Abstract
High-risk human papillomavirus (HR-HPV) infection is a major risk factor of head and neck cancers (HNCs). Despite the rising prevalence of HPV-driven HNC (HPV-HNC), biomarkers for detection, prognostication, and disease monitoring are lacking. To evaluate the capacity of salivary HR-HPV DNA as a biomarker of HPV-HNC, the salivary HR-HPV statuses of 491 and 10 patients with primary and recurrent HNC, respectively, were determined at diagnosis, using quantitative real-time PCR and MassARRAY. Tumor cyclin-dependent kinase inhibitor 2A (p16) expression was determined by IHC analysis. Patients with oropharyngeal cancer (OPC) (n = 215) were followed up for ≤5 years. Survival characteristics were evaluated in terms of event-free and cause-specific survival. Of the primary-HNC cohort, 43.2% were positive for salivary HR-HPV DNA, with most having OPC. Salivary HR-HPV DNA was detected in 81.4% of tumor p16–positive OPC patients at diagnosis. Prognosis in salivary HR-HPV–positive OPC patients was favorable compared with that in salivary HR-HPV–negative patients (event-free survival, hazard ratio = 0.42 [95% CI, 0.21–0.81, P = 0.010]; cause-specific survival, hazard ratio = 0.39 [95% CI, 0.18–0.86, P = 0.019]). In the recurrent-HNC cohort, salivary HR-HPV DNA was detected in 83.3% of those who previously had tumor p16–positive HNC. These findings indicate that this liquid biopsy–based, noninvasive biomarker can play an essential role in the detection and management of HPV-HNC.
Infectious agents, including viruses such as human papillomavirus (HPV), account for over 15% of the worldwide cancer burden.1 Despite all of the undesirable aspects of cancer being transmissible, the association of these cancers with infectious agents renders them preventable through eradication of the infectious agent. Furthermore, it allows these cancers to be identified early by coupling of the detection of the involved organism with other investigations. Even though significant progress has been made over the years in terms of the identification and prevention of these cancers, certain cancers such as HPV-driven head and neck cancers (HPV-HNCs) are still usually diagnosed at an advanced stage.2
Early detection of HPV-HNC is hindered by several obstacles. The overwhelming majority of HPV-HNC arises in the oropharynx, which is a relatively difficult site to assess.3 Detection is further complicated if these smaller lesions are hidden in areas such as tonsillar crypts. Additionally, these cancers have the capacity to metastasize into regional lymph nodes at an early stage, even when the primary cancer is still undetectable in size.4, 5, 6 Consequently, most patients present with nodal metastasis at the time of diagnosis. Currently, there are no established early-detection methods or screening programs for HPV-HNC despite the rise in prevalence.7,8
Even though the mere presence of a carcinogenic infectious agent does not confirm the presence of associated cancer, it can be used to identify individuals at risk. As such, many studies have investigated the utility of high-risk (HR)-HPV DNA as a biomarker for the early detection of HPV-HNC.8 Considering the proximity of and access to these tumors, salivary HPV detection has been given major emphasis. Population studies suggest that most individuals clear HPV infection within 2 years.9, 10, 11, 12 However, by evaluating the current literature, it can be hypothesized that many of those who go on to develop, or who already have, HPV-HNC also have, and continue to have, HPV DNA in their saliva. This hypothesis is attested to in two recent studies in which it was confirmed that longitudinal salivary HR-HPV testing, coupled with clinical investigations, identified HPV-HNC in asymptomatic individuals.13,14
However, varying degrees of sensitivity of salivary HPV detection in HPV-HNC patients have been reported.15, 16, 17, 18, 19, 20 A general consensus, therefore, has not been reached regarding the utility of HPV detection for HPV-HNC identification and monitoring. The present study investigated the salivary HPV statuses of HNC patients in a larger-scale patient cohort to determine the efficacy of salivary HPV detection as a biomarker of HPV-HNC. Considering that the vast majority of HPV-driven cancers arise from oropharyngeal sites, this study also investigated survival patterns in patients with oropharyngeal cancer (OPC) to evaluate the utility of salivary HR-HPV as a prognostic biomarker of OPC.
Materials and Methods
Ethics Considerations
The study protocol was approved by the Metro South Human Research Ethics Committee (HREC/12/QPAH/381; Brisbane, QLD, Australia). Informed consent was obtained from all of the participants.
Patient Recruitment
HNC patients were recruited from two major hospitals in Brisbane, Australia, from 2012 to 2019. Patients who were >18 years of age were included in the study, and patients who had conditions limiting saliva secretion were excluded. A total of 523 patients were enrolled. Clinical information was obtained from the relevant hospital clinic and tumor board (Supplemental Figure S1).
p16 IHC Analysis
Queensland pathologists performed immunohistochemistry (IHC) analysis for cyclin-dependent kinase inhibitor 2A (p16) as part of the routine clinical assessment of HNC patients using the CINtec p16INK4a histologic examination kit (E6H4 clone; Roche MTM Laboratories, Heidelberg, Germany). Samples were considered positive when strong diffused nuclear and cytoplasmic staining was present in over 70% of tumor tissue.
Saliva Collection and Processing
Saliva samples were collected according to a previously described procedure.21,22 Briefly, patients were requested to tilt the head down to facilitate the passive pooling of saliva for 2 to 5 minutes before expectorating into the collection container. The procedure was continued until 3 to 5 mL of saliva was collected. Samples were transported on ice and immediately aliquoted into 1.5-mL Eppendorf tubes before storage at −80°C.
DNA Extraction
DNA was extracted using the QIAamp DNA Mini Kit (Qiagen, Germantown, MD) according to a previously published procedure.21
HR-HPV Detection
DNA isolated from salivary samples was tested for HPV type 16 by quantitative real-time (qPCR). PCR primers designed to amplify specific regions in the HPV16 E2 gene (forward primer, 5′-AACGAAGTATCCTCTCCTGAAATTATTAG-3′; reverse primer, 5′-CCAAGGCGACGGCTTTG-3′) and HPV E6/7 genes (forward primer, 5′-ACCGGTCGATGTATGTCTTGTTG-3′; reverse primer, 5′-GATCAGTTGTCTCTGGTTGCAAATC-3′) were used for the assay. Hemoglobin subunit β (HBB) (forward primer, 5′-CAACTTCCACGGTTCACC-3′; reverse primer, 5′-GAAGAGCCAAGGACAGGTAC-3′) was used as the internal quality control.23 PCR reaction was performed using PowerUp SYBR Green Master Mix (Applied Biosystems, Waltham, MA). The QuantStudio 7 Flex real-time PCR system (Applied Biosystems) was used for the assay. PCR conditions were 50°C for 2 minutes, 95°C for 10 minutes, 40 cycles at 95°C for 15 seconds and at 60°C for 60 seconds, and a final melting curve analysis at 95°C for 15 seconds, 60°C for 60 seconds, and 95°C for 15 seconds. All samples were tested in duplicate. qPCR results were analyzed using QuantStudio real-time PCR software version 1.3 (Applied Biosystems) using 0.02 as the cutoff value. A cycle threshold value of <40 for any of the HPV16 targets was considered as positivity for salivary HPV16.
Samples that were HPV16 negative and tumor p16 positive were tested for other HR-HPV types by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (Agena Bioscience, San Diego, CA). This assay detects 17 HR-HPV types, namely, 16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, 68, 53, 66, 67, and 73, and 2 low-risk HPV types, 6 and 11.24 The iPlex Pro reagent set (catalog number 10303; Agena Bioscience) was used for the assay. Primers targeting the E6/7 regions of 19 HPV types were used in the initial multiplex PCR assay (catalog number 06010; Agena Bioscience). PCR conditions were 95°C for 2 minutes; 45 cycles at 95°C, 56°C, and 72°C for 30 seconds each; and a final extension at 72°C for 5 minutes. Glyceraldehyde-3-phosphate dehydrogenase was used as the internal quality control. The reaction mixture was incubated with shrimp alkaline phosphatase for 40 minutes at 37°C and 5 minutes at 85°C to de-phosphorylate the remaining additional dNTPs in the mixture. Extension primers designed to bind a specific sequence of each HPV type were used in the extension reaction, together with mass-modified nucleotides with di-deoxynucleotide terminators. The reaction was performed at 94°C for 30 seconds; 40 cycles at 94°C for 5 seconds, 5 cycles at 52°C for 5 seconds, and 80°C for 5 seconds in each cycle within 40 cycles; and finally 72°C for 3 minutes. The MassArray Nanodispenser RS1000 instrument and MassArray analyzer (Agena Bioscience) were used for sample loading and analysis.
HPV16 Copy Number Determination
HPV16 copy number was determined using a calibration curve (cycle threshold value against the logarithm of the copy number) generated with pHPV16 purified plasmid DNA (catalog number 45113D; ATCC, Manassas, VA). A serial dilution prepared using the plasmid DNA spiked into DNA obtained from the SCC9 (HPV-negative HNC) cell line was used.
Virus copy numbers were determined for HPV16 considering the qPCR cycle threshold values obtained for 50 ng of DNA extracted from saliva samples. Cycle threshold values resulted from the amplification of the HPV16 E6/7 region were considered in the calculation.
Patient Follow-Up Data
OPC patients who were undergoing treatments were clinically followed up for ≤5 years. Follow-up data available up to June 2020 were considered in the study. Event-free survival (EFS) and cause-specific survival (CSS) were used as the primary end points. EFS was defined as the period of time without experiencing an event (recurrence or death) after the completion of the primary treatment. CSS was defined as the time period of survival after the completion of primary treatment without experiencing death specifically due to OPC.
Statistical Analysis
JMP Pro software version 15.2.0 (SAS Institute, Cary, NC) was used for the data analysis. All statistical tests were two-sided, and 0.05 was considered as the level of significance. The χ2 test and the Fisher exact test were used to analyze nominal and ordinal data. Continuous and ratio data were analyzed using the independent-samples t-test and analysis of variance. Kohen κ was used to estimate the inter-rater agreement. Survival data were analyzed using Kaplan-Meier analysis and Cox regression. Median survival times were estimated using parametric survival methods with exponential distribution, as follow-up was insufficient to observe the >50% mortality needed for a Kaplan-Meier or Cox proportional hazards analysis.
Results
Patient Characteristics
Patients with primary HNC (n = 491) (Supplemental Tables S1 and S2) or recurrent HNC (n = 10) (Supplemental Table S3) and who had sufficient DNA in their salivary samples were considered in this study. The recurrent-HNC cohort was considered separately as a pilot cohort to investigate the capacity of salivary HR-HPV DNA as a biomarker for disease monitoring.
Salivary HR-HPV Prevalence in HNC and Associated Factors
In the primary-HNC patient cohort, 43.2% were positive for salivary HR-HPV (Table 1). Among the HR-HPV types detected, the overwhelming majority (92.0%) were HPV16 (Supplemental Table S4). However, only the samples that were HPV16 negative and tumor p16 positive were investigated for other HR-HPV types, and thus co-infections with HPV16 were not investigated.
Table 1.
Salivary HR-HPV Status versus Site of HNC
| HNC site | Salivary HR-HPV status |
|
|---|---|---|
| Positive, n (%) | Negative, n (%) | |
| Oropharynx | 194 (72.4) | 74 (27.6) |
| Oral | 11 (8.2) | 123 (91.8) |
| Unknown primary | 4 (50.0) | 4 (50.0) |
| Larynx | 0 | 47 (100) |
| Other | 3 (8.8) | 31 (91.2) |
| Total | 212 (43.2) | 279 (56.8) |
Salivary HR-HPV–positive HNC patients were younger compared to the HR-HPV–negative HNC patients [mean age, 60.2 versus 62.4 years; t(489) = –2.275, P = 0.023]. The majority of the HNC cohort were men (82.5%), and men also had a higher percentage with positive salivary HR-HPV status compared to women [47.6% versus 21%; χ2(1, n = 491) = 19.467, P < 0.001]. Salivary HR-HPV status was not associated with other major HNC risk factors such as smoking [χ2(1, n = 475) = 0.002, P = 0.962] and alcohol consumption status [χ2(2, n = 461) = 0.700, P = 0.403].
Oropharynx as the Hotspot for HPV-HNC
In the primary-HNC cohort, 91.5% of salivary HR-HPV–positive HNC patients were diagnosed with OPC. Within the OPC cohort, salivary HR-HPV was almost exclusively detected from cancers involving tonsils and the base of the tongue (Supplemental Table S5). Considering the high salivary HR-HPV prevalence in OPC patients, a comprehensive investigation, including a survival analysis, was conducted, with a focus on the OPC cohort (n = 268) (Supplemental Table S6).
Concordance between Salivary HR-HPV and Tumor p16 in OPC Patients
Salivary HR-HPV had a positive predictive value of 99.5% for tumor p16 status. However, the negative predictive value was only 38.0% (Table 2). The Cohen κ value indicated moderate inter-rater agreement between tumor p16 status and salivary HR-HPV DNA positivity (κ = 0.464; 95% CI, 0.34–0.59).
Table 2.
Agreement between Salivary HR-HPV Positivity and p16 Status of OPC
| Tumor p16 Status | Salivary HR-HPV status |
||
|---|---|---|---|
| Positive, n (%) | Negative, n (%) | Total, n (%) | |
| Positive | 193 (81.4) | 44 (18.6) | 237 (100) |
| Negative | 1 (3.57) | 27 (96.4) | 28 (100) |
| Total | 194 (73.20) | 71 (26.79) | 265 (100) |
Salivary HR-HPV and Tumor p16 as Prognostic Biomarkers of OPC
Follow-up data were available from 215 OPC patients. Patients who refused treatment, as well as those who died of other causes, were excluded from the survival analysis. The mean follow-up period was 38 months (range, 6 to 60 months for censored patients).
OPC patient follow-up data revealed that both EFS [χ2(1, n = 211) = 7.072, P = 0.008 (log-rank test)] and CSS [χ2(1, n = 211) = 5.827, P = 0.016 (log-rank test)] were significantly associated with salivary HR-HPV status (Figure 1, A and B). Salivary HR-HPV–positive patients had a clear survival advantage over their salivary HR-HPV–negative counterparts [hazard ratio (HR) (95% CIs): EFS, 0.42 (0.21–0.81, P = 0.010); CSS, 0.39 (0.18–0.86, P = 0.019)]. The estimated median EFSs were 205 months (95% CI, 132–318) in salivary HR-HPV–positive OPC patients and 82 months (95% CI, 50–136) in salivary HR-HPV–negative OPC patients. Similar patterns were observed when estimated median CSS was considered [304 months (95% CI, 180–514) versus 119 months (95% CI, 66–215)].
Figure 1.
Kaplan-Meier estimates of OPC patient survival by salivary HR-HPV and tumor p16. EFS (A, C, and E) and CSS (B, D, and F), by salivary HR-HPV (A and B), tumor p16 (C and D), and salivary HR-HPV and tumor p16 (E and F) status. Survival differences were evaluated using the log-rank test.
The currently recommended prognostic biomarker of OPC, tumor p16, was also able to stratify patients in terms of EFS [χ2(1, n = 208) = 16.410, P < 0.001 (log-rank test)] and CSS [χ2(1, n = 208) = 11.104, P = 0.001 (log-rank test)] (Figure 1, C and D). Tumor p16–positive patients were observed to have a considerably lower risk for an undesirable event or death due to the disease compared to tumor p16–negative patients [HRs (95% CIs): EFS, 0.25 (0.12–0.51, P < 0.001); CSS, 0.25 (0.10–0.60, P = 0.002)]. Similarly, the estimated median (95% CI) EFS [200 months (134–298) versus 48 months (26–90)] and CCS [293 months (182–472) versus 75 months (36–157)] were considerably higher in tumor p16–positive OPC patients. Moreover, American Joint Committee on Cancer (AJCC) staging25 based on tumor p16 status predicted both EFS and CSS in OPC patients [EFS, χ2(3, n = 206) = 52.289, P < 0.001; CSS, χ2(3, n = 206) = 39.040, P < 0.001 (log-rank test)] (Supplemental Figure S2, and Supplemental Table S7).
To further evaluate the associations, a three-group comparison was performed between the salivary HR-HPV–positive and tumor p16–positive group (salivary HR-HPV+/p16+), the salivary HR-HPV–negative and tumor p16–positive group (salivary HR-HPV–/p16+), and the salivary HR-HPV–negative and tumor p16–negative group (salivary HR-HPV–/p16–) (Figure 1, E and F). Only one OPC patient was positive for salivary HR-HPV and negative for tumor p16 (salivary HR-HPV+/p16–) and therefore the survival patterns of salivary HR-HPV+/p16– OPC patients could not be evaluated in the study.
Salivary HR-HPV–/p16– OPC patients had a poor EFS compared to those in salivary HR-HPV+/p16+ OPC patients (HR = 4.16; 95% CI, 1.95–8.90; P < 0.001) and salivary HR-HPV–/p16+ patients (HR = 3.64; 95% CI, 1.14–11.61; P = 0.029). The difference in EFS between the salivary HR-HPV+/p16+ group and the salivary HR-HPV–/p16+ group was not statistically significant (HR = 0.87; 95% CI, 0.30–2.56; P = 0.807). In contrast, when CSS was considered, the difference in survival patterns was statistically significant only between the salivary HR-HPV–/p16– group and the salivary HR-HPV+/p16+ group (HR = 4.14; 95% CI, 1.67–10.27; P = 0.002). CSS was not significantly different between the salivary HR-HPV–/p16– group and the salivary HR-HPV–/p16+ group (HR = 3.28; 95% CI, 0.85–12.71; P = 0.085). The estimated median survival revealed a similar trend (Supplemental Table S8).
HPV16 Viral Load in OPC
Salivary HPV16 copy numbers appeared to be heterogeneous across OPC samples and were not associated with either the AJCC stage25 [F(3,170) = 0.254, P = 0.858] or the T stage [F(3,170) = 0.465, P = 0.707] and N stage [F(3,170) = 0.195, P = 0.900] of the TNM staging system (Supplemental Figure S3). Furthermore, HPV16 copy number was not significantly associated with EFS [χ2(3, n = 141) = 2.066, P = 0.559 (log-rank test)] or CSS [χ2(3, n = 141) = 1.166, P = 0.761 (log-rank test)] in OPC patients (Supplemental Figure S4).
HPV16 Physical Status in OPC
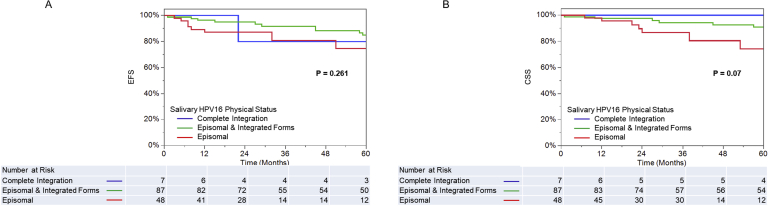
HPV integration into the host genome often results in partial deletion of the HPV genome, including the region that is coding its regulatory protein E2, leading to uncontrolled oncoprotein expression.26,27 As such, the ratio between E2 and E6/7 copy numbers can be used as an indirect indicator of the physical status of HPV.15 Among salivary HPV16–positive OPC patients, 36.5% had only episomal forms, 58.4% had episomal and integrated forms, and 5.1% had only integrated forms (Supplemental Figure S5). Significant associations were not observed between HPV16 physical status and OPC survival [EFS, χ2(2, n = 142) = 2.685, P = 0.261; CSS, χ2(2, n = 142) = 5.308, P = 0.07 (log-rank test)] (Supplemental Figure S6).
Other Factors Associated With OPC Patient Survival
OPC patient survival was also observed to be significantly associated with the cancer site [EFS, χ2(3, n = 210) = 14.204, P = 0.003; CSS, χ2(3, n = 210) = 20.922, P < 0.001 (log-rank test)], where tonsillar-cancer patients had considerably greater survival compared to other OPC patients (Supplemental Figure S7 and Supplemental Table S9). Significant associations were also observed between CSS and patients' age group [≤55, 56 to 65, or >65 years; χ2(2, n = 211) = 6.046, P = 0.049 (log-rank test)]. However, the association between EFS and age categories of OPC patients was not statistically significant [χ2(2, n = 211) = 5.138, P = 0.077 (log-rank test)] (Supplemental Figure S8). Patients who were older than 65 years appeared to have poorer survival compared with those in younger age groups (Supplemental Table S10). Conversely, associations with sex (EFS, χ2(1, n = 211) = 0.003, P = 0.955; CSS, χ2(1, n = 211) = 0.653, P = 0.419 (log-rank test)], smoking [EFS, χ2(1, n = 201) = 0.011, P = 0.916; CSS, χ2(1, n = 201) = 0.185, P = 0.667 (log-rank test)], and alcohol consumption [EFS, χ2(1, n = 196) = 1.914, P = 0.167; CSS, χ2(1, n = 196) = 1.967, P = 0.161 (log-rank test)], did not reach significant levels.
Salivary HR-HPV in Patients with Recurrent HNC
This study also included a cohort of 10 patients with recurrent HNC (Table 3). Salivary HR-HPV testing was performed after the identification of recurrence. Six of these 10 patients (60%) were positive for salivary HR-HPV at the point of recurrence. Five of them previously had known p16-positive primary HNC, while the other patient's p16 status of the primary cancer was not available.
Table 3.
Demographic and Clinical Characteristics of Recurrent HNC Cohort
| Sample no. | Age | Sex | Primary HNC |
Recurrent HNC |
||||
|---|---|---|---|---|---|---|---|---|
| Anatomic site | Tumor p16 status | Anatomic site of recurrent HNC | TNM stage | Tumor p16 | Salivary HR-HPV | |||
| SSR01 | 71 | Male | Tonsil | Unknown | Piriform sinus | T2N2c | Negative | Positive (HPV16) |
| SSR02 | 76 | Male | Tonsil | Positive | Cervical lymph node | T0N2 | Positive | Positive (HPV16) |
| SSR03 | 60 | Male | Tonsil | Positive | Oropharyngeal wall | T1N0 | Positive | Positive (HPV16) |
| SSR04 | 34 | Female | Tonsil | Positive | Cervical lymph nodes | T0N2 | Positive | Positive (HPV16) |
| SSR05 | 78 | Male | BOT | Positive | Cervical lymph node | T0N1 | Not Available | Negative |
| SSR06 | 47 | Male | BOT | Positive | Supraglottis | T1N0 | Positive | Positive (HPV16) |
| SSR07 | 55 | Male | BOT | Positive | Oral tongue and floor of the mouth | T2N0 | Positive | Positive (HPV16) |
| SSR08 | 63 | Male | Pharyngeal wall | Negative | Oropharyngeal wall | T1N0 | Negative | Negative |
| SSR09 | 44 | Male | Glottis | Unknown | Glottis | T1N0 | Positive | Negative |
| SSR10 | 61 | Male | Upper lip | Unknown | Cervical lymph node | T0N2 | Positive | Negative |
BOT, base of the tongue; TNM, tumor-node-metastasis cancer staging.
Discussion
Similar to current trends in other developed countries, this study reports that HR-HPV types are associated with a considerable number of HNC cases (43.2% in the studied HNC patient cohort, based on the presence of salivary HR-HPV DNA).28,29 HPV16 was detected in 92% of HPV-positive saliva samples, followed by other HR-HPV types, such as 33, 18, and 35. Salivary HR-HPV–positive HNC patients were observed to be comparatively younger, as reported in previous studies.7,30 Even though male preponderance was observed in the overall sample, the male sex had a higher risk for HPV-HNC. Salivary HR-HPV status was not observed to have been associated with other major HNC risk factors, such as smoking and alcohol consumption.
Consistent with other studies, the vast majority of HPV-HNC had arisen from the oropharynx, especially from palatine tonsils and the base of the tongue.3,6 A total of 72.4% of OPC patients were positive for HR-HPV DNA in their saliva, and tumor p16 overexpression was observed in 89.3%. The present study could not detect salivary HR-HPV in 18.6% of the tumor p16–positive OPC patients. Even though p16 is often overexpressed in HPV-driven cancers, it is not a unique marker for these cancers.31, 32, 33, 34 Previous studies have reported that in 5% to 20% of tumor p16–positive OPCs, the tumor is not positive for HPV.15,35, 36, 37 As such, it can be concluded that salivary HR-HPV can be detected in the vast majority of HPV-HNC patients, at the point of diagnosis, indicating its possible utility as a biomarker for facilitating early detection and screening of HPV-HNC. Salivary HR-HPV DNA also appears to have value as a surveillance biomarker. Even though the sample of patients with recurrent HNC considered in the study was quite small, findings indicate that salivary HR-HPV tends to be positive in the majority of the HPV-HNC patients at the point of locoregional recurrence, indicating its possible utility as a surveillance biomarker.
Furthermore, the current study confirms the utility of tumor p16 as a prognostic biomarker of OPC, further supporting its use in current clinical practice.3,38 Moreover, AJCC staging25 based on tumor p16 expression and TNM staging was representative of the survival patterns in OPC patients. However, this study also highlights the capacity of salivary HR-HPV as a robust OPC prognosticator. Consistent with previous studies, salivary HR-HPV+/p16+ OPC patients had better survival compared to salivary HR-HPV–/p16– OPC patients.36,39 Even though survival patterns in HR-HPV–/p16+ OPC patients have been controversial and contradictory findings have been reported previously, the present study did not observe a statistically significant difference in survival between salivary HR-HPV+/p16+ OPC patients and salivary HR-HPV–/p16+ OPC patients.36,39, 40, 41 However, only EFS but not CSS in salivary HR-HPV–/p16+ OPC patients was significantly different from that in salivary HR-HPV–/p16– OPC patients. As such, it can be speculated that additional salivary HR-HPV testing can add value in OPC prognostication.
This study also evaluated variations in salivary HPV16 copy number and physical status in HPV-OPC patients. Salivary HPV16 copy numbers appeared to be markedly heterogeneous among the samples, and no association with the stage of the cancer was observed. Available literature indicates that HPV copies per cell vary considerably among HNC patients.42,43 Hence, salivary HPV copy number across patients is not just a function involving tumor size or spread of the tumor. It can be reasoned that HPV copy number is possibly representative of the tumor growth when individual patients are considered. However, longitudinal follow-up studies are necessary to establish these associations. HPV16 physical status was evaluated considering the ratio between HPV E6/7 and E2 copy numbers. Confirming that HPV16 DNA integration is common in OPC, integration was observed in 63.5% of OPC patients, 5.1% of whom had exclusively integrated forms.44,45 However, statistically significant associations between these factors and OPC patient survival were not identified despite previous observations.46,47
Our findings also indicate that there are apparent dissimilarities between cancers originating from different oropharyngeal subsites, even though they are collectively evaluated as related oncologic entities. Our findings indicate that tonsillar-cancer patients had considerably greater survival compared to patients with cancers originating in other oropharyngeal sites. Similar observations were also reported previously.48 Furthermore, OPC survival was observed to be linked with patients' age, where the patient group aged >65 years had poor survival compared to younger age groups. Despite previous reports, smoking and alcohol consumption were not observed to have been associated with OPC patient survival.49, 50, 51 As such, the effects of these risk behaviors may possibly be synergistic, rather than mutually exclusive, in the carcinogenic process of OPC, as suggested previously.52 However, the present study used only a binomial categorization for risk behaviors (ever or never). The extent of the risk behaviors, which would have been more representative, was not evaluated.
Difficulty in locating early cancerous lesions is a major drawback of HPV-HNC detection. Consequently, a minimal number of HPV-HNC patients are diagnosed in the early stages. This study demonstrates that salivary HR-HPV DNA can be detected in the vast majority of HPV-HNC patients at diagnosis. Salivary HR-HPV positivity at the very early stages of HNC has also been proven in two recent studies in which longitudinal salivary HPV testing was coupled with clinical investigations to detect HNC in asymptomatic individuals.13,14 These studies, together with the present findings, highlight the possible utility of screening programs based on longitudinal salivary HR-HPV DNA testing for the early detection of HPV-HNC. Furthermore, the present findings indicate the potential of salivary HR-HPV DNA testing for HNC patient monitoring. When the noninvasive nature and convenience of the collection are considered, salivary HR-HPV testing can be considered as an ideal mode of screening asymptomatic individuals and of long-term monitoring in HPV-driven HNC patients. Moreover, this study emphasizes the capacity of salivary HR-HPV DNA as a prognostic biomarker of OPC.
Strengths and Limitations
The large-scale sample size and a follow-up period of up to 5 years were the strengths of this study. However, tumor samples were characterized for HPV status only by p16 IHC analysis. As such, a direct comparison between tumor HPV status and salivary HPV status was not possible.
Acknowledgments
We thank Trang Le, Jennifer Edmunds, Charmaine Micklewright, Jacqui Keller, and Dana Middleton for assistance with patient recruitment; and Dr. Darryl Irwin and Vandhana Bharti (Agena Bioscience, St. Lucia, Australia) for contributing to HPV genotyping.
Footnotes
Supported by Cancer Australia grant APP1145657 (C.P.), the Royal Brisbane and Women's Hospital Foundation (C.P.), the Garnett Passé and Rodney Williams Foundation (C.P.), a Sri Lanka University Grants Commission scholarship (C.E.W.), and Queensland University of Technology (C.E.W.).
Disclosures: None declared.
Supplemental material for this article can be found at https://doi.org/10.1016/j.jmoldx.2021.07.005.
Contributor Information
Kai D. Tang, Email: kai.tang@qut.edu.au.
Chamindie Punyadeera, Email: chamindie.punyadeera@qut.edu.au.
Author Contributions
K.D.T. and C.P. conceptualized and designed the study; C.E.W., Z.L., and Y.L. performed the experiments; S.V., J.L.-L., L.K., L.M., I.F., and K.D.T. validated the data; G.H. and C.E.W. analyzed the data; C.E.W. and G.H. drafted the manuscript; S.V., J.L.-L., L.K., L.M., I.F., K.D.T., and C.P. reviewed and edited the manuscript and supervised the project; and C.P. secured funding for the project.
Supplemental Data
Supplemental Figure S1.
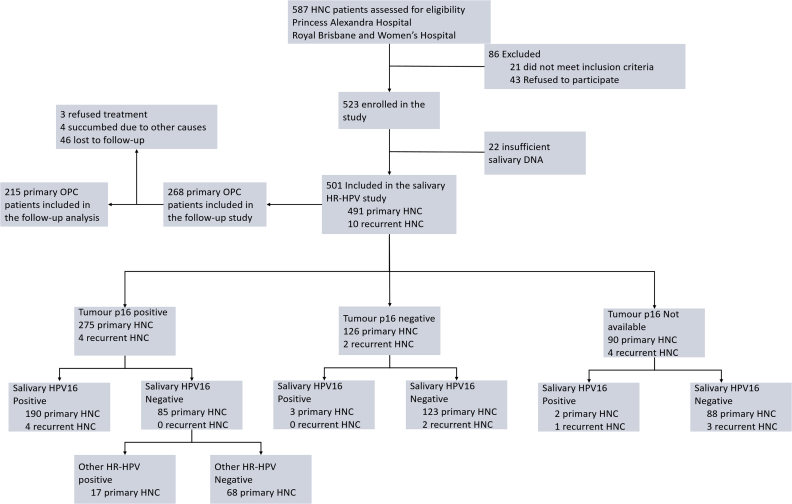
Study flow chart. A total of 523 HNC patients were recruited for the study from Princess Alexandra Hospital and Royal Brisbane and Women's Hospital (Brisbane, QLD, Australia). After the exclusion of 22 patients due to unsatisfactory salivary DNA yield, 491 primary-HNC patients and 10 recurrent-HNC patients were considered in the study. Tumor p16 expression was evaluated as a part of the routine clinical investigations. Saliva samples were collected before the initiation of the treatment. Considering that HPV16 is the HPV type that most commonly causes HPV-HNC, all of the salivary samples were initially tested for HPV16 DNA. Samples that were salivary HPV16 DNA negative and p16 positive were tested for other HR-HPV types. The OPC cohort (n = 268) considered in the study was followed up for up to 5 years and 215 OPC patients were considered in the survival analysis after the exclusion of patients who refused the treatment, died from other causes, and/or were lost to follow-up.
Supplemental Figure S2.
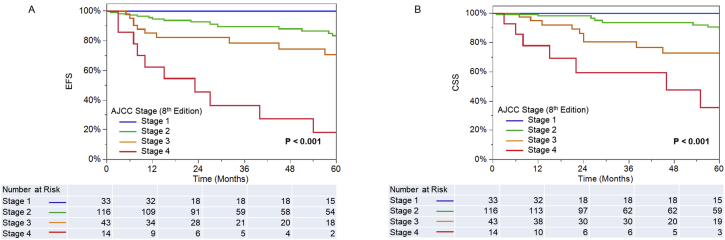
Kaplan-Meier estimates of OPC patient survival, by American Joint Committee on Cancer (AJCC) stage.25 EFS (A) and CSS (B), by AJCC stage. Survival differences were evaluated using the log-rank test.
Supplemental Figure S3.
Associations between salivary HPV16 copy number and T stage (A), N stage of the tumor-node-metastasis (TNM) staging system (B), and American Joint Committee on Cancer (AJCC) stage (C).25
Supplemental Figure S4.
Kaplan-Meier estimates of OPC patient survival, by salivary HPV16 copy number. EFS (A) and CSS (B), by salivary HPV16 copy number. Survival differences were evaluated using the log-rank test.
Supplemental Figure S5.
Salivary HPV16 physical status by OPC site. BOT, base of the tongue.
Supplemental Figure S6.
Kaplan-Meier estimate of OPC patient survival, by salivary HPV16 physical status. EFS (A) and CSS (B), by salivary HPV16 physical status. Survival differences were evaluated using the log-rank test.
Supplemental Figure S7.
Salivary HR-HPV status and tumor p16 status, by OPC site. Salivary HR-HPV status (A) and tumor p16 status (B), by OPC site. BOT, base of the tongue.
Supplemental Figure S8.
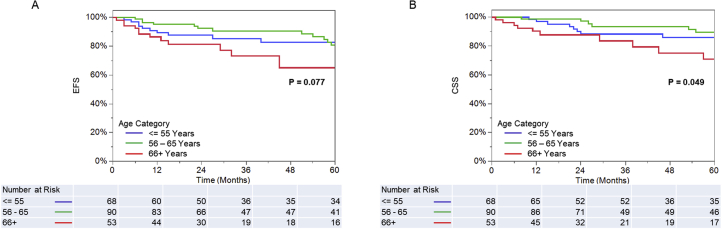
Kaplan-Meier estimates of OPC patient survival, by age category. EFS (A) and CSS (B), by age category. Survival differences were evaluated using the log-rank test.
References
- 1.Zapatka M., Borozan I., Brewer D.S., Iskar M., Grundhoff A., Alawi M., Desai N., Sültmann H., Moch H., Alawi M., Cooper C.S., Eils R., Ferretti V., Lichter P., Borozan I., Brewer D.S., Cooper C.S., Desai N., Eils R., Ferretti V., Grundhoff A., Iskar M., Kleinheinz K., Lichter P., Nakagawa H., Ojesina A.I., Pedamallu C.S., Schlesner M., Su X., Zapatka M., Pathogens P., PCAWG Consortium The landscape of viral associations in human cancers. Nat Genet. 2020;52:320–330. doi: 10.1038/s41588-019-0558-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Chai R.C., Lambie D., Verma M., Punyadeera C. Current trends in the etiology and diagnosis of HPV-related head and neck cancers. Cancer Med. 2015;4:596–607. doi: 10.1002/cam4.424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chow L.Q.M. Head and neck cancer. N Engl J Med. 2020;382:60–72. doi: 10.1056/NEJMra1715715. [DOI] [PubMed] [Google Scholar]
- 4.Schroeder L., Boscolo-Rizzo P., Dal Cin E., Romeo S., Baboci L., Dyckhoff G., Hess J., Lucena-Porcel C., Byl A., Becker N., Alemany L., Castellsagué X., Quer M., León X., Wiesenfarth M., Pawlita M., Holzinger D. Human papillomavirus as prognostic marker with rising prevalence in neck squamous cell carcinoma of unknown primary: a retrospective multicentre study. Eur J Cancer. 2017;74:73–81. doi: 10.1016/j.ejca.2016.12.020. [DOI] [PubMed] [Google Scholar]
- 5.Bussu F., Sali M., Gallus R., Petrone G., Autorino R., Santangelo R., Pandolfini M., Miccichè F., Delogu G., Almadori G., Galli J., Sanguinetti M., Rindi G., Tommasino M., Valentini V., Paludetti G. HPV and EBV infections in neck metastases from occult primary squamous cell carcinoma: another virus-related neoplastic disease in the head and neck region. Ann Surg Oncol. 2015;22:979–984. doi: 10.1245/s10434-015-4808-5. [DOI] [PubMed] [Google Scholar]
- 6.Cramer J.D., Burtness B., Le Q.T., Ferris R.L. The changing therapeutic landscape of head and neck cancer. Nat Rev Clin Oncol. 2019;16:669–683. doi: 10.1038/s41571-019-0227-z. [DOI] [PubMed] [Google Scholar]
- 7.Berman T.A., Schiller J.T. Human papillomavirus in cervical cancer and oropharyngeal cancer: one cause, two diseases. Cancer. 2017;123:2219–2229. doi: 10.1002/cncr.30588. [DOI] [PubMed] [Google Scholar]
- 8.Kreimer A.R., Shiels M.S., Fakhry C., Johansson M., Pawlita M., Brennan P., Hildesheim A., Waterboer T. Screening for human papillomavirus-driven oropharyngeal cancer: considerations for feasibility and strategies for research. Cancer. 2018;124:1859–1866. doi: 10.1002/cncr.31256. [DOI] [PubMed] [Google Scholar]
- 9.Rautava J., Willberg J., Louvanto K., Wideman L., Syrjanen K., Grenman S., Syrjanen S. Prevalence, genotype distribution and persistence of human papillomavirus in oral mucosa of women: a six-year follow-up study. PLoS One. 2012;7:e42171. doi: 10.1371/journal.pone.0042171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Beachler D.C., Lang Kuhs K.A., Struijk L., Schussler J., Herrero R., Porras C., Hildesheim A., Cortes B., Sampson J., Quint W., Gonzalez P., Kreimer A.R. The natural history of oral human papillomavirus in young Costa Rican women. Sex Transm Dis. 2017;44:442–449. doi: 10.1097/OLQ.0000000000000625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Beachler D.C., Sugar E.A., Margolick J.B., Weber K.M., Strickler H.D., Wiley D.J., Cranston R.D., Burk R.D., Minkoff H., Reddy S., Xiao W., Guo Y., Gillison M.L., D'Souza G. Risk factors for acquisition and clearance of oral human papillomavirus infection among HIV-infected and HIV-uninfected adults. Am J Epidemiol. 2014;181:40–53. doi: 10.1093/aje/kwu247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhang C., Liu F., Pan Y., Deng Q., Li X., He Z., Liu M., Ning T., Guo C., Liang Y., Xu R., Zhang L., Cai H., Ke Y. Incidence and clearance of oral human papillomavirus infection: a population-based cohort study in rural China. Oncotarget. 2017;8:59831–59844. doi: 10.18632/oncotarget.16306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tang K.D., Vasani S., Taheri T., Walsh L.J., Hughes B.G.M., Kenny L., Punyadeera C. An occult HPV-driven oropharyngeal squamous cell carcinoma discovered through a saliva test. Front Oncol. 2020;10:408. doi: 10.3389/fonc.2020.00408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.D'Souza G., Clemens G., Strickler H.D., Wiley D.J., Troy T., Struijk L., Gillison M., Fakhry C. Long-term persistence of oral HPV over 7 years of follow-up. JNCI Cancer Spectr. 2020;4:pkaa047. doi: 10.1093/jncics/pkaa047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tang K.D., Baeten K., Kenny L., Frazer I.H., Scheper G., Punyadeera C. Unlocking the potential of saliva-based test to detect HPV-16-driven oropharyngeal cancer. Cancers (Basel) 2019;11:473. doi: 10.3390/cancers11040473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Qureishi A., Ali M., Fraser L., Shah K.A., Moller H., Winter S. Saliva testing for human papilloma virus in oropharyngeal squamous cell carcinoma: a diagnostic accuracy study. Clin Otolaryngol. 2018;43:151–157. doi: 10.1111/coa.12917. [DOI] [PubMed] [Google Scholar]
- 17.Wasserman J.K., Rourke R., Purgina B., Caulley L., Dimitroulakis J., Corsten M., Johnson-Obaseki S. HPV DNA in saliva from patients with SCC of the head and neck is specific for p16-positive oropharyngeal tumours. J Otolaryngol Head Neck Surg. 2017;46:3. doi: 10.1186/s40463-016-0179-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chai R.C., Lim Y., Frazer I.H., Wan Y., Perry C., Jones L., Lambie D., Punyadeera C. A pilot study to compare the detection of HPV-16 biomarkers in salivary oral rinses with tumour p16(INK4a) expression in head and neck squamous cell carcinoma patients. BMC Cancer. 2016;16:178. doi: 10.1186/s12885-016-2217-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Rettig E.M., Wentz A., Posner M.R., Gross N.D., Haddad R.I., Gillison M.L., Fakhry C., Quon H., Sikora A.G., Stott W.J., Lorch J.H., Gourin C.G., Guo Y., Xiao W., Miles B.A., Richmon J.D., Andersen P.E., Misiukiewicz K.J., Chung C.H., Gerber J.E., Rajan S.D., D'Souza G. Prognostic implication of persistent human papillomavirus type 16 DNA detection in oral rinses for human papillomavirus-related oropharyngeal carcinoma. JAMA Oncol. 2015;1:907–915. doi: 10.1001/jamaoncol.2015.2524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Koslabova E., Hamsikova E., Salakova M., Klozar J., Foltynova E., Salkova E., Rotnaglova E., Ludvikova V., Tachezy R. Markers of HPV infection and survival in patients with head and neck tumors. Int J Cancer. 2013;133:1832–1839. doi: 10.1002/ijc.28194. [DOI] [PubMed] [Google Scholar]
- 21.Tang K.D., Kenny L., Frazer I.H., Punyadeera C. High-risk human papillomavirus detection in oropharyngeal cancers: comparison of saliva sampling methods. Head Neck. 2019;41:1484–1489. doi: 10.1002/hed.25578. [DOI] [PubMed] [Google Scholar]
- 22.Sun C.X., Bennett N., Tran P., Tang K.D., Lim Y., Frazer I., Samaranayake L., Punyadeera C. A pilot study into the association between oral health status and human papillomavirus-16 infection. Diagnostics (Basel) 2017;7:11. doi: 10.3390/diagnostics7010011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Peitsaro P., Johansson B., Syrjänen S. Integrated human papillomavirus type 16 is frequently found in cervical cancer precursors as demonstrated by a novel quantitative real-time PCR technique. J Clin Microbiol. 2002;40:886–891. doi: 10.1128/JCM.40.3.886-891.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kriegsmann M., Wandernoth P., Lisenko K., Casadonte R., Longuespée R., Arens N., Kriegsmann J. Detection of HPV subtypes by mass spectrometry in FFPE tissue specimens: a reliable tool for routine diagnostics. J Clin Pathol. 2017;70:417–423. doi: 10.1136/jclinpath-2016-204017. [DOI] [PubMed] [Google Scholar]
- 25.Amin M.B., Greene F.L., Edge S.B., Compton C.C., Gershenwald J.E., Brookland R.K., Meyer L., Gress D.M., Byrd D.R., Winchester D.P. The Eighth Edition AJCC Cancer Staging Manual: continuing to build a bridge from a population-based to a more "personalized" approach to cancer staging. CA Cancer J Clin. 2017;67:93–99. doi: 10.3322/caac.21388. [DOI] [PubMed] [Google Scholar]
- 26.Woodman C.B., Collins S.I., Young L.S. The natural history of cervical HPV infection: unresolved issues. Nat Rev Cancer. 2007;7:11–22. doi: 10.1038/nrc2050. [DOI] [PubMed] [Google Scholar]
- 27.Gammoh N., Isaacson E., Tomaic V., Jackson D.J., Doorbar J., Banks L. Inhibition of HPV-16 E7 oncogenic activity by HPV-16 E2. Oncogene. 2009;28:2299–2304. doi: 10.1038/onc.2009.78. [DOI] [PubMed] [Google Scholar]
- 28.de Martel C., Plummer M., Vignat J., Franceschi S. Worldwide burden of cancer attributable to HPV by site, country and HPV type. Int J Cancer. 2017;141:664–670. doi: 10.1002/ijc.30716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gillison M.L., Chaturvedi A.K., Anderson W.F., Fakhry C. Epidemiology of human papillomavirus-positive head and neck squamous cell carcinoma. J Clin Oncol. 2015;33:3235–3242. doi: 10.1200/JCO.2015.61.6995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Castellsagué X., Alemany L., Quer M., Halec G., Quirós B., Tous S., et al. HPV involvement in head and neck cancers: comprehensive assessment of biomarkers in 3680 patients. J Natl Cancer Inst. 2016;108:djv403. doi: 10.1093/jnci/djv403. [DOI] [PubMed] [Google Scholar]
- 31.Klingenberg B., Hafkamp H.C., Haesevoets A., Manni J.J., Slootweg P.J., Weissenborn S.J., Klussmann J.P., Speel E.-J.M. p16INK4A overexpression is frequently detected in tumour-free tonsil tissue without association with HPV. Histopathology. 2010;56:957–967. doi: 10.1111/j.1365-2559.2010.03576.x. [DOI] [PubMed] [Google Scholar]
- 32.Gould V.E., Schmitt M., Vinokurova S., Reddy V.B., Bitterman P., Alonso A., Gattuso P. Human papillomavirus and p16 expression in inverted papillomas of the urinary bladder. Cancer Lett. 2010;292:171–175. doi: 10.1016/j.canlet.2009.11.022. [DOI] [PubMed] [Google Scholar]
- 33.Shi S.s., Wang X., Xia Q.y., Rao Q., Shen Q., Ye S.b., Li R., Shi Q.l., Lu Z.f., Ma H.h., Zhou X.j. P16 overexpression in BRAF-mutated gastrointestinal stromal tumors. Expert Rev Mol Diagn. 2017;17:195–201. doi: 10.1080/14737159.2017.1272413. [DOI] [PubMed] [Google Scholar]
- 34.Prigge E.S., Arbyn M., von Knebel Doeberitz M., Reuschenbach M. Diagnostic accuracy of p16INK4a immunohistochemistry in oropharyngeal squamous cell carcinomas: a systematic review and meta-analysis. Int J Cancer. 2017;140:1186–1198. doi: 10.1002/ijc.30516. [DOI] [PubMed] [Google Scholar]
- 35.Grønhøj Larsen C., Gyldenløve M., Jensen D.H., Therkildsen M.H., Kiss K., Norrild B., Konge L., von Buchwald C. Correlation between human papillomavirus and p16 overexpression in oropharyngeal tumours: a systematic review. Br J Cancer. 2014;110:1587–1594. doi: 10.1038/bjc.2014.42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Nauta I.H., Rietbergen M.M., van Bokhoven A., Bloemena E., Lissenberg-Witte B.I., Heideman D.A.M., Baatenburg de Jong R.J., Brakenhoff R.H., Leemans C.R. Evaluation of the eighth TNM classification on p16-positive oropharyngeal squamous cell carcinomas in The Netherlands and the importance of additional HPV DNA testing. Ann Oncol. 2018;29:1273–1279. doi: 10.1093/annonc/mdy060. [DOI] [PubMed] [Google Scholar]
- 37.Rietbergen M.M., Snijders P.J.F., Beekzada D., Braakhuis B.J.M., Brink A., Heideman D.A.M., Hesselink A.T., Witte B.I., Bloemena E., Baatenburg-De Jong R.J., René Leemans C., Brakenhoff R.H. Molecular characterization of p16-immunopositive but HPV DNA-negative oropharyngeal carcinomas. Int J Cancer. 2014;134:2366–2372. doi: 10.1002/ijc.28580. [DOI] [PubMed] [Google Scholar]
- 38.Lydiatt W.M., Patel S.G., O'Sullivan B., Brandwein M.S., Ridge J.A., Migliacci J.C., Loomis A.M., Shah J.P. Head and neck cancers-major changes in the American Joint Committee on cancer eighth edition cancer staging manual. CA Cancer J Clin. 2017;67:122–137. doi: 10.3322/caac.21389. [DOI] [PubMed] [Google Scholar]
- 39.Craig S.G., Anderson L.A., Schache A.G., Moran M., Graham L., Currie K., Rooney K., Robinson M., Upile N.S., Brooker R., Mesri M., Bingham V., McQuaid S., Jones T., McCance D.J., Salto-Tellez M., McDade S.S., James J.A. Recommendations for determining HPV status in patients with oropharyngeal cancers under TNM8 guidelines: a two-tier approach. Br J Cancer. 2019;120:827–833. doi: 10.1038/s41416-019-0414-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Coordes A., Lenz K., Qian X., Lenarz M., Kaufmann A.M., Albers A.E. Meta-analysis of survival in patients with HNSCC discriminates risk depending on combined HPV and p16 status. Eur Arch Oto Rhino Laryngol. 2016;273:2157–2169. doi: 10.1007/s00405-015-3728-0. [DOI] [PubMed] [Google Scholar]
- 41.Lewis J.S., Jr., Thorstad W.L., Chernock R.D., Haughey B.H., Yip J.H., Zhang Q., El-Mofty S.K. p16 positive oropharyngeal squamous cell carcinoma: an entity with a favorable prognosis regardless of tumor HPV status. Am J Surg Pathol. 2010;34:1088–1096. doi: 10.1097/PAS.0b013e3181e84652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Conway C., Chalkley R., High A., Maclennan K., Berri S., Chengot P., Alsop M., Egan P., Morgan J., Taylor G.R., Chester J., Sen M., Rabbitts P., Wood H.M. Next-generation sequencing for simultaneous determination of human papillomavirus load, subtype, and associated genomic copy number changes in tumors. J Mol Diagn. 2012;14:104–111. doi: 10.1016/j.jmoldx.2011.10.003. [DOI] [PubMed] [Google Scholar]
- 43.Nulton T.J., Olex A.L., Dozmorov M., Morgan I.M., Windle B. Analysis of the Cancer Genome Atlas sequencing data reveals novel properties of the human papillomavirus 16 genome in head and neck squamous cell carcinoma. Oncotarget. 2017;8:17684–17699. doi: 10.18632/oncotarget.15179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Cancer Genome Atlas Network Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015;517:576–582. doi: 10.1038/nature14129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Peter M., Rosty C., Couturier J., Radvanyi F., Teshima H., Sastre-Garau X. MYC activation associated with the integration of HPV DNA at the MYC locus in genital tumors. Oncogene. 2006;25:5985–5993. doi: 10.1038/sj.onc.1209625. [DOI] [PubMed] [Google Scholar]
- 46.Worden F.P., Kumar B., Lee J.S., Wolf G.T., Cordell K.G., Taylor J.M.G., Urba S.G., Eisbruch A., Teknos T.N., Chepeha D.B., Prince M.E., Tsien C.I., D'Silva N.J., Yang K., Kurnit D.M., Mason H.L., Miller T.H., Wallace N.E., Bradford C.R., Carey T.E. Chemoselection as a strategy for organ preservation in advanced oropharynx cancer: response and survival positively associated with HPV16 copy number. J Clin Oncol. 2008;26:3138–3146. doi: 10.1200/JCO.2007.12.7597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Koneva L.A., Zhang Y., Virani S., Hall P.B., McHugh J.B., Chepeha D.B., Wolf G.T., Carey T.E., Rozek L.S., Sartor M.A. HPV integration in HNSCC correlates with survival outcomes, immune response signatures, and candidate drivers. Mol Cancer Res. 2018;16:90–102. doi: 10.1158/1541-7786.MCR-17-0153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Mäkitie A.A., Pukkila M., Laranne J., Pulkkinen J., Vuola J., Bäck L., Koivunen P., Grénman R. Oropharyngeal carcinoma and its treatment in Finland between 1995–1999: a nationwide study. Eur Arch Oto Rhino Laryngol Head Neck. 2006;263:139–143. doi: 10.1007/s00405-005-0975-5. [DOI] [PubMed] [Google Scholar]
- 49.Gillison M.L., Zhang Q., Jordan R., Xiao W., Westra W.H., Trotti A., Spencer S., Harris J., Chung C.H., Ang K.K. Tobacco smoking and increased risk of death and progression for patients with p16-positive and p16-negative oropharyngeal cancer. J Clin Oncol. 2012;30:2102–2111. doi: 10.1200/JCO.2011.38.4099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kumar B., Cordell K.G., Lee J.S., Prince M.E., Tran H.H., Wolf G.T., Urba S.G., Worden F.P., Chepeha D.B., Teknos T.N., Eisbruch A., Tsien C.I., Taylor J.M.G., D'Silva N.J., Yang K., Kurnit D.M., Bradford C.R., Carey T.E. Response to therapy and outcomes in oropharyngeal cancer are associated with biomarkers including human papillomavirus, epidermal growth factor receptor, gender, and smoking. Int J Radiat Oncol Biol Phys. 2007;69:S109–S111. doi: 10.1016/j.ijrobp.2007.05.072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Lassen P., Lacas B., Pignon J.-P., Trotti A., Zackrisson B., Zhang Q., Overgaard J., Blanchard P. Prognostic impact of HPV-associated p16-expression and smoking status on outcomes following radiotherapy for oropharyngeal cancer: the MARCH-HPV project. Radiother Oncol. 2018;126:107–115. doi: 10.1016/j.radonc.2017.10.018. [DOI] [PubMed] [Google Scholar]
- 52.Anantharaman D., Muller D.C., Lagiou P., Ahrens W., Holcátová I., Merletti F., et al. Combined effects of smoking and HPV16 in oropharyngeal cancer. Int J Epidemiol. 2016;45:752–761. doi: 10.1093/ije/dyw069. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.