Abstract
Coronaviruses (CoVs) have the largest genome among RNA viruses and store large amounts of information without genome integration as they replicate in the cell cytoplasm. The replication of the virus is a continuous process, whereas the transcription of the subgenomic mRNAs is a discontinuous one, involving a template switch, which resembles a high frequency recombination mechanism that may favor virus genome variability. The origin of the three deadly human CoVs SARS-CoV, MERS-CoV and SARS-CoV-2 are zoonotic events. SARS-CoV-2 has incorporated in its spike protein a furine proteolytic site that facilitates the activation of the virus in any tissue, making this CoV strain highly polytropic and pathogenic. Using MERS-CoV as a model, a propagation-deficient RNA replicon was generated by removing E protein gene (essential for viral morphogenesis and involved in virulence), and accessory genes 3, 4a, 4b and 5 (responsible for antagonism of the innate immune response) to attenuate the virus: MERS-CoV-Δ[3,4a,4b,5,E]. This RNA replicon is strongly attenuated and elicits sterilizing protection after a single immunization in transgenic mice with the receptor for MERS-CoV, making it a promising vaccine candidate for this virus and an interesting platform for vector-based vaccine development. A strategy could be developed for the design of RNA replicon vaccines for other human pathogenic coronaviruses.
Keywords: Coronavirus, MERS-CoV, SARS-CoV-2, Transcription, RNA replicon, Vaccine
Graphical abstract
Highlights
-
•
CoV RNA transcription is a discontinuous high-frequency recombination process.
-
•
Human CoVs have a zoonotic origin.
-
•
Replication-competent propagation-defective CoVs are the bases for vaccines.
-
•
Single immunization with CoV RNA replicon vaccines elicits sterilizing immunity.
1. Introduction
Coronaviruses (CoVs) are a family of enveloped, positive strand RNA viruses of the Nidovirales order. The Orthocoronavirinae subfamily is divided into the Alpha, Beta-, Gamma- and Deltacoronavirus genera (de Groot et al., 2013; Gorbalenya et al., 2020; ICTV, 2019). Viruses of these four genera can infect a wide range of birds and mammals, including humans (Banerjee et al., 2019; Colina et al., 2021; Corman et al., 2018; Cui et al., 2019), producing different clinical signs depending on the infected tissue and organ (Corman et al., 2018; Polak et al., 2020; van den Brand et al., 2015; Zappulli et al., 2020). Seven CoVs infecting humans have been identified. Of them, HCoV-229E, HCoV-OC43, HCoV-NL63, and HCoV-HUK1 cause mild infections similar to common colds (Corman et al., 2018; Perlman and Netland, 2009). In contrast, severe acute respiratory syndrome coronavirus (SARS-CoV) (Drosten et al., 2003; Rota et al., 2003), Middle East respiratory syndrome coronavirus (MERS-CoV) (de Groot et al., 2013; Zaki et al., 2012) and SARS-CoV-2 (Hu et al., 2021; Zhu et al., 2020) cause a severe respiratory infection that can be fatal. The three highly pathogenic human CoVs differ in transmissibility and mortality rate: SARS-CoV in 2002 (number of cases 8437, average mortality 10%) (Drosten et al., 2003; Rota et al., 2003), MERS-CoV in 2012 (2650 cases, around 800 deaths) (de Groot et al., 2013; Zaki et al., 2012) and SARS-CoV-2 (530 million cases, more than 6 million deaths) (https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports, June 9th, 2022).
In this review, essential concepts of CoV biology, such as viral RNA synthesis and origin, which influence the development of antiviral strategies, will be summarized. In addition, novel CoV vaccine candidate approaches, based on CoV-derived RNA replicons, will be reviewed.
2. Replication and transcription in CoVs and stability of their genome
CoVs are spherical viruses with a single-strand positive-sense RNA genome with around 30 thousand nucleotides, among the largest virus RNA genomes identified so far. The 5′-proximal two-thirds of the coronavirus genome encode the replicase gene, which contains two open reading frames, ORF1a and ORF1b. Translation of ORF1a yields polyprotein 1a (pp1a), and −1 ribosomal frameshifting allows translation of ORF1b to yield pp1ab (Firth and Brierley, 2012; Masters, 2006). Together, these polyproteins are co- and post-translationally processed into 16 nonstructural proteins (nsps), most of them driving viral genome replication and subgenomic mRNA (sgmRNA) synthesis. The 3′ third of the genome encodes the structural and accessory proteins, which vary in number among the different coronaviruses (Fig. 1).
Fig. 1.
CoV genome. Genome organization of the three highly pathogenic human CoVs: SARS-CoV, MERS-CoV and SARS-CoV-2. The letters above the boxes indicate the viral genes. Leader sequence (L), replicase genes (ORF1a, ORF1b), spike (S), envelope (E), membrane (M) and nucleocapsid (N) genes. Accessory genes are indicated by numbers and are different for each CoV. An, poly A sequence.
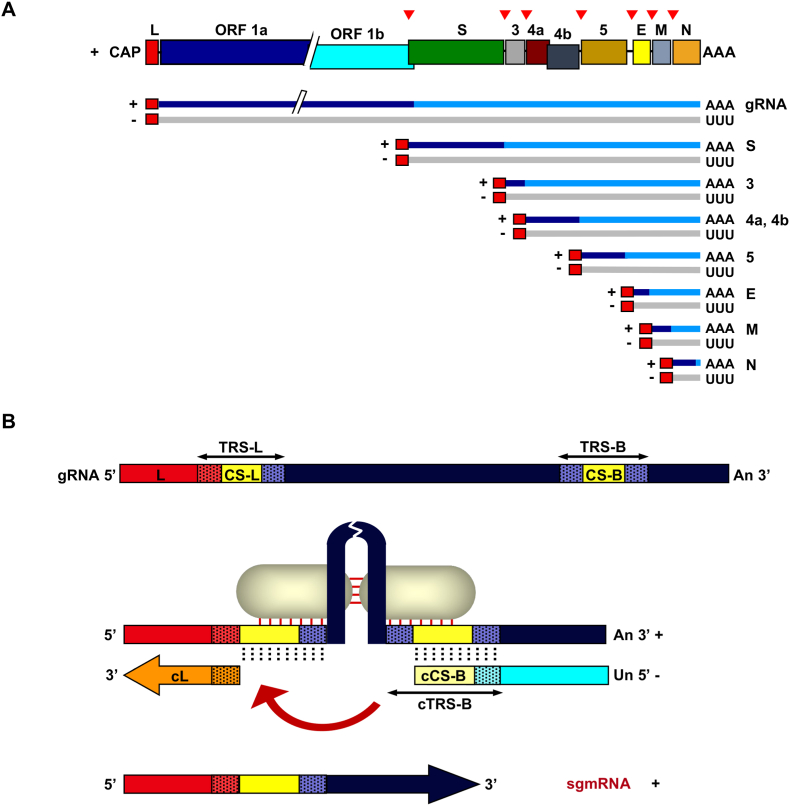
CoV RNA synthesis is performed by a replication-transcription complex that includes viral and cell proteins that recognize cis-acting RNA elements mainly located in the highly structured 5′ and 3′ untranslated genomic regions. Replication of the coronavirus genome requires continuous RNA synthesis, whereas transcription is a discontinuous process unique among RNA viruses (Sola et al., 2015). Similar to other positive-strand RNA viruses, continuous RNA synthesis during genome replication utilizes a full-length complementary negative-strand RNA as the template for the production of progeny virus genomes (Fig. 2A).
Fig. 2.
CoV RNA synthesis. (A) Upper panel represents the MERS-CoV genome as in Fig. 1. During infection, the viral polymerase produces a series of viral RNAs (lower panel), of positive (+) polarity and other complementary RNAs of negative (−) polarity. The dark blue line indicates the portion of the gRNA or mRNA that is translated. Numbers and letter in the right indicated the viral proteins translated from each viral mRNA. (B) CoV transcription. The coding gRNA of positive polarity is shown as a dark blue line, in which leader sequence (red) and TRSs from leader (TRS-L) and body (TRS-B) are indicated, highlighting the core sequences in each case (yellow). The negative polarity RNA is shown as a light blue line, were complementary sequences (cL, cTRS-B and cCS-B) are indicated. RNA-RNA and RNA-protein interactions (represented by the RNA folding and grey ovals, respectively) control the template switch (red arrow). (For interpretation of the references to colour in this figure legend, the reader is referred to the Web version of this article.)
In contrast to replication, CoV transcription includes a discontinuous step during the production of sgmRNAs (Enjuanes et al., 2008; Sawicki et al., 2007). This process, unique among known RNA viruses, is a hallmark of the order Nidovirales and ultimately generates a nested set of sgmRNAs that are 5′ and 3′ coterminal with the virus genome. All sgmRNAs include at their 5′ end a common leader sequence, whose length ranges from 65 to 98 nt in different coronaviruses (van der Most and Spaan, 1995). This common leader sequence is present only once at the very 5′ end of the genome, which implies that sgmRNAs are synthesized by the fusion of noncontiguous sequences, the leader and the 5′ end of each mRNA coding sequence, named the “body” (B) (Fig. 2B). Transcription includes a template switch during the synthesis of subgenomic negative-strand RNAs to add a copy of the leader sequence to the nascent negative RNA. Coronavirus transcription is regulated by multiple factors, including the extent of base-pairing between transcription-regulating sequences (TRSs) of positive and negative polarity, viral and cell protein–RNA binding, and high-order RNA-RNA interactions (Fig. 2B).
The discontinuous synthesis adds a high risk in terms of variability of the RNA genome of coronavirus. CoV transcription resembles high-frequency, similarity-assisted copy-choice RNA recombination, requiring sequence identity between donor and acceptor RNAs and hairpin structures present in the acceptor RNA (Nagy and Simon, 1997), in which the TRS-L would act as an acceptor for the complementary cTRS-B donor sequence in the nascent negative RNA. This high frequency recombination event, is a procedure that increases the possibility of recombination among two coronaviruses infecting the same cell and facilitates the observed evolution of CoVs.
CoVs encode a proofreading machinery, unique in the RNA virus world. The proof-reading system maps in the nsp14 of the polymerase, and it was discovered by introducing point mutations in two domains of this nsp14 (Denison et al., 2011; Smith et al., 2014). A proof-reading system has only been described in nidoviruses with the largest known RNA genomes (above 20 thousand nt). In contrast, this activity is not present in the Arteriviridae, with a replication-transcription strategy similar to that of CoVs, but with a genome size of around 14 thousand nt, suggesting that the proof-reading system was selected in CoVs and other nidoviruses to maintain their large genome sizes and to avoid an error catastrophe by accumulation of mutations.
3. Human CoVs origin
Nowadays, the zoonotic origin of the three human deadly CoVs is known, including that of the SARS-CoV-2. The natural reservoir for these CoVs are bats and the viruses were transmitted to humans by an intermediate host (Fig. 3). SARS-CoV and MERS-CoV have been transmitted to people by civet cats and camels, respectively (de Wit et al., 2016). In fact, MERS-CoV prevalence in camels is very high, 80–99% depending on the country (Omrani et al., 2015). SARS-CoV-2 has been most-likely transmitted to humans from racoons, as a virus with a 99.998% sequence identity to SARS-CoV-2 has been found in the feces present in the metal cages of the Huanan market at Wuhan (Gao et al., 2022; Holmes et al., 2021; Maxmen, 2022; Pekar et al., 2022; Worobey et al., 2022).
Fig. 3.
Human CoVs origin. The three pathogenic human CoVs, SARS-CoV, MERS-CoV and SARS-CoV-2 were derived from natural reservoirs in bats (black), using intermediate hosts (orange) before jumping to humans (blue). Wild and domestic animals (grey) can be infected by SARS-CoV-2 or its variants, with possibility of virus circulating and adapting to these animal species, and posing the thread of creating novel virus reservoirs from which the virus could emerge to infect humans (dashed lines). (For interpretation of the references to colour in this figure legend, the reader is referred to the Web version of this article.)
This information is essential to control the reemergence of the virus from natural reservoirs. Unfortunately, the pandemic caused by the SARS-CoV-2 will have as additional problems for its eradication that the virus also infects wild, captive and domestic animals, such as white tail deer and ferrets in nature (Hale et al., 2022; Sharun et al., 2021a). Virus adaptation to these animal hosts has been described and, in some cases, such as with minks, the animal-adapted virus can cross species barriers and infect humans, which poses a thread for novel pathogenic variants emergence (Fig. 3). It has been proposed that a One-Health approach is needed to control SARS-CoV-2 pandemic and viral surveillance in animals is required, as novel potential animal reservoirs are being described (Sharun et al., 2021b).
4. Replicon-based vaccine candidate against MERS-CoV
To protect against MERS-CoV at least four vaccines have been developed at present (three based on viral-vectors and one on DNA) (Folegatti et al., 2020; Koch et al., 2020; Modjarrad et al., 2019; NCT, 2019a, b, c). Most of these vaccine candidates are based on the spike (S) protein, as this protein is the major inducer of virus neutralizing antibodies (Mou et al., 2013), and it includes the receptor binding domain (RBD) interacting with dipeptidyl peptidase 4 (DPP4) in the host cell (Raj et al., 2013). Our laboratory has developed CoV derived RNA replicons as promising platforms for vaccine generation (Gutiérrez-Álvarez et al., 2021; Lundstrom, 2020) (Fig. 4A). These replicons are replication-competent and propagation-defective, using an RNA dose 60-fold lower (Vogel et al., 2018) than conventional mRNA-based vaccines. Most frequently developed replicons are based on Alphaviruses such as Semliki forest virus (SFV) (Liljestrom and Garoff, 1991; Sanchez-Paulete et al., 2018), Sindbis virus (SIN) (Lundstrom, 2016), and Venezuelan equine encephalitis virus (VEE) (Agnihothram et al., 2018; Jurgens et al., 2012), among others (Lundstrom, 2020).
Fig. 4.
VLPs containing CoV-derived RNA replicons. (A) RNA replicon is introduced in packaging cell lines, providing the missing proteins (protein X) required for VLP formation. Then cells produce VLPs encapsulating the RNA replicon. (B) Upper panel: when cells are infected with WT virus, membrane vesicles containing virions were observed (EM image) and progeny viruses were produced. In contrast (lower panel), when replicon VLPs enter into the cells, some membrane rearrangements with VLPs were observed (EM image), but no infectious virus is produced.
The development of the first strategy to engineer CoV genomes described in this review, was reported by our laboratory in 2000 (Almazan et al., 2000), and was based in the cloning of a CoV cDNA copy into BACs. This strategy has been essential for the development of self-replicating RNAs derived from CoV RNA genomes by the deletion of a set of genes (Almazan et al., 2013; Castaño-Rodriguez et al., 2018; Ortego et al., 2007).
4.1. Virus-like particles assembly by rMERS -CoV-ΔE replicon in the absence of the E protein
Deletion of E gene from MERS-CoV lead to the formation of a replication-competent propagation defective CoV (Almazan et al., 2013). Packaging cell lines expressing E protein were required for optimum rescue and evaluation of RNA replicons derived from MERS-CoV mutants lacking the E gene (Fig. 4A). In order to select the appropriate rescue strategy, two different systems were evaluated: constitutive expression using a pcDNA3.1-E-MERS-CoV plasmid (Almazan et al., 2013), and an inducible expression system based on a Tetracycline (Tet-On) plasmid (TRE-Auto-rtTA-V10-2T-E-MERS-CoV), which included a positive feedback loop (Das et al., 2016). In the absence of E protein, the rMERS-CoV-ΔE replicon did not propagate, whereas in cells expressing E protein, virus production was observed at 24 h post-infection (hpi). At 72 hpi, MERS-CoV-ΔE titers increased 100-fold in cells transfected with pcDNA3.1-E-MERS-CoV compared to non-transfected cells. However, in cells transfected with TRE-Auto-rtTA-V10-2T-E-MERS-CoV, the titers of rMERS-CoV-ΔE increased more than 1000-fold, reaching levels close to those of the parental virus. These results showed that the inducible expression system facilitated optimal production of large amounts of MERS-CoV mutants lacking E gene, which can be implemented for vaccine candidate production in a factory.
The morphogenesis of parental rMERS-CoV-WT virus and rMERS-CoV-ΔE replicon was studied in the absence of E protein supplementation in Huh-7 cells (Fig. 4B). Cells infected with MERS-CoV-WT virus showed greater morphological alterations, with vesicles full of virions with a spherical shape. In contrast, in cells infected with rMERS-CoV-ΔE replicon, vesicles were less abundant, elongated, and included a lower number of virus-like particles (VLPs) (Fig. 4B). However, these polymeric structures had high immunogenic potential. In addition, the VLPs harboring RNA replicons, in the absence of E protein complementation, were non-infectious, even when this VLPs were artificially released from infected cells by freeze-thawing, reinforcing their safety.
4.2. Attenuation of rMERS-CoV-ΔE replicon
Highly susceptible K18-hDPP4 transgenic mice (Li et al., 2016) were intranasally inoculated with rMERS-CoV-ΔE replicon. All mice infected with the parental virus lost weight and died. In contrast, mice inoculated with rMERS-CoV-ΔE replicon survived, and the initial small weight loss was quickly regained, indicating that rMERS-CoV-ΔE replicon was attenuated. Moreover, mice immunized with rMERS-CoV-ΔE survived to a lethal MERS-CoV challenge, and none of them suffered a significant weight loss. These results demonstrated that a single immunization with the rMERS-CoV-ΔE replicon was sufficient to protect against lethal infection in this highly susceptible model of disease.
The deletion of the envelope E protein is key for virus attenuation, since rMERS-CoV-ΔE is propagation deficient (Almazan et al., 2013; Ortego et al., 2007). This characteristic of E− replicons is a consequence of the role of this protein in intracellular transport, virus morphogenesis, and virion release from the cell (DeDiego et al., 2007; Fischer et al., 1998; Gutierrez-Alvarez et al., 2021; Schoeman and Fielding, 2019). Another contribution of the deletion of E gene to biosafety is the elimination of several virulence motifs. One is related to the ability of E protein to form pentamers with ion channel activity (Nieto-Torres et al., 2015). We have previously shown that the introduction of point mutations in the transmembrane domain of SARS-CoV E protein destroys E protein ion channel activity (Nieto-Torres et al., 2014). A second virulence factor present in E protein is the PDZ-binding motif (PBM), which maps at the end of the carboxy-terminus domain of E protein (Jimenez-Guardeño et al., 2014; Regla-Nava et al., 2015). The PBM has a core sequence of four amino acids that can potentially bind to more than 400 cellular proteins, including syntenin (Castaño-Rodriguez et al., 2018; Jimenez-Guardeño et al., 2014). Binding of SARS-CoV E protein to syntenin leads to the activation of p38MAPK by phosphorylation, activating the expression of several cytokines and exacerbating proinflammatory responses (Jimenez-Guardeño et al., 2014). The replacement of these four amino acids by glycine, or its deletion, leads to an attenuated virus (Jimenez-Guardeño et al., 2014; Regla-Nava et al., 2015). Recently, we described that a forkhead-associated binding motif (FHA-BM), present in the carboxy-terminus domain of MERS-CoV E protein, is also probably implicated in virulence (Gutierrez-Alvarez et al., 2021).
4.3. Engineering of MERS-CoV RNA-replicons missing several non-essential genes
In order to increase the safety of the RNA replicon based on rMERS-CoV-ΔE, additional safety features were included by deleting up to five different genes in a mouse-adapted virus background (rMERS-MA30-Δ[3,4a,4b,5,E]). The deletion of additional non-essential genes, such as 3 and 5, in the RNA reduced the possibility of recombination with other human CoVs, such as HCoV-229-E, HCoV-OC43, HCoV-NL-63, HCoV-HKU-1, or SARS-CoV-2 and MERS-like CoVs circulating in the field (Chen et al., 2017; Corman et al., 2014; Lau et al., 2018; Luo et al., 2018; Moreno et al., 2017; Woo et al., 2012; Yang et al., 2014). In addition, the accessory genes are implicated in virulence (Canton et al., 2018; Lee et al., 2019a, b; Nakagawa et al., 2018; Niemeyer et al., 2013; Rabouw et al., 2016; Siu et al., 2014), and their deletion resulted in increased attenuation by a loss of function. In the presence of E protein provided in trans, the absence of the accessory proteins delayed but did not prevent maximal replication. However, in the absence of E protein, the rMERS-MA30-Δ[3,4a,4b,5,E] replicon was propagation-defective, as expected. Furthermore, the stability of the rMERS-MA30-Δ[3,4a,4b,5,E] replicon in cell culture was assessed by evaluating whether it could recombine with an RNA encoding E protein transcribed from an expression plasmid. After five passages in Huh-7, there was no evidence that rMERS-MA30-Δ[3,4a,4b,5,E] regained the E gene by recombining with the RNA encoding the E protein.
The hDPP4-KI mice model (Li et al., 2017) was used to evaluate the pathogenicity of rMERS-MA30-Δ[3,4a,4b,5,E] replicon, compared with the parental rMERS-MA30 virulent virus. All mice inoculated with rMERS-MA30 virus lost weight and died. In contrast, none of the mice infected rMERS-MA30-Δ [3,4a,4b,5,E] RNA replicon lost weight and all of them survived. In line with this data, no significant pathological changes were observed in the lungs of mice infected with rMERS-MA30-Δ[3,4a,4b,5,E], while the lungs of mice infected with the parental virulent virus showed alveolar wall thickening, hyaline membrane formation, generalized infiltration and parenchyma consolidation, as well as edema in the airspaces. Altogether, these results indicated that this replicon was fully attenuated.
4.4. Protection elicited by MERS-MA30 derived replicon in hDPP4-KI mice
To assess whether rMERS-MA30-Δ[3,4a,4b,5,E] would be a useful vaccine, hDPP4-KI mice were intranasally immunized and challenged 21 days post-immunization (dpim) with a lethal dose of MERS-MA30. Non-immunized mice lost weight and died. However, all mice immunized with the replicon survived the challenge, and none of them suffered significant weight loss. Interestingly, no infectious virus was detected in the lungs of mice immunized with rMERS-MA30-Δ[3,4a,4b,5,E] replicon at any time after challenge, indicating that it conferred sterilizing immunity. In fact, while the lungs of non-immunized mice showed hyaline membranes, edema, extensive cellular infiltration and focal hemorrhages, the lungs of immunized mice remained nearly normal in appearance.
Levels of neutralizing antibodies were determined in the serum of mice. At 21 dpim, mice immunized with rMERS-MA30-Δ[3,4a,4b,5,E] replicon showed significative levels of neutralizing antibodies compared with non-immunized mice after one single immunization. The results demonstrated that rMERS-MA30-Δ[3,4a,4b,5,E] replicon induced protection against a lethal dose of MERS-MA30 virus and promoted sterilizing immunity with significant levels of neutralizing antibodies.
5. Next generation vaccine candidates against SARS-CoV-2 based in efficient and safe replication-competent propagation-defective RNA replicons
Several highly effective and licensed SARS-CoV-2 vaccines have been based in the administration of an mRNA, preferentially encoding the S protein of the virus, the major inducer of CoV neutralizing antibodies (Moderna and Pfizer vaccines) (Corbett et al., 2020; Walsh et al., 2020). Unfortunately, these mRNAs are not self-amplifying and we decided to develop a self-replicating one, based on previous results obtained for MERS-CoV, as a vaccine to prevent infections by this virus. These replicons were derived from SARS-CoV-2 genome by deleting genes responsible for virus virulence and dissemination, which are non-essential for virus replication, as in the case of MERS-CoV replicon. To increase the biosafety of SARS-CoV-2 derived replicon, we aimed to the total or partial deletion of replicase and non-replicase genes, mapping at distal domains of the genome, in order to prevent that a single recombination event with circulating CoVs could restore virus virulence and the possibility of dissemination. Previous publications of our group with MERS-CoV replicon as a vaccine supported the viability of the project (Almazan et al., 2013; Gutiérrez-Álvarez et al., 2021). Nevertheless, the identity of the genes deleted in SARS-CoV-2 to render the replicon propagation-deficient was different from those deleted in the case of MERS-CoV replicon (J. M. Honrubia, I. Sola, S. Zuñiga, and L. Enjuanes, manuscript in preparation). Similarly, the set of genes deleted to attenuate SARS-CoV-2 virus was different from those deleted in the case of MERS-CoV, because the nature of the accessory genes in MERS-CoV and SARS-CoV-2 genome is different (Fig. 1).
The most efficient strategy for the generation of an RNA replicon derived from SARS-CoV-2 was the development of VLPs generated in packaging cell lines that trans-complement RNA replicons with the proteins required for replicon propagation (Fig. 4). These replicons included modifications of the replicase nsp1 gene, and the S protein of the virus, additional deletions of two structural virus genes, and other four non-structural genes non-essential for virus replication. A deletion of the furin proteolytic site present in S protein was introduced, in order to drastically reduce the multiorgan tropism of the virus (Johnson et al., 2021). These modifications lead to a multi-attenuated and propagation defective RNA replicon (J. M. Honrubia, I. Sola, S. Zuñiga, and L. Enjuanes, manuscript in preparation).
VLPs containing the self-replicating propagation-defective RNA replicons were used for evaluation in preclinical immunogenicity and protection assays. Evaluation of the protection induced by RNA replicons has been performed in humanized K18-hACE2 transgenic mice by intranasal immunization, as usually done in our laboratory (Castaño-Rodriguez et al., 2018; Gutierrez-Alvarez et al., 2021). The immune response induced at different days post-immunization was evaluated by analyzing antibodies (by ELISA and virus neutralization), and virus-specific CD4+ and CD8+ T cells (by flow cytometry and ELISPOT). The results have shown that SARS-CoV-2 VLPs provided full-protection against virus infections. These results have already been protected by two file applications.
In addition to the transgenic mouse model, hamster and non-human primate animal models are being used to further confirm the efficacy and safety of RNA replicon vaccine candidates. The definitive vaccine candidate will be selected based on the protection elicited against different SARS-CoV-2 variants, including the Omicron ones, and by the absence of adverse effects, and by evaluating the genetic stability of the engineered constructs, both in cell culture and in vivo, in the different animal models.
6. Concluding remarks and future perspectives
Viruses are the major generators of genetic variability in different species, including humans, by interchanging sequences with the infected cells (Hamilton, 2006; Petersen et al., 2020). CoVs, with one of the largest genomes among known RNA viruses, have incorporated a proof-reading system within their replication-transcription machinery (Denison et al., 2011), but still generate a high variability probably favored by the discontinuous RNA synthesis during the production of subgenomic RNAs, a process mediated by a high frequency recombination event (Enjuanes et al., 2006). This mechanism may facilitate the evolution of CoVs, leading to a variety of novel animal and human variants. In fact, the emergence of novel animal and highly pathogenic human CoVs has been widely documented. Since different species of bats that fly across all continents act as a natural reservoir for these CoVs, the emergence or reemergence of deadly CoVs is a likely event for which we should be prepared, among other things, by the development of efficient vaccines. In this manuscript it has been shown how through the application of reverse genetics procedures the engineering of highly safe, CoV-derived RNA replicons resulted in vaccine candidates inducing sterilizing immunity, by a single intranasal dose administration. A similar strategy could be applied, in principle, to the development of vaccines against other highly pathogenic coronaviruses.
RNA replicons combine the advantages of two classic vaccine types. They are almost as safe as inactivated vaccines, as they cannot propagate, and their ability to amplify their genomes generated a protective response as high as the one elicited by live attenuated vaccines, as shown in this review. The attenuation and safety of rMERS-MA30-Δ[3,4a,4b,5,E] replicon could be improved by the introduction of partial deletions within the nsp1 nonstructural protein, as we have shown for the full-length viruses MERS-CoV and SARS-CoV, in which these deletions further attenuated the virus (Regla-Nava et al., 2015). Similarly, additional attenuation of the virus could be increased by mutations in the viral nsp16 protein, the 2-O’ methyl transferase, which is important for immune evasion (Menachery et al., 2017).
The safety of MERS-CoV-based RNA replicons was reinforced by the observation that VLPs harboring RNA replicons in the absence of E protein complementation were non-infectious, even when this VLPs were artificially released from infected cells by freeze-thawing. However, they may induce small syncytia formation in Huh-7 cells fusing less than 16 cells (Gutierrez-Alvarez et al., 2021). It has been shown that the membrane fusion peptide domain of the S2 subunit of the S protein is involved in the induction of syncytia formation (Alsaadi et al., 2019). Mutation of this site in rMERS-MA30-Δ[3,4a,4b,5,E] would most likely block S protein processing and syncytia formation, reducing cell fusion-dependent spread and increasing biosafety.
The insertion of micro-RNAs may modulate or enhance the immune response elicited by the replicon (Waring et al., 2018), opening new avenues for the design of advanced vaccines that may improve the immune response in those individuals with a weakened immune system, such as that of the elderly (Waring et al., 2018). We have previously shown that certain miRNAs can modulate the growth of the virus and attenuate its pathogenicity in the context of SARS-CoV infection (Morales et al., 2017, 2022).
CRediT authorship contribution statement
Conceptualization (L.E., I.S., S.Z.), Validation (L.E., J.M.H.), Investigation (J.M.H., A.S.B., E.G.M., M.B.P., J.H. L.W., R.R., D.M., C.S., A.E, J.R.), Writing – Original draft (L.E., I.S., S.Z.), Writing – Review & editing (L.E., I.S., S.Z., J.M.H., A.S.B., E.G.M., M.B.P., J.H., R.R., L.W., D.M., C.S., A.E, J.R.), Visualization (S.Z.), Supervision (L.E.), Funding acquisition (L.E., I.S.).
Declaration of competing interest
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Luis Enjuanes has patent issued to PCT/ES2021/070378. Luis Enjuanes has patent issued to PCT/EP21383072.2.
Acknowledgements
We thank Marga Gonzalez (CNB–CSIC) for her technical assistance. In vivo experiments were performed at CISA, INIA-CSIC (Madrid, Spain). This work was supported by grants from the Government of Spain (PID2019-107001RB-I00 AEI/FEDER, UE; SEV 2017-0712 and PIE_INTRAMURAL_LINEA 1-202020E079), the CSIC (PIE_INTRAMURAL-202020E043), the European Commission (ISOLDA_848166 H2020-SC1-2019-Two-Stage-RTD, RIA; MANCO_101003651 H2020-SC1-PHE-CORONAVIRUS-2020 RIA), and the U.S. National Institutes of Health (NIH_2P01AI060699).The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- Agnihothram S., Menachery V.D., Yount B.L., Jr., Lindesmith L.C., Scobey T., Whitmore A., Schafer A., Heise M.T., Baric R.S. Development of a broadly accessible Venezuelan equine encephalitis virus replicon particle vaccine platform. J. Virol. 2018;92 doi: 10.1128/JVI.00027-18. 18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Almazan F., DeDiego M.L., Sola I., Zuñiga S., Nieto-Torres J.L., Marquez-Jurado S., Andres G., Enjuanes L. Engineering a replication-competent, propagation-defective Middle East respiratory syndrome coronavirus as a vaccine candidate. mBio. 2013;4 doi: 10.1128/mBio.00650-13. 13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Almazan F., Gonzalez J.M., Penzes Z., Izeta A., Calvo E., Plana-Duran J., Enjuanes L. Engineering the largest RNA virus genome as an infectious bacterial artificial chromosome. Proc. Natl. Acad. Sci. USA. 2000;97:5516–5521. doi: 10.1073/pnas.97.10.5516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alsaadi E.A.J., Neuman B.W., Jones I.M. A fusion peptide in the spike protein of MERS coronavirus. Viruses. 2019;11:825. doi: 10.3390/v11090825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banerjee A., Kulcsar K., Misra V., Frieman M., Mossman K. Bats and coronaviruses. Viruses. 2019;11:41. doi: 10.3390/v11010041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Canton J., Fehr A.R., Fernandez-Delgado R., Gutierrez-Alvarez F.J., Sanchez-Aparicio M.T., Garcia-Sastre A., Perlman S., Enjuanes L., Sola I. MERS-CoV 4b protein interferes with the NF-kappaB-dependent innate immune response during infection. PLoS Pathog. 2018;14 doi: 10.1371/journal.ppat.1006838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castaño-Rodriguez C., Honrubia J.M., Gutierrez-Alvarez J., DeDiego M.L., Nieto-Torres J.L., Jimenez-Guardeno J.M., Regla-Nava J.A., Fernandez-Delgado R., Verdia-Baguena C., Queralt-Martin M., Kochan G., Perlman S., Aguilella V.M., Sola I., Enjuanes L. Role of severe acute respiratory syndrome coronavirus viroporins E, 3a, and 8a in replication and pathogenesis. mBio. 2018;9 doi: 10.1128/mBio.02325-17. 17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen N., Li S., Zhou R., Zhu M., He S., Ye M., Huang Y., Li S., Zhu C., Xia P., Zhu J. Two novel porcine epidemic diarrhea virus (PEDV) recombinants from a natural recombinant and distinct subtypes of PEDV variants. Virus Res. 2017;242:90–95. doi: 10.1016/j.virusres.2017.09.013. [DOI] [PubMed] [Google Scholar]
- Colina S.E., Serena M.S., Echeverria M.G., Metz G.E. Clinical and molecular aspects of veterinary coronaviruses. Virus Res. 2021;297 doi: 10.1016/j.virusres.2021.198382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corbett K.S., Edwards D., Leist S.R., Abiona O.M., Boyoglu-Barnum S., Gillespie R.A., Himansu S., Schafer A., Ziwawo C.T., DiPiazza A.T., Dinnon K.H., Elbashir S.M., Shaw C.A., Woods A., Fritch E.J., Martinez D.R., Bock K.W., Minai M., Nagata B.M., Hutchinson G.B., Bahl K., Garcia-Dominguez D., Ma L., Renzi I., Kong W.P., Schmidt S.D., Wang L., Zhang Y., Stevens L.J., Phung E., Chang L.A., Loomis R.J., Altaras N.E., Narayanan E., Metkar M., Presnyak V., Liu C., Louder M.K., Shi W., Leung K., Yang E.S., West A., Gully K.L., Wang N., Wrapp D., Doria-Rose N.A., Stewart-Jones G., Bennett H., Nason M.C., Ruckwardt T.J., McLellan J.S., Denison M.R., Chappell J.D., Moore I.N., Morabito K.M., Mascola J.R., Baric R.S., Carfi A., Graham B.S. SARS-CoV-2 mRNA vaccine development enabled by prototype pathogen preparedness. Nature. 2020;586:567–571. doi: 10.1038/s41586-020-2622-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corman V.M., Ithete N.L., Richards L.R., Schoeman M.C., Preiser W., Drosten C., Drexler J.F. Rooting the phylogenetic tree of middle East respiratory syndrome coronavirus by characterization of a conspecific virus from an African bat. J. Virol. 2014;88:11297–11303. doi: 10.1128/JVI.01498-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corman V.M., Muth D., Niemeyer D., Drosten C. Hosts and sources of endemic human coronaviruses. Adv. Virus Res. 2018;100:163–188. doi: 10.1016/bs.aivir.2018.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cui J., Li F., Shi Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019;17:181–192. doi: 10.1038/s41579-018-0118-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das A.T., Tenenbaum L., Berkhout B. Tet-on systems for doxycycline-inducible gene expression. Curr. Gene Ther. 2016;16:156–167. doi: 10.2174/1566523216666160524144041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Groot R.J., Baker S.C., Baric R.S., Brown C.S., Drosten C., Enjuanes L., Fouchier R.A., Galiano M., Gorbalenya A.E., Memish Z.A., Perlman S., Poon L.L., Snijder E.J., Stephens G.M., Woo P.C., Zaki A.M., Zambon M., Ziebuhr J. Middle East respiratory syndrome coronavirus (MERS-CoV): announcement of the Coronavirus Study Group. J. Virol. 2013;87:7790–7792. doi: 10.1128/JVI.01244-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Wit E., van Doremalen N., Falzarano D., Munster V.J. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016;14:523–534. doi: 10.1038/nrmicro.2016.81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeDiego M.L., Alvarez E., Almazan F., Rejas M.T., Lamirande E., Roberts A., Shieh W.J., Zaki S.R., Subbarao K., Enjuanes L. A severe acute respiratory syndrome coronavirus that lacks the E gene is attenuated in vitro and in vivo. J. Virol. 2007;81:1701–1713. doi: 10.1128/JVI.01467-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denison M.R., Graham R.L., Donaldson E.F., Eckerle L.D., Baric R.S. Coronaviruses: an RNA proofreading machine regulates replication fidelity and diversity. RNA Biol. 2011;8:270–279. doi: 10.4161/rna.8.2.15013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drosten C., Gunther S., Preiser W., van der Werf S., Brodt H.R., Becker S., Rabenau H., Panning M., Kolesnikova L., Fouchier R.A., Berger A., Burguiere A.M., Cinatl J., Eickmann M., Escriou N., Grywna K., Kramme S., Manuguerra J.C., Muller S., Rickerts V., Sturmer M., Vieth S., Klenk H.D., Osterhaus A.D., Schmitz H., Doerr H.W. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003;348:1967–1976. doi: 10.1056/NEJMoa030747. [DOI] [PubMed] [Google Scholar]
- Enjuanes L., Almazan F., Sola I., Zuñiga S. Biochemical aspects of coronavirus replication and virus-host interaction. Annu. Rev. Microbiol. 2006;60:211–230. doi: 10.1146/annurev.micro.60.080805.142157. [DOI] [PubMed] [Google Scholar]
- Enjuanes L., Gorbalenya A.E., de Groot R.J., Cowley J.A., Ziebuhr J., Snijder E.J. In: Encyclopedia of Virology. third ed. Mahy B.W.J., Van Regenmortel M., Walker P., Majumder-Russell D., editors. Elsevier Ltd.; Oxford: 2008. The Nidovirales; pp. 419–430. [Google Scholar]
- Firth A.E., Brierley I. Non-canonical translation in RNA viruses. J. Gen. Virol. 2012;93:1385–1409. doi: 10.1099/vir.0.042499-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fischer F., Stegen C.F., Masters P.S., Samsonoff W.A. Analysis of constructed E gene mutants of mouse hepatitis virus confirms a pivotal role for E protein in coronavirus assembly. J. Virol. 1998;72:7885–7894. doi: 10.1128/jvi.72.10.7885-7894.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Folegatti P.M., Bittaye M., Flaxman A., Lopez F.R., Bellamy D., Kupke A., Mair C., Makinson R., Sheridan J., Rohde C., Halwe S., Jeong Y., Park Y.S., Kim J.O., Song M., Boyd A., Tran N., Silman D., Poulton I., Datoo M., Marshall J., Themistocleous Y., Lawrie A., Roberts R., Berrie E., Becker S., Lambe T., Hill A., Ewer K., Gilbert S. Safety and immunogenicity of a candidate Middle East respiratory syndrome coronavirus viral-vectored vaccine: a dose-escalation, open-label, non-randomised, uncontrolled, phase 1 trial. Lancet Infect. Dis. 2020;20:816–826. doi: 10.1016/S1473-3099(20)30160-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao G., Liu W.J., Liu P., Lei W., Jia Z., He X., Liu L.-L., Shi W., Tan Y., Zou S., Zhao X., Wong G., Wang J., Wang F., Wang G., Qin K., Gao R., Zhang J., Li M., Xiao W., Guo Y., Xu Z., Zhao Y., Song J., Zhang J., Zhen W., Zhou W., Ye B., Song J., Yang M., Zhou W., Bi Y., Cai K., Wang D., Tan W., Han J., Xu W., Gao G.F., Wu G. Surveillance of SARS-CoV-2 in the environment and animal samples of the huanan seafood market. Preprint Res. Square. 2022 doi: 10.21203/rs.21203.rs-1370392/v1370391. [DOI] [Google Scholar]
- Gorbalenya A.E., Baker S.C., Baric R.S., de Groot R.J., Drosten C., Gulyaeva A.A., Haagmans B.L., Lauber C., Leontovich A.M., Neuman B.W., Penzar D., Perlman S., Poon L.L.M., Samborskiy D.V., Sidorov I.A., Sola I., Ziebuhr J., Coronaviridae Study Group of the International Committee on Taxonomy of, V The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020;5:536–544. doi: 10.1038/s41564-020-0695-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gutierrez-Alvarez J., Honrubia J.M., Fernandez-Delgado R., Wang L., Castaño-Rodriguez C., Zuñiga S., Sola I., Enjuanes L. Genetically engineered live-attenuated Middle East Respiratory Syndrome Coronavirus viruses confer full protection against lethal infection. mBio. 2021;12 doi: 10.1128/mBio.00103-21. e00121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gutiérrez-Álvarez J., Honrubia J.M., Sanz-Bravo A., González-Miranda E., Fernández-Delgado R., Rejas M.T., Zúñiga S., Sola I., Enjuanes L. Middle East respiratory syndrome coronavirus vaccine based on a propagation-defective RNA replicon elicited sterilizing immunity in mice. Proc. Natl. Acad. Sci. USA. 2021;118 doi: 10.1073/pnas.2111075118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hale V.L., Dennis P.M., McBride D.S., Nolting J.M., Madden C., Huey D., Ehrlich M., Grieser J., Winston J., Lombardi D., Gibson S., Saif L., Killian M.L., Lantz K., Tell R.M., Torchetti M., Robbe-Austerman S., Nelson M.I., Faith S.A., Bowman A.S. SARS-CoV-2 infection in free-ranging white-tailed deer. Nature. 2022;602:481–486. doi: 10.1038/s41586-021-04353-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamilton G. Virology: the gene weavers. Nature. 2006;441:683–685. doi: 10.1038/441683a. [DOI] [PubMed] [Google Scholar]
- Holmes E.C., Goldstein S.A., Rasmussen A.L., Robertson D.L., Crits-Christoph A., Wertheim J.O., Anthony S.J., Barclay W.S., Boni M.F., Doherty P.C., Farrar J., Geoghegan J.L., Jiang X., Leibowitz J.L., Neil S.J.D., Skern T., Weiss S.R., Worobey M., Andersen K.G., Garry R.F., Rambaut A. The origins of SARS-CoV-2: a critical review. Cell. 2021;184:4848–4856. doi: 10.1016/j.cell.2021.08.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu B., Guo H., Zhou P., Shi Z.L. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 2021;19:141–154. doi: 10.1038/s41579-020-00459-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ICTV . 2019. International Committee on Taxonomy of Viruses: Master Species List 2019.V1.https://talk.ictvonline.org/files/master-species-lists/m/msl/9601 [Google Scholar]
- Jimenez-Guardeño J.M., Nieto-Torres J.L., DeDiego M.L., Regla-Nava J.A., Fernandez-Delgado R., Castaño-Rodriguez C., Enjuanes L. The PDZ-binding motif of severe acute respiratory syndrome coronavirus envelope protein Is a determinant of viral pathogenesis. PLoS Pathog. 2014;10 doi: 10.1371/journal.ppat.1004320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson B.A., Xie X., Bailey A.L., Kalveram B., Lokugamage K.G., Muruato A., Zou J., Zhang X., Juelich T., Smith J.K., Zhang L., Bopp N., Schindewolf C., Vu M., Vanderheiden A., Winkler E.S., Swetnam D., Plante J.A., Aguilar P., Plante K.S., Popov V., Lee B., Weaver S.C., Suthar M.S., Routh A.L., Ren P., Ku Z., An Z., Debbink K., Diamond M.S., Shi P.Y., Freiberg A.N., Menachery V.D. Loss of furin cleavage site attenuates SARS-CoV-2 pathogenesis. Nature. 2021;591:293–299. doi: 10.1038/s41586-021-03237-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jurgens C.K., Young K.R., Madden V.J., Johnson P.R., Johnston R.E. A novel self-replicating chimeric lentivirus-like particle. J. Virol. 2012;86:246–261. doi: 10.1128/JVI.05191-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koch T., Dahlke C., Fathi A., Kupke A., Krahling V., Okba N.M.A., Halwe S., Rohde C., Eickmann M., Volz A., Hesterkamp T., Jambrecina A., Borregaard S., Ly M.L., Zinser M.E., Bartels E., Poetsch J.S.H., Neumann R., Fux R., Schmiedel S., Lohse A.W., Haagmans B.L., Sutter G., Becker S., Addo M.M. Safety and immunogenicity of a modified vaccinia virus Ankara vector vaccine candidate for Middle East respiratory syndrome: an open-label, phase 1 trial. Lancet Infect. Dis. 2020;20:827–838. doi: 10.1016/S1473-3099(20)30248-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lau S.K.P., Zhang L., Luk H.K.H., Xiong L., Peng X., Li K.S.M., He X., Zhao P.S., Fan R.Y.Y., Wong A.C.P., Ahmed S.S., Cai J.P., Chan J.F.W., Sun Y., Jin D., Chen H., Lau T.C.K., Kok R.K.H., Li W., Yuen K.Y., Woo P.C.Y. Receptor usage of a novel bat lineage C betacoronavirus reveals evolution of Middle East respiratory syndrome-related coronavirus spike proteins for human dipeptidyl peptidase 4 binding. J. Infect. Dis. 2018;218:197–207. doi: 10.1093/infdis/jiy018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee J.Y., Bae S., Myoung J. Middle East respiratory syndrome coronavirus-encoded accessory proteins impair MDA5-and TBK1-mediated activation of NF-kappaB. J. Microbiol. Biotechnol. 2019;29:1316–1323. doi: 10.4014/jmb.1908.08004. [DOI] [PubMed] [Google Scholar]
- Lee J.Y., Bae S., Myoung J. Middle East respiratory syndrome coronavirus-encoded ORF8b strongly antagonizes IFN-beta promoter activation: its implication for vaccine design. J. Microbiol. 2019;57:803–811. doi: 10.1007/s12275-019-9272-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li K., Wohlford-Lenane C., Perlman S., Zhao J., Jewell A.K., Reznikov L.R., Gibson-Corley K.N., Meyerholz D.K., McCray P.B., Jr. Middle East respiratory syndrome coronavirus causes multiple organ damage and lethal disease in mice transgenic for human dipeptidyl peptidase 4. J. Infect. Dis. 2016;213:712–722. doi: 10.1093/infdis/jiv499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li K., Wohlford-Lenane C.L., Channappanavar R., Park J.E., Earnest J.T., Bair T.B., Bates A.M., Brogden K.A., Flaherty H.A., Gallagher T., Meyerholz D.K., Perlman S., McCray P.B., Jr. Mouse-adapted MERS coronavirus causes lethal lung disease in human DPP4 knockin mice. Proc. Natl. Acad. Sci. USA. 2017;114:E3119–E3128. doi: 10.1073/pnas.1619109114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liljestrom P., Garoff H. A new generation of animal cell expression vectors based on the Semliki Forest virus replicon. Biotechnology. 1991;9:1356–1361. doi: 10.1038/nbt1291-1356. [DOI] [PubMed] [Google Scholar]
- Lundstrom K. Alphavirus-based vaccines. Methods Mol. Biol. 2016;1404:313–328. doi: 10.1007/978-1-4939-3389-1_22. [DOI] [PubMed] [Google Scholar]
- Lundstrom K. Self-amplifying RNA viruses as RNA vaccines. Int. J. Mol. Sci. 2020;21:5130. doi: 10.3390/ijms21145130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo C.M., Wang N., Yang X.L., Liu H.Z., Zhang W., Li B., Hu B., Peng C., Geng Q.B., Zhu G.J., Li F., Shi Z.L. Discovery of novel bat coronaviruses in south China that use the same receptor as Middle East respiratory syndrome coronavirus. J. Virol. 2018;92 doi: 10.1128/JVI.00116-18. e00118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masters P.S. The molecular biology of coronaviruses. Adv. Virus Res. 2006;66:193–292. doi: 10.1016/S0065-3527(06)66005-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxmen A. Wuhan market was epicentre of pandemic's start, studies suggest. Nature. 2022;603:15–16. doi: 10.1038/d41586-022-00584-8. [DOI] [PubMed] [Google Scholar]
- Menachery V.D., Gralinski L.E., Mitchell H.D., Dinnon K.H., 3rd, Leist S.R., Yount B.L., Jr., Graham R.L., McAnarney E.T., Stratton K.G., Cockrell A.S., Debbink K., Sims A.C., Waters K.M., Baric R.S. Middle East respiratory syndrome coronavirus nonstructural protein 16 is necessary for interferon resistance and viral pathogenesis. mSphere. 2017;2 doi: 10.1128/mSphere.00346-17. 17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Modjarrad K., Roberts C.C., Mills K.T., Castellano A.R., Paolino K., Muthumani K., Reuschel E.L., Robb M.L., Racine T., Oh M.D., Lamarre C., Zaidi F.I., Boyer J., Kudchodkar S.B., Jeong M., Darden J.M., Park Y.K., Scott P.T., Remigio C., Parikh A.P., Wise M.C., Patel A., Duperret E.K., Kim K.Y., Choi H., White S., Bagarazzi M., May J.M., Kane D., Lee H., Kobinger G., Michael N.L., Weiner D.B., Thomas S.J., Maslow J.N. Safety and immunogenicity of an anti-Middle East respiratory syndrome coronavirus DNA vaccine: a phase 1, open-label, single-arm, dose-escalation trial. Lancet Infect. Dis. 2019;19:1013–1022. doi: 10.1016/S1473-3099(19)30266-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morales L., Oliveros J.C., Enjuanes L., Sola I. Contribution of host miRNA-223-3p to SARS-CoV-induced lung inflammatory pathology. mBio. 2022;13 doi: 10.1128/mbio.03135-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morales L., Oliveros J.C., Fernandez-Delgado R., tenOever B.R., Enjuanes L., Sola I. SARS-CoV-encoded small RNAs contribute to infection-associated lung pathology. Cell Host Microbe. 2017;21:344–355. doi: 10.1016/j.chom.2017.01.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moreno A., Lelli D., de Sabato L., Zaccaria G., Boni A., Sozzi E., Prosperi A., Lavazza A., Cella E., Castrucci M.R., Ciccozzi M., Vaccari G. Detection and full genome characterization of two beta CoV viruses related to Middle East respiratory syndrome from bats in Italy. Virol. J. 2017;14:239. doi: 10.1186/s12985-017-0907-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mou H., Raj V.S., van Kuppeveld F.J., Rottier P.J., Haagmans B.L., Bosch B.J. The receptor binding domain of the new MERS coronavirus maps to a 231-residue region in the spike protein that efficiently elicits neutralizing antibodies. J. Virol. 2013;87:9379–9383. doi: 10.1128/JVI.01277-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagy P.D., Simon A.E. New insights into the mechanisms of RNA recombination. Virology. 1997;235:1–9. doi: 10.1006/viro.1997.8681. [DOI] [PubMed] [Google Scholar]
- Nakagawa K., Narayanan K., Wada M., Makino S. Inhibition of stress granule formation by Middle East respiratory syndrome coronavirus 4a accessory protein facilitates viral translation, leading to efficient virus replication. J. Virol. 2018;92 doi: 10.1128/JVI.00902-18. e00918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- NCT . 2019. Safety and Immunogenicity of the Candidate Vaccine MVA-MERS-S_DF-1 against MERS (MVA-MERS-S)https://clinicaltrials.gov/ct2/show/NCT04119440?term=vaccine&recrs=abe&cond=MERS+%28Middle+East+Respiratory+Syndrome%29&draw=2 [Google Scholar]
- NCT . 2019. Study of Safety and Immunogenicity of BVRS‐GamVac‐Combi.https://clinicaltrials.gov/ct2/show/NCT04128059?term=vaccine&recrs=abe&cond=MERS+%28Middle+East+Respiratory+Syndrome%29&draw=2&rank=6 [Google Scholar]
- NCT . 2019. Study of Safety and Immunogenicity of BVRS‐GamVac.https://clinicaltrials.gov/ct2/show/NCT04130594?term=vaccine&recrs=abe&cond=MERS+%28Middle+East+Respiratory+Syndrome%29&draw=2&rank=5 [Google Scholar]
- Niemeyer D., Zillinger T., Muth D., Zielecki F., Horvath G., Suliman T., Barchet W., Weber F., Drosten C., Muller M.A. Middle East respiratory syndrome coronavirus accessory protein 4a is a type I interferon antagonist. J. Virol. 2013;87:12489–12495. doi: 10.1128/JVI.01845-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nieto-Torres J.L., Dediego M.L., Verdia-Baguena C., Jimenez-Guardeño J.M., Regla-Nava J.A., Fernandez-Delgado R., Castaño-Rodriguez C., Alcaraz A., Torres J., Aguilella V.M., Enjuanes L. Severe acute respiratory syndrome coronavirus envelope protein ion channel activity promotes virus fitness and pathogenesis. PLoS Pathog. 2014;10 doi: 10.1371/journal.ppat.1004077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nieto-Torres J.L., Verdia-Baguena C., Castaño-Rodriguez C., Aguilella V.M., Enjuanes L. Relevance of viroporin ion channel activity on viral replication and pathogenesis. Viruses. 2015;7:3552–3573. doi: 10.3390/v7072786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Omrani A.S., Al-Tawfiq J.A., Memish Z.A. Middle East respiratory syndrome coronavirus (MERS-CoV): animal to human interaction. Pathog. Glob. Health. 2015;109:354–362. doi: 10.1080/20477724.2015.1122852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ortego J., Ceriani J.E., Patino C., Plana J., Enjuanes L. Absence of E protein arrests transmissible gastroenteritis coronavirus maturation in the secretory pathway. Virology. 2007;368:296–308. doi: 10.1016/j.virol.2007.05.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pekar J.E., Magee A., Parker E., Moshiri N., Izhikevich K., Havens J.L., Gangavarapu K., Malpica Serrano L.M., Crits-Christoph A., Matteson N.L., Zeller M., Levy J.I., Wang J.C., Hughes S., Lee J., Park H., Park M.-S., Ching Zi Yan K., Tzer Pin Lin R., Mat Isa M.N., Muhammad Noor Y., Vasylyeva T.I., Garry R.F., Holmes E.C., Rambaut A., Suchard M.A., Andersen K.G., Worobey M., Wertheim J.O. SARS-CoV-2 emergence very likely resulted from at least two zoonotic events. 2022. Zenodo. [DOI]
- Perlman S., Netland J. Coronaviruses post-SARS: update on replication and pathogenesis. Nat. Rev. Microbiol. 2009;7:439–450. doi: 10.1038/nrmicro2147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petersen E., Koopmans M., Go U., Hamer D.H., Petrosillo N., Castelli F., Storgaard M., Al Khalili S., Simonsen L. Comparing SARS-CoV-2 with SARS-CoV and influenza pandemics. Lancet Infect. Dis. 2020;20:e238–e244. doi: 10.1016/S1473-3099(20)30484-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polak S.B., Van Gool I.C., Cohen D., von der Thusen J.H., van Paassen J. A systematic review of pathological findings in COVID-19: a pathophysiological timeline and possible mechanisms of disease progression. Mod. Pathol. 2020;33:2128–2138. doi: 10.1038/s41379-020-0603-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rabouw H.H., Langereis M.A., Knaap R.C., Dalebout T.J., Canton J., Sola I., Enjuanes L., Bredenbeek P.J., Kikkert M., de Groot R.J., van Kuppeveld F.J. Middle East respiratory coronavirus accessory protein 4a inhibits PKR-mediated antiviral stress responses. PLoS Pathog. 2016;12 doi: 10.1371/journal.ppat.1005982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raj V.S., Mou H., Smits S.L., Dekkers D.H., Muller M.A., Dijkman R., Muth D., Demmers J.A., Zaki A., Fouchier R.A., Thiel V., Drosten C., Rottier P.J., Osterhaus A.D., Bosch B.J., Haagmans B.L. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature. 2013;495:251–254. doi: 10.1038/nature12005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Regla-Nava J.A., Nieto-Torres J.L., Jimenez-Guardeno J.M., Fernandez-Delgado R., Fett C., Castaño-Rodriguez C., Perlman S., Enjuanes L., DeDiego M.L. Severe acute respiratory syndrome coronaviruses with mutations in the E protein are attenuated and promising vaccine candidates. J. Virol. 2015;89:3870–3887. doi: 10.1128/JVI.03566-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rota P.A., Oberste M.S., Monroe S.S., Nix W.A., Campganoli R., Icenogle J.P., Peñaranda S., Bankamp B., Maher K., Chen M.-H., Tong S., Tamin A., Lowe L., Frace M., DeRisi J.L., Chen Q., Wang D., Erdman D.d., Peret T.C.T., Burns C., Ksiazek T.G., Rollin P.E., Sanchez A., Liffick S., Holloway B., Limor J., McCaustland K., Olsen-Rassmussen M., Fouchier R., Gunther S., Osterhaus A.D.M.E., Drosten C., Pallansch M.A., Anderson L.J., Bellini W.J. Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science. 2003;300:1394–1399. doi: 10.1126/science.1085952. [DOI] [PubMed] [Google Scholar]
- Sanchez-Paulete A.R., Teijeira A., Quetglas J.I., Rodriguez-Ruiz M.E., Sanchez-Arraez A., Labiano S., Etxeberria I., Azpilikueta A., Bolanos E., Ballesteros-Briones M.C., Casares N., Quezada S.A., Berraondo P., Sancho D., Smerdou C., Melero I. Intratumoral immunotherapy with XCL1 and sFlt3L encoded in recombinant Semliki Forest Virus-derived vectors fosters dendritic cell-mediated T-cell cross-priming. Cancer Res. 2018;78:6643–6654. doi: 10.1158/0008-5472.CAN-18-0933. [DOI] [PubMed] [Google Scholar]
- Sawicki S.G., Sawicki D.L., Siddell S.G. A contemporary view of coronavirus transcription. J. Virol. 2007;81:20–29. doi: 10.1128/JVI.01358-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schoeman D., Fielding B.C. Coronavirus envelope protein: current knowledge. Virol. J. 2019;16:69. doi: 10.1186/s12985-019-1182-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharun K., Dhama K., Pawde A.M., Gortazar C., Tiwari R., Bonilla-Aldana D.K., Rodriguez-Morales A.J., de la Fuente J., Michalak I., Attia Y.A. SARS-CoV-2 in animals: potential for unknown reservoir hosts and public health implications. Vet. Q. 2021;41:181–201. doi: 10.1080/01652176.2021.1921311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharun K., Tiwari R., Saied A.A., Dhama K. SARS-CoV-2 vaccine for domestic and captive animals: an effort to counter COVID-19 pandemic at the human-animal interface. Vaccine. 2021;39:7119–7122. doi: 10.1016/j.vaccine.2021.10.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siu K.L., Yeung M.L., Kok K.H., Yuen K.S., Kew C., Lui P.Y., Chan C.P., Tse H., Woo P.C., Yuen K.Y., Jin D.Y. Middle East respiratory syndrome coronavirus 4a protein is a double-stranded RNA-binding protein that suppresses PACT-induced activation of RIG-I and MDA5 in innate antiviral response. J. Virol. 2014;88:4866–4876. doi: 10.1128/JVI.03649-13. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- Smith E.C., Sexton N.R., Denison M.R. Thinking outside the triangle: replication fidelity of the largest RNA viruses. Ann. Rev. Virol. 2014;1:111–132. doi: 10.1146/annurev-virology-031413-085507. [DOI] [PubMed] [Google Scholar]
- Sola I., Almazán F., Zúñiga S., Enjuanes L. Continuous and discontinuous RNA synthesis in coronaviruses. Annu. Rev. Virol. 2015;2:265–288. doi: 10.1146/annurev-virology-100114-055218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van den Brand J.M., Smits S.L., Haagmans B.L. Pathogenesis of Middle East respiratory syndrome coronavirus. J. Pathol. 2015;235:175–184. doi: 10.1002/path.4458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Most R.G., Spaan W.J.M. In: The Coronaviridae. Siddell S.G., editor. Plenum Press; New York: 1995. Coronavirus replication, transcription, and RNA recombination; pp. 11–31. [Google Scholar]
- Vogel A.B., Lambert L., Kinnear E., Busse D., Erbar S., Reuter K.C., Wicke L., Perkovic M., Beissert T., Haas H., Reece S.T., Sahin U., Tregoning J.S. Self-aplifying RNA vaccines give equivalent protection against Influenza to mRNA vaccines but at much lower doses. Mol. Ther. 2018;26:446–455. doi: 10.1016/j.ymthe.2017.11.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walsh E.E., Frenck R.W., Jr., Falsey A.R., Kitchin N., Absalon J., Gurtman A., Lockhart S., Neuzil K., Mulligan M.J., Bailey R., Swanson K.A., Li P., Koury K., Kalina W., Cooper D., Fontes-Garfias C., Shi P.Y., Tureci O., Tompkins K.R., Lyke K.E., Raabe V., Dormitzer P.R., Jansen K.U., Sahin U., Gruber W.C. Safety and immunogenicity of two RNA-based covid-19 vaccine candidates. N. Engl. J. Med. 2020;383:2439–2450. doi: 10.1056/NEJMoa2027906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waring B.M., Sjaastad L.E., Fiege J.K., Fay E.J., Reyes I., Moriarity B., Langlois R.A. MicroRNA-based attenuation of influenza virus across susceptible hosts. J. Virol. 2018;92 doi: 10.1128/JVI.01741-17. 17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woo P.C., Lau S.K., Li K.S., Tsang A.K., Yuen K.Y. Genetic relatedness of the novel human group C betacoronavirus to Tylonycteris bat coronavirus HKU4 and Pipistrellus bat coronavirus HKU5. Emerg. Microb. Infect. 2012;1:e35. doi: 10.1038/emi.2012.45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Worobey M., Levy J.I., Malpica Serrano L.M., Crits-Christoph A., Pekar J.E., Goldstein S.A., Rasmussen A.L., Kraemer M.U.G., Newman C., Koopmans M.P.G., Suchard M.A., Wertheim J.O., Lemey P., Robertson D.L., Garry R.F., Holmes E.C., Rambaut A., Andersen K.G. The Huanan market was the epicenter of SARS-CoV-2 emergence. Preprint at Zenodo. 2022 doi: 10.5281/zenodo.6299600. [DOI] [Google Scholar]
- Yang L., Wu Z., Ren X., Yang F., Zhang J., He G., Dong J., Sun L., Zhu Y., Zhang S., Jin Q. MERS-related betacoronavirus in Vespertilio superans bats, China. Emerg. Infect. Dis. 2014;20:1260–1262. doi: 10.3201/eid2007.140318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaki A.M., van Boheemen S., Bestebroer T.M., Osterhaus A.D., Fouchier R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012;367:1814–1820. doi: 10.1056/NEJMoa1211721. [DOI] [PubMed] [Google Scholar]
- Zappulli V., Ferro S., Bonsembiante F., Brocca G., Calore A., Cavicchioli L., Centelleghe C., Corazzola G., De Vreese S., Gelain M.E., Mazzariol S., Moccia V., Rensi N., Sammarco A., Torrigiani F., Verin R., Castagnaro M. Pathology of coronavirus infections: a review of lesions in animals in the one-health perspective. Animals (Basel) 2020;10:2377. doi: 10.3390/ani10122377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu N., Zhang D., Wang W., Li X., Yang B., Song J., Zhao X., Huang B., Shi W., Lu R., Niu P., Zhan F., Ma X., Wang D., Xu W., Wu G., Gao G.F., Tan W., China Novel Coronavirus I., Research T. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020;382:727–733. doi: 10.1056/NEJMoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]