Abstract
We have constructed strains that allow a direct selection for mutators of Escherichia coli on a single plate medium. The plate selection is based on using two different markers whose reversion is enhanced by a given mutator. Plates containing limiting amounts of each respective nutrient allow the growth of ghost colonies or microcolonies that give rise to full-size colonies only if a reversion event occurs. Because two successive mutational events are required, mutator cells are favored to generate full-size colonies. Reversion of a third marker allows direct visualization of the mutator phenotype by the large number of blue papillae in the full-size colonies. We also describe plate selections involving three successive nutrient markers followed by a fourth papillation step. Different frameshift or base substitution mutations are used to select for mismatch-repair-defective strains (mutHLS and uvrD). We can detect and monitor mutator cells arising spontaneously, at frequencies lower than 10−5 in the population. Also, we can measure a mutator cascade, in which one type of mutator (mutT) generates a second mutator (mutHLS) that then allows stepwise frameshift mutations. We discuss the relevance of mutators arising on a single medium as a result of cells overcoming successive growth barriers to the development and progression of cancerous tumors, some of which are mutator cell lines.
Several successive genetic alterations are required for a cancer cell to develop, proliferate, and be able to metastasize (70). In fact, as many as 6 to 10 mutations may be needed to turn a normal cell into a full-blown invasive tumor line (2), which is one reason why many cancers take decades to develop. Loeb (35) has postulated that an early step in the progression of some cancers may be the creation of a mutator cell, since the low spontaneous mutation rate might not account for the incidence of cancer. However, the elevated mutation rates in mutators would greatly increase the probability of cells accumulating all of the necessary mutations (27, 35), especially if repetitive rounds of clonal expansion and somatic selection occurred (48). In accord with this idea was the exciting finding that the inherited susceptibility to human nonpolyposis colon cancer (HNPCC) and ovarian cancer is due to a defect in one copy of one of the genes involved in the human counterpart to the bacterial mismatch repair system (3, 18, 30, 49). Presumably, when a somatic cell loses the other copy, the resulting cell is completely defective for mismatch repair and has a higher mutation rate. In fact, tumor lines from HNPCC patients are mutators with greatly increased repeat-tract or microsatellite instability (1, 26, 30, 36, 52, 53, 69), a propensity for frequent additions or deletions at repetitive nucleotide sequences (such as repetitive mono-, di-, tri-, and tetranucleotide repeats), in analogy with the repeat-tract instability seen in mismatch-repair-deficient strains of bacteria and yeast (11, 57, 66). These tumor lines also show elevated mutation rates in genes such as hprt (1, 3, 21). The realization that mutator cells are cancer prone (1, 30, 50, 51) has led to increased research in this area, including a search for new types of mutators and for a better understanding of how mutators proliferate.
Experiments with bacteria aimed at finding mutators (12, 23, 24, 34) or at examining the behavior of wild-type or mixed populations in chemostats (6, 9, 20, 46, 61, 65) showed that continuous selection increases the proportion of mutators in cell populations. In a previous paper, we demonstrated how easily a population of Escherichia coli cells could become principally or totally mutators in response to several successive selections (39). Here, we ask whether instead of changing medium in successive selections a single medium that selects directly for mutator colonies could be devised. By asking cells to overcome several barriers to growth on a medium with limiting amounts of each required nutrient, we developed a plate medium on which only mutator cells grow. We have employed this medium to examine the occurrence of mutators under several conditions, and we discuss how this mimics the situation in mammalian cells that need to overcome several growth restrictions before becoming proliferating cancer cells. We also show that one mutator can induce a second mutator that then stimulates useful genetic changes, as part of a mutator cascade.
MATERIALS AND METHODS
Bacterial strains and plasmids.
The strain CC107 carries an F′lacpro episome in the P90C (11, 43) strain background ara Δ(gpt-lac)5. The lac region on the F factor carries a lacI mutation and also a frameshift in the lacZ gene that reverts by the addition of a GC base pair to a monotonous run of GC base pairs. This strain is described by Cupples et al. (11). AS18 is a Met− derivative of CC107 carrying an ICR-191-induced frameshift mutation in the metE (or possibly metR) gene. AS18-29 is a Bgl− derivative of AS18 carrying an additional ICR-191-induced frameshift mutation, this time in the blgA gene. AS210 is a Leu− derivative of AS18-29, carrying a mutH-induced frameshift mutation at the leu locus and a Tn10kan insert near the wild-type mutH gene (zgh-3159 [62]). We constructed specific mutator derivatives of certain strains by using P1 transduction from strains in which a mini-Tn10 had integrated into either the mutT, mutH, mutS, mutL, or uvrD gene (45).
Genetic methods.
Mapping experiments were carried out with P1 cotransduction with Tn10 transposons that had integrated near either the mutH, mutL, mutS, or uvrD gene, or near various nutritional markers (62). Auxotrophs were detected after ICR-191 mutagenesis by replicating Luria-Bertani (LB) plates spread with 100 to 300 colonies onto minimal medium plates and recognizing those colonies which failed to grow. Combinations of supplements were used to restore growth, and then individual supplements were employed. All other strains and bacterial genetic methods, such as determination of rifampin resistance (Rifr), are described by Miller (43). We initially examined colonies for strong mutator activity by gridding them onto an LB plate and growing them overnight at 32°C before replicating them onto a second LB plate. The second plate was grown overnight, and then colonies were replicated onto an LB plate with 100 μg of rifampin per ml. After overnight growth, mutators defective in mismatch repair showed many Rifr colonies growing out of the replicated patch, whereas nonmutator strains did not. More quantitative measurements were then carried out when relevant.
Preparation of cultures.
Unless otherwise stated, all cultures for the experiments reported here were prepared by inoculating a portion of a single colony into LB medium (43) or other medium and growing it overnight. A different single colony was used for each culture. Therefore, all mutants occurring in different cultures are of independent origin, since each single colony is derived from a single isolated cell. Typically, 5-ml cultures were used.
Mutagenesis.
All methods of mutagenesis were exactly as described in the work of Miller (43). 2-Aminopurine (2AP; Sigma) was used at concentrations of 700 μg/ml. Cultures were prepared by subculturing 104 to 105 cells into 4 ml of LB with 2AP, and these were grown for 12 to 16 generations in LB with 2AP before plating. ICR-191 (Sigma) was used at concentrations of 10 μg/ml in minimal A medium (43) supplemented with 1 ml of LB broth, 2 ml of 20% glucose, 0.1 ml of 1 M MgSO4, and 0.5 ml of B1 (thiamine hydrochloride) per 100 ml of medium.
Specialized media.
Mutators were selected on lactose minimal A medium (43) supplemented with limiting amounts of required sugars or nutrients. We used 150 μg of glucose per ml as a limiting carbon source and 0.5 μg of required amino acids, purines, or pyrimidines per ml. For instance, in conjunction with strain AS18 or AS18-29 we used minimal A plates containing lactose, B1, and MgSO4 (33), supplemented with 150 μg of lactose per ml and 0.5 μg of methionine per ml. When called for, these plates were also supplemented with 40 μg of X-Glu (5-bromo-4-chloro-3-indolyl-β-d-glucoside; Research Organics) per ml. This dye stains colonies with active β-glucosidase deep blue. We find that colonies of Bgl− strains on plates containing glucose as a carbon source and X-Glu will yield blue papillae without the addition of another substrate for β-glucosidase. Thus, in the blue papillation method (47) as originally described for the lac system, phenyl-β-d-galactoside is added as a carbon source for papillae stained blue by X-Gal (5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside). Here, this second carbon source is not necessary. Perhaps as a result of the lower concentration of the X-Glu in the role of carbon source, sugars are utilized in the following order by revertants of the Lac− Bgl− strains: glucose > lactose > X-Glu.
Determination of mutator frequency.
The data from Tables 3 and 4 were used to calculate the spontaneous frequency of mismatch-repair-deficient mutants. Since the majority of cultures had no mutators, the fraction of cultures with no mutators was used to calculate the mean, m, for the Poisson distribution describing the distribution of mutants. In the case of cells at a density of 105 cells per plate, the zero fraction is 327/355 = 0.921. Since for the Poisson distribution Po = e−m, 0.921 is e−m, and m is 0.082, or, in this case, 0.082 × 10−5. Accounting for the 10% plating efficiency gives a value of m as 0.82 × 10−5. Similar calculations for cells at a density of 104 cells per plate give 0.63 × 10−5, for a plating efficiency of 100%. We find that the plating efficiency at 104 cells varies somewhat more than that for 105 cells, so we take 0.8 × 10−5 as a more probable value. This value is for the fraction of mutators (or mutant frequency) in a culture grown for about 33 generations.
TABLE 3.
Efficiency of plating of mutH derivative of AS18-29 in the presence of different densities of cells of AS18-29a
| Strain | Value/plate | ||
|---|---|---|---|
| AS18-29 (no. of cells) | 106 | 105 | 104 |
| mutH derivative (% plating efficiency) | 1 | 10 | 50–100 |
See Materials and Methods for details.
TABLE 4.
Spontaneous mutators in cultures of AS18-29 detected on selection plates with different cell densitiesa
| No. of AS18-29 cells | No. of cultures | No. of cul-tures with no mutators | Mutator plating efficiency (%) | No. of mutators found | No. of nonmu-tators | % Muta-tors |
|---|---|---|---|---|---|---|
| 105 | 355 | 327 | 10 | 43 | 48 | 47 |
| 104 | 344 | 323 | 50–100 | 21 | 27 | 44 |
Close to 700 cultures were plated at two different cell densities, and the number of mutators found in samples of each culture was recorded. Each sample represented either 104 or 105 cells. The frequency of mutators in the population was determined from the fraction of cultures with no mutators and the plating efficiency at the respective cell density (see Materials and Methods).
Plating efficiency of mutators.
Strain AS18-29 was plated at different densities on selective medium, and then dilutions of a mutH derivative were plated on the same plates and the percentages of mutator cells forming colonies were determined.
RESULTS
Finding mutants with useful selective markers.
We looked for mutations which could be used in combination with other mutations to provide useful selections for mutator strains. We initially targeted mutators lacking the mismatch repair system. These strains have greatly enhanced rates of transitions and also of frameshifts at runs of single base pairs or repeating dinucleotides (11, 33, 57). Therefore, strains such as CC107 (11), which reverts from Lac− to Lac+ via the addition of a G to a run of six G’s, are useful for selecting for mutators, as we described previously (39). In order to find additional markers to use in concert with Lac− strains such as CC107, we mutagenized CC107 and derivatives of CC107 and looked for auxotrophs and then characterized the mutants with regard to reversion rates in both wild-type and mismatch-repair-deficient (mutH or mutS) strain backgrounds. We also determined the nutritional requirement and mapped the mutation to one of the known loci on the chromosome in most cases (see Materials and Methods).
Auxotrophs found after treatment with ICR-191 are usually caused by additions or deletions at monotonous runs of GC base pairs (5). Some of these should create monotonous runs of 6 bp or more and thus show greatly enhanced reversion rates in a mismatch-repair-deficient strain compared to reversion in a wild-type strain. Approximately 1% of the survivors of ICR-191 treatment were auxotrophs, and about 75% of these could be identified with respect to the nutritional requirement. Mutants that were leaky or which did not show a significantly increased reversion in a mutH strain were discarded. Of 70 mutants derived by ICR-191 mutagenesis, 22 merited further study. These mutants are shown in Table 1. Many of these mutants have low reversion rates but show greatly increased rates in a mutH background. Thus, they represent promising indicators for mutators. Several of the strains, including AS18, which carries a mutation in the metE (or metR) gene, were selected for additional experiments.
TABLE 1.
Mutants detected after ICR-191 mutagenesis of strain CC107
| Strain | Requirement | Locusa | No. of revertants/ 108 cellsb
|
|
|---|---|---|---|---|
| WT | mutH or mutS | |||
| AS1 | Methionine | metBFJL | 30 | ∼700 |
| AS2 | Adenosine | 2 | ∼500 | |
| AS3 | ? | 10 | ∼10,000 | |
| AS4 | Cysteine | 2 | ∼500 | |
| AS8 | Adenine | 3 | ∼2,000 | |
| AS10 | Leucine | leu | 46 | ∼2,000 |
| AS11 | Guanine, adenosine | 5 | ∼1,000 | |
| AS18 | Methionine | metER | 10 | ∼1,000 |
| AS101 | Methionine | metER | 15 | ∼1,000 |
| AS107 | Uracil, cytosine | pyrE | 30 | ∼10,000 |
| AS112 | Adenosine | purA | 59 | ∼8,000 |
| AS114 | Threonine | thr | 33 | ∼2,500 |
| AS115 | Serine | serA | 6 | ∼1,500 |
| AS121 | Guanine, adenosine | purDH | 3 | ∼5,000 |
| AS129 | Histidine | his | 0 | ∼1,000 |
| AS130 | Tryptophan | trp | 7 | ∼5,000 |
| AS131 | Guanine, adenosine | purDH | 26 | ∼25,000 |
| AS132 | Guanine, adenosine | purF | 2 | ∼10,000 |
| AS137 | Methionine | metBFJL | 8 | ∼1,000 |
| AS139 | Tryptophan | trp | 25 | ∼4,000 |
| AS141 | Tyrosine | tyr | 7 | ∼3,500 |
| AS150 | Methionine, cysteine | 20 | ∼20,000 | |
Mutational locus responsible for the auxotrophic phenotype when determined.
The revertants per 108 cells were determined in multiple cultures. In the case of mutH or mutS derivatives, the number of revertants is difficult to determine exactly, since colonies continue to arise after plating. Therefore, the numbers given should be considered estimates. WT, wild type.
Detection of mutations in the bgl operon.
As an additional indicator of mutator strains, we sought mutations in the bgl operon that revert in response to mismatch-repair-defective backgrounds. Since the bgl operon is cryptic in most E. coli K-12 strains (56, 58), we first scored spontaneous mutants of both CC107 and AS18 that could grow on salicin as a sole carbon source for intense blue color on glucose minimal plates with X-Glu (see Materials and Methods). We selected one Bgl+ mutant from each strain, mutagenized them with ICR-191, and screened for white colonies on X-Glu plates. These were then tested for enhanced reversion in a mutH strain in a variation of the blue papillation assay (47), adapted for X-Glu (see Materials and Methods). One Bgl− mutant was selected for further use. The mutant derived from CC107 was termed CC107-17, and the mutant derived from AS18 was named AS18-29. In each case, the frequency of revertants to Bgl+ goes from approximately 10 per 108 cells in a wild-type strain to approximately 5,000 per 108 cells in a mutH strain. These strains form numerous blue papillae in a mismatch-repair-deficient background on X-Glu plates and thus serve as indicators for mutHLS and uvrD strains.
Sequential selection on a single medium detects mutators.
We initially used strain AS18-29, which carries frameshift mutations in both lacZ and metE, to detect mutators. We experimented with different media to select colonies of strain AS18-29 that overcome both the Lac− and the Met− defects. We applied the principles depicted in Fig. 1 and 2. Lactose medium with limiting amounts of glucose and methionine should allow cells to grow to a small population size of approximately 105 cells before exhausting the limiting carbon source (glucose). To grow further would require metabolizing the lactose, and this would require a mutation from Lac− to Lac+. Then, the Lac+ cell could grow until it exhausted the methionine and again reach 105 cells. Now, a second mutation to Met+ would be required to allow further colony growth. In cases where a mutator population of 105 cells would have a Lac+ or a Met+ revertant, we would expect each mutator cell to give rise to a full colony under the right conditions, whereas only a fraction (about 10−5 to 10−4) of the nonmutator cells would form colonies (Fig. 1). We therefore examined the plating efficiency of a mutH derivative of AS18-29 on medium with different limiting amounts of glucose and methionine and found the best results with 150 μg of glucose per ml and 0.5 μg of methionine per ml (see Materials and Methods). On these plates, the mutH strain formed colonies with a 50 to 100% plating efficiency, while the wild-type derivative formed no colonies at all. We determined mutator colonies by either testing for enhanced frequency of Rifr colonies or including X-Glu in the medium to indicate frequent mutation to Bgl+ by the appearance of many blue papillae. (In the absence of X-Glu, colonies are usually picked after 3 days, whereas in the presence of X-Glu an extra day is needed for optimal papillae formation.)
FIG. 1.
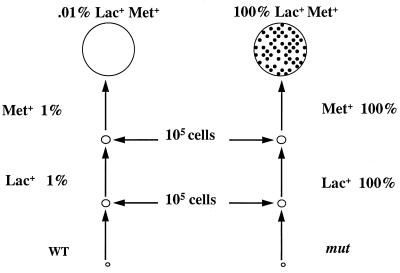
Selection for mutator colonies. Lac− Met− cells are plated on lactose medium with limiting amounts of glucose and methionine. Cells form a microcolony before exhausting the glucose. Revertants to Lac+ can grow further. Only 1% of the nonmutator microcolonies (left) will have a Lac+ cell, whereas 100% of the mutator microcolonies (right) will have a Lac+ cell. Further growth of the Lac+ cells yields microcolonies that exhaust the methionine. Again, 1% of the nonmutator Lac+ microcolonies (left) will have a Met+ cell, whereas 100% of the mutator Lac+ microcolonies (right) will. The very rare nonmutator Lac+ Met+ colonies have no Bgl+ papillae (left), but the Lac+ Met+ mutator colonies show microsatellite instability and give Bgl+ papillae (right; black dots). WT, wild type.
FIG. 2.
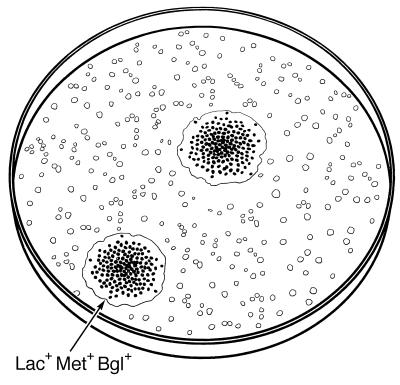
A representation of mutator colonies growing to full size after the selection described in the legend to Fig. 1. Many small microcolonies (open dots) are seen in the background, but only two full-grown colonies appear, and these have many Bgl+ papillae (dark dots).
To determine whether the selective medium could detect mutators from a mixed population, we grew AS18-29 overnight in 2AP and first measured the mutators by direct visualization on minimal glucose plates supplemented with methionine and X-Glu, as well as on the selective plates. The glucose–X-Glu plates with methionine allow the growth of all colonies, with mutators displaying a large number of blue papillae. This can be seen in Fig. 3A, which shows that of several hundred colonies, one prominent mutator is evident. In these experiments, mutators deficient in mismatch repair are found in between 1/1,000 and 1/300 cells after 2AP treatment. Figure 3B shows the result of plating an even larger sample of cells on the selective medium. Now, only the mutator colonies grow, as indicated by the blue color of the colonies, which is, upon magnification (Fig. 3C), really due to thousands of Bgl+ papillae from the X-Glu indicator. Figure 3C also allows the visualization of the microcolonies that failed to develop into true colonies. We verified that the surviving colonies were predominantly mutators by testing for increased frequency of Rifr mutants. Table 2 shows the results from several experiments. It can be seen that approximately 90% of the full-size colonies that grow on the selective plates after 2AP mutagenesis of strains AS18 and AS18-29 are strong mutators. Because both mutations that need to revert to restore the Lac+ Met+ phenotype are frameshifts, it is expected that the mutators are deficient in the mismatch repair system. We have examined sample mutators and found that all of the mutants tested carry a mutation in one of the four mismatch repair loci (mutHLS or uvrD [see below]).
FIG. 3.
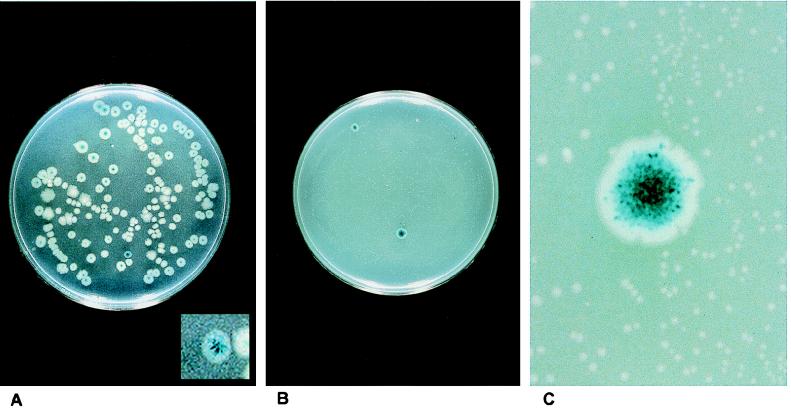
Selection for mutators. (A) Strain AS18-29 was grown overnight in LB broth with 2AP and plated on X-Glu plates. Here, one mutator colony (see insert) is evident, as indicated by the numerous blue Bgl+ papillae. (B) A photograph of a plate like that diagrammed in Fig. 2. Two full-size mutator colonies grow on selective plates onto which several thousand cells have been spread. Minute microcolonies in the background are nonmutator colonies. The mutator colonies appear blue because of thousands of blue Bgl+ papillae (see magnification in panel C). (C) A magnification of part of the plate shown in panel B. Here, the nonmutator microcolonies are evident, as are the blue Bgl+ papillae, representing the microsatellite instability of the full-size mutator colony.
TABLE 2.
Percentage of mutators among Lac+ Met+ colonies in 2AP cultures of AS18 and AS18-29 after the plate selection described in the text and for Fig. 1 to 5
| Strain | No. of 2AP cultures tested | No. of colonies tested | No. of mutator colonies | % Mutator |
|---|---|---|---|---|
| AS18 | 5 | 91 | 81 | 89 |
| AS18-29 | 5 | 136 | 127 | 93 |
We have carried out reconstruction experiments by mixing different proportions of wild-type and mutH derivatives of AS18-29 and then plating them on selective medium. As Table 3 shows, the selective plates are sensitive to cell density. The cell lawn eats up some of the limiting nutrients, and the efficiency of recovering a mutator goes down as the cell density increases. Placing up to 10,000 cells on a plate gives optimal results. As the number of cells approaches and then exceeds 105 per plate, the efficiency of recovery of mutators drops significantly. With 105 cells per plate, the efficiency drops from 50 to 100% to 10%, and with 106 cells, the efficiency drops to 1% recovery. Despite this, we can still detect mutators that occur spontaneously, at levels near 1 per 105 cells (see below).
Detection of spontaneous mutators.
We employed strain AS18-29 to detect spontaneous mutators in cultures grown for three generations. We examined 355 cultures, by plating 105 cells from each culture, and an additional 344 cultures, by plating 104 cells from each culture. Colonies were detected on selection medium without X-Glu and purified once before being tested for the frequency of Rifr colonies. Because the frequency of spontaneous mutators is on the order of 10−5 in the population, some Lac+ Met+ colonies are due to spontaneous mutations occurring in nonmutator cells. These colonies also appear on the order of 10−5 in the population. As Table 4 shows, about 50% of the colonies detected were mutators. About 92% of the plates with 105 cells yielded no mutator colonies. Applying the 10% plating efficiency determined from reconstruction experiments (Table 3), we can calculate the mutator frequency from the fraction of cultures with no mutators. This gives a frequency of mutators of 0.8 × 10−5 (see Materials and Methods). For the cultures plated with only 105 cells per plate, which gives a plating efficiency of 50 to 100%, approximately 94% of the cultures yielded no mutator colonies, which translates to 0.6 × 10−5 to 1.2 × 10−5 for the mutator frequency. These values are close to estimates based on other measurements we have made, as described in the Discussion.
Identification of mutators detected spontaneously.
We mapped the mutation causing the mutator phenotype in 45 of the spontaneous mutators detected in the above experiments, by P1 transduction (see Materials and Methods). All of the mutations fell into one of the four mismatch repair genes. However, as Table 5 shows, the mutations were not distributed equally among mutL, mutH, mutS, and uvrD. Instead, 19 of the mutations were in mutH, 18 were in mutL, 7 were in mutS, and 1 was in uvrD.
TABLE 5.
Position of the mutation resulting in the mutator phenotype in each of the 45 independent mutators of spontaneous origin
| Mutated gene | No. of spontaneous mutators from AS18-29 with mutation |
|---|---|
| mutH | 19 |
| mutL | 18 |
| mutS | 7 |
| uvrD | 1 |
Different combinations of markers are effective in selection.
The strain AS18-29 carries a mutation in the lac region on an F′ plasmid and a met mutation in the chromosome. The selective medium is not limited to those markers, nor does it depend on having a mutation on an F′ plasmid (data not shown). Several different combinations of markers were made for use in the successive selection medium. Some of the markers are derived from the experiment whose results are shown in Table 1. A second set of markers was derived by first making a mutH derivative of AS18-29 and obtaining auxotrophs (in the presence of methionine) induced by the mutator effect of the mutH allele. Then, the mutH allele was crossed out, and the reversion rates were compared. The new strains from this selection carry mutations that are not limited to additions or deletions at monotonous runs of G’s or C’s. They may contain additions or deletions at runs of A’s or T’s, or at repeating dinucleotides, as well as certain base substitutions. Table 6 shows the useful strains obtained from this selection. One additional mutation used in some experiments (data not shown) is the argE amber mutation derived from strain XAC (43). This mutation reverts via base substitutions that either restore the UAG codon to a sense codon or else create an amber or ocher suppressor.
TABLE 6.
Auxotrophic requirement of ICR-191-induced mutants of strain AS18-29a
| Strain | Requirement | Locus | No. of revertants/ 108 cells
|
|
|---|---|---|---|---|
| WT | mutH | |||
| AS201 | Cysteine | cys | 300 | ∼10,000 |
| AS210 | Leucine | leu | 40 | ∼5,000 |
| AS212 | Isoleucine | ile | 700 | ∼10,000 |
| AS213 | Arginine | arg | 20 | ∼10,000 |
| AS220 | Adenine | pur | 5 | ∼10,000 |
Each of the auxotrophic requirements is in addition to the Met− phenotype of AS18-29 (see legend to Fig. 1). WT, wild type.
Triple selection.
We utilized strain AS210 (Table 6), which carries frameshift mutations in lacZ, metE, and leu. Even though three successive selections occur, colonies were obtained on plates with limiting amounts of glucose, methionine, and leucine. After mutagenesis with 2AP, 97% of the colonies (26 of 27) were found to be mutators, and 90% of the colonies detected spontaneously were mutators. All of the mutator colonies tested had defects in one of the mismatch repair genes. The plating efficiency of a mutator on this medium, however, does not exceed 10%.
Mutator cascade.
Some mutators may not exhibit an increased rate of mutations at certain sequences, such as frameshifts at runs of identical bases of repeated sequences. However, they might show an increased rate of mutation at a second mutator locus, the resulting secondary mutators now being able to stimulate the specific sequence change. This type of mutator cascade might be significant in creating certain phenotypes. We can examine this phenomenon by looking at our tester strain, AS18-29, into which we have crossed a mutation at the mutT locus. The resulting mutators show an increased incidence of only one specific transversion, A:T→C:G (10), because of the failure to hydrolyze the oxidatively damaged DNA synthesis precursor 8-oxo-dGTP (38). That mutT cells do not greatly increase mutations at the frameshifts we used to monitor mutations in AS18-29 is evidenced in Table 7. Comparing the wild-type and mutT derivatives of AS18-29 with respect to reversion of either the lac, met, or bgl frameshift mutation shows either only slight or no differences compared with the mutH derivative of AS18-29, which shows an enormous increase (Table 7). Yet, when we look at the incidence of mutations in the mutHLS pathway that create mutators that can now stimulate the frameshifts in lac, met, and bgl, we see that mutT increases the spontaneous level of these mutators from 8 × 10−6 to 3 × 10−4.
TABLE 7.
Mutant frequencies for wild type and derivativesa
| Strain | No. of cultures with characteristic:
|
|||
|---|---|---|---|---|
| Lac+ | Met+ | Bgl+ | MMR− | |
| WT | 79 ± 4 | 8.3 ± 3.8 | 6.9 ± 4.3 | ∼800 |
| mutT | 76 ± 18 | 9.1 ± 3.4 | 8.1 ± 3.7 | ∼30,000 |
| mutH | ∼10,000 | ∼1,000 | ∼5,000 | |
Mutant frequencies determined as the averages of multiple cultures are shown for wild type (WT) (strain AS18-29; Lac− Met− Bgl−) and mutT or mutH derivatives prepared by transducing into AS18-29 a mini-Tn10 inserted into either the mutT or mutH locus. Mismatch repair deficiency (MMR−) results from loss of mutHLS or uvrD.
In the mutT derivative, approximately 1 of each 3,300 cells has a mutation in the mutHLS system. As a single cell divides and forms a colony, on average at some point between the 12th and 16th cell division the first mutator arises and forms a very thin sector, representing the lineage of daughter cells in the colony. Occasionally, a mutHLS mutator arises in the first cell division, yielding a colony that is half mutT alone and half mutT mutH (or mutL or mutS). We can visualize these sectors by decorating them with blue papillae arising from the Bgl+ reversion event at the frameshift in the bglA gene in AS18-24. Figure 4 shows some of these sectors. Recall that the starting mutT derivative of AS18-29 cannot give rise to frequent Bgl+ papillae. Figure 4A shows a colony that is half sectored for the mutHLS phenotype. Figure 4B shows a colony with a smaller mutator sector.
FIG. 4.
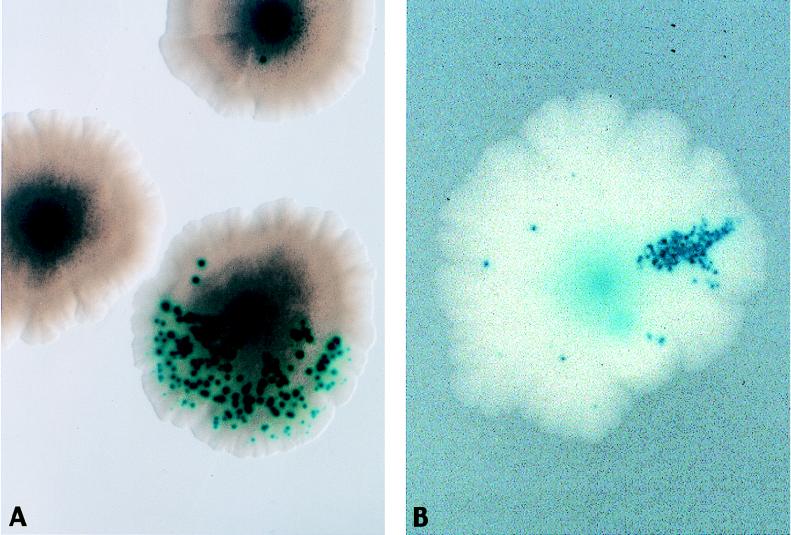
Appearance of mutator subpopulations as colony sectors. A mutT derivative of strain AS18-29 was plated on minimal glucose plates with methionine and X-Glu. Mismatch-repair-deficient (MMR−) sectors (e.g., mutH) are revealed by frequent blue Bgl+ papillae (microsatellite instability). (A) An MMR− mutant has arisen at the first cell division, resulting in half of the colony having microsatellite instability. (B) An MMR− mutant has arisen at approximately the fifth cell division, resulting in a thinner sector of the colony displaying microsatellite instability.
When the subpopulation of mutT-generated mutHLS mutators within a colony sector reaches a sufficient size, it can give rise to the first revertant able to overcome one of the growth restrictions imposed by the Lac− Met− Bgl− phenotype of the AS18-29 strain background. As depicted in Fig. 1 and 2, this can then lead to a progression that leads to the development of a full-size mutator colony. As Fig. 5 diagrams, in the case shown here, the starting strain can grow to only a certain size on medium with limiting methionine and limiting glucose. Then, the mutHLS mutator cell sector generates a Lac+ revertant (for instance), which can grow further, and then a Met+ cell that can grow without restrictions arises. The rapidly growing colony thus arises from within a slow-growing colony, extends out beyond the original colony, and displays microsatellite instability by throwing off frequent blue papillae in the presence of X-Glu. Figure 6 shows some of these colonies. Figure 6A and B show single colonies outgrowing the original slow-growing colony, and Fig. 6C shows how the unrestricted growth can ultimately invade other surrounding slow-growing or nongrowing microcolonies.
FIG. 5.
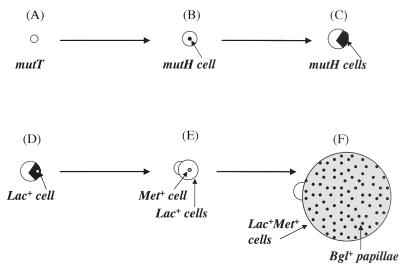
The effects of a mutator cascade are depicted in a schematic diagram of a mutT derivative of strain AS18-29 (Lac− Met− Bgl−) growing on lactose minimal medium with trace amounts of glucose and methionine and the indicator X-Glu. The starting cell (A) forms a microcolony, in which a mutH cell arises (B). As the microcolony grows, exhausting the glucose in the medium, the subpopulation of mutH cells also slowly proliferates (C), until a Lac+ cell arises within the mutH subpopulation (D). The Lac+ cells can grow further on the lactose in the medium until the trace methionine is exhausted. If a Met+ cell appears (E), it can now grow without the restriction of limiting methionine or glucose, expanding rapidly and showing microsatellite instability by the appearance of many blue Bgl+ papillae (F).
FIG. 6.
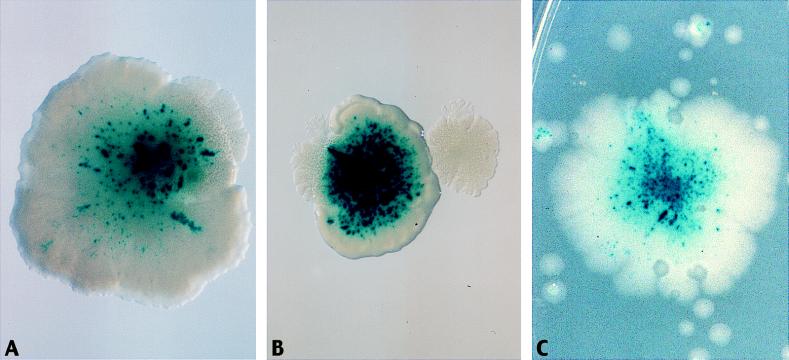
Mutator cascade. The figure shows selection for the growth of a mutator arising from a microcolony. The starting microcolony is a mutT derivative of strain AS18-29. The mutT mutator cannot revert the frameshift mutations in lacZ and metE, but can generate mutations in the mutH, -L, or -S genes, resulting in mismatch-repair-deficient (MMR−) mutators that can revert the frameshifts and break free of growth restrictions. (A and B) MMR− mutator colonies growing out from a microcolony. The MMR− mutators have many blue papillae, demonstrating microsatellite instability (see legend to Fig. 4). (C) An MMR− mutator generated on a more crowded plate will grow in an unrestricted fashion, proliferating over the plate and overrunning the nonmutator microcolonies.
DISCUSSION
Mutator cells, defined as cells with higher rates of mutation than those of wild type, were first detected in Drosophila melanogaster in the early 1940’s (54) and in bacteria in the early 1950’s (68). Now, mutations in more than 20 different genes in bacteria such as E. coli have been shown to result in the mutator phenotype (see reviews in references 8, 13, 19, 44, and 64). Most of these cause defects in repair or damage avoidance systems, although some affect less well defined pathways (7, 42, 63). Higher cells also display mutator phenotypes when certain genes are inactivated, and this can lead to increased incidence of disease. For instance, inheriting one copy of a defective gene involved in the human counterpart to the bacterial mismatch repair system leads to increased susceptibility to colon cancer, HNPCC-related endometrial cancer, and ovarian cancer (4, 18, 30, 49; see review in reference 37). The resulting tumor lines are mutators that show increased repeat-tract instability (see reviews in references 27 and 35) and higher mutation rates in genes such as hprt (1, 3, 21). Loeb had already postulated that creation of a mutator cell would be an early step in some cases of carcinogenesis, since a mutator can generate the multistep mutations faster than can a normal cell with lower mutation rates (27, 36). How then do mutators arise in a population of cells, and how do they proliferate?
Clearly, spontaneous mutants constantly arise in a population, and some of these are mutators. The balance of selective advantage and disadvantage will affect the proportion of mutators in any given environment, but the continued generation of new spontaneous mutations will provide a constant source. Using the selection system described in this work and discussed further below allows us to measure the proportion of cells in a growing population of E. coli cells that have defects in one of the four mismatch repair genes. This number of just under 10−5 (0.8 × 10−5) mutants is in a population growing in broth for about 33 generations. This number is in agreement with estimates from our previous study (39), in which Lac+ frameshift mutants detected at 2 × 10−7 were scored to reveal that 0.5% (5 × 10−3) were mutators. These values predict that mutants that are both Lac+ and defective in one of the four mismatch repair loci are present at 10−9 in the population (the product of these latter two frequencies). However, since the Lac+ revertant rate in each mutator subpopulation is close to 10−4 (9, 30), then the mutators are estimated to be in the population at 10−9 divided by 10−4, or close to 10−5. (Interestingly, a subsequent analysis of mismatch repair mutators in Salmonella typhimurium shows that they occur spontaneously an order of magnitude less frequently than found here for E. coli [32]).
Looking at the distribution of mutations among the four mismatch repair loci (Table 5) that leads to a combined rate of 0.8 × 10−5 mutators in the population makes us realize how these measurements can vary, depending on the specific gene being monitored. The mutant frequencies for each of the four loci are as follows: mutH, 3.4 × 10−6; mutL, 3.2 × 10−6; mutS, 1.2 × 10−6; uvrD, 1.8 × 10−7. It is not clear why only 1 of 45 mutations is in the uvrD gene. Reconstruction experiments indicate that the efficiency of plating of strains with uvrD mutations in the selection employed is the same as for strains with mutations in mutH (data not shown), suggesting that the effect is due to rates of mutations themselves, but it is possible that other hidden experimental biases conspire to reduce the appearance of UvrD mutants.
We should note at this point that mutagenesis from any of a number of sources serves to increase the proportion of mutators to near or above 10−3 (see, for instance, reference 39). However, in addition to mutagenesis, selection for mutant phenotypes can serve to increase the frequency of mutators in a population. Chemostat experiments show that extensive continued growth in a full-nutrient environment can still lead to the selection of fitter strains and increase the proportion of mutators, since they can give rise to fitter variants more rapidly than can the wild type (6, 9, 20, 46, 61, 65). Computer simulations also argue that mutators are selected for in continued growth in chemostats (67). In fact, some mutators were originally found by procedures designed to screen mutagenized cells after single or successive selections (12, 23, 24, 34) or by observation of cells selected for a specific phenotype (60, 61). Several investigators have found that natural isolates of bacteria have several percent mutators (22, 28, 31), suggesting that populations in the wild might be undergoing constant selection. Cebula and coworkers have argued that the several percent mutators found among E. coli and Salmonella strains isolated from patients might be related to pathogenicity (31), although others have disputed this correlation (41).
In a previous study (39), we showed how quickly the proportion of mutators can rise in a population, and how several successive selections can result in the entire surviving population being mutator. This underscores the consequences of certain chemotherapies, in which the resistant cells may be enriched for mutators. In the previous work (39), the successive selections involved transferring cells to different media. We have extended that study in the work reported here by designing a single medium that selects for several phenotypes in succession. Cells with multiple nutrient requirements are challenged to form colonies by eventually overcoming each of the growth barriers. In the most studied case, strain AS18-29, carrying frameshifts in the lacZ gene on the F plasmid and in the chromosomal metE gene, must revert both mutations in order to form full-size Lac+ Met+ colonies. A second strain adds a third growth requirement via a frameshift mutation in the chromosomal leu operon. The medium contains very small amounts of glucose as a carbon source and small amounts of methionine (and also of leucine when relevant). This permits a microcolony of nearly 105 cells to form, which if derived from a mutator cell will contain enough cells to have a mutant that can now use, for example, the lactose in the medium as a carbon source to initiate a new microcolony of 105 cells before it exhausts the methionine. A mutator cell will now have a second mutation in this new microcolony that reverses the metE mutation, allowing it to form a full-size colony (in the case of the derivative with the mutation in leu, a third round of successive microcolony formation would be required). Full-size colonies can be picked and analyzed or directly visualized by using reversion of a frameshift mutation in the bgl operon to decorate mutators with scores of blue papillae. The results, shown in Fig. 3, reveal that cells having to overcome several growth requirements in succession on a single defined medium give rise to mutator colonies. After mild mutagenesis, 90 to 100% of the colonies on this medium are mutators with defects in the mismatch repair system, depending on whether two or three growth requirements are employed. Even without a mutagen, spontaneous mutators constitute 50% of the colonies of cells that break through the growth restrictions.
It is interesting to compare the emergence of mutator colonies on plates selecting for overcoming several successive growth restrictions with the emergence of a cancer cell that undergoes successive mutations to break free of growth restrictions, particularly in the case of colon cancer. In both cases, frameshift mutations at runs of mono- or dinucleotides are involved in creating mutants. In the case of the bacterial strain shown here, the frameshift mutations restore the normal gene, whereas in the carcinogenesis model, frameshifts inactivate certain genes. Many colon cancer cell lines have mutations in the APC gene (25), the rII gene encoding the negative growth suppressor transforming growth factor β II (40, 50), and the apoptosis-associated BAX gene (55). The APC gene contains runs of A residues and an AG dinucleotide repeat that are frameshifted in sequenced mismatch-repair-deficient tumor lines (25). In these lines, most of the mutations inactivating transforming growth factor β are at a run of 10 A’s or at a threefold repeat of a GT sequence in the rII gene (40), and frameshifts at a run of eight G’s are found in the BAX gene (55). It is easy to see the parallels between successive selections that enrich for mismatch-repair-deficient strains in bacteria and those in human tumor lines that have to overcome several growth restrictions.
Figures 4 to 6 portray the events that occur when a mutHLS mutator cell arises within a colony of cells growing very slowly under restrictive conditions. As all the cells grow, the patch of cells derived from the mutHLS mutator cell (Fig. 4) reaches a sufficient size to allow the appearance of a mutant that can overcome subsequent growth restrictions (Fig. 5), proliferate during unrestricted growth, and exhibit repeat-tract instability (Fig. 6). In this series of experiments, the appearance of the mutHLS mutator is accelerated by the presence of a different mutator, in this case mutT, that cannot revert the frameshifts (Table 7) to overcome the growth restrictions but that can generate mutations that inactivate the mutH, mutL, or mutS gene. This produces a mutator cascade, where one mutator induces a second one. We have also found similar results with the mutA mutators, which result from miscoding tRNAs (data not shown). It will be interesting to see whether any examples of such a mutator cascade are found among tumor lines, or whether any cancer susceptibilities are found to result from the inheritance of a defective copy of a different repair gene that by itself leads to a mutator effect without microsatellite instability but which can generate mismatch-repair-deficient mutators. There is precedent for inactivation of the mismatch repair genes as a second step in the development of colon cancer. A significant fraction of sporadic colon cancer tumor lines that show microsatellite instability suffer inactivation of mismatch repair genes as a consequence of the epigenetic gene silencing produced by hypermethylation of both copies of the relevant promoters (29).
ACKNOWLEDGMENT
This work was supported by grant GM32184 to J.H.M. from the National Institutes of Health.
REFERENCES
- 1.Aaltonen L A, Peltomaki P, Leach F S, Sistonen P, Pylkkanen L, Mecklin J-P, Jarvinen H, Powell S M, Jen J, Hamilton S R, Petersen G M, Kinzler K W, Vogelstein B, de la Chapelle A. Clues to the pathogenesis of familial colorectal cancer. Science. 1993;260:812–816. doi: 10.1126/science.8484121. [DOI] [PubMed] [Google Scholar]
- 2.Armitage P, Doll R. The age distribution of cancer and multi-stage theory of carcinogenesis. Br J Cancer. 1954;8:1–12. doi: 10.1038/bjc.1954.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bhattacharyya N P, Skandalis A, Ganesh A, Groden J, Meuth M. Mutator phenotypes in human colorectal carcinoma cell lines. Proc Natl Acad Sci USA. 1994;91:6319–6323. doi: 10.1073/pnas.91.14.6319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bronner C E, Baker S M, Morrison P T, Warren G, Smith L G, Lescoe M K, Kane M, Earabino C, Lipford J, Lindblom A, Tannergard P, Bollag, Godwin A R, Ward D C, Nordenskjold M, Fishel R, Kolodner R, Liskay R M. Mutation in the DNA mismatch repair gene homologue hMLH1 is associated with hereditary nonpolyposis colon cancer. Nature. 1994;368:258–261. doi: 10.1038/368258a0. [DOI] [PubMed] [Google Scholar]
- 5.Calos M P, Miller J H. Genetic and sequence analysis of frameshift mutations induced by ICR-191. J Mol Biol. 1981;153:39–66. doi: 10.1016/0022-2836(81)90525-8. [DOI] [PubMed] [Google Scholar]
- 6.Chao L, Cox E C. Competition between high and low mutating strains of Escherichia coli. Evolution. 1983;37:125–134. doi: 10.1111/j.1558-5646.1983.tb05521.x. [DOI] [PubMed] [Google Scholar]
- 7.Connolly D M, Winkler M E. Genetic and physiological relationships among the miaA gene, 2-methylthio-N6-(Δ2-isopentenyl)-adenosine tRNA modification, and spontaneous mutagenesis in Escherichia coli K-12. J Bacteriol. 1989;171:3233–3246. doi: 10.1128/jb.171.6.3233-3246.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cox E C. Bacterial mutator genes and the control of spontaneous mutation. Annu Rev Genet. 1976;10:135–156. doi: 10.1146/annurev.ge.10.120176.001031. [DOI] [PubMed] [Google Scholar]
- 9.Cox E C, Gibson T C. Selection for high mutation rates in chemostats. Genetics. 1974;77:169–184. doi: 10.1093/genetics/77.2.169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cox E C, Yanofsky C. Altered base ratios in the DNA of an Escherichia coli mutator strain. Proc Natl Acad Sci USA. 1967;58:1895–1902. doi: 10.1073/pnas.58.5.1895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cupples C G, Cabrera M, Cruz C, Miller J H. A set of lacZ mutations in Escherichia coli that allow rapid detection of specific frameshift mutations. Genetics. 1990;125:275–280. doi: 10.1093/genetics/125.2.275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Degnen G E, Cox E C. A conditional mutator gene in Escherichia coli: isolation, mapping, and effector studies. J Bacteriol. 1974;117:477–487. doi: 10.1128/jb.117.2.477-487.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Drake J W. A constant rate of spontaneous mutation in DNA-based microbes. Proc Natl Acad Sci USA. 1991;88:7160–7164. doi: 10.1073/pnas.88.16.7160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Drake J W. Spontaneous mutation. Annu Rev Genet. 1991;25:125–146. doi: 10.1146/annurev.ge.25.120191.001013. [DOI] [PubMed] [Google Scholar]
- 15.Drake J W. Rates of spontaneous mutation among RNA viruses. Proc Natl Acad Sci USA. 1993;90:4171–4175. doi: 10.1073/pnas.90.9.4171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Eigen M. Viral quasispecies. Sci Am. 1993;269:42–49. doi: 10.1038/scientificamerican0793-42. [DOI] [PubMed] [Google Scholar]
- 17.Eshleman J R, Markowitz S D. Microsatellite instability in inherited and sporadic neoplasms. Curr Opin Oncol. 1995;7:83–89. [PubMed] [Google Scholar]
- 18.Fishel R, Lescoe M K, Rao M R S, Copeland N G, Jenkins N A, Garber J, Kane M, Kolodner R. The human mutator gene homolog MSH2 and its association with hereditary nonpolyposis colon cancer. Cell. 1993;75:1027–1038. doi: 10.1016/0092-8674(93)90546-3. [DOI] [PubMed] [Google Scholar]
- 19.Friedberg E C, Walker G C, Seide W. DNA repair and mutagenesis. Washington, D.C: American Society for Microbiology; 1995. [Google Scholar]
- 20.Gibson T C, Scheppe M L, Cox E C. Fitness of an Escherichia coli mutator gene. Science. 1970;169:686–688. doi: 10.1126/science.169.3946.686. [DOI] [PubMed] [Google Scholar]
- 21.Glaab W E, Tindall K R. Mutation rate at the hprt locus in human cancer cell lines with specific mismatch repair-gene defects. Carcinogenesis. 1997;18:1–8. doi: 10.1093/carcin/18.1.1. [DOI] [PubMed] [Google Scholar]
- 22.Gross M D, Siegel E C. Incidence of mutator strains in Escherichia coli and coliforms in nature. Mutat Res. 1981;91:107–110. doi: 10.1016/0165-7992(81)90081-6. [DOI] [PubMed] [Google Scholar]
- 23.Helling R B. Selection of a mutant of Escherichia coli which has high mutation rates. J Bacteriol. 1968;96:975–980. doi: 10.1128/jb.96.4.975-980.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hoess R H, Herman R K. Isolation and characterization of muator strains of Escherichia coli K-12. J Bacteriol. 1975;122:474–484. doi: 10.1128/jb.122.2.474-484.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Huang J, Papadopoulos N, McKinley A J, Farrington S M, Curtis L J, Wyllie A H, Zheng S, Willson J K V, Markowitz S D, Morin P, Kinzler K W, Vogelstein B, Dunlop M G. APC mutations in colorectal tumors with mismatch repair deficiency. Proc Natl Acad Sci USA. 1996;93:9049–9054. doi: 10.1073/pnas.93.17.9049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ionov Y, Peinado M A, Malkhosyan S, Shibata D, Perucho M. Ubiquitous somatic mutations in simple repeated sequences reveal a new mechanism for colonic carcinogenesis. Nature. 1995;363:558–561. doi: 10.1038/363558a0. [DOI] [PubMed] [Google Scholar]
- 27.Jackson A L, Loeb L A. The mutation rate and cancer. Genetics. 1998;148:1483–1490. doi: 10.1093/genetics/148.4.1483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jyssum K. Observations on two types of genetic instability in Escherichia coli. Acta Pathol Microbiol Scand. 1960;48:113–120. doi: 10.1111/j.1699-0463.1960.tb04747.x. [DOI] [PubMed] [Google Scholar]
- 29.Kane M F, Loda M, Gaida G M, Lipman J, Mishra R, Goldman H, Jessup J M, Kolodner R. Methylation of the hMLH1 promoter correlates with lack of expression of hMLH1 in sporadic colon tumors and mismatch repair-defective human tumor cell lines. Cancer Res. 1997;57:808–811. [PubMed] [Google Scholar]
- 30.Leach F S, Nicolaides N C, Papadopoulos N, Liu B, Jen J, Parsons R, Petlomaki P, Sistonene P, Aaltonen L A, Nystrom-Lahti M, Guan X-Y, Zhang J, Meltzer P S, Yu J-W, Kao F-T, Chen D J, Cerosaletti K M, Fournier R E, Rodd S, Lewis T, Leach R J, Naylor S L, Weissenbach J, Mecklin J-P, Jarvinen H, Petersen G M, Hamilton S R, Green J, Jass J, Watson P, Lynch H T, Trent J M, de la Chapelle A, Kinzler K W, Vogelstein B. Mutations of a mutS homolog in hereditary nonpolyposis colorectal cancer. Cell. 1993;75:1215–1225. doi: 10.1016/0092-8674(93)90330-s. [DOI] [PubMed] [Google Scholar]
- 31.LeClerc J E, Li B, Payne W L, Cebula T A. High mutation frequencies among Escherichia coli and Salmonella pathogens. Science. 1996;274:1208–1211. doi: 10.1126/science.274.5290.1208. [DOI] [PubMed] [Google Scholar]
- 32.LeClerc J E, Payne W L, Kupchella E, Cebula T A. Detection of mutator subpopulations in Salmonella typhimurium LT2 by reversion of his alleles. Mutat Res. 1998;400:89–97. doi: 10.1016/s0027-5107(98)00069-4. [DOI] [PubMed] [Google Scholar]
- 33.Leong P M, Hsia H C, Miller J H. Analysis of spontaneous base substitutions generated in mismatch-repair-deficient strains of Escherichia coli. J Bacteriol. 1986;168:412–416. doi: 10.1128/jb.168.1.412-416.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liberfarb R M, Bryson V. Isolation, characterization and genetic analysis of mutator genes in Escherichia coli B and K-12. J Bacteriol. 1970;104:363–375. doi: 10.1128/jb.104.1.363-375.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Loeb L A. Mutator phenotype may be required for multistage carcinogenesis. Cancer Res. 1991;51:3075–3079. [PubMed] [Google Scholar]
- 36.Loeb L A. Microsatellite instability: marker of mutator phenotype in cancer. Cancer Res. 1994;54:5059–5063. [PubMed] [Google Scholar]
- 37.Lynch H T, Smyrk T, Lynch J. An update of HNPCC (Lynch syndrome) Cancer Genet Cytogenet. 1997;93:84–99. doi: 10.1016/s0165-4608(96)00290-7. [DOI] [PubMed] [Google Scholar]
- 38.Maki H, Sekiguchi M. MutT protein specifically hydrolyses a potent mutagenic substrate for DNA synthesis. Nature. 1992;355:273–275. doi: 10.1038/355273a0. [DOI] [PubMed] [Google Scholar]
- 39.Mao E F, Lane L, Lee J, Miller J H. Proliferation of mutators in a cell population. J Bacteriol. 1997;179:417–422. doi: 10.1128/jb.179.2.417-422.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Markowitz S, Wang J, Myeroff L, Parsons R, Sun L, Lutterbaugh J, Fan R S, Zborowska E, Kinzler K W, Vogelstein B, Brattain M, Willson J K V. Inactivation of the type II TGF-β receptor in colon cancer cells with microsatellite instability. Science. 1995;268:1336–1338. doi: 10.1126/science.7761852. [DOI] [PubMed] [Google Scholar]
- 41.Matic I, Radman M, Taddei F, Picard B, Doit C, Bingen E, Denamur E, Elion J. Highly variable mutation rates in commensal and pathogenic Escherichia coli. Science. 1997;277:1833–1834. doi: 10.1126/science.277.5333.1833. [DOI] [PubMed] [Google Scholar]
- 42.Michaels M L, Cruz C, Miller J H. mutA and mutC; two mutator loci in Escherichia coli that stimulate transversions. Proc Natl Acad Sci USA. 1990;89:9211–9215. doi: 10.1073/pnas.87.23.9211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Miller J H. A short course in bacterial genetics: a laboratory manual and handbook for Escherichia coli and related bacteria. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1992. pp. 194–195. [Google Scholar]
- 44.Miller J H. Spontaneous mutators in bacteria: insights into pathways of mutagenesis and repair. Annu Rev Microbiol. 1996;50:625–643. doi: 10.1146/annurev.micro.50.1.625. [DOI] [PubMed] [Google Scholar]
- 45.Miller, J. H. Unpublished data.
- 46.Nestman E R, Hill R F. Population changes in continuously growing mutator cultures of Escherichia coli. Genetics. 1973;73(Suppl.):41–44. [PubMed] [Google Scholar]
- 47.Nghiem Y, Cabrera M, Cupples C G, Miller J H. The mutY gene: a mutator locus in Escherichia coli that generates GC→TA transversions. Proc Natl Acad Sci USA. 1988;85:2709–2713. doi: 10.1073/pnas.85.8.2709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Nowell P C. The clonal evolution of tumor cell populations. Science. 1976;194:23–28. doi: 10.1126/science.959840. [DOI] [PubMed] [Google Scholar]
- 49.Papadopoulos N, Nicolaides N C, Wei Y F, Ruben S M, Carter K C, Rosen C A, Haseltine W A, Fleischmann R D, Fraser C M, Adams M S. Mutation of a mutL homolog in hereditary nonpolyposis colon cancer. Science. 1994;263:1625–1629. doi: 10.1126/science.8128251. [DOI] [PubMed] [Google Scholar]
- 50.Parsons R, Li G-M, Longley M J, Fang W-H, Papadopoulos N, Jen J, de la Chapelle A, Kinzler K W, Vogelstein B, Modrich P. Hypermutability and mismatch repair deficiency in RER tumor cells. Cell. 1993;75:1227–1236. doi: 10.1016/0092-8674(93)90331-j. [DOI] [PubMed] [Google Scholar]
- 51.Parsons R, Myeroff L, Liu B, Willson J K V, Markowitz S D, Kinzler K W, Vogelstein B. Microsatellite instability and mutations of the transforming growth factor beta type II receptor gene in colorectal cancer. Cancer Res. 1995;55:5548–5550. [PubMed] [Google Scholar]
- 52.Peinado M A, Malkhosyan S, Velazquez A, Perucho M. Isolation and characterization of allelic losses and gains in colorectal tumors by arbitrarily primed polymerase chain reaction. Proc Natl Acad Sci USA. 1992;89:10065–10069. doi: 10.1073/pnas.89.21.10065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Peltomaki P, Lothe R A, Aaltonen L A, Pylkkanen L, Nystrom-Lahti M, Seruca R, David L, Holm R, Ryberg D, Gaugen A, Brogger A, Borresen A-L, de la Chapelle A. Microsatellite instability is associated with tumors that characterize the hereditary non-polyposis colorectal carcinoma syndrome. Cancer Res. 1993;53:5853–5855. [PubMed] [Google Scholar]
- 54.Plough H H. Spontaneous mutability in Drosophila. Cold Spring Harbor Symp Quant Biol. 1941;9:127–137. [Google Scholar]
- 55.Rampino N, Yamamoto H, Ionov Y, Li Y, Sawai H, Reed J C, Perucho M. Somatic frameshift mutations in the BAX gene in colon cancers of the microsatellite mutator phenotype. Science. 1997;275:967–969. doi: 10.1126/science.275.5302.967. [DOI] [PubMed] [Google Scholar]
- 56.Reynolds A E, Felton J, Wright A. Insertion of DNA activates the cryptic bgl operon in E. coli K12. Nature. 1981;293:625–629. doi: 10.1038/293625a0. [DOI] [PubMed] [Google Scholar]
- 57.Schaaper R M, Dunn R L. Spectra of spontaneous mutations in Escherichia coli strains defective in mismatch correction: the nature of in vivo DNA replication errors. Proc Natl Acad Sci USA. 1987;84:6220–6224. doi: 10.1073/pnas.84.17.6220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Schaefler S. Inducible system for the utilization of betaglucosides in Escherichia coli. I. Active transport and utilization of beta-glucosides. J Bacteriol. 1967;93:254–263. doi: 10.1128/jb.93.1.254-263.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Shibata D, Navidi W, Salovaara R, Li Z H, Aaltonen L A. Somatic microsatellite mutations as molecular tumor clocks. Nat Med. 1996;2:676–681. doi: 10.1038/nm0696-676. [DOI] [PubMed] [Google Scholar]
- 60.Siegel E C, Bryson V. Antimicrob. Agents Chemother. 1963. 1964. Selection of resistant strains of Escherichia coli by antibiotics and antibacterial agents: role of normal and mutator strains; pp. 629–634. [PubMed] [Google Scholar]
- 61.Siegel E C, Bryson V. Mutator gene of Escherichia coli B. J Bacteriol. 1967;94:38–47. doi: 10.1128/jb.94.1.38-47.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Singer M, Baker T A, Schnitzler G, Deischel S M, Goel M, Dove W, Jaacks K J, Grossman A D, Erickson J W, Gross C A. A collection of strains containing genetically linked alternating antibiotic resistance elements for genetic mapping of Escherichia coli. Microbiol Rev. 1989;53:1–24. doi: 10.1128/mr.53.1.1-24.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Slupska M S, Baikalov C, Lloyd R, Miller J H. Mutator tRNAs are encoded by the Escherichia coli mutator genes mutA and mutC: a novel pathway for mutagenesis. Proc Natl Acad Sci USA. 1995;93:4380–4385. doi: 10.1073/pnas.93.9.4380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Smith K C. Spontaneous mutagenesis: experimental, genetic, and other factors. Mutat Res. 1992;277:139–162. doi: 10.1016/0165-1110(92)90002-q. [DOI] [PubMed] [Google Scholar]
- 65.Sniegowski P D, Gerrish P J, Lenski R E. Evolution of high mutation rates in experimental populations of E. coli. Nature. 1997;387:703–705. doi: 10.1038/42701. [DOI] [PubMed] [Google Scholar]
- 66.Strand M, Prolla T A, Liskay R M, Petes T D. Destabilization of tracts of simple repetitive DNA in yeast by mutations affecting DNA mismatch repair. Nature. 1993;365:274–276. doi: 10.1038/365274a0. [DOI] [PubMed] [Google Scholar]
- 67.Taddei F, Radman M, Maynard-Smith J, Toupance B, Gouyon P H, Godelle B. Role of mutator alleles in adaptive evolution. Nature. 1997;387:700–702. doi: 10.1038/42696. [DOI] [PubMed] [Google Scholar]
- 68.Treffers H P, Spinelli V, Belser N O. A factor (or mutator gene) influencing mutation rates in Escherichia coli. Proc Natl Acad Sci USA. 1954;40:1064–1071. doi: 10.1073/pnas.40.11.1064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Umar A, Kunkel T A. DNA-replication fidelity, mismatch repair and genome instability in cancer cells. Eur J Biochem. 1996;238:297–307. doi: 10.1111/j.1432-1033.1996.0297z.x. [DOI] [PubMed] [Google Scholar]
- 70.Weinberg R A. How cancer arises: an explosion of research is uncovering the long-hidden molecular underpinnings of cancer—and suggesting new therapies. Sci Am. 1996;275:62–70. doi: 10.1038/scientificamerican0996-62. [DOI] [PubMed] [Google Scholar]