Abstract
A PCR approach was used to construct a database of nasA genes (called narB genes in cyanobacteria) and to detect the genetic potential for heterotrophic bacterial nitrate utilization in marine environments. A nasA-specific PCR primer set that could be used to selectively amplify the nasA gene from heterotrophic bacteria was designed. Using seawater DNA extracts obtained from microbial communities in the South Atlantic Bight, the Barents Sea, and the North Pacific Gyre, we PCR amplified and sequenced nasA genes. Our results indicate that several groups of heterotrophic bacterial nasA genes are common and widely distributed in oceanic environments.
The importance of inorganic N (NH4+ or NO3−) for the nutrition and growth of marine phytoplankton has long been recognized (5, 7, 8), while the utilization of inorganic N by bacteria has historically received less attention (11, 13, 15, 17, 43). The primary role of heterotrophic bacteria is classically considered to be the decomposition and mineralization of dissolved and particulate organic nitrogen (27). Bacterial NO3− assimilation is not a pathway currently considered in pelagic carbon and nitrogen cycle models (1, 6, 10). A recent review of freshwater and marine studies, however, reported that bacteria may rely on both NH4+ and NO3− for growth and biomass synthesis, and overall they may be significant consumers of inorganic N; mean consumption values of 30 and 40% have been reported for NH4+ and NO3−, respectively (14). Under certain conditions, such as in the presence of high concentrations of dissolved organic carbon relative to the concentration of dissolved organic nitrogen, bacteria may be responsible for most, if not all, of the observed NO3− uptake and disappearance (16, 24, 25). Significant heterotrophic bacterial utilization of dissolved inorganic nitrogen likely would have profound effects on the fluxes of N and C in the water column.
Bacterial nitrate utilization in aquatic communities, however, is difficult to study by conventional tracer approaches. Within the bacterial size class, autotrophic cyanobacteria (picoplankton) are often abundant (4, 41) and are likely to complicate conclusions regarding the total flux of labeled nitrogen into the heterotrophic fraction of the bacterial community. Also, size fractionation does not allow for examination of nitrate uptake by attached bacteria or large cells caught in filters.
It is known that some, but not all, heterotrophic bacteria are capable of growth on NO3− as a sole N source (28). In Klebsiella pneumoniae, the structural genes for nitrate assimilation form an operon, nasFEDCBA (19–21). The NASC protein is thought to mediate electron transfer from NADH to NASA, which contains the active site for nitrate reduction (19, 20). The NASA protein has also been purified from a phototrophic member of the alpha subclass of the class Proteobacteria, Rhodobacter capsulatus, and characterized (2, 22). Examination of currently available prokaryotic genome sequences suggests that nasA is present in a wide diversity of organisms, although these observations need verification (28).
Molecular techniques can be employed to illuminate factors which control the rates of fluxes and transformations in nitrogen-cycling processes (40, 46, 47, 51). Molecular approaches have been successfully used to detect and characterize bacteria and the genes that are important in several aspects of the nitrogen cycle, including nitrification, dentrification, and nitrogen fixation (18, 29, 30, 37–39, 45, 48–50).
Here we describe the design and optimization of a series of nested heterotrophic bacterium-specific nasA PCR primers. The detection of nasA genes in a variety of marine environments provided a basis for the hypothesis that the potential for NO3− utilization by heterotrophic bacteria is significant. Phylogenetic analysis of nasA genes cloned from diverse samples indicated that there are several distinct clades and suggests that there is a clear genetic distinction between nasA genes from heterotrophic bacteria and nasA genes from autotrophic cyanobacteria.
Initially, three nested universal degenerate nasA primers were designed based on five previously determined sequences from cyanobacteria and one sequence from a heterotrophic bacterium (3, 12, 20, 23, 34). The sequences from cyanobacterial strains were from Oscillatoria chalybea, Anabaena sp. strain PCC7120, Synechocystis sp. strain PCC6803, a Synechococcus sp., and Synechococcus sp. strain 7942, and the sequence from a heterotrophic strain was from Klebsiella oxytoca. The GenBank accession numbers for these sequences are X89445, L49163, BAA17488, CAA52675, P39458, and L06800, respectively. An alignment of the inferred amino acid sequences encoded by nasA indicated that there were conserved regions suitable for targeting by PCR oligonucleotide primers. Such primers have been used to amplify nasA sequences in other heterotrophic bacteria, and a group-specific degenerate primer was designed to specifically amplify the nasA gene from heterotrophic bacteria. All of the primers used in this study are listed in Table 1.
TABLE 1.
Oligonucleotide primers used in this study
| Primer | Sequence (5′ to 3′) | Amino acid sequence | Application |
|---|---|---|---|
| nas22 | TGYCCNTAYTGYGGNGT | CPYCG | nasA/narB PCR amplification |
| nas964 | CARCCNAAYGCNATGGG | QPNAM | nasA/narB PCR amplification |
| nasA1735 | ATNGTRTGCCAYTGRTC | DQWHT | nasA PCR amplification |
| nas1933 | CARTGCATNGGNAYRAA | F/L V/I/M PMH | nasA/narB PCR amplification |
| fd1 | AGAGTTTGATCCTGGCTCAG | 16S rRNA forward primer | |
| rp2 | ACGGCTACCTTGTTACGACTT | 16S rRNA reverse primer | |
| M13F | GTAAAACGACGGCCAG | Forward sequencing primer, M13 vector | |
| 522F | CAGCCGCGGTAATAC | Forward sequencing primer, 16S rRNA | |
| 1056F | TGGCTGTCGTCAGCTCGTGT | Forward sequencing primer, 16S rRNA | |
| M13R | CAGGAAACAGCTATGAC | Reverse sequencing primer, M13 vector | |
| 1056R | ACACGAGCTGACGACAGCCA | Reverse sequencing primer, 16S rRNA | |
| 522R | GTATTACCGCGGCTG | Reverse sequencing primer, 16S rRNA |
Surface water samples were collected during two cruises in the South Atlantic Bight (SAB) off the Georgia coast aboard the R/V Bluefin during October 1998 and aboard the R/V Cape Hatteras during March 1999 (31 to 33°N, 78 to 81°W). SAB samples were also collected from docks located on the Skidaway River (March 1999) and the Wilmington River (July 1998). Additional water samples used in this study were collected at depths of 5, 30, and 80 m in the Barents Sea (70 to 78°N, 30°E) aboard the R/V Jan Mayen during July 1999 and from the surface of the North Pacific Gyre at Hawaii Ocean Time Series stations (22°45′N, 158°W) during May 1997 and aboard the R/V Melville during June 1999. For DNA extraction, bacteria were collected from 40 liters of water. To remove eukaryotic plankton, the water was prefiltered under a vacuum through a 3-μm-pore-size polycarbonate cartridge filter (Gelman Sciences, Inc., Ann Arbor, Mich.) and then through a 142-mm-diameter, 0.8-μm-pore-size polycarbonate Supor filter (Gelman) with a custom-manufactured acrylic filter holder. Bacterial cells in the filtrate were collected on a 142-mm-diameter, 0.2-μm-pore-size polycarbonate Supor filter (Gelman) and stored at −20°C aboard ship and then transferred to storage at −80°C in the lab. DNA in the SAB samples collected in October 1998 was extracted as described by Gonzalez et al. (9), and DNA in all other samples was extracted with an UltraClean Mega Prep soil DNA kit (Mo Bio Laboratories, Inc., Solana Beach, Calif.). For the latter procedure, frozen filters were crushed inside Whirl-Pak bags (Nasco, Fort Atkinson, Wis.) and put directly into the lysing matrix used for step one of the soil DNA extraction procedure. Visualization of purified DNA by gel electrophoresis revealed the presence of high-molecular-weight DNA with little shearing and no RNA contamination. From 40 liters of seawater, this method typically yielded an average of 100 to 110 μg of DNA. If it was assumed that the average concentration bacterial cells was 106 cells per ml of seawater and the average DNA content was 3 fg/cell (31), the approximate extraction efficiency of this method was between 80 and 90%.
PCR was performed by using a nested format to improve specificity and sensitivity. The PCR products obtained with the outermost degenerate nasA/narB universal primers (nas22, nas1933) were subsequently used as templates in PCR with the heterotroph-specific internal primer set (nas964, nasA1735). Amplification was accomplished by using the Qiagen Taq PCR Master Mix System and the standard protocol recommended by the vendor (Qiagen, Valencia, Calif.); a hot start at 94°C for 5 min was followed by 35 cycles consisting of 94°C for 5 s, 55.5°C for 10 s, and 72°C for 1 min, with a 7-min final extension step at 72°C. DNA template (10 to 100 ng of community DNA or 0.1 to 10 ng of genomic DNA from a pure culture) was added to each 25-μl PCR mixture. First-round reaction mixtures contained 1 μM primer nas22, 1 μM primer nas1933, and 3.5 mM MgCl2. Second-round nested PCR mixtures contained 1 to 2 μl of product from the first round, 2.5 mM MgCl2, 1 μM primer nas964, and 1 μM primer nasA1735, and the extension time in each cycle was decreased to 30 s. The nasA-specific primers yielded a PCR product that was 750 to 800 bp long. 16S rRNA amplification of the nearly complete 16S rRNA gene was facilitated by using eubacterial primers fd1 and rp2 (Table 1) (42) (100 nM each). Thermal cycling was performed with a model 2400 or 9700 thermal cycler (Perkin-Elmer Corp., Norwalk, Conn.).
Although nasA PCR could be optimized for specific community DNA samples by raising the annealing temperature to 57 to 60°C, a somewhat less stringent annealing temperature, 55°C, was used during the initial construction of clone libraries to increase the yield and the likelihood of amplification with most primer-template combinations.
The PCR product of the desired size was excised from the gel and purified by using GenElute agarose spin columns (Supelco, Bellefonte, Pa.). PCR products were ligated and cloned by using either a TOPO TA Cloning kit (for pure cultures) or an Original TA Cloning kit (for community DNA samples). In both cases, the PCR product was ligated into a pCR 2.1 plasmid vector and cloned into TOP10 One Shot competent Escherichia coli cells (Invitrogen, Carlsbad, Calif.). Plasmid DNA was extracted and purified by using the Wizard Plus Minipreps DNA purification system (Promega, Madison, Wis.).
Sequences were determined by automated sequencing at the Molecular Genetics Facility (University of Georgia) with ABI automated sequencers (models 373 and 377). Sequencing reactions were facilitated by using an ABI Big Dye Prism dideoxy sequencing dye terminator kit as recommended by the manufacturer. Sequence analysis was accomplished by using ABI software, version 3.3 (ABI, Foster City, Calif.). The sequencing primers used are listed in Table 1.
Bacteria were isolated from seawater samples collected from the SAB continental shelf during March and June 1999 (31 to 33°N, 78 to 81°W). Bacteria were also isolated from Barents Sea water samples collected during July 1999 (70 to 80°N, 30°E). Bacteria were isolated by using either organic nitrogen or nitrate as the sole nitrogen source. Selected colonies were axenically transferred to new plates twice to ensure that pure cultures were obtained. For long-term storage each isolate was maintained in a 15% (vol/vol) glycerol freezer stock preparation at −80°C.
To screen isolated strains for the presence of nasA, PCR-amenable DNA was extracted from each of the isolates by using the FastDNA spin protocol and a Fast Prep instrument (both from BIO 101, Vista, Calif.). In all PCRs, appropriate negative controls without DNA and positive controls were included.
To test isolates for the ability to grow on nitrate as the sole N source, two tubes containing 5 ml of NFG medium (33) were prepared. To one of the tubes a sterile NaNO3− solution was added to obtain a final NO3− concentration of 10 mM. The second tube did not receive such an addition and served as a negative control. The two NFG medium tubes were then inoculated 1:100 with a stationary-phase culture grown in peptone- and yeast-enriched artificial seawater (26). After 72 h, the optical densities of the two tubes were compared to the optical density of a NFG medium tube that had not been inoculated. Additionally, several isolates were selected for batch culture growth assays. These experiments were conducted in axenic 100-ml NFG medium cultures containing 80 μM or 10 mM NO3− as the sole N source. Doubling rates were determined by estimating cell density at a minimum of four time points during the exponential growth phase. Cell densities were determined by direct epiflourescent microscopy after staining with DAPI (4′,6′-diamidino-2-phenylindole) (44).
Phylogenetic relationships based on nasA gene sequences were determined. Nucleotide sequences were translated into approximately 264 unambiguous amino acids. All of the available narB/nasA amino acid sequences were then aligned by using the CLUSTAL W (version 1.7) multiple-sequence-alignment algorithm (32). Phylogenetic trees were inferred and drawn by using the TREECON software package (version 1.3b) (35, 36) and the Kimura two-parameter model for inferring evolutionary distances. Bootstrap estimates (100 replicates) of confidence intervals were also made by using the algorithms available in the TREECON package.
For 16S rRNA analysis, 464 unambiguously alignable nucleotide positions were used. The nucleotide sequences were compared to 16S rRNA gene sequences available in the GenBank database by using the Blast program to determine the degrees of sequence similarity to known organisms. All of the nasA and 16S rRNA sequences determined in this study (Table 2) have been deposited in the GenBank database.
TABLE 2.
Srains used in this study and 16S rRNA accession numbers
| Strain | Closest relative | nasA PCR amplification | 16S rRNA accession no. |
|---|---|---|---|
| Known cultures | NAc | + | |
| Fischerella sp.a | NA | + | |
| Plectonema boryanuma | NA | + | |
| Trichodesmium sp. strain IMS101a | NA | + | |
| Klebsiella pneumoniae ATCC 13883a | NA | + | |
| Clostridium oceanicaa | NA | + | |
| Pseudomonas sp.a | NA | + | |
| Vibrio diazotrophicusa | NA | + | |
| Bacillus sp. | NA | − | |
| Micrococcus luteus | NA | − | |
| Vibrio sp. strain S-14 | NA | − | |
| Pseudomonas stutzeri | NA | − | |
| Sagittula stellata E37 | NA | − | |
| Strains isolated in this study | − | ||
| South Atlantic Bight Ab | Cytophaga sp. | + | AF300973 |
| South Atlantic Bight Bb | Aerococcus viridans | − | AF300974 |
| South Atlantic Bight Cb | Alteromonas sp. strain MS23 | + | AF300975 |
| Barents Sea isolate 4b | Pseudoalteromonas sp. strain ANG.ro2 | + | AF300976 |
| Barents Sea isolate 10b | Marinobacter sp. strain DS40M8 | + | AF300977 |
| Barents Sea isolate 23 | Pseudoalteromonas haloplanktis | − | AF300978 |
| Barents Sea isolate 25b | Pseudoalteromonas citrea | + | AF300979 |
| Barents Sea isolate 32 | Erythrobacter citreus | − | AF300980 |
| Skidaway River 1b | Vibrio carchariae | + | AF300981 |
| Skidaway River 2b | Vibrio furnissii ATCC 35016T | + | AF300982 |
| Skidaway River 3 | Micrococcus luteus FO-084a | − | AF300983 |
Sequence amplified in this study by using oligonucleotide primers nas964 and nas1934.
Sequence amplified in this study by using oligonucleotide primers nas964 and nas1735.
NA, not applicable.
Results.
Using the universal nasA nested primer set, we amplified, cloned, and sequenced a 1,000-bp fragment from a group of phylogenetically diverse bacteria, including Clostridium oceanica, Vibrio diazotrophicus, a Pseudomonas sp., Trichodesmium sp. strain IMS101, a Fischerella sp., and Plectonema boryanum, as well as from DNA extracted from the bacterial size fraction of seawater collected at a Hawaii Ocean Times Series station near Hawaii. Also, we attempted to amplify the 1,000-bp nasA fragment from Bacillus sp., Micrococcus luteus, Vibrio sp. strain S-14, and Pseudomonas stutzeri. These templates, however, did not yield a PCR product, and we concluded that they were nasA negative (Table 2).
Using the expanded database of nasA sequences, we targeted an additional reverse primer, at amino acid position 579, for heterotrophic organisms. The heterotroph-specific primer was nasA1735 (Table 1) and was approximately 200 bp downstream from the universal nasA reverse primer.
Using a collection of isolates obtained during a cruise in the Barents Sea, we examined the relationship between the presence of the nasA gene and the ability to utilize NO3− as a sole N source during aerobic growth. Of the 30 isolates screened, 17 were able to grow by using NO3− as a sole N source. All of these strains were PCR positive for the nasA gene fragment. Thirteen of the isolates screened could not grow on NO3− alone, and none of these strains contained a nasA gene fragment. Three isolates from SAB water and three isolates from the Skidaway River estuary in Georgia were also screened. Two of these six isolates were NO3− growth positive and nasA PCR positive, two were NO3− growth negative and nasA PCR negative, and two were NO3− growth negative and nasA PCR positive. Therefore, of 36 isolates examined, 19 were PCR positive for nasA and had the ability to utilize NO3− as a sole N source, 15 were PCR negative for nasA and were not able to utilize NO3−, and two displayed somewhat contradictory results because they were PCR positive for nasA and apparently not able to grow on NO3− as a sole N source (Table 3). The results of the batch growth assays indicated that there was some variability between isolates in terms of their affinity for NO3− (Table 3). Data are reported here only for experiments conducted with 10 mM NO3−. Experiments conducted with 80 μM NO3− generated similar doubling times for the different strains tested, but the final cell yields were lower.
TABLE 3.
Abilities of bacterial isolates to grow on nitrate media and detection of nasA PCR gene product
| Isolate | Growth on NO3− as sole nitrogen source | nasA PCR amplification |
|---|---|---|
| Barents Sea isolates | ||
| Barents Sea isolate 4a | Y | + |
| Barents Sea isolate 10a | Y | + |
| Barents Sea isolate 23 | N | − |
| Barents Sea isolate 25a | Y | + |
| Barents Sea isolate 32 | N | − |
| Barents Sea isolate 1 | N | − |
| Barents Sea isolate 2 | Y | + |
| Barents Sea isolate 3 | N | − |
| Barents Sea isolate 5 | N | − |
| Barents Sea isolate 6 | Y | + |
| Barents Sea isolate 7 | N | − |
| Barents Sea isolate 8 | Y | + |
| Barents Sea isolate 9 | Y | + |
| Barents Sea isolate 12 | Y | + |
| Barents Sea isolate 13 | Y | + |
| Barents Sea isolate 14 | Y | + |
| Barents Sea isolate 15 | Y | + |
| Barents Sea isolate 16 | Y | + |
| Barents Sea isolate 18 | Y | + |
| Barents Sea isolate 19 | N | − |
| Barents Sea isolate 20 | N | − |
| Barents Sea isolate 21 | N | − |
| Barents Sea isolate 22 | N | − |
| Barents Sea isolate 24 | Y | + |
| Barents Sea isolate 26 | N | − |
| Barents Sea isolate 27 | Y | + |
| Barents Sea isolate 28 | N | − |
| Barents Sea isolate 29 | Y | + |
| Barents Sea isolate 30 | Y | + |
| Barents Sea isolate 31 | N | − |
| SAB isolates | ||
| South Atlantic Bight A | N | + |
| South Atlantic Bight B | N | − |
| South Atlantic Bight C | N | + |
| Skidaway River 1 | Y | + |
| Skidaway River 2 | Y | + |
| Skidaway River 3 | N | − |
The doubling times of Barents Sea isolates 4, 10, and 25 on 10 mM NO3− as the sole N source were 3.78, 5.48, and 5.16 h, respectively.
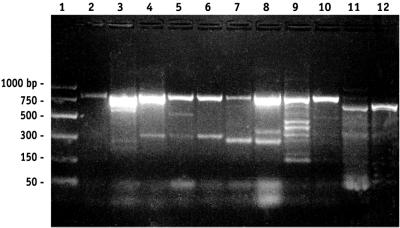
nasA was detected in all of the environments examined, including the SAB, the North Pacific Gyre, a Norwegian fjord, and the Barents Sea (Fig. 1). The sensitivity of nasA detection by PCR was initially estimated by amending filtered seawater with 103 and 102 K. pneumoniae cells per ml and detecting nasA in the samples (Fig. 1). This approach did not establish a minimum level of detection but demonstrated that a concentration of at least 102 cells positive for the nasA gene per ml could be detected. Since a strong signal was detected in a wide range of marine samples, heterotrophic bacteria with the capacity for NO3− assimilation appear to be very common and well distributed.
FIG. 1.
PCR amplification of the expected 750- to 800-bp nasA gene fragment from various marine samples. Lane 2 contained a PCR product from a sample that was prepared by amending filtered seawater with 103 K. pneumoniae cells/ml (final concentration). nasA PCR products were amplified from samples collected from a Norwegian fjord (lane 3), the Barents Sea (lanes 4 to 6), the Skidaway River (lanes 7 and 8), the SAB (lanes 9 and 10), and the North Pacific Gyre (lanes 11 and 12). Lane 1 contained a molecular weight standard PCR marker (Promega).
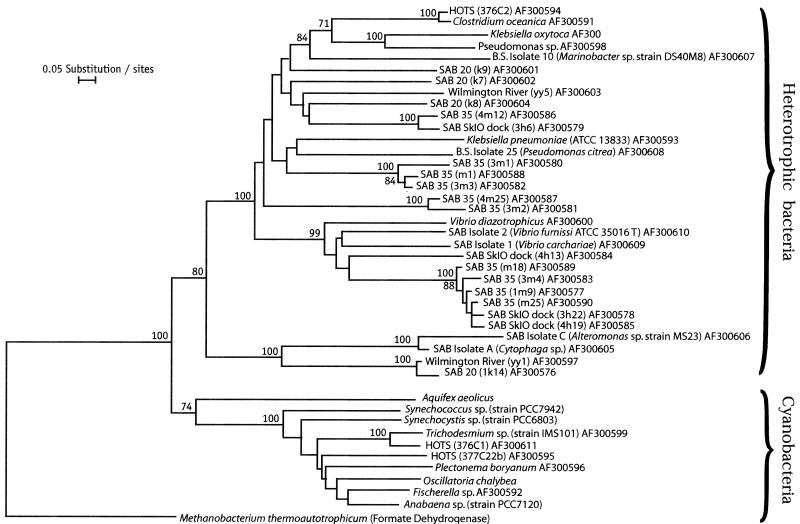
In general, nasA genes from uncultured organisms do not form clades separate from the clades of cultured bacteria. Also, nasA genes in taxonomically related bacteria are not necessarily similar, except in the case of Vibrio representatives. V. diazotrophicus and two Vibrio isolates form a separate clade which includes seven clones from samples collected in the Skidaway River and the SAB midshelf (35 miles offshore). K. pneumoniae ATCC 13883 and a Pseudoalteromonas isolate typify another clade, which includes three clones from SAB midshelf samples. A third discrete cluster includes C. oceanica, K. oxytoca, a Pseudomonas sp., and a Marinobacter sp. isolate. Also in this cluster are clones from the midshelf (25 and 35 miles offshore), the Skidaway River, and the adjacent Wilmington River (Fig. 2).
FIG. 2.
Inferred phylogenetic relationships of nasA- and narB-encoded amino acid sequences from heterotrophic bacteria and cyanobacteria, respectively. The scale bar indicates 0.05 fixed amino acid substitution per site. The numbers at the nodes are bootstrap values. Bootstrap values less than 70 (of 100) are not shown. The amino acid sequence of formate dehydrogenase from Methanobacterium thermoautotrophicum (GenBank accession number U52681), a putative evolutionary ancestor of the proteins encoded by the nasA and narB genes, was used to root the tree. HOTS, Hawaii Ocean Time Series.
Among the nitrate-assimilating (nasA-positive) and non-nitrate-assimilating (nasA-negative) isolates whose 16S rRNA genes have been sequenced, there are more different types of taxa associated with the nasA-negative strains. For example, nasA-negative isolates include organisms such as a Micrococcus sp. (gram-positive phylum, high-G+C-content subdivision), an Erythrobacter sp. (alpha subclass of the Proteobacteria), an Aerococcus sp. (gram-positive phylum, low-G+C-content subdivision), Sagittula stellata E37 (alpha subclass of the Proteobacteria), and a Pseudoalteromonas sp. (gamma subclass of the Proteobacteria) (Table 2). By contrast, the majority of the nasA-positive strains whose 16S rRNA have been sequenced are members of the gamma subclass of the Proteobacteria. In particular, members of a Pseudoalteromonas sp. and a Vibrio sp. account for five of seven of the nasA-positive strains that we isolated.
Conclusions.
The correlation between the presence of nasA and nitrate utilization assay results for individual isolates (34 of 36 isolates tested) supports the hypothesis that the nasA-specific primer sets developed in this study provide a reliable assay for functional assimilatory nitrate reductase genes. Sequences derived from isolates that were unable to utilize NO3− in culture (Table 3) are more closely related to dehydrogenases and proteins encoded by members of other gene families and can be distinguished phylogenetically from functional assimilatory nitrate reductases (Fig. 2). This illustrates the fact that although degenerate primers are powerful and able to retrieve gene fragments from very diverse organisms, it is important to sequence and phylogenetically analyze PCR products from as many different types of organisms as possible in order to identify potential nonspecific PCR products.
We demonstrated that genetic probes that recognize the functional assimilatory nitrate reductase genes of specific groups of bacteria can be constructed. Heterotrophic nasA genes have been detected in every marine sample tested thus far, indicating that bacteria capable of assimilating nitrate are ubiquitous in diverse ocean margins and open water. These observations suggest that heterotrophic bacteria are potentially important consumers of NO3− in marine environments.
Nucleotide sequence accession numbers.
The nasA and 16S rRNA sequences determined in this study have been deposited in the GenBank database under the accession numbers shown in Table 2 and Fig. 2.
ACKNOWLEDGMENTS
We thank G. P. Paffenhoffer, D. Bronk, J. Bower, and P. Wassman for providing ship time, and we thank the crews of the R/V Bluefin, the R/V Hatteras, and the R/V Jan Mayen. We also thank M. A. Moran for donating bacterial strains. In addition, we thank H. Howard-Jones for help with microscopy, S. McIntosh and A. Boyette for preparing figures, and Dee Peterson for preparing the manuscript.
This research was supported by grants DE FG02-88ER62531 and DE-FG02-98ER62531 from the U.S. Department of Energy.
REFERENCES
- 1.Bissett W, Walsh J, Dieterle D, Carder K. Carbon cycling in the upper waters of the Sargasso Sea. 1. Numerical simulation of differential carbon and nitrogen fluxes. Deep-Sea Res. 1999;46:205–269. [Google Scholar]
- 2.Blasco R, Castillo F, Matinez-Luque M. The assimilatory nitrate reductase from the phototrophic bacterium, Rhodobacter capsulatusE1F1, is a flavoprotein. FEBS Lett. 1997;414:45–49. doi: 10.1016/s0014-5793(97)00968-x. [DOI] [PubMed] [Google Scholar]
- 3.Cai Y, Wolk C P. Nitrogen deprivation of Anabaenasp. strain PCC 7120 elicits rapid activation of a gene cluster that is essential for uptake and utilization of nitrate. J Bacteriol. 1997;179:258–266. doi: 10.1128/jb.179.1.258-266.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Chisholm S W, Olsen R J, Zehler E R, Goericke R, Waterbury J B, Welschmeyer N A. A novel free-living prochlorophyte abundant in the oceanic euphotic zone. Nature. 1988;334:340–343. [Google Scholar]
- 5.Dugdale R C, Goering J J. Uptake of new and regenerated forms of nitrogen in primary productivity. Limnol Oceanogr. 1967;12:196–206. [Google Scholar]
- 6.Fasham M, Ducklow H, McKelvie S. A nitrogen-based model of plankton dynamics in the oceanic mixed layer. J Mar Res. 1990;48:591–639. [Google Scholar]
- 7.Glibert P M, McCarthy J J. Uptake and assimilation of ammonium and nitrate by phytoplankton: indices of nutritional status for natural assemblages. J Plankton Res. 1984;6:677–697. [Google Scholar]
- 8.Goldman J C, Gilbert P M. Kinetics of inorganic nitrogen uptake by phytoplankton. In: Carpenter E J, Capone D G, editors. Nitrogen in the marine environment. New York, N.Y: Academic Press; 1983. pp. 233–276. [Google Scholar]
- 9.Gonzalez J M, Whitman W B, Hodson R E, Moran M A. Identifying numerically abundant culturable bacteria from complex communities: an example from a lignin enrichment culture. Appl Environ Microbiol. 1996;62:4433–4440. doi: 10.1128/aem.62.12.4433-4440.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Haupt O, Wolf U, Bodungen B. Modelling the pelagic nitrogen cycle and vertical particle flux in the Norwegian Sea. J Mar Syst. 1999;19:173–199. [Google Scholar]
- 11.Horrigan S G, Hagstrom A, Koike I, Azam F. Inorganic nitrogen utilization by assemblages of marine bacteria in seawater culture. Mar Ecol Prog Ser. 1988;50:147–150. [Google Scholar]
- 12.Kaneko T, Sato S, Kotani H, Tanaka A, Asamizu E, Nakamura Y, Miyajima N, Hirosawa M, Sugiura M, Sasamoto S, Kimura T, Hosouchi T, Matsuno A, Muraki A, Nakazaki N, Naruo K, Okumura S, Shimpo S, Takeuchi C, Wada T, Watanabe A, Yamada M, Yasuda M, Tabata S. Sequence analysis of the genome of the unicellular cyanobacterium Synechocystissp. strain PCC6803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions. DNA Res. 1996;3:109–136. doi: 10.1093/dnares/3.3.109. [DOI] [PubMed] [Google Scholar]
- 13.Keil R G, Kirchman D L. Contribution of dissolved free amino acids and ammonium to the nitrogen requirements of heterotrophic bacterioplankton. Mar Ecol Prog Ser. 1991;73:1–10. [Google Scholar]
- 14.Kirchman D L. Uptake and regeneration of inorganic nutrients by marine heterotrophic bacteria. In: Kirchman D L, editor. Microbial ecology of the oceans. New York, N.Y: John Wiley & Sons; 2000. pp. 261–288. [Google Scholar]
- 15.Kirchman D L. The uptake of inorganic nutrients by heterotrophic bacteria. Microb Ecol. 1994;28:255–271. doi: 10.1007/BF00166816. [DOI] [PubMed] [Google Scholar]
- 16.Kirchman D L, Suzuki Y, Garside C, Ducklow H W. High turnover rates of dissolved organic carbon during a spring phytoplankton bloom. Nature. 1991;352:612–614. [Google Scholar]
- 17.Kirchman D L, Wheeler P A. Uptake of ammonium and nitrate by heterotrophic bacteria and phytoplankton in the sub-arctic Pacific. Deep-Sea Res. 1998;45:347–365. [Google Scholar]
- 18.Kirshtein J D, Paerl H W, Zehr J. Amplification, cloning, and sequencing of a nifHsegment from aquatic microorganisms and natural communities. Appl Environ Microbiol. 1991;57:2645–2650. doi: 10.1128/aem.57.9.2645-2650.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lin J T, Glodman B S, Stewart V. The nasFEDCBA operon for nitrate and nitrite assimilation in Klebsiella pneumoniaeM5a1. J Bacteriol. 1994;176:2551–2559. doi: 10.1128/jb.176.9.2551-2559.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lin J T, Goldman B S, Stewart V. Structures of genes nasA and nasB, encoding assimilatory nitrate and nitrite reductases in Klebsiella pneumoniaeM5a1. J Bacteriol. 1993;175:2370–2378. doi: 10.1128/jb.175.8.2370-2378.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lin J T, Stewart V. Nitrate- and nitrite-mediated transcription antitermination control of nasF (nitrate assimilation) operon expression in Klebsiella pneumoniaeM5a1. J Mol Biol. 1996;256:423–435. doi: 10.1006/jmbi.1996.0098. [DOI] [PubMed] [Google Scholar]
- 22.Moreno-Vivan C, Cabello P, Martinez-Luque M, Blasco R, Castillo F. Prokaryotic nitrate reduction: molecular properties and functional distinction among bacterial nitrate reductases. J Bacteriol. 1999;181:6573–6584. doi: 10.1128/jb.181.21.6573-6584.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Omata T, Andriesse X, Hirano A. Identification and characterization of a gene cluster involved in nitrate transport in the cyanobacterium SynechococcusPCC7942. Mol Gen Genet. 1993;236:193–202. doi: 10.1007/BF00277112. [DOI] [PubMed] [Google Scholar]
- 24.Parker R R, Sibert J, Brown T J. Inhibition of primary productivity through heterotrophic competition for nitrate in a stratified estuary. J Fish Res Board Can. 1975;32:72–77. [Google Scholar]
- 25.Parsons T R, Albright L J, Whitney F, Wong C S, Williams P J. The effect of glucose on the productivity of seawater: an experimental approach using controlled aquatic ecosystems. Mar Environ Res. 1980;4:229–242. [Google Scholar]
- 26.Paul J. The use of Hoechst dyes 33258 and 33342 for enumeration of attached and planktonic bacteria. Appl Environ Microbiol. 1982;43:939–944. doi: 10.1128/aem.43.4.939-944.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pomeroy L R. The ocean's food web, a changing paradigm. BioScience. 1974;24:499–504. [Google Scholar]
- 28.Richardson D J, Berks B C, Russell D A, Spiro S, Taylor C J. Functional, biochemical and genetic diversity of prokaryotic nitrate reductases. Cell Mol Life Sci. 2001;58:165–178. doi: 10.1007/PL00000845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Scala D J, Kerkhof L J. Diversity of nitrous oxide reductase (nosZ) genes in continental shelf sediments. Appl Environ Microbiol. 1999;65:1681–1687. doi: 10.1128/aem.65.4.1681-1687.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Scala D J, Kerkhof L J. Nitrous oxide reductase (nosZ) gene-specific PCR primers for detection of denitrifiers and three nosZgenes from marine sediments. FEMS Microbiol Lett. 1988;162:61–68. doi: 10.1111/j.1574-6968.1998.tb12979.x. [DOI] [PubMed] [Google Scholar]
- 31.Simon M, Azam F. Protein content and protein synthesis rates of planktonic marine bacteria. Mar Ecol Prog Ser. 1989;51:201–213. [Google Scholar]
- 32.Thompson J D, Higgins D G, Gibson T J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tibbles B J, Rawlings D E. Characterization of nitrogen-fixing bacteria from a temperate saltmarsh lagoon, including isolates that produce ethane from acetylene. Microb Ecol. 1994;27:65–80. doi: 10.1007/BF00170115. [DOI] [PubMed] [Google Scholar]
- 34.Unthan M, Klipp W, Schmid G H. Nucleotide sequence of the nar beta gene encoding assimilatory nitrate reductase from the cyanobacterium Oscillatoria chalybea. Biochim Biophys Acta. 1996;1305:19–24. doi: 10.1016/0167-4781(95)00210-3. [DOI] [PubMed] [Google Scholar]
- 35.Van de Peer Y, De Wachter R. Construction of evolutionary distance trees with TREECON for Windows: accounting for variation in nucleotide substitution rate among sites. Comput Applic Biosci. 1997;13:227–230. doi: 10.1093/bioinformatics/13.3.227. [DOI] [PubMed] [Google Scholar]
- 36.Van de Peer Y, De Wachter R. TREECON for Windows: a software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment. Comput Applic Biosci. 1994;10:569–570. doi: 10.1093/bioinformatics/10.5.569. [DOI] [PubMed] [Google Scholar]
- 37.Voytek M A, Priscu J C, Ward B B. The distribution and relative abundance of ammonium-oxidizing bacteria in lakes of the McMurdo Dry Valley, Antarctica. Hydrobiologia. 1999;401:113–130. [Google Scholar]
- 38.Voytek M A, Ward B B. Detection of ammonium-oxidizing bacteria in the beta-subclass of the class Proteobacteriain aquatic samples with the PCR. Appl Environ Microbiol. 1995;61:1444–1450. doi: 10.1128/aem.61.4.1444-1450.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Voytek M A, Ward B B, Priscu J C. Ecosystem dynamics in a polar desert: the McMurdo Dry Valleys, Antarctica. Washington, D.C.: American Geophysical Union; 1997. The abundance of ammonium-oxidizing bacteria in Lake Bonney, Antarctica, determined by immunofluorescence, PCR and in situ hybridization; pp. 217–228. [Google Scholar]
- 40.Ward B B. Nitrification and denitrification: probing the nitrogen cycle in aquatic environments. Microb Ecol. 1996;32:247–261. doi: 10.1007/BF00183061. [DOI] [PubMed] [Google Scholar]
- 41.Waterbury J B, Watson S W, Guillard R R, Brane L E. Widespread occurrence of a unicellular, marine, planktonic cyanobacterium. Nature. 1979;277:293–294. [Google Scholar]
- 42.Weisburg W G, Barns S M, Pelletier D A, Lane D J. 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol. 1991;173:697–703. doi: 10.1128/jb.173.2.697-703.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wheeler P A, Kirchman D L. Utilization of inorganic and organic nitrogen by bacteria in marine systems. Limnol Oceanogr. 1986;31:998–1009. [Google Scholar]
- 44.Williams S C, Hong Y, Danavall D C A, Howard-Jones M H, Gibson D, Frischer M E, Verity P G. Distinguishing between living and nonliving bacteria: evaluation of the vital stain propidium iodide and its combined use with molecular probes in aquatic samples. J Micobiol Methods. 1998;35:225–236. [Google Scholar]
- 45.Zani S, Mellon M T, Collier J T, Zehr J P. Expression of nifHgenes in natural microbial assemblages in Lake George, New York, detected by reverse transcriptase PCR. Appl Environ Microbiol. 2000;66:3119–3124. doi: 10.1128/aem.66.7.3119-3124.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zehr J P, Capone D G. Problems and promises of assaying the genetic potential for nitrogen fixation in the marine environment. Microb Ecol. 1996;32:263–281. doi: 10.1007/BF00183062. [DOI] [PubMed] [Google Scholar]
- 47.Zehr J P, Hiorns W. Molecular approach to studies of the activities of marine organisms. In: Cooksey K E, editor. Molecular approaches to the study of the ocean. London, United Kingdom: Chapman and Hall; 1998. pp. 91–111. [Google Scholar]
- 48.Zehr J P, McReynolds L A. Use of degenerate oligonucleotides for amplification of the nifH gene from the marine cyanobacterium Trichodesmium thiebautii. Appl Environ Microbiol. 1989;55:2522–2526. doi: 10.1128/aem.55.10.2522-2526.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zehr J P, Mellon M T, Zani S. New nitrogen-fixing microorganisms detected in oligotrophic oceans by amplification of nitrogenase (nifH) genes. Appl Environ Microbiol. 1998;64:3444–3450. doi: 10.1128/aem.64.9.3444-3450.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zehr J P, Paerl H W. Nitrogen fixation in the marine environment: genetic potential and nitrogenase expression. In: Cooksey K E, editor. Molecular approaches to the study of the ocean. London, United Kingdom: Chapman and Hall; 1997. pp. 285–301. [Google Scholar]
- 51.Zehr J P, Voytek M A. Molecular ecology of aquatic communities: reflections and future directions. Hydrobiologia. 1999;401:1–8. [Google Scholar]