | 244 (23.99%) | |
| Positive | 859 (69.78%) | 773 (76.01%) | |
| PR | .076 | ||
| Negative | 553 (44.92%) | 419 (41.20%) | |
| Positive | 678 (55.08%) | 598 (58.80%) | |
| HER2 | <.001 | ||
| Negative | 896 (72.79%) | 808 (79.45%) | |
| Positive | 335 (27.21%) | 209 (20.55%) | |
| Bone involvement | <.001 | ||
| No | 557 (45.25%) | 222 (21.83%) | |
| Yes | 674 (54.75%) | 795 (78.17%) | |
| Liver involvement | <.001 | ||
| No | 818 (66.45%) | 502 (49.36%) | |
| Yes | 413 (33.55%) | 515 (50.64%) | |
The Stage IVindolent and Stage IVaggressive subgroups are characterized by significantly different characteristics both in terms of tumor biology and clinical behavior.
Abbreviations: ER, estrogen receptor; PR, progesterone receptor.
The model was then trained based on estrogen receptor (ER) status (positive vs negative, 1% threshold), progesterone receptor (PR) status (positive vs negative, 1% threshold), HER2 status (positive vs negative), treatment line (continuous variable), bone and liver involvement (yes vs no).
Patients with all the necessary features (2248) were then 3:1 randomly assigned to a training set (1687) and a validation set (561) (Supplementary Fig. 1).
The model was built using R (The R foundation for Statistical Computing. version 3.3.1) and the “caret” package.10
CTCs Simulation on an Independent Database
The classifiers’ performance was tested on an independent retrospective database of 446 consecutive HR-positive HER2-negative MBC patients treated with first-line ET or CT at the University Hospitals of Naples and Udine, Italy, between 2004 and 2014. Patients’ characteristics were previously published.11 This study was previously approved by the Review Committees of each center. CTCs risk stratification was simulated through the “caret” package.10
Statistical Analysis
Categorical variables were reported as frequency distribution, whereas continuous variables were described through median and interquartile range (IQR).
Overall survival (OS) was defined as the time from baseline CTCs enumeration to death from any cause or date of last follow-up. Progression-free survival (PFS) was defined as the time from baseline CTCs enumeration to disease progression (according to RECIST criteria) or death from any cause or date of last follow-up. Censoring was applied to patients without an endpoint event at the last follow-up visit. Survival was represented by Kaplan-Meier estimator plot and analyzed by log-rank test and by uni- and multivariable Cox regression models.
Changes in the predictive power of the Cox regression models using the simulated CTCs enumeration were investigated through Harrell’s c concordance statistics.
Differences in distribution of CTCs enumeration, according to the simulated CTCs status across MBC subtypes, were tested through the Mann-Whitney U test.
Statistical analysis was conducted using StataCorp 2016 Stata Statistical Software: Release 15.1 (College Station, TX, USA), and R (The R foundation for Statistical Computing. version 3.3.1).
Data Availability
The datasets supporting the conclusions of this article are available from the corresponding author on reasonable request.
Results
From the initial cohort of 2436 patients, 2248 (92.4%) had no missing data and were therefore eligible for the model training (Table 1).4 Consistent with previously reported data, CTCs enumeration was associated with specific baseline characteristics. In particular, Stage IVaggressive patients were more likely to be ER positive (P = .001), HER2 negative (P < .001), and have bone or liver metastasis (P < .001).
CTCs Enumeration Can Be Simulated Through Machine Learning
Based on these premises, a KNN model was trained with a resulting 65.1% accuracy (95% CI [CI]: 61.0%-69.0%), 72.6% sensitivity (95%CI: 68.95-76.33%) and a 55.9% specificity (95%CI: 51.8-60.0%) (Supplementary Table S1).
The proportion of correctly classified observations was higher in patients without detectable CTCs (78.3% classified as Simulated IVindolent) than in patients with CTCs enumeration higher than the 75th percentile (21 CTCs; 62.6% classified as Simulatedaggressive).
In the validation cohort, the prognostic impact of the CTCs enumeration was hazard ratio (HR) 2.76 (95%CI 2.18-3.49; P < .001) for Stage IVaggressive vs Stage IVindolent. Consistently, the classifier resulted in a HR of 1.89 (95%CI 1.50-2.38; P < .001) for Simulatedaggressive vs Simulated IVindolent (Fig. 1).
Figure 1.
Comparison between the CTC-based risk stratification (Stage IVindolent vs Stage IVaggressive) and the KNN simulation (Simulatedindolent vs Simulatedaggressive). The model was capable to simulate a comparable risk stratification with respect to CTCs enumeration both for StageIVindolent (CTCs vs Simulation HR 1.18, 95%CI 0.93-1.51, P = .177) and StageIVaggressive (CTCs vs Simulation HR 0.88, 95%CI 0.70-1.09, P = .242).
Patients classified as Simulatedaggressive had a significantly higher number of CTCs with respect to Simulated IVindolent (median CTCs 11.5, IQR: 1-44 vs 2, IQR 0-9; P = .0001, respectively, for Simulatedaggressive and Simulated IVindolent).
CTCs Classifier and Breast Cancer Subtypes
The classifier’s performance was then tested in the overall population across different MBC subtypes.
Patients classified as Simulatedaggressive had a significantly higher CTCs enumeration with respect to Simulated IVindolent in HR-positive MBC (median CTCs 10, IQR: 1-60.5 and 1, IQR 0-11; P < .0001, respectively) (Fig. 2A), HER2-positive MBC (median CTCs 8, IQR: 1-32 and 1, IQR 0 – 6; P = .0091, respectively) (Fig. 2B) and TNBC (median CTCs 11, IQR: 2-52 and 2, IQR 0 – 16; P = .0403, respectively) (Fig. 2C). Patients classified as Simulatedaggressive experienced a significantly worse prognosis (Fig. 2D, E), especially in the HR-positive subgroup (Fig. 2D).
Figure 2.
CTCs enumeration simulation across MBC subtypes. Patients classified as Simulatedaggressive (Agg) had a significantly higher CTCs enumeration with respect to Simulated IVindolent (Ind) in HR-positive MBC (A), HER2-positive MBC (B) and TNBC (C). Patients classified as Simulatedaggressive experienced a significantly worse prognosis (D).
CTC-Based Risk Stratification Can Be Simulated on an Independent Real-World Dataset
To test the consistency and applicability of the classifier in a real-world MBC cohort, a proof-of-concept analysis was performed on an independent database comprising 446 HR-positive, HER2-negative MBC patients. Patients’ characteristics were previously published. Three patients were excluded from the analysis due to missing PR status.11
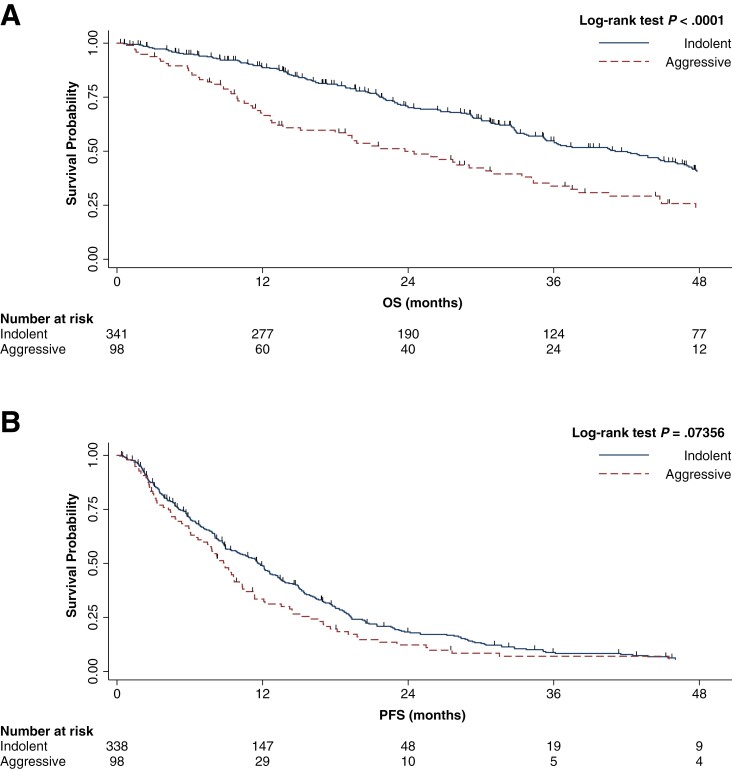
Of the 443 eligible patients, 345 (77.9%) were classified as Simulated IVindolent and, among them, 126 (36.5%) received first-line CT, whereas 219 (63.5%) received ET without targeted companions (eg, PI3K inhibitors, CDK 4/6 inhibitors (CDK4/6i)). On the other hand, 98 patients (22.2%) were classified as Simulatedaggressive and among them 46 (46.9%) received first-line CT, while 48 (53.1%) received ET. Patients classified as Simulatedaggressive had a significantly worse outcome compared with the Simulated IVindolent subgroup, in terms of OS (HR: 1.82; 95%CI 1.38-2.39; P < .001; median: 23.74 vs 41.06 months) (Fig. 3A) and a non-significant numerical difference with respect to PFS (HR: 1.24; 95%CI 0.98-1.59; P = .074, median: 8.91 vs 11.70 months) (Fig. 3B).
Figure 3.
CTCs enumeration simulation of on an independent cohort comprising HR-positive, HER2-negative first-line MBC patients. The classifier was capable to stratify patients both in terms of OS (A) and PFS (B).
The prognostic impact for OS was also investigated through multivariable analysis to test the independent role of the classifier on outcome. Simulatedaggressive retained its prognostic significance in terms of OS (HR: 1.58; 95%CI 1.17-2.15; P = .003; Table 2) and its addition increased the model’s overall concordance index (Harrell’s C = 0.6381 and 0.6581, respectively, before and after the addition of the classifier to the multivariable model).
Table 2.
Main prognostic factors in terms of OS both on uni and multivariable analysis
| Variable | Univariate | Multivariate | ||||
|---|---|---|---|---|---|---|
| HR | 95% CI | P-value | HR | 95% CI | P-value | |
| BC subtype | ||||||
| Luminal A | 1 | 1 | ||||
| Luminal B | 1.57 | 1.16-2.12 | 0.003 | 1.46 | 1.06-2.00 | .020 |
| BMI | ||||||
| ≤25 | 1 | |||||
| >25 | 1.15 | 0.87-1.53 | 0.327 | |||
| CTCs simulation | ||||||
| Simulatedindolent | 1 | 1 | ||||
| Simulatedaggressive | 1.82 | 1.38-2.39 | < 0.001 | 1.58 | 1.17-2.14 | .003 |
| Age at treatment start | ||||||
| <70 years | 1 | |||||
| ≥70 years | 1.84 | 1.43-2.36 | < 0.001 | 2.08 | 1.57-2.76 | <.001 |
| ET naïve | ||||||
| No | 1 | 1 | ||||
| Yes | 0.71 | 0.55-0.90 | 0.005 | 0.51 | 0.29-0.89 | .017 |
| CT naïve | ||||||
| No | 1 | |||||
| Yes | 0.81 | 0.63 -1.02 | 0.077 | |||
| Stage IV onset | ||||||
| Relapsed | 1 | 1 | ||||
| De novo | 0.73 | 0.57 - 0.94 | 0.016 | 1.03 | 0.58-1.82 | .931 |
| ECOG PS | ||||||
| 0-1 | 1 | 1 | ||||
| ≥2 | 1.62 | 1.17-2.25 | 0.004 | 1.58 | 1.11-2.25 | .011 |
The model was also able to further stratify clinical subgroups usually considered prognostically homogeneous. Similarly to what was observed with CTCs enumeration in the pooled population (Fig. 4A), patients with bone only metastases classified as Simulated IVaggressive had a significantly worse prognosis with respect to the Simulated IVindolent counterpart (P < .0001) (Fig. 4C). Consistently, patients with liver metastases classified as Simulated IVindolent had a significantly better prognosis than the Simulated IVaggressive counterpart (P < .0001; Fig. 4D).
Figure 4.
CTCs stratification and simulation among patients with bone-only (BO) disease (A, C) and liver (Liv) involvement (B, D). In the pooled population, patients with bone only metastases classified as StageIVaggressive had a significantly worse prognosis with respect to StageIVindolent (P < .0001) (A). Similar results were observed in the Simulated counterpart (P < .0001) (C). Consistently, patients with liver metastases classified as StageIVindolent had a significantly better prognosis than the StageIVaggressive counterpart (P < .0001) (B) Similar results were observed in Simulatedindolent patients with liver metastases (P < .0001) (D).
CTCs Classifier and Impact of Treatment Type
As a proof of principle, the differential prognostic impact of first-line ET and CT was explored across the Simulatedaggressive and Simulatedindolent subgroups. Consistently with previously published results, no significant differences were observed between ET and CT in the overall population, both in terms of PFS (HR: 1; 95%CI 0.81-1.23; P = .998) and OS (HR: HR: 0.92; 95%CI 0.72-1.18; P = .511) (Fig. 5A, D). Similar results were obtained in the Simulatedindolent subgroup (HR: 1.13; 95%CI 0.89-1.44; P = .301 and HR: 0.95; 95%CI 0.72-1.28; P = .759 respectively for PFS and OS) (Fig. 5B, E). On the other hand, a statistically significant difference, favoring CT over ET was observed among Simulatedaggressive patients (HR: 0.62; 95%CI 0.40 – 0.96; P = .030 and HR: 0.60; 95%CI 0.37 – 0.97; P = .037, respectively, for PFS and OS; Fig. 5C, F).
Figure 5.
Impact of first-line ET and CT in terms of OS and PFS according to the classifier’s stratification. No significant impact was observed in the total population (A, D) nor in the Simulatedindolent subgroup (B, E) in terms of OS and PFS, while a significantly different outcome was observed in the Simulatedaggressive subgroup (C, F).
Discussion
The present study explored the concept of simulating the CTC-based prognostication to investigate the impact of different therapeutic approaches in existing databases that are lacking for this characterization. A KNN supervised machine learning model was trained on a pooled dataset of 2436 MBC patient from EPAC and MDACC with a resulting 57.1% sensitivity (95%CI: 50.8-63.3%), a 61.6% specificity (95%CI: 55.9-67.0%) and a notably comparable risk stratification with respect to the real CTCs enumeration (StageIVindolent vs Simulatedindolent HR 1.18, 95%CI 0.93-1.51 P = .177; StageIVaggressive vs Simulatedaggressive HR 0.88, 95%CI 0.70-1.09, P = .242).
As a proof of concept, the classifier was applied to a real-world cohort of 446 patients affected by HR-positive HER2-negative MBC to investigate the differential prognostic impact of first-line ET and CT across the Simulatedaggressive and the Simulatedindolent subgroups in a clinically homogeneous population. The dataset was previously analyzed through a propensity score matching approach to explore the prognostic impact of CT vs ET as first-line treatment showing no significant differences.11 As expected, Simulatedaggressive patients experienced a worse outcome both in terms of PFS and OS (Fig. 3), the latter confirmed also in multivariable analysis. Intriguingly, a differential prognostic impact of ET and CT was observed within the 2 simulated cohorts. In the Simulatedindolent subgroup, no difference in outcome was observed when patients were treated with ET or CT. In contrast, the Simulatedaggressive subgroup had a significantly better outcome in patients receiving CT over ET, both in terms of PFS and OS (Fig. 5).
The use of ET-based treatments with respect to CT, is supported by a therapeutic benefit burdened by lower toxicities and better quality of life. Moreover, novel combinations of ET plus targeted treatments, including CDK4/6i, have shown remarkable efficacy in patients with HR-positive HER2-negative MBC.12,13 Therefore, current guidelines recommend ET-based treatments with CDK4/6i as first-line therapy for HR-positive HER2-negative MBC, whereas CT should be considered as the preferred treatment strategy in patients with visceral disease that acutely threaten organ function.11
Bone-only disease is often regarded as a distinct clinical subgroup characterized by a favorable prognosis and a prolonged OS, and therefore these patients are potentially eligible to receive a lower-intensity treatment and disease monitoring.14 However, we observed a subset of patients with CTC-defined bone-only disease that experienced a significantly worse prognosis (Fig. 4) which accounted for 49.6% of patients with bone-only disease in the pooled cohort (198 out of the total 399). The present study, therefore, suggests that additional biomarkers, such as CTCs enumeration, could be useful to further stratify this subpopulation and identify patients that may benefit from a different therapeutic approach. Many studies have been conducted so far to evaluate clinical features such as disease-free interval, number, and type of metastatic sites as clinical markers potentially useful to guide treatment decision. However, none of these have been shown to be a useful predictive marker.15 In this scenario, CTCs enumeration could provide a potentially useful tool.
A similar concept was explored by the phase III STIC CTCs trial.8 The study randomized 761 MBC patients between a clinically-driven choice or a CTC-driven choice defined on the established ≥5 CTC/7.5ml cutoff.4 Patients classified as “high risk” based on the assigned approach received CT, while those classified as “low-risk” received ET. PFS was showed to be not inferior in the CTC-driven with respect to the clinically-driven one (HR 0.98, 90%CI 0.84-1.13).8 Intriguingly, StageIVaggressive patients that were clinically defined as “low-risk” had a significantly longer PFS when treated with CT (in the CTCs arm) with respect to those treated with ET (in the clinically driven arm), highlighting the impact of treatment type on patient outcome (PFS HR 0.67, 95%CI 0.49-0.92 P = .01). Importantly, these results are consistent with those generated by the classifier in our study, further supporting its reliability and potential utility.
One limitation of this study is the lack of inclusion of patients treated with CDK4/6i. On the other hand, this is the first proposed “in silico” approach capable to stratify patients according to the simulation of CTCs-based staging.
Of note, since the classifier was trained using the widely established ≥5 CTC/7.5 ml cutoff and a large un-selected MBC cohort, it offers a generalizable platform for hypothesis generation that can be transferred to a broad variety of real-world or clinical trial databases.3-6,8,9
Previous attempts have been made to explore new subgroups with differential treatment benefits through machine learning algorithms. Patient-level data from 4580 breast cancer patients enrolled in 8 randomized clinical trials treated with CDK4/6i were analyzed through random survival forest models based on clinical baseline characteristics with a resulting 69.2% accuracy.16
The present study designed a classifier with a 65.1% accuracy based on a strong, setting-independent biological biomarker, enabling its application on a broader set of clinical questions.
Although the present study provided evidence of an in silico simulation of the CTC-based stratification, its main objective was not to replace the real CTCs enumeration, which has specific biological implications and is certainly more solid in PFS and OS prognostication. It rather identifies patients with comparable prognostic characteristics for hypothesis generation and the subsequent design of prospective, biomarker-driven, clinical trials with the ultimate goal of catalyzing sample size optimization and clinical trials optimization by exploring different levels of treatment intensity and the impact of methodological aspects in subpopulations with different risk profiles.17
Conclusion
The present study showed the feasibility of a KNN machine learning classifier to simulate a baseline CTCs-based staging. This model could be used for hypothesis generation in specific case scenarios of interest to identify subpopulations which may benefit from higher intensity treatments due to a more aggressive outcome, representing a valuable tool for future clinical trials design and prospective, biomarker-driven, validation studies.
Supplementary Material
Contributor Information
Lorenzo Gerratana, Department of Medical Oncology, Centro di Riferimento Oncologico (CRO), IRCCS, Aviano (PN), Italy.
Jean-Yves Pierga, Department of Medical Oncology, Institut Curie, Paris & Saint-Cloud, Paris University, Paris, France.
James M Reuben, Department of Hematopathology, The University of Texas MD Anderson Cancer Center, Houston, TX, USA.
Andrew A Davis, Division of Hematology and Oncology, Department of Medicine, Northwestern University Feinberg School of Medicine, Chicago, IL, USA; Department of Medicine, Division of Oncology, Washington University School of Medicine in St. Louis, MO, USA.
Firas H Wehbe, Division of Hematology and Oncology, Department of Medicine, Northwestern University Feinberg School of Medicine, Chicago, IL, USA.
Luc Dirix, Translational Cancer Research Unit, GZA Hospitals Sint-Augustinus, Antwerp, Belgium.
Tanja Fehm, Department of Gynecology and Obstetrics, Heinrich Heine University Düsseldorf, Düsseldorf, Germany.
Franco Nolé, Medical Oncology Division of Urogenital and Head & Neck Tumours IEO, European Institute of Oncology IRCCS, Milan, Italy.
Rafael Gisbert-Criado, Clinical Laboratory, Hospital Arnau de Vilanova, Valencia, Spain.
Dimitrios Mavroudis, Laboratory of Translational Oncology, School of Medicine, University of Crete, Heraklion, Greece; Department of Medical Oncology, University Hospital of Heraklion, Greece.
Salvatore Grisanti, epartment of Transfusion Medicine, Laboratory for Stem Cells Manipulation and Cryopreservation, AO Spedali Civili di Brescia, Brescia, Italy.
Jose A Garcia-Saenz, Instituto de Investigación Sanitaria Hospital Clinico San Carlos (IdISSC), CIBERONC, Madrid, Spain.
Justin Stebbing, Division of Cancer, Department of Surgery and Cancer, Imperial College London, London, UK.
Carlos Caldas, Cancer Research UK Cambridge Institute and Department of Oncology Li Ka Shing Centre, University of Cambridge, Cambridge, UK.
Paola Gazzaniga, Department of Molecular Medicine, Sapienza University of Rome, Rome, Italy.
Luis Manso, Hospital 12 de Octubre, Madrid, Spain.
Rita Zamarchi, Veneto Institute of Oncology IOV - IRCCS, Padua, Italy.
Marta Bonotto, Department of Oncology, ASUFC University Hospital, Udine, Italy.
Angela Fernandez de Lascoiti, Hospital de Navarra, Pamplona, Spain.
Leticia De Mattos-Arruda, Val d’Hebron Institute of Oncology, Val d’Hebron University Hospital, and Universitat Autònoma de Barcelona, Barcelona, Spain.
Michail Ignatiadis, Department of Medical Oncology and Breast Cancer Translational Research Laboratory, Institut Jules Bordet, Université Libre de Bruxelles, Brussels, Belgium.
Maria-Teresa Sandri, Division of Laboratory Medicine, Humanitas Reseach Hospital, Rozzano, Milan, Italy.
Daniele Generali, Women Cancer Center, Azienda Socio-Sanitaria Territoriale di Cremona, Cremona, Italy; University of Trieste, Trieste, Italy.
Carmine De Angelis, Department of Clinical Medicine and Surgery, University Federico II, Naples, Italy; Lester and Sue Smith Breast Center, Baylor College of Medicine, Houston, TX, USA.
Sarah-Jane Dawson, Centre for Cancer Research and Sir Peter MacCallum Department of Oncology, The University of Melbourne, VIC, Australia.
Wolfgang Janni, Frauenklinik, University of Ulm, Ulm, Germany.
Vicente Carañana, Clinical Oncology, Hospital Arnau de Vilanova, Valencia, Spain.
Sabine Riethdorf, Department of Tumor Biology, Center of Experimental Medicine, University Cancer Center Hamburg, University Medical Center Hamburg-Eppendorf, Hamburg, Germany.
Erich-Franz Solomayer, Saarland University, Homburg, Germany.
Fabio Puglisi, Department of Medical Oncology, Centro di Riferimento Oncologico (CRO), IRCCS, Aviano (PN), Italy; Department of Medicine, University of Udine, Udine, UD, Italy.
Mario Giuliano, Department of Clinical Medicine and Surgery, University Federico II, Naples, Italy.
Klaus Pantel, Department of Tumor Biology, Center of Experimental Medicine, University Cancer Center Hamburg, University Medical Center Hamburg-Eppendorf, Hamburg, Germany.
François-Clément Bidard, Department of Medical Oncology, Institut Curie, Paris & Saint-Cloud, Paris University, Paris, France.
Massimo Cristofanilli, Division of Hematology and Oncology, Department of Medicine, Northwestern University Feinberg School of Medicine, Chicago, IL, USA; Department of Medicine, Division of Hematology-Oncology, Weill Cornell Medicine/New York-Presbyterian Hospital, New York, NY, USA.
Funding
The study was supported by Lynn Sage Cancer Research Foundation and the the CRO Aviano 5x1000 2014 per la Ricerca Sanitaria, Cancer Specific Intramural Grant. The funding sources had no role in the study design, data collection, data analysis, interpretation, or writing of the manuscript.
Compliance with Ethical Standards
Ethical approval: The anonymized EPAC/MDACC data were transferred to the Robert H. Lurie Comprehensive Cancer Center-Bioinformatics and a retrospective Institutional Review Board-approved protocol was used to access and analyze the data. The combined Udine/Neaples cohort was previously approved by the Review Committees of each participant center.
Conflict of Interest
Lorenzo Gerratana: Eli Lilly and Novartis (H), Menarini Silicon Biosystems (RF); Jean-Yves Pierga: Seagen, Daiichi Sankyo, Pfizer, AstraZeneca, MSD, Gilead, Roche, Novartis, Exact Sciences (C/A), Menarini, Servier (RF), Viatris, Lilly, Pierre Fabre Oncology (H); Andrew A. Davis: Menarini Silicon Biosystems (RF); Justin Stebbing: Vaccitech, Heat Biologics, Eli Lilly, Alveo Technologies, Pear Bio, Agenus, Equilibre Biopharmaceuticals, Graviton Bioscience Corporation, Celltrion, Volvox, Certis Oncology Solutions, Greenmantle, Zedsen, Bryologyx and Benevolent AI (SAB), Lansdowne partners and Vitruvian (C/A); Xerion and BB Biotech Healthcare Trust PLC (Other—Leadership, Board of Directors); Carlos Caldas: AstraZeneca (SAB), AstraZeneca, Genentech, Roche, Servier, Cycle Therapeutics (RF—inst); Michail Ignatiadis: Novartis, Seattle Genetics (C/A), Roche, Pfizer, Natera Inc (RF—inst); Carmine De Angelis: Roche, AstraZeneca, Lilly, GSK, Novartis, Pfizer (C/A), Novartis, Pfizer, Lilly (H), Novartis (RF—inst); Sarah Jane Dawson: AstraZeneca, Inivata (SAB), Wolgang Janni: Menarini (RF); Erich-Franz Solomayer: Amgen, AstraZeneca, Celgene, Clovis Oncology, Eisai, Erbe, Gedeon Richter, Genomic Health, Jenapharm, Johnson Johnson, Matramed, Medac, Mentor, Novartis Pfizer, Pharma Mar, MSD, Roche, Samsung, Storz, Tewa, Vifor (H); Mario Giuliano: Lilly, Novartis, Pfizer, Roche, MSD, Seagen, AstraZeneca (C/A, H); Fabio Puglisi: Amgen, Astrazeneca, Daichii-Sankyo, Eisai, Eli Lilly, MSD, Novartis, Pierre-Fabre, Roche, Seagen (C/A), AstraZeneca, Eisai, Roche Other, Celgene, GlaxoSmithKline, Roche (RF); Massimo Cristofanilli: Pfizer, Merus, Novartis, CytoDyn (H), Menarini, Olaris, Lilly, Celcuity, AZ (C/A), Lilly, Pfizer, Menarini, Guardant, AZ (RF). The other authors indicated no financial relationships.
(C/A) Consulting/advisory relationship; (RF) Research funding; (E) Employment; (ET) Expert testimony; (H) Honoraria received; (OI) Ownership interests; (IP) Intellectual property rights/inventor/patent holder; (SAB) Scientific advisory board.
Author Contributions
Conception/design: L. G., A.A.D., F.P., M.C. Provision of study material/patients: L.G., J.Y.P., J.R., A.A.D., F.H.W., L.D., T.F., F.N., R.G.-C., D.M., S.G., J.A.G-S., J.S., C.C., P.G., L.M., R.Z., M.B., A.F.d.L., L.D.M.A., M.I., M.-T.S., D.G., C.D.A., S.-J.D., W.J., V.C., S.R., E.-F.S., F.P., M.G., K.P., F. -C.B., M.C. Collection and/or assembly of data: L.G., J.-Y.P., J.R., A.A.D., F.H.W., L.D., T.F., F.N., R.G.-C., D.M., S.G., J.A.G.-S., J.S., C.C., P.G., L.M., R.Z., M.B., A.F.d.L., L.D.M.-A., M.I., M.-T.S., D.G., C.D.A., S.-J.D., W.J., V.C., S.R., E.-F.S., F.P., M.G., K.P., F.-C.B., M.C. Data analysis and interpretation: L.G., A.A.D., F.H.W., M.B., C.D.A., F.P., M.G., F.-C.B., M.C. Manuscript writing: L.G., J.-Y.P., J.R., A.A.D., F.H.W., L.D., T.F., F.N., R.G.-C., D.M., S.G., J.A.G.-S., J.S., C.C., P.G., L.M., R.Z., M.B., A.F.d.L., L.D.M.-A., M.I., M.-T.S., D. G., C.D.A., S.-J.D., W.J., V.C., S.R., E.-F.S., F.P., M.G., K.P., F.-C.B., M.C. Final approval of manuscript: All authors.
Data Availability
The data underlying this article will be shared on reasonable request to the corresponding author.
References
- 1. Bonotto M, Gerratana L, Poletto E, et al. Measures of outcome in metastatic breast cancer: insights from a real-world scenario. Oncologist. 2014;19(6):608-615. 10.1634/theoncologist.2014-0002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Kiely BE, Soon YY, Tattersall MHN, Stockler MR.. How long have I got? Estimating typical, best-case, and worst-case scenarios for patients starting first-line chemotherapy for metastatic breast cancer: a systematic review of recent randomized trials. J Clin Oncol. 2011;29(4):456-463. 10.1200/JCO.2010.30.2174. [DOI] [PubMed] [Google Scholar]
- 3. Cristofanilli M, Budd GT, Ellis MJ, et al. Circulating tumor cells, disease progression, and survival in metastatic breast cancer. N Engl J Med. 2004;351(8):781-791. 10.1056/NEJMoa040766. [DOI] [PubMed] [Google Scholar]
- 4. Cristofanilli M, Pierga J-Y, Reuben J, et al. The clinical use of circulating tumor cells (CTCs) enumeration for staging of metastatic breast cancer (MBC): international expert consensus paper. Crit Rev Oncol Hematol. 2019;134(134):39-45. 10.1016/j.critrevonc.2018.12.004. [DOI] [PubMed] [Google Scholar]
- 5. Bidard F-C, Peeters DJ, Fehm T, et al. Clinical validity of circulating tumour cells in patients with metastatic breast cancer: a pooled analysis of individual patient data. Lancet Oncol. 2014;15(4):406-414. 10.1016/S1470-2045(14)70069-5. [DOI] [PubMed] [Google Scholar]
- 6. Smerage JB, Barlow WE, Hortobagyi GN, et al. Circulating tumor cells and response to chemotherapy in metastatic breast cancer: SWOG S0500. J Clin Oncol. 2014;32(31):3483-3489. 10.1200/JCO.2014.56.2561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Gerratana L, Davis AA, Zhang Q, et al. Longitudinal dynamics of circulating tumor cells and circulating tumor DNA for treatment monitoring in metastatic breast cancer. JCO Precis Oncol 2021;943(5):52. 10.1200/po.20.00345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Bidard F-C, Jacot W, Kiavue N, et al. Efficacy of circulating tumor cell count–driven vs clinician-driven first-line therapy choice in hormone receptor–positive, ERBB2-negative metastatic breast cancer. JAMA Oncol. 2021;7(1):34-41. 10.1001/jamaoncol.2020.5660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Zeune LL, Wit S, Berghuis AMS, et al. How to agree on a CTC: evaluating the consensus in circulating tumor cell scoring. Cytom Part A. 2018;93(12):1202-1206. 10.1002/cyto.a.23576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Kuhn M. Building predictive models in R using the caret package. J Stat Softw. 2008;28(5):1-26. 10.18637/jss.v028.i0527774042 [DOI] [Google Scholar]
- 11. Bonotto M, Gerratana L, Di Maio M, et al. Chemotherapy versus endocrine therapy as first-line treatment in patients with luminal-like HER2-negative metastatic breast cancer: a propensity score analysis. The Breast. 2017;31(31):114-120. 10.1016/j.breast.2016.10.021. [DOI] [PubMed] [Google Scholar]
- 12. Sledge GW, Toi M, Neven P, et al. The effect of abemaciclib plus fulvestrant on overall survival in hormone receptor–positive, ERBB2-negative breast cancer that progressed on endocrine therapy—MONARCH 2. JAMA Oncol. 2020;6(1):116-124. 10.1001/jamaoncol.2019.4782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Turner NC, Slamon DJ, Ro J, et al. Overall survival with palbociclib and fulvestrant in advanced breast cancer. N Engl J Med. 2018;379(20):1926-1936. 10.1056/NEJMoa1810527. [DOI] [PubMed] [Google Scholar]
- 14. Toss A, Venturelli M, Sperduti I, et al. First-line treatment for endocrine-sensitive bone-only metastatic breast cancer: systematic review and meta-analysis. Clin Breast Cancer. 2019;19(6): e701-e716. 10.1016/j.clbc.2019.06.011. [DOI] [PubMed] [Google Scholar]
- 15. Cristofanilli M, DeMichele A, Giorgetti C, et al. Predictors of prolonged benefit from palbociclib plus fulvestrant in women with endocrine-resistant hormone receptor–positive/human epidermal growth factor receptor 2–negative metastatic breast cancer in PALOMA-3. Eur J Cancer. 2018;104(104):21-31. 10.1016/j.ejca.2018.08.011. [DOI] [PubMed] [Google Scholar]
- 16. Mason J, Gong Y, Amiri-Kordestani L, Wedam S, Gao JJ, Singh H, et al. Abstract PD2-07: Prediction of CDK inhibitor efficacy in ER+/HER2- breast cancer using machine learning algorithms. Poster Spotlight Sess. Abstr., American Association for Cancer Research; 2020; (4_Suppl), p. PD2-07-PD2-07. 10.1158/1538-7445.SABCS19-PD2-07. [DOI] [Google Scholar]
- 17. Broglio KR, Berry D.. Detecting AN overall survival benefit that is derived from progression-free survival. JNCI J Natl Cancer Inst. 2009;101:1642-1649. 10.1093/jnci/djp369. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets supporting the conclusions of this article are available from the corresponding author on reasonable request.
The data underlying this article will be shared on reasonable request to the corresponding author.