Highlights
-
•
SEM-based method can be applied for rapid phenotypic AST on Gram-positive cocci towards 2 vancomycin based on morphological changes 3.
-
•
The ratio of septa as a marker of bacterial division and size of grape-like clusters enabled the 4 profiling of E. faecalis, E. faecium and S. aureus after brief incubation with vancomycin 5.
-
•
SEM-AST strategy showed the feasibility of detecting antibiotic susceptibility or 6 resistance on Gram-positive cocci within one hour of exposition to vancomycin 7.
Keywords: Gram-positive cocci, Staphylococcus, Enterococcus, Vancomycin, Microbiology, Rapid AST, Scanning electron microscopy
Abstract
The rapid detection of resistant bacteria has become a challenge for microbiologists worldwide. Numerous pathogens that cause nosocomial infections are still being treated empirically and have developed resistance mechanisms against key antibiotics. Thus, one of the challenges for researchers has been to develop rapid antimicrobial susceptibility testing (AST) to detect resistant isolates, ensuring better antibiotic stewardship. In this study, we established a proof-of-concept for a new strategy of phenotypic AST on Gram-positive cocci towards vancomycin using scanning electron microscopy (SEM). Our study evaluated the profiling of Enterococcus faecalis, Enterococcus faecium and Staphylococcus aureus after brief incubation with vancomycin. Sixteen isolates were analysed aiming to detect ultrastructural modifications at set timepoints, comparing bacteria with and without vancomycin. After optimising slides preparation and micrographs acquisition, two analytical strategies were used. The high magnification micrographs served to analyse the division of cocci based on the ratio of septa, along with the bacterial size. Susceptible strains with vancomycin showed a reduced septa percentage and the average surface area was consequently double that of the controls. The resistant bacteria revealed multiple septa occurring at advanced timepoints. Low magnification micrographs made it possible to quantify the pixels at different timepoints, confirming the profiling of cocci towards vancomycin. This new phenotypic AST strategy proved to be a promising tool to discriminate between resistant and susceptible cocci within an hour of contact with vancomycin. The analysis strategies applied here would potentially allow the creation of artificial intelligence algorithms for septa detection and bacterial quantification, subsequently creating a rapid automated SEM-AST assay.
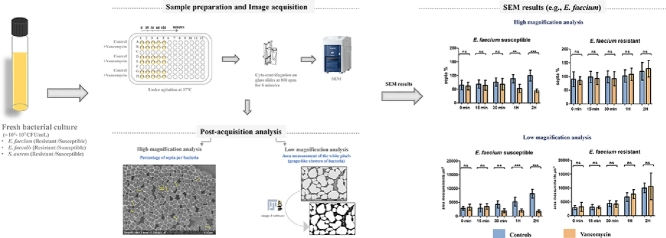
Graphical abstract
1. Introduction
The early detection of bacterial pathogens and their antimicrobial susceptibility is urgently needed to treat infections with appropriate and faster targeted therapy (Khatib et al., 2006; Lodise et al., 2003; Wolk and Johnson, 2019). One of the major worldwide causes of nosocomial and community-acquired bacteriemia is Gram-positive bacteria, particularly Gram-positive cocci. Within these are vancomycin-resistant Enterococcus (VRE) and Staphylococcus aureus, especially methicillin-resistant Staphylococcus aureus (MRSA) (Asokan et al., 2019; Cornaglia, 2009; Dweba et al., 2018; Tong et al., 2015). Patients with infectious diseases are currently still treated empirically, while awaiting a diagnosis and the subsequent early targeted administration of antibiotics (Wolk and Johnson, 2019). Therefore, antimicrobial susceptibility testing (AST) diagnostics have a direct impact on patient management and accurate antibiotic stewardship (Khatib et al., 2006; Lodise et al., 2003; Nathwani et al., 2019; Wolk and Johnson, 2019). Conventional AST methods such as broth micro-dilution or agar disk diffusion remain the gold standard, but lack speed and rapid results (Khan et al., 2019; Wilson et al., 2015). Recent advances in AST systems in recent decades, mainly molecular and genotypic methods (Datar et al., 2021; Emonet et al., 2016; Fluit et al., 2001; Sundsfjord et al., 2004; van Belkum et al., 2020), seek to accelerate the time to provide AST results, yet, they are often costly and only available in well-equipped labs. In contrast, most phenotypic systems are usually performed on isolated and pre-identified sub-cultures (Barman et al., 2018; Kaprou et al., 2021a), while others consist of fully automated systems capable of performing identification followed by AST (Lutgring et al., 2018; Humphries and Di Martino, 2019; Chapot et al., 2021). However, the high costs of these platforms are a major barrier to their widespread use for small-budget institutions.
Rapid and affordable AST has a crucial role to play in clinical microbiology and the appropriate choice of treatment. For this reason, in this study, we evaluated a new preliminary strategy for the direct AST of Gram-positive cocci towards vancomycin using SEM, with a view to accelerating the time it takes to reach the phenotypic AST results. Knowing that vancomycin has a direct impact on bacterial morphology through its ability to inhibit bacterial cell wall biosynthesis, thus affecting bacterial cell division (Lorian and Fernandes, 1997; Pinho and Errington, 2003), we performed a detailed analysis on the most common Gram-positive pathogens, based on the morphological changes that occurred after contact with vancomycin.
2. Materials and methods
2.1. Bacterial selection and profiling
In this study, we analysed 16 Gram-positive cocci strains: Staphylococcus aureus (four isolates), Enterococcus faecalis (six isolates) and Enterococcus faecium (six isolates) with different resistance profiles towards vancomycin (Table 1). Due to the lack of clinical vancomycin-resistant- S. aureus isolates in our country, we tested only susceptible strains. All bacterial isolates were obtained from the “Collection de Souches de l'Unité des Rickettsies” (CSUR, WDCM 875). The bacterial identification and antibiotic susceptibility profiling for each isolate was carried out using MALDI TOF MS (matrix-assisted laser desorption/ionization time-of-flight mass spectrometry; Microflex, Bruker Daltonics, Germany) (Seng et al., 2009), followed by antimicrobial susceptibility tests (AST) using the E-test technique (bioMérieux, Marcy l'Etoile, France) and broth micro-dilution assays (Baker et al., 1991). Briefly, each of the bacterial strains were cultured on Mueller-Hinton broth liquid media (MHB) (Sigma-Aldrich, St. Louis, MO, USA) overnight at 37 °C under aerobic atmosphere, then the bacterial suspensions were adjusted to 0.5 McFarland standard. For E-test method, the bacterial suspensions were inoculated onto Muller-Hinton agar plates (Becton Dickinson, Sparks, MD), then vancomycin strips (antibiotic gradient from 0.016 to 256 μ g/ml) were applied in the plate and incubated at 37 °C for 18 to 24 h. After the incubation, minimum inhibitory concentration (MIC) of each strain was read directly from the graduated E-test strip at the point of intersection of inhibition ellipse and the test strip. Isolates were also evaluated by the broth micro-dilution assay, where, the same bacterial suspensions were loaded on 96-well microplate and 20 incremental dilutions of vancomycin from 0.25 to 256 µg/mL (0.25, 0.50, 0.75, 1, 1.5, 2, 3, 4, 6, 8, 12, 16, 24, 32, 48, 64, 96, 128, 192 and 256 µg/mL) were tested. In addition, positive (MHB with bacteria) and negative (serial dilutions of vancomycin with MHB without bacteria) controls were prepared. The plates were incubated at 37 °C for 18 h and the MIC was read manually after incubation by adding iodonitrotetrazolium chloride colorimetric assay (Sigma-Aldricht, Illkirch, France). Results of vancomycin were interpreted according to the Clinical and Laboratory Standards Institute (CLSI) and European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines.
Table 1.
List of isolates analysed.
| Strain | Resistance profile (Vancomycin) | MIC breakpoint (mg/L) | E-test (mg/L) | |
|---|---|---|---|---|
| Staphylococcus aureus | Q3211 | susceptible | 2 mg/L | 1 |
| Staphylococcus aureus | Q3208 | susceptible | 2 mg/L | 1 |
| Staphylococcus aureus | Q5877 | susceptible | 2 mg/L | 1 |
| Staphylococcus aureus | Q5878 | susceptible | 2 mg/L | 1 |
| Enterococcus faecium | Q3193 | resistant | 4 mg/L | >256 |
| Enterococcus faecium | Q3195 | resistant | 4 mg/L | >256 |
| Enterococcus faecium | Q3194 | resistant | 4 mg/L | 32 |
| Enterococcus faecium | Q3189 | susceptible | 4 mg/L | 0.75 |
| Enterococcus faecium | Q3191 | susceptible | 4 mg/L | 0.75 |
| Enterococcus faecium | Q3188 | susceptible | 4 mg/L | 0.5 |
| Enterococcus faecalis | Q3206 | resistant | 4 mg/L | >256 |
| Enterococcus faecalis | Q3205 | resistant | 4 mg/L | 8 |
| Enterococcus faecalis | Q3203 | resistant | 4 mg/L | >256 |
| Enterococcus faecalis | Q3202 | susceptible | 4 mg/L | 2 |
| Enterococcus faecalis | Q3201 | susceptible | 4 mg/L | 0.75 |
| Enterococcus faecalis | Q3200 | susceptible | 4 mg/L | 0.75 |
2.2. Sample preparation for antimicrobial susceptibly testing (AST) for vancomycin
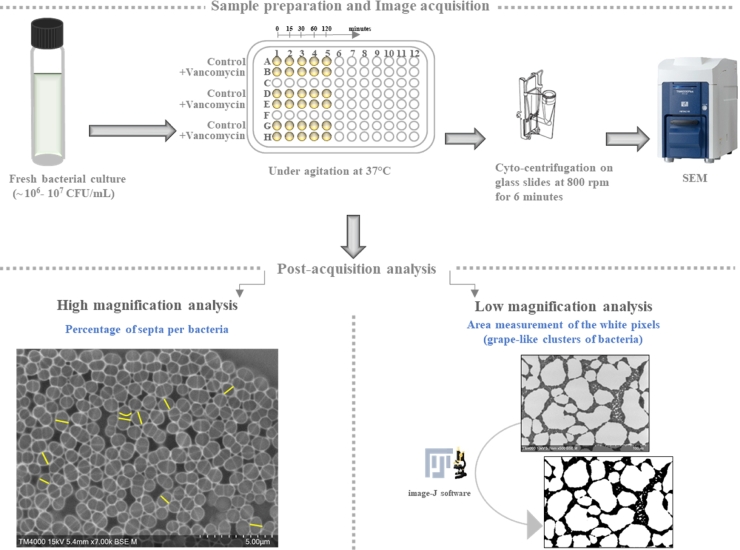
The samples were prepared as described in our recent publication, for AST of Gram-negative bacteria (Haddad et al., 2021b). A patent application related to this strategy and methodology was also deposited (WO2020/183,072 A1). Bacterial species were grown overnight on Mueller-Hinton broth liquid media (MHB) (Sigma-Aldrich, St. Louis, MO, USA) under aerobic conditions at 37 °C for 16–18 h. The bacterial concentration was adjusted using MHB to reach approximately ∼106 −107 CFU/mL, then distributed in 96-well microplates for AST (Fig. 1). Vancomycin was added at the standard MIC recovered from EUCAST for each of the tested species. Bacterial suspensions were continually agitated and were analysed with and without antibiotics at different timepoints (0, 15, 30, 60 and 120 min). Cell staining was performed by directly adding phosphotungstic acid (PTA) (Sigma-Aldrich, St. Louis, MO, USA) at 10% in all wells (with and without the antibiotic), followed by five minutes of agitation. Stained suspensions were then cyto-centrifuged on glass slides at 800 rpm for six minutes, before being processed by scanning electron microscopy (SEM) (Fig. 1).
Fig. 1.
Detailed strategy applied to a selection of Gram-positive cocci incubated with and without vancomycin, then imaged at selected timepoints (0, 15, 30, 60, and 120 min) using TM4000 Plus tabletop SEM, followed by two analyses strategies. Yellow lines represent the septa.
2.3. AST using SEM
2.3.1. Imaging process
Micrographs were acquired on the TM4000 Plus tabletop SEM (Hitachi, Tokyo, Japan) at each timepoint using Zigzag software that allows for a rapid and automated acquisition. Low magnification micrographs were recorded at 10 kV accelerating voltage with a 500X magnification, whereas higher magnifications (5000X and 7000X) were recorded with an accelerating voltage of 15 kV, as previously described (Haddad et al., 2021b).
2.3.2. Post-acquisition analysis and cell measurements
Images were analysed using the image-J software (Wayne Rasband, National Institutes of Health, USA, version 1.52i, Java 1.8.0_172 (64-bit)) and all analyses were performed in a double-blind assay for greater accuracy. At least three hundred bacteria and their septa were counted and measured on between two and five randomly selected micrographs in each condition, as described in Fig. 1. We aimed to find morphological modifications at early timepoints, then extended our investigations for up to two hours when no changes were detected. Image analysis consisted of detecting deep modifications at the morphological and ultrastructural levels on high magnification micrographs. We targeted the septum formation as a sign of bacterial division, as well as bacterial engulfment, cell wall modification and amorphous shapes. The cell surface area was also calculated using the following equation: S = π td, where d is half of the equatorial length (µm) and t half of the axial length (µm).
As for the low magnification micrographs used to assess the overall distribution of the bacteria, we measured the total area in µm2 of grape-like clusters visible on the micrographs. We adjusted the threshold and removed the background noise on Image-J. Bacterial pixels are represented in black and the background pixels in white. This analysis enabled us to make an overall assessment of the bacterial distribution.
2.4. Statistical analysis
Data were analysed by one-way-ANOVA followed by Turkey's test using GraphPad Prism software 5.03 (GraphPad, San Diego, CA). The septa formation was percentage-transformed and expressed as mean ± standard error of septa formation per bacterial phenotypic group (total count of 300 bacteria per isolate, per condition, in triplicate). Significant differences between bacteria incubated with the antibiotic and controls were statistically significant when the p-value <0.05. All experiments were conducted in triplicate.
3. Results
To study the morphological changes in cocci occurring after contact with vancomycin, we used PTA staining, which enabled us to better visualise various morphological modifications in the bacteria, notably the septa formation at high magnifications. All analyses presented below were based on the percentage of septa along with the bacterial surface areas and the grape-like cluster size (Fig. 1).
3.1. Enterococcus faecalis
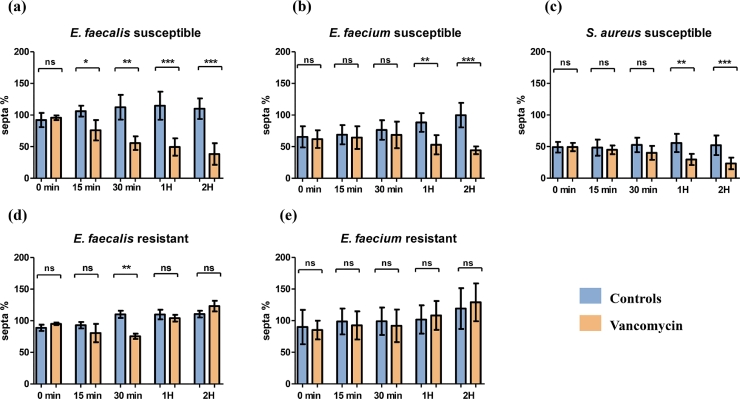
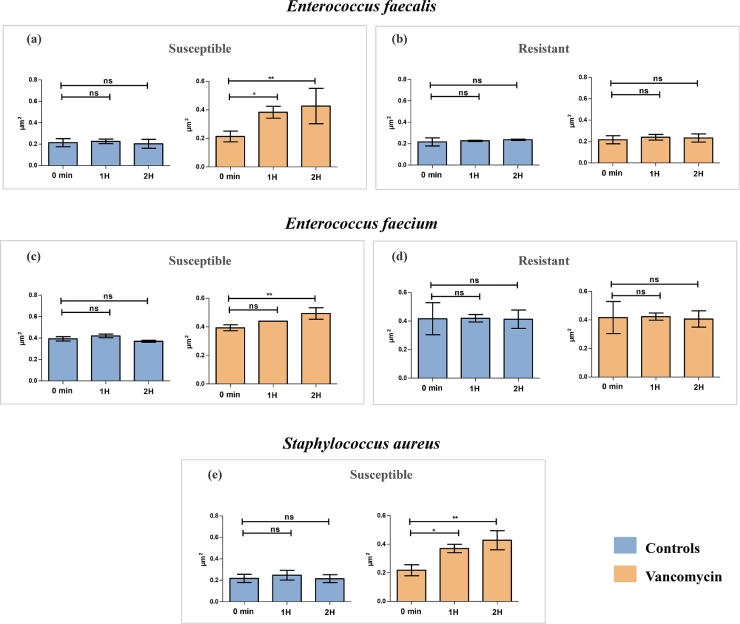
Susceptible E. faecalis strains incubated without any antibiotics showed a subtle increase in the septa percentage starting from 92±11% to 110±16% (p>0.05) after two hours of incubation (Fig. 3a). SEM micrographs showed rugby-ball-shaped and hourglass-shaped bacteria with an apparent septum (Fig. 2a-c). The mean surface area of all the susceptible strains remained the same throughout the two hours of incubation (0.21± 0.04 µm2 and 0.20±0.04 µm2 at 0 and 120 min, respectively) (Fig. 4a). Interestingly, in the presence of vancomycin, the septum markedly decreased until it disappeared completely, and rugby-ball-shaped bacteria with incomplete septa or without any visible septa in most cases subsequently emerged. The septa percentage significantly decreased by 30% starting 15 min (p<0.01), and by 72% after two hours (p < 0.0001) (Fig. 3a). Thus, the mean surface areas of the susceptible strains significantly increased, by 1.8 to 2 times, after one hour (0.38± 0.04 µm2, p < 0.01) and two hours (0.43± 0.12 µm2, p < 0.001) of incubation with vancomycin, respectively (Fig. 4a). The septa in vancomycin-resistant isolates were intensely stained and visible on the micrographs (Fig. 2d-f). After 30 min of contact with vancomycin, a statistically significant decrease of 35% in the septa proportion was observed (p < 0.001) (Fig. 3d). However, from the 60-minute timepoint onwards, the percentage increased again in all three resistant isolates (104±16%), with no significant difference compared to the controls (110±23%) (Fig. 3d, S1a-c). In contrast to susceptible bacteria, the mean surface area of all the resistant strains remained constant (∼0.23 µm2) over time, despite the presence of vancomycin (Fig. 4b).
Fig. 3.
Graphs representing the mean ±SD of total septa percentage for each phenotypic group (Susceptible and Resistant) conducted in triplicate.
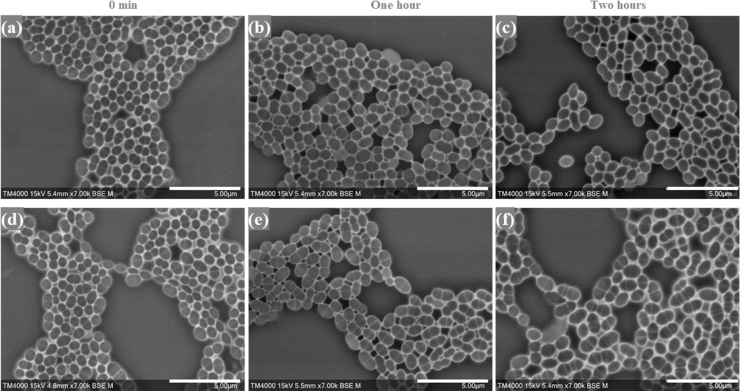
Fig. 2.
Micrographs of E. faecalis incubated with vancomycin at the starting time and at the one- and two-hour timepoints. a, b, c: high magnification micrographs of the vancomycin-susceptible E. faecalis isolate (Q3202). d, e, f: high magnification micrographs of the vancomycin-resistant E. faecalis (Q3205). Micrographs were recorded using TM4000 Plus SEM at a 7,000x magnification. Imaging settings are visible on the micrographs. Scale bars: 5 µm.
Fig. 4.
Graphs representing the mean ±SD of the surface area (µm2) for each phenotypic group (Susceptible and Resistant) conducted in triplicate.
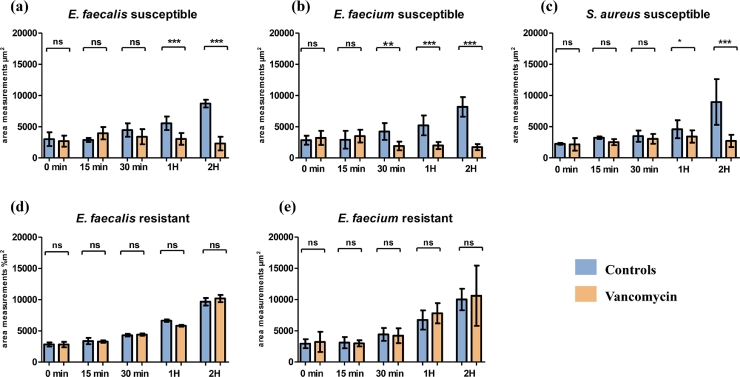
The low magnification micrograph analysis made it possible to quantify bacteria and monitor their proliferation over time, showing matching results when measuring the cluster areas. Susceptible bacteria presented a significative decrease in bacterial density from 60 min onwards with vancomycin, compared to the controls (p < 0.0001) (Figs. 5, 6a). Surface area measurements at 0, 15 and 30 min showed no significant alterations between the controls and vancomycin incubations, although a slight increase was observed in both. Nevertheless, an exponential increase in bacterial density was observed in the resistant isolates, in both the control and vancomycin incubations with no apparent decrease in grape-like clusters at 30 min, as perceived using the high magnification analysis (Fig. 5, 6d, S2a-c).
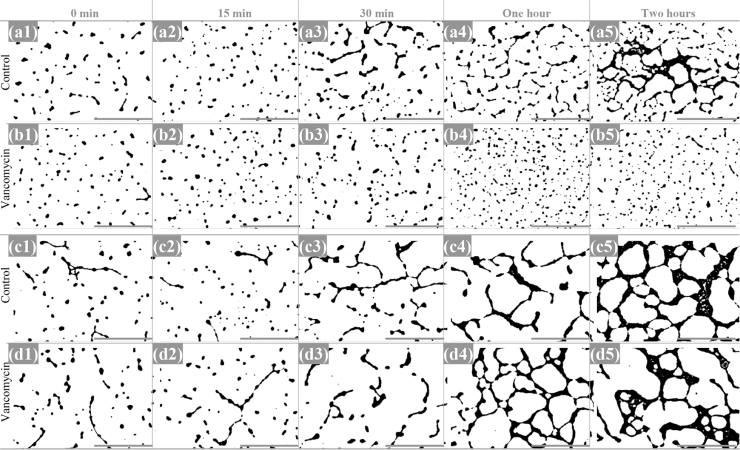
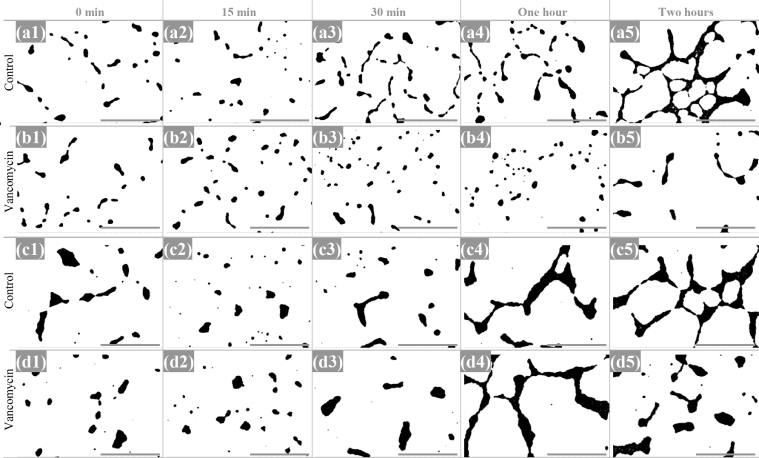
Fig. 5.
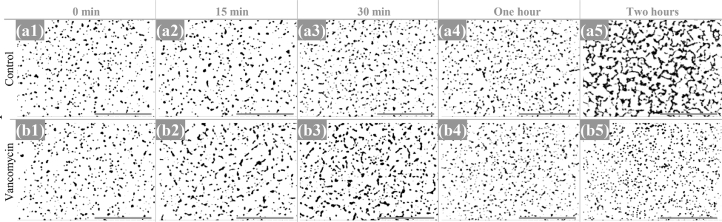
Low magnification micrographs of E. faecalis at the studied timepoints, after threshold adjustment on Image-J. a, b: Vancomycin-susceptible E. faecalis isolate (Q3202) incubated without (a) and with vancomycin (b). c, d: vancomycin-resistant E. faecalis isolate (Q3205) incubated without (c) and with vancomycin (d). Micrographs were recorded at a 500x magnification. Scale bars: 100 µm.
Fig. 6.
Graphs representing the mean ±SD of total cluster area measurement for each phenotypic group (Susceptible and Resistant) conducted in triplicate.
3.2. Enterococcus faecium
In the absence of vancomycin, susceptible isolates showed a slight increase in the septa percentage at the 30-minute timepoint, then increased by 23% after one hour of incubation (88%±15, p > 0.01) to reach a significative septa amplification of 34% (100±19%, p < 0.001) after two hours (Fig. 3b). The bacterial cells appeared mostly as rugby-ball-shaped or hour-glass-shaped, similar to E. faecalis strains but more elongated with complete septa (Fig. 7) and an unchanged surface area (∼0.39 µm2) throughout the course of the two hours of incubation (Fig. 4c). However, when in contact with vancomycin, the susceptible strains had a significant decrease of 35% (53±15%, p<0.001), starting at the 60-minute timepoint and reaching 56% (44±6%, p < 0.0001) after two hours of incubation (Fig. 3b, 7a-c). The morphological effects observed with the vancomycin-susceptible strains were very similar to those found in the E. faecalis susceptible strains, where the majority of bacteria were ovoid-shaped with no visible septum (Fig. 7a-c). In addition, the mean surface area progressively increased by 0.10 µm2 (0.49±0.04, p < 0.001) after two hours of contact with vancomycin (Fig. 4c). For the resistant isolates of E. faecium, a slight, non-significant increase in septa percentage over time was observed, regardless of the presence of vancomycin (Fig. 3e, S1a-c). No clear morphological changes occurred regardless of whether or not the strains were in contact with vancomycin. Under both conditions, the resistant strains were ovoid-shaped with visible septa (Figure D-f) and a surface area which was constant (∼0.42 µm2) over time (Fig. 4d).
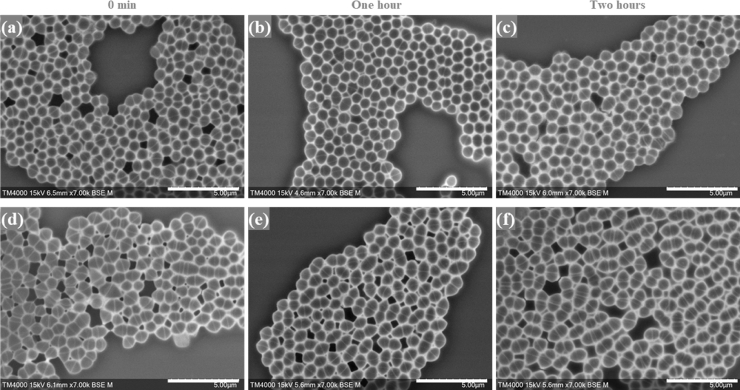
Fig. 7.
Micrographs of E. faecium incubated with vancomycin at the starting time and at the one- and two-hour timepoints. a, b, c: High magnification micrographs of the vancomycin-susceptible isolate (Q3188). d, e, f: High magnification micrographs of the vancomycin-resistant E. faecium (Q3193). Micrographs were recorded using TM4000 Plus SEM at a 7,000x magnification. Imaging settings are visible on the micrographs. Scale bars: 5 µm.
When analysing the low magnification micrographs, the susceptible isolates showed a significant decrease starting at 30 min (p < 0.001) in the bacteria-covered portion of the micrographs when incubated with vancomycin (Fig. 6b, 8b, S2d-f). This decrease was more noticeable at later timepoints (p < 0.0001). In contrast, the resistant E. faecium cluster areas increased exponentially, regardless of the presence of antibiotics (Figs. 6e, 8c, 8d).
Fig. 8.
Low magnification micrographs of E. faecium at the studied timepoints, after threshold adjustment on Image-J. a, b: vancomycin-susceptible E. faecium isolate (Q3188) incubated without (a) and with vancomycin (b). c, d: vancomycin-resistant E. faecium isolate (Q3193) incubated without (c) and with vancomycin (d). Micrographs were recorded at a 500x magnification. Scale bars: 100 µm.
3.3. Staphylococcus aureus
The susceptible S. aureus isolates showed minor, non-significant increases in septa percentage throughout the whole vancomycin-free incubation period. The mean surface area remained unchanged (∼0.23 µm2) and the majority of susceptible bacteria were d-shaped and roughly spherical with a clear septum. However, after one hour of incubation with vancomycin, the septa of susceptible strains (30±9%) showed a significant decrease of 26% (p < 0.0001) compared to the controls (56±14%) and remained low after two hours of incubation with vancomycin (23±9%, p < 0.0001) (Figs. 3c, 9b). In addition, the bacterial cells appeared ellipsoidal in shape, sometimes spherical, with incomplete or missing septa. Their mean surface areas also significantly increased 1.7 to 2-fold after one hour (0.38± 0.03 µm2, p < 0.01) and two hours (0.43± 0.8 µm2, p < 0.001) incubation with vancomycin, respectively (Figs. 3c, 9b). The area measurements of grape-like clusters significantly decreased one- to two-fold after one and two hours of incubation with vancomycin (p < 0.01 and p < 0.0001, respectively), as opposed to the control which continued to proliferate (Figs. 6c, 10, S2m-p).
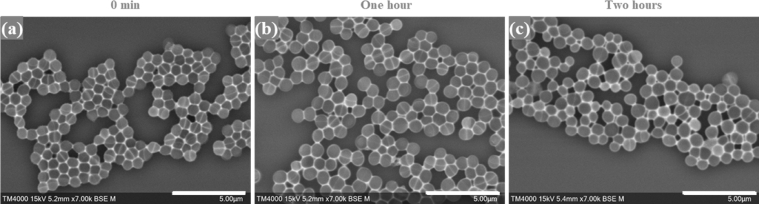
Fig. 9.
Micrographs of S. aureus incubated with vancomycin at the starting timepoint and at the one- and two-hour timepoints. a, b, c: High magnification micrographs of the vancomycin-susceptible S. aureus isolate (Q5878). Micrographs were recorded using TM4000 Plus SEM at a 7,000x magnification. Imaging settings are visible on the micrographs. Scale bars: 5 µm.
Fig. 10.
Low magnification micrographs of S. aureus at the studied timepoints, after threshold adjustment on Image-J. a, b: Vancomycin-susceptible S. aureus isolate (Q5878) incubated without (a) and with vancomycin (b). Micrographs were recorded at a 500x magnification. Scale bars: 100 µm.
4. Discussion
The lack of rapid and accurate AST calls for the development of innovative strategies and systems aiming to improve patient outcomes by ensuring appropriate antibiotic therapy (Perez et al., 2014; Vickers et al., 2019). In this study, we assessed the morphological modification patterns of three frequently occurring clinical pathogens, S. aureus, E. faecalis, and E. faecium, under exposure to vancomycin using a SEM assay.
Analysis of the micrographs enabled us to assign distinct morphological profiles to the resistant and susceptible strains, mainly the formation of septa and the size of grape-like clusters yielding correlating results. The susceptible strains affected by vancomycin exhibited a missing or incomplete septum at high magnifications, and hence the average surface area was almost twice that of the controls. On the contrary the vancomycin-resistant cocci appeared normal with a constant area size regardless the presence of vancomycin at high magnifications. Therefore, the septum was apparent and intensely stained in both conditions. For both Enterococcus sp., bacteria with multiple septa occurred on the micrographs at advanced time points.
The loss of visible septa could result from vancomycin inhibiting cell-wall-synthesis (Lorian and Fernandes, 1997; Pereira et al., 2007; Pinho and Errington, 2003), and thereby inhibiting further cell division, as confirmed herein by the low magnification analysis. Similar effects on the septum have been described for MRSA and VRE on ultra-thin sections (Belley et al., 2009).
Although there have been vast improvements in rapid AST automation, the turn-around time for AST results of many culture-based ASTs remains high (Doern, 2018; van Belkum et al., 2019; World Health Organization, 2019; Avesar et al., 2017; Maugeri et al., 2019). Some of these systems, which can provide identification and AST results, are costly and yield varied results in terms of time or accuracy (Kaprou et al., 2021b; Wolk and Johnson, 2019). While conventional AST methods remain the gold standard, our AST-SEM strategy for testing cocci towards vancomycin provides results within one hour. Together with the straightforward way in which samples are prepared, the AST-SEM strategy has demonstrated its potential to be a rapid and low-cost sample preparation.
It is important to point out that these findings are preliminary. One major limitation of our strategy is that AST was performed on pure cultures, and should eventually be applied directly on blood cultures. Additional investigations, testing various pathogenic strains against a large panel of antibiotics, are essential for clinical applications. The advantages of the tabletop SEM used in this study relies in its ease of use and user-friendly interface, the rapid acquisition of micrographs, and being an affordable tool (€ 50k) that can be installed in clinical labs. A similar strategy for clinically relevant Gram-negative bacteria against imipenem was investigated by our team, showing morphological changes after one hour of incubation (Haddad et al., 2021b). Previously, we also developed an SEM strategy for bacterial identification from blood cultures (Haddad et al., 2021a). Merging these studies could make it possible to develop an automated system coupled with artificial intelligence for rapid identification and AST directly from blood cultures, which would mark a milestone in bacteriological diagnosis.
CRediT authorship contribution statement
Sara Bellali: Writing – original draft, Writing – review & editing, Formal analysis, Visualization. Gabriel Haddad: Writing – original draft, Writing – review & editing, Formal analysis, Visualization. Rim Iwaza: Formal analysis. Anthony Fontanini: Investigation, Formal analysis. Akiko Hisada: Methodology. Yusuke Ominami: Methodology. Didier Raoult: Conceptualization, Supervision. Jacques Bou Khalil: Validation, Writing – review & editing, Supervision.
Declaration of Competing Interest
Authors would like to declare that D.R. was a consultant in microbiology for the Hitachi High-Tech Corporation from March 2018 until March 2021. Y.O. is employed by the company Hitachi High-Tech Corporation. AH is employed by the company Hitachi, Ltd. Personal fees of G.H., S.B., A.F. and J.B. are paid through a collaborative contract from the company Hitachi High-Tech Corporation. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Funding
This work was supported by a grant from the French Government managed by the National Research Agency under the “Investissements d'avenir (Investments for the Future)” programme with the reference ANR-10-IAHU-03 (Méditerranée Infection), by the Région Provence-Alpes-Côte-d'Azur and the European funding FEDER PRIMI. In addition, collaborative study conducted by IHU Méditerranée Infection and the Hitachi High-Tech Corporation is funded by the Hitachi High-Tech Corporation.
Acknowledgments
We sincerely thank Takashi Irie, Kyoko Imai, Shigeki Matsubara, Taku Sakazume, Toshihide Agemura and the Hitachi team in Japan for the collaborative study conducted together with IHU Méditerranée Infection, and the installation of TM4000 Plus microscopes at the IHU Méditerranée Infection facility.
Footnotes
Supplementary material associated with this article can be found, in the online version, at doi:10.1016/j.crmicr.2022.100154.
Contributor Information
Didier Raoult, Email: didier.raoult@gmail.com.
Jacques Bou Khalil, Email: boukhaliljacques@gmail.com.
Appendix. Supplementary materials
References
- Asokan G., Ramadhan T., Ahmed E., Sanad H. WHO global priority pathogens list: a bibliometric analysis of medline-pubmed for knowledge mobilization to infection prevention and control practices in Bahrain. OMJ. 2019;34(3):184–193. doi: 10.5001/omj.2019.37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Avesar J., Rosenfeld D., Truman-Rosentsvit M., Ben-Arye T., Geffen Y., Bercovici M., Levenberg S. Rapid phenotypic antimicrobial susceptibility testing using nanoliter arrays. Proc. Natl. Acad. Sci. USA. 2017;114:E5787–E5795. doi: 10.1073/pnas.1703736114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker C.N., Stocker S.A., Culver D.H., Thornsberry C. Comparison of the E Test to agar dilution, broth microdilution, and agar diffusion susceptibility testing techniques by using a special challenge set of bacteria. J. Clin. Microbiol. 1991;29:533–538. doi: 10.1128/jcm.29.3.533-538.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barman P., Chopra S., Thukral T. Direct testing by VITEK® 2: a dependable method to reduce turnaround time in Gram-negative bloodstream infections. J. Lab. Physicians. 2018;10:260–264. doi: 10.4103/JLP.JLP_11_18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belley A., Harris R., Beveridge T., Parr T., Moeck G. Ultrastructural Effects of Oritavancin on Methicillin-Resistant Staphylococcus aureus and Vancomycin-Resistant Enterococcus. Antimicrob. Agents Chemother. 2009;53:800–804. doi: 10.1128/AAC.00603-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chapot V., Effenberg L., Dohmen-Ruetten J., Buer J., Kehrmann J. Evaluation of the accelerate pheno system for rapid identification and antimicrobial susceptibility testing of positive blood culture bottles inoculated with primary sterile specimens from patients with suspected severe infections. J. Clin. Microbiol. 2021;59 doi: 10.1128/JCM.02637-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cornaglia G. Fighting infections due to multidrug-resistant Gram-positive pathogens. Clin. Microbiol. Infect. 2009;15:209–211. doi: 10.1111/j.1469-0691.2009.02737.x. [DOI] [PubMed] [Google Scholar]
- Datar R., Orenga S., Pogorelcnik R., Rochas O., Simner P.J., van Belkum A. Recent advances in rapid antimicrobial susceptibility testing. Clin. Chem. 2021;68:91–98. doi: 10.1093/clinchem/hvab207. [DOI] [PubMed] [Google Scholar]
- Doern C.D. The slow march toward rapid phenotypic antimicrobial susceptibility testing: are we there yet? J. Clin. Microbiol. 2018;56 doi: 10.1128/JCM.01999-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dweba C.C., Zishiri O., El Zowalaty M. Methicillin-resistant Staphylococcus aureus: livestock-associated, antimicrobial, and heavy metal resistance. IDR Volume. 2018;11:2497–2509. doi: 10.2147/IDR.S175967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emonet S., Charles P.G., Harbarth S., Stewardson A.J., Renzi G., Uckay I., Cherkaoui A., Rougemont M., Schrenzel J. Rapid molecular determination of methicillin resistance in staphylococcal bacteraemia improves early targeted antibiotic prescribing: a randomized clinical trial. Clin. Microbiol. Infect. 2016;22 doi: 10.1016/j.cmi.2016.07.022. 946.e9-946.e15. [DOI] [PubMed] [Google Scholar]
- Fluit A.C., Visser M.R., Schmitz F.-.J. Molecular detection of antimicrobial resistance. Clin. Microbiol. Rev. 2001;14:836–871. doi: 10.1128/CMR.14.4.836-871.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haddad G., Bellali S., Takakura T., Fontanini A., Ominami Y., Bou Khalil J., Raoult D. Scanning Electron Microscope: a New Potential Tool to Replace Gram Staining for Microbe Identification in Blood Cultures. Microorganisms. 2021;9:1170. doi: 10.3390/microorganisms9061170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haddad G., Fontanini A., Bellali S., Takakura T., Ominami Y., Hisada A., Hadjadj L., Rolain J.-.M., Raoult D., Bou Khalil J.Y. Rapid detection of imipenem resistance in gram-negative bacteria using tabletop scanning electron microscopy: a preliminary evaluation. Front. Microbiol. 2021;12 doi: 10.3389/fmicb.2021.658322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Humphries R., Di Martino T. Effective implementation of the Accelerate Pheno™ system for positive blood cultures. J. Antimicrob. Chemother. 2019;74 doi: 10.1093/jac/dky534. i40–i43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaprou G.D., Bergšpica I., Alexa E.A., Alvarez-Ordóñez A., Prieto M. Rapid methods for antimicrobial resistance diagnostics. Antibiotics. 2021;10:209. doi: 10.3390/antibiotics10020209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaprou G.D., Bergšpica I., Alexa E.A., Alvarez-Ordóñez A., Prieto M. Rapid methods for antimicrobial resistance diagnostics. Antibiotics. 2021;10:209. doi: 10.3390/antibiotics10020209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khan Z.A., Siddiqui M.F., Park S. Current and emerging methods of antibiotic susceptibility testing. Diagnostics (Basel) 2019;9:49. doi: 10.3390/diagnostics9020049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khatib R., Saeed S., Sharma M., Riederer K., Fakih M.G., Johnson L.B. Impact of initial antibiotic choice and delayed appropriate treatment on the outcome of Staphylococcus aureus bacteremia. Eur. J. Clin. Microbiol. Infect. Dis. 2006;25:181–185. doi: 10.1007/s10096-006-0096-0. [DOI] [PubMed] [Google Scholar]
- Lodise T.P., McKinnon P.S., Swiderski L., Rybak M.J. Outcomes analysis of delayed antibiotic treatment for hospital-acquired Staphylococcus aureus bacteremia. Clin. Infect. Dis. 2003;36:1418–1423. doi: 10.1086/375057. [DOI] [PubMed] [Google Scholar]
- Lorian V., Fernandes F. The effect of vancomycin on the structure of vancomycin-susceptible and -resistant Enterococcus faecium strains. Antimicrob. Agents Chemother. 1997;41:1410–1411. doi: 10.1128/aac.41.6.1410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lutgring J.D., Bittencourt C., McElvania TeKippe E., Cavuoti D., Hollaway R., Burd E.M. Evaluation of the accelerate pheno system: results from two academic medical centers. J. Clin. Microbiol. 2018;56 doi: 10.1128/JCM.01672-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maugeri G., Lychko I., Sobral R., Roque A.C.A. Identification and antibiotic-susceptibility profiling of infectious bacterial agents: a review of current and future trends. Biotechnol. J. 2019;14 doi: 10.1002/biot.201700750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nathwani D., Varghese D., Stephens J., Ansari W., Martin S., Charbonneau C. Value of hospital antimicrobial stewardship programs [ASPs]: a systematic review. Antimicrob. Resist. Infect. Control. 2019;8:35. doi: 10.1186/s13756-019-0471-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pereira P.M., Filipe S.R., Tomasz A., Pinho M.G. Fluorescence Ratio imaging microscopy shows decreased access of vancomycin to cell wall synthetic sites in vancomycin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 2007;51:3627–3633. doi: 10.1128/AAC.00431-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perez K.K., Olsen R.J., Musick W.L., Cernoch P.L., Davis J.R., Peterson L.E., Musser J.M. Integrating rapid diagnostics and antimicrobial stewardship improves outcomes in patients with antibiotic-resistant Gram-negative bacteremia. J. Infect. 2014;69:216–225. doi: 10.1016/j.jinf.2014.05.005. [DOI] [PubMed] [Google Scholar]
- Pinho M.G., Errington J. Dispersed mode of Staphylococcus aureus cell wall synthesis in the absence of the division machinery. Mol. Microbiol. 2003;50:871–881. doi: 10.1046/j.1365-2958.2003.03719.x. [DOI] [PubMed] [Google Scholar]
- Seng P., Drancourt M., Gouriet F., La Scola B., Fournier P.-.E., Rolain J.M., Raoult D. Ongoing revolution in bacteriology: routine identification of bacteria by matrix-assisted laser desorption ionization time-of-flight mass spectrometry. Clin. Infect. Dis. 2009;49:543–551. doi: 10.1086/600885. [DOI] [PubMed] [Google Scholar]
- Sundsfjord A., Simonsen G.S., Haldorsen B.C., Haaheim H., Hjelmevoll S.-.O., Littauer P., Dahl K.H. Genetic methods for detection of antimicrobial resistance. APMIS. 2004;112:815–837. doi: 10.1111/j.1600-0463.2004.apm11211-1208.x. [DOI] [PubMed] [Google Scholar]
- Tong S.Y.C., Davis J.S., Eichenberger E., Holland T.L., Fowler V.G. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin. Microbiol. Rev. 2015;28:603–661. doi: 10.1128/CMR.00134-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Belkum A., Bachmann T.T., Lüdke G., Lisby J.G., Kahlmeter G., Mohess A., Becker K., Hays J.P., Woodford N., Mitsakakis K., Moran-Gilad J., Vila J., Peter H., Rex J.H., Dunne Wm.M. Developmental roadmap for antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 2019;17:51–62. doi: 10.1038/s41579-018-0098-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Belkum A., Burnham C.-A.D., Rossen J.W.A., Mallard F., Rochas O., Dunne W.M. Innovative and rapid antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 2020;18:299–311. doi: 10.1038/s41579-020-0327-x. [DOI] [PubMed] [Google Scholar]
- Vickers R.J., Bassetti M., Clancy C.J., Garey K.W., Greenberg D.E., Nguyen M.-.H., Roblin D., Tillotson G.S., Wilcox M.H. Combating resistance while maintaining innovation: the future of antimicrobial stewardship. Future Microbiol. 2019;14:1331–1341. doi: 10.2217/fmb-2019-0227. [DOI] [PubMed] [Google Scholar]
- Wilson, M.L., Weinstein, M.P., Reller, B., 2015. Laboratory detection of bacteremia and fungemia, 11th ed. [PubMed]
- Wolk D.M., Johnson K. Rapid diagnostics for blood cultures: supporting decisions for antimicrobial therapy and value-based care. J. Appl. Lab. Med. 2019;3:686–697. doi: 10.1373/jalm.2018.028159. [DOI] [PubMed] [Google Scholar]
- World Health Organization . World Health Organization; 2019. Landscape of Diagnostics Against Antibacterial resistance, Gaps and Priorities. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.