Abstract
Bone diseases such as otosclerosis (conductive hearing loss) and osteoporosis (low bone mineral density) can result from the abnormal expression of genes that regulate cartilage and bone development. The forkhead box transcription factor FOXL1 has been identified as the causative gene in a family with autosomal dominant otosclerosis and has been reported as a candidate gene in GWAS meta-analyses for osteoporosis. This potentially indicates a novel role for foxl1 in chondrogenesis, osteogenesis, and bone remodelling. We created a foxl1 mutant zebrafish strain as a model for otosclerosis and osteoporosis and examined jaw bones that are homologous to the mammalian middle ear bones, and mineralization of the axial skeleton. We demonstrate that foxl1 regulates the expression of collagen genes such as collagen type 1 alpha 1a and collagen type 11 alpha 2, and results in a delay in jawbone mineralization, while the axial skeleton remains unchanged. foxl1 may also act with other forkhead genes such as foxc1a, as loss of foxl1 in a foxc1a mutant background increases the severity of jaw calcification phenotypes when compared to each mutant alone. Our zebrafish model demonstrates atypical cartilage formation and mineralization in the zebrafish craniofacial skeleton in foxl1 mutants and demonstrates that aberrant collagen expression may underlie the development of otosclerosis.
Keywords: foxl1, foxc1, zebrafish, otosclerosis, collagen, osteoporosis, iron binding, bone mineral density
1. Introduction
Vertebrate skeletal development requires extracellular signalling cues and transcription factor expression to direct the development of cartilage, bone, and other connective tissues [1,2,3,4]. In vertebrates, cranial neural crest cells (NCCs) derived from a specialized population within the first, second, and third pharyngeal arches are responsible for developing into the progenitor cells of the craniofacial skeleton, while the paraxial mesoderm condenses into epithelial somites that give rise to the sclerotome from which the axial skeleton develops [3,5]. This cellular commitment is induced by external paracrine factors leading to pax1 and scleraxis expression [1,6], followed by subsequent expression of transcription factors (TFs), such as those of the SOX gene family in skeletal progenitors that aid in chondrogenesis and osteogenesis [7,8]. Many other well-known signaling cascades such as Hedgehog (Hh) [9,10,11,12,13,14,15], Fibroblast growth factor (Fgf) [16,17], Jagged-Notch, and Bone Morphogenic Protein (Bmp) signalling [16,18,19,20] are also required to direct progenitor cells into their specialized skeletal fates. Mutation of genes in these pathways can result in developmental defects in cartilage and bone formation/remodelling and can result in bone diseases such as otosclerosis (conductive hearing loss from abnormal middle ear bone growth) or osteoporosis (low bone density) [10,16,17,18,20].
Forkhead box transcription factors (FOX) have been previously shown to be crucial in craniofacial patterning as well as in the differentiation of chondrocytes and osteoblasts in humans, mice, and zebrafish [17,21,22,23,24,25,26]. FOX TFs contain highly conserved DNA-binding domains consisting of 110 amino acids, and may enhance or repress downstream gene expression by binding to forkhead consensus sequences in regulatory regions of the DNA [27,28]. Several FOX genes, such as those in classes C and F [25], have been studied regarding craniofacial and axial skeletal development. This prior work suggests an inherent FOX expression map resulting from the partial overlap of FOX genes to create distinct boundaries that position bone and cartilage fields in the head during development. Genes such as Foxc1 [mouse, (foxc1a in zebrafish)] and Foxc2 [mouse (foxc1b in zebrafish)] are required for chondrocyte and osteoblast differentiation, contributing to much of the expression map necessary for dermal bone formation (derived from intramembranous and endochondral ossification) in the upper and lower facial cartilages [23,25,29,30,31]. As such, they are associated with diseases such as Axenfeld–Rieger syndrome, which often presents with craniofacial abnormalities and hearing loss [21,32,33,34].
Recently, a mutation in the forkhead gene (FOXL1) was identified as the causative gene of autosomal dominant otosclerosis in a large Newfoundland family [35], and previous studies have associated this gene with osteoporosis [36,37,38,39]. In animal models, foxl1 is expressed in the paraxial mesoderm and NCCs of pharyngeal arches, and is a downstream target of both BMP and Hh signalling [40,41], yet few studies have examined foxl1’s role in bone development and remodelling of the craniofacial and axial skeleton. Given its association with two bone remodelling disorders, further study into its role in regulating skeletal development and remodelling in conjunction with other forkhead transcription factors is warranted.
Herein, we utilize a zebrafish model to determine the role of foxl1 in cartilage and bone development in the embryonic head and axial skeleton to further understand its role in disease pathogenesis. We also utilize two previously studied zebrafish forkhead mutants (foxc1a and foxc1b) [42,43] in conjunction with a new foxl1 mutant line to examine the importance of foxl1 in the FOX map. As key cartilage/bone developmental and remodelling pathways are conserved between teleosts and mammals [4,10,16,31,44,45,46,47,48,49,50], as well as aspects of the zebrafish jaw being homologous to the middle ear bones of mammals [49,51,52,53], zebrafish make an excellent model for the study of otosclerosis and osteoporosis [54,55,56,57,58]. We find that while foxl1 regulates the expression of known markers of chondro/osteogenesis, CRISPR-induced mutation of foxl1 in zebrafish only results in a delay in the formation of craniofacial cartilages and subsequent calcification, with no apparent effects on axial skeletal development. As loss of foxl1 is insufficient to overtly alter skeletal development, we assessed skeletal patterning in foxl1 mutants crossed onto foxc1a or foxc1b mutant backgrounds, which overlap in expression domains with foxl1. We find that foxl1/foxc1a double mutants exhibit an exacerbated phenotype with more severe calcification defects in the craniofacial skeleton when compared to either mutant alone, and that foxc1a and foxl1 share gene targets involved in skeletal development. Thus, we propose that foxl1 is dispensable for overall craniofacial and axial skeleton development but may act as a modifier locus for foxc1a.
2. Materials and Methods
2.1. Zebrafish Husbandry
As per Kimmel et al. [59], both wildtype (strain AB) and mutant zebrafish were reared under standard conditions and staged in terms of hours post fertilization (hpf) or days post fertilization (dpf), as described. All experiments were performed following the regulations and procedures outlined by Memorial University of Newfoundland’s Animal Care Committee and the Canadian Council on Animal Care. To prevent pigmentation and ensure optic clarity, all embryos 24 hpf and older were subject to 0.003% 1-phenyl 2-thiourea (PTU; Sigma-Aldrich, St. Louis, MO, USA). A total of 0.168 mg/mL tricaine was used to anesthetize embryos 48 hpf and older during experiments or prior to fixation with 4% paraformaldehyde (PFA; Sigma-Aldrich, St. Louis, MO, USA).
2.2. Zebrafish Strains Used in This Study
CRISPR-Cas9 sgRNAs were designed to target zebrafish foxl1 zebrafish. Two sgRNAs were designed to bind before the forkhead DNA binding domain (Supplemental Table S1) and were co-injected inducing a 52 bp deletion and shift in the open reading frame before this critical region (Supplemental Figure S1). We have designated this allele foxl1n1001. Experiments were performed by crossing heterozygous fish to produce homozygous embryos and wildtype (WT) sibling controls, or by crossing homozygous fish and WT (AB strain fish) for comparison. Previously generated foxc1aua101 and foxc1bua1018 alleles [42,43] were utilized for single and double mutant analysis.
2.3. Alcian Blue and Alizarin Red Staining
Larvae were stained at 6 and 10 dpf using an acid-free alcian blue and alizarin red double stain as previously reported [60]. Up to 50 larvae were collected in a 1.5 mL microcentrifuge tube and fixed for 2 h in 4% w/v paraformaldehyde in phosphate buffered saline pH 7.4 with gentle shaking at room temperature. After washing with 1 mL 50% v/v ethanol, larvae were gently shaken in 1 mL 50% ethanol for 10 min. The ethanol solution was removed and replaced with 1 mL of alcian blue (0.01% w/v alcian blue 8 GX, 100 mmol·L−1 MgCl2, 70% v/v ethanol) and 25 µL of alizarin red (5 mg·mL−1 alizarin red in ultrapure water). Larvae were incubated in the staining solution for 1 h at room temperature with gentle rocking. The stain solution was removed, and larvae were washed with 1 mL ultrapure water, followed by addition of 1 mL bleaching solution (1.5% v/v H2O2, 1% w/v KOH in ultrapure water). Larvae were incubated in bleaching solution for 20 min at room temperature with the microcentrifuge tubes uncapped. After removing the bleaching solution, a solution of 20% w/v glycerol and 0.25% w/v KOH was added. Larvae were gently rocked at room temperature for 30 min, before the solution was replaced with 50% w/v glycerol and 0.25% w/v KOH. Larvae were incubated overnight at 4 °C before being imaged and transferred to a storage solution (50% w/v glycerol and 0.1% w/v KOH), before being genotyped. For genotyping, individual larvae were washed twice with 100 µL PBSTw (phosphate buffered saline 0.1% w/v Tween 20) and once with 100 µL PCR-clean water. Larvae were then boiled in 50 µL 50 mmol·L−1 NaOH for 20 min at 95 °C and cooled to room temperature before addition of 5 µL 1 mol·L−1 Tris-HCl pH 8. The resulting solution was used as the template for PCR.
2.4. Whole Mount In Situ Hybridizations
In accordance with Thisse and Thisse [61], an in situ hybridization probe (foxl1) was synthesized from a pooling of whole-body RNA. PCR amplicons for probe synthesis were generated using the One Step Superscript IV RT-PCR Kit (Invitrogen). Antisense probes labelled with DIG (Roche) were constructed using an incorporated T7 RNA polymerase promoter (added at the 5′ end of the antisense primer). Proteinase K was used to permeabilize older embryos before incubation with probe: 3 min for 24 hpf, 18 min (48 hpf). DIG labelled probes were detected using alkaline phosphatase coupled anti-DIG FAB fragments, with subsequent coloration via nitro-blue tetrazolium (NBT; Roche) and 5-bromo-4-chloro-3-indolyl phosphate (BICP; Roche). Primer sequences are listed in Supplemental Table S1.
2.5. Calcein Staining
Live embryos previously growing in 0.003% PTU embryo media at ages 6 and 10 dpf were incubated in a 0.2% (w/v) calcein solution, pH 7.5 (Sigma-Aldrich, St. Louis, MO, USA), for 12 min followed by three 5-min washes in 0.003% PTU embryo media. Embryos were subsequently anesthetized in tricaine (see Zebrafish Husbandry) and mounted in 6% methylcellulose for imaging. Images were collected in the dark using a Nikon SMZ18 microscope equipped with a long-pass green filter (excitation 480 ± 40 nm; emission 510 nm). Significance testing for delayed craniofacial cartilage and bone at 6 dpf was calculated using Fisher’s exact test. Delay of cartilage development and bone calcification was characterized by decreased staining intensity, shape, and size of area (in the case of primary ossification centres), and the presence or lack thereof in comparison to their WT siblings in their respective experimental groups.
2.6. TaqMan Real-Time Quantitative PCR
Total RNA was isolated using Trizol (Invitrogen) at 6 and 10 dpf from WT sibling controls and foxl1-/- mutants, as well as at 6 dpf for foxc1a-/- and foxc1b-/- mutants and WT siblings as per Peterson and Freeman [62]. cDNA was then generated using the High-Capacity cDNA Reverse-Transcription Kit (Applied Biosystems). A total of 50–100 ng of cDNA was used in each reaction. Two biological replicates (containing a pool of 20 embryos) with three technical replicates were used with each probe and normalized to expression levels of the TATA-box binding protein (tbp) housekeeping gene. RT-qPCR runs were completed on an Applied Biosystems 7500 Real-Time PCR System or a ViiA 7 system. Data were analyzed using the ΔΔCT method [63,64]. Data bars are given as means ± standard error of the mean (SEM) with significance testing (p-values) calculated using a two-tailed t-test. Previously validated TaqMan assays were purchased from Thermo Fisher as follows: matn1 (Dr_03092841), col1a1a (Dr_03150834), col1a1b (Dr03074863), col11a2 (Dr03085627), hbae3 (Dr03125483), hpx (Dr03430535), and sp7 (Dr03133254).
2.7. DXA Scanning
Whole-body BMD assessment was performed using DXA scanning and was completed as per Green et al. [64] using a PIXImus Scanner (GE/Lunar; Madison, WI, USA) that was calibrated daily with a standard phantom. Data bars are given as means ± standard error of the mean (SEM) with significance testing (p-values) calculated using a one-way ANOVA to compare WT, foxl1 heterozygotes, and homozygotes, and a two-tailed t-test to compare WT to foxl1+/- and foxc1b+/- mutants.
3. Results
3.1. foxl1 Mutant Generation
A CRISPR-Cas9 induced mutant zebrafish strain was generated for foxl1, creating a 52 base-pair deletion in the lone foxl1 exon. Although not degraded through the nonsense mediated decay pathway due to the gene containing a single exon, the predicted protein was expected to have a shift in the reading frame that would disrupt the forkhead DNA binding domain with a premature termination codon (PTC) after amino acid 148, removing over half the protein sequence (Supplemental Figure S1). Homozygous mutants were viable and fertile, with heterozygotes and homozygotes produced at expected Mendelian frequencies.
3.2. Craniofacial Cartilage Formation and Calcification in foxl1 Mutants
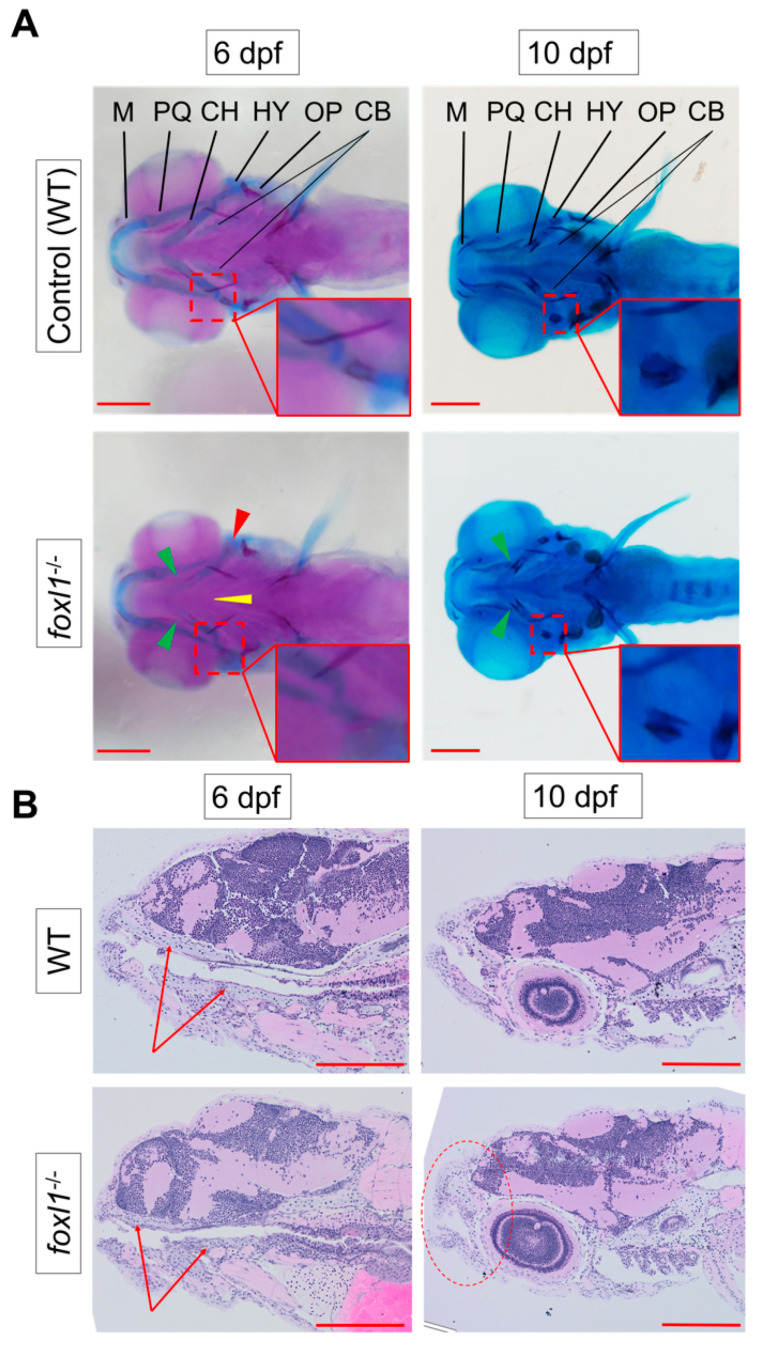
Given the association of otosclerosis with aberrant cartilage formation [65,66,67,68,69], we assessed the underlying cartilage formation of the zebrafish jaw by using alcian blue staining combined with alizarin red at 6 and 10 dpf. Homozygous foxl1 mutants exhibited a visible reduction in cartilage (alcian blue) staining of the ceratohyal (Figure 1A, green arrowheads), ceratobranchials (Figure 1A, yellow arrowhead), and hyomandibular cartilages at 6 dpf (Figure 1A, red arrowheads), which mostly recovered by 10 dpf. At 10 dpf, all skeletal elements were well stained, indicating a recovery in chondrogenesis (Figure 1A, 10 dpf).
Figure 1.
Reduction in cartilage in foxl1 mutants. (A) Alcian blue (staining cartilage) of WT and foxl1-/- (6 dpf n = 24, 10 dpf n = 20) embryos, indicating a delay in ceratohyal cartilages at 6 dpf (green arrowhead), but a recovery begins by 10 dpf. Alizarin red indicates a delay in the ossification of the hyomandibula (red arrowhead and inserts) in foxl1 mutants at 6 dpf, which also begins to normalize by 10 dpf. (B) Longitudinal sections through jaw, demonstrating a reduced presence of cartilage formation in the upper and lower jaw as indicated by the red arrows at 6 dpf, along with a shortened and round jaw structure that progresses by 10 dpf (red circle). PQ, palatoquadrate; M, Meckel; HY, hyomandibula (hyosympletic); CH, ceratohyal; CB, ceratobranchials (yellow arrowhead); OP, operculum. Zoomed-in inserts in each panel focus on the cartilage and primary centres of ossification of the hyomandibula. Scale bars are 200 µm.
Similarly, at 6 dpf, alizarin red staining for calcified bone demonstrated a reduction in calcification in foxl1 mutants. Normal calcification of the operculum is evident, while the calcification of the hyomandibula (Figure 1A, red arrowheads) and ceratohyal cartilages (Figure 1A, 6 dpf green arrowheads) were reduced at their respective primary centres of ossification, as shown by the reduced alizarin red staining. By 10 dpf, foxl1 mutants appear to have recovered with no major morphological differences observed at this developmental stage, but they still had less calcification at the primary centres of ossification in the ceratohyals than their WT counterparts, as indicated by the lesser staining (alizarin red, Figure 1A, 10 dpf green arrowheads).
As we observed a reduction in cartilage formation of the jaw in foxl1 mutants via alcian blue staining, further analysis in the form of histology with an H&E stain was used to compare jaw morphology between WT and foxl1-/- larvae at 6 and 10 dpf (Figure 1B). It was evident that the foxl1-/- mutants had reduced cartilage elements (arrows Figure 1B) and a shortened and rounded jawline that became more prominent as the embryos aged from 6 to 10 dpf (circle, Figure 1B).
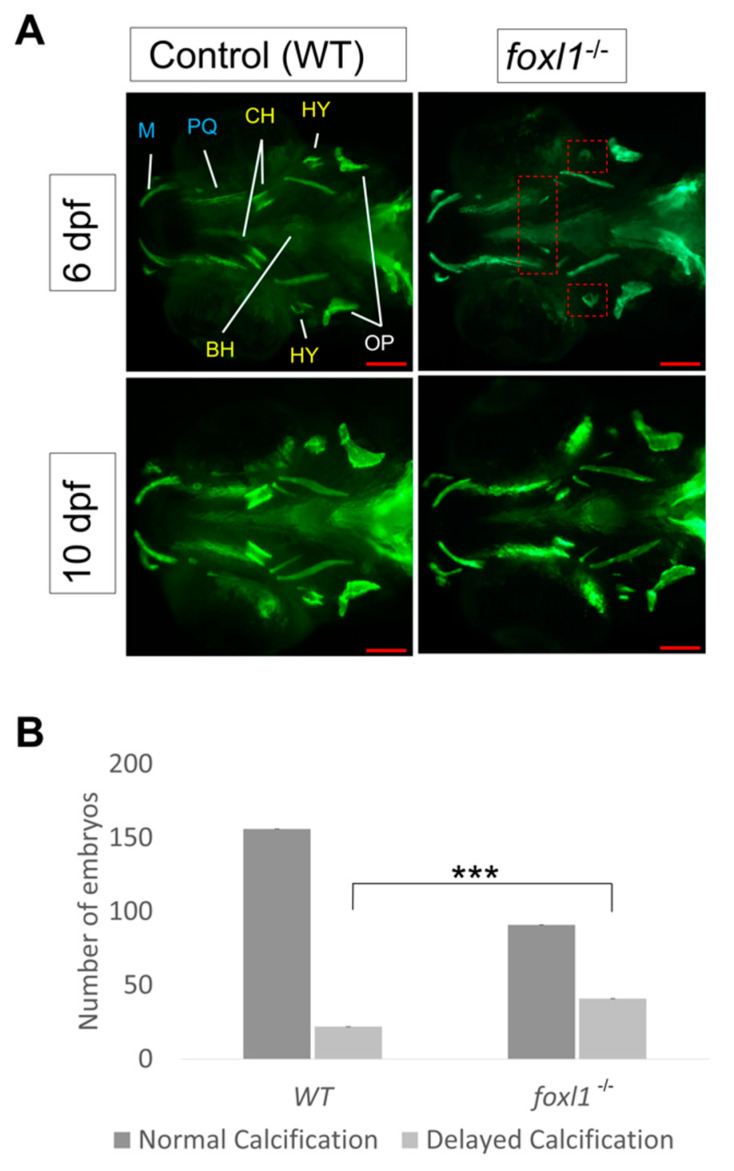
To further visualize the effect that the foxl1 mutation has on bone formation and calcification, calcein staining was performed on live embryos at both 6 and 10 dpf. The hyomandibula and ceratohyal were examined closely as they are the homologous structures of the stapes (middle ear bone) in mammals, which is often the afflicted structure in otosclerosis and was delayed in its calcification according with alizarin red staining. At 6 dpf, fewer foxl1 homozygous mutant embryos had complete calcification of the hyomandibula and ceratohyal bones when compared to wildtype siblings, as indicated by the reduced staining/presence of calcein (p = 0.0001, Figure 2A,B). Both ossification centres of the hyomandibula were observed at by 10 dpf, albeit with less intense staining, indicating a partial recovery of calcification (Figure 2A). These data agree with the alizarin red staining and indicate a delay in calcification specifically at primary centres of ossification in the developing jaw.
Figure 2.
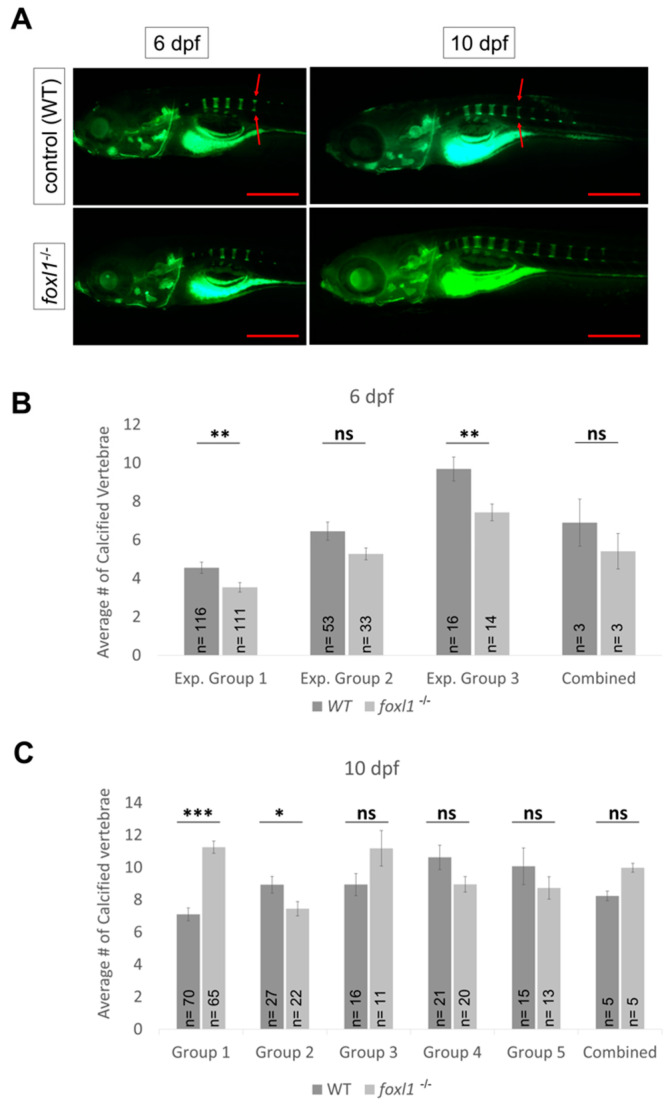
Calcein staining illustrating the impact of foxl1 loss on craniofacial development and calcification. (A) Embryos of WT aged 6 and 10 dpf (n = 173 and 99, respectively) as well as those of foxl1-/- (n = 116 and 65, respectively). WT embryos at both 6 and 10 dpf exhibit normal craniofacial development and calcification of all jaw structures. foxl1-/- embryos show a delayed calcification in the ceratohyal and hyomandibula (red boxes) at 6 dpf yet appear mostly recovered by 10 dpf. (B) The proportion of embryos with delayed calcification in foxl1 mutants is statistically significant (Fisher’s *** p = 0.0001) when compared to wildtype siblings. PQ, palatoquadrate; M, Meckel; HY, hyomandibula (hyosympletic); BH, basihyal; OP, opercula; CH, ceratohyal. Scale bars are 100 µm.
3.3. Overlapping Expression of foxl1, foxc1a, and foxc1b in Zebrafish
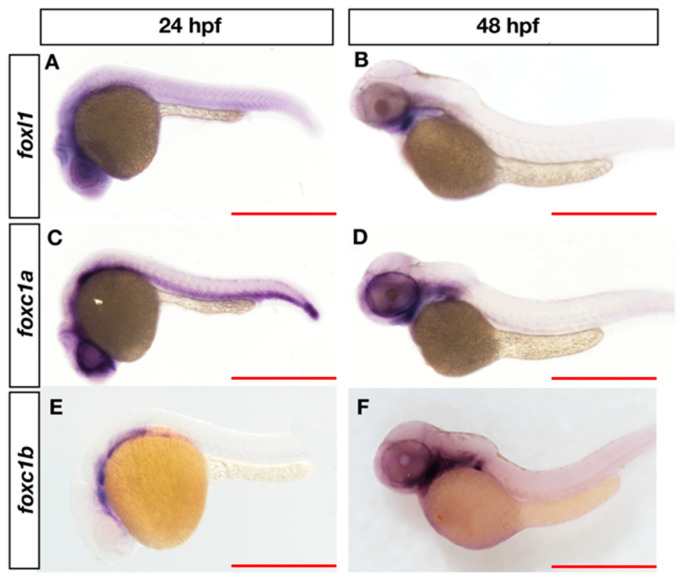
Given that phenotypes in jaw development recover in foxl1 mutants as development proceeds, we asked whether genetic redundancy could allow for overtly normal jaw development in foxl1 mutants. We assessed the expression of other forkhead genes that can cause craniofacial and axial skeletal defects in zebrafish and humans when mutated, mainly foxc1a and foxc1b (FOXC2) [24,25]. All three fox genes are expressed in the head in a pattern consistent with NCC development at 24 hpf (Figure 3A,C,E) and in the pharyngeal arches (Figure 3B,D,F) that are predominantly responsible for giving rise to Meckel’s, palatoquadrate, ceratohyal, and hyomandibular cartilages by 48 hpf. Expression in the ventral somite domains is observed for both foxl1 and foxc1a at 24 hpf.
Figure 3.
Expression of foxl1, foxc1a, and foxc1b in wildtype zebrafish at 24 and 48 hpf. At 24 hpf, foxl1 is expressed at in the brain and trunk (A), similar to foxc1a (C), while foxc1b is observed in the ventral head/brain regions (E). At 48 hpf, all three forkhead genes are expressed in the pharyngeal arches (B,D,F). Scale bars are 500 µm.
3.4. foxl1 Mutation in foxc1a and foxc1b Mutant Backgrounds
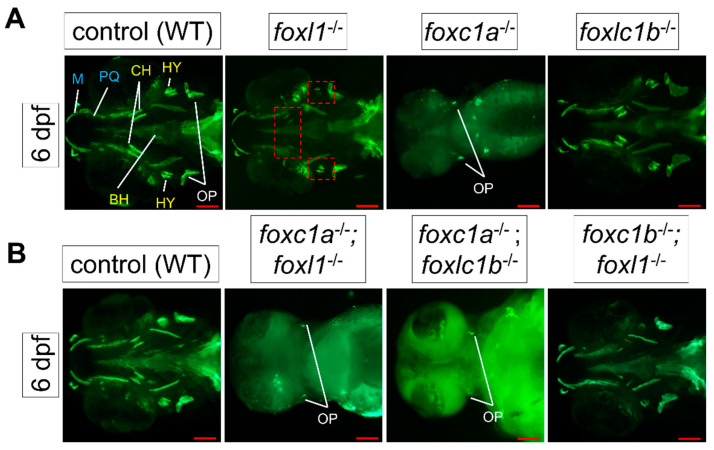
To assess the effect that foxl1 mutation has in combination with other forkhead mutants, calcein staining was performed on live embryos at 6 dpf. In agreement with other studies [24,25], foxc1b mutants did not exhibit any morphological changes in comparison to WT embryos; however, loss of foxc1a in zebrafish embryos clearly resulted in the lack of development of most major craniofacial structures such as the hyomandibula, palatoquadrate, and Meckel’s composing the developing jaw (Figure 4A). foxc1a mutants also display hydrocephalous and cardiac/abdominal edema compared to wildtype siblings. foxc1a homozygous mutants die around 7 dpf.
Figure 4.
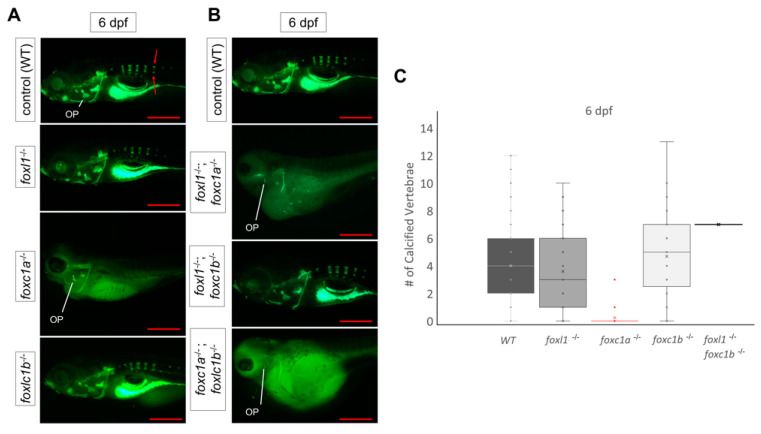
Calcein staining illustrating the impact of foxl1, foxc1a, and foxc1b lone and combined loss on craniofacial development and calcification. (A) foxc1a-/- embryos have a lack of bone development in all craniofacial bones resulting from the first (blue) and second (yellow) pharyngeal arches and diminished formation of the opercula bones at 6 dpf. foxc1b-/- embryos exhibit no change. foxl1 panels are included for reference. (B) WT (n = 173); foxc1a-/-; foxl1-/- (n = 6); foxc1b-/-; foxl1-/- (n = 2); and foxc1a-/-; foxc1b-/- (n = 4) embryos at 6 dpf. foxc1a-/-; foxl1-/- embryos exhibit a further loss of craniofacial bone formation and calcification with increased size in the cardiac edema present, while foxc1b-/-; foxl1-/- embryos exhibit no change from their respective individual knockout models. foxc1a-/-; foxc1b-/- double mutants show a similar phenotype as foxc1a-/-; foxl1-/- embryos. Red boxes highlight areas of interest. Small red squares highlight the hyomandibula and the large rectangular box isolates the ceratohyal bone. Scale bars are 100 µm.
Double foxc1a-/-; foxl1-/- mutants (Figure 4B) exhibited reduced bone formation, including a lack of calcification of the operculum, when compared to foxc1a mutants alone which exhibit mostly normal operculum calcification. Unlike foxc1a-/-; foxl1-/- double mutants, foxl1-/-; foxc1b-/- embryos did not exhibit an abnormal phenotype in comparison to the wildtype controls (Figure 4B). foxc1a-/-; foxc1b-/- double mutant embryos had similar phenotypes as the foxc1a-/-; foxl1-/- embryos at 6 dpf, which included further loss of bone calcification in the face (Figure 4B).
3.5. Axial Skeletal Calcification in Forkhead Mutants
In humans, variants in FOXL1 and FOXC2 are associated with a reduced bone mineral density (BMD) and a risk of osteoporosis [35,36,39]. Furthermore, the co-occurrence of otosclerosis and osteoporosis has been noted [65], suggesting similar aetiologies for both bone remodelling diseases. We thus tested foxl1, foxc1a, and foxc1b for defects in the calcification of the zebrafish spine to determine if either gene affects calcification of the axial skeleton. Mutation of foxl1 has little effect on the patterning and formation of the vertebrae of the skeleton at 6 and 10 dpf when compared with WT controls (Figure 5), as vertebral calcification appeared to be variable within experimental groups (Figure 5B,C). This could possibly be due to environmental changes or natural variability in bone formation. This led to an overall decrease in the number of calcified vertebrae at 6 dpf, but an increased trend in the rate of vertebrae calcification on average in foxl1 mutants in comparison with WT siblings at 10 dpf (Figure 5B,C). The average number of vertebrates calcified (based on primary centre of ossification) for WT and foxl1-/- was 5 and 4 approximately by 6 dpf (with no change in body length; Supplemental Figure S3) and 8 and 10 approximately by 10 dpf (Figure 5B,C).
Figure 5.
Calcein staining illustrating the impact of foxl1 loss on axial skeletal development and calcification. (A) WT and foxl1-/- embryos at 6 and 10 dpf showing normal formation of the vertebrae in foxl1. (B,C) Experimental groups of calcein staining illustrating the variability in calcification of the zebrafish vertebrate between WT and foxl1 homozygotes at 6 and 10 dpf, respectively (* p < 0.05, ** p < 0.01, *** p < 0.001), showing an increased trend in number of calcified vertebrae in foxl1 mutants by 10 dpf. Total number of calcified vertebrae were counted by including both complete and partially calcified vertebrae [partially calcified vertebrae are indicated by disconnected primary and secondary centres of ossification (red arrows)]. Scale bars are 500 µm.
foxc1a mutants lack any axial skeleton formation at 6 dpf while presenting with edemas around the heart and body cavity that grew over the course of time (Figure 6A), thus preventing the analysis of spine calcification in foxl1-/-, foxc1a-/- mutants entirely (Figure 6B). It should be noted, however, that the lateral views presented in Figure 6 reiterate the increased severity of craniofacial phenotypes observed in foxl1-/-, foxc1a-/- mutants, including decreased calcification of the operculum previously shown in in Figure 4. foxc1b mutant embryos did not exhibit any abnormal phenotypes regarding axial skeletal development at 6 dpf (Figure 6A), nor did double foxl1-/-, foxc1b-/- mutants (Figure 6B).
Figure 6.
Calcein staining illustrating the impact of foxl1, foxc1a, and foxc1b lone and combined loss on axial skeleton development and calcification. (A) WT, foxl1-/-, foxc1a-/-, and foxc1b-/- embryos at 6 dpf showing normal formation of the vertebrae in foxl1 and foxc1b mutants. foxc1a-/- embryos did not develop or calcify vertebrae and had large abdomen cavity edemas that grew over time. (B) WT; foxc1a-/-; foxl1-/-, foxc1b-/-; foxl1-/-, and foxc1a-/-; foxc1b-/- embryos at 6 dpf. Double foxc1a-/-; foxl1-/- embryos exhibit larger edemas and a lack of most calcified structures at 6 dpf, including the operculum (OP). foxc1b-/-; foxl1-/- embryos had no significant change. foxc1a-/-; foxc1b-/- mutants elicited similar results as foxc1a-/-; foxl1-/- embryos, including edemas and lack of craniofacial calcification. (C) Boxplots illustrating the variability in the number of vertebrae calcified in foxl1-/-, foxc1a-/-, foxc1b-/-,and foxl1-/-; foxc1b-/- mutants in comparison with WT embryos at 6 dpf. Red arrows indicate primary centres of ossification within the vertebrae. Scale bars are 500 µm.
To further investigate the calcification of the zebrafish skeleton, dual-energy X-ray absorptiometry (DXA) was performed on foxl1 mutants to measure bone mineral density as per Green et al. (Supplemental Figure S2) [70]. No significant difference was found between WT siblings and heterozygous or homozygous foxl1 mutants alone at 5.5 months old (Supplemental Figure S2A). As both foxl1 and foxc1b have been associated with osteoporosis, we also assessed foxl1+/-; foxc1b+/- double heterozygous mutants and WT controls at 1 year of age, and similarly, no difference in their BMD was observed (Supplemental Figure S2B). However, our foxl1 homozygous mutants exhibited wider heads and jaws along with longer jaws in comparison to their WT counterparts (Supplemental Figure S3A,B).
3.6. Expression of Genes Required for Bone Formation in Forkhead Mutants
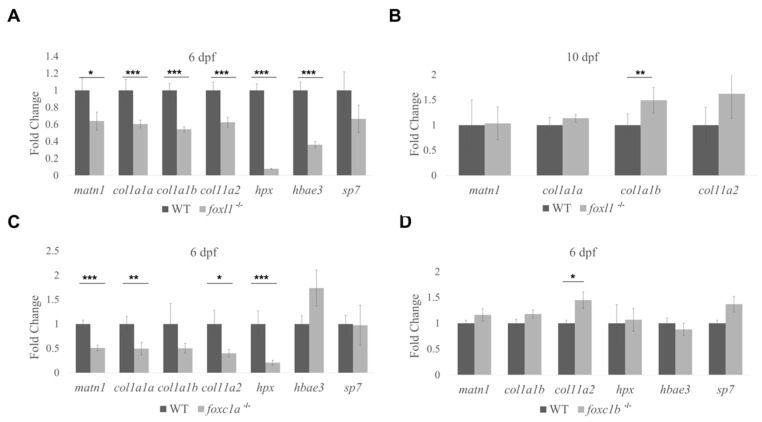
To further demonstrate the role of foxl1 on skeletal development, we assessed the expression of genes with known roles in cartilage and bone formation in zebrafish foxl1, foxc1a, and foxc1b mutants. To determine candidate target genes of foxl1, an RNA-Seq screen was completed on WT and foxl1-/- embryos at 6 dpf. Prominent genes with differential expression were then validated using TaqMan Real-Time quantitative PCR (RT-qPCR). At 6 dpf, we identified markers of cartilage and heme-binding proteins as differentially expressed (Figure 7). Chondrocyte markers such as matrilin 1 (matn1, 36% reduction), collagen type 1 alpha 1a (col1a1a, 40% reduction), collagen type 1 alpha 1b (col1a1b, 46% reduction), and collagen type 11 alpha 2 (col11a2, 38% reduction) saw statistically significant (p < 0.05) reductions in gene expression based on total whole embryo RNA (Figure 7A). hemoglobin alpha embryonic-3 (hbae3, 64% reduction) and hemopexin (hpx, 92% reduction), both important factors in heme-binding and transport, were even more so reduced in foxl1 mutants. osterix (sp7), a marker of osteoblast differentiation, was downregulated in foxl1 mutants but did not reach statistical significance (Figure 7A). These data demonstrate the reduction in genes required for collagen and bone formation at 6 dpf, consistent with the delay in chondrogenesis and subsequent calcification of aspects of the craniofacial skeleton at this developmental timepoint. To further illustrate the genetic redundancy that may allow overtly normal jaw development in the absence of foxl1, we tested the expression of the same genes in foxc1a and foxc1b mutants. Much like foxl1 mutants, foxc1a-/- embryos saw significant reduction in matrilin 1, collagen type 1 alpha 1a, collagen type 11 alpha 2, and hemopexin, indicating target gene overlap (Figure 7C). Intriguingly foxc1b-/- embryos saw an increase in collagen type 11 alpha 2 (Figure 7C,D), suggesting it acts as a repressive regulator in comparison to the enhancing mechanism of foxl1 and foxc1a.
Figure 7.
Differential gene expression in forkhead mutants. foxl1-/- (A), foxc1a-/- (C), and foxc1b-/- (D) embryos at 6 dpf using TaqMan RT-qPCR to confirm genes of interest from RNA-Seq screen. Targets of interest were examined at 10 dpf in foxl1 mutants (B), demonstrating a recovery in gene expression. Fold change between WT and foxl1-/- plotted with WT values set to 1 (* p < 0.05, ** p < 0.01, *** p < 0.001).
As a recovery was observed in the formation of craniofacial cartilages and bone in foxl1-/- embryos by 10 dpf, the gene expression of matrilin 1, collagen type 1 alpha 1a, and collagen type 11 alpha 2 were assessed to determine the source of recovery. All genes saw a return to WT level gene expression, notably with col1a1b significantly overexpressed in direct contrast to its previous reduction at 6 dpf (Figure 7B). This indicates that despite foxl1 loss, increased gene expression of necessary factors is upregulated by other means to achieve a largely normal skeleton throughout the maturation of the zebrafish larvae.
4. Discussion
Here, we report on a foxl1 mutant strain in zebrafish as a potential model of otosclerosis and osteoporosis. Structures in the mammalian middle ear are homologous with the ceratohyal, palatoquadrate, and hyomandibular jaw bones in zebrafish [48,51,52,53,71] and as such, the malformation of these bones leads to the loss of sound conduction in mammals and jaw formation in zebrafish. Despite abundant foxl1 expression in the pharyngeal arches and a delay in cartilage formation and calcification in the jaw, loss of foxl1 does not significantly affect the overall structure of the craniofacial skeleton, consistent with recent reports using strain harboring a foxl1 nonsense mutation [25]. In agreement with a delay in cartilage formation and calcification at 6 dpf, we do observe a reduction in the expression of cartilage markers such as matrillin1 and col1a1a/b in foxl1 homozygous mutants at this developmental time point. In accordance with the partial phenotypic recovery seen by 10 dpf, we also see a recovery in gene expression. Conversely, previous foxl1 morpholino studies have shown much more pronounced phenotypes including abnormalities in craniofacial skeleton, midbrain, eye, and pectoral fin [72]. While these discrepancies may be due to morpholino off-target effects, genetic compensation in CRISPR-generated INDEL mutants [73,74,75] may also play a role.
While FOXL1 (and FOXC2) have been associated with osteoporosis in humans, and the co-occurrence of otosclerosis and osteoporosis has been reported [65,66,76], we did not find any defects in development of the larval zebrafish axial skeleton in foxl1 or foxc1b (FOXC2 homolog) mutants. The axial skeleton in foxl1-/- mutants showed an increased trend of vertebrae calcified by 10 dpf but remained largely unaffected by the loss of foxl1. It is also possible that foxl1 may play a specific role in endochondral ossification, which includes the jaw, but may not regulate intramembranous bone development that occurs in the zebrafish spine. No differences in BMD are observed in adult foxl1 mutant zebrafish using DXA scanning; however, detection of BMD in fish using this method may not be sensitive enough to determine small changes in BMD, and only relatively young fish (5.5 months of age) were tested in this study. Additional studies assessing BMD in varying ages of adult fish through more sensitive µCT imaging could determine whether foxl1 mutants have defects in maintaining BMD as they age.
Since foxl1 has been shown to be expressed in cartilage and bone-specific tissues and is associated with bone remodelling diseases such as otosclerosis and osteoporosis [35,36,57], it is interesting that the loss of foxl1 does not cause such profound bone phenotypes in the head or axial skeleton in the developing zebrafish. This indicates that, as with many other FOX genes, foxl1 may only act as minor genetic modifier in development [28,72,77,78]. Our data indicate that when mutated alone, the development of bone and cartilage is predominantly normal in zebrafish larvae, which only exhibit minor delays by 10 dpf. However, when mutated in conjunction with foxc1a, another FOX gene known to be central in upper craniofacial cartilage formation and somite patterning, further loss of bone is observed than in foxc1a mutants alone. In zebrafish, bones in the craniofacial skeleton of foxc1a mutants that began to calcify, such as the operculum, did not form or were even more malformed in foxl1-/-; foxc1a-/- mutants. Taken together with qPCR data indicating that foxl1 shares similar targets as foxc1a, the results support a model whereby foxl1 may act as a modifier locus for foxc1a. Alternatively, the increased severity of phenotypes through the combined loss of foxl1 and foxc1a may represent reduced growth and failure to thrive when compared to foxc1a mutants alone, as opposed to a specific effect on bone development. This possibility should be explored further. Heterozygous loss of FOXC1 in humans results in Axenfeld–Rieger syndrome (ARS), which results in increased risk for glaucoma and variable systemic anomalies including defects in craniofacial bone development [79]. Recently, an atypical FOXC1-attributable ARS patient was described with clinical otosclerosis [33], again highlighting the potential overlapping functions of foxl1 and foxc1a.
Another novel hypothesis indicating foxl1’s possible involvement in bone development and maintenance is its regulation of hemopexin, which is responsible for high-affinity heme-binding and transport to the liver for degradation, preventing oxidative stress and iron loss [80,81] as well as hemoglobin alpha embryonic-3 that is involved up-stream of the oxygen transport chain as an early marker of erythrocyte development [82]. Both genes were significantly downregulated in foxl1-/- mutants, and there is a known association between heme-transport/blood diseases such as anemia and low BMD leading to fracture over time [83,84,85]. This could suggest that a loss of foxl1 may cause an increase in oxidative stress, which in turn may exacerbate the mild cartilage and bone phenotypes observed during growth and remodelling. This hypothesis warrants further investigation.
5. Conclusions
In zebrafish, foxl1 is required for the expression of key collagen genes but plays only a minor role in the formation and maintenance of zebrafish jaw cartilages and bones. Given the homology of zebrafish jawbones and mammalian middle ear bones, foxl1 mutant zebrafish serve as a useful model to gain an understanding of the mechanistic insights regarding the pathophysiology of otosclerosis. Additional analyses will be needed to determine if BMD defects occur in foxl1 mutant zebrafish and whether they may serve as a useful model of osteoporosis. Our study supports the hypothesis that foxl1 and foxc1a function together to regulate bone development; however, the manner in which foxl1 and foxc1a function together to achieve optimal expression of cartilage and bone markers has yet to be determined and needs further exploration. This study highlights a new foxl1 mutant zebrafish line, finding novel target genes of foxl1 that could be utilized for new therapies to treat otosclerosis.
Acknowledgments
We would like to thank the animal care staff at Memorial University for maintaining zebrafish stocks, Jules Dore (Memorial University) for help with qPCR, and Ordan Lehmann (University of Alberta) for providing foxc1a and foxc1b mutant lines. We would also like to thank the Histology Services Unit at Memorial University for help with sectioning and H&E staining.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/genes13071107/s1, Table S1: Primer and sgRNA sequences used in this study, Figure S1: CRISPR-induced mutation of foxl in zebrafish, Figure S2: Loss of foxl1 does not affect bone density in adult zebrafish, Figure S3: Loss of foxl1 results in increased head and jaw width, and jaw length, but does not affect body length.
Author Contributions
A.H.-N. performed calcein staining in all mutants, qPCR, and histological analysis and wrote the manuscript. J.A.P. created the foxl1 mutant strain and performed a preliminary phenotypic analysis and RNA sequencing screen. R.K. performed alcian blue/alizarin red staining. M.F. performed in situ analysis in Supplemental Figure S1. A.S.M. and C.S.K. aided in BMD analysis in Supplemental Figure S2. T.-L.Y. was involved in creating foxl1 mutants and edited the manuscript. C.R.F. is the Principal Investigator and participated in study design and manuscript editing. All authors have read and agreed to the published version of the manuscript.
Institutional Review Board Statement
This study was approved by Memorial University’s Animal Ethics Committee.
Informed Consent Statement
Not applicable.
Data Availability Statement
All available raw data are available from the Principal Investigator, Cutis R. French.
Conflicts of Interest
The authors declare no conflict of interest.
Funding Statement
This research was funded through the Medical Research Fund at Memorial University (Curtis French) and the Atlantic Canada Opportunities Agency—Community Innovation Fund (Terry-Lynn Young). Alexia Hawkey-Noble is supported by the Dean’s Fellowship (Memorial University).
Footnotes
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Berendsen A.D., Olsen B.R. Bone Development. Bone. 2015;80:14–18. doi: 10.1016/j.bone.2015.04.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Fleming A., Kishida M.G., Kimmel C.B., Keynes R.J. Building the Backbone: The Development and Evolution of Vertebral Patterning. Development. 2015;142:1733–1744. doi: 10.1242/dev.118950. [DOI] [PubMed] [Google Scholar]
- 3.Javidan Y., Schilling T.F. Methods in Cell Biology. Volume 76. Elsevier; Amsterdam, The Netherlands: 2004. Development of Cartilage and Bone; pp. 415–436. [DOI] [PubMed] [Google Scholar]
- 4.Kozhemyakina E., Lassar A.B., Zelzer E. A Pathway to Bone: Signaling Molecules and Transcription Factors Involved in Chondrocyte Development and Maturation. Development. 2015;142:817–831. doi: 10.1242/dev.105536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dash S., Trainor P.A. The Development, Patterning and Evolution of Neural Crest Cell Differentiation into Cartilage and Bone. Bone. 2020;137:115409. doi: 10.1016/j.bone.2020.115409. [DOI] [PubMed] [Google Scholar]
- 6.Winnier G.E., Hargett L., Hogan B.L. The Winged Helix Transcription Factor MFH1 Is Required for Proliferation and Patterning of Paraxial Mesoderm in the Mouse Embryo. Genes Dev. 1997;11:926–940. doi: 10.1101/gad.11.7.926. [DOI] [PubMed] [Google Scholar]
- 7.Lefebvre V. Roles and Regulation of SOX Transcription Factors in Skeletogenesis. Curr. Top. Dev. Biol. 2019;133:171–193. doi: 10.1016/bs.ctdb.2019.01.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Long F., Ornitz D.M. Development of the Endochondral Skeleton. Cold Spring Harb. Perspect. Biol. 2013;5:a008334. doi: 10.1101/cshperspect.a008334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Abe Y., Tanaka N., Abe Y., Tanaka N. Roles of the Hedgehog Signaling Pathway in Epidermal and Hair Follicle Development, Homeostasis, and Cancer. J. Dev. Biol. 2017;5:12. doi: 10.3390/jdb5040012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ahlgren S.C., Bronner-Fraser M. Inhibition of Sonic Hedgehog Signaling in Vivo Results in Craniofacial Neural Crest Cell Death. Curr. Biol. 1999;9:1304–1314. doi: 10.1016/S0960-9822(00)80052-4. [DOI] [PubMed] [Google Scholar]
- 11.Armstrong B.E., Henner A., Stewart S., Stankunas K. Shh Promotes Direct Interactions between Epidermal Cells and Osteoblast Progenitors to Shape Regenerated Zebrafish Bone. Development. 2017;144:1165–1176. doi: 10.1242/dev.143792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Avaron F., Smith A., Akimenko M.-A. Sonic Hedgehog Signaling in the Developing and Regenerating Fins of Zebrafish. Landes Bioscience; Austin, TX, USA: 2013. [Google Scholar]
- 13.Carballo G.B., Honorato J.R., de Lopes G.P.F., Spohr T.C.L. de S. e A Highlight on Sonic Hedgehog Pathway. Cell Commun. Signal. 2018;16:11. doi: 10.1186/s12964-018-0220-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chung U., Schipani E., McMahon A.P., Kronenberg H.M. Indian Hedgehog Couples Chondrogenesis to Osteogenesis in Endochondral Bone Development. J. Clin. Investig. 2001;107:295–304. doi: 10.1172/JCI11706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Eberhart J.K., Swartz M.E., Crump J.G., Kimmel C.B. Early Hedgehog Signaling from Neural to Oral Epithelium Organizes Anterior Craniofacial Development. Development. 2006;133:1069–1077. doi: 10.1242/dev.02281. [DOI] [PubMed] [Google Scholar]
- 16.Katoh M. Networking of WNT, FGF, Notch, BMP, and Hedgehog Signaling Pathways during Carcinogenesis. Stem Cell Rev. Totowa. 2007;3:30–38. doi: 10.1007/s12015-007-0006-6. [DOI] [PubMed] [Google Scholar]
- 17.Rice R., Rice D.P.C., Thesleff I. Foxc1 Integrates Fgf and Bmp Signalling Independently of Twist or Noggin during Calvarial Bone Development. Dev. Dyn. 2005;233:847–852. doi: 10.1002/dvdy.20430. [DOI] [PubMed] [Google Scholar]
- 18.Asharani P.V., Keupp K., Semler O., Wang W., Li Y., Thiele H., Yigit G., Pohl E., Becker J., Frommolt P., et al. Attenuated BMP1 Function Compromises Osteogenesis, Leading to Bone Fragility in Humans and Zebrafish. Am. J. Hum. Genet. 2012;90:661–674. doi: 10.1016/j.ajhg.2012.02.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Katoh M., Katoh M. Transcriptional Regulation of WNT2B Based on the Balance of Hedgehog, Notch, BMP and WNT Signals. Int. J. Oncol. 2009;34:1411–1415. doi: 10.3892/ijo_00000269. [DOI] [PubMed] [Google Scholar]
- 20.Wan M., Cao X. BMP Signaling in Skeletal Development. Biochem. Biophys. Res. Commun. 2005;328:651–657. doi: 10.1016/j.bbrc.2004.11.067. [DOI] [PubMed] [Google Scholar]
- 21.Chrystal P.W., French C.R., Jean F., Havrylov S., van Baarle S., Peturson A.-M., Xu P., Crump J.G., Pilgrim D.B., Lehmann O.J., et al. The Axenfeld-Rieger Syndrome Gene FOXC1 Contributes to Left-Right Patterning. Genes. 2021;12:170. doi: 10.3390/genes12020170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kim S.H., Cho K.-W., Choi H.S., Park S.J., Rhee Y., Jung H.-S., Lim S.-K. The Forkhead Transcription Factor Foxc2 Stimulates Osteoblast Differentiation. Biochem. Biophys. Res. Commun. 2009;386:532–536. doi: 10.1016/j.bbrc.2009.06.071. [DOI] [PubMed] [Google Scholar]
- 23.Kume T., Jiang H., Topczewska J.M., Hogan B.L.M. The Murine Winged Helix Transcription Factors, Foxc1 and Foxc2, Are Both Required for Cardiovascular Development and Somitogenesis. Genes Dev. 2001;15:2470–2482. doi: 10.1101/gad.907301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Xu P., Yu H.V., Tseng K.-C., Flath M., Fabian P., Segil N., Crump J.G. Foxc1 Establishes Enhancer Accessibility for Craniofacial Cartilage Differentiation. eLife. 2021;10:e63595. doi: 10.7554/eLife.63595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Xu P., Balczerski B., Ciozda A., Louie K., Oralova V., Huysseune A., Crump J.G. Fox Proteins Are Modular Competency Factors for Facial Cartilage and Tooth Specification. Development. 2018;145:dev.165498. doi: 10.1242/dev.165498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.You W., Gao H., Fan L., Duan D., Wang C., Wang K. Foxc2 Regulates Osteogenesis and Angiogenesis of Bone Marrow Mesenchymal Stem Cells. BMC Musculoskelet. Disord. 2013;14:199. doi: 10.1186/1471-2474-14-199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Dai S., Qu L., Li J., Chen Y. Toward a Mechanistic Understanding of DNA Binding by Forkhead Transcription Factors and Its Perturbation by Pathogenic Mutations. Nucleic Acids Res. 2021;49:10235–10249. doi: 10.1093/nar/gkab807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hannenhalli S., Kaestner K.H. The Evolution of Fox Genes and Their Role in Development and Disease. Nat. Rev. Genet. 2009;10:233–240. doi: 10.1038/nrg2523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nifuji A., Miura N., Kato N., Kellermann O., Noda M. Bone Morphogenetic Protein Regulation of Forkhead/Winged Helix Transcription Factor Foxc2 (Mfh1) in a Murine Mesodermal Cell Line C1 and in Skeletal Precursor Cells. J. Bone Miner. Res. Off. J. Am. Soc. Bone Miner. Res. 2001;16:1765–1771. doi: 10.1359/jbmr.2001.16.10.1765. [DOI] [PubMed] [Google Scholar]
- 30.Sun J., Ishii M., Ting M.-C., Maxson R. Foxc1 Controls the Growth of the Murine Frontal Bone Rudiment by Direct Regulation of a Bmp Response Threshold of Msx2. Dev. Camb. Engl. 2013;140:1034–1044. doi: 10.1242/dev.085225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Mariotti M., Carnovali M., Banfi G. Danio Rerio: The Janus of the Bone from Embryo to Scale. Clin. Cases Miner. Bone Metab. 2015;12:188–194. doi: 10.11138/ccmbm/2015.12.2.188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Tümer Z., Bach-Holm D. Axenfeld–Rieger Syndrome and Spectrum of PITX2 and FOXC1 Mutations. Eur. J. Hum. Genet. 2009;17:1527–1539. doi: 10.1038/ejhg.2009.93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang R., Wang W.-Q., Li X.-Q., Zhao J., Yang K., Feng Y., Guo M.-M., Liu M., Liu X., Wang X., et al. A Novel Variant in FOXC1 Associated with Atypical Axenfeld-Rieger Syndrome. BMC Med. Genom. 2021;14:277. doi: 10.1186/s12920-021-01130-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.French C.R. Mechanistic Insights into Axenfeld-Rieger Syndrome from Zebrafish Foxc1 and Pitx2 Mutants. Int. J. Mol. Sci. 2021;22:10001. doi: 10.3390/ijms221810001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Abdelfatah N., Mostafa A.A., French C.R., Doucette L.P., Penney C., Lucas M.B., Griffin A., Booth V., Rowley C., Besaw J.E., et al. A Pathogenic Deletion in Forkhead Box L1 (FOXL1) Identifies the First Otosclerosis (OTSC) Gene. Hum. Genet. 2021;141:965–979. doi: 10.1007/s00439-021-02381-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kim B.-J., Ahn S.H., Kim H.-M., Ikegawa S., Yang T.-L., Guo Y., Deng H.-W., Koh J.-M., Lee S.H. Replication of Caucasian Loci Associated with Osteoporosis-Related Traits in East Asians. J. Bone Metab. 2016;23:233–242. doi: 10.11005/jbm.2016.23.4.233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kung A.W.C. Novel Genetic Loci Associated with Osteoporosis. Bone. 2010;47:S376. doi: 10.1016/j.bone.2010.09.058. [DOI] [Google Scholar]
- 38.Liu Y.-J., Zhang L., Papasian C.J., Deng H.-W. Genome-Wide Association Studies for Osteoporosis: A 2013 Update. J. Bone Metab. 2014;21:99–116. doi: 10.11005/jbm.2014.21.2.99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.the Genetic Factors for Osteoporosis (GEFOS) Consortium. Rivadeneira F., Styrkársdottir U., Estrada K., Halldórsson B.V., Hsu Y.-H., Richards J.B., Zillikens M.C., Kavvoura F.K., Amin N., et al. Twenty Bone-Mineral-Density Loci Identified by Large-Scale Meta-Analysis of Genome-Wide Association Studies. Nat. Genet. 2009;41:1199–1206. doi: 10.1038/ng.446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Aoki R., Shoshkes-Carmel M., Gao N., Shin S., May C.L., Golson M.L., Zahm A.M., Ray M., Wiser C.L., Wright C.V.E., et al. Foxl1-Expressing Mesenchymal Cells Constitute the Intestinal Stem Cell Niche. Cell. Mol. Gastroenterol. Hepatol. 2016;2:175–188. doi: 10.1016/j.jcmgh.2015.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Madison B.B., McKenna L.B., Dolson D., Epstein D.J., Kaestner K.H. FoxF1 and FoxL1 Link Hedgehog Signaling and the Control of Epithelial Proliferation in the Developing Stomach and Intestine. J. Biol. Chem. 2009;284:5936–5944. doi: 10.1074/jbc.M808103200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Umali J., Hawkey-Noble A., French C.R. Loss of Foxc1 in Zebrafish Reduces Optic Nerve Size and Cell Number in the Retinal Ganglion Cell Layer. Vision Res. 2019;156:66–72. doi: 10.1016/j.visres.2019.01.008. [DOI] [PubMed] [Google Scholar]
- 43.Whitesell T.R., Chrystal P.W., Ryu J.-R., Munsie N., Grosse A., French C.R., Workentine M.L., Li R., Zhu L.J., Waskiewicz A., et al. Foxc1 Is Required for Embryonic Head Vascular Smooth Muscle Differentiation in Zebrafish. Dev. Biol. 2019;453:34–47. doi: 10.1016/j.ydbio.2019.06.005. [DOI] [PubMed] [Google Scholar]
- 44.Mackay E.W., Apschner A., Schulte-Merker S. A Bone to Pick with Zebrafish. BoneKEy Rep. 2013;2:445. doi: 10.1038/bonekey.2013.179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Witten P.E., Harris M.P., Huysseune A., Winkler C. Chapter 13—Small Teleost Fish Provide New Insights into Human Skeletal Diseases. In: Detrich H.W., Westerfield M., Zon L.I., editors. Methods in Cell Biology. Volume 138. Academic Press; London, UK: 2017. pp. 321–346. The Zebrafish. [DOI] [PubMed] [Google Scholar]
- 46.Nikaido M., Tada M., Saji T., Ueno N. Conservation of BMP Signaling in Zebrafish Mesoderm Patterning. Mech. Dev. 1997;61:75–88. doi: 10.1016/S0925-4773(96)00625-9. [DOI] [PubMed] [Google Scholar]
- 47.Wada N., Javidan Y., Nelson S., Carney T.J., Kelsh R.N., Schilling T.F. Hedgehog Signaling Is Required for Cranial Neural Crest Morphogenesis and Chondrogenesis at the Midline in the Zebrafish Skull. Development. 2005;132:3977–3988. doi: 10.1242/dev.01943. [DOI] [PubMed] [Google Scholar]
- 48.Raterman S.T., Metz J.R., Wagener F.A.D.T.G., Von den Hoff J.W. Zebrafish Models of Craniofacial Malformations: Interactions of Environmental Factors. Front. Cell Dev. Biol. 2020;8:1346. doi: 10.3389/fcell.2020.600926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Mork L., Crump G. Zebrafish Craniofacial Development: A Window into Early Patterning. Curr. Top. Dev. Biol. 2015;115:235–269. doi: 10.1016/bs.ctdb.2015.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Bergen D.J.M., Kague E., Hammond C.L. Zebrafish as an Emerging Model for Osteoporosis: A Primary Testing Platform for Screening New Osteo-Active Compounds. Front. Endocrinol. 2019;10:6. doi: 10.3389/fendo.2019.00006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Anthwal N., Joshi L., Tucker A.S. Evolution of the Mammalian Middle Ear and Jaw: Adaptations and Novel Structures. J. Anat. 2013;222:147–160. doi: 10.1111/j.1469-7580.2012.01526.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Takechi M., Kuratani S. History of Studies on Mammalian Middle Ear Evolution: A Comparative Morphological and Developmental Biology Perspective. J. Exp. Zoolog. B Mol. Dev. Evol. 2010;314B:417–433. doi: 10.1002/jez.b.21347. [DOI] [PubMed] [Google Scholar]
- 53.Ankamreddy H., Bok J., Groves A.K. Uncovering the Secreted Signals and Transcription Factors Regulating the Development of Mammalian Middle Ear Ossicles. Dev. Dyn. Off. Publ. Am. Assoc. Anat. 2020;249:1410–1424. doi: 10.1002/dvdy.260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Apschner A., Schulte-Merker S., Witten P.E. Chapter 10—Not All Bones Are Created Equal—Using Zebrafish and Other Teleost Species in Osteogenesis Research. In: Detrich H.W., Westerfield M., Zon L.I., editors. Methods in Cell Biology. Volume 105. Academic Press; London, UK: 2011. pp. 239–255. The Zebrafish: Disease Models and Chemical Screens. [DOI] [PubMed] [Google Scholar]
- 55.Chang Z., Chen P.-Y., Chuang Y.-J., Akhtar R. Zebrafish as a Model to Study Bone Maturation: Nanoscale Structural and Mechanical Characterization of Age-Related Changes in the Zebrafish Vertebral Column. J. Mech. Behav. Biomed. Mater. 2018;84:54–63. doi: 10.1016/j.jmbbm.2018.05.004. [DOI] [PubMed] [Google Scholar]
- 56.Du S.J., Frenkel V., Kindschi G., Zohar Y. Visualizing Normal and Defective Bone Development in Zebrafish Embryos Using the Fluorescent Chromophore Calcein. Dev. Biol. 2001;238:239–246. doi: 10.1006/dbio.2001.0390. [DOI] [PubMed] [Google Scholar]
- 57.Huang H., Lin H., Lan F., Wu Y., Yang Z., Zhang J. Application of Bone Transgenic Zebrafish in Anti-Osteoporosis Chemical Screening. Anim. Models Exp. Med. 2018;1:53–61. doi: 10.1002/ame2.12000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Kwon R.Y., Watson C.J., Karasik D. Using Zebrafish to Study Skeletal Genomics. Bone. 2019;126:37–50. doi: 10.1016/j.bone.2019.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kimmel C.B., Ballard W.W., Kimmel S.R., Ullmann B., Schilling T.F. Stages of Embryonic Development of the Zebrafish. Dev. Dyn. 1995;203:253–310. doi: 10.1002/aja.1002030302. [DOI] [PubMed] [Google Scholar]
- 60.Walker D.M., Kimmel C. A Two-Color Acid-Free Cartilage and Bone Stain for Zebrafish Larvae. Biotech. Histochem. 2007;82:23–28. doi: 10.1080/10520290701333558. [DOI] [PubMed] [Google Scholar]
- 61.Thisse C., Thisse B. High-Resolution in Situ Hybridization to Whole-Mount Zebrafish Embryos. Nat. Protoc. 2008;3:59–69. doi: 10.1038/nprot.2007.514. [DOI] [PubMed] [Google Scholar]
- 62.Peterson S.M., Freeman J.L. RNA Isolation from Embryonic Zebrafish and CDNA Synthesis for Gene Expression Analysis. J. Vis. Exp. JoVE. 2009;30:1470. doi: 10.3791/1470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Livak K.J., Schmittgen T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- 64.Rao X., Huang X., Zhou Z., Lin X. An Improvement of the 2ˆ (–Delta Delta CT) Method for Quantitative Real-Time Polymerase Chain Reaction Data Analysis. Biostat. Bioinforma. Biomath. 2013;3:71–85. [PMC free article] [PubMed] [Google Scholar]
- 65.Atan D., Atan T., Özcan K.M., Ensari S., Dere H. Relation of Otosclerosis and Osteoporosis: A Bone Mineral Density Study. Auris. Nasus. Larynx. 2016;43:400–403. doi: 10.1016/j.anl.2015.11.001. [DOI] [PubMed] [Google Scholar]
- 66.Markou K., Goudakos J. An Overview of the Etiology of Otosclerosis. Eur. Arch. Otorhinolaryngol. 2008;266:25. doi: 10.1007/s00405-008-0790-x. [DOI] [PubMed] [Google Scholar]
- 67.Niedermeyer H.P., Arnold W. Etiopathogenesis of Otosclerosis. ORL. 2002;64:114–119. doi: 10.1159/000057789. [DOI] [PubMed] [Google Scholar]
- 68.Schrauwen I., Khalfallah A., Ealy M., Fransen E., Claes C., Huber A., Murillo L.R., Masmoudi S., Smith R.J.H., Van Camp G. COL1A1 Association and Otosclerosis: A Meta-Analysis. Am. J. Med. Genet. A. 2012;158A:1066–1070. doi: 10.1002/ajmg.a.35276. [DOI] [PubMed] [Google Scholar]
- 69.Niedermeyer H.P., Becker E.T., Arnold W. Expression of Collagens in the Otosclerotic Bone. Adv. Otorhinolaryngol. 2007;65:45–49. doi: 10.1159/000098668. [DOI] [PubMed] [Google Scholar]
- 70.Green J., Taylor J.J., Hindes A., Johnson S.L., Goldsmith M.I. A Gain of Function Mutation Causing Skeletal Overgrowth in the Rapunzel Mutant. Dev. Biol. 2009;334:224–234. doi: 10.1016/j.ydbio.2009.07.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Kitazawa T., Fujisawa K., Narboux-Nême N., Arima Y., Kawamura Y., Inoue T., Wada Y., Kohro T., Aburatani H., Kodama T., et al. Distinct Effects of Hoxa2 Overexpression in Cranial Neural Crest Populations Reveal That the Mammalian Hyomandibular-Ceratohyal Boundary Maps within the Styloid Process. Dev. Biol. 2015;402:162–174. doi: 10.1016/j.ydbio.2015.04.007. [DOI] [PubMed] [Google Scholar]
- 72.Nakada C., Satoh S., Tabata Y., Arai K., Watanabe S. Transcriptional Repressor Foxl1 Regulates Central Nervous System Development by Suppressing Shh Expression in Zebra Fish. Mol. Cell. Biol. 2006;26:7246–7257. doi: 10.1128/MCB.00429-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.El-Brolosy M.A., Kontarakis Z., Rossi A., Kuenne C., Günther S., Fukuda N., Kikhi K., Boezio G.L.M., Takacs C.M., Lai S.-L., et al. Genetic Compensation Triggered by Mutant MRNA Degradation. Nature. 2019;568:193–197. doi: 10.1038/s41586-019-1064-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.El-Brolosy M.A., Stainier D.Y.R. Genetic Compensation: A Phenomenon in Search of Mechanisms. PLoS Genet. 2017;13:e1006780. doi: 10.1371/journal.pgen.1006780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Salanga C.M., Salanga M.C. Genotype to Phenotype: CRISPR Gene Editing Reveals Genetic Compensation as a Mechanism for Phenotypic Disjunction of Morphants and Mutants. Int. J. Mol. Sci. 2021;22:3472. doi: 10.3390/ijms22073472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Clayton A.E., Mikulec A.A., Mikulec K.H., Merchant S.N., McKenna M.J. Association between Osteoporosis and Otosclerosis in Women. J. Laryngol. Otol. 2004;118:617–621. doi: 10.1258/0022215041917790. [DOI] [PubMed] [Google Scholar]
- 77.Cuesta I., Zaret K.S., Santisteban P. The Forkhead Factor FoxE1 Binds to the Thyroperoxidase Promoter during Thyroid Cell Differentiation and Modifies Compacted Chromatin Structure. Mol. Cell. Biol. 2007;27:7302–7314. doi: 10.1128/MCB.00758-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Sekiya T., Muthurajan U.M., Luger K., Tulin A.V., Zaret K.S. Nucleosome-Binding Affinity as a Primary Determinant of the Nuclear Mobility of the Pioneer Transcription Factor FoxA. Genes Dev. 2009;23:804–809. doi: 10.1101/gad.1775509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Seifi M., Walter M.a. Axenfeld-Rieger Syndrome. Clin. Genet. 2018;93:1123–1130. doi: 10.1111/cge.13148. [DOI] [PubMed] [Google Scholar]
- 80.Delanghe J.R., Langlois M.R. Hemopexin: A Review of Biological Aspects and the Role in Laboratory Medicine. Clin. Chim. Acta. 2001;312:13–23. doi: 10.1016/S0009-8981(01)00586-1. [DOI] [PubMed] [Google Scholar]
- 81.Tolosano E., Altruda F. Hemopexin: Structure, Function, and Regulation. DNA Cell Biol. 2002;21:297–306. doi: 10.1089/104454902753759717. [DOI] [PubMed] [Google Scholar]
- 82.Kobayashi I., Kobayashi-Sun J., Hirakawa Y., Ouchi M., Yasuda K., Kamei H., Fukuhara S., Yamaguchi M. Dual Role of Jam3b in Early Hematopoietic and Vascular Development. Development. 2020;147:dev181040. doi: 10.1242/dev.181040. [DOI] [PubMed] [Google Scholar]
- 83.Cesari M., Pahor M., Lauretani F., Penninx B.W.H.J., Bartali B., Russo R., Cherubini A., Woodman R., Bandinelli S., Guralnik J.M., et al. Bone Density and Hemoglobin Levels in Older Persons: Results from the InCHIANTI Study. Osteoporos. Int. J. Establ. Result Coop. Eur. Found. Osteoporos. Natl. Osteoporos. Found. USA. 2005;16:691–699. doi: 10.1007/s00198-004-1739-6. [DOI] [PubMed] [Google Scholar]
- 84.Chuang M.-H., Chuang T.-L., Koo M., Wang Y.-F. Low Hemoglobin Is Associated with Low Bone Mineral Density and High Risk of Bone Fracture in Male Adults: A Retrospective Medical Record Review Study. Am. J. Mens Health. 2019;13:1557988319850378. doi: 10.1177/1557988319850378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Steer K., Stavnichuk M., Morris M., Komarova S.V. Bone Health in Patients with Hematopoietic Disorders of Bone Marrow Origin: Systematic Review and Meta-Analysis. J. Bone Miner. Res. 2017;32:731–742. doi: 10.1002/jbmr.3026. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All available raw data are available from the Principal Investigator, Cutis R. French.