Abstract
Hybridization with oligonucleotide microchips (microarrays) was used for discrimination among strains of Escherichia coli and other pathogenic enteric bacteria harboring various virulence factors. Oligonucleotide microchips are miniature arrays of gene-specific oligonucleotide probes immobilized on a glass surface. The combination of this technique with the amplification of genetic material by PCR is a powerful tool for the detection of and simultaneous discrimination among food-borne human pathogens. The presence of six genes (eaeA, slt-I, slt-II, fliC, rfbE, and ipaH) encoding bacterial antigenic determinants and virulence factors of bacterial strains was monitored by multiplex PCR followed by hybridization of the denatured PCR product to the gene-specific oligonucleotides on the microchip. The assay was able to detect these virulence factors in 15 Salmonella, Shigella, and E. coli strains. The results of the chip analysis were confirmed by hybridization of radiolabeled gene-specific probes to genomic DNA from bacterial colonies. In contrast, gel electrophoretic analysis of the multiplex PCR products used for the microarray analysis produced ambiguous results due to the presence of unexpected and uncharacterized bands. Our results suggest that microarray analysis of microbial virulence factors might be very useful for automated identification and characterization of bacterial pathogens.
In recent years, DNA and oligonucleotide microchip (microarray) technology has played an increasingly important role in genomic studies, drug discovery, and toxicological research. Unlike other hybridization formats (hybridization with microplates or dot blot hybridization with membrane-bound probes), glass microchips allow significant miniaturization so that thousands of individual probes can be arranged on one glass slide. As a result, this technology is ideal for an extensive parallel identification of nucleic acids and analysis of gene expression. Simultaneous analysis for the presence of multiple markers makes it possible to determine a complete genetic profile of a single strain or distinguish one strain from a very large collection of possible alternatives in one experiment. Therefore, this approach is potentially useful for the screening of multiple microbial isolates in a diagnostic assay.
Oligonucleotide microchips containing multiple oligonucleotides are spotted on the chip surface. DNA samples for analysis are labeled with fluorescent dyes and hybridized with the oligonucleotide spots on the chip. The fluorescence pattern is then recorded by a scanner, quantified, and analyzed. While DNA microchips have been used mostly for gene expression studies, the technique has great potential to be used for the discrimination of genotypes, point mutants, and other closely related sequences by employing oligonucleotides specific for each sequence variant.
Microarray technology has great potential for use in diagnostic microbiology. Microbial pathogens are currently identified by using surrogate biochemical and immunological markers. An alternative approach developed in recent years makes use of ribosomal DNA (rDNA) as surrogate markers for bacterial identification. These conventional approaches are well established and effective; however, they are often time-consuming and do not directly characterize virulence factors of the organism identified. It is desirable to be able to identify both the organism and its virulence factors, and this may be feasible by using oligonucleotide microchips specifically recognizing and discriminating bacterial rDNA and various virulence factors.
We have tested this technology by using oligonucleotide arrays to identify the presence of specific markers in bacterial genomes associated with pathogenesis. As a model system, we chose to analyze the enteric bacteria Shigella, Salmonella, and Escherichia coli.
E. coli O157:H7 is a leading cause of hemorrhagic colitis and is one of the most important food-borne human pathogens (1, 23). Traditionally, E. coli O157:H7 is identified by using microbiological culture techniques (4, 15, 16, 22) followed by immunological methods to detect the O157 and H7 antigens. The presence of the shiga-like toxins (SLT) that characterize E. coli O157:H7 is then confirmed, using antibodies to the toxins (24). This method is slow and complicated and sometimes yields false-positive results because of cross-reactivity of the antibodies or false-negative results when other variants of E. coli O157:H7 are inadvertently isolated in the initial culture step (7).
As an alternative, many PCR-based assays have been developed for the detection of E. coli O157:H7. Some of the common target genes for PCR amplification are the conserved regions of slt-I and -II (19) and eaeA (intimin) (28), which mediates the adherence of the organism to host cells.
Several PCR-based assays have been developed for O157:H7 detection using slt-I and -II and eaeA (6, 8, 9, 11, 12, 17, 18, 20, 21, 25, 26). Because these genes are not unique to serotype O157:H7, more specific target genes have also been used, including rfbE, which encodes an enzyme involved in the biosynthesis of the O157 antigen (2, 5) in combination with fliC, encoding the H7 antigen. A multiplex PCR has been developed which directly detects genes that are involved in the biosynthesis of the O157 and H7 antigens, along with SLT-I, SLT-II, and intimin (14). This multiplex PCR was shown to be effective in the analysis of bovine feces. There was no cross-reaction with the background bacterial flora, and the researchers were able to detect a single O157:H7 organism per gram of feces when an enrichment step was used.
An additional virulence factor is the invasion plasmid antigen (ipaH) (13) associated with the invasive phenotype of Shigella and enteroinvasive E. coli (EIEC). ipaH was used to identify Shigella species or EIEC among 154 patients with dysentery and family contacts (10). The ipaH PCR system increased the detection of Shigella species and EIEC from 58 to 79% among patients with dysentery and from 6 to 22% among 527 family contacts; 75% of infections in family members were asymptomatic. Detection of ipaH was statistically associated with dysentery.
Here, we used eaeA, slt-I, slt-II, fliC, rfbE, and ipaH, encoding bacterial antigenic determinants and virulence factors, for a multiplex PCR for microarray analysis.
MATERIALS AND METHODS
Bacterial strains.
Bacterial strains from the Food and Drug Administration Center for Food Safety and Applied Nutrition collection used in this study are shown in Table 1.
TABLE 1.
Strains used in this work
| Sample no. | Strain | Organism | Type |
|---|---|---|---|
| 1 | EC536 | E. coli | EHEC |
| 2 | EC488 | E. coli | EHEC |
| 3 | 567 | Shigella dysenterii | |
| 4 | SENT | Salmonella enterica serovar Enteritidis | |
| 5 | ECOR48 | E. coli | O157-like |
| 6 | DEC5A | E. coli | Pathogen |
| 7 | DEC5C | E. coli | Pathogen |
| 8 | M90T | Shigella flexneri | |
| 9 | EC503 | E. coli | EHEC |
| 10 | TB285 | E. coli | EHEC |
| 11 | KL10035 | E. coli | EPEC |
| 12 | KL10016 | E. coli | EIEC |
| 13 | W3110 | E. coli | K-12 |
| 14 | TB334 | E. coli | EHEC |
| 15 | EC535 | E. coli | EHEC |
Primers.
Primers used to amplify the fragments of virulence factor genes (Table 2) eaeA, rfbE, slt-I, slt-II, and fliC were described previously (14). Primers for the amplification of ipaH were provided by K. Lampel.
TABLE 2.
Oligonucleotides and PCR primers for virulence factor targets
| Target gene | GenBank accession no. | PCR product size (bp) | Primer sequences for PCR amplificationa | Oligonucleotide targets adsorbed to array chipb | Annealing temp (°C) |
|---|---|---|---|---|---|
| rfbE | AF061251 | 292 | L, TGTCCATTTATACGGACATCCATG | GGTTGTCACGAATGACAAAAC | 50 |
| R, CCTATAACGTCATGCCAATATTGCC | CTATTACTACAGGTGAAGGTG | 50 | |||
| fliC | AF169323 | 625 | L, GCGCTGTCGAGTTCTATCGAGC | AACGGTTAGCAATCGCCTGA | 52 |
| R, CAACGGTGACTTATCGCCATTCC | CGTCATCCTTCGCGCTGTTA | 54 | |||
| eaeA | AF081184 | 368 | L, GACTGTCGATGCATCAGGCAAAG | GCCTATTATGCTGATGCTATG | 50 |
| R, TTGGAGTATTAACATTAACCCCAGG | GTTCTATGAACTCAATAACTGCTTGG | 55 | |||
| slt-I | L04539 | 210 | L, TGTAACTGGAAAGGTGGAGTATAC | GGATGACGTAACCATTAAAACT | 49 |
| R, GCTATTCTGAGTCAACGAAAAATAAC | CTAATGCCTGTCATAATGGAGG | 53 | |||
| slt-II | X65949 | 484 | L, GTTTTTCTTCGGTATCCTATTCCG | TCAGTCGTCACTCACTGGTT | 52 |
| R, GATGCATCTCTGGTCATTGTATTAC | CTGGAACGTTCCGGAATGCAA | 54 | |||
| ipaH | M76445 | 269 | L, CCACTGAGAGCTGTGAGGAC | GTGGATGAGATAGAAGTCTACC | 53 |
| R, ATGGCGTGTCGGGAGTGACA | GACCATGCTCGCAGAGAAACT | 54 |
L, left; R, right.
Oligonucleotide sequences are independent sequences corresponding to the top and bottom spots in each set of rows in Fig. 3.
Target sequence design.
BLAST search (at the website of the National Center for Biotechnology Information at the National Library of Medicine) was used to find and retrieve the sequences of homologues of each of the six genes analyzed. Included in the procedure were sequences (with accession numbers) of rfbE (AF061251, S83460, and AB008676), fliC (AF169323, AF169321, AF169320, AF128958, AF128956, AF128955, and AF128958), eaeA (AF081184, AF81183, AF81182, AF071034, and X60439), slt-I (L04539, AB015056, and AB035142), slt-II (X65949, Z37725, X81415, X81416, X81417, X81418, X67514, and M36727), and ipaH (M76445, M76444, M32063, AF047365, and M76443). The retrieved sequences were aligned using ClustalX software. For each gene, two conserved regions of approximately 21 nucleotides with a low annealing temperature were chosen as target sequences. The oligonucleotide targets and their respective melting temperatures (http://www.basic.nwu.edu/biotools/OligoCalc.html) are presented in Table 2.
The 5′ end of each oligonucleotide was aminated during the synthesis using TFA Aminolink CE reagent (PE Applied Biosystems, Davis, Calif.) to increase the efficiency of the immobilization of oligonucleotide to silylated slides (CEL Associates, Inc., Houston, Tex.).
Microchip design and analysis.
Microchips were printed by using a contact micro-spotting robotic system, PIXSYS 5500 (Cartesian Technologies, Inc., Irvine, Calif.) and ChipMaker Micro-spotting device (TeleChem International, Inc., Sunnyvale, Calif.). The average size of the spots was 200 μm. The concentration of oligonucleotides before printing was adjusted to 50 μM in 15% glycerol–0.25 M acetic acid. Printed slides were incubated for at least 30 min at 75°C to evaporate glycerol completely, followed by a 5-min incubation in a fresh 0.25% solution of NaBH4 in water. Slides were washed once for 1 min with 0.2% sodium dodecyl sulfate (SDS) in water and two times for 1 min each with distilled water to remove unbound oligonucleotides. To reduce the nonspecific adsorption of fluorescent probe to the surface, slides were incubated in 4× Denhardt's solution containing 1% SDS for 10 min and dried in air before hybridization. Control spots used for marking array position on slides were generated using 1× Spotting Solution (ArrayIt, Sunnyvale, Calif.) in 0.25 M acetic acid.
Multiplex PCR and synthesis of Cy5-labeled probes.
Multiplex PCR was performed as previously described (14), with minor modifications. Briefly, 50 μl of reaction mixture contained 1× AmpliTaq PCR buffer with 1.5 mM MgCl2, 200 nM concentrations of each primer, 100 μM concentrations of each of the four deoxynucleoside triphosphates, and 1 U of AmpliTaq polymerase. Amplification was performed using 35 cycles (30 s at 94°C, 1 min at 59°C, 1 min at 72°C). The PCR products were analyzed by electrophoresis in a 2% agarose gel. Cy5-labeled fluorescent probe was generated by using PCR conditions described above except that the annealing temperature was 45°C. The concentration of dCTP in the reaction mixture was reduced to 20 μM, and the fluorescent analogs of dCTP (Cy5-dCTP) and Triton X-100 were added to 20 μM and 1%, respectively. Labeled PCR products were separated from nonincorporated Cy5-dCTP by centrifugation on CentriSep columns (Princeton Separations, Adelphia, N.J.), dried in vacuum, and solubilized in 10 μl of water.
Hybridization conditions.
Hybridization of the fluorescent probe to the microchip was performed in 1× UniHyb solution (ArrayIt) at 37°C for 30 min. Probe DNA was denatured before hybridization at 95°C for 1 min and chilled on ice. A 2- to 3-μl spot from each probe was applied to the microarray and covered with a 5- by 5-mm plastic coverslip to prevent drying of the probe during incubation in the hybridization cassette (TeleChem International, Inc.). After hybridization, the slides were washed once with 6× SSC–0.2% SDS for 1 min at room temperature, once with 6× SSC for 1 min, and once with 2× SSC and were dried (1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate).
Scanning and quantitation of microarrays.
Fluorescent images of the microarrays were generated by scanning the slides by using a ScanArray 4000 (GSI Lumonics, Billerica, Mass.). The fluorescent signals from each spot were measured and compared by using QuantArray software (GSI Lumonics). Analysis of collected data was performed on the basis of total fluorescence intensities measured from a fixed circular area of each oligonucleotide spot. Fluorescent signals with a statistically significant difference (P < 0.01) from the background level were considered to be positive.
Colony hybridization.
The presence or absence of the six genes was confirmed by using hybridization of radiolabeled probes to bacterial colonies. The probes were PCR products labeled by using 32P-dCTP and random octomers (NEBLOT; New England Biolabs, Beverly, Mass.). Filter hybridization was performed by using a standard method (3). Briefly, overnight cultures in Luria broth were used to inoculate Luria broth-agar plates. After overnight growth, the colonies were adsorbed onto Whatman 541 paper, lysed, and dried. The probes were hybridized to the filters in hybridization solution (Life Technologies, Grand Island, N.Y.) at 68°C for 4 h followed by serial washes with 2× and 0.5× SSC at the same temperature. Detection was by autoradiography overnight. Each probe was hybridized against two replicate filters and each filter had several repeated spots for quality control.
RESULTS AND DISCUSSION
Multiplex PCR of bacterial virulence factor genes.
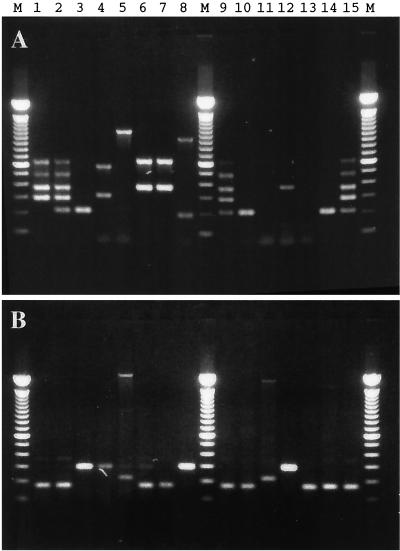
To evaluate the use of microarrays for detection of bacterial virulence factors, we used primers directed against these genes to generate fluorescent probes. Primers for amplification of ipaH were also included since the shigellae are genetically similar to enterohemorrhagic E. coli (EHEC) and have overlapping virulence factors although they are epidemiologically and clinically distinct. We used 15 different bacterial strains with various combinations of virulence factor genes (Table 1). The PCR products generated by using bacterial DNA and either five pairs of primers or ipaH-specific primers alone are presented in Fig. 1A and Fig. 1B, respectively. The PCR products generated using relaxed reaction conditions (the annealing temperature was reduced from 59 to 45°C) and all six primer pairs in the same reaction mixture are presented in Fig. 2.
FIG. 1.
Identification of virulence factor genes by multiplex PCR. The PCR (annealing temperature, 59°C) was performed in the presence of primers designed to detect the rfbE, fliC, eaeA, slt-I, and slt-II genes (A) or the ipaH gene (B). Lane numbers correspond to the bacterial sample numbers in Table 1. Lanes M, 100-bp molecular size DNA marker.
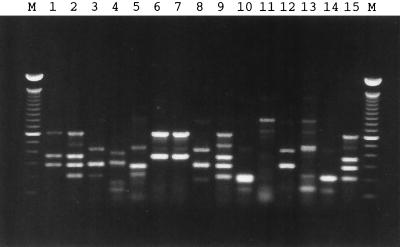
FIG. 2.
Synthesis of fluorescent probes using multiplex PCR. The Cy5-labeled PCR products were generated in the presence of six pairs of primers required for the simultaneous amplification of all target microbial virulence factor genes. The annealing temperature of the PCR was 45°C. Lane numbers correspond to the bacterial sample numbers in Table 1. Lanes M, 100-bp molecular size DNA marker.
The pitfalls of agarose gel analysis of PCR products, particularly multiplex PCR, are well illustrated in Fig. 1 and 2. Several lanes in Fig. 1 and 2 had bands that did not match the size of the positive control bands and were presumably nonspecifically amplified products containing short sequences similar to those of the primers among the set of five or six pairs. More troubling was the fact that some of these nonspecific bands had locations on the gel that were quite similar to positions of the bands in the positive control. Thus, the resolving power of the agarose gel was poor. Moreover, adapting the method to include a sixth primer pair (to detect a sixth gene) was difficult. Simply adding the extra primers led to poor amplification of some products (data not shown). Relaxed PCR conditions, those using a lower annealing temperature, resulted in the amplification of all genes, thus reducing the possibility of a false-negative result. However, this modification of PCR conditions increased amplification of additional nontargeted genes. Even when the conditions of PCR were more stringent (Fig. 1), some bands were difficult to interpret. For example, the minor bands in Fig. 1B (lanes 4, 5, and 6), with molecular sizes near 300 bp, caused uncertainty regarding the presence of ipaH. Extraneous DNA bands in Fig. 1A, lanes 8 and 12 (containing fragments >200 and 360 bp, respectively) could be misinterpreted as PCR products related to slt-I and eaeA. In summary, the detection and identification of PCR products by agarose gel electrophoresis was unsatisfactory due to the interference caused by false-positive bands.
Design and fabrication of microarray for discrimination between virulence factors.
To overcome the difficulties in the interpretation of the results of the multiplex PCR, we used oligonucleotide microarray analysis. This method has several advantages over other techniques. First, it is more specific because the sequence of a substantial portion of the amplified PCR product must match the control sequence rather than relying on a gross size estimate for identification. This permitted the use of less-stringent PCR conditions, reliably generating the desired products and reducing the possibility of a false-negative result. Second, while the specificity of detection is similar to that of methods based on the adsorption of colonies onto paper or polymer membranes, this method is much less time-consuming and labor-intensive. For each gene, two conserved regions of approximately 20 nucleotides with a low annealing temperature were chosen as target sequences. The oligonucleotide targets used in this experiment to discriminate among virulence factors, their positions, and their respective melting temperatures are presented in Table 2.
Synthesis of fluorescent probes and their hybridization.
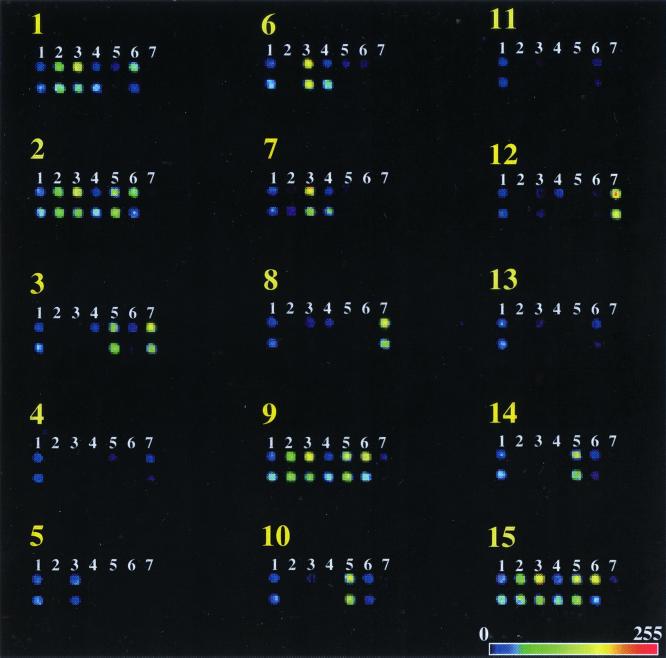
Fluorescent probes were synthesized using PCR in the presence of a Cy5-labeled analog of dCTP. The molar ratio of 1:1 of dCTP and Cy5-dCTP that was used in this study not only had no effect on the yield of PCR products (Fig. 2) but also produced higher levels of incorporation of fluorescent label into PCR product (data not shown). To generate fluorescent images of the microarrays after hybridization, a confocal scanner with a red laser operating at 633 nm was employed. Since the thermodynamic parameters of the duplex between oligonucleotide and template were slightly different for each pair, the fluorescent signals were not equal at any single temperature. Nonetheless, in most cases, the fluorescent signals were much higher (up to 80-fold) than the background level for both spots in each pair when the strain being tested contained the gene. A rainbow palette presentation of the fluorescent images of the 15 arrays hybridized at 37°C with the different fluorescent probes generated for each bacterial DNA is presented in Fig. 3.
FIG. 3.
Detection of the presence of virulence factor genes in the bacterial DNAs by using microarray technology. Yellow numbers correspond to the sample numbers of the bacterial strains listed in Table 1. Shown are nonbacterial marker spots (lanes 1) and oligonucleotide targets for specific detection of the rfbE (lanes 2), fliC (lanes 3), eaeA (lanes 4), sltI (lanes 5), sltII (lanes 6), and ipaH (lanes 7) genes. Each spot is one of two independent sequences chosen from the gene. The horizontal bar at the bottom is the scale for the color representation of fluorescent-signal intensity.
Comparison of hybridization, PCR, and microarray tests.
Three methods for detection and discrimination among bacterial virulence factors were compared. These methods were colony hybridization of target bacterial DNA with radioactive probes specific for each gene and amplification of those genes by PCR, followed by either separation of the products on agarose gel or hybridization with microchip. The results of the comparison are presented in Table 3. Data from the three different methods showed good correlation. Colony hybridization was a time-consuming but reliable method. However, the background hybridization was high for fliC. Since all the strains were motile, it is likely that similarities of flagellar genes among the related species caused this cross-reactivity. A high background for fliC was not observed in the chip assay that relied on probes that were shorter but employed more-stringent hybridization conditions. In contrast to the radioactive hybridization test, use of an oligonucleotide microarray in combination with multiplex PCR allowed the detection of several genes simultaneously. The major advantage of the microarray assay over agarose gel analysis of PCR products was that the microarray did not rely solely on the length of the PCR products but also required the internal sequences of these DNA fragments to be complementary to the oligonucleotide probes on the microchip. Since the main goal of the present study was the evaluation and comparison of methods for detection of microbial virulence factors, we used only two oligonucleotide targets for each selected gene. The number of oligonucleotides used for the detection of the same gene can be extended to make a more robust analysis. Microarray analysis is sequence sensitive. Even minimal differences between target and probe can be easily detected. This feature of the microarray was successfully used for the detection of single nucleotide polymorphisms in the human genome (27). Thus, multiple targets can be used on each chip to account for minor polymorphisms in bacteria being screened.
TABLE 3.
Comparison of three different methods for detection of bacterial virulence factors
| Sample no. | Strain | Results of methods detecting bacterial virulence factors in genea:
|
|||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
rfbE
|
fliC
|
eaeA
|
slt-I
|
slt-II
|
ipaH
|
||||||||||||||||||||
| H | P | M1 | M2 | H | P | M1 | M2 | H | P | M1 | M2 | H | P | M1 | M2 | H | P | M1 | M2 | H | P | M1 | M2 | ||
| 1 | EC536 | + | + | + | + | + | + | + | + | + | + | + | + | − | − | − | − | + | + | + | + | − | − | − | − |
| 2 | EC488 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | − | − | − | − |
| 3 | S. dysenterii 567 | − | − | − | − | + | − | − | − | − | − | − | − | + | + | + | + | − | − | − | − | + | + | + | + |
| 4 | S. enterica serovar Enteritidis | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ? | + | − |
| 5 | ECOR48 | − | − | − | − | − | − | + | + | − | − | − | − | − | − | − | − | − | − | − | − | ? | − | − | |
| 6 | DEC5A | − | − | − | − | + | + | + | + | + | + | + | + | − | − | − | − | − | − | − | − | − | ? | − | − |
| 7 | DEC5C | − | − | − | − | + | + | + | + | + | + | + | + | − | − | − | − | − | − | − | − | − | − | − | − |
| 8 | S. flexneri M90T | − | − | − | − | − | − | − | − | − | − | − | − | − | ? | − | − | − | − | − | − | + | + | + | + |
| 9 | EC503 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | − | − | − | − |
| 10 | TB285 | − | − | − | − | − | − | − | − | − | − | − | − | + | + | + | + | − | − | + | − | − | − | − | − |
| 11 | KL10035 | − | − | − | − | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − |
| 12 | KL10016 | − | − | − | − | + | − | − | − | − | ? | − | − | − | − | − | − | − | − | − | − | + | + | + | + |
| 13 | K-12 W3110 | − | − | − | − | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − |
| 14 | TB334 | − | − | − | − | − | − | − | − | − | − | − | − | + | + | + | + | − | − | + | − | − | − | − | − |
| 15 | EC535 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | − | − | − | − |
Positive (+), ambiguous (?), and negative (−) results of microbial gene detection are given. H, colony hybridization test; P, PCR product analyzed using agarose gel electrophoresis; M1 and M2, microarray tests, each using one of two independent oligonucleotide targets.
Use of PCR-generated fluorescent probes for hybridization with the microarray revealed an advantage of this approach to detect PCR products. DNA product intensities from multiplex PCR are typically highly variable and unrelated to the quantity of target sequence in the sample (14). We observed uniform fluorescent signals from the chip, signaling the qualitative presence (or absence) of the target. For example, relative amounts of slt-II-specific PCR product (484 bp) in samples 1, 2, 9, and 15 (Fig. 2) varied over a wide range, whereas the corresponding fluorescent signals did not show a detectable difference among those samples (Fig. 3). In this connection, it is very interesting to note the statistically significant detection of fliC-related sequence in the fifth sample (ECOR48). Neither radioactive probe hybridization nor agarose gel analysis of DNA products of multiplex PCR showed the presence of fliC-related sequences in this bacterial strain. Nevertheless, both fliC-specific oligonucleotide probes in the microarray clearly matched the presence of short sequences in PCR product(s) which are, if not identical, homologous to fliC-specific targets. Finally, the nonspecific PCR products visible in Fig. 2 clearly did not produce aberrant signals in the array (Fig. 3), indicating the specificity of the assay.
It is noteworthy that all oligonucleotide probes complementary to the middle part of the PCR fragment showed a lower efficiency of hybridization than did targets located closer to the end of the fragment. This observation is likely to be related to the use of double-stranded fluorescent probes. Complementary strands of the PCR product may have been able to reanneal during hybridization, displacing the oligonucleotide probe. The probability of this process seems to depend on the distance of the oligonucleotide position from the end of the DNA probe.
Specificity and sensitivity of the assay.
Thus, use of a DNA array greatly improves the specificity of an assay based on multiplex PCR. The hybridization of the fluorescently labeled PCR products was much more specific as an identifier than the size of a band on an agarose gel. Nonspecifically amplified PCR products, which might have had sequences similar to that of the targeted genes, did not interfere with the assay (Fig. 3). The sensitivity of the assay is similar to that of the original multiplex PCR. We started from pure cultures and did not attempt to develop methods to deal with interference found in clinical or environmental samples.
Thus, the hybridization of PCR products to glass microchips has been demonstrated to be a relatively fast, flexible, and reliable method to characterize a group of genetic characteristics in bacterial strains. Previously developed techniques, such as multiplex PCR, can be rapidly adapted to take advantage of the specificity and speed of hybridization analysis. The main advantage of DNA array technology is the ability to provide efficient access to vast genetic information using miniaturized DNA chips. This is particularly suited to the simultaneous determination of the presence or absence of a relatively large number of analytes by including various virulence factors and genetic markers in a bacterial strain.
Microbial pathogens are frequently identified using surrogate biochemical and immunological markers which identify the pathogen but do not always provide information about virulence factors, which frequently move horizontally on plasmids, phage, integrons, and other vectors that are independent of the surrogate chromosomal markers. Direct parallel analysis of virulence factors can be used for the identification and characterization of microbial pathogens. One of the main advantages of the microarray-based analysis for microbial identification presented here is that it can be computerized and automated. Multiple specific genes can be used to identify each organism, thus turning microbial identification into a pattern recognition process, a process that is amenable to automated, computer-based analysis. These results demonstrate the feasibility of using this approach to obtain a comprehensive snapshot of the genetic makeup of a bacterial sample that includes both phylogenetic classification and specific biochemical or pathogenic markers.
ACKNOWLEDGMENT
This work was supported in part by a grant to Konstantin Chumakov from the U.S. Defense Advanced Research Project Agency (DARPA).
REFERENCES
- 1.Altekruse S F, Cohen M L, Swerdlow D L. Emerging foodborne diseases. Emerg Infect Dis. 1997;3:285–293. doi: 10.3201/eid0303.970304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bilge S S, Vary J C, Jr, Dowell S F, Tarr P I. Role of the Escherichia coli O157:H7 O side chain in adherence and analysis of an rfb locus. Infect Immun. 1996;64:4795–4801. doi: 10.1128/iai.64.11.4795-4801.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Cebula T A, Koch W H. Analysis of spontaneous and psoralen-induced Salmonella typhimurium hisG46 revertants by oligodeoxyribonucleotide colony hybridization: use of psoralens to cross-link probes to target sequences. Mutat Res. 1990;229:79–87. doi: 10.1016/0027-5107(90)90010-2. [DOI] [PubMed] [Google Scholar]
- 4.Chart H, Cheasty T, Cope D, Gross R J, Rowe B. The serological relationship between Yersinia enterocolitica O9 and Escherichia coli O157 using sera from patients with yersiniosis and haemolytic uraemic syndrome. Epidemiol Infect. 1991;107:349–356. doi: 10.1017/s0950268800048986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Desmarchelier P M, Bilge S S, Fegan N, Mills L, Vary J C, Jr, Tarr P I. A PCR specific for Escherichia coli O157 based on the rfb locus encoding O157 lipopolysaccharide. J Clin Microbiol. 1998;36:1801–1804. doi: 10.1128/jcm.36.6.1801-1804.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Franck S M, Bosworth B T, Moon H W. Multiplex PCR for enterotoxigenic, attaching and effacing, and Shiga toxin-producing Escherichia coli strains from calves. J Clin Microbiol. 1998;36:1795–1797. doi: 10.1128/jcm.36.6.1795-1797.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Fratamico P M, Buchanan R L, Cooke P H. Virulence of an Escherichia coli O157:H7 sorbitol-positive mutant. Appl Environ Microbiol. 1993;59:4245–4252. doi: 10.1128/aem.59.12.4245-4252.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gannon V P, King R K, Kim J Y, Thomas E J. Rapid and sensitive method for detection of Shiga-like toxin-producing Escherichia coli in ground beef using the polymerase chain reaction. Appl Environ Microbiol. 1992;58:3809–3815. doi: 10.1128/aem.58.12.3809-3815.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gannon V P, D'Souza S, Graham T, King R K. Specific identification of Escherichia coli O157:H7 using a multiplex PCR assay. Adv Exp Med Biol. 1997;412:81–82. doi: 10.1007/978-1-4899-1828-4_10. [DOI] [PubMed] [Google Scholar]
- 10.Gaudio P A, Sethabutr O, Echeverria P, Hoge C W. Utility of a polymerase chain reaction diagnostic system in a study of the epidemiology of shigellosis among dysentery patients, family contacts, and well controls living in a shigellosis-endemic area. J Infect Dis. 1997;176:1013–1018. doi: 10.1086/516531. [DOI] [PubMed] [Google Scholar]
- 11.Gooding C M, Choudary P V. Comparison of different primers for rapid detection of Salmonella using the polymerase chain reaction. Mol Cell Probes. 1999;13:341–347. doi: 10.1006/mcpr.1999.0257. [DOI] [PubMed] [Google Scholar]
- 12.Gooding C M, Choudary P V. Rapid and sensitive immunomagnetic separation-polymerase chain reaction method for the detection of Escherichia coli O157:H7 in raw milk and ice-cream. J Dairy Res. 1997;64:87–93. doi: 10.1017/s0022029996001896. [DOI] [PubMed] [Google Scholar]
- 13.Hartman A B, Venkatesan M, Oaks E V, Buysse J M. Sequence and molecular characterization of a multicopy invasion plasmid antigen gene, ipaH, of Shigella flexneri. J Bacteriol. 1990;172:1905–1915. doi: 10.1128/jb.172.4.1905-1915.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hu Y, Zhang Q, Meitzler J C. Rapid and sensitive detection of Escherichia coli O157:H7 in bovine faeces by a multiplex PCR. J Appl Microbiol. 1999;87:867–876. doi: 10.1046/j.1365-2672.1999.00938.x. [DOI] [PubMed] [Google Scholar]
- 15.March S B, Ratnam S. Latex agglutination test for detection of Escherichia coli serotype O157. J Clin Microbiol. 1989;27:1675–1677. doi: 10.1128/jcm.27.7.1675-1677.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.March S B, Ratnam S. Sorbitol-MacConkey medium for detection of Escherichia coli O157:H7 associated with hemorrhagic colitis. J Clin Microbiol. 1986;23:869–872. doi: 10.1128/jcm.23.5.869-872.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Meng J, Zhao S, Doyle M P, Mitchell S E, Kresovich S. A multiplex PCR for identifying Shiga-like toxin-producing Escherichia coli O157:H7. Lett Appl Microbiol. 1997;24:172–176. doi: 10.1046/j.1472-765x.1997.00375.x. [DOI] [PubMed] [Google Scholar]
- 18.Meng J, Zhao S, Doyle M P, Mitchell S E, Kresovich S. Polymerase chain reaction for detecting Escherichia coli O157: H7. Int J Food Microbiol. 1996;32:103–113. doi: 10.1016/0168-1605(96)01110-5. [DOI] [PubMed] [Google Scholar]
- 19.Nagano I, Kunishima M, Itoh Y, Wu Z, Takahashi Y. Detection of verotoxin-producing Escherichia coli O157:H7 by multiplex polymerase chain reaction. Microbiol Immunol. 1998;42:371–376. doi: 10.1111/j.1348-0421.1998.tb02297.x. [DOI] [PubMed] [Google Scholar]
- 20.Paton A W, Paton J C. Detection and characterization of Shiga toxigenic Escherichia coli by using multiplex PCR assays for stx1, stx2, eaeA, enterohemorrhagic E. coli hlyA, rfbO111, and rfbO157. J Clin Microbiol. 1998;36:598–602. doi: 10.1128/jcm.36.2.598-602.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Paton J C, Paton A W. Pathogenesis and diagnosis of Shiga toxin-producing Escherichia coli infections. Clin Microbiol Rev. 1998;11:450–479. doi: 10.1128/cmr.11.3.450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sanderson M W, Gay J M, Hancock D D, Gay C C, Fox L K, Besser T E. Sensitivity of bacteriologic culture for detection of Escherichia coli O157:H7 in bovine feces. J Clin Microbiol. 1995;33:2616–2619. doi: 10.1128/jcm.33.10.2616-2619.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Slutsker L, Altekruse S F, Swerdlow D L. Foodborne diseases. Emerging pathogens and trends. Infect Dis Clin N Am. 1998;12:199–216. doi: 10.1016/s0891-5520(05)70418-9. [DOI] [PubMed] [Google Scholar]
- 24.Smith H R, Scotland S M. ACP Broadsheet 135: January 1993. Isolation and identification methods for Escherichia coli O157 and other Vero cytotoxin producing strains. J Clin Pathol. 1993;46:10–17. doi: 10.1136/jcp.46.1.10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Takeshi K, Ikeda T, Kubo A, Fujinaga Y, Makino S, Oguma K, Isogai E, Yoshida S, Sunagawa H, Ohyama T, Kimura H. Direct detection by PCR of Escherichia coli O157 and enteropathogens in patients with bloody diarrhea. Microbiol Immunol. 1997;41:819–822. doi: 10.1111/j.1348-0421.1997.tb01934.x. [DOI] [PubMed] [Google Scholar]
- 26.Venkateswaran K, Kamijoh Y, Ohashi E, Nakanishi H. A simple filtration technique to detect enterohemorrhagic Escherichia coli O157:H7 and its toxins in beef by multiplex PCR. Appl Environ Microbiol. 1997;63:4127–4131. doi: 10.1128/aem.63.10.4127-4131.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wang D G, Fan J B, Siao C J, Berno A, Young P, Sapolsky R, Ghandour G, Perkins N, Winchester E, Spencer J, Kruglyak L, Stein L, Hsie L, Topaloglou T, Hubbell E, Robinson E, Mittmann M, Morris M S, Shen N, Kilburn D, Rioux J, Nusbaum C, Rozen S, Hudson T J, Lander E S, et al. Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome. Science. 1998;280:1077–1082. doi: 10.1126/science.280.5366.1077. [DOI] [PubMed] [Google Scholar]
- 28.Yu J, Kaper J B. Cloning and characterization of the eae gene of enterohaemorrhagic Escherichia coli O157:H7. Mol Microbiol. 1992;6:411–417. doi: 10.1111/j.1365-2958.1992.tb01484.x. [DOI] [PubMed] [Google Scholar]