Abstract
Despite low mutational burden, immune checkpoint inhibitors have demonstrated promising results in a significant minority of hepatocellular carcinoma (HCC) patients with advanced disease. We hypothesized that HCC patients with higher levels of CD8+ T cell infiltration reflect an immune-inflamed cohort which has improved oncologic outcomes. 355 HCC patients with clinical and transcriptome data in the Cancer Genome Atlas (TCGA) and 151 HCC patients from cohort GSE7624 were analyzed. xCell computational algorithm was used to analyze immune cell infiltration in these patients. Each cohort was divided into high and low expression by the highest 2 terciles value. Gene Set Enrichment Analysis was performed to identify enriched gene sets. High CD8 score associated with improved overall survival in both cohorts (both P < 0.05). High score correlates with early BCLC stage (P = 0.035) but not AJCC stage. High CD8 also correlated with increased IFN-γ response (p = 0.038), lymphocyte infiltration (P < 0.001), and leukocyte fraction (P < 0.001). It was associated with increased polyclonality of T cell (P < 0.001) and B cell response (P = 0.017). High CD8 score correlated with increased cytolytic activity score (P < 0.001) and expression of multiple immune checkpoints including PD-1, PD-L1, CTLA-4 and Lag3 (all P < 0.001). There was no correlation to tumor mutational burden and neoantigens. GSEA demonstrated upregulation of several gene sets involved in inflammatory response and IFN-γ response. In conclusion, HCC patients with high CD8 score demonstrated favorable oncologic outcomes, which may be due to immune-mediated tumor cell attack. Furthermore, CD8 score may be a potentially useful biomarker to select patients for immune checkpoint inhibition.
Keywords: Hepatocellular carcinoma, CD8 T cell score, T cell immunity, checkpoint blockade, tumor mutational burden
Introduction
Hepatocellular carcinoma (HCC) is a primary liver malignancy with the second highest lethality of all cancers [1,2]. In its advanced form prognosis is dismal and even when localized, curative-intent liver-directed therapy is followed by recurrence more often than not. Recently, a novel combination of immune checkpoint inhibition with antiangiogenic therapy led to an unprecedented 30% response rate and a 5% complete response rate in patients with unresectable HCC [3]. Following T-cell immune attack, tumor cells develop ‘adaptive immunoresistance’, upregulating inhibitory pathways to limit further T cell effector function [4-6]. Antibodies targeting immune checkpoint pathways - called checkpoint blockade (CPB) - disrupt this T cell inhibition, ‘releasing the brake’ on the tumor-targeted T cell response. CPB is the primary tool for immune activation against a wide range of solid tumors and has led to dramatic, durable responses in clinical scenarios previously considered uniformly fatal [7-9].
Biomarkers of response to CPB include tumor tissue expression of Programmed Death-Ligand 1 (PD-L1) [10-12], immunohistochemical identification of tumor-infiltrating T cells [13,14], and various measures of tumor mutational burden [15]. A particularly strong correlation exists between CPB response and both quantity and quality of neoantigens [16,17], which are uniquely capable of strong T cell activation due to their recognition as foreign antigen. Yet for HCC, the tumor mutational burden is low. That nonetheless there is a significant minority of patients who respond to CPB speaks to a cohort of patients with ‘hot’ immune inflamed tumors with an initiated and active yet suppressed T-cell mediated immune attack. CD8+ T cells are a primary effector of tumor-targeted immunity and one of the major T cell subsets responsible for clinical response to CPB therapy. We have previously demonstrated that RNA expressional measures of T cell immunity are correlated to oncologic outcomes in multiple solid tumor histologies [18-20]. These measures of T cell immunity correlate to T cell effector gene sets and other immune cell subsets as part of a comprehensive evaluation of the tumor environment that informs prognostic significance.
In this manuscript, we identify a cohort of patients for whom a tumor-targeted immune response exists in the absence of high tumor mutational burden, identifying a cohort of HCC patients with favorable overall survival and which may by particularly susceptible to treatment with immune therapies.
Methods
Clinical and transcriptomic data collection for hepatocellular carcinoma patients
In this study, 355 patients with HCC in The Cancer Genome Atlas (TCGA) [21] cohort were analyzed. Normalized and log2-transformed gene expression data were obtained from cBio Cancer Genomic Portal. We obtained the pathological grade data for the TCGA tumors using Text Information extraction System (TIES) Cancer Research Network, as described previously [22-25]. Further, in order to validate findings in a second cohort, 115 patients of the GSE76427 were obtained from the Gene Expression Omnibus (GEO) database (GEO: https://www.ncbi.nlm.nih.gov/geo/). CD8 score association with degree of liver pathology (normal, pre-malignant, and malignant liver) was assessed in both GSE6764 [26] and GSE89377 [27] cohorts. Probe-level expression values were summarized using mean to obtain gene expression values. Given that the patient data used in this study, TCGA and GEO cohorts, are all de-identified and are in the public domain, Institutional Review Board approval was waived.
Tumor immune microenvironment analysis
xCell algorithm [28] was used to examine whole-tumor gene expression data to score the relative abundance across tumors of 64 types of immune and stromal cells, as we previously described [29-34]. The CD8 T cell of xCell algorithm was used as the CD8 T cell score in this study in the same manner as was done for fibroblast [29] and adipocyte [33] previously.
Gene set enrichment analysis
To explore signaling pathways enrichment, Gene Set Enrichment Analysis (GSEA) [35] was performed between low and high CD8 T cells score groups using GSEA Java software (https://www.gsea-msigdb.org/gsea/index.jsp version 4.0) with MSigDb Hallmark gene sets [36]. A false discovery rate (FDR) of less than 0.25 was used to deem statistical significance, as recommended by the GSEA.
Other statistical analyses
All analyses and data plotting were performed using R software (https://www.r-project.org/version 4.0.1, R Project for Statistical Computing) and Microsoft Excel (version 16, Redmond, WA, USA) for Windows. All depicted boxplots are of Tukey type, showing medians and inter-quartile ranges. One-way analysis of variance (ANOVA) or Fisher’s exact tests were used to compare group means. The third tertile of the CD8 T cell score was used to divide patients into low and high groups (high = upper 1/3 tertile). Survival among groups was compared using the Kaplan-Meier plot with the log-rank test. Cox proportional-hazards regression models tested hazard ratios of CD8 T cell score and measures of tumor mutational burden association with overall survival. A p value less than 0.05 was considered statistically significant. All experimental protocols were approved by institutional IRB and meet the guidelines of their responsible governmental agency.
Results
CD8 score is associated with overall survival, and with very early stage HCC and pre-malignant liver
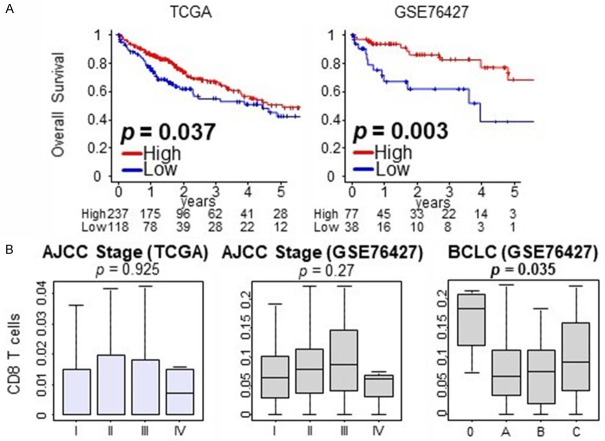
We analyzed gene expression of bulk tumors using the xCell algorithm [28]. The genes that xCell uses for scoring CD8 T cells are listed in Table S1. To establish the clinical relevance of CD8 T-cell infiltration as measured by CD8 score, we first examined the association of the CD8 score with overall survival (OS). After dichotomizing CD8 score to high and low groups using the score’s lower tertile value for each cohort (bottom tertile vs top two tertiles), we found a positive association between high CD8 score and OS in both TCGA and GSE76427 cohorts (Figure 1A, both P < 0.05).
Figure 1.
A. Correlation of CD8 T Cell score with overall survival in TCGA and GSE76427 patient cohorts in hepatocellular carcinoma. B. Correlation of CD8 T cell score with AJCC and BCLC staging systems.
Upon evaluating CD8 score (as a continuous variable) association with HCC stage - either American Joint Committee on Cancer staging (AJCC, available for both cohorts) or Barcelona-Clinic Liver Cancer staging (available for GSE76427 cohort), CD8 score has a statistically significant association with very early HCC (BCLC stage 0, P = 0.035) but not with AJCC staging in neither cohort (Figure 1B).
To evaluate an association between a potential loss of immune surveillance as measured by CD8 score with development of a malignant phenotype, we compared CD8 score along a spectrum of pre-malignant histologic change (chronic liver disease/cirrhosis, dysplasia and HCC). CD8 score is associated with pre-malignant cirrhotic livers (Figure 2) in both GSE6764 and GSE89377 cohorts.
Figure 2.
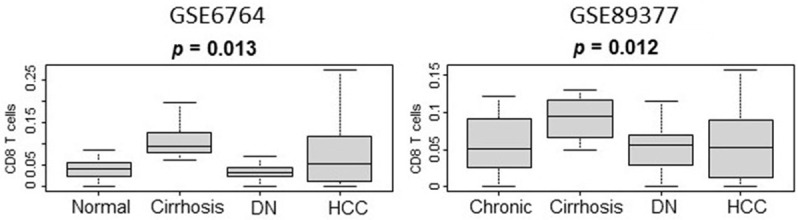
Correlation of CD8 T cell score with liver histology along a spectrum of pre-malignant change including chronic liver disease, cirrhosis, dysplasia (DN) and hepatocellular carcinoma (HCC) in patient cohorts GSE67674 and GSE89377.
CD8 score is associated with measurements of elevated immune activity
We next analyzed the extent to which CD8 score reflects a favorable anti-tumor inflamed tumor environment. We evaluated immune activity by multiple established methods of measuring immune response at the transcription level, to include cell populations, immune-related scores, and gene set enrichment analysis.
Analyzing immune cell subsets, high CD8 score is associated with CD4+ memory T cells (P < 0.001 in both TCGA and GSE76427 cohorts), dendritic cells (P < 0.001 TCGA, P = 0.002 GSE76427), B cells (P < 0.001 TCGA, P = 0.005 GSE76427), regulatory T cells (P < 0.001 TCGA, P = 0.019 GSE76427), Th2 helper T cells (P = 0.005 TCGA, P < 0.001 GSE76427), and M1 macrophages (P = 0.069 TCGA, P < 0.001, GSE76427). Figure 3A depicts cell subsets typically associated with a pro-cancer environment whereas Figure 3B depicts cell subsets typically representing an anti-cancer environment.
Figure 3.
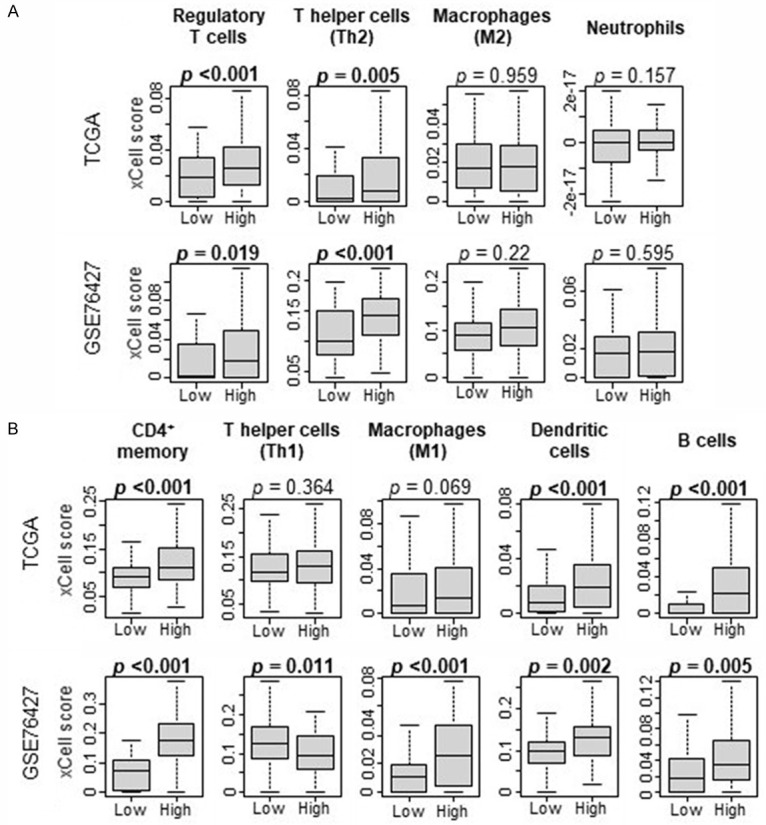
A. Correlation of CD8 xCell score with pro-tumorigenic immune cell populations. B. Correlation of CD8 xCell score with anti-tumorigenic immune cell populations.
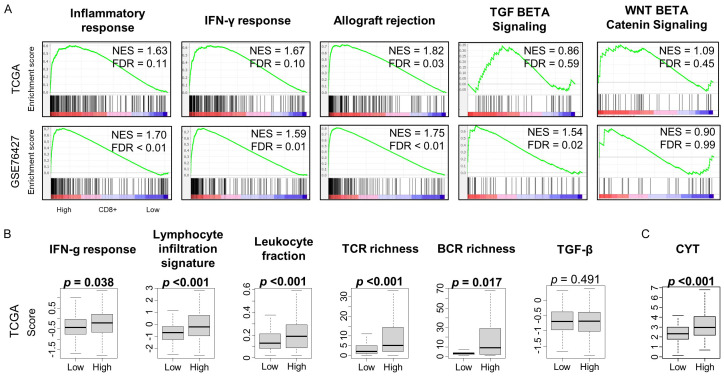
Gene set enrichment analysis (GSEA) of MSigDb Hallmark gene sets demonstrated high CD8 score significantly enriched for effector immunity gene sets depicting inflammatory response, IFN-gamma response, and allograft rejection gene sets in both cohorts (Figure 4A; normalized enrichment score (NES) = 1.66, NES = 1.58, and NES = 1.54; false discovery rate (FDR) = 0.03, FDR = 0.19, and FDR = 0.03 in TCGA, NES = 1.56, NES = 1.66, NES = 1.72, FDR = 0.03, FDR ≤ 0.01, FDR < 0.01 in GSE76427, respectively). The TGF-β [37,38] and Wnt/β-catenin [39,40] signaling pathways contribute to HCC carcinogenesis and also inhibit T cell immune activation [41-44]; we did not find a consistent association with CD8 score in both cohorts (Figure 4A).
Figure 4.
A. Gene Set Enrichment Analysis of MSigDb Hallmark gene sets involved in immune signalling and correlation with CD8 score; NES - Normalized Enrichment Score, FDR - False Discovery Rate. B. CD8 T Cell score correlation with tumor immune-related factors in the TCGA cohort. C. Cytolytic Activity Score (CYT) and correlation with CD8 T Cell score.
Next, we investigated the relationship between CD8 score and several tumor immune-related features in the TCGA cohort as previously quantified by Thorsson et al. [45]. HCC tumors with a high CD8 score had significantly increased levels of IFN-gamma response (P = 0.038), lymphocyte infiltration (P < 0.001), and leukocyte fraction (P < 0.001). Antigen-specific T-cell receptors (TCRs) and B-cell receptors (BCRs) serve as determinants of tumor antigen recognition. A high TCR and BCR repertoire polyclonality- or richness - may reflect a more robust anti-tumor immune cell response, with multiple different T cell and B cell clones recognizing tumor antigen and undergoing expansion. We found that CD8 score is associated with polyclonality of the T cell and B cell immune response, as measured by T cell receptor (TCR, P < 0.001) and B cell receptor (BCR, P = 0.017) richness (Figure 4B). Furthermore, CD8 score is positively associated with cytolytic activity score (CYT) - reflecting those genes involved in CD8+ T cell-mediated cytotoxicity/tumor cell killing (P < 0.001, Figure 4C). These data together suggest that the CD8 score reflects an anti-tumor immune microenvironment that potentially is the biological basis for the positive association with favorable oncologic outcomes.
Tumor mutation CD8 T cell score does not correlate to tumor mutation burden and mutation burden does not correlate with oncologic outcomes in HCC
Inflamed tumor environments such as those represented by high CD8 scores are commonly associated with high tumor mutational burden (TMB) [15]. Nonsilent (or non-synonymous) mutations in tumor coding regions can generate immunogenic neoantigens recognized by T cells as foreign peptides [16,17]. Insertion and deletion (Indel) mutations create novel open reading frames and as such generate more highly immunogenic peptides than single nucleotide variant (SNV) type mutations [46]. Measurements of DNA damage (intratumor heterogeneity, DNA chromosomal copy number alterations (CNA), and deficiencies in homologous recombination DNA damage repair proteins as measured by the HRD score) can represent predisposition to form immunogenic mutations [47-49]. We assessed correlation of CD8 score to measurements of genomic instability (Figure 5A), and measurements of mutational rate (total non-silent mutation rate, as well as neoantigens arising from either Indel or SNV mutations, Figure 5B) and did not find any association. High microsatellite mutation rate, or microsatellite instability, also often associates with an inflamed tumor environment and response to checkpoint blockade immunotherapy [50]. We also correlated CD8 score to MSI sensor score, which is a measure of mutation rate in DNA microsatellite regions, and again found no correlation (Figure 5B).
Figure 5.
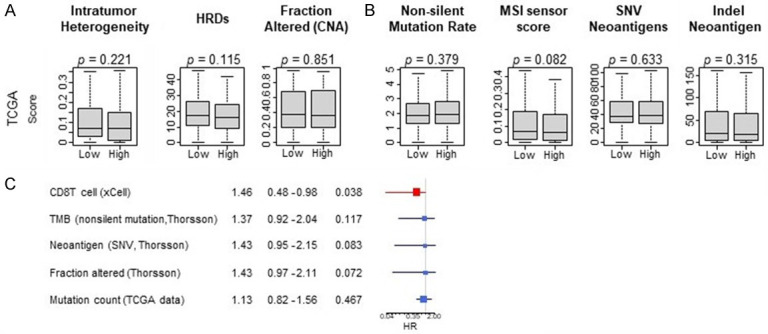
CD8 T cell score correlation to measures of tumor mutational burden. Bottom panel shows Cox proportional hazards for Overall Survival for individual variables.
Furthermore, tumor mutational burden, number of neoantigens, copy number alterations, and mutation count did not associate with OS (Figure 5C). This represents a novel framework within which to classify HCC, identifying an immune inflamed yet mutation poor patient subset with favorable oncologic outcomes.
CD8 score is associated with expression of multiple immune checkpoints
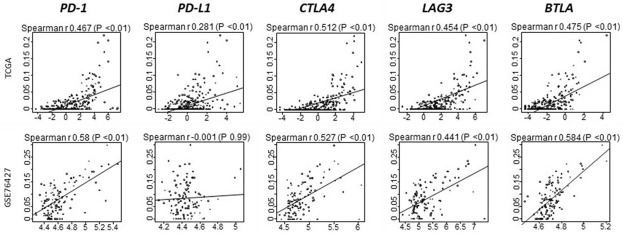
We hypothesized that an immune inflamed tumor environment as defined by CD8 score would trigger immune checkpoint receptor and ligand overexpression, a mechanism of adaptive immune resistance that serves to limit the anti-tumor immune response [4-6]. High CD8 score was associated with expression of well characterized checkpoint molecules (PD-1, r = 0.467, P < 0.01; PD-L1, r = 0.281, P < 0.01; CTLA-4, r = 0.512, P < 0.01; LAG3, r = 0.454, P < 0.01; BTLA, r = 0.475, P < 0.01; TIM3, r = 0.275, P < 0.01; Figure 6). Association with immune checkpoint expression reinforces the presence of an immune activated tumor microenvironment. Since PD-L1 predicts checkpoint blockade treatment response in other malignancies [10-12], high CD8 score may represent a testable biomarker to select HCC patients for immune checkpoint blockade.
Figure 6.
CD8 T Cell score correlation with concentration of immune checkpoint markers.
Discussion
Hepatocellular carcinoma is a primary liver malignancy with the second highest lethality of all cancers [1,2]. In its advanced form prognosis is dismal and even when localized, curative-intent liver-directed therapy is followed by recurrence more often than not. Recently, a novel combination of immune checkpoint inhibition with antiangiogenic therapy led to an unprecedented 30% response rate and a 5% complete response rate in patients with unresectable HCC [3]. Following T-cell immune attack, tumor cells develop ‘adaptive immunoresistance’, upregulating inhibitory pathways to limit further T cell effector function [4-6]. Antibodies targeting immune checkpoint pathways disrupt this T cell inhibition, ‘releasing the brake’ on the tumor-targeted T cell response. Checkpoint blockade (CPB) is the primary tool for immune activation against a wide range of solid tumors and has led to dramatic, durable responses in clinical scenarios previously considered uniformly fatal [7-9]. A particularly strong correlation exists between CPB response and the presence of neoantigens, which are uniquely capable of strong T cell activation due to their recognition as foreign antigen [16,17]. Yet for HCC, the tumor mutational burden is low. That nonetheless there is a significant minority of patients who respond to CPB speaks to a mechanism of T-cell mediated immune attack with specificity to antigens other than neoantigens and calls for a readily obtainable biomarker to identify those patients who would stand to benefit from CPB therapy so that we may better select patients for therapy.
In this manuscript, we characterize a cohort of patients by a high CD8 T cell RNA expression signature reflecting a primary component of the anti-tumor T cell mediated immune response - CD8+ T cells. That these patients associate with improved overall survival is perhaps due to a relatively favorable immune-mediated tumor cell attack. We demonstrate that these immune inflamed tumors can be characterized by anti-tumor immune cell subsets, gene set enrichment for effector T cell immunity and expression signatures consistent with activated T- and B-cell mediated immunity, and a corresponding upregulation of inhibitory checkpoint pathways potentially reflecting an adaptive tumor response to an active cancer-specific immunity. Other studies including at our institution have concluded a positive association of HCC survival with other measurements of anti-tumor immune response, including Th1 cytokine producing CD8+ T cells in peripheral blood detected by flow cytometry [51], and immune inflamed gene sets [20,52]. We extend these observations employing the CD8 T cell score to represent a primary immune cell subset responsible for immune attack and confirm an absence of correlation to multiple measures of tumor mutational burden and neoantigen loads.
That there may be an immune inflamed cohort of patients responsive to immune checkpoint blockade with low mutational burden is a conclusion made by Jaffee and colleagues, identifying such a cohort by immunohistochemistry-based detection of PD-L1 expression within tumor tissue [53]. While we do not have data on treatment response or protein level detection - which is the major limitation of our RNA level expression based analysis - we extend these findings and demonstrate an alternative identifying biomarker of a specifically HCC cohort of patients, a histology typically characterized by an overall low tumor mutational burden yet with a significant minority of patients attaining clinical response to CPB. A significant limitation of our study is that we do not have information on treatments given and treatment response, including any immunotherapy. A next step to be performed in the context of a clinical trial or institutional review would be to test the association of CD8 score with immunotherapy treatment response to assess the value of this marker either alone or in combination with other accepted markers of treatment response (ie. PD-L1 score, tumor mutational burden, and tumor infiltrating lymphocytes [54]).
Other studies identified a CD8 T cell exclusion expression signature correlating to dense fibrosis and HCC disease progression, a signature that may be reversed by blocking TGF-beta signaling [55,56]. While we similarly detected a decrease in CD8 T cell activity as defined by CD8 T cell score as disease state progressed along the cirrhotic/pre-neoplastic to cancer spectrum, we did not find an association with TGF-beta signaling gene set or Wnt/B-catenin pathway, which is another signaling pathway whose activation has been identified in HCC as critical for both carcinogenesis and T cell immune inhibition [39,42,44]. This may reflect as of yet unidentified oncogenic signaling pathways underlying a state of relative T cell dysfunction.
More studies are needed to explain the mechanisms by which HCC patients generate tumor-targeted immunity despite a neoantigen-poor substrate. Immunity to self-antigens - also designated tumor-associated antigens - are a potential target of CD8+ T cell immunity. Such T cell populations have been detected in a multitude of cancer patients across histologies, including for HCC [57]. Whether these T cells merely reflect a correlation and are ultimately in a state of tolerance given the multiple protective mechanisms limiting self-antigen specific T cell responses has not been studied. The wide breadth of research extensively characterizing the role of neoantigens as the target of high-affinity anti-tumor immune responses and as the target of CPB activated T cell activity serves as an example of how self-antigen specific responses can be studied in cohorts with low tumor mutational burden such as HCC.
Conclusion
In conclusion, a significant fraction of HCC tumor are ‘hot’ and can respond to immune therapies. We propose CD8 score to identify this immune inflamed population which can be characterized by improved overall survival, infiltration by anti-tumor immune cell subsets, gene set enrichment for effector T cell immunity, expression signatures consistent with activated T- and B-cell mediated immunity, and a corresponding upregulation of inhibitory checkpoint pathways potentially reflecting an adaptive tumor response to an active cancer-specific immunity. CD8 score may be a potentially useful biomarker to select patients for CPB therapy.
Disclosure of conflict of interest
None.
Supporting Information
References
- 1.Jemal A, Ward EM, Johnson CJ, Cronin KA, Ma J, Ryerson B, Mariotto A, Lake AJ, Wilson R, Sherman RL, Anderson RN, Henley SJ, Kohler BA, Penberthy L, Feuer EJ, Weir HK. Annual report to the nation on the status of cancer, 1975-2014, featuring survival. J Natl Cancer Inst. 2017;109:djx030. doi: 10.1093/jnci/djx030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Llovet JM, Zucman-Rossi J, Pikarsky E, Sangro B, Schwartz M, Sherman M, Gores G. Hepatocellular carcinoma. Nat Rev Dis Primers. 2016;2:16018. doi: 10.1038/nrdp.2016.18. [DOI] [PubMed] [Google Scholar]
- 3.Finn RS, Qin S, Ikeda M, Galle PR, Ducreux M, Kim TY, Kudo M, Breder V, Merle P, Kaseb AO, Li D, Verret W, Xu DZ, Hernandez S, Liu J, Huang C, Mulla S, Wang Y, Lim HY, Zhu AX, Cheng AL IMbrave150 Investigators. Atezolizumab plus bevacizumab in unresectable hepatocellular carcinoma. N Engl J Med. 2020;382:1894–1905. doi: 10.1056/NEJMoa1915745. [DOI] [PubMed] [Google Scholar]
- 4.Grosser R, Cherkassky L, Chintala N, Adusumilli PS. Combination immunotherapy with CAR T cells and checkpoint blockade for the treatment of solid tumors. Cancer Cell. 2019;36:471–482. doi: 10.1016/j.ccell.2019.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Spranger S, Spaapen RM, Zha Y, Williams J, Meng Y, Ha TT, Gajewski TF. Up-regulation of PD-L1, IDO, and T(regs) in the melanoma tumor microenvironment is driven by CD8(+) T cells. Sci Transl Med. 2013;5:200ra116. doi: 10.1126/scitranslmed.3006504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Tumeh PC, Harview CL, Yearley JH, Shintaku IP, Taylor EJ, Robert L, Chmielowski B, Spasic M, Henry G, Ciobanu V, West AN, Carmona M, Kivork C, Seja E, Cherry G, Gutierrez AJ, Grogan TR, Mateus C, Tomasic G, Glaspy JA, Emerson RO, Robins H, Pierce RH, Elashoff DA, Robert C, Ribas A. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature. 2014;515:568–571. doi: 10.1038/nature13954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Janjigian YY, Wolchok JD, Ariyan CE. Eradicating micrometastases with immune checkpoint blockade: strike while the iron is hot. Cancer Cell. 2021;39:738–742. doi: 10.1016/j.ccell.2021.05.013. [DOI] [PubMed] [Google Scholar]
- 8.Keenan TE, Burke KP, Van Allen EM. Genomic correlates of response to immune checkpoint blockade. Nat Med. 2019;25:389–402. doi: 10.1038/s41591-019-0382-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zappasodi R, Merghoub T, Wolchok JD. Emerging concepts for immune checkpoint blockade-based combination therapies. Cancer Cell. 2018;34:690. doi: 10.1016/j.ccell.2018.09.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Garon EB, Rizvi NA, Hui R, Leighl N, Balmanoukian AS, Eder JP, Patnaik A, Aggarwal C, Gubens M, Horn L, Carcereny E, Ahn MJ, Felip E, Lee JS, Hellmann MD, Hamid O, Goldman JW, Soria JC, Dolled-Filhart M, Rutledge RZ, Zhang J, Lunceford JK, Rangwala R, Lubiniecki GM, Roach C, Emancipator K, Gandhi L KEYNOTE-001 Investigators. Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med. 2015;372:2018–2028. doi: 10.1056/NEJMoa1501824. [DOI] [PubMed] [Google Scholar]
- 11.Robert C, Long GV, Brady B, Dutriaux C, Maio M, Mortier L, Hassel JC, Rutkowski P, McNeil C, Kalinka-Warzocha E, Savage KJ, Hernberg MM, Lebbe C, Charles J, Mihalcioiu C, Chiarion-Sileni V, Mauch C, Cognetti F, Arance A, Schmidt H, Schadendorf D, Gogas H, Lundgren-Eriksson L, Horak C, Sharkey B, Waxman IM, Atkinson V, Ascierto PA. Nivolumab in previously untreated melanoma without BRAF mutation. N Engl J Med. 2015;372:320–330. doi: 10.1056/NEJMoa1412082. [DOI] [PubMed] [Google Scholar]
- 12.Topalian SL, Hodi FS, Brahmer JR, Gettinger SN, Smith DC, McDermott DF, Powderly JD, Carvajal RD, Sosman JA, Atkins MB, Leming PD, Spigel DR, Antonia SJ, Horn L, Drake CG, Pardoll DM, Chen L, Sharfman WH, Anders RA, Taube JM, McMiller TL, Xu H, Korman AJ, Jure-Kunkel M, Agrawal S, McDonald D, Kollia GD, Gupta A, Wigginton JM, Sznol M. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med. 2012;366:2443–2454. doi: 10.1056/NEJMoa1200690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Danilova L, Wang H, Sunshine J, Kaunitz GJ, Cottrell TR, Xu H, Esandrio J, Anders RA, Cope L, Pardoll DM, Drake CG, Taube JM. Association of PD-1/PD-L axis expression with cytolytic activity, mutational load, and prognosis in melanoma and other solid tumors. Proc Natl Acad Sci U S A. 2016;113:E7769–E7777. doi: 10.1073/pnas.1607836113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gajewski TF, Louahed J, Brichard VG. Gene signature in melanoma associated with clinical activity: a potential clue to unlock cancer immunotherapy. Cancer J. 2010;16:399–403. doi: 10.1097/PPO.0b013e3181eacbd8. [DOI] [PubMed] [Google Scholar]
- 15.Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, Ibrahim F, Bruggeman C, Gasmi B, Zappasodi R, Maeda Y, Sander C, Garon EB, Merghoub T, Wolchok JD, Schumacher TN, Chan TA. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015;348:124–128. doi: 10.1126/science.aaa1348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Balachandran VP, Łuksza M, Zhao JN, Makarov V, Moral JA, Remark R, Herbst B, Askan G, Bhanot U, Senbabaoglu Y, Wells DK, Cary CIO, Grbovic-Huezo O, Attiyeh M, Medina B, Zhang J, Loo J, Saglimbeni J, Abu-Akeel M, Zappasodi R, Riaz N, Smoragiewicz M, Kelley ZL, Basturk O Australian Pancreatic Cancer Genome Initiative; Garvan Institute of Medical Research; Prince of Wales Hospital; Royal North Shore Hospital; University of Glasgow; St Vincent’s Hospital; QIMR Berghofer Medical Research Institute; University of Melbourne, Centre for Cancer Research; University of Queensland, Institute for Molecular Bioscience; Bankstown Hospital; Liverpool Hospital; Royal Prince Alfred Hospital, Chris O’Brien Lifehouse; Westmead Hospital; Fremantle Hospital; St John of God Healthcare; Royal Adelaide Hospital; Flinders Medical Centre; Envoi Pathology; Princess Alexandria Hospital; Austin Hospital; Johns Hopkins Medical Institutes; ARC-Net Centre for Applied Research on Cancer. Gönen M, Levine AJ, Allen PJ, Fearon DT, Merad M, Gnjatic S, Iacobuzio-Donahue CA, Wolchok JD, DeMatteo RP, Chan TA, Greenbaum BD, Merghoub T, Leach SD. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature. 2017;551:512–516. doi: 10.1038/nature24462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.McGranahan N, Furness AJ, Rosenthal R, Ramskov S, Lyngaa R, Saini SK, Jamal-Hanjani M, Wilson GA, Birkbak NJ, Hiley CT, Watkins TB, Shafi S, Murugaesu N, Mitter R, Akarca AU, Linares J, Marafioti T, Henry JY, Van Allen EM, Miao D, Schilling B, Schadendorf D, Garraway LA, Makarov V, Rizvi NA, Snyder A, Hellmann MD, Merghoub T, Wolchok JD, Shukla SA, Wu CJ, Peggs KS, Chan TA, Hadrup SR, Quezada SA, Swanton C. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science. 2016;351:1463–1469. doi: 10.1126/science.aaf1490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Narayanan S, Kawaguchi T, Yan L, Peng X, Qi Q, Takabe K. Cytolytic activity score to assess anticancer immunity in colorectal cancer. Ann Surg Oncol. 2018;25:2323–2331. doi: 10.1245/s10434-018-6506-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Oshi M, Asaoka M, Tokumaru Y, Yan L, Matsuyama R, Ishikawa T, Endo I, Takabe K. CD8 T cell score as a prognostic biomarker for triple negative breast cancer. Int J Mol Sci. 2020;21:6968. doi: 10.3390/ijms21186968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Takahashi H, Kawaguchi T, Yan L, Peng X, Qi Q, Morris LGT, Chan TA, Tsung A, Otsuji E, Takabe K. Immune cytolytic activity for comprehensive understanding of immune landscape in hepatocellular carcinoma. Cancers (Basel) 2020;12:1221. doi: 10.3390/cancers12051221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Liu J, Lichtenberg T, Hoadley KA, Poisson LM, Lazar AJ, Cherniack AD, Kovatich AJ, Benz CC, Levine DA, Lee AV, Omberg L, Wolf DM, Shriver CD, Thorsson V Cancer Genome Atlas Research Network. Hu H. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell. 2018;173:400–416. e411. doi: 10.1016/j.cell.2018.02.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Oshi M, Katsuta E, Yan L, Ebos JML, Rashid OM, Matsuyama R, Endo I, Takabe K. A novel 4-gene score to predict survival, distant metastasis and response to neoadjuvant therapy in breast cancer. Cancers (Basel) 2020;12:1148. doi: 10.3390/cancers12051148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Takahashi H, Oshi M, Asaoka M, Yan L, Endo I, Takabe K. Molecular biological features of nottingham histological grade 3 breast cancers. Ann Surg Oncol. 2020;27:4475–4485. doi: 10.1245/s10434-020-08608-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Takeshita T, Asaoka M, Katsuta E, Photiadis SJ, Narayanan S, Yan L, Takabe K. High expression of polo-like kinase 1 is associated with TP53 inactivation, DNA repair deficiency, and worse prognosis in ER positive Her2 negative breast cancer. Am J Transl Res. 2019;11:6507–6521. [PMC free article] [PubMed] [Google Scholar]
- 25.Tokumaru Y, Katsuta E, Oshi M, Sporn JC, Yan L, Le L, Matsuhashi N, Futamura M, Akao Y, Yoshida K, Takabe K. High expression of miR-34a associated with less aggressive cancer biology but not with survival in breast cancer. Int J Mol Sci. 2020;21:3045. doi: 10.3390/ijms21093045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wurmbach E, Chen YB, Khitrov G, Zhang W, Roayaie S, Schwartz M, Fiel I, Thung S, Mazzaferro V, Bruix J, Bottinger E, Friedman S, Waxman S, Llovet JM. Genome-wide molecular profiles of HCV-induced dysplasia and hepatocellular carcinoma. Hepatology. 2007;45:938–947. doi: 10.1002/hep.21622. [DOI] [PubMed] [Google Scholar]
- 27.Shen Q, Eun JW, Lee K, Kim HS, Yang HD, Kim SY, Lee EK, Kim T, Kang K, Kim S, Min DH, Oh SN, Lee YJ, Moon H, Ro SW, Park WS, Lee JY, Nam SW. Barrier to autointegration factor 1, procollagen-lysine, 2-oxoglutarate 5-dioxygenase 3, and splicing factor 3b subunit 4 as early-stage cancer decision markers and drivers of hepatocellular carcinoma. Hepatology. 2018;67:1360–1377. doi: 10.1002/hep.29606. [DOI] [PubMed] [Google Scholar]
- 28.Aran D, Hu Z, Butte AJ. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017;18:220. doi: 10.1186/s13059-017-1349-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Katsuta E, Rashid OM, Takabe K. Fibroblasts as a biological marker for curative resection in pancreatic ductal adenocarcinoma. Int J Mol Sci. 2020;21:3890. doi: 10.3390/ijms21113890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Oshi M, Takahashi H, Tokumaru Y, Yan L, Rashid OM, Matsuyama R, Endo I, Takabe K. G2M cell cycle pathway score as a prognostic biomarker of metastasis in estrogen receptor (ER)-positive breast cancer. Int J Mol Sci. 2020;21:2921. doi: 10.3390/ijms21082921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Oshi M, Takahashi H, Tokumaru Y, Yan L, Rashid OM, Nagahashi M, Matsuyama R, Endo I, Takabe K. The E2F pathway score as a predictive biomarker of response to neoadjuvant therapy in ER+/HER2- breast cancer. Cells. 2020;9:1643. doi: 10.3390/cells9071643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Tokumaru Y, Asaoka M, Oshi M, Katsuta E, Yan L, Narayanan S, Sugito N, Matsuhashi N, Futamura M, Akao Y, Yoshida K, Takabe K. High expression of microRNA-143 is associated with favorable tumor immune microenvironment and better survival in estrogen receptor positive breast cancer. Int J Mol Sci. 2020;21:3213. doi: 10.3390/ijms21093213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tokumaru Y, Oshi M, Katsuta E, Yan L, Huang JL, Nagahashi M, Matsuhashi N, Futamura M, Yoshida K, Takabe K. Intratumoral adipocyte-high breast cancer enrich for metastatic and inflammation-related pathways but associated with less cancer cell proliferation. Int J Mol Sci. 2020;21:5744. doi: 10.3390/ijms21165744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Tokumaru Y, Oshi M, Katsuta E, Yan L, Satyananda V, Matsuhashi N, Futamura M, Akao Y, Yoshida K, Takabe K. KRAS signaling enriched triple negative breast cancer is associated with favorable tumor immune microenvironment and better survival. Am J Cancer Res. 2020;10:897–907. [PMC free article] [PubMed] [Google Scholar]
- 35.Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liberzon A, Birger C, Thorvaldsdottir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1:417–425. doi: 10.1016/j.cels.2015.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chen J, Zaidi S, Rao S, Chen JS, Phan L, Farci P, Su X, Shetty K, White J, Zamboni F, Wu X, Rashid A, Pattabiraman N, Mazumder R, Horvath A, Wu RC, Li S, Xiao C, Deng CX, Wheeler DA, Mishra B, Akbani R, Mishra L. Analysis of genomes and transcriptomes of hepatocellular carcinomas identifies mutations and gene expression changes in the transforming growth factor-beta pathway. Gastroenterology. 2018;154:195–210. doi: 10.1053/j.gastro.2017.09.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wu K, Ding J, Chen C, Sun W, Ning BF, Wen W, Huang L, Han T, Yang W, Wang C, Li Z, Wu MC, Feng GS, Xie WF, Wang HY. Hepatic transforming growth factor beta gives rise to tumor-initiating cells and promotes liver cancer development. Hepatology. 2012;56:2255–2267. doi: 10.1002/hep.26007. [DOI] [PubMed] [Google Scholar]
- 39.de La Coste A, Romagnolo B, Billuart P, Renard CA, Buendia MA, Soubrane O, Fabre M, Chelly J, Beldjord C, Kahn A, Perret C. Somatic mutations of the beta-catenin gene are frequent in mouse and human hepatocellular carcinomas. Proc Natl Acad Sci U S A. 1998;95:8847–8851. doi: 10.1073/pnas.95.15.8847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Inagawa S, Itabashi M, Adachi S, Kawamoto T, Hori M, Shimazaki J, Yoshimi F, Fukao K. Expression and prognostic roles of beta-catenin in hepatocellular carcinoma: correlation with tumor progression and postoperative survival. Clin Cancer Res. 2002;8:450–456. [PubMed] [Google Scholar]
- 41.Chen CH, Seguin-Devaux C, Burke NA, Oriss TB, Watkins SC, Clipstone N, Ray A. Transforming growth factor beta blocks Tec kinase phosphorylation, Ca2+ influx, and NFATc translocation causing inhibition of T cell differentiation. J Exp Med. 2003;197:1689–1699. doi: 10.1084/jem.20021170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ding Y, Shen S, Lino AC, Curotto de Lafaille MA, Lafaille JJ. Beta-catenin stabilization extends regulatory T cell survival and induces anergy in nonregulatory T cells. Nat Med. 2008;14:162–169. doi: 10.1038/nm1707. [DOI] [PubMed] [Google Scholar]
- 43.Li MO, Sanjabi S, Flavell RA. Transforming growth factor-beta controls development, homeostasis, and tolerance of T cells by regulatory T cell-dependent and -independent mechanisms. Immunity. 2006;25:455–471. doi: 10.1016/j.immuni.2006.07.011. [DOI] [PubMed] [Google Scholar]
- 44.Spranger S, Bao R, Gajewski TF. Melanoma-intrinsic beta-catenin signalling prevents anti-tumour immunity. Nature. 2015;523:231–235. doi: 10.1038/nature14404. [DOI] [PubMed] [Google Scholar]
- 45.Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA, Ziv E, Culhane AC, Paull EO, Sivakumar IKA, Gentles AJ, Malhotra R, Farshidfar F, Colaprico A, Parker JS, Mose LE, Vo NS, Liu J, Liu Y, Rader J, Dhankani V, Reynolds SM, Bowlby R, Califano A, Cherniack AD, Anastassiou D, Bedognetti D, Mokrab Y, Newman AM, Rao A, Chen K, Krasnitz A, Hu H, Malta TM, Noushmehr H, Pedamallu CS, Bullman S, Ojesina AI, Lamb A, Zhou W, Shen H, Choueiri TK, Weinstein JN, Guinney J, Saltz J, Holt RA, Rabkin CS Cancer Genome Atlas Research Network. Lazar AJ, Serody JS, Demicco EG, Disis ML, Vincent BG, Shmulevich I. The immune landscape of cancer. Immunity. 2019;51:411–412. doi: 10.1016/j.immuni.2019.08.004. [DOI] [PubMed] [Google Scholar]
- 46.Turajlic S, Litchfield K, Xu H, Rosenthal R, McGranahan N, Reading JL, Wong YNS, Rowan A, Kanu N, Al Bakir M, Chambers T, Salgado R, Savas P, Loi S, Birkbak NJ, Sansregret L, Gore M, Larkin J, Quezada SA, Swanton C. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 2017;18:1009–1021. doi: 10.1016/S1470-2045(17)30516-8. [DOI] [PubMed] [Google Scholar]
- 47.Kim K, Kim HS, Kim JY, Jung H, Sun JM, Ahn JS, Ahn MJ, Park K, Lee SH, Choi JK. Predicting clinical benefit of immunotherapy by antigenic or functional mutations affecting tumour immunogenicity. Nat Commun. 2020;11:951. doi: 10.1038/s41467-020-14562-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Roh W, Chen PL, Reuben A, Spencer CN, Prieto PA, Miller JP, Gopalakrishnan V, Wang F, Cooper ZA, Reddy SM, Gumbs C, Little L, Chang Q, Chen WS, Wani K, De Macedo MP, Chen E, Austin-Breneman JL, Jiang H, Roszik J, Tetzlaff MT, Davies MA, Gershenwald JE, Tawbi H, Lazar AJ, Hwu P, Hwu WJ, Diab A, Glitza IC, Patel SP, Woodman SE, Amaria RN, Prieto VG, Hu J, Sharma P, Allison JP, Chin L, Zhang J, Wargo JA, Futreal PA. Integrated molecular analysis of tumor biopsies on sequential CTLA-4 and PD-1 blockade reveals markers of response and resistance. Sci Transl Med. 2017;9:eaah3560. doi: 10.1126/scitranslmed.aah3560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Samstein RM, Krishna C, Ma X, Pei X, Lee KW, Makarov V, Kuo F, Chung J, Srivastava RM, Purohit TA, Hoen DR, Mandal R, Setton J, Wu W, Shah R, Qeriqi B, Chang Q, Kendall S, Braunstein L, Weigelt B, Blecua Carrillo Albornoz P, Morris LGT, Mandelker DL, Reis-Filho JS, de Stanchina E, Powell SN, Chan TA, Riaz N. Mutations in BRCA1 and BRCA2 differentially affect the tumor microenvironment and response to checkpoint blockade immunotherapy. Nat Cancer. 2021;1:1188–1203. doi: 10.1038/s43018-020-00139-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK, Lu S, Kemberling H, Wilt C, Luber BS, Wong F, Azad NS, Rucki AA, Laheru D, Donehower R, Zaheer A, Fisher GA, Crocenzi TS, Lee JJ, Greten TF, Duffy AG, Ciombor KK, Eyring AD, Lam BH, Joe A, Kang SP, Holdhoff M, Danilova L, Cope L, Meyer C, Zhou S, Goldberg RM, Armstrong DK, Bever KM, Fader AN, Taube J, Housseau F, Spetzler D, Xiao N, Pardoll DM, Papadopoulos N, Kinzler KW, Eshleman JR, Vogelstein B, Anders RA, Diaz LA Jr. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science. 2017;357:409–413. doi: 10.1126/science.aan6733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kalathil SG, Hutson A, Barbi J, Iyer R, Thanavala Y. Augmentation of IFN-gamma+ CD8+ T cell responses correlates with survival of HCC patients on sorafenib therapy. JCI Insight. 2019;4:e130116. doi: 10.1172/jci.insight.130116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Sia D, Jiao Y, Martinez-Quetglas I, Kuchuk O, Villacorta-Martin C, Castro de Moura M, Putra J, Camprecios G, Bassaganyas L, Akers N, Losic B, Waxman S, Thung SN, Mazzaferro V, Esteller M, Friedman SL, Schwartz M, Villanueva A, Llovet JM. Identification of an immune-specific class of hepatocellular carcinoma, based on molecular features. Gastroenterology. 2017;153:812–826. doi: 10.1053/j.gastro.2017.06.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Yarchoan M, Albacker LA, Hopkins AC, Montesion M, Murugesan K, Vithayathil TT, Zaidi N, Azad NS, Laheru DA, Frampton GM, Jaffee EM. PD-L1 expression and tumor mutational burden are independent biomarkers in most cancers. JCI Insight. 2019;4:e126908. doi: 10.1172/jci.insight.126908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Meng X, Huang Z, Teng F, Xing L, Yu J. Predictive biomarkers in PD-1/PD-L1 checkpoint blockade immunotherapy. Cancer Treat Rev. 2015;41:868–876. doi: 10.1016/j.ctrv.2015.11.001. [DOI] [PubMed] [Google Scholar]
- 55.Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, Kadel EE III, Koeppen H, Astarita JL, Cubas R, Jhunjhunwala S, Banchereau R, Yang Y, Guan Y, Chalouni C, Ziai J, Senbabaoglu Y, Santoro S, Sheinson D, Hung J, Giltnane JM, Pierce AA, Mesh K, Lianoglou S, Riegler J, Carano RAD, Eriksson P, Hoglund M, Somarriba L, Halligan DL, van der Heijden MS, Loriot Y, Rosenberg JE, Fong L, Mellman I, Chen DS, Green M, Derleth C, Fine GD, Hegde PS, Bourgon R, Powles T. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature. 2018;554:544–548. doi: 10.1038/nature25501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Tauriello DVF, Palomo-Ponce S, Stork D, Berenguer-Llergo A, Badia-Ramentol J, Iglesias M, Sevillano M, Ibiza S, Canellas A, Hernando-Momblona X, Byrom D, Matarin JA, Calon A, Rivas EI, Nebreda AR, Riera A, Attolini CS, Batlle E. TGFbeta drives immune evasion in genetically reconstituted colon cancer metastasis. Nature. 2018;554:538–543. doi: 10.1038/nature25492. [DOI] [PubMed] [Google Scholar]
- 57.Schmidt N, Flecken T, Thimme R. Tumor-associated antigen specific CD8(+) T cells in hepatocellular carcinoma - a promising target for immunotherapy. Oncoimmunology. 2014;3:e954919. doi: 10.4161/21624011.2014.954919. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.