Abstract
The Ophiostomatales was erected in 1980. Since that time, several of the genera have been redefined and others have been described. There are currently 14 accepted genera in the Order. They include species that are the causal agents of plant and human diseases and common associates of insects such as bark beetles. Well known examples include the Dutch elm disease fungi and the causal agents of sporotrichosis in humans and animals. The taxonomy of the Ophiostomatales was confused for many years, mainly due to the convergent evolution of morphological characters used to delimit unrelated fungal taxa. The emergence of DNA-based methods has resolved much of this confusion. However, the delineation of some genera and the placement of various species and smaller lineages remains inconclusive. In this study we reconsidered the generic boundaries within the Ophiostomatales. A phylogenomic framework constructed from genome-wide sequence data for 31 species representing the major genera in the Order was used as a guide to delineate genera. This framework also informed our choice of the best markers from the currently most commonly used gene regions for taxonomic studies of these fungi. DNA was amplified and sequenced for more than 200 species, representing all lineages in the Order. We constructed phylogenetic trees based on the different gene regions and assembled a concatenated data set utilising a suite of phylogenetic analyses. The results supported and confirmed the delineation of nine of the 14 currently accepted genera, i.e. Aureovirgo, Ceratocystiopsis, Esteya, Fragosphaeria, Graphilbum, Hawksworthiomyces, Ophiostoma, Raffaelea and Sporothrix. The two most recently described genera, Chrysosphaeria and Intubia, were not included in the multi-locus analyses. This was due to their high sequence divergence, which was shown to result in ambiguous taxonomic placement, even though the results of phylogenomic analysis supported their inclusion in the Ophiostomatales. In addition to the currently accepted genera in the Ophiostomatales, well-supported lineages emerged that were distinct from those genera. These are described as novel genera. Two lineages included the type species of Grosmannia and Dryadomyces and these genera are thus reinstated and their circumscriptions redefined. The descriptions of all genera in the Ophiostomatales were standardised and refined where this was required and 39 new combinations have been provided for species in the newly emerging genera and one new combination has been provided for Sporothrix. The placement of Afroraffaelea could not be confirmed using the available data and the genus has been treated as incertae sedis in the Ophiostomatales. Paleoambrosia was not included in this study, due to the absence of living material available for this monotypic fossil genus. Overall, this study has provided the most comprehensive and robust phylogenies currently possible for the Ophiostomatales. It has also clarified several unresolved One Fungus-One Name nomenclatural issues relevant to the Order.
Taxonomic novelties: New genera: Harringtonia Z.W. de Beer & M. Procter, Heinzbutinia Z.W. de Beer & M. Procter, Jamesreidia Z.W. de Beer & M. Procter, Masuyamyces Z.W. de Beer & M. Procter. New species: Masuyamyces massonianae M. Procter & Z.W. de Beer. New combinations: Dryadomyces montetyi (M. Morelet) M. Procter & Z.W. de Beer, Dryadomyces quercivorus (Kubono & Shin. Ito) M. Procter & Z.W. de Beer, Dryadomyces quercus-mongolicae (K.H. Kim et al.) M. Procter & Z.W. de Beer, Dryadomyces sulphureus (L.R. Batra) M. Procter & Z.W. de Beer, Graphilbum pusillum (Masuya) M. Procter & Z.W. de Beer, Grosmannia abieticolens (K. Jacobs & M.J. Wingf.) M. Procter & Z.W. de Beer, Grosmannia altior (Paciura et al.) M. Procter & Z.W. de Beer, Grosmannia betulae (Jankowiak et al.) M. Procter & Z.W. de Beer, Grosmannia curviconidia (Paciura et al.) M. Procter & Z.W. de Beer, Grosmannia euphyes (K. Jacobs & M.J. Wingf.) M. Procter & Z.W. de Beer, Grosmannia fenglinhensis (R. Chang et al.) M. Procter & Z.W. de Beer, Grosmannia gestamen (de Errasti & Z.W. de Beer) M. Procter & Z.W. de Beer, Grosmannia innermongolica (X.W. Liu et al.) M. Procter & Z.W. de Beer, Grosmannia pistaciae (Paciura et al.) M. Procter & Z.W. de Beer, Grosmannia pruni (Masuya & M.J. Wingf.) M. Procter & Z.W. de Beer, Grosmannia taigensis (Linnak. et al.) M. Procter & Z.W. de Beer, Grosmannia trypodendri (Jankowiak et al.) M. Procter & Z.W. de Beer, Harringtonia aguacate (D.R. Simmons et al.) M. Procter & Z.W. de Beer, Harringtonia brunnea (L.R. Batra) M. Procter & Z.W. de Beer, Harringtonia lauricola (T.C. Harr. et al.) Z.W. de Beer & M. Procter, Heinzbutinia grandicarpa (Kowalski & Butin) Z.W. de Beer & M. Procter, Heinzbutinia microspora (Arx) M. Procter & Z.W. de Beer, Heinzbutinia solheimii (B. Strzałka & Jankowiak) Z.W. de Beer & M. Procter, Jamesreidia coronata (Olchow. & J. Reid) M. Procter & Z.W. de Beer, Jamesreidia nigricarpa (R.W. Davidson) M. Procter & Z.W. de Beer, Jamesreidia rostrocoronata (R.W. Davidson & Eslyn) M. Procter & Z.W. de Beer, Jamesreidia tenella (R.W. Davidson) Z.W. de Beer & M. Procter, Leptographium cainii (Olchow. & J. Reid) M. Procter & Z.W. de Beer, Leptographium europioides (E.F. Wright & Cain) M. Procter & Z.W. de Beer, Leptographium galeiforme (B.K. Bakshi) M. Procter & Z.W. de Beer, Leptographium pseudoeurophioides (Olchow. & J. Reid) M. Procter & Z.W. de Beer, Leptographium radiaticola (J.J. Kim et al.) M. Procter & Z.W. de Beer, Masuyamyces acarorum (R. Chang & Z.W. de Beer) M. Procter & Z.W. de Beer, Masuyamyces ambrosius (B.K. Bakshi) M. Procter & Z.W. de Beer, Masuyamyces botuliformis (Masuya) Z.W. de Beer & M. Procter, Masuyamyces jilinensis (R. Chang et al.) M. Procter & Z.W. de Beer, Masuyamyces lotiformis (Z. Wang & Q. Lu) M. Procter & Z.W. de Beer, Masuyamyces pallidulus (Linnak. et al.) M. Procter & Z.W. de Beer, Masuyamyces saponiodorus (Linnak. et al.) M. Procter & Z.W. de Beer, Sporothrix longicollis (Massee & E.S. Salmon) M. Procter & Z.W. de Beer.
Citation: de Beer W, Procter M, Wingfield MJ, Marincowitz S, Duong TA (2022). Generic boundaries in the Ophiostomatales reconsidered and revised. Studies in Mycology 101: 57–120. doi: 10.3114/sim.2022.101.02
Keywords: Generic boundaries, new taxa, nomenclature, Ophiostomataceae, Ophiostomatales, Sordariomycetidae, taxonomy
INTRODUCTION
The Ophiostomatales (Sordariomycetidae, Ascomycota), was described by Benny & Kimbrough (1980) accommodating the single family Ophiostomataceae. The Order includes species that are the causal agents of plant and human diseases as well as common associates of wood infesting insects such as bark beetles (Wingfield et al. 1993, Seifert et al. 2013). Well known examples are the Dutch elm disease fungi and the causal agents of sporotrichosis in humans and animals (Figs 1, 2). As described, this Order originally accommodated four genera, Ophiostoma, Ceratocystiopsis, Sphaeronaemella and Ceratocystis. These authors considered Ceratocystis distinct from Ophiostoma based on cell wall constituents and conidiogenesis (Weijman & De Hoog 1975, Seifert et al. 2013). Upadhyay (1981) treated Ophiostoma as a synonym of Ceratocystis based on morphological similarities such as their long-necked ascocarps that produce sheathed ascospores in sticky droplets to facilitate arthropod dispersal (Upadhyay 1981, Malloch & Blackwell 1993). In the 1990’s, DNA sequence data confirmed that Ophiostoma and Ceratocystis resided in two distinct Orders of the fungi (Hausner et al. 1993c, Spatafora & Blackwell 1994). Ceratocystis has subsequently been shown to represent several morphologically and ecologically distinct genera in the Ceratocystidaceae (Microascales) (De Beer et al. 2013a, 2014, Nel et al. 2018, Mayers et al. 2015, 2020).
Fig. 1.

Ecological niches in which genera and species residing in the Ophiostomatales are found. A. Ulmus americana street trees dying as a result of Dutch Elm Disease (photo: D.W. French). B. Symptoms of infection by the human pathogen Sporothix schenckii (photo: Prof. Dr Flávio de Queiroz Telles Filho, Federal University of Paraná, Brazil). C. Signage in Yellowstone National Park emphasising the important role that bark beetles (and by extension their fungal symbionts) play in the ecology of conifer ecosystems. D. Blue stain in conifer timber caused by numerous species of Ophiostomatoid fungi. E. Hylobius rhizophagus (root collar weevil) squashed onto the surface of agar medium containing cycloheximide selective for many genera and species of Ophiostomatales and in this case Leptographium procerum. F. Infructescences of a Protea species in which numerous species of Ophiostomatoid fungi can be found. G. Douglas fir (Pseudotsuga menzesii) trees dying as a result of black stain root disease caused by Leptographium wageneri var. pseudotsugae (Photo: F.W. Cobb). H. Pinus resinosa trees dying as a result of mass infestation by Ips pini and associated Ophiostoma minus.
Fig. 2.
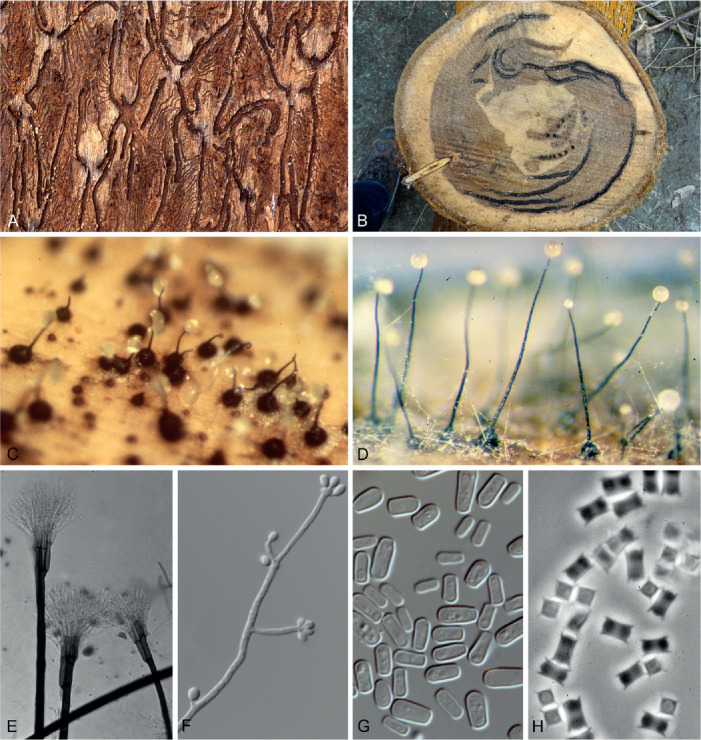
Ecological niches of Ophiostomatoid fungi and micrographs providing examples of structures typical of these fungi. A. Transverse stellate gallery systems of Ips schmutzenhoferi in the bark of a Pinus spinulosa tree, showing blue-stain around nuptial chambers and female galleries. B. Section through a Eucalyptus stem infested by the ambrosia beetle Megaplatypus mutatus showing tunnels in which species of Ophiostomatales occur. C, D. Ascomata of Ophiostoma ulmi (C) (photo: D.W. French) and O. pilliferum (D) (photo: Z.W. de Beer) with sticky ascospores masses at their apices, illustrating the manner by which these fungi easily attach to the insects that carry them. E. Conidiophores of Leptographium procerum, illustrating asexual structures well suited to being vectored by insects. F. Conidiogenous cells of a Sporothrix sp. G. Typical single-celled conidia found in most species of Ophiostomatales. H. Many species of Ophiostomatales have ascospores with sheaths such as these pillow-shaped spores in Ophiostoma ips.
The Ophiostomatales as it is currently defined accommodates a single family, the Ophiostomataceae (De Beer et al. 2013a), which was initially described in 1932 (Nannfeldt 1932). At the time, it included Ophiostoma, with Endoconidiophora and Ceratocystis as synonyms (Melin & Nannfeldt 1934). The family was treated in various Orders prior to 1980 (De Beer et al. 2013a). Apparently unaware of Benny & Kimbrough’s (1980) study, Upadhyay (1981) re-defined the Ophiostomataceae with Ceratocystis as type genus, and Ophiostoma, Sphaeronaemella, Grosmannia and Europhium as synonyms, and with Ceratocystiopsis as a distinct new genus. However, the DNA-based distinction between Ceratocystis and Ophiostoma led to their inevitable separation and emended definitions of the Ceratocystidaceae and Ophiostomataceae (Wingfield et al. 1993, Réblová et al. 2011, De Beer et al. 2013a, 2016a).
The first attempt to resolve generic boundaries within the Ophiostomatales subsequent to its separation from the Ceratocystidaceae was made by Zipfel et al. (2006). Based on phylogenies constructed from ribosomal DNA and β-tubulin sequences and including 55 taxa, they recognised Ophiostoma, Ceratocystiopsis and Grosmannia as distinct sexual genera. Following the dual nomenclature system at the time, asexual Sporothrix and Leptographium species retained their names in these genera, although they respectively grouped in Ophiostoma and Grosmannia.
After 20 years of DNA-based taxonomy and in the wake of the abandonment of a dual nomenclature for the fungi (Hawksworth et al. 2011, Hawksworth 2012), De Beer & Wingfield (2013) revised the Ophiostomatales, considering all published ribosomal large subunit (LSU) and internal transcribed spacer (ITS) sequences, including 266 species. They redefined Ophiostoma sensu stricto, Raffaelea s.s., Ceratocystiopsis, Fragosphaeria and Graphilbum, and recognised 18 species complexes within the various genera of the Ophiostomatales. However, they concluded that the rDNA-based phylogenies were not sufficiently robust to resolve the generic status of the Sporothrix schenckii-Ophiostoma stenoceras species complex, or that of various lineages within what they defined as Leptographium sensu lato. Their phylogenies also suggested that ambrosial species previously treated as Raffaelea did not form a monophyletic group.
After the revision of the Ophiostomatales by De Beer & Wingfield (2013), the majority of studies focused on new species descriptions (Romón et al. 2014a, b, Musvuugwa et al. 2015, 2016, De Errasti et al. 2016, Simmons et al. 2016, Chang et al. 2017, Marincowitz et al. 2017, 2020, etc.) and on resolving issues within species complexes (Ando et al. 2016, Linnakoski et al. 2016, Jankowiak et al. 2017, Yin et al. 2019, 2020, etc.).
De Beer et al. (2016a) reconsidered the status of the S. schenckii-O. stenoceras complex, providing sequences for four gene regions of 65 species with sporothrix-like asexual morphs. They concluded that Sporothrix represented a distinct genus including 51 species and incorporated the characters of the sexual morphs of many of the species, previously treated as Ophiostoma, in the emended definition of Sporothrix. A lineage including some of the remaining sporothrix-like species that did not form part of the newly defined genus were provided with the new genus name, Hawksworthiomyces, in a subsequent paper (De Beer et al. 2016b). In addition, Van der Linde et al. (2016) and Bateman et al. (2017) described Aureovirgo and Afroraffaelea respectively as novel, monotypic genera in the Order. In 2018, a fungus was discovered preserved in amber alongside an ambrosia beetle, leading to the description of Paleoambrosia. The genus was treated in the Ophiostomatales based on morphological characters resembling Raffaelea species (Poinar & Vega 2018). Most recently, Nel et al. (2021) described two new genera, Chrysosphaeria and Intubia, from the abandoned combs of fungus-growing termites (Termitomyces) in South Africa.
In this study, we reconsidered and redefined the unresolved boundaries of genera including Leptographium, Raffaelea and some smaller lineages in the Ophiostomatales. To achieve this goal, we selected four gene regions based on a phylogenomic framework constructed from genome-wide sequence data for representative ophiostomatalean species. Sequence data for these four gene regions were then generated for as many species in the Order as possible, and phylogenetic analyses were conducted.
MATERIALS AND METHODS
Fungal isolates and DNA extraction
Fungal cultures used in this study were obtained from the Culture Collection (CMW) of the Forestry and Agricultural Biotechnology Institute (FABI), University of Pretoria, South Africa, and the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, the Netherlands. Isolates were grown on 2 % malt extract agar (MEA: 20 g malt extract, 20 g agar, 1 L dH2O) at room temperature; initially with streptomycin (0.4 g/L, Sigma-Aldrich, Kempton Park, South Africa) and cycloheximide (0.5 g/L, Sigma-Aldrich, Kempton Park, South Africa) supplemented in the media, then sub-cultured to MEA, and maintained at 4 °C after optimal growth had occurred. DNA was extracted following the protocol described by Duong et al. (2012).
Phylogenomic analyses
Available genome sequences for 31 species representing 11 of 14 currently recognised genera (excluding Afroraffaelea, Aureovirgo and Paleoambrosia) in the Ophiostomatales (Supplementary Table S1) were used to construct a phylogenomic tree. Two species of Diaporthales (Cryphonectria parasitica and Diaporthe ampelina), two species of Magnaporthales (Magnaporthe grisea and Magnaporthe poae), and one species of Togniniales (Phaeoacremonium minimum) were included in the analyses as outgroup taxa. Genome assemblies for all species were subjected to BUSCO v. 4.0.5 (Seppey et al. 2019) runs using the Sordariomycetes_odb10 dataset to obtain BUSCO genes. The amino acid sequences were extracted from BUSCO results and datasets were compiled for each of the BUSCO orthologous groups. All BUSCO orthologous groups with duplicated BUSCO genes were excluded from the analysis. PRANK (Löytynoja 2014) was used to align the datasets with default parameters.
Trimal v. 1.4 (Capella-Gutiérrez et al. 2009) was used for trimming of the alignments with “-resoverlap 0.8 -seqoverlap 75” parameters. Only datasets with lengths equal or larger than 100 aa after trimming step were retained for further analysis. Permutation Tail Probability (PTP) tests were conducted in PAUP v. 4.0a (Swofford 2003) to identify and remove datasets having no phylogenetic signal as well as those with less than 50 parsimony-informative characters. Individual gene trees for each BUSCO orthologous group were constructed using IQ-TREE v. 2 with an optimal substitution model automatically determined and 1 000 ultrafast bootstraps (Hoang et al. 2018, Minh et al. 2020). TreeShrink v. 1.3.7 (Mai & Mirarab 2018) was used to remove outliers (taxa with abnormal branch length) from all trees with default parameters. Newick utilities (Junier & Zdobnov 2010) was used to collapse branches with less than 10 % bootstrap support. A species tree was then constructed from the final set of trees under the multi-species coalescent model using ASTRAL v. 5.7.7 (Mirarab et al. 2014). Branch length of the species tree was optimized with RAxML v. 8.2.11 (Stamatakis 2014) using only BUSCO orthogroups that have all 36 taxa present and without any outliner taxon as identified with TreeShrink analysis.
Selection of gene regions for phylogenetic analyses
Sequences for eight gene regions commonly used in phylogenetic studies of the fungi were extracted from 31 draft genome sequences for species in the Ophiostomatales, as well as from those used as outgroups in the phylogenomic analysis (Supplementary Table S1). These included β-tubulin (β-tub), translation elongation factor 1 alpha (TEF-1α), internal transcribed spacer region (ITS), ribosomal large subunit (LSU), mini chromosome maintenance protein complex 7 (MCM7), DNA-directed RNA polymerase II second largest subunit (RPBII), DNA-directed RNA polymerase II largest subunit (RPBI) and ribosomal small subunit (SSU). A phylogenetic tree was constructed for each of these datasets using IQ-TREE v. 2 as indicated above. Based on the level of congruency between individual gene trees and the phylogenomic tree as well as the phylogenetic signal of these gene regions, the LSU, ITS, TEF-1α and RPBII gene regions were selected as markers to delineate genera in the Ophiostomatales. These regions included the primary (ITS) and secondary (TEF-1α) barcodes for fungi (Schoch et al. 2012, Stielow et al. 2015).
Primer selection
Existing primers were used to amplify and sequence the ITS (ITS1F: Gardes & Bruns 1993, ITS4: White et al. 1990), TEF-1α (EF2F: Marincowitz et al. 2015, EF2R: Jacobs et al. 2004) and LSU (LR5, LROR: Vilgalys & Hester 1990) gene regions. Since previously available primers for RPBII did not consistently amplify the targeted region in most of the isolates investigated, we designed new primers for this gene region based on the available genome sequences: Oph-RPB2F1 (5’ - GAYGAYCGIGAYCAYTTYGG - 3’), Oph-RPB2F2 (5’ - TICTGGCIAARCTNTTCCG - 3’) and Oph-RPB2R1 (5’ - CCCATRGCYTGYTTRCCCAT - 3’). A combination of Oph-RPBF1 and Oph-RPBR1 was used in most instances, while Oph-RPBF2 and Oph-RPBR1 were used in cases where the former combination was not successful.
PCR
For the ITS, TEF-1α and LSU regions, FastStart Taq DNA Polymerase (Roche, Germany) was used. For the RPBII gene region the Platinum® Multiplex PCR Master Mix (Applied Biosystems, Foster City, California) was used. The ITS, LSU and TEF-1α gene regions were amplified following the protocol described by Duong et al. (2012). For the RPBII gene region, the protocol provided with the Platinum® Multiplex PCR Master Mix was used but amended as follows: the PCR mixture was made up to a final volume of 12.5 μL, primers were added to a concentration of 1 μM each, and PCR was carried out with 40 cycles of denaturing at 94 °C for 30 s, annealing at 58 °C for 30 s, and elongation at 72 °C for 1 min.
Agarose gel electrophoresis (1 % agarose) was performed on all PCR products to confirm the success of amplification. PCR products were treated with ExoSAP (a mixture of exonuclease I and alkaline phosphatase; one unit of each enzyme was used for approximately 20 μL of PCR product). The mixture was then subjected to two incubation steps at 37 °C for 15 min (for enzymatic action) and 80 °C for 15 min (to deactivate the enzymes). The treated products were stored at 4 °C until PCR sequencing was carried out.
DNA sequencing
Sanger sequencing was performed for all ExoSAP treated PCR products. The PCR sequencing setup reaction (12 μL) consisted of 6.4 μL dH2O, 2.1 μL 5× sequencing buffer, 0.5 μL BigDye v 3.1, 1 μL of the forward or reverse primer (10 mM), and 2 μL ExoSAP treated PCR product. The reaction was performed under the following conditions: 25 cycles of a denaturing step at 96 °C for 10 s, an annealing step at 55 °C for 5 s, and an elongation step at 60 °C for 4 min. The products were maintained at 4 °C until being used for precipitation.
PCR sequencing products were precipitated using the ethanol/NaOAc precipitation method. For each of the PCR sequencing products (total volume of 12 μL) 8 μL dH2O, 2 μL NaOAc (3 M, pH 5.2) and 50 μL absolute ethanol (EtOH) was added. The tubes were incubated on ice for 10 min, then centrifuged at 13 400 rpm for 30 min at room temperature. After centrifugation, the supernatant was removed, and the pellet washed twice with 150 μL of 70 % EtOH and centrifuged for 10 min at 13 400 rpm at room temperature. After the final wash, the supernatant was removed, and the pellet was air-dried for approximately 15 min. The samples were kept at -20 °C until they could be analysed. The fragment separations were performed using an ABI PRISM® 3100 Genetic Analyzer (Applied Biosystems). Consensus sequences were derived from sequences obtained with forward and reverse primers. All sequences generated in this study have been submitted to GenBank, and those from ex-type isolates will be included in the RefSeq Targeted Loci (RTL) database in GenBank (Schoch et al. 2014).
Phylogenetic analyses
For species with available genome sequences (Supplementary Table S1), gene region data were extracted from assembled genome sequences. Sequences were downloaded from GenBank for newly described species for which cultures were not available during the study period, as well as for species for which our sequence data were incomplete. Datasets were aligned using the online version of MAFFT v. 7 (Katoh & Stanley 2013) with default parameters. Alignments were refined with an online version of Gblocks v. 0.91b (Castresana 2000) using default parameters – i.e. no alternative options for more or less stringent selection were selected. Datasets for the various gene regions obtained from Gblocks were concatenated using FASconCAT-G (Kück & Meusemann 2010). Partitionfinder v. 2.1.1 (Lanfear et al. 2017) was used to determine best substitution models for the combined dataset.
Maximum Likelihood (ML) analyses using RAxML (Stamatakis 2014) were performed separately for all gene regions and for the concatenated dataset with raxmlGUI v. 1.3 (Silvestro & Michalak 2012) using the GTR+G+I substitution model and 1 000 thorough bootstrap replicates. Bayesian analysis was conducted using PhyloBayes-MPI v. 1.8 (Lartillot et al. 2013); two chains were run in parallel under the CAT-GTR model. The program bpcomp was used to assess the convergence in tree space. Runs were terminated when the maxdiff value obtained between two chains reached 0.1 or lower.
Morphology
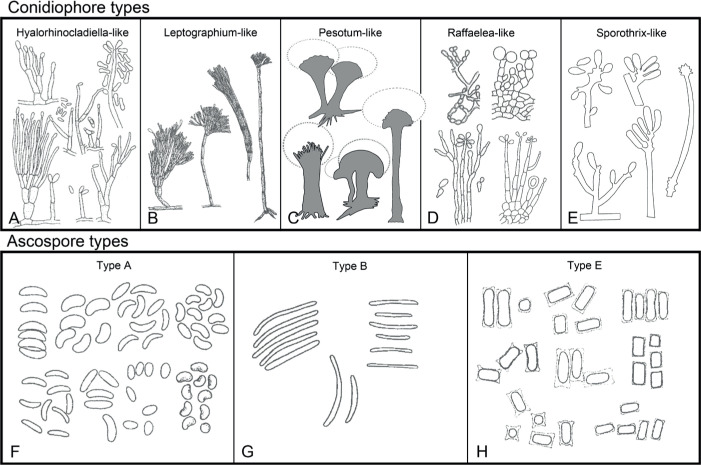
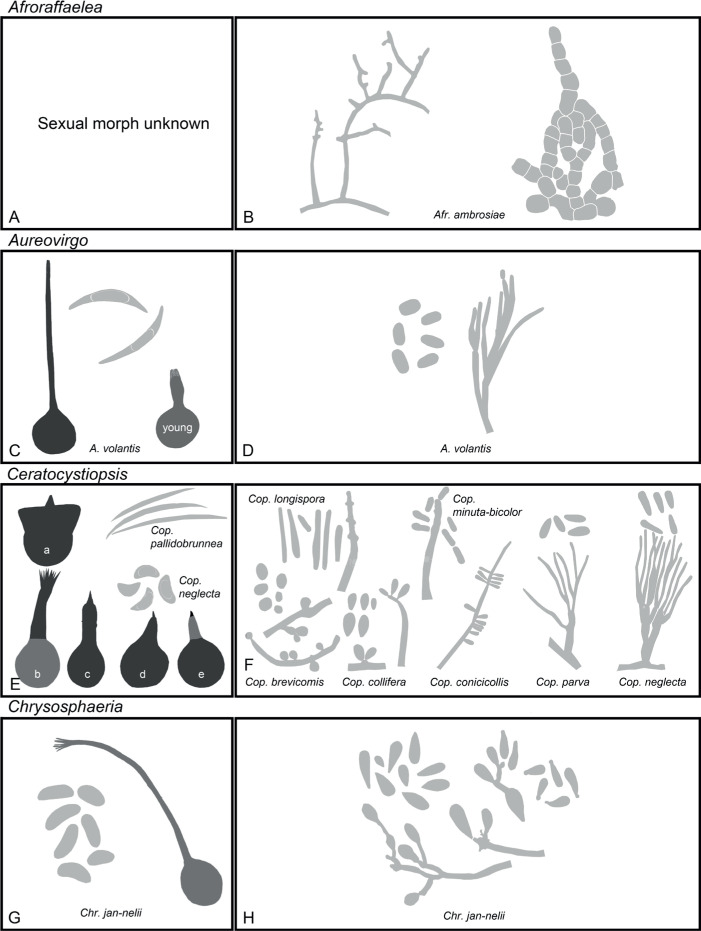
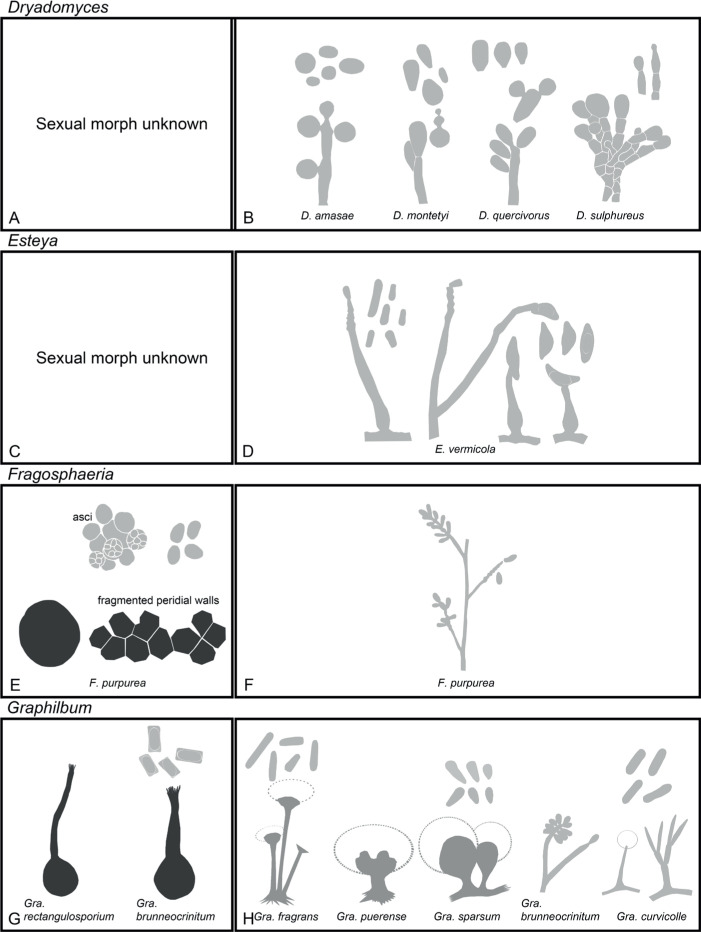
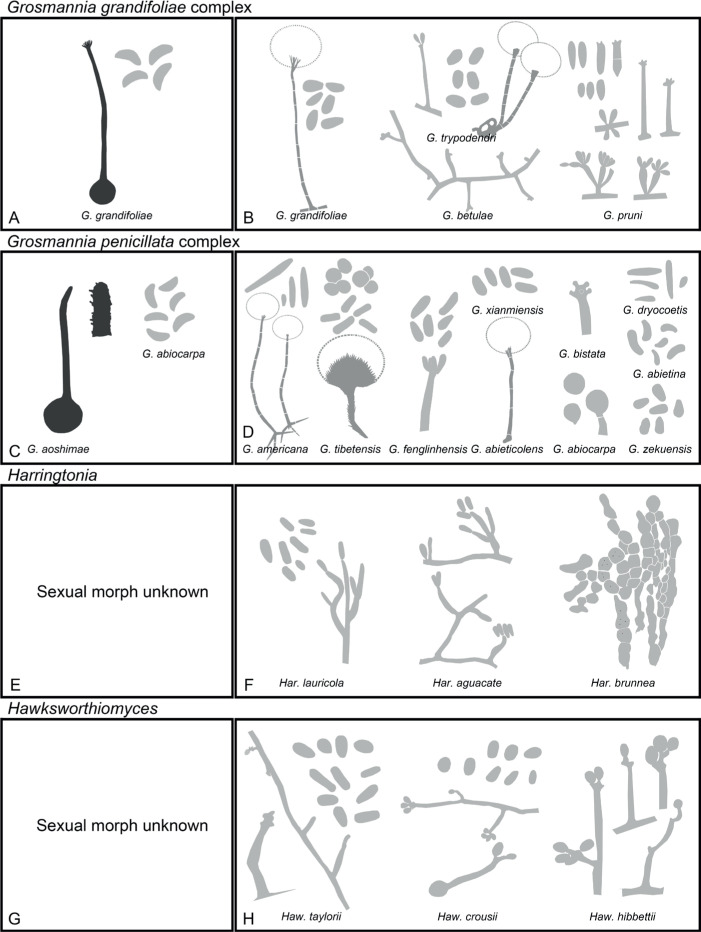
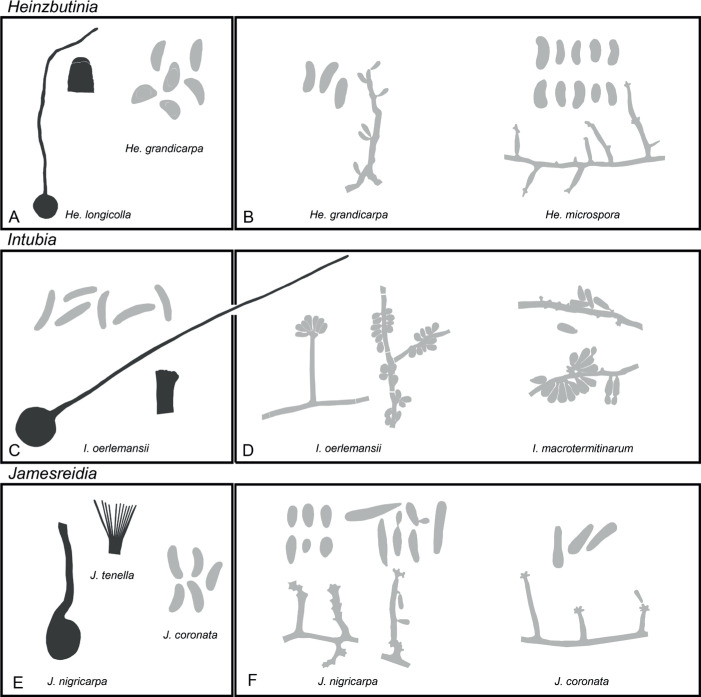
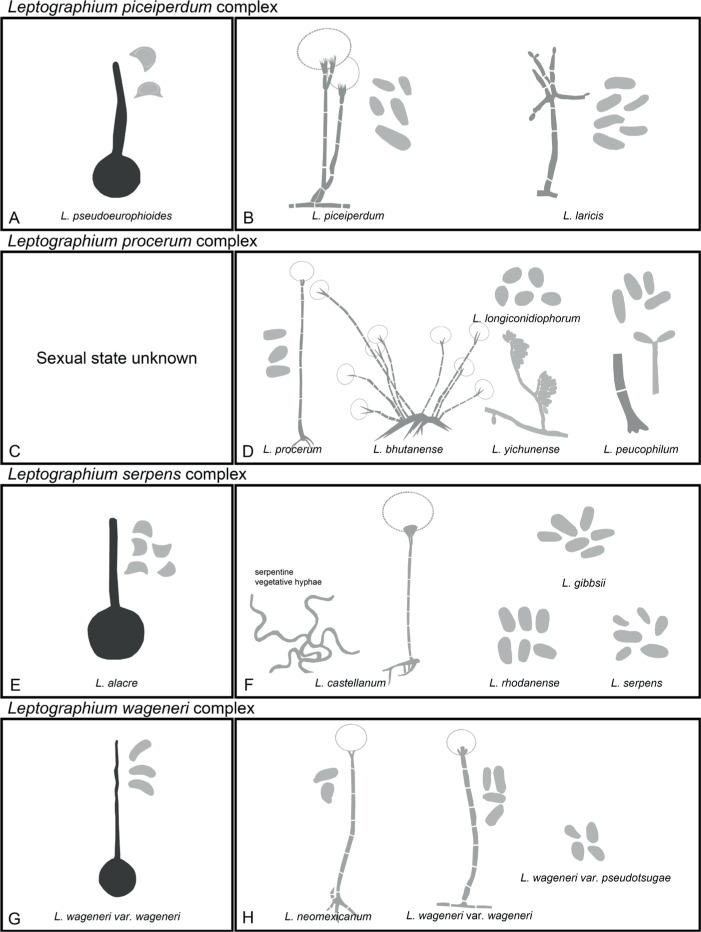
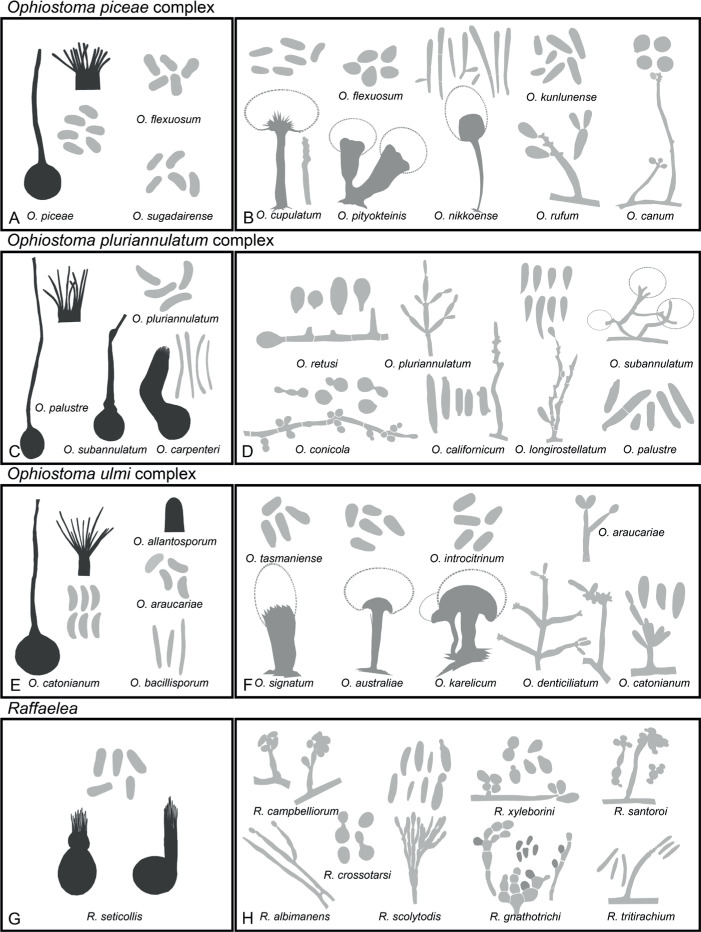
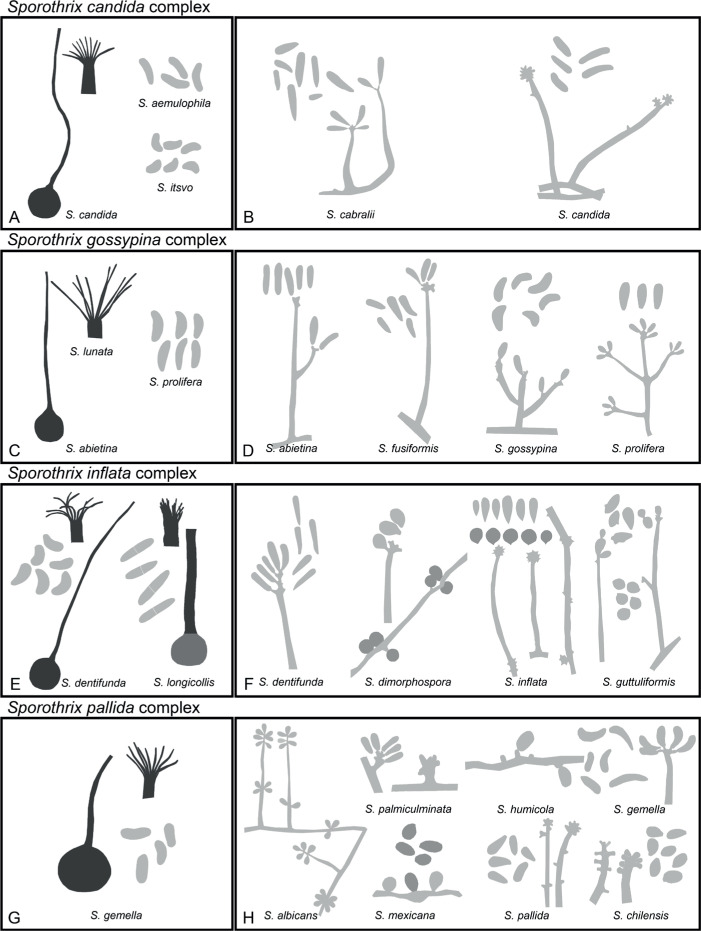
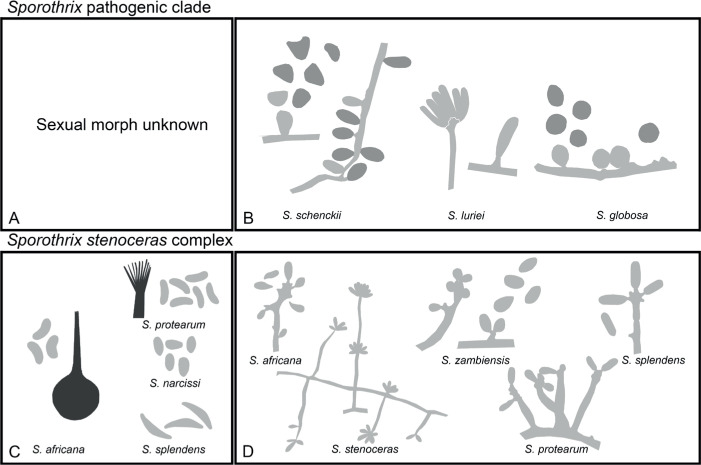
Of the 11 ascospore morphotypes defined by De Beer & Wingfield (2013) for the Ophiostomatales, three (Type A, B, E) were used to describe ascospore types in the present study (Fig. 3). Type A ascospores are those described as allantoid, bean-shaped, crescent to sickle-shaped, clavate to ovate, curved, cylindrical and slightly curved, orange section, lunate or reniform. Type B ascospores are those that are bacilliform, elongate filiform or narrow clavate. Type E ascospores are those with sheaths and are box-shaped, cylindrical, oblong, pillow-shaped, rectangular or rod-shaped. For the asexual morphs, five conidiophore types, hyalorhinocladiella-like, leptographium-like, pesotum-like, raffaelea-like and sporothrix-like, were used as descriptors where applicable (Fig. 3). Three shades of grey were applied in the figures to depict the colours of structures. Thus, hyaline to subhyaline structures were shaded in pale grey. Brown to dark brown structures were a medium-tone grey and fuscous black to black structures were presented in dark grey.
Fig. 3.
Conidiophore and ascospore types mentioned in the paper. A–E. Conidiophore types. F–H. Ascospore types sensu De Beer & Wingfield (2013). A, B, D, F–H. Adapted from illustrations in De Beer & Wingfield (2013). Shades of grey depict colours of various structures ranging from hyaline to dematiaceous.
RESULTS
The phylogenomic tree constructed from the genome-wide sequence data for 31 ophiostomatalean species (Fig. 4) showed a similar topology to those obtained in previous studies (Nel et al. 2021, Vanderpool et al. 2018). The placement of Graphilbum fragrans was, however, different from that suggested by Vanderpool et al. (2018). This inconsistent placement for Gra. fragrans was also observed by Nel et al. (2021) where different phylogenomic approaches were applied. Species of Leptographium s.l. grouped in two distinct lineages; one of these accommodated the Grosmannia penicillata complex and the other the L. lundbergii, L. procerum and L. galeiforme complexes. Species of Raffaelea s.l. resolved in three distinct clades, which is consistent with the findings in two previous studies (Nel et al. 2021, Vanderpool et al. 2018). Otherwise, all remaining species and genera included in the analysis resided in the clades consistent with their current recognition as separate genera in the Ophiostomatales.
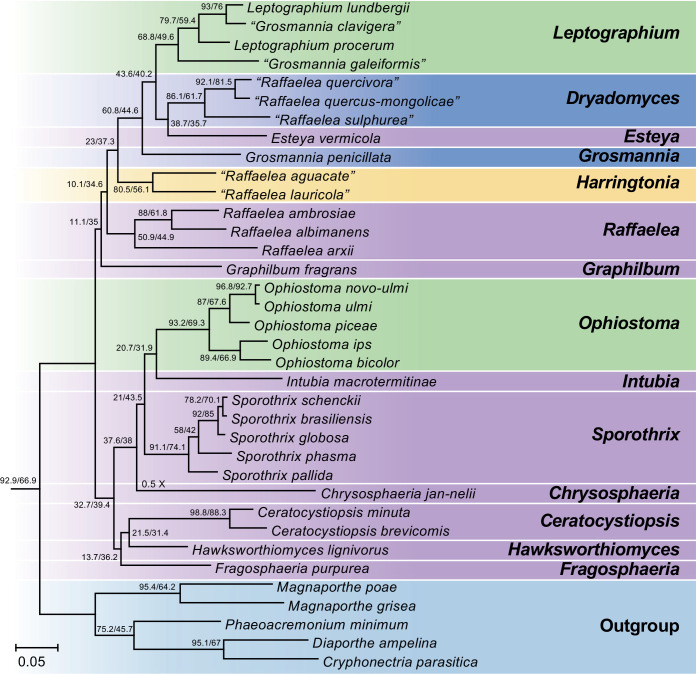
Fig. 4.
Phylogenomic tree obtained from supertree analysis with ASTRAL using gene trees constructed from 3 548 BUSCO genes (identified using the sordariomycetes_odb10 dataset; BUSCO v. 4.0.5). All 31 species in the Ophiostomatales for which genome sequence currently available were included in the analysis. Cryphonectria parasitica, Diaporthe ampelina, Magnaporthe grisea, Magnaporthe poae and Phaeoacremonium minimum were included as outgroup taxa. Gene concordance factors (gCF) and site concordance factors (sCF), which indicate the percentage of genes and sites that support a parcular nodes respectively, were determined using IQ-TREE2 are presented at nodes as gCF/sCF. Species names presented in double quotes denote old names which have been changed to their new respective genera subsequent to this study.
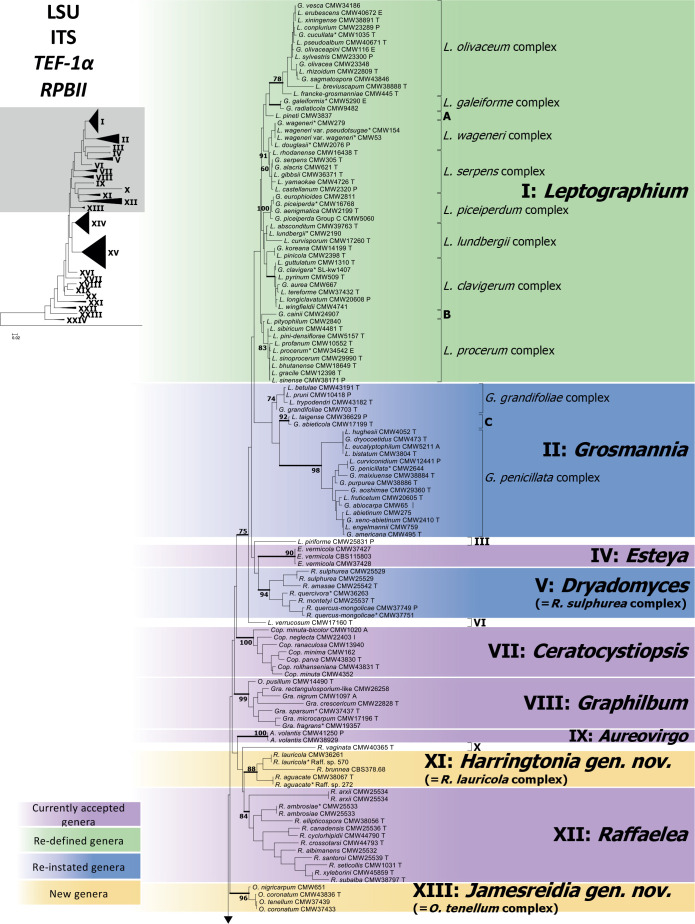
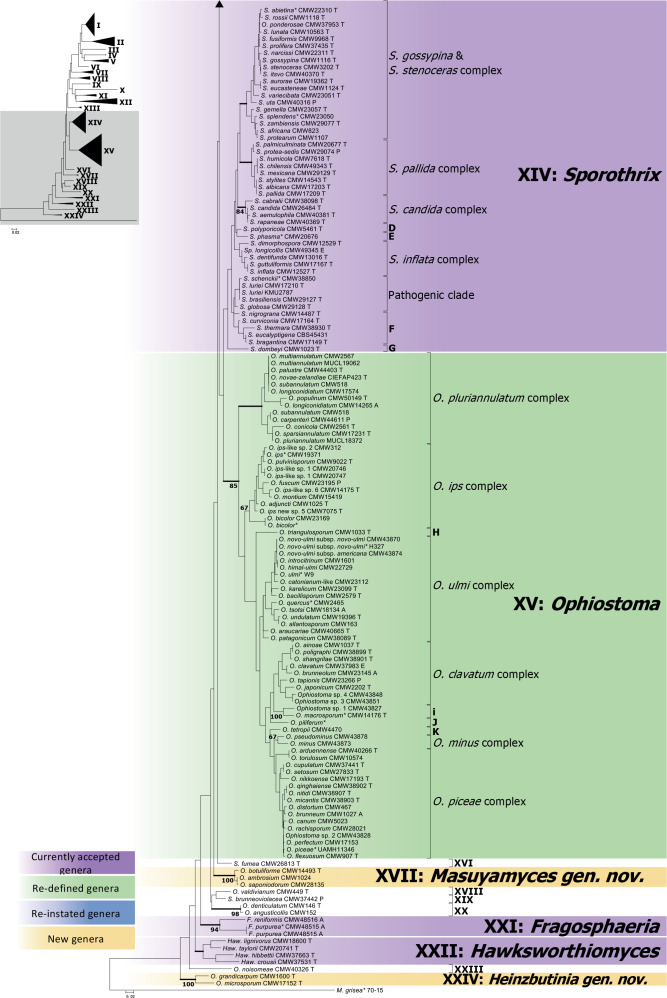
The maximum likelihood tree (Fig. 5) resulting from analyses of the concatenated dataset (LSU, ITS, TEF-1α and RPBII) for 264 isolates representing 249 species revealed 24 distinct lineages. Posterior probability values generated from Bayesian Inference analysis are indicated at the genus-level nodes (Fig. 5). Although the topologies of the individual gene trees (Figs S1–S4) were different to one another and to those in the combined tree, (apart from a few exceptions discussed below), the terminal clades were mostly consistent for the gene regions. To facilitate a discussion of the emerging results, the 24 lineages that represent genera or smaller groups were annotated using Roman numerals (I–XXIV). These were applied in the order of appearance in the concatenated tree (Fig. 5). Single species that grouped within major lineages, but not within defined species complexes, or not consistent within the same lineage in different trees, were labelled alphabetically (A–K).
Fig. 5.
Phylogenetic tree depicting the boundaries of currently accepted genera in the Ophiostomatales. This tree was generated using maximum likelihood analysis of the concatenated dataset of LSU, ITS, TEF1-α and RPBII gene regions. The dataset consisted of 264 isolates and 2 360 characters (including gaps). Bootstrap values above 60 % are shown. Bold lines indicate Bayesian posterior probabilities values above 0.8. Bootstrap values and Bayesian posterior probabilities values below the species complex level were removed for simplification. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks, genera that we redefine here, and blue blocks indicate genera that that have been reinstated and re-defined. (T = ex-type, E = ex-epitype, P = ex-paratype; L = ex-lectotype; A = authentic isolate, used in the original study; * Genome sequenced).
The overall topology of the LSU tree (Fig. S1) showed some differences from that of the concatenated tree, but 22 of the lineages corresponded between the two trees. The exceptions were Lineages III and XIX. Lineage III consisted of a single species in the concatenated tree, for which no LSU data were available, while Lineage XIX grouped outside the other genera in the concatenated tree, but as part of Sporothrix (Lineage XIV) in the LSU tree. Species complexes within Leptographium, Sporothrix and Ophiostoma were generally less well-defined in the LSU tree than in the concatenated tree.
The ITS tree (Fig. S2) showed little resolution below the genus level. Due to the variable nature of the ITS1 and ITS2 regions, the dataset was subjected to a strict Gblocks treatment (using automated parameters). The dataset consisted of 1 132 characters (including gaps) prior to treatment with Gblocks, and only 169 characters thereafter. The remaining dataset on which the tree (Fig. S2) is based, consisted predominantly of the 5.8S region. Nevertheless, we retained this analysis in the study because the ITS region is the officially recognised barcode for the fungi (Schoch et al. 2012). The ITS sequences were submitted to the GenBank Refseq database. The ITS tree (Fig. S2) supported separation of most of the genera, but Lineages XI, XII and XXII were not monophyletic in this tree, when compared to the concatenated tree. The relatively small dataset also failed to resolve most of the species complexes for the larger genera.
The TEF-1α tree (Fig. S3) resolved almost all the lineages representing species complexes, but failed to support monophyly of Lineages I, II, VIII, XI and XXII.
The RPBII tree (Fig. S4) supported the separation of all genera, species complexes and smaller lineages apart from Lineage XII that separated in four clades, and Lineage VIII that grouped within Lineage I.
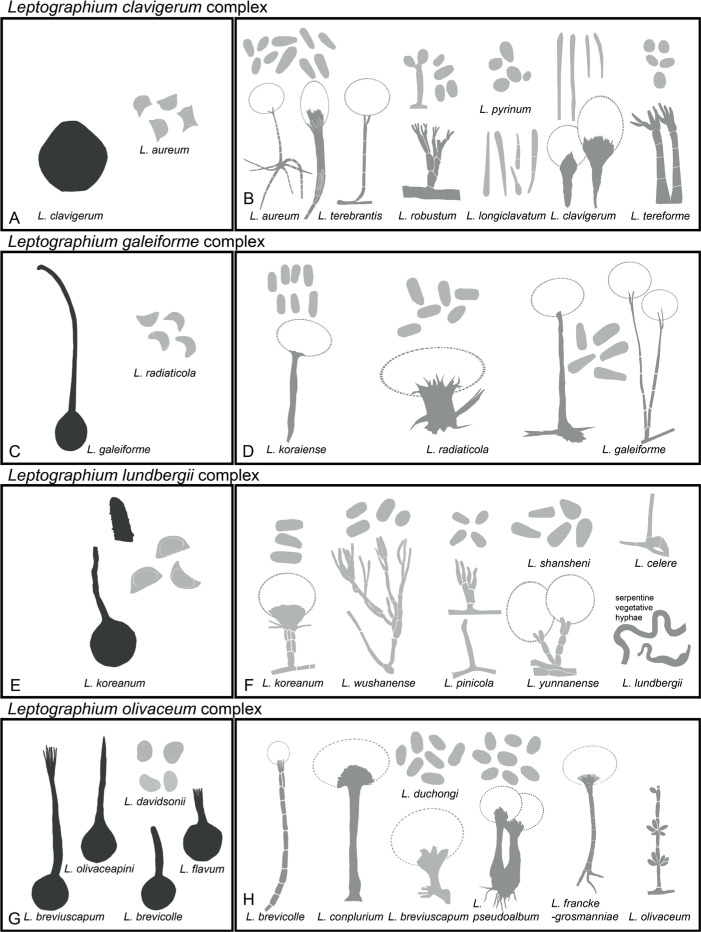
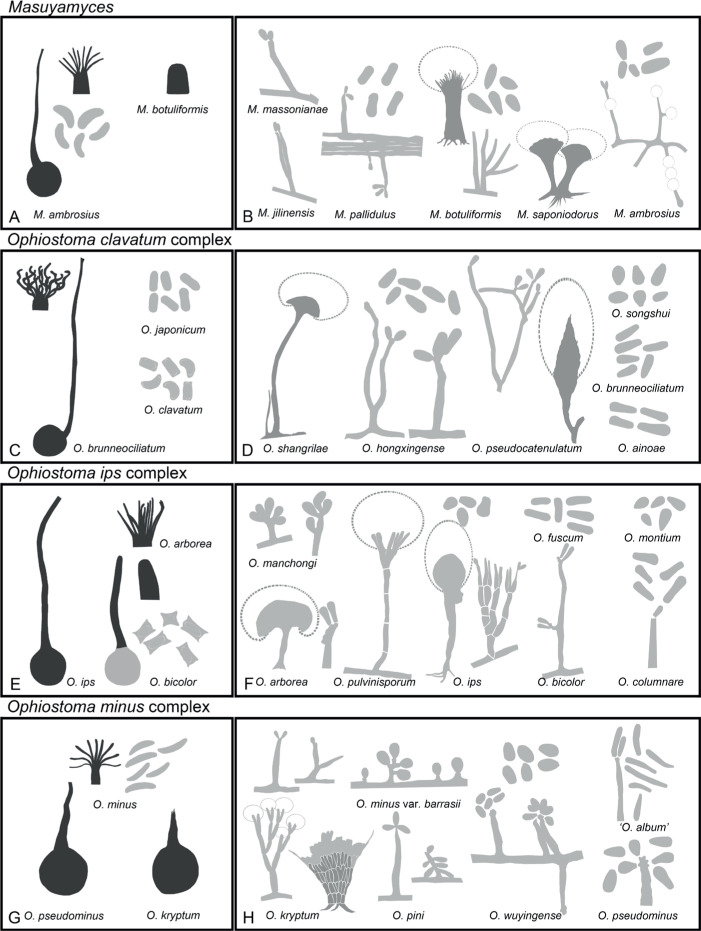
Based on the concatenated tree (Fig. 5), Lineage I included species complexes previously defined in Leptographium s.l. (De Beer & Wingfield 2013), namely the Leptographium clavigerum, L. galeiforme, L. lundbergii, L. olivaceum, L. piceiperdum, L. procerum, L. serpens and L. wageneri complexes; as well as two species not forming part of these species complexes (A & B). Lineage II included the G. grandifoliae and G. penicillata species complexes and a smaller lineage (C). Leptographium piriforme was labelled as Lineage III and L. verrucosum as Lineage VI, both grouping outside of Leptographium (Lineage I). Lineage IV consisted of three isolates of the monotypic genus Esteya. The Raffaelea sulphurea complex formed Lineage V. Ceratocystiopsis spp. formed Lineage VII, Graphilbum formed Lineage VIII and Aureovirgo formed Lineage IX. Raffaelea vaginata grouped outside of Raffaelea (Lineage XII) and was labelled Lineage X. Lineage XI consisted of the R. lauricola complex, distinct from Raffaelea spp., which formed Lineage XII. The Ophiostoma tenellum complex formed Lineage XIII. Sporothrix and Ophiostoma, as defined by De Beer et al. (2016a), formed Lineages XIV and XV, respectively. Lineage XIV consisted of the S. gossypina and S. stenoceras species complexes (which grouped inseparably from each other), the S. candida, S. inflata and S. pallida species complexes, the pathogenic clade (including the type species of Sporothrix, S. schenckii), as well as groups D to G. Lineage XV included the O. clavatum, O. ips, O. minus, O. piceae, O. pluriannulatum and O. ulmi complexes as well as groups H to K. Sporothrix fumea and S. brunneoviolacea both grouped separate from Sporothrix, and were labelled as Lineages XVI and XIX, respectively. Three Ophiostoma species consistently grouped together and distinct from Ophiostoma and were labelled Lineage XVII. Lineage XVIII included O. valdivianum and Lineage XX consisted of O. denticulatum and O. angusticollis. Lineage XXI represented Fragosphaeria, and Lineage XXII represented Hawksworthiomyces. Ophiostoma noisomeae was labelled Lineage XXIII, while O. grandicarpum and O. microsporum together constituted Lineage XXIV.
Afroraffaelea was excluded from the final analyses because the placement of the type for this monotypic species, Afr. ambrosiae, was completely incongruent among the separate gene trees (data not shown), most often forming long branches, distinct from all other groups. This impacted negatively on the support for several lineages in the concatenated tree, which prompted the decision to exclude the species from the analyses. Likewise, the two species of Intubia and the monotypic Chrysosphaeria were excluded from the analyses due to their ambiguous generic placement when using traditionally applied phylogenetic markers (Nel et al. 2021).
TAXONOMY
Phylogenetic analyses for four gene regions revealed 16 lineages within the Ophiostomatales, which we now recognise as valid genera. Seven of these represent genera currently known and defined in the Order. They include Esteya (Lineage IV), Ceratocystiopsis (Lineage VII), Graphilbum (Lineage VIII), Aureovirgo (Lineage IX), Sporothrix (Lineage XIV), Fragosphaeria (Lineage XXI) and Hawksworthiomyces (Lineage XXII). Two Lineages (Lineage II and V), which were clearly distinct from all other genera, included the ex-type cultures of Grosmannia penicillata and Dryadomyces amasae respectively. These species have previously been treated in the genera Grosmannia/Leptographium and Raffaelea respectively and we have consequently reinstated and redefined the genera Grosmannia and Dryadomyces with emended descriptions. Four lineages (Lineage XI, XIII, XVII and XXIV) are recognised as representing new genera and are described as such. Characters of these new genera were previously included in the descriptions of Leptographium, Ophiostoma and Raffaelea, and we have consequently emended their descriptions. Although Afroraffaelea, Chrysosphaeria, Intubia, and the fossil genus Paleoambrosia were not included in our analyses, we recognise these genera as valid, retaining them in the Ophiostomatales.
The placement of the remaining lineages (III, VI, X, XVI, XVIII, XIX, XX and XXIII) remains uncertain, as most of these lineages were represented by single species that grouped inconsistently in our analyses. We have chosen not to describe new monotypic genera for these lineages, but rather to delay this decision until additional taxa are discovered that support establishing novel genera. In addition to describing and redefining genera, new combinations have been provided for species where necessary.
Circumscription of the Ophiostomatales and Ophiostomataceae
At present there is no need to revise the description of the Ophiostomatales. This is because the emended description by De Beer et al. (2013a) broadly encompasses the morphologies of all genera, including the novel genera described in the present study. With more than 300 species and clearly distinct broad morphological groups in the Order, it would make sense to provide a narrower definition for the Ophiostomataceae, and to introduce one or more additional families. However, in view of the lack of support for the deeper nodes in our analyses, we have refrained from doing so at present. We suggest that this should be done only when related taxa outside the Ophiostomatales, such as those included in the LSU and SSU phylogenies of De Beer et al. (2013a), can be incorporated in multigene phylogenies to provide more robust context within the Sordariomycetidae.
Currently accepted genera in the Ophiostomatales
All currently accepted genera in the Ophiostomatales are defined below, and based on our results, descriptions have been emended where necessary. Genera and species complexes are discussed in alphabetical order, with lineage numbers corresponding to their appearance in the concatenated phylogenetic tree (Fig. 5).
New combinations
Where required, new combinations have been provided, and these are listed under the relevant genera. Species that have been treated in a particular genus but were shown based on our data to reside in a different genus, for which a name already exists in the appropriate genus, have been listed under ‘current name’ in Table 1, and thus not in the following section For example, Grosmannia serpens is now treated as Leptographium serpens but did not require a new combination and is listed in Table 1.
Table 1.
Taxa described in the Ophiostomatales, based on currently published data.
| Previous name | Current name | CMW1 | CBS or other1 | Type2 | Isolated from | Country | Collector |
GenBank Accession Numbers3
|
|||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ITS | LSU | TEF1-α | RPBII | ||||||||
| Afroraffaelea | |||||||||||
|
| |||||||||||
| Afr. ambrosiae | Afr. ambrosiae | 48331 | 141678 | T | Premnobius cavipennis | Florida, USA | C. Bateman | OM632703 | OM584293 | OM631576 | OM631577 |
|
| |||||||||||
| Aureovirgo | |||||||||||
|
| |||||||||||
| A. volantis | A. volantis | 41250 | 139649 | P | Cyrtogenius africus on Euphorbia ingens | South Africa | J.A. van der Linde | OM501369 | OM514700 | OM631743 | OM631579 |
| A. volantis | A. volantis | 38929 | 140081 | Cyrtogenius africus on Euphorbia ingens | South Africa | J.A. van der Linde | OM501368 | OM514699 | OM631742 | OM631578 | |
|
| |||||||||||
| Ceratocystiopsis | |||||||||||
|
| |||||||||||
| Cop. brevicomis | Cop. brevicomis | 40952 | 333.97 | T | Dendroctonus brevicomis | California, USA | T. Harrington | EU913722 | EU913683 | – | – |
| Cop. collifera | Cop. collifera | 7074 | 126.89 | T | Dendroctonus valens on Pinus teocote | Mexico | J. Marmolejo | EU913721 | EU913681 | – | – |
| Cop. concentrica | Cop. concentrica | – | WIN(M)71-07 | Pinus banksiana | Canada | J. Reid, A. Olchowecki | – | AF135571 | – | – | |
| Cop. conicicollis | Cop. conicicollis | – | WIN(M)69-25 | Abies balsemea | Canada | J. Reid, A. Olchowecki | – | – | – | – | |
| Cop. longispora | Cop. longispora | – | UM48 | Pinus sp. | Canada | A. Olchowecki | EU913723 | EU913684 | – | – | |
| Cop. lunata # | Cop. lunata | 55897 | 47171 | T | Xylosandrus crassciusculus | South Africa | W.J. Nel | MW028169 | MW028141 | – | – |
| Cop. manitobensis | Cop. manitobensis | 13792 | UAMH9813 | T | Manitoba beetle gallery in Pinus resinosa | Canada | J. Reid | EU913714 | EU913674 | – | – |
| Cop. minima | Cop. minima | 162 | 182.86 | Pinus banksiana | Wisconsin, USA | M.J. Wingfield | OM501370 | OM514701 | OM631744 | – | |
| Cop. minuta* | Cop. minuta | 4352 | 138717 | Ips cembrae | Poland | T. Kirisits | OM501372 | OM514703 | OM631745 | OM631581 | |
| Cop. minuta-bicolor * | Cop. minuta-bicolor | 1020 | 635.66 | A | Gallery of Ips sp. in Pinus contorta | USA | R.W. Davidson | OM501371 | OM514702 | – | OM631580 |
| Cop. neglecta | Cop. neglecta | 22403 | 100596 | I | Hylurgops palliatus | Germany | R. Kirschner | OM501373 | OM514704 | OM631746 | OM631582 |
| Cop. ochracea | Cop. ochracea | – | DAOM100148 | Picea mariana | Canada | H.D. Griffin | – | – | – | – | |
| Cop. pallidobrunnea | Cop. pallidobrunnea | – | UM51 | Populus tremuloides | Canada | J. Reid | – | EU913682 | – | – | |
| Cop. parva | Cop. parva | 43830 | UAMH9650 | T | Abies balsamea | Canada | A. Olchowecki | OM501374 | – | OM631747 | OM631583 |
| Cop. ranaculosa | Cop. ranaculosa | 13940 | 119683 | Pinus echinata | North Carolina, USA | F. Hains | OM501375 | OM514705 | OM631748 | OM631584 | |
| Cop. rollhanseniana | Cop. rollhanseniana | 43831 | UAMH9774 | T | Unknown beetle on Pinus sylvestris | Norway | J. Reid | OM501376 | OM514706 | OM631749 | – |
| Cop. spinulosa | Cop. spinulosa | DAOM110151 | T | Tilia americana | Canada | H.D. Griffin | – | – | – | – | |
| Cop. synnemata # | Cop. synnemata | – | NRIF 16918DA | T | Dryocoetes alni infesting Populus tremula | Poland | K. Miœkiewicz | MN900988 | MN900988 | MN901018 | – |
| Cop. yantaiensis # | Cop. yantaiensis | – | SNM650 | T | Pinus thunbergii | China | R. Chang | MW989411 | MZ819924 | MZ853080 | – |
| Cop. weihaiensis # | Cop. weihaiensis | – | SNM649 | T | Pinus thunbergii | China | R. Chang | MW989413 | MZ819926 | MZ853082 | – |
|
| |||||||||||
| Chrysosphaeria | |||||||||||
|
| |||||||||||
| Chr. jan-nelii # | Chr. jan-nelii | 47058 | 141570 | T | Termitomyces fungal comb of Macrotermes natalensis | South Africa | W.J. Nel | MT637038 | MT637006 | – | – |
|
| |||||||||||
| Dryadomyces (R. sulphurea complex) | |||||||||||
|
| |||||||||||
| R. amasae | D. amasae | 25542 | 116694 | T | Amasa concitatus on Angiosperms | Taiwan | H. Gebhardt | – | MT629750 | OM631750 | OM631585 |
| R. montetyi | D. montetyi | 25537 | 463.94 | T | Platypus cylindrus on Quercus suber | France | D. Vouland | – | MT629761 | OM631751 | – |
| R. quercivora * | D. quercivorus | 36263 | 122982 | Quercus mongolica | Japan | T. Kubono | MT633072 | MT629762 | OM631752 | OM631586 | |
| R. quercus-mongolicae | D. quercus-mongolicae | 37749 | KACC44403 | P | Quercus mongolica | South Korea | K.H. Kim | MT633073 | MT629764 | OM631754 | OM631588 |
| R. quercus-mongolicae * | D. quercus-mongolicae | 37751 | KACC44405 | Platypus koryoensis-infested Quercus | South Korea | K.H. Kim | MT633074 | MT629763 | OM631753 | OM631587 | |
| R. sulphurea* | D. sulphureus | 25529 | 380.68 | Xyleborus saxesenii gallery in Populus deltoides | Kansas, USA | L.R. Batra | MT633077 | MT629768 | OM631755 | OM631589 | |
|
| |||||||||||
| Esteya | |||||||||||
|
| |||||||||||
| E. vermicola | E. vermicola | 37427 | 100821 | Olea europeae | Italy | S. Frisullo | – | OM514708 | OM631757 | OM631591 | |
| E. vermicola | E. vermicola | 37428 | 156.82 | Pinus sp. | Taiwan | T. Tatsuno | OM501378 | OM514709 | OM631758 | – | |
| E. vermicola* | E. vermicola | – | 115803 | S. intricatus and its galleries in oak trees | Czech Republic | L. Marvanova | OM501377 | OM514707 | OM631756 | OM631590 | |
|
| |||||||||||
| Fragosphaeria | |||||||||||
|
| |||||||||||
| F. purpurea* | F. purpurea | 48515 | 133.34 | A | Fagus sp. | England, UK | C.G.C. Chesters | OM501379 | OM514710 | OM631759 | OM631592 |
| F. reniformis | F. reniformis | 48516 | 134.34 | A | Fagus sp. | England, UK | E.W. Mason | OM501381 | – | OM631760 | OM631593 |
|
| |||||||||||
| Graphilbum | |||||||||||
|
| |||||||||||
| Gra. acuminatum # | Gra. acuminatum | 54769 | 145828 | T | Ips acuminatus gallery on Pinus sylvestris | Poland | R. Jankowiak | MN548902 | – | MN548952 | – |
| Gra. brunneocrinitum | Gra. brunneocrinitum | – | TRTC34581 | T | Abies balsamea | Canada | – | – | – | – | – |
| Gra. carpaticum # | Gra. carpaticum | 43141 | 145835 | T | Pissodes piceae gallery on Abies alba | Poland | P. Majka | KY568116 | – | MN548956 | – |
| Gra. curvidentis # | Gra. curvidentis | 54779 | 145832 | T | Pitokteines curvidens gallery on Abies alba | Poland | P. Bilański | KY568111 | – | KY56850 | – |
| Gra. crescericum | Gra. crescericum | 22828 | 130864 | T | Hylurgops palliatus on Pinus radiata | Spain | P. Romón | OM501403 | OM514749 | OM631779 | OM631604 |
| Gra. curvicolle | Gra. curvicolle | – | WIN(M)70-25 | T | Abies balsamea | Canada | J. Reid, A. Olchowecki | – | – | – | – |
| Gra. fragrans * | Gra. fragrans | 19357 | 138720 | Pinus patula | South Africa | X. D. Zhou | OM501404 | OM514750 | OM631780 | OM631605 | |
| Gra. furuicola # | Gra. furuicola | 44770 | 145813 | T | Tomicus piniperda in Pinus sylvestris | Norway | R.H. Lindseth, T.H. Sundt | MN548907 | – | MN548961 | – |
| Gra. gorcense # | Gra. gorcense | 34153 | 146203 | T | Tetropium sp. in Picea abies | Poland | R. Jankowiak | MN548919 | – | MN548972 | – |
| Gra. interstitiale # | Gra. interstitiale | 54780 | 145816 | T | Hylurgops interstitialis in Pinus sylvestris | Russia | H. Solheim | MN548909 | – | MN548963 | – |
| Gra. ipis-grandicollis # | Gra. ipis-grandicollis | – | VPRI43762 | Ips grandicollis gallery on Pinus radiata | Australia | A.J. Carnegie | MW046071 | MW046117 | MW066405 | – | |
| Gra. kesiyae | Gra. kesiyae | 41729 | 139652 | T | Polygraphus szemaoensis on Pinus kesiya | China | S. Taerum | MG205669 | – | – | – |
| Gra. microcarpum | Gra. microcarpum | 17196 | YCC439 | T | Cryphalus montanus | Japan | Y. Yamaoka | OM501405 | OM514751 | OM631781 | OM631606 |
| Gra. nigrum | Gra. nigrum | 1097 | 163.61 | A | Abies lasiocarpa | Colorado, USA | R.W. Davidson | OM501406 | OM514752 | – | OM631607 |
| Gra. niveum # | Gra. niveum | – | SNM145 | T | Pinus thunbergii | China | R. Chang | MW989418 | – | MZ019548 | – |
| Gra. puerense | Gra. puerense | 41673 | 139640 | T | Ips acuminatus on Pinus kesiya | China | S. Taerum | MG205671 | – | – | – |
| Gra. rectangulosporium | Gra. rectangulosporium | 29364 | MAFF 238952 | T | Polygraphus proximus on Abies mariesii | Japan | N. Ohtaka | MG205671 | – | – | – |
| Gra. roseum # | Gra. roseum | 40349 | 141074 | T | Curtisia dentata | South Africa | T. Musvuugwa | KY050751 | – | – | – |
| Gra. sexdentatum # | Gra. sexdentatum | 54773 | 145814 | T | Ips sexdentatus in Pinus sylvestris | Norway | H. Solheim, M.E. Waalberg | MN548915 | – | MN548968 | – |
| Gra. sparsum * | Gra. sparsum | 37437 | 405.77 | T | Bark beetle gallery on Picea glauca | Alaska, USA | R.W. Davidson | OM501409 | OM514755 | – | OM631608 |
| Gra. translucens # | Gra. translucens | – | SNM144 | T | Pinus thunbergii | China | R. Chang | MW989416 | – | MZ019546 | – |
| Gra. tsugae | Gra. tsugae | – | UAMH11701 | T | Tsuga heterophylla | Canada | J. Reid, B. Reid | KJ661745 | – | – | – |
| Gra. tubicolle | Gra. tubicolle | 43837 | UAMH9686 | T | Pinus banksiana | Canada | A. Olchowecki | – | – | – | – |
| O. pusillum | Gra. pusilllum | 14490 | – | T | Pinus densiflora | Japan | H. Masuya | OM501407 | OM514753 | – | – |
|
| |||||||||||
| Grosmannia: G. penicillata complex | |||||||||||
|
| |||||||||||
| G. abiocarpa | G. abiocarpa | 65 | 594.85 | L | Ips sp. on Picea engelmannii | Colorado, USA | R.W. Davidson | OM501384 | OM514715 | OM631764 | – |
| G. americana | G. americana | 495 | 497.96 | T | Dendroctonus simplex on Larix laricina | Vermont, USA | D. Bergdalh | OM501385 | OM514718 | OM631765 | OM631595 |
| G. aoshimae | G. aoshimae | 29360 | MAFF238948 | T | Polygraphus proximus on Abies mariesii | Japan | N. Ohtaka | OM501386 | OM514719 | OM631766 | OM631596 |
| G. crassifolia # | G. crassifolia | 38885 | 136505 | T | Polygraphus poligraphus in Pinus crassifolia | China | X.D. Zhou, S. Taerum | – | MN644475 | MN647897 | – |
| G. dryocoetis | G. dryocoetis | 473 | 376.66 | T | Dryocoetes confusus on Abies lasiocarpa | Canada | A.C. Molnar | OM501392 | OM514727 | OM631768 | OM631597 |
| L. fenglinhense | G. fenglinhensis | 44579 | 141896 | T | Ips typographus on Pinus sp. | China | R. Chang, S.F. Chen | MH144128 | – | MH124404 | – |
| G. maixiuense | G. maixiuense | 38884 | 136502 | T | Polygraphus poligraphus, Ips shangrila in Picea crassifolia | China | M. Yin, S. Taerum, X.D. Zhou | OM501396 | MN644474 | MN647900 | OM631599 |
| G. penicillata * | G. penicillata | 2644 | 116008 | Picea abies | Norway | H. Solheim | OM501397 | OM514737 | OM631774 | OM631600 | |
| G. purpurea | G. purpurea | 38886 | 136975 | T | Ips shangrila on Picea purpurea | China | M. Yin, S. Taerum, X.D. Zhou | OM501399 | MN644476 | MN647914 | – |
| G. tibetensis | G. tibetensis | – | CFCC53415 | T | Orthotomicus sp. on Pinus likiangensis var. balfouriana | Tibet | Z. Wang, Q. Lu | MT269759 | – | MT268756 | – |
| G. xeno-abietinum | G. xeno-abietinum | 2410 | 136514 | T | Pinus ponderosa | California, USA | T. Harrington | OM501402 | MN644471 | MN647894 | OM631603 |
| G. xianmiense # | G. xianmiense | 38892 | 136500 | T | Polygraphus poligraphus in Pinus crassifolia | China | X.D. Zhou, S. Taerum | – | MN644479 | MN647911 | – |
| G. zekuensis # | G. zekuensis | 41876 | 141901 | T | Bakerdania sp. in gallery of Ips nitidus on Picea crassifolia | China | S.J. Taerum | MH121683 | MH121683 | MH124546 | – |
| L. abieticolens | G. abieticolens | 2865 | 115248 | T | Abies balsamea | Vermont, USA | D. Bergdalh | AF343701 | – | – | – |
| L. abietinum | G. abietina | 275 | 118590 | Picea engelmannii | Canada | A. Molnar | OM501383 | OM514713 | OM631763 | OM631594 | |
| L. altius | G. altior | 12471 | 123619 | T | Picea koraiensis | China | X.D. Zhou, Z.W. de Beer | HQ406851 | – | HQ406875 | – |
| L. bistatum | G. bistata | 3804 | 120192 | T | Pinus radiata | Japan | J.J. Kim | OM501388 | OM514721 | OM631795 | – |
| L. chlamydatum | G. chlamydata | 36631 | 128840 | Pityogenes chalcographus on Picea abies | Finland | Z.W. de Beer | JF279965 | – | JF280080 | – | |
| L. curviconidium | G. curviconidia | 12441 | 123617 | P | Ips typographus on Picea koraiensis | China | X.D. Zhou, Z.W. de Beer | OM501390 | OM514725 | OM631767 | – |
| L. curvisporum | G. curvispora | 17260 | 123914 | T | Picea abies | Norway | M.J. Wingfield, H. Solheim | OM501391 | OM514726 | EU979347 | – |
| L. engelmannii | = G. abietina | 759 | – | Picea engemannii | Canada | R.W. Davidson | – | OM631763 | OM631762 | – | |
| L. eucalyptophilum | G. eucalyptophila | 5211 | – | A | Eucalyptus urophylla × E. pellita | Democratic Republic of Congo | J. Roux | OM501393 | OM514728 | OM631769 | – |
| L. euphyes | G. euphyes | 259 | 109701 | T | Pinus strobus | New Zealand | M. Dick | AF343686 | – | – | – |
| L. fruticetum | G. fruticetum | 20605 | – | T | Picea engelmannii × P. glauca | Canada | S. Massoumi Alamouti | OM501394 | OM514730 | OM631770 | – |
| L. hughesii | G. hughesii | 4052 | 109709 | T | Aquilaria sp. | Vietnam | B. Lanchette | OM501395 | OM514732 | OM631772 | OM631598 |
| L. pistaciae | G. pistaciae | 12499 | 123626 | T | Pistacia chinensis | China | X.D. Zhou, Z.W. de Beer | HQ406846 | – | HQ406870 | – |
|
| |||||||||||
| Grosmannia: G. grandifoliae complex | |||||||||||
|
| |||||||||||
| G. grandifoliae | G. grandifoliae | 703 | ATCC28746 | T | Fagus grandifolia | Iowa, USA | R.W. Davidson | – | – | OM631771 | – |
| L. betulae | G. betulae | 43191 | 142734 | T | Scolytus ratzeburgi on Betula verrucosa | Poland | R. Jankowiak | KY801840 | – | KY801817 | – |
| L. pruni | G. pruni | 10418 | 120197 | P | Polygraphus ssiori on Prunus jamasakura | Japan | H. Masuya | OM501398 | OM514740 | OM631775 | – |
| L. trypodendri | G. trypodendri | 43182 | 142724 | T | Trypodendron domesticum on Fagus sylvatica | Norway | R. Jankowiak | KY801828 | – | KY801805 | – |
|
| |||||||||||
| Grosmannia: Group C | |||||||||||
|
| |||||||||||
| G. abieticola | G. abieticola | 17199 | – | T | Dryocoetes hectographus on Abies mariesii | Japan | Y. Yamaoka | OM501382 | OM514712 | OM631761 | – |
| L. innermongolicum | G. innermongolica | – | MUCL55158 | T | Ips subelongatus on Larix sp. | China | Q. Lu | KM236107 | – | KM981763 | – |
| L. taigense | G. taigensis | 36629 | – | P | Ips typographus on Picea abies | Russia | Z.W. de Beer | OM501400 | OM514744 | OM631777 | – |
| L. gestamen | G. gestamen | 38096 | CIEFAP453 | T | Nothofagus dombeyi | Argentina | A. de Errasti | KT362234 | KT362232 | KT381300 | – |
|
| |||||||||||
| Harringtonia gen. nov. (previously Raffalea lauricola complex) | |||||||||||
|
| |||||||||||
| R. aguacate | Har. aguacate | 38067 | 141672 | T | Persea americana | Florida, USA | C.L. Harmon | – | KJ909296 | – | – |
| R. aguacate * | Har. aguacate | – | Raff. sp. 272 | Persea americana | Florida, USA | C.L. Harmon | MT633065 | MT629748 | OM631783 | OM631613 | |
| R. brunnea | Har. brunnea | – | 378.68 | Monarthrum sp. | USA | L.R. Batra | – | EU177457 | – | – | |
| R. lauricola * | Har. lauricola | – | Raff. sp. 570 | Xyleborus sp. on Persea sp. | Florida, USA | J. Smith | MT633071 | MT629759 | OM631784 | OM631614 | |
| R. lauricola | Har. lauricola | 36261 | PL159 | Xyleborus glabratus | Georgia, USA | S. Fraedrich | OM501411 | MT629760 | OM631785 | OM631615 | |
|
| |||||||||||
| Hawksworthiomyces | |||||||||||
|
| |||||||||||
| Haw. crousii | Haw. crousii | 37531 | MUCL55928 | T | Bamboo chips | South Korea | J.J. Kim | KX396551 | KX396548 | OM652622 | OM631609 |
| Haw. hibbettii | Haw. hibbettii | 37663 | MUCL55929 | T | Trachymyrmex sp. | Texas, USA | U. Mueller | KX396550 | KX396547 | OM652623 | OM631610 |
| Haw. lignivorus* | Haw. lignivorus | 18600 | 119148 | T | Eucalyptus pole | South Africa | E.M. de Meyer | OM501410 | OM514756 | OM631782 | OM631611 |
| Haw. taylorii | Haw. taylorii | 20741 | MUCL55927 | T | Eucalyptus pole | South Africa | E.M. de Meyer | KX396549 | KX396546 | OM652624 | OM631612 |
| ‘Haw. sequentia ENAS’ | ‘Haw. sequentia ENAS’ | – | nik62104a_03C_19 | T | Picea log | Sweden | – | HQ611296 | – | – | – |
|
| |||||||||||
| Heinzbutinia gen. nov. | |||||||||||
|
| |||||||||||
| O. grandicarpum | He. grandicarpa | 1600 | 250.88 | T | Quercus robor | Poland | H. Butin | OM501412 | OM514757 | OM631786 | OM631616 |
| O. longicollum | He. longicolla | – | JCM10198 | T | Quercus mongolica var. grosseserrata infested by Platypus quercivorus | Japan | H. Masuya | – | – | – | – |
| O. microsporum | He. microspora | 17152 | 440.69 | T | Quercus sp. | Virginia, USA | R.W. Davidson | – | OM514758 | OM631787 | OM631617 |
| O. solheimii # | He. solheimii | 52050 | 144881 | T | Anisandrus dispar infesting Quercus robur | Poland | P.Wieczorek | MH283134 | – | MH283488 | – |
|
| |||||||||||
| Intubia | |||||||||||
|
| |||||||||||
| I. macrotermitinarum # | I. macrotermitinarum | 46496 | 141560 | T | Termitomyces fungal comb of Macrotermes natalensis | South Africa | W.J. Nel | MT637025 | – | – | – |
| I. oerlemansii # | I. oerlemansii | 47048 | 141564 | T | Termitomyces fungal comb of Macrotermes natalensis | South Africa | W.J. Nel | MT637024 | – | – | – |
|
| |||||||||||
| Jamesreidia gen. nov. (previously O. tenellum complex) | |||||||||||
|
| |||||||||||
| O. coronatum | J. coronata | 43836 | UAMH9685 | T | Pinus sp. | Canada | A. Olchowecki | OM501413 | OM514759 | OM631788 | OM631618 |
| O. nigricarpum | J. nigricarpa | 651 | 638.66 | Pseudotsuga menziesii | Idaho, USA | R.W. Davidson | AY280490 | DQ294356 | – | – | |
| O. rostrocoronata | J. rostrocoronata | 456 | 434.77 | Pulpwood chips | Wisconsin, USA | R.W. Davidson | AY194509 | KX590871 | – | – | |
| O. tenellum | J. tenella | 37439 | 189.86 | Pinus sp. | Colorado, USA | R.W. Davidson | OM501414 | OM514760 | OM631789 | – | |
|
| |||||||||||
| Leptographium: L. clavigerum complex | |||||||||||
|
| |||||||||||
| G. aurea | L. aureum | 667 | 438.69 | A | Pinus contorta var. latifolia | Canada | R. W. Davidson | OM501387 | OM514720 | OM631793 | OM631621 |
| G. clavigera * | L. clavigerum | – | SL-Kw1407 | Sapwood associated with Dendroctonus ponderosae | Canada | S. Lee | OM501421 | OM514723 | OM631799 | OM631624 | |
|
| |||||||||||
| G. robusta | L. robustum | 668 | – | T | Pinus ponderosa | Idaho, USA | R.C.R. Jeffrey, R.W. Davidson | – | AY544619 | JF798465 | – |
| L. longiclavatum | L. longiclavatum | 20608 | – | P | Pinus contorta | Canada | S. Lee | – | OM514771 | – | – |
| L. pyrinum | L. pyrinum | 509 | 120181 | T | Dendroctonus adjunctus | USA | R.W. Davidson | OM501445 | OM514781 | OM631819 | OM631642 |
| L. terebrantis | L. terebrantis | 29841 | 337.70 | T | Dendoctronus terebrantis | Louisiana, USA | S.J. Baras | JF798477 | – | JF798470 | – |
| L. tereforme | L. tereforme | 37432 | 125736 | T | Hylurgus ligniperda | California, USA | S.J. Kim | OM501455 | OM514786 | OM631828 | OM631649 |
| L. wingfieldii | L. wingfieldii | 4741 | – | Pinus densiflora | Japan | H. Masuya | OM501461 | OM514789 | OM631834 | OM631654 | |
|
| |||||||||||
| Leptographium: L. galeiforme complex | |||||||||||
|
| |||||||||||
| G. galeiformis* | L. galeiforme | 5290 | 115711 | E | Pinus sylvestris | Scotland | T. Kirisits | OM501428 | OM514731 | OM631806 | OM631631 |
| G. radiaticola | L. radiaticola | 9482 | – | Hylurgus ligniperda on Pinus radiata | Chile | X.D. Zhou | OM501446 | OM514742 | OM631820 | OM631643 | |
| L. doddsii # | L. doddsii | 34479 | 143470 | T | Dendroctonus valens | California, USA | M.J. Wingfield | MT637215 | MT637212 | MT637205 | – |
| L. gordonii # | L. gordonii | 34619 | 143477 | T | Dendroctonus valens in Pinus resinosa | New Hampshire, USA | M.J. Wingfield | MT637226 | MT637213 | MT637207 | – |
| L. koraiense | L. koraiense | 44461 | 141898 | T | Ips typographus on Pinus koraiensis | China | R. Chang | MH144096 | – | MH124372 | – |
| L. owenii # | L. owenii | 34448 | 143467 | T | Dendroctonus valens | California, USA | M.J. Wingfield | MT637217 | KF515912.1 | KF515884.1 | – |
| L. seifertii # | L. seifertii | 34620 | 143478 | T | Dendroctonus valens on Pinus resinosa | New Hampshire, USA | M.J. Wingfield | MT637224 | KF515911.1 | KF515885.1 | – |
|
| |||||||||||
| Leptographium: L. lundbergii complex | |||||||||||
|
| |||||||||||
| G. koreana | L. koreanum | 14199 | KUC2078 | T | Tomicus piniperda on Pinus koraiensis | South Korea | J.J. Kim | OM501431 | OM514733 | OM631808 | OM631633 |
| G. yunnanensis | L. yunnanense | 5152 | – | Pinus yunnanensis | China | X.D. Zhou | AY707207 | – | – | – | |
| L. absconditum | L. absconditum | 39763 | 136527 | T | Orthotomicus laricis on Pinus nigra | Spain | P. Romón | OM501415 | OM514761 | OM631790 | – |
| L. celere | L. celere | 12422 | 123628 | T | Pinus semaonensis | China | X.D. Zhou, Z.W. de Beer | HQ406834 | – | HQ406858 | – |
| L. conjunctum | L. conjunctum | 12473 | 123631 | T | Pinus yunnanensis | China | X.D. Zhou, Z.W. de Beer | HQ406831 | – | HQ406855 | – |
| L. lundbergii * | L. lundbergii | 2190 | 138716 | Pinus sylvestris | Norway | H. Roll-Hansen | OM501432 | OM514772 | OM631809 | OM631634 | |
| L. manifestum | L. manifestum | 12436 | 123622 | T | Larix olgensis | China | X.D. Zhou, Z.W. de Beer | HQ406839 | – | HQ406863 | – |
| L. pinicola | L. pinicola | 2398 | – | T | Hylastes sp. on Pinus sp. | Canada | J. Juzwik | OM501439 | OM514775 | – | – |
| L. shansheni | L. shansheni | 44462 | 141895 | T | Ips typographus on Picea sp. | China | R. Chang, S.F. Chen | MH144097 | – | MH124373 | – |
| L. sosnaicola # | L. sosnaicola | 52084 | 147023 | T | Pinus sylvestris | Poland | D. Jazłowiecka | MT210337 | MT210353 | MT210397 | – |
| L. truncatum | L. truncatum | 28 | 929.85 | T | Pinus taeda | South Africa | M.J. Wingfield | DQ062052/AY935626 | – | DQ062019 | – |
| L. wushanense | L. wushanense | – | YMF1.04936 | T | Pinus sp. | China | J. Lu | MG878407 | – | MG878409 | – |
|
| |||||||||||
| Leptographium: L. olivaceum complex | |||||||||||
|
| |||||||||||
| G. cucullata * | L. cucullatum | 1035 | 218.83 | T | Ips typographus | Norway | H. Solheim | OM501423 | OM514724 | OM631801 | OM631626 |
| G. davidsonii | L. davidsonii | – | YCC611 | Logs of Larix kaempferi infested with Ips subelongatus | Japan | Y. Yamaoka | GU134165 | – | – | – | |
| G. olivacea | L. olivaceum | 23348 | – | Pinus sylvestris | Finland | Z.W. de Beer, P. Niemelä | OM501434 | OM514735 | OM631811 | – | |
| G. olivaceapini | L. olivaceapini | 116 | 504.86 | E | Pinus ponderosa in Dendroctonus sp. | Arizona, USA | T. Hinds | OM501433 | OM514736 | OM631810 | OM631635 |
| G. sagmatospora | L. sagmatosporum | 43846 | UAMH6971 | Pinus strobus | Canada | B. Grylls, K. Seifert | OM501449 | OM514743 | OM631823 | – | |
| G. vesca | L. vescum | 34186 | 800.73 | Ips pilifrons, Dendroctonus engelmanni in Picea engelmannii | Colorado, USA | F.F. Lombard, R. W. Davidson | OM501457 | OM514745 | – | – | |
| L. brevicolle | L. brevicolle | – | 150.78 | Beetle gallery in Populus tremuloides | Colorado, USA | R.W. Davidson | MH055549 | AF155670 | MH055635 | – | |
| L. breviuscapum | L. breviuscapum | 38888 | 136507 | T | Picea crassifolia infested with Polygraphus poligraphus | China | M.L. Yin, X.D. Zhou | OM501419 | OM514763 | MN517742 | – |
| L. conplurium | L. conplurium | 23289 | 128834 | P | Pinus sylvestris | Finland | Z.W. de Beer, P. Niemelä | OM501422 | OM514765 | OM631800 | OM631625 |
| L. duchongi | L. duchongi | 44455 | 141897 | T | Ips typographus on Pinus koraiensis | China | R. Chang | MH144122 | – | MH124398 | – |
| L. erubescens | L. erubescens | 40672 | 278.54 | E | Pinus sylvestris | Sweden | A. Mathiesen-Käärik | OM501425 | OM514767 | OM631803 | OM631628 |
| L. flavum | L. flavum | 51797 | 144099 | T | Quercus robur | Poland | R. Jankowiak | MH055548 | – | MH055634 | – |
| L. francke-grosmanniae | L. francke-grosmanniae | 445 | 356.77 | T | Hylecoetus dermestoides on Quercus sp. | Germany | H. Francke-Grosmann | OM501427 | OM514768 | OM631805 | OM631630 |
| L. sylvestris | L. sylvestris | 23300 | 128833 | P | Picea abies | Finland | Z.W. de Beer, P. Niemelä | OM501454 | OM514777 | OM631827 | – |
| L. pseudoalbum | L. pseudoalbum | 40671 | 276.54 | T | Blastophagus piniperda in Pinus sylvestris | Sweden | A. Mathiesen-Käärik | MN516723 | MN516723 | MN517755 | OM631641 |
| L. raffai # | L. raffai | 34451 | 143468 | T | Dendroctonus valens | California, USA | M.J. Wingfield | MT637219 | MT637211 | MT637206 | – |
| L. rhizoidum | L. rhizoidum | 22809 | 136512 | T | Hylastes ater on Pinus radiata | Spain | P. Romón, X.D. Zhou | MN516724 | MN516724 | MN517748 | OM631644 |
| L. tardum | L. tardum | 51789 | 144091 | T | Trypodendron domesticum on Fagus sylvatica | Poland | R. Jankowiak | MH055529 | – | MH055615 | – |
| L. vulnerum | L. vulnerum | 51794 | 144096 | T | Fagus sylvatica | Poland | R. Jankowiak | MH055534 | – | MH055620 | – |
| L. xiningense# | L. xiningense | 38891 | 136509 | T | Polygraphus poligraphus in Picea crassifolia | China | M.L. Yin, X.D. Zhou | MN516732 | MN516732 | MN517752 | OM631637 |
|
| |||||||||||
| Leptographium: L. piceiperdum complex | |||||||||||
|
| |||||||||||
| G. aenigmatica | L. aenigmaticum | 2199 | – | T | Ips typographus japonicus on Picea jezoensis | Japan | Y. Yamaoka | OM501416 | OM514716 | OM631791 | OM631619 |
| G. europhioides | L. europhioides | 2811 | 115245 | Picea rubens | New York, USA | T. Harrington | OM501426 | OM514729 | OM631804 | OM631629 | |
| G. laricis | L. laricis | 1913 | 120188 | Larix sp. | Japan | Y. Yamaoka | – | DQ294393 | – | – | |
| G. piceiperdum * | L. piceiperdum | 16768 | 138719 | Picea glauca | Canada | K. Harrison | OM501435 | OM514738 | OM631812 | OM631636 | |
| G. pseudoeurophioides | L. pseudoeurophioides | – | WIN(M)42 | – | Canada | J. Reid | EU879136 | – | – | – | |
| L. heilongjiangense | L. heilongjiangense | 44456 | 141702 | T | Ips typographus on Pinus koraiensis | China | R. Chang | MH144098 | – | MH124374 | – |
| L. zhangii | L. zhangii | – | MUCL55162 | T | Ips subelongatus on Larix gmelinii | China | X. Liu | KM236108 | – | KM974275 | – |
|
| |||||||||||
| Leptographium: L. procerum complex | |||||||||||
|
| |||||||||||
| L. bhutanense | L. bhutanense | 18649 | 122076 | T | Pinus wallichiana | Bhutan | M.J. Wingfield | OM501418 | OM514762 | OM631794 | – |
| L. gracile | L. gracile | 12398 | 123623 | T | Pinus armandii | China | X.D. Zhou | OM501430 | OM514769 | – | – |
| L. latens | L. latens | 12438 | 124023 | T | Picea koraiensis | China | X.D. Zhou, Z.W. de Beer | HQ406845 | – | HQ406869 | – |
| L. longiconidiophorum | L. longiconidiophorum | 2004 | 135624 | T | Hylastes sp. on Pinus densiflora | Japan | M.J. Wingfield | KM491421 | – | KM491471 | – |
| L. peucophilum | L. peucophilum | 2876 | 120191 | Picea sp. | New York, USA | D. Bergdalh | – | – | – | – | |
| L. pini-densiflorae | L. pini-densiflorae | 5157 | 115261 | T | Pinus densiflora | Japan | H. Masuya | OM501438 | OM514774 | OM631815 | |
| L. procerum * | L. procerum | 34542 | 138288 | E | Dendroctonus valens on Pinus resinosa | Maine, USA | M.J. Wingfield | OM501442 | OM514778 | OM631816 | OM631639 |
| L. profanum | L. profanum | 10552 | 120307 | T | Carya sp. | Alabama, USA | L. Eckhardt | OM501443 | OM514779 | OM631817 | OM631640 |
| L. sibiricum | L. sibiricum | 4481 | 115260 | T | Monochamus urussoni on Abies sibitica | Russia | V.P. Vetrova | OM501451 | OM514783 | – | – |
| L. sinense | L. sinense | 38171 | 316515 | P | Pinus elliottii | China | M. Yin, R. Chang, X.D. Zhou | OM501452 | OM514784 | OM631825 | OM631647 |
| L. sinoprocerum | L. sinoprocerum | 29990 | MUCL46532 | T | Pinus tabuliformis | China | Q. Lu | OM501453 | OM514785 | OM631826 | OM631648 |
| L. yichunense | L. yichunense | 44464 | 141705 | T | Ips typographus on Picea sp. | China | R. Chang, S.F. Chen | MH144114 | – | MH124390 | – |
|
| |||||||||||
| Leptographium: L. serpens complex | |||||||||||
|
| |||||||||||
| G. alacris | L. alacre | 621 | 128830 | T | Pinus pinaster | Portugal | M. de Famtima Moniz | OM501417 | OM514717 | OM631792 | OM631620 |
| G. serpens | L. serpens | 305 | 141.36 | T | Pinus sylvestris | Italy | G. Goidànich | OM501450 | – | OM631824 | OM631646 |
| L. castellanum | L. castellanum | 2320 | 128698 | P | Pinus occidentalis | Dominican Republic | R. Webb | OM501420 | OM514764 | OM631798 | OM631623 |
| L. gibbsii | L. gibbsii | 36371 | – | T | Hylastes opacus on Pinus sylvestris | England, UK | J. Gibbs | OM501429 | – | OM631807 | OM631632 |
| L. rhodanense | L. rhodanense | 16438 | 138284 | T | Pinus sylvestris | Switzerland | U. Heiniger | OM501448 | – | OM631822 | OM631645 |
| L. yamaokae | L. yamaokae | 4726 | 129732 | T | Pinus densiflora | Japan | H. Masuya | – | – | OM631835 | OM631655 |
|
| |||||||||||
| Leptographium: L. wageneri complex | |||||||||||
|
| |||||||||||
| G. wageneri * | L. wageneri var. ponderosum | 279 | – | Pinus sp. | USA | T. Harrington | OM501458 | OM514746 | OM631831 | OM631651 | |
| L. douglasii * | L. douglasii | 2076 | – | P | Pseudotsuga menziesii | New Mexico, USA | M. Midke | OM501424 | OM514766 | OM631802 | OM631627 |
| L. neomexicanum | L. neomexicanum | 2079 | 168.93 | T | Pinus ponderosa | New Mexico, USA | T. Harrington, W Livingston | AY553382 | – | AY536176 | – |
| L. reconditum | L. reconditum | 15 | 116348 | Zea mays | South Africa | W. Jooste | AF343690 | – | AY536177 | – | |
| L. wageneri var. pseudotsugae* | L. wageneri var. pseudotsugae | 154 | 115246 | Pseudotsuga menziesii | USA | T. Harrington | OM501459 | OM514747 | OM631832 | OM631652 | |
| L. wageneri var. wageneri* | L. wageneri var. wageneri | 53 | 139665 | Pinus ponderosa | California, USA | T. Harrington | OM501460 | OM514788 | OM631833 | OM631653 | |
|
| |||||||||||
| Leptographium: Group A | |||||||||||
|
| |||||||||||
| L. pineti | L. pineti | 3837 | 115257 | Pinus sp. | Indonesia | M.J. Wingfield | – | – | OM631814 | OM631638 | |
| L. ningerense | L. ningerense | 41786 | 139663 | T | Coccotrypes cyperi on Pinus kesiya | China | S. Taerum | MG205674 | – | MG205765 | – |
|
| |||||||||||
| Leptographium: Group B | |||||||||||
|
| |||||||||||
| G. cainii | L. cainii | 24907 | – | Picea sp. | Canada | C. Breuil | OM501389 | OM514722 | OM631797 | OM631622 | |
|
| |||||||||||
| Leptographium Incertae sedis (based on our data) | |||||||||||
|
| |||||||||||
| G. huntii | G. huntii | 2868 | 118780 | Pinus strobus | North Carolina, USA | V. Lackner | AY553394 | – | DQ354938 | – | |
| G. leptographioides | G. leptographioides | 481 | 144.59 | Quercus sp. | New York, USA | R.W. Davidson | – | DQ294382 | – | – | |
| G. truncicola | G. truncicola | – | – | Dendroctonus sp. on Picea sp. | USA | – | – | – | – | – | |
| L. albopini | L. albopini | ^26 | – | Hylastes sp. on Pinus sp. | USA | – | AF343695 | – | – | – | |
| L. alethinum | L. alethinum | ^3763 | – | Galleries of Hylobus abietis on log of Pinus nigra var. maritima | England, UK | A. Uzunovic | AY553391 | – | AY536185 | ||
| L. calophylli | L. calophylli | 752 | 277.51 | Cryphalus sp. on Calophyllum sp. | Mauritius | J.A. Stevenson | MH856855 | MH868375 | – | – | |
| L. microsporum | L. microsporum | – | – | T | Fagus sp. | Mississippi, USA | R.W. Davidson | – | – | – | – |
| L. obscurum | L. obscurum | 37429 | 125.39 | Pinus sp. | USA | R.W. Davidson | – | – | – | – | |
| L. pityophilum | L. pityophilum | 2840 | 109706 | Pinus nigra | Italy | S. Frisullo | OM501441 | OM514776 | – | – | |
| L. rostrocylindricum | L. rostrocylindricum | – | – | Quercus sp. | Connecticut, USA | R.W. Davidson | – | – | – | – | |
|
| |||||||||||
| Leptographium & Grosmannia incertae sedis (based on our data) | |||||||||||
|
| |||||||||||
| L. guttulatum | L. guttulatum | 1310 | 120185 | T | Tomicus piniperda on Pinus sylvestris | England, UK | J. Gibbs | – | OM514770 | – | – |
|
| |||||||||||
| Lineage III | |||||||||||
|
| |||||||||||
| G. crassivaginata | G. crassivaginata | 90 | 120178 | – | – | T. Hinds | AF343673 | – | – | – | |
| L. alneum# | L. alneum | 52076 | 144901 | T | Dryocoetes alni infesting Populus tremula | Poland | K. Miœkiewicz | MN900997 | MN900997 | MN901024 | – |
| L. piriforme | L. piriforme | 25381 | UAMH10681 | P | Beetle caught in a trap baited with coyote dung | Canada | M.D. Greif | OM501440 | – | – | – |
|
| |||||||||||
| Lineage VI | |||||||||||
|
| |||||||||||
| L. verrucosum | L. verrucosum | 17160 | 112420 | T | Xyleborus dryographus | Germany | H. Gebhardt | OM501456 | OM514787 | OM631830 | OM631650 |
|
| |||||||||||
| Masuyamyces gen. nov. | |||||||||||
|
| |||||||||||
| O. ambrosium | M. ambrosius | 1024 | 210.64 | Wood of Pinus sylvestris | Netherlands | J.A. von Arx | OM501465 | OM514793 | OM631836 | OM631656 | |
| O. acarorum | M. acarorum | 41850 | 139748 | T | Orthotomicus angulatus on Pinus kesiya | China | S. Taerum | MG205657 | – | – | – |
| O. botuliforme | M. botuliformis | 14493 | – | Cryphalus jeholensis | Japan | H. Masuya | OM501471 | OM514799 | OM631837 | – | |
| O. jilinense | M. jilinensis | 40491 | 141894 | T | Ips typographus on Picea sp. | China | X. D. Zhou | MH144094 | – | MH124370 | – |
| O. lotiforme # | M. lotiformis | – | MUCL55165 | T | Ips subelongatus on Pinus sylvestris var. mongolica | China | X. Meng | MK748185 | – | – | – |
| O. massoniana | M. massonianae | – | MUCL55179 | T | Monochamus sp. on Pinus sp. | China | Q. Lu | KY094067 | – | – | – |
| O. pallidulum | M. pallidulus | 23278 | 128118 | T | Pinus sylvestris | Russia | Z.W. de Beer, P. Niemelä | HM031510 | – | – | – |
| O. saponiodorum | M. saponiodorus | 28135 | 128302 | Pinus sylvestris | Russia | R. Linnakoski | OM501512 | OM514838 | OM631838 | OM631657 | |
|
| |||||||||||
| Ophiostoma: O. clavatum complex | |||||||||||
|
| |||||||||||
| O. ainoae | O. ainoae | 1037 | 205.83 | T | Picea abies | Norway | H. Solheim | OM501463 | OM514791 | – | – |
| O. brevipilosi | O. brevipilosi | 41873 | 139660 | T | Tomicus brevipilosus on Pinus kesiya | China | S. Taerum | MG205660 | – | MG205732 | – |
| O. brunneociliatum | O. brunneociliatum | 5212 | – | Larix sp. | Scotland, UK | T. Kirisits | KU184422 | – | KU184379 | – | |
| O. brunneolum | O. brunneolum | 23145 | – | A | Picea abies | Russia | J. Ahtiainen, P. Niemelä | OM501472 | OM514800 | OM631843 | OM631663 |
| O. clavatum | O. clavatum | 37983 | 141080 | E | Ips acuminatus on Pinus sylvestris | Sweden | C. Villari | OM501477 | OM514805 | OM631848 | – |
| O. hongxingense # | O. hongxingense | – | CFCC52695 | T | Ips subelongatus on Larix gmelinii | China | Q. Lu | MK748194 | – | MN896068 | – |
| O. japonicum | O. japonicum | 2202 | YCC099 | T | Ips typographus japonicus on Picea jezoensis | Japan | Y. Yamaoka | OM501492 | – | OM631855 | OM631673 |
| O. jiamusiensis | O. jiamusiensis | 40512 | 141893 | T | Ips typographus on Picea sp. | China | X.D. Zhou | MH144064 | – | MH124343 | – |
| O. macroclavatum | O. macroclavatum | 23115 | 141081 | T | Pinus sylvestris | Russia | Z.W. de Beer | HM031499 | – | KU094765 | – |
| O. peniculi # | O. peniculi | – | CFCC52687 | T | Ips subelongatus infesting Larix gmelinii | China | Q. Lu | MK748198 | – | MN896063 | – |
| O. poligraphi | O. poligraphi | 38899 | 136517 | T | Polygraphus poligraphus on Picea crassifolia | China | M. Yin, S. Taerum, X.D. Zhou | OM501507 | OM514832 | OM631871 | OM631688 |
| O. pseudocatenulatum | O. pseudocatenulatum | 43103 | 141276 | T | Ips cembrae on Larix decidua | Poland | R. Jankowiak | KU094686 | – | KU094774 | – |
| O. shangrilae | O. shangrilae | 38901 | 136519 | T | Ips shangrila on Picea purpurea | China | M. Yin, S. Taerum, X.D. Zhou | OM501514 | OM514840 | OM631879 | OM631695 |
| O. songshui | O. songshui | 44473 | 141707 | T | Ips typographus on Picea sp. | China | R. Chang, S.F. Chen | MH144065 | – | MH124344 | – |
| O. subelongati # | O. subelongati | – | CFCC52693 | T | Ips subelongatus infesting Larix gmelinii | China | Q. Lu | MK748200 | – | MN896064 | – |
| O. tapionis | O. tapionis | 23266 | 128122 | P | Picea abies | Russia | Z.W. de Beer, P. Niemelä | OM501516 | OM514842 | OM631881 | OM631697 |
| Ophiostoma sp. 3 (Hyalorhinocladiella sp. 2) | Ophiostoma sp. 3 (Hyalorhinocladiella sp. 2) | 43851 | UAMH10642 | Ips sp. | Canada | S. Massoumi Alamouti | OM501524 | OM514851 | OM631888 | – | |
| Ophiostoma sp. 4 (Hyalorhinocladiella sp. 1) | Ophiostoma sp. 4 (Hyalorhinocladiella sp. 1) | 43848 | UAMH10639 | Ips sp. | Canada | S. Massoumi Alamouti | OM501525 | OM514852 | OM631889 | OM631704 | |
|
| |||||||||||
| Ophiostoma: O. ips complex | |||||||||||
|
| |||||||||||
| O. adjuncti | O. adjuncti | 1025 | 314.77 | T | Stained sapwood in Pinus ponderosa | New Mexico, USA | R.W. Davidson | OM501462 | OM514790 | – | OM631658 |
| ‘O. arborea’ | ‘O. arborea’ | – | WIN(M)69-23 | Picea mariana | Canada | A. Olchowecki, J. Reid | – | – | – | – | |
| O. bicolor | O. bicolor | 23169 | – | Pinus sylverstris | Russia | J. Ahtiainen, P. Niemelä | OM501470 | OM514798 | OM631842 | OM631662 | |
| O. columnare | O. columnare | – | WIN(M)71-27 | Pinus banksiana | Canada | A. Olchowecki, J. Reid | – | – | – | – | |
| O. fuscum | O. fuscum | 23195 | 128124 | P | Pinus sylverstris | Finland | Z.W. de Beer, P. Niemelä | OM501483 | OM514810 | – | OM631670 |
| O. gilletteae # | O. gilletteae | 30681 | 143458 | T | Dendroctonus valens | Washington, USA | N. Gillette | MT637227 | – | – | – |
| O. guatemalensis | O. guatemalensis | 44221 | – | T | Pinus patula | Guatemala | I. Barnes, J. Garnas | – | – | – | – |
| O. hyalothecium | O. hyalothecium | – | ATCC28825 | Pinus contorta | Wyoming, USA | R.W. Davidson | – | AF137284 | – | – | |
| Tuberculariella ips | O. ips-like sp. 6 | 14175 | 435.34 | T | Ips sp. on Pinus sp. | Minnesota, USA | J.G. Leach | OM501487 | OM514813 | OM631854 | OM631672 |
| O. ips * | O. ips | 19371 | 138721 | Pinus taeda | Louisiana, USA | X.D. Zhou | OM501486 | OM514812 | OM631853 | OM631671 | |
| O. manchongi # | O. manchongi | 41954 | 141906 | Uropodoidea sp. in Ips shangrila gallery on Picea purpurea | China | S.J. Taerum | MH121662 | – | – | – | |
| O. montium | O. montium | 15419 | – | Pinus contorta | Idaho, USA | B. Bentz | OM501498 | OM514822 | OM631861 | OM631678 | |
| O. pseudobicolor # | O. pseudobicolor | – | CFCC52683 | T | Ips subelongatus in Larix gmelinii | China | Q. Lu | MK748188 | – | – | – |
| O. pulvinisporum | O. pulvinisporum | 9022 | 118673 | T | Pinus pseudostrobus | Mexico | X.D. Zhou | OM501509 | OM514835 | OM631874 | OM631691 |
|
| |||||||||||
| Ophiostoma: O. minus complex | |||||||||||
|
| |||||||||||
| ‘O. album’ | ‘O. album’ | – | MUCL55189 | T | Monochamus alternatus gallery on Pinus massoniana | China | Q. Lu, Y.Y. Lun | KY094073 | – | – | – |
| O. exiguum | O. exiguum | – | – | Pinus virginiana | West Virginia, USA | G.G. Hedgcock | – | – | – | – | |
| O. kryptum | O. kryptum | – | 116190 | T | Tetropium on Picea | Austria | T. Kirisits | AY305685 | – | – | – |
| O. minus | O. minus | 43873 | UAMH4917 | Dendroctonus ponderosae on Pinus flexillis | Canada | P. Muruyama | OM501497 | OM514821 | OM631860 | OM631677 | |
| ‘O. olgensis’ | ‘O. olgensis’ | – | CXY1410 | T | Ips subelongatus on Larix olgensis | China | Q. Lu | KU551303 | – | KU551297 | – |
| O. minus (in Europe O. pini) | O. minus (in Europe O. pini) | 43346 | – | Pinus sylvestris | Poland | T. Tomasz | – | – | – | – | |
| O. pseudominus | O. pseudominus | 43878 | UAMH9721 | Pseudotsuga menziesii | Canada | J. Reid, B Reid | OM501508 | OM514834 | OM631873 | OM631690 | |
| O. pseudotsugae | O. pseudotsugae | – | D48/3 | – | Canada | H. Solheim | AY542501 | – | – | – | |
| O. wuyingense | O. wuyingense | 44474 | 141706 | T | Ips typographus on Picea sp. | China | R. Chang, S.F. Chen | MH144061 | – | MH124340 | – |
|
| |||||||||||
| Ophiostoma: O. piceae complex | |||||||||||
|
| |||||||||||
| O. arduennense (= O. distortum) | O. distortum | 40266 | MUCL44866 | T | Fagus sylvatica | Belgium | F.X. Carlier, T. Defrance | OM501468 | OM514796 | KU184376 | – |
| O. brunneum | O. brunneum | 1027 | 161.61 | A | Standing dead Abies sp. | Colorado, USA | R.W. Davidson | OM501473 | OM514801 | OM631844 | OM631664 |
| O. canum (Pachnodium) | O. canum (Pachnodium) | 5023 | 118668 | Tomicus minor on Pinus sylvestris | Austria | T. Kirisits | OM501474 | OM514802 | OM631845 | OM631665 | |
| O. cupulatum | O. cupulatum | 37441 | 102358 | T | Pseudotsuga menziesii | Washington, USA | T. Harrington | OM501479 | OM514806 | OM631849 | OM631667 |
| O. distortum | O. distortum | 467 | 397.77 | Picea engelmannii | Arizona, USA | R.W. Davidson | OM501481 | OM514808 | OM631850 | OM631668 | |
| O. flexuosum | O. flexuosum | 907 | 208.83 | T | Picea abies | Norway | H. Solheim | OM501482 | OM514809 | OM631851 | OM631669 |
| O. floccosum | O. floccosum | 34182 | 799.73 | T | Wood | Sweden | A. Mathiesen-Käärik | KU184431 | – | KU184388 | – |
| O. genhense # | O. genhense | – | CFCC52675 | T | Ips subelongatus infesting Larix gmelinii | China | Q. Lu | MK748199 | – | MN896074 | – |
| O. kunlunense # | O. kunlunense | 41927 | 141903 | T | Uropodoidea sp. in gallery of Ips shangrila on Picea purpurea | China | S.J. Taerum | MH121648 | – | MH124515 | – |
| O. micantis | O. micantis | 38903 | 136523 | T | Dendroctonus micans on Picea crassifolia | China | M. Yin, S. Taerum, X.D. Zhou | OM501496 | OM514820 | OM631859 | OM631676 |
| O. multisynnematum # | O. multisynnematum | – | CFCC52677 | Ips subelongatus infesting Larix gmelinii | China | Q. Lu | MK748196 | – | MN896071 | – | |
| O. nikkoense | O. nikkoense | 17193 | YCC430 | T | Polygraphus proximus | Japan | Y. Yamaoka | OM501500 | OM514824 | OM631863 | OM631679 |
| O. nitidi | O. nitidi | 38907 | 136525 | T | Picea crassifolia | China | M. Yin, S. Taerum, X.D. Zhou | OM501501 | OM514825 | OM631864 | OM631680 |
| O. perfectum | O. perfectum | 17153 | 600.85 | Cat hairs | Germany | G.S. de Hoog | OM501506 | OM514831 | OM631868 | OM631685 | |
| O. peregrinum | O. peregrinum | – | CIEFAP426 | T | Pinus radiata | Argentina | A. de Errasti | MG345116 | – | – | – |
| O. piceae * | O. piceae | – | UAMH11346 | Pine saw timber | Canada | A. Uzunovic | MT633062 | MT629745 | OM631869 | OM631686 | |
| O. pityokteinis # | O. pityokteinis | 52056 | 144879 | T | Pityokteines curvidens infesting Abies alba | Poland | P. Majka | MH837046 | – | MH837055 | – |
| O. qinghaiense | O. qinghaiense | 38902 | 136521 | T | Picea crassifolia | China | M. Yin, S. Taerum, X.D. Zhou | OM501510 | OM514836 | OM631875 | – |
| O. rachisporum | O. rachisporum | 28021 | – | Picea abies | Russia | R. Linnakoski | OM501511 | OM514837 | OM631877 | OM631693 | |
| O. rufum # | O. rufum | 52062 | 144871 | T | Ips cembrae galleries on Larix decidua | Czech Republic | K. Lukášová | MH837040 | – | KY568647 | – |
| O. setosum | O. setosum | 27833 | AU160-38 | T | Tsuga sp. | Canada | A. Uzunovic | OM501513 | OM514839 | OM631878 | OM631694 |
| O. shanziensis # | O. shanziensis | 48329 | MUCL46456 | T | Phloem adjacent to Dendroctonus valens gallery in Pinus tabuliformis | China | Q. Lu, C. Decock | MT637221 | – | – | – |
| O. sugadairense | O. sugadairense | – | YCC589 | T | Polygraphus sp. on Larix sp. | Japan | Y. Yamaoka | LC090227 | – | AB934343 | – |
| O. taphrorychi # | O. taphrorychi | 52045 | 144891 | T | Taphrorychus bicolor infesting Fagus sylvatica | Poland | P. Bilánski | MH837052 | – | MH837062 | – |
| O. typographi | O. typographi | 44483 | 141709 | T | Picea sp. | China | R. Chang, S.F. Chen | MH144059 | – | MH124337 | – |
| O. torulosum | O. torulosum | 10574 | CTK106 | Fagus sylvatica | Germany | T. Kirisits | OM501518 | OM514844 | OM631883 | OM631699 | |
| O. xinganense # | O. xinganense | – | CFCC52679 | Ips subelongatus infesting Larix gmelinii | China | Q. Lu | MK748186 | – | MN896078 | – | |
|
| |||||||||||
| Ophiostoma: O. pluriannulatum complex | |||||||||||
|
| |||||||||||
| O. californicum | O. californicum | 143 | 796.73 | Prunus domestica | California, USA | R.W. Davidson | KU756602 | AF137280 | – | – | |
| O. carpenteri | O. carpenteri | 44611 | UAMH9696 | P | Trypodendron lineatum | Oregon, USA | S. Carpenter | OM501475 | OM514803 | OM631846 | – |
| O. conicola | O. conicola | 2561 | 127.89 | T | Pinus cembroides | Mexico | H. Butin | OM501478 | – | – | – |
| O. longiconidiatum | O. longiconidiatum | 14265 | 121350 | A | Faurea saligna | South Africa | G. Kamgan Nkuekam | OM501494 | OM514818 | OM631857 | – |
| O. longirostellatum | O. longirostellatum | – | 134.51 | T | Quercus sp. | Scotland, UK | B.K. Bakshi | – | AF155688 | – | – |
| O. multiannulatum | O. multiannulatum | 2567 | 357.77 | Pinus sp. | North Carolina, USA | R.W. Davidson | OM501499 | OM514823 | OM631862 | – | |
| O. novae-zelandiae | O. novae-zelandiae | – | CIEFAP423 | T | Dead fallen wood | New Zealand | – | KT362249 | KT362226 | – | – |
| O. palustre | O. palustre | 44403 | 140596 | T | Barringtonia racemosa | South Africa | J.A. Osorio | KU865593 | – | – | – |
| O. pluriannulatum | O. pluriannulatum | – | MUCL18372 | Quercus sp. | Oregon, USA | R.W. Davidson | AY934517 | DQ294365 | – | – | |
| O. populinum | O. populinum | 50149 | 212.67 | T | Populus tremuloides | Colorado, USA | R.W. Davidson | – | OM514833 | OM631872 | OM631689 |
| O. retusi | O. retusi | ATCC22324 | Populus sp. | Colorado, USA | R.W. Davidson | – | L05783 | – | – | ||
| O. sparsiannulatum | O. sparsiannulatum | 17231 | 122815 | T | Pinus taeda | Georgia, USA | L. Eckhardt | OM501515 | OM514841 | OM631880 | OM631696 |
| O. subannulatum | O. subannulatum | 518 | 188.86 | Pinus sp. | USA | W.H. Livingston, R.W. Davidson | AY934522 | DQ294364 | – | – | |
|
| |||||||||||
| Ophiostoma: O. ulmi complex | |||||||||||
|
| |||||||||||
| O. allantosporum | O. allantosporum | 163 | 185.86 | Pinus resinosa | Wisconsin, USA | M.J. Wingfield | OM501464 | OM514792 | OM631839 | OM631659 | |
| O. araucariae | O. araucariae | 40665 | 114.68 | T | Araucaria araucana | Chile | H. Butin | OM501467 | OM514795 | – | – |
| O. australiae | O. australiae | 6606 | 121025 | T | Acacia mearnsii | Australia | M.J. Wingfield | EF408603 | – | – | – |
| O. bacillisporum | O. bacillisporum | 2579 | MUCL45378 | T | Xyloterus domesticus on Fagus sylvatica | Germany | F.X. Carlier | OM501469 | OM514797 | OM631841 | OM631661 |
| O. borealis | O. borealis | 18966 | 123222 | T | Betula logs | Norway | G. Kamgan Nkuekam, H. Solheim | EF408593 | – | – | – |
| O. catonianum | O. catonianum | 11535 | 263.35 | A | Pyrus communis | Italy | G. Goidanich | AF198243 | – | – | – |
| O. denticiliatum | O. denticiliatum | 29493 | 124497 | T | Betula pendula | Norway | R. Linnakoski | FJ804490 | – | – | – |
| O. himal-ulmi | O. himal-ulmi | 22729 | ATCC36204 | Ulmus sp. | India | H. Rebel | OM501484 | OM514811 | OM631852 | – | |
| O. hylesini | O. hylesini | 51680 | 144296 | T | Hylesinus crenatus on Fraxinus excelsior | Poland | P. Wieczorek | MH055636 | MH055675 | MH062835 | – |
| O. introcitrinum | O. introcitrinum | – | UAMH9549 | T | Betula sp. | Canada | A. Olchowecki | OM501485 | – | – | – |
| O. karelicum | O. karelicum | 23099 | 123219 | T | Scolytus sp. on Betula sp. | Russia | Z.W. de Beer, P. Niemelä | OM501493 | OM514817 | OM631856 | OM631674 |
| O. novo-ulmi subsp. americana | O. novo-ulmi subsp. americana | 43874 | UAMH5030 | Ulmus americana | – | – | – | OM514828 | – | OM631683 | |
| O. novo-ulmi subsp. novo-ulmi* | O. novo-ulmi subsp. novo-ulmi | – | H327 | Ulmus sp. | Slovakia | H. Jamnicky | OM501503 | OM514827 | OM631865 | OM631682 | |
| O. novo-ulmi subsp. novo-ulmi | O. novo-ulmi subsp. novo-ulmi | 43870 | UAMH10443 | Ulmus sp. | Iran | M. Rahju, K. Rahnama | OM501504 | OM514829 | OM631866 | OM631684 | |
| O. patagonicum | O. patagonicum | 38089 | CIEFAP431 | T | Nothofagus pumilio | Argentina | A. de Errasti | OM501505 | OM514830 | OM631867 | – |
| O. pseudokarelicum | O. pseudokarelicum | 51704 | 144281 | T | Trypodendron domesticum on Alnus incana | Norway | T. Aas | MH055659 | MH055693 | MH062859 | – |
| O. quercus * | O. quercus | 2465 | 117912 | Quercus robur | France | M. Morelet | MT633064 | MT629747 | OM631876 | OM631692 | |
| O. signatum | O. signatum | 51689 | 144269 | T | Trypodendron signatum on Alnus incana | Norway | G. Kvammen | MH055645 | MH055682 | MH062844 | – |
| O. tasmaniense | O. tasmaniense | 29088 | 127212 | T | Eucalyptus nitens stumps | Australia | G. Kamgan Nkuekam | GU797211 | – | GU797223 | – |
| O. tsotsi | O. tsotsi | 18134 | 123599 | A | Julbenardia globiflora | Malawi | J. Roux | OM501520 | OM514846 | – | OM631701 |
| O. ulmi * | O. ulmi | – | W9 | Ulmus sp. | – | – | OM501521 | OM514847 | OM631885 | OM631702 | |
| O. undulatum | O. undulatum | 19396 | 127183 | Eucalyptus grandis | Australia | M.J. Wingfield | – | OM514848 | – | – | |
| O. villosum | O. villosum | 51694 | 144274 | T | Dryocoetes villosus on Quercus robur | Norway | T. Aas, K.D. Hansen | MH055650 | MH055685 | MH062849 | – |
|
| |||||||||||
| Ophiostoma: Group H | |||||||||||
|
| |||||||||||
| O. triangulosporum | O. triangulosporum | 1033 | 138.77 | T | Araucaria angustifolia | Brazil | H. Butin | OM501519 | OM514845 | OM631884 | OM631700 |
|
| |||||||||||
| Ophiostoma: Group I | |||||||||||
|
| |||||||||||
| O. macrosporum * | O. macrosporum | 14176 | 367.53 | T | Pinus sylvestris | Sweden | H. Francke-Grosmann | OM501495 | OM514819 | OM631858 | OM631675 |
| O. tingens | O. tingens | 25530 | 366.53 | Xyleborus saxesenii gallery in Populus deltoides | Sweden | H. Francke-Grosmann | – | EU177474 | – | – | |
| Ophiostoma sp. 1 (Ambrosiella sp. 1) | Ophiostoma sp. 1 (Ambrosiella sp. 1) | 43827 | UAMH10633 | Ips sp. | Canada | S. Massoumi Alamouti | OM501522 | OM514850 | OM631887 | OM631703 | |
| Ophiostoma sp. 2 (Ambrosiella sp. 2) | Ophiostoma sp. 2 (Ambrosiella sp. 2) | 43828 | UAMH10634 | Ips sp. | Canada | S. Massoumi Alamouti | OM501523 | – | – | – | |
|
| |||||||||||
| Ophiostoma: Group J | |||||||||||
|
| |||||||||||
| O. piliferum * | O. piliferum | – | – | – | – | – | MT633063 | MT629746 | OM631870 | OM631687 | |
| O. ponderosae | O. ponderosae | 37953 | ATCC26665 | T | Blue stain of Pinus ponderosa | Arizona, USA | R.W. Davidson | OM501554 | OM514882 | – | – |
|
| |||||||||||
| Ophiostoma: Group K | |||||||||||
|
| |||||||||||
| O. tetropii | O. tetropii | 4470 | 428.94 | Picea abies | Austria | T. Kirisits | OM501517 | OM514843 | OM631882 | OM631698 | |
|
| |||||||||||
| Ophiostoma incertae sedis (based on our data) | |||||||||||
|
| |||||||||||
| Lineage XVIII | |||||||||||
| O. valdivianum | O. valdivianum | 449 | 454.83 | T | Nothofagus alpiva | Chile | H. Butin | – | OM514849 | OM631886 | – |
| Lineage XX | |||||||||||
| O. angusticollis | O. angusticollis | 152 | – | Pinus banksiana | Wisconsin, USA | M.J. Wingfield | OM501466 | OM514794 | OM631840 | OM631660 | |
| O. denticulatum | O. denticulatum | 146 | ATCC38087 | T | Gnathotrichus sp. on Pinus sp. | Colorado, USA | R.W. Davidson | OM501480 | OM514807 | – | – |
| O. sejunctum | O. sejunctum | – | Ophi 1A | T | Tomicus sp. on Pinus sp. | Spain | M.R. Bareal | AY934519 | – | – | – |
|
| |||||||||||
| Lineage XXIII | |||||||||||
|
| |||||||||||
| O. noisomeae | O. noisomeae | 40326 | – | T | Rapanea melanophloeos | South Africa | T. Musvuugwa | OM501502 | OM514826 | – | OM631681 |
|
| |||||||||||
| Paleoambrosia | |||||||||||
|
| |||||||||||
| P. entomophila | P. entomophila | – | No. B-F-7 | T | Palaeotylus femoralis in burmese amber | Myanmar | G.O. Poinar, F.E. Vega | – | – | – | – |
|
| |||||||||||
| Raffaelea | |||||||||||
|
| |||||||||||
| R. albimanens* | R. albimanens | 25532 | 271.70 | T | Platypus externedentatus in Ficus sycomorus | South Africa | D.B. Scott | MT633066 | MT629749 | OM631890 | OM631705 |
| R. ambrosiae* | R. ambrosiae | 25533 | 185.64 | T | Platypus cylindrus tunnel in Quercus sp. | England, UK | J.M. Bakeer | MT633067 | MT629751 | OM631891 | OM631706 |
| R. arxii | R. arxii | 25534 | 273.70 | Xyleborus torquatus on Cussonia sp. | South Africa | D.B. Scott | – | MT629754 | OM631892 | OM631708 | |
| R. borbonica # | R. borbonica | 51553 | PPRI27953 | T | Leucaena leucocephala | Réunion, France | M.J. Wingfield, P.W. Crous | MT633059 | MT629742 | – | – |
| R. campbelliorum | R. campbelliorum | 44800 | 139943 | T | Xyleborus glabratus on Persea palustris | Florida, USA | A.S. Campbell | – | KR018414 | – | – |
| R. canadensis | R. canadensis | 25536 | 168.66 | T | Platypus wilsonii in Pseudotsuga menziessii | Canada | A. Funk | GQ225699 | MT629755 | – | – |
| R. canadensis (=A. sulcati) | R. canadensis | 25528 | 805.70 | T | Gmnathotrichus sulcatus (mycangium) on Pseudotsuga menziesii | Canada | A. Funk | – | EU177459 | – | – |
| R. crossotarsi | R. crossotarsi | 44793 | 141675 | T | Mycangium extract of Crossotarsus emancipatus in Lithocarpus sp. | Taiwan | J. Hulcr, A. Black, D.R. Simmons | KX267138 | MT629756 | OM631893 | OM631709 |
| R. cyclorhipidii | R. cyclorhipidii | 44790 | 141676 | T | Cyclorhipidion sp. on Lithocarpus sp. | Taiwan | J. Hulcr, A. Black, D.R. Simmons | MT633069 | MT629757 | – | OM631710 |
| R. ellipticospora | R. ellipticospora | 38056 | 121569 | T | Xyleborus glabratus on Persea sp. | South Carolina, USA | S. Fraedrich | MT633070 | MT629758 | OM631894 | – |
| R. fusca | R. fusca | 38798 | 121570 | T | Xyleborus glabratus on Persea sp. | South Carolina, USA | S. Fraedrich | – | EU177449 | – | – |
| R. gnathotrichi | R. gnathotrichi | 25523 | 379.68 | T | Gnathotrichus retusus on Picea engelmannii | Colorado, USA | L.R. Batra | – | EU177460 | – | – |
| R. promiscua # | R. promiscua | 55899 | 147173 | T | Xyleborinus saxesenii | South Afica | W.J. Nel | MW028176 | – | – | – |
| R. rapaneae | R. rapaneae | 40357 | 140084 | T | Platypodinae sp. on Rapanea melanophloeos | South Africa | T. Musvuugwa | KT192596 | KT182930 | – | – |
| R. santoroi | R. santoroi | 25539 | 399.67 | T | Platypus sp. bore hole | Argentina | J. Wright | MT633075 | MT629765 | – | – |
| R. scolytodis | R. scolytodis | 23001 | CCF3566 | A | Scolytodes sp. on Cecropia sp. | Costa Rica | M. Kolarik | AM267264 | – | – | – |
| R. seticollis | R. seticollis | 1031 | 634.66 | T | Tsuga canadensis | New York, USA | R.W. Davidson | MT633076 | MT629766 | OM631895 | OM631711 |
| R. subalba | R. subalba | 38797 | 121568 | T | Xyleborus sp. on Persea sp. | South Carolina, USA | S. Fraedrich | – | MT629767 | – | OM631712 |
| R. subfusca | R. subfusca | 38055 | 121571 | T | Xyleborus glabratus | South Carolina, USA | S. Fraedrich | – | EU177450 | – | – |
| R. sulcati | R. sulcati | 25540 | 806.70 | T | Gnatotrichus sulculcattus mycangium in Pseudotsuga menziesii | Canada | A. Funk | – | EU177462 | – | – |
| R. tritirachium | R. tritirachium | 25541 | 726.69 | T | Monarthrum mali tunnel in Quercus sp. | Pennsylvania, USA | D.B. Scott | – | EU177464 | – | – |
| R. xyleborini | R. xyleborini | 45859 | Hulcr6099 | T | Xyleborinus andrewesii | Florida, USA | C. Bateman | MT633078 | MT629769 | – | – |
| Raffaelea sp. A (PL1001) | Raffaelea sp. A (PL1001) | 38062 | – | Persea sp. | California, USA | A. Eskalen | – | KJ909293 | – | – | |
|
| |||||||||||
| Raffaelea incertae sedis (based on our data) | |||||||||||
|
| |||||||||||
| R. deltoideospora | R. deltoideospora | – | WIN(M)41 | Pinus sp. | Canada | J. Reid | EU879121 | KT182932 | – | – | |
| Lineage X | |||||||||||
| R. vaginata | R. vaginata | 40365 | 140086 | T | Lanurgus sp. on Olea capensis | South Africa | T. Musvuugwa | KT192602 | – | – | – |
|
| |||||||||||
| Sporothrix: S. candida complex | |||||||||||
|
| |||||||||||
| S. aemulophila | S. aemulophila | 40381 | 140087 | T | Rapanea melanophloeos | South Africa | T. Musvuugwa | OM501527 | OM514854 | OM631897 | – |
| S. cabralii | S. cabralii | 38098 | CIEFAP456 | T | Nothofagus pumilio | Argentina | A. de Errasti | OM501533 | OM514861 | OM631903 | OM631716 |
| S. candida | S. candida | 26484 | 129713 | T | Eucalyptus cloeziana | South Africa | G. Kamgan Nkuekam | – | OM514862 | OM631904 | OM631717 |
| S. itsvo | S. itsvo | 40370 | 141063 | T | Rapanea melanophloeos | South Africa | T. Musvuugwa | KX590840 | – | – | – |
| S. oleae # | S. oleae | 40362 | 142082 | T | Olea capensis ssp. macrocarpa wound | South Africa | T. Musvuugwa | MN298851 | – | – | – |
| S. rapaneae | S. rapaneae | 40369 | 141060 | T | Rapanea melanophloeos | South Africa | T. Musvuugwa | OM501558 | OM514885 | OM631926 | OM631735 |
|
| |||||||||||
| Sporothrix: S. inflata complex | |||||||||||
|
| |||||||||||
| S. dentifunda | S. dentifunda | 13016 | 115790 | T | Quercus wood | Hungary | C. Delatour | OM501535 | OM514865 | OM631907 | OM631718 |
| S. dimorphospora | S. dimorphospora | 12529 | 553.74 | T | Soil | Canada | R.A.A. Morall | OM501536 | OM514866 | OM631908 | OM631719 |
| S. guttuliformis | S. guttuliformis | 17167 | 437.76 | T | Soil | Malaysia | T. Furukawa | OM501542 | OM514873 | OM631915 | OM631725 |
| S. inflata | S. inflata | 12527 | 239.68 | T | Wheat field soil | Canada | W. Gams | OM501544 | – | OM631917 | OM631727 |
| Spumatoria longicollis | S. longicollis | 49345 | 141464 | E | Dung | Netherlands | J. vd Lee | – | OM514895 | OM631933 | – |
|
| |||||||||||
| Sporothrix: S. stenoceras & S. gossypina complexes | |||||||||||
|
| |||||||||||
| S. abietina * | S. abietina | 22310 | 125.89 | T | Pseudohylesinus gallery on Abies vejari | Mexico | J.G. Marmolejo | OM501526 | OM514853 | OM631896 | OM631713 |
| S. africana | S. africana | 823 | 116571 | Protea gaguedi | South Africa | M.J. Wingfield | OM501528 | OM514855 | OM631898 | OM631714 | |
| S. aurorae | S. aurorae | 19362 | 118837 | T | Hylastes angustatus on Pinus elliottii | South Africa | X.D. Zhou | – | OM514857 | OM631900 | – |
| S. cantabriensis | S. cantabriensis | 39766 | 136529 | T | Hylastes attenuates on Pinus sylvestris | Spain | P. Romón | KF951554 | – | – | – |
| S. cracoviensis # | S. cracoviensis | – | 147942 | T | Adult of Tryopodendron domesticum beetle on Fagus sylvaticum | Poland | R. Jankowiak | MW768964 | – | – | – |
| S. eucastaneae | S. eucastaneae | 1124 | 424.77 | T | Canker on Castanea dentata | North Carolina, USA | R.W. Davidson | OM501539 | OM514868 | OM631910 | OM631720 |
| S. euskadiensis | S. euskadiensis | 27318 | 122138 | T | Hylurgops palliatus on Pinus radiata | Spain | X.D. Zhou | DQ674369 | – | – | – |
| S. fraxini # | S. fraxini | – | 147936 | T | Gallery of Hylesinus varius on Fraxinus excelsior | Poland | R. Jankowiak | MH283150 | – | – | – |
| S. fusiformis | S. fusiformis | 9968 | 112912 | T | Populus nigra | Azerbaijan | D. Aghayeva | – | OM514870 | OM631912 | OM631722 |
| S. gossypina | S. gossypina | 1116 | ATCC18999 | T | Pinus ponderosa | New Mexico, USA | R.W. Davidson | – | – | OM631914 | OM631724 |
| S. villosa # | S. villosa | – | SNM188 | T | Pinus thunbergii | China | R. Chang | MW989428 | – | MZ853078 | – |
| S. lunata | S. lunata | 10563 | 112927 | T | Carpinus betulus | Austria | T. Kirisits | OM501545 | – | OM631918 | OM631728 |
| S. narcissi | S. narcissi | 22311 | 138.50 | T | Narcissus sp. | Netherlands | D.P. Limber | OM501548 | OM514876 | OM631920 | – |
| S. nsini | S. nsini | 28602 | 143281 | T | Protea caffra | South Africa | F. Roets | EU660458 | – | – | – |
| S. prolifera | S. prolifera | 37435 | 251.88 | T | Quercus robur | Poland | T. Kowalski | OM501555 | – | OM631924 | OM631732 |
| S. protearum | S. protearum | 1107 | 116654 | Protea caffra | South Africa | M.J. Wingfield | OM501557 | OM514884 | OM631925 | OM631734 | |
| S. resoviensis # | S. resoviensis | – | 147927 | T | Wound on Betula pendula | Poland | R. Jankowiak | MH740962 | – | – | – |
| S. rossii | S. rossii | 1118 | 116.78 | T | Dendroctonus adjunctus gallery on Pinus ponderosa | New Mexico, USA | R.W. Davidson | OM501559 | OM514886 | OM631927 | OM631736 |
| S. splendens * | S. splendens | 23050 | 138722 | Oodinychus sp. on Protea repens | South Africa | F. Roets | OM501561 | OM514888 | OM631929 | OM631738 | |
| S. stenoceras | S. stenoceras | 3202 | 237.32 | T | Pine pulp | Norway | H. Robak | AF484462 | DQ294350 | – | – |
| S. uta | S. uta | 40316 | 141069 | P | Rapanea melanophloeos | South Africa | T. Musvuugwa | OM501565 | OM514892 | OM631930 | OM631739 |
| S. variecibata | S. variecibata | 23051 | 121961 | T | Trichouropoda sp. from Protea repens | South Africa | F. Roets | OM501566 | OM514893 | OM631931 | OM631740 |
| S. zambiensis | S. zambiensis | 29077 | 124914 | T | Protea caffra | Zambia | F. Roets | OM501567 | OM514894 | OM631932 | OM631741 |
|
| |||||||||||
| Sporothrix: S. pallida complex | |||||||||||
|
| |||||||||||
| S. albicans (syn S. pallida) | S. albicans (syn S. pallida) | 17203 | 302.73 | T | Soil | England, UK | S.B. Saksena | OM501529 | OM514856 | OM631899 | OM631715 |
| S. chilensis | S. chilensis | 49343 | 139890 | T | Soil | Chile | R. Cruz Choappa | – | OM514863 | OM631905 | – |
| S. gemella | S. gemella | 23057 | 121959 | T | Tarsonemus sp. on Protea caffra | South Africa | F. Roets | DQ821560 | DQ821531 | – | – |
| S. humicola | S. humicola | 7618 | 118129 | T | Soil | South Africa | H.F. Vismer | OM501543 | – | OM631916 | OM631726 |
| S. mexicana | S. mexicana | 29129 | 120341 | T | Soil | Mexico | A. Espinosa | OM501547 | OM514875 | OM631919 | – |
| S. pallida | S. pallida | 17209 | 131.56 | T | Stemonitis fusca | Japan | K. Tubaki | OM501550 | OM514878 | OM631921 | OM631729 |
| S. palmiculminata | S. palmiculminata | 20677 | 119590 | T | Protea repens | South Africa | F. Roets | OM501551 | OM514879 | OM631922 | OM631730 |
| S. protea-sedis | S. protea-sedis | 29074 | 124911 | P | Protea caffra | Zambia | F. Roets | OM501556 | OM514883 | – | OM631733 |
| S. stylites | S. stylites | 14543 | 118848 | T | Pine utility poles | South Africa | E.M. de Meyer | OM501563 | OM514890 | – | – |
|
| |||||||||||
| Sporothrix: Pathogenic clade | |||||||||||
|
| |||||||||||
| S. brasiliensis | S. brasiliensis | 29127 | 120339 | T | Human skin | Brazil | M.D.S. Lazera | OM501531 | OM514859 | OM631901 | |
| S. globosa | S. globosa | 29128 | 120340 | T | Human face | Spain | C. Rubio | OM501541 | OM514872 | OM631913 | OM631723 |
| S. luriei | S. luriei | 17210 | 937.72 | T | Human skin | South Africa | L. Lurie | OM501546 | – | – | – |
| S. schenckii* | S. schenckii | 38850 | 138723 | Clinical isolate | South Africa | H.F. Vismer | OM501560 | OM514887 | OM631928 | OM631737 | |
|
| |||||||||||
| Sporothrix: Group D | |||||||||||
|
| |||||||||||
| S. cavum # | S. cavum | – | 147943 | T | Cavity of Dendrocopos major on Salix fragilis | Poland | R. Jankowiak | MF782813 | MF782813 | – | – |
| S. polyporicola | S. polyporicola | 5461 | 669.88 | T | Fomitopsis pinicola | Sweden | S. Ryman | OM501553 | OM514881 | – | – |
|
| |||||||||||
| Sporothrix: Group E | |||||||||||
|
| |||||||||||
| S. phasma* | S. phasma | 20676 | 119722 | T | Protea laurifolia | South Africa | F. Roets | OM501552 | OM514880 | OM631923 | OM631731 |
|
| |||||||||||
| Sporothrix: Group F | |||||||||||
|
| |||||||||||
| S. bragantina | S. bragantina | 17149 | 474.91 | T | Soil | Brazil | W. Gams | OM501530 | OM514858 | – | – |
| S. curviconia | S. curviconia | 17164 | 959.73 | T | Terminalia ivorensis | Ivory Coast | J. Devois | OM501534 | OM514864 | OM631906 | – |
| S. epigloea | S. epigloea | 22308 | 573.63 | T | Tremella fuciformis | Argentina | R. T. Guerrero | KX590817 | KX590854 | – | – |
| S. eucalyptigena | S. eucalyptigena | 45431 | 139899 | T | Eucalyptus sp. | Australia | P. A. Barber | OM501538 | – | OM631909 | – |
| S. nebularis | S. nebularis | 27319 | 122135 | T | Hylastes attenuatus on Pinus radiata | Spain | P. Romón | KX590823 | KX590862 | – | – |
| S. nigrograna | S. nigrograna | 14487 | MAFF410943 | T | Pinus densiflora | Japan | H. Masuya | OM501549 | OM514877 | – | – |
| S. smangaliso | S. smangaliso | 50502 | 143341 | T | Protea gaguedi | South Africa | N.P. Ngubane | MF103773 | – | – | – |
| S. thermara | S. thermara | 38930 | 139747 | T | Cyrtogenius africus on Euphorbia ingens | South Africa | J.A. van der Linde | OM501564 | OM514891 | – | – |
| S. zhejiangensis | S. zhejiangensis | – | MUCL55183 | T | Monochamus sp. on Pinus sp. | China | Q. Lu, Y.Y. Lun | KY094071 | – | – | – |
|
| |||||||||||
| Sporothrix: Group G | |||||||||||
|
| |||||||||||
| S. dombeyi | S. dombeyi | 1023 | 455.83 | T | Nothofagus sp. | Chile | H. Butin | OM501537 | OM514867 | – | – |
|
| |||||||||||
| Sporothrix incertae sedis (based on our data) | |||||||||||
|
| |||||||||||
| Lineage XVI | |||||||||||
|
| |||||||||||
| S. fumea | S. fumea | 26813 | 129712 | T | Phoracantha sp. galleries on Eucalyptus | South Africa | C. Perez | OM501540 | OM514869 | OM631911 | OM631721 |
|
| |||||||||||
| Lineage XIX | |||||||||||
|
| |||||||||||
| S. brunneoviolacea | S. brunneoviolacea | 37442 | 124560 | P | Soil | Spain | C. Silvera | OM501532 | OM514860 | OM631902 | – |
|
| |||||||||||
| Sporothrix incertae sedis (based on published data) | |||||||||||
|
| |||||||||||
| S. cryptarchum # | S. cryptarchum | – | 147934 | T | Adult of Cryptarcha undata | Poland | R. Jankowiak | MW768966 | – | – | – |
| S. hypoxyli # | S. hypoxyli | 47441 | 141569 | T | Hypoxylon petriniae on Fraxinus wood | Netherlands | E. Osieck & W.J. Nel | MT637058 | MW012948 | – | – |
| S. undulata # | S. undulata | – | 147929 | T | Adult of Epuraea guttata | Poland | R. Jankowiak | MH740976 | – | – | – |
|
| |||||||||||
| Ophiostomatales insertae sedis (based on published data) | |||||||||||
|
| |||||||||||
| L. antibioticum | L. antibioticum | 2777 | DAOM84338 | T | Pinus taeda | Georgia, USA | S. Alexander | AF343677 | – | – | – |
| L. brachiatum | L. brachiatum | 2855 | C388 | T | Picea rubens | New York, USA | S. Alexander | AF343676 | – | – | – |
| L. elegans | L. elegans | 2245 | 115241 | Chamaecyparis / Hinoke | Taiwan | M.J. Wingfield | AF343675 | – | – | – | |
| O. crenulatum | O. crenulatum | – | WIN(M)70-17 | T | Pinus banksiana | Canada | J. Reid | – | AF135589 | – | – |
| O. fasciatum | O. fasciatum | – | UM56 | Pseudotsuga menziesii | Canada | A. Olchowecki | EU913720 | EU913680 | – | – | |
| O. breviusculum | O. breviusculum | – | YCC522 | Single ascospore isolate from YCC-494 | Japan | Y. Yamaoka | AB200423 | – | – | – | |
| O. ssiori | O. ssiori | – | MAFF410973 | T | Polygraphus ssiori on Prunus sp. | Japan | H. Masuya | AB096209 | – | – | – |
| O. subalpinum | O. subalpinum | – | YCC408 | Cryphalus sp. on Abies sp. | Japan | Y. Yamaoka | AB200424 | – | LC090750 | – | |
| O. pseudonigrum | O. pseudonigrum | – | WIN(M)71-13 | Picea mariana | Canada | J. Reid | – | AF135577 | – | – | |
| O. pehueninum | O. pehueninum | – | 142995 | T | Araucaria araucana | Chile | V. Sepúlveda | MF576438 | – | MF576446 | – |
| O. tremulo-aureum | O. tremulo-aureum | – | 361.65 | Black canker on Populus tremuloides | Colorado, USA | R.W. Davidson | – | AF135573 | – | – | |
ATCC = American Type Culture Collection, Maryland, USA; CBS = the culture collection of Westerdijk Fungal Biodiversity Institute, Utrecht, the Netherlands; CFCC = China Forestry Culture Collection Center, Research Institute of Forest Ecology, Environment and Protection, Chinese Academy of Forestry, Beijing, China; CIEFAP = the culture collection of the Centro de Investigación y Extensión Forestal Andino Patagónico, Argentina; CMW = the culture collection of the Forestry and Agricultural Biotechnology Institute (FABI), University of Pretoria, South Africa; CTK = the culture collection of the Institute of Forest Entomology, Forest Pathology and Forest Protection, Department of Forest and Soil Sciences, University of Natural Resources and Applied Life Sciences, Vienna, Austria; CXY = the culture collection of the Chinese Academy of Forestry, China; DAOM = Canadian National Mycological Herbarium, Ottawa, Canada; KACC = Korean Agricultural Culture Collection, Suwon, Korea; KUC = Korea University Culture Collection, Seoul, Korea; MAFF = the culture collection of National Institute of Agrobiological Resources, Japan; MUCL = BCCM/MUCL Agro-food & Environmental Fungal Collection, Université catholique de Louvain, Belgium; PPRI = National Collection of Fungi, Pretoria, South Africa; RWD = the private collection of R.W. Davidson; SNM = the microbial culture collection of Shandong Normal University, Jinan, Shandong, China; TRTC = Royal Ontario Museum Fungarium, Toronto, Canada; UAMH = University of Alberta Mold Herbarium and Culture Collection, Edmonton, Canada; VPRI = Victorian Plant Pathology Herbarium, Victoria, Australia; WIN(M) = University of Manitoba, Microbiology and Botany (J. Reid’s personal collection); YCC = the private collection of Y. Yamaoka.
2 T = ex-type, E = ex-epitype, P = ex-paratype; L = ex-lectotype; A = authentic isolate, used in the original study.
* Genome data used in this study.
# Species published after 2018, thus not included in the current analyses.
^ CMW culture replaced with a different fungus.
3 Sequences with accession numbers preceded with “OM” were generated in the current study.
Afroraffaelea (not included in the phylogenetic analyses)
Afroraffaelea C.C. Bateman et al., Fungal Ecol. 25: 46. 2017. MycoBank MB 816236. Fig. 6A, B.
Fig. 6.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Afroraffaelea. C, D. Aureovirgo. E, F. Ceratocystiopsis (a. Cop. collifera; b. Cop. ochracea; c. Cop. manitobensis; d. Cop. concentrica; e. Cop. rollhanseniana). G, H. Chrysosphaeria. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Etymology: ‘Prefix Afro - indicating its likely African origin, and ‘raffaelea’ to recognise the ecological similarity to the closely related (though probably not monophyletic) ambrosia fungus genus Raffaelea’ (Bateman et al. 2017).
Sexual morph: Unknown.
Asexual morph: No reproductive structures observed. Colonies white becoming brown with age, aerial hyphae present, abundant, subhyaline, extensively branched often perpendicularly, thickened and melanized intercalarily, distal hyphae denticulate, submerged hyphae subhyaline, constricted at septa, frequently branched, with tapering distal ends, monillioid mycelia occurring in peptone-containing medium, fragmented unicellularly or bicellularly.
Type species: Afroraffaelea ambrosiae C.C. Bateman et al., Fungal Ecol. 25: 46. 2017. MycoBank MB 816237.
No other species known.
Notes: Afroraffaelea was described by Bateman et al. (2017) and has a unique morphology relative to the rest of the Ophiostomatales. Within a few days of sub-culturing, Afr. ambrosiae (the only species in this genus) produces masses of aerial hyphae, unlike any other ophiostomatalean species, and changes colour from white to brown as the culture matures. The fungus is the dominant fungal symbiont of the ambrosia beetle Premnobius cavipennis, presumably introduced into North America from Africa (Bateman et al. 2017). The phylogenies of Bateman et al. (2017) showed Afr. ambrosiae grouping close to Fragosphaeria, but there was insufficient phylogenetic support for these authors to determine the taxonomic placement of Afroraffaelea in the Ophiostomatales.
The taxonomic placement of Afroraffaelea in our phylogenetic trees is not conclusive. The four known isolates of Afr. ambrosiae consistently grouped together as one species. However, the clade grouped inconsistently in the single gene trees. For example, in the LSU and TEF-1α trees, Afr. ambrosiae isolates grouped in Lineage VIII, but with Lineage V in ITS, and with Lineage XXIII in RPBII (data not shown). We consequently excluded these isolates from our datasets. We believe that the taxonomic placement of this fungus will be resolved when additional species are found, or possibly with whole genome data. Until such time, we treat Afroraffaelea as incertae sedis in the Ophiostomatales.
Aureovirgo (Lineage IX)
Aureovirgo J.A. van der Linde et al., Antonie van Leeuwenhoek 109: 593. 2016. MycoBank MB 813870. Fig. 6C, D.
Etymology: ‘Refers to the golden appearance of the immature ascomata and the pure white colour of the cultures (‘‘Aureovirgo’’ refers to a golden maiden with an unstated overtone of virginal whiteness)’ (Van der Linde et al. 2016).
Sexual morph: Ascomatal bases subglobose to globose, pale brown when young, becoming darker with age; necks cylindrical, dark brown. Ostiolar hyphae hyaline, parallel. Ascospores hyaline, 1-celled, enclosed in a sheath, falcate, endospores (the body of ascospores) allantoid.
Asexual morph: Leptographium-like; conidiophores macronema-tous, mononematous, hyaline; conidiogenous cells cylindrical, hyaline; conidia hyaline, 1-celled, oblong to ellipsoidal.
Type species: Aureovirgo volantis J.A. van der Linde et al., Antonie van Leeuwenhoek 109: 593. 2016. MycoBank MB 813872.
No other species known.
Notes: Aureovirgo was described from galleries of the ambrosia beetle Cyrtogenius africus in dead and dying Euphorbia ingens trees in South Africa. It is a monotypic genus accommodating A. volantis (Van der Linde et al. 2016). The ascomatal bases are honey-coloured when immature, while the rest of the culture is white. This is the only genus in the Ophiostomatales associated with Eurphorbia trees. In our dataset, Aureovirgo formed a well-supported, distinct lineage.
Ceratocystiopsis (Lineage VII)
Ceratocystiopsis H.P. Upadhyay & W.B. Kendr., Mycologia 67: 799. 1975. MycoBank MB 889. Fig. 6E, F.
Etymology: ‘Resembling the genus Ceratocystis’ (Upadhyay & Kendrick 1975, Upadhyay 1981). At the time of the description, Upadhyay & Kendrick (1975) and others treated Ophiostoma as a synonym of Ceratocystis in the Ophiostomatales. Ceratocystis was later shown to form part of the Microascales, and Ceratocystiopsis to be related to Ophiostoma in the Ophiostomatales (Zipfel et al. 2006). Although somewhat of a misnomer, the name Ceratocystiopsis remains valid and represents a distinct genus in the Ophiostomatales.
Synonym: Hyalorhinocladiella H.P. Upadhyay & W.B. Kendr., Mycologia 67: 800. 1975. MycoBank MB 8582. [Type species Hyalorhinocladiella minuta-bicolor (R.W. Davidson) H.P. Upadhyay & W.B. Kendr.]
Sexual morph: Ascomatal bases subglobose, globose, obpyriform, black or pale brown, upper part surrounded with collar-like structure or corona of globose cells; necks short conical to elongate, cylindrical, black or paler than base. Ostiolar hyphae absent or present, convergent, parallel, divergent. Asci evanescent, clavate, broadly fusiform, 8-spored. Ascospores hyaline, 1-celled, enclosed in a sheath, falcate in side view, fusiform, acicular in face view; endospores elongate orange segment-shaped in side view, cylindrical in face view; cucullate, sometimes forming bulbous swelling toward one end.
Asexual morph: Hyalorhinocladiella-like; conidiophores micronema-tous, semimacronematous, macronematous, mononematous, hyaline; conidiogenous cells integrate or discrete, polyblastic, sympodial, with denticle scars; conidia hyaline, 1-celled, obovoid, ellipsoidal, oval to globose, clavate, T-shaped, Y-shaped, cylindrical.
Type species: Ceratocystiopsis minuta (Siemaszko) H.P. Upadhyay & W.B. Kendr., Mycologia 67: 800. 1975. MycoBank MB 310480.
Other species: Listed in Table 1.
Notes: In their revision of the genus Ceratocystis, Upadhyay & Kendrick (1975) erected Ceratocystiopsis for species with falcate ascospores and a Hyalorhinocladiella asexual morph. Hausner et al. (1993a) reduced Ceratocystiopsis to synonymy with Ophiostoma based on rDNA sequences, but Zipfel et al. (2006) included additional taxa in their study and reinstated the genus that is typified by Ceratocystiopsis minuta. Species in this genus have elongated sickle-shaped (falcate) ascospores that are sheathed, produced in very short necked ascomata, characters not seen in any other genus in the Ophiostomatales (Upadhyay & Kendrick 1975, Zipfel et al. 2006, De Beer & Wingfield 2013). Species of Ceratocystiopsis are found in the galleries of conifer-infesting bark beetles (Dendroctonus species in North America or Ips species in Japan and Europe) in the Northern Hemisphere (Plattner et al. 2009).
Ceratocystiopsis spp. formed a well-supported monophyletic clade in all our phylogenies, supporting the decision of Zipfel et al. (2006) to reinstate the genus. Our data also supported the placement of Cop. neglecta (previously Ophiostoma neglectum) in Ceratocystiopsis, as shown by De Beer & Wingfield (2013).
Chrysosphaeria
Chrysosphaeria W.J. Nel et al., Mycologia 113: 1206. 2021. MycoBank MB 837564. Fig. 6G, H.
Etymology: ‘From Latin chryso-, golden, and -sphaera, sphere or orb, referring to the light colour of the ascoma bases in the type species’ (Nel et al. 2021).
Sexual morph: Ascomatal bases globose, light to golden brown; necks light brown, cylindrical, tapering towards apex, flexible. Ostiolar hyphae present, slightly divergent, hyaline. Ascospores hyaline, 1-celled, no sheath, short cylindrical to bean-shaped.
Asexual morph: Sporothrix-like; conidiophores micronematous, mononematous, hyaline; conidiogenous cells hyaline, denticulate; conidia hyaline, 1-celled, oblong, occasionally producing secondary spores hyaline, 1-celled, obovoid.
Type species: Chrysosphaeria jan-nelii W.J. Nel et al., Mycologia 113: 1206. 2021. MycoBank MB 837566.
Notes: Unlike most species in the Ophiostomatales, which occur on wood and have associations with bark or ambrosia beetles, Chyrysosphaeria was isolated from abandoned combs of the fungus-growing termite Macrotermes natalensis.
Dryadomyces (Lineage V)
Dryadomyces Gebhardt, Mycol. Res. 109: 693. 2005. MycoBank MB 28937, emend. Z.W. de Beer & M. Procter Fig 7A, B.
Fig. 7.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Dryadomyces. C, D. Esteya. E, F. Fragosphaeria. G, H. Graphilbum. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Etymology: ‘dryads, the tree nymphs in Greek mythology; referring to the habitat of these fungi in woody plants’ (Gebhardt et al. 2005).
Sexual morph: Unknown.
Asexual morph: Colony confluent, mucilaginous. Conidiophores single or aggregated in sporodochia, macronematous, mononematous, hyaline, smooth. Conidiogenous cells monillioid or oblong, sympodial, with denticles or inconspicuous scars. Conidia hyaline to subhyaline, 1-celled, globose to subglobose, obovoid to pyriform with truncate base, smooth, producing secondary spores. Aleuriospores absent or present, hyaline, globose to subglobose, terminal.
Type species: Dryadomyces amasae Gebhardt, Mycol. Res. 109: 693. 2005. MycoBank MB 369332.
Notes: Based on our analyses, Dryadomyces now includes five ambrosia beetle-associated species, previously described in either Ambrosiella, Raffaelea or Dryadomyces (Gebhardt et al. 2005, De Beer & Wingfield 2013). Raffaelea sulphurea was isolated from hardwood-infesting ambrosia beetles, as was its sister species, R. montetyi (Massoumi Alamouti et al. 2009). Two Asian species in this complex, R. quercivora from Japan and R. quercus-mongolicae from Korea, have been implicated in contributing to the death of large numbers of Quercus spp. in their native ranges (Dreaden et al. 2014).
The fifth species in Clade V of our analyses (Fig. 5), Drydomyces amasae, was initially described as the type species of the monotypic genus Dryadomyces. It was isolated from the ambrosia beetle Amasae concitatus in Taiwan (Gebhardt et al. 2005). Although the conidia of D. amasae differ from other Raffaelea species, Harrington et al. (2008) suggested that all ambrosial fungi in the Ophiostomatales should be treated in Raffaelea, until further studies revealed the taxonomic identities of these fungi. The more robust phylogenies constructed by Massoumi Alamouti et al. (2009) revealed a monophyletic lineage including D. amasae, R. sulphurea and R. montetyi. However, Harrington et al. (2010) treated D. amasae in Raffaelea, not considering the differences between conidia of D. amasae and other Raffaelea species as sufficient to separate Dryadomyces from Raffaelea.
De Beer & Wingfield (2013) named the lineage containing R. amasae, R. montetyi, R. suphurea, R. quercus-mongolicae and R. quercivora as the R. sulphurea complex, referring to the oldest of the five described species. This complex grouped within Leptographium s.l. in their phylogenies. They did, however, note that the R. sulphurea complex is most probably not part of the larger Leptographium s.l., but their data were insufficient to show otherwise. They suggested that when more robust phylogenies support the R. sulphurea complex as a monophyletic lineage separate from other genera, Dryadomyces would be a suitable name as the complex includes D. amasae.
Dreaden et al. (2014) investigated the monophyly of Raffaelea and showed that the R. sulphurea complex grouped close to isolates of E. vermicola within Leptographium s.l. as defined by De Beer & Wingfield (2013). They concluded that a larger study including additional Leptographium and Raffaelea species should provide clarity on the placement of these outlying Raffaelea lineages.
The R. sulphurea complex consistently formed a well-defined lineage distinct from Leptographium in our phylogenies. This supported re-instating the name Dryadomyces for this lineage. The description of Dryadomyces by Gebhardt et al. (2005) is emended here to accommodate other species in this lineage.
New combinations:
1) Dryadomyces montetyi (M. Morelet) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840313.
Basionym: Raffaelea montetyi M. Morelet, Ann. Soc. Sci. Nat. Archéol. Toulon Var 50: 189. 1998. MycoBank MB 445315.
Description: Morelet (1998: 189–191, fig. A).
Phylogenetic data: Gebhardt et al. (2005), Massoumi Alamouti et al. (2009), Harrington et al. (2010), Matsuda et al. (2010), De Beer & Wingfield (2013), Dreaden et al. (2014), Musvuugwa et al. (2015), De Beer et al. (2016a, b), Vanderpool et al. (2017), Li et al. (2018), Saucedo-Carabez et al. (2018).
Notes: It forms part of Dryadomyces (Fig. 5), previously designated as the R. sulphurea complex (De Beer & Wingfield 2013).
2) Dryadomyces quercivorus (Kubono & Shin. Ito) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840316.
Basionym: Raffaelea quercivora Kubono & Shin. lto, Mycoscience 43: 256. 2002. MycoBank MB 483997.
Description: Kubono & lto (2002: 256–259, figs 1–11).
Phylogenetic data: Kim et al. (2009), Seo et al. (2010), Matsuda et. al. (2010), Endoh et al. (2011), Dreaden et al. (2014), Musvuugwa et al. (2015), Simmons et al. (2016), Van der Linde et al. (2016), De Beer et al. (2016a, b), Vanderpool et al. (2017), De Errasti et al. (2018), Li et al. (2018), Saucedo-Carabez et al. (2018).
Notes: Dryadomyces quercivorus groups in Dryadomyces and was previously accommodated in the R. sulphurea complex (Fig. 5; De Beer & Wingfield 2013, De Errasti et al. 2018, Li et al. 2018, Saucedo-Carabez et al. 2018).
3) Dryadomyces quercus-mongolicae (K.H. Kim et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840318.
Basionym: Raffaelea quercus-mongolicae K.H. Kim et al., Mycotaxon 110: 193. 2009. MycoBank MB 515072.
Description: Kim et al. (2009: 193–195, fig. 2).
Phylogenetic data: Kim et al. (2009), Seo et al. (2010), De Beer et al. (2016a), Vanderpool et al. (2017), De Errasti et al. (2018), Li et al. (2018), Saucedo-Carabez et al. (2018).
Notes: Dryadomyces quercus-mongolicae groups in Dryadomyces, and was formerly in the R. sulphurea complex (Fig. 5; De Beer & Wingfield 2013, Li et al. 2018, Saucedo-Carabez et al. 2018).
4) Dryadomyces sulphureus (L.R. Batra) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840321.
Basionym: Ambrosiella sulphurea L.R. Batra, Mycologia 59: 992. 1967. MycoBank MB 326145.
Synonyms: Raffaelea sulphurea (L.R. Batra) T.C. Harr., Mycotaxon 111: 353. 2010. MycoBank MB 515298.
Cephalosporium luteum Verrall, J. Agric. Res. 66: 141. 1943. MycoBank MB 284848.
Description: Batra (1967: 992–998, figs 20, 21, 26–29), Verrall (1943: 141, 142. fig 4).
Phylogenetic data: Cassar & Blackwell (1996), Rollins et al. (2001), Gebhardt et al. (2005), Massoumi Alamouti et al. (2009), Harrington et al. (2010), Matsuda et al. (2010), De Beer & Wingfield (2013), Dreaden et al. (2014), Musvuugwa et al. (2015), Simmons et al. (2016), De Beer et al. (2016a, b), Vanderpool et al. (2017), Saucedo-Carabez et al. (2018).
Notes: This taxon groups in Dryadomyces (Fig. 5), previously designated as the R. sulphurea species complex (De Beer & Wingfield 2013), and has a heterotypic synonym, Cephalosporium luteum (Verrall 1943). Gharabigloozare (2015) proposed C. luteum as conspecific with R. sulphurea. These two species were isolated from Xyleborinus saxeseni (syn. Xyleborus pecanis) and their galleries. The culture morphologies and conidial dimensions of Batra (1967) and Verrall (1943) match and we support their conspecificity. In terms of nomenclatural priority, Cephalosporium luteum (1943) has precedence over A. sulphurea (1967) and the former epithet has a priority over the latter as a basionym. However, since De Beer & Wingfield (2013) introduced R. sulphurea complex, the name R. sulphurea has been widely used and recognised, and we have chosen not to introduce a new epithet for this species.
Esteya (Lineage IV)
Esteya J.Y. Liou et al., Mycol. Res. 103: 243. 1999. MycoBank MB 28256, emend. Z.W. de Beer & M. Procter Fig. 7C, D.
Etymology: ‘Named in honour of Prof. Ralph H. Estey (Macdonald College, McGill University, Canada), in recognition of his contribution to the study of nematophagous fungi and nematology’ (Liou et al. 1999).
Sexual morph: Unknown.
Asexual morph: Conidiophores macronematous, mononematous, simple, subhyaline to pigmented, smooth to verrucous. Conidiogenous cells flask-shaped. Conidia hyaline, 1-celled, smooth, asymmetrically ellipsoidal in face view, concave, lunate in side view, ends moderately apiculate, with a layer of adhesive mucus on concave surface, containing ovoid endospore-like structures. Hyalorhinocladiella-like; conidiophores macronematous, mononematous, simple or branched, subhyaline to pigmented, smooth to verrucous; conidiogenous cells cylindrical to subulate; conidia hyaline, 1-celled, cylindrical, smooth.
Type species: Esteya vermicola J.Y. Liou et al., Mycol. Res. 103: 243. 1999. MycoBank MB 450702.
No other species known.
Notes: The genus Esteya was first described by Liou et al. (1999) and includes the single species, E. vermicola (Wang et al. 2015). Esteya vermicola produces two different types of conidia depending on abiotic and biotic factors, with one of the forms lethal to nematodes when consumed (Liou et al. 1999). This species was first isolated from the pine wood nematode Bursaphelencus xylophilus. It is the only nematode-associated fungus in the Ophiostomatales, clearly distinguishing Esteya from other genera in the Order. Wang et al. (2008) discovered another isolate of E. vermicola, and upon performing a BLAST search in GenBank found that members of the Ophiostomatales were the closest phylogenetic relatives. They subsequently performed molecular phylogenetic analyses including various Ophiostomatales species and found that Esteya grouped close to but separately from a clade containing two Leptographium and two Grosmannia species, and distinct from any other genus in the Ophiostomatales. The phylogenies of De Beer & Wingfield (2013) did not resolve the generic placement of Esteya in the Ophiostomatales, showing that Esteya grouped within Leptographium s.l. based on both ITS and LSU sequence data. These authors chose not to reduce Esteya to synonymy with Leptographium and recommended that Esteya should not be treated in Leptographium but be retained as a distinct genus until further studies could be conducted on the group. Later studies using LSU data again placed Esteya in Leptographium s.l. (Musvuugwa et al. 2015, De Beer et al. 2016a). Dreaden et al. (2014) used SSU, LSU and β-tub genes, which also placed Esteya in Leptographium s.l. In the phylogenies of the TEF-1α and β-tub genes generated by Wang et al. (2014), the separate treatment of Esteya from Leptographium s.l. was well supported.
In our datasets, Esteya grouped within what was previously known as Leptographium s.l. The separation of Grosmannia and Leptographium based on our data, allows the recognition of Esteya as a distinct genus, forming a well-supported clade including two of the known E. vermicola isolates. Our data did not include the type specimen of E. vermicola, because cultures of this fungus were not available in the culture collections from which isolates were sought. It is interesting that a nematode-associated fungus groups within a family of fungi not associated with nematodes. The discovery of other Esteya isolates may shed light on the evolution of this unique lifestyle within the Ophiostomatales.
Fragosphaeria (Lineage XXI)
Fragosphaeria Shear, Mycologia 15: 124. 1923. MycoBank MB 2011, emend. Z.W. de Beer & M. Procter Fig. 7E, F.
Etymology: Name is derived from the word “fragor” meaning “breaking to pieces” and “sphaera” (Shear 1923), referring to the spherical ascomata of the two species in the genus that easily break into fragments when handled.
Sexual morph: Ascomatal bases dark, globose, cleistothecial, walls easily fragmented. Asci subglobose. Ascospores yellowish-brown, 1-celled, broadly bean-shaped.
Asexual morph: Conidia hyaline, 1-celled, oblong to ellipsoidal, subalantoid, inequilateral.
Type species: Fragosphaeria purpurea Shear, Mycologia 15: 124. 1923. MycoBank MB 275760.
Other species: Listed in Table 1.
Notes: Fragosphaeria was first associated with the Ophiostomatales when a study by Suh & Blackwell (1999) showed that F. purpurea, the type species of this genus, grouped with some Ophiostoma species. The genus currently includes only two species, F. purpurea (Shear 1923) and F. reniformis (Saccardo 1881). The latter species was originally described in Cephalotheca, which Chesters (1935) reduced to synonymy with Fragosphaeria based on the similarities between F. purpurea, Cephalotheca sulfurea and C. reniformis. Malloch & Cain (1970) reinstated Fragosphaeria, transferring Cephalotheca reniformis to Fragosphaeria.
Both Fragosphaeria species are associated with stained wood within and surrounding bark beetle galleries in hardwoods and have been isolated in Britain and North America (Chesters 1935, De Beer & Wingfield 2013). Drawings by Chesters (1935) show a sporothrix-like asexual morph and allantoid ascospores produced by the cephalothecoid ascomata, which link this genus with other ophiostomatalean genera. However, the spherical cephalothecoid ascomata distinguish it from other Ophiostomatales (De Beer & Wingfield 2013). A study by Yaguchi et al. (2006) showed that there was no evident relationship between species of Cephalotheca and Fragosphaeria and confirmed that the two Fragosphaeria isolates group together. Our data confirm the treatment of Fragosphaeria as a distinct, monophyletic genus in the Ophiostomatales.
Graphilbum (Lineage VIII)
Graphilbum H.P. Upadhyay & W.B. Kendr., Mycologia 67: 800. 1975. MycoBank MB 8393, emend. Z.W. de Beer et al. In The Ophiostomatoid Fungi: Expanding Frontiers: 268. 2013. Fig. 7G, H.
Etymology: Graphilbum was described as the hyaline analogue of the genus Graphium (Upadhyay & Kendrick 1975). Graphium was later shown to form part of the Microascales (Okada et al. 1998), while Graphilbum resides in the Ophiostomatales (Zipfel et al. 2006). Although somewhat of a misnomer, the name Graphilbum remains valid and represents a distinct genus in the Ophiostomatales.
Sexual morph: Ascomatal bases globose, subglobose, black; necks dark brown to black, nearly cylindrical, straight or slightly curved. Ostiolar hyphae present or absent, parallel or divergent. Asci evanescent, broadly clavate to subglobose. Ascospores hyaline, 1-celled, cylindrical to oblong in side view, globose in end view, enclosed in rectangular sheath.
Asexual morph: Conidiophores micronematous, semimacrone-matous, macronematous, mononematous, synnematous, branched, unbranched. Hyalorhinocladiella-like; conidiophores simple or sparingly branched; conidiogenous cells sympodial, subulate; conidia hyaline, 1-celled, cylindrical or clavate to broadly clavate, oblong or ellipsoidal, sometimes slightly curved. Pesotum-like; stipes pale to dark pigmented, biverticillate; conidiogenous cells sympodial.
Type species: Graphilbum sparsum H.P. Upadhyay & W.B. Kendr., Mycologia 67: 800. 1975. MycoBank MB 314730.
Other species: Listed in Table 1.
Notes: Graphilbum was originally described to accommodate the asexual morph of some Ophiostoma (then Ceratocystis) species (Upadhyay & Kendrick 1975). Okada et al. (1998) treated Graphilbum together with other genera in the Ophiostomatales, which have synnematous asexual morphs, as a synonym of Pesotum. However, De Beer et al. (2013a) showed that the type species of Pesotum, O. ulmi, grouped within Ophiostoma s.s., while the type species of Graphilbum, O. sparsum, formed a lineage distinct from Ophiostoma s.s. They consequently reduced Pesotum to synonymy with Ophiostoma, and reinstated Graphilbum with Gra. sparsum as the type species, including seven additional species in the genus. Graphilbum species are characterised by Type E ascospores (Fig. 3H) and either a hyalorhinocladiella- or pesotum-like asexual morph (Fig. 3A, C; sensu De Beer & Wingfield 2013). This is with the exception of Gra. tsugae that produces a continuum of hyalorhinocladiella-like and pesotum-like asexual morphs. Graphilbum species are commonly found in galleries of conifer-infesting bark beetles in the Northern Hemisphere (De Beer et al. 2013b, Romón et al. 2014a, Reid & Hausner 2015, Jankowiak et al. 2020).
In our analyses, Graphilbum formed a well-supported monophyletic lineage, distinct from other genera. Interestingly, Ophiostoma pusillum consistently grouped with Graphilbum, supporting previous suggestions that the species might form part of this genus (De Beer & Wingfield 2013).
New combination:
1) Graphilbum pusillum (Masuya) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840324.
Basionym: Ophiostoma pusillum Masuya, Mycoscience 44: 302. 2003. MycoBank MB 489291.
Description: Masuya et al. (2003: 302, figs 1–10).
Phylogenetic data: Fig. 5.
Notes: Masuya et al. (2013) treated this species as part of the O. ips complex based on morphology. However, due to morphological similarities with the fungus known as Ophiostoma nigrum and Ceratocystis tubicollis (Masuya et al. 2003), De Beer et al. (2013b) suggested that it may rather belong in Graphilbum. Both O. nigrum (now Gra. nigrum) and C. tubicollis (now Gra. tubicolle) were treated in Graphilbum by De Beer et al. (2013b), and our sequence data also confirm the placement of O. pusillum in this genus (Fig. 5). The name should not be confused with S. pusilla U. Braun & Crous [= Quambalaria pusilla (U. Braun & Crous) J.A. Simpson] (De Beer et al. 2006) or Graphium pusillum (Wallr.) Sacc. (De Beer et al. 2013b).
Grosmannia (Lineage II)
Grosmannia Goid., Boll. Staz. Patol. Veg. Roma 16: 27. 1936. MycoBank MB 2141, emend. Z.W. de Beer & M. Procter. Fig. 8A–D.
Fig. 8.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Grosmannia grandifoliae complex. C, D. Grosmannia penicillata complex. E, F. Harringtonia. G, H. Hawksworthiomyces. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Etymology: Goidànich (1935) named the genus after the German forest pathologist, Helene Grosmann, later Francke-Grosmann (1900–1990), who described both the asexual morph (as Leptographium penicillatum Grosmann) and the sexual morph (as Ceratostomella penicillata Grosmann) of the type species of this genus.
Synonym: Verticicladiella S. Hughes, Canad. J. Bot. 31: 653. 1953. MycoBank MB 10394. [Type species V. abietina (Peck) S. Hughes].
Sexual morph: Ascomatal bases subglobose to globose, black; necks nearly cylindrical, straight or slightly curved, black. Ostiolar hyphae absent or present, divergent or variable. Asci evanescent. Ascospores hyaline, 1-celled, enclosed in sheath or not, endospores orange segment-shaped in side view, cylindrical in face view, globose to ellipsoidal in end view.
Asexual morph: Conidiophores macronematous, micronematous, mononematous, synnematous, singly or in groups. Hyalorhinocladiella-like; conidia hyaline, 1-celled, oblong to obovoid. Leptographium-like; conidiophores branched; stipes pale brown, branches pale brown, becoming hyaline towards apex, bases simple or rhizoid-like; conidiogenous cells hyaline; conidia hyaline, 1-celled, oblong, broadly ellipsoidal to slightly obovoid, sometimes allantoid, narrowly obovoid to clavate, straight or distinctly curved. Pesotum-like; conidiophores single or in groups; conidia hyaline, oblong, 1-celled. Sporothrix-like; conidiophores simple; conidiogenous cells hyaline, cylindrical, sympodial, denticulate; conidia hyaline, 1-celled, oblong, ovoid to ellipsoidal, clavate, sometimes developing into larger ramoconidia, producing secondary spores. Ramoconidia hyaline, 1-celled, clavate. Single ascospore cultures yeasty, budding hyaline conidia, ovoid to elongate, colony later darkened. Aleuriospores hyaline or pigmented, globose to subglobose, oval to ellipsoid.
Type species: Grosmannia penicillata (Grosmann) Goid., Boll. Staz. Patol. Veg. Roma 16: 46. 1936. MycoBank MB 253870.]
Notes: Grosmannia was originally erected by Goidànich (1935) to accommodate sexual species of Ceratostomella with leptographium-like asexual morphs (Davidson 1942). The genus was later treated as synonym of both Ophiostoma (Siemaszko 1939) and Ceratocystis (Bakshi 1951). Zipfel et al. (2006) showed that Ophiostoma and Grosmannia were distinct from each other based on ITS and LSU sequences, and separated the two genera based on these and morphological differences. However, the focus of the Zipfel et al. (2006) study was primarily on sexually reproducing species.
De Beer & Wingfield (2013) included sequence data for many more asexual Leptographium spp. in their study and showed that Leptographium and Grosmannia spp. grouped together, along with other previously unassociated Ophiostoma spp. They applied the older name, Leptographium to this group, rather than Grosmannia, which following the dual nomenclature system (McNeill et al. 2012) had preference because it was considered a sexual genus (De Beer & Wingfield 2013). They referred to the lineage as Leptographium s.l., even though the lineage did not show strong monophyletic support. The type species of Grosmannia (G. penicillata) grouped in a lineage distinct from the type species of Leptographium (L. lundbergii). However, they recommended that novel species grouping in what they referred to as the G. penicillata complex should be treated as Grosmannia species. This was until more robust analyses could confirm whether the G. penicillata complex should be treated as a distinct genus.
Based on the data emerging from the present study, we have reinstated Grosmannia as a genus distinct from Leptographium, for species that produce leptographium-like asexual morphs and have allantoid, hyaline and aseptate ascospores. These species are commonly associated with conifer-infesting bark beetles in the Northern Hemisphere (Jacobs & Wingfield 2001, Linnakoski et al. 2012). Based on our phylogenetic analyses, Grosmannia includes the fungus previously known as G. penicillata s.s. (referred to here as the G. penicillata complex), G. abieticola and L. taigense (Lineage C), and the newly recognised G. grandifoliae complex (Jankowiak et al. 2017).
Species complexes:
The G. penicillata complex
The G. penicillata complex as defined by Six et al. (2011), Linnakoski et al. (2012) and De Beer & Wingfield (2013) included 18 species. Our results and those emerging from other studies add another seven species to the complex within a well-supported lineage. All 25 species are associated in some way with conifer-infesting bark and ambrosia beetles in the Northern Hemisphere. The complex is named based on the first species in the complex to be described, G. penicillata [as Ceratostomella penicillata (Grosmann 1932)], which is also the type species of Grosmannia.
The G. grandifoliae complex
The second species complex included in our definition of Grosmannia, the G. grandifoliae complex, emerged when two new species of Leptographium were described from Poland (Jankowiak et al. 2017). Both species formed a well-supported lineage with G. grandifoliae and L. pruni, sister to the G. penicillata complex. Prior to the description of the new species, L. pruni and G. grandifoliae did not form part of the G. penicillata complex or any of the other species complexes recognised by De Beer et al. (2013b) in Leptographium s.l. Jankowiak et al. (2017) suggested that G. grandifoliae should be the name-bearing species of this small complex, because it is the oldest known name among the species that it accommodates.
Grosmannia grandifoliae is the only species for which a sexual morph has been observed and that is characterised by ascomata with very long necks (Jacobs & Wingfield 2001). With the addition of the two new species from Poland, L. trypodendri and L. betulae, it became evident that this complex represents a well-supported hardwood-infesting lineage in Grosmannia (Fig. 5) (Jankowiak et al. 2017). Leptographium trypodendri was isolated mostly from Trypodendron (beetle) species and L. betulae in association with Scolytus species (Jankowiak et al. 2017). This is the first species complex in Leptographium and Grosmannia that accommodates only hardwood-infesting species (Jankowiak et al. 2017).
In our datasets, the G. grandifoliae complex grouped within Lineage II in all except the TEF-1α tree (Fig. S3) where it grouped with Group C (below) among species complexes in Leptographium. The clear ecological distinction and the well-supported monophyletic lineage of the G. grandifoliae complex, supports its treatment as novel genus. In the future, this will need to be verified using more robust phylogenies including additional gene regions, and if possible, inclusion of a greater number of species in the complex.
Group C
In the phylogenies of Linnakoski et al. (2012), L. taigense grouped in a distinct lineage between the G. grandifoliae and G. penicillata complexes. In the LSU tree generated by De Beer & Wingfield (2013), L. taigense also grouped outside of the G. penicillata complex, close to, but not in a well-supported lineage with G. abieticola. With the additional gene regions included in this study, L. taigense and G. abieticola emerged together in a well-supported lineage distinct from, but between the two complexes defined above. Both these species are associated with conifers, as is true for all species in the G. penicillata complex. For the present, we have chosen to treat them in Grosmannia, but inclusion of newly discovered species in this lineage in the future could justify the description of a new genus to accommodate these species.
New combinations:
1) Grosmannia abieticolens (K. Jacobs & M.J. Wingf.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840331.
Basionym: Leptographium abieticolens K. Jacobs & M.J. Wingf., Mycoscience 41: 599. 2000. MycoBank MB 466545.
Description: Jacobs & Wingfield (2001: 46–48, figs 19–21).
Phylogenetic data: Jacobs et al. (2001b), Kim et al. (2004, 2005a), Masuya et al. (2004), Massoumi Alamouti et al. (2006), Paciura et al. (2010), Six et al. (2011), Duong et al. (2012), De Beer & Wingfield (2013), Huang & Chen (2014), Musvuugwa et al. (2015), Chang et al. (2019).
Notes: Sexual morph unknown. While it was not included in our study, G. abieticolens groups in the G. penicillata complex based on previously published data (Six et al. 2011, De Beer & Wingfield 2013, Chang et al. 2019). The name should not be confused with Grosmannia abieticola that formed part of Group C in our analyses.
Notes on Grosmannia abietina: Sexual morph unknown. Grosmannia abietina groups in the G. penicillata species complex (Fig. 4; Six et al. 2011, Linnakoski et al. 2012, De Beer & Wingfield 2013). The species should not be confused with Sporothrix abietina, that resides in Sporothrix. Jacobs et al. (1998) proposed the synonymy of G. engelmannii (as L. engelmannii) with G. abietina (as L. abietinum), which was supported by Jacobs & Wingfield (2001). In our analyses, an isolate of L. engelmannii (CMW 759) grouped with G. abietina in the TEF1-α tree (Fig. S3) and separate from that species in the LSU tree (Fig. S1).
2) Grosmannia altior (Paciura, Z.W. de Beer & M.J. Wingf.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840369.
Basionym: Leptographium altius Paciura et al., Persoonia 25: 106. 2010. MycoBank MB 516740.
Description: Paciura et al. (2010: 106, fig. 7h–m).
Phylogenetic data: Paciura et al. (2010), Duong et al. (2012), Linnakoski et al. (2012), De Beer & Wingfield (2013), Liu et al. (2017), Chang et al. (2019).
Notes: Sexual morph unknown. This species forms part of the G. penicillata complex based on previously published data (Paciura et al. 2010, Linnakoski et al. 2012, De Beer & Wingfield 2013, Liu et al. 2017, Chang et al. 2019).
3) Grosmannia betulae (Jankowiak et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840370.
Basionym: Leptographium betulae Jankowiak et al., Antonie van Leeuwenhoek 110: 1550. 2017. MycoBank MB 821670.
Description: Jankowiak et al. (2017: 1550, fig. 6).
Phylogenetic data: Jankowiak et al. (2017, 2018).
Notes: Sexual morph unknown. Groups in the G. grandifoliae species complex (Fig. 5; Jankowiak et al. 2017).
4) Grosmannia curviconidia (Paciura et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840373.
Basionym: Leptographium curviconidium Paciura et al., Persoonia 25: 104. 2010. MycoBank MB 516739.
Description: Paciura et al. (2010: 104–105, figs 7a–g).
Phylogenetic data: Paciura et al. (2010), Duong et al. (2012), Linnakoski et al. (2012), De Beer & Wingfield (2013), Huang & Chen (2014), Musvuugwa et al. (2015), Jankowiak et al. (2017, 2018), Liu et al. (2017), Chang et al. (2019).
Notes: Sexual morph unknown. Grosmannia curviconidia groups in the G. penicillata species complex (Fig. 5; Linnakoski et al. 2012, De Beer & Wingfield 2013, Huang & Chen 2014, Musvuugwa et al. 2015, Jankowiak et al. 2017, 2018, Liu et al. 2017, Chang et al. 2019).
5) Grosmannia euphyes (K. Jacobs & M.J. Wingf.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840376.
Basionym: Leptographium euphyes K. Jacobs & M.J. Wingf., Mycol. Res. 105: 497. 2001. MycoBank MB 467761.
Descriptions: Jacobs et al. (2001c: 496–498, figs 15–21), Jacobs & Wingfield (2001: 96–99, figs 70–72).
Phylogenetic data: Jacobs et al. (2001a), Kim et al. (2004, 2005a), Masuya et al. (2004), Massoumi Alamouti et al. (2006), Paciura et al. (2010), Six et al. (2011), Duong et al. (2012), De Beer & Wingfield (2013), Jankowiak et al. (2017, 2018), Liu et al. (2017), Chang et al. (2019).
Notes: Sexual morph unknown. This species grouped in the G. penicillata complex based on previous data (Six et al. 2011, De Beer & Wingfield 2013, Jankowiak et al. 2017, 2018, Liu et al. 2017, Chang et al. 2019).
6) Grosmannia fenglinhensis (R. Chang et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840377.
Basionym: Leptographium fenglinhense R. Chang et al., Persoonia 42: 67. 2018 (2019). MycoBank MB 825092.
Description: Chang et al. (2019: 66–67, fig. 21).
Phylogenetic data: Chang et al. (2019)
Notes: Sexual morph unknown. Groups in the G. penicillata species complex (Fig. 5; Chang et al. 2019).
7) Grosmannia gestamen (Errasti & Z.W. de Beer) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840381.
Basionym: Leptographium gestamen Errasti & Z.W. de Beer, Mycol. Prog. 15: 11. 2016. MycoBank MB 814177.
Descriptions: De Errasti et al. (2016: 11–13, fig. 7).
Phylogenetic data: De Errasti et al. (2016), Jankowiak et al. (2017, 2018), Chang et al. (2019).
Notes: Sexual morph unknown. This species grouped close to G. taigensis (as L. taigense) in the phylogenies of Jankowiak et al. (2017) and Chang et al. (2019), and with G. abieticola (Jankowiak et al. 2018). We have consequently included it in Grosmannia, as part of the complex defined by the genus in group C (Fig. 5).
8) Grosmannia innermongolica (X.W. Liu et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840384.
Basionym: Leptographium innermongolicum X.W. Liu et al., Mycol. Prog. 16: 8. 2017. MycoBank MB 811204.
Description: Liu et al. (2017: 8–10, fig. 6).
Phylogenetic data: Liu et al. (2017), Chang et al. (2019).
Notes: Sexual morph unknown. This species grouped with L. taigense (Group C in Fig. 5) in the phylogenies of Liu et al. (2017) and Chang et al. (2019). It is consequently included in Grosmannia (Fig. 5).
9) Grosmannia pistaciae (Paciura et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840385.
Basionym: Leptographium pistaciae Paciura et al., Persoonia 25: 104. 2010. MycoBank MB 516738.
Description: Paciura et al. (2010: 104, figs 6g–l).
Phylogenetic data: Paciura et al. (2010), Duong et al. (2012), Linnakoski et al. (2012), De Beer & Wingfield (2013), Jankowiak et al. (2017, 2018), Liu et al. (2017), Chang et al. (2019).
Notes: Sexual morph unknown. Leptographium pistaciae has previously been shown to reside in the G. penicillata complex (Paciura et al. 2010, Linnakoski et al. 2012, De Beer & Wingfield 2013, Jankowiak et al. 2017, 2018, Liu et al. 2017, Chang et al. 2019).
10) Grosmannia pruni (Masuya & M.J. Wingf.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840386.
Basionym: Leptographium pruni Masuya & M.J. Wingf., Mycologia 96: 553. 2004. MycoBank MB 488574.
Description: Masuya et al. (2004: 553–555, figs 1–16).
Phylogenetic data: Masuya et al. (2004, 2013), Massoumi Alamouti et al. (2006), Matsuda et al. (2010), Duong et al. (2012), De Beer & Wingfield (2013), De Errasti et al. (2016), Jankowiak et al. 2017, 2018), Chang et al. (2019).
Notes: Sexual morph unknown. This species grouped with G. grandifoliae and two other species to form the G. grandifoliae species complex (Jankowiak et al. 2017).
11) Grosmannia taigensis (Linnak. et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840387.
Basionym: Leptographium taigense Linnak. et al., Antonie van Leeuwenhoek 102: 387. 2012. MycoBank MB 564881.
Description: Linnakoski et al. (2012: 387–388, fig. 7).
Phylogenetic data: Linnakoski et al. (2012), De Beer & Wingfield (2013), Jankowiak et al. (2017, 2018), Liu et al. (2017), Chang et al. (2019).
Notes: Sexual morph unknown. This species grouped with G. abieticola, separated from other species complexes in Grosmannia (Fig. 5), based on LSU (Fig. S1) and TEF1-α (Fig. S3) sequence data.
12) Grosmannia trypodendri (Jankowiak et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840388.
Basionym: Leptographium trypodendri Jankowiak et al., Antonie van Leeuwenhoek 110: 1546. 2017. MycoBank MB 821669.
Description: Jankowiak et al. (2017: 1546–1550, fig. 5).
Phylogenetic data: Jankowiak et al. (2017, 2018).
Notes: Sexual morph unknown. Groups in the G. grandifoliae species complex (Fig. 5).
Other species: Listed in Table 1.
Harringtonia (Lineage XI)
Harringtonia Z.W. de Beer & Procter, gen. nov. MycoBank MB 840400. Fig. 8E, F.
Etymology: Named for the American mycologist Prof. Thomas C. Harrington, who described the type species of this genus as Raffaelea lauricola (Harrington et al. 2008), and who has contributed substantially to our understanding of the biology and taxonomy of the Ophiostomatales (Harrington 1981, 1988, Harrington & Cobb 1987, Harrington et al. 2001, 2008, 2010, 2011).
Synonym: Raffaelea lauricola complex sensu Z.W. de Beer & M.J. Wingf., In The Ophiostomatoid Fungi: Expanding Frontiers: 34. 2013.
Sexual morph: Unknown.
Asexual morph: Colonies initially yeast-like at centre becoming cottony in 2 wk (Har. lauricola), cream-coloured, having submerged mycelium at margins, aging to dark green on MEA (Har. aguacate). Colonies when repeatedly subcultured forming a filamentous mycelium, with surface becoming finely tomentose (Har. brunnea). Conidiophores single or aggregated in sporodochia, hyaline, macronematous, semimacronematous, mononematous, simple or branched. Conidiogenous cells with inconspicuous annellations at point of conidial dehiscence. Conidia hyaline, 1-celled, oblong to ellipsoidal, obovoid, truncated at base, occasionally borne sessile, in some species producing secondary budding cells. Associated with ambrosia beetles.
Type species: Harringtonia lauricola (T.C. Harr. et al.) Z.W. de Beer & M. Procter, comb. nov. MycoBank MB 840401.
Basionym: Raffaelea lauricola T.C. Harr. et al., Mycotaxon 104: 401. 2008. MycoBank MB 511590.
Description: Fraedrich et al. (2008: 219–220, fig. 5).
Phylogenetic data: Fraedrich et al. (2008), Harrington et al. (2008, 2010, 2011), Kim et al. (2009), Massoumi Alamouti et al. (2009), Matsuda et al. (2010), De Beer & Wingfield (2013), Dreaden et al. (2014), Musvuugwa et al. (2015), Simmons et al. (2016), De Beer et al. (2016a, b), Bateman et al. (2017), Vanderpool et al. (2018), Wingfield et al. (2017), Li et al. (2018), Saucedo-Carabez et al. (2018).
Notes: Harringtonia lauricola consistently forms a clade together with Har. brunnea and Har. aguacate, that has previously been referred to as the R. lauricola complex (De Beer & Wingfield 2013). This complex has always formed a monophyletic lineage close to, but distinct from Raffaelea s.s. in previous studies (Massoumi Alamouti et al. 2009, De Beer & Wingfield 2013, Dreaden et al. 2014, Simmons et al. 2016). When De Beer & Wingfield (2013) first defined this species complex, they chose R. lauricola as the name bearing species, because it is the best-known species in this complex. This is despite the fact that it was not the first species in the complex to be described. They suggested that this lineage would likely emerge as a new genus when more robust phylogenetic analyses were performed, as is the case in the present study.
In the analyses of our datasets, the R. lauricola complex grouped close to Raffaelea s.s. and together they formed a monophyletic clade, although this relationship was not suppoted. Phylogenomic analyses conducted in the current study and previous studies (Nel et al. 2021; Vanderpool et al. 2018) showed that the R. lauricola complex and Raffaelea s.s. did not form a monophyletic lineage. Based on these results, we have removed the R. lauricola complex from Raffaelea and have described the complex as a new genus. While fully cogniscient of the fact that describing a new genus to accommodate an important tree pathogen might result in some early discomfort, this change is based on robust taxonomic data, not only from the present study. Failing to make this change perpetuates a taxonomically confused situation. In our view this decision will also provide opportunities to better understand the biology and evolutionary characteristics of two phylogenetically distinct and important groups of fungi.
All species in the R. lauricola complex produce raffaelea-like asexual morphs (De Beer & Wingfield 2013). Harringtonia lauricola is the causal agent of laurel wilt, a vascular wilt disease affecting Lauraceae species in the south-eastern USA. It is believed to have been introduced into the USA from Asia in the mycangia of Xyleborus glabratus, an ambrosia beetle invasive to the USA (Harrington et al. 2008). The other species in this complex are Har. brunnea, associated with ambrosia beetles infesting hardwoods (De Beer & Wingfield 2013), and Har. aguacate, originally isolated from avocado (Persea americana) (Simmons et al. 2016) and later shown to be vectored by X. bispinatus (Saucedo-Carabez et al. 2018).
Other new combinations:
1) Harringtonia aguacate (D.R. Simmons et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840402.
Basionym: Raffaelea aguacate D.R. Simmons et al., IMA Fungus 7: 269. 2016. MycoBank MB 817170.
Description: Simmons et al. (2016: 269, fig. 2).
Phylogenetic data: Simmons et al. (2016), Vanderpool et al. (2018), Li et al. (2018), Saucedo-Carabez et al. (2018).
Notes: This species forms part of Harringtonia (Fig. 5), previously the R. lauricola complex (De Beer & Wingfield 2013).
2) Harringtonia brunnea (L.R. Batra) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840403.
Basionym: Ambrosiella brunnea L.R. Batra, Mycologia 59: 980. 1968 (1967). MycoBank MB 326140.
Synonyms: Monilia brunnea Verrall, J. Agric. Res. 66: 142. 1943. MycoBank MB 440994. nom. illegit., Art. 53. 1.
Raffaelea brunnea (L.R. Batra) T.C. Harr., Mycotaxon 111: 351. 2010. MycoBank MB 515296.
Descriptions: Verrall (1943: 142–143, fig. 5), Batra (1967: 1004–1007, figs 43, 45, 46).
Phylogenetic data: Cassar & Blackwell (1996), Rollins et al. (2001), Gebhardt et al. (2005), Massoumi Alamouti et al. (2009), Harrington et al. (2010), Matsuda et al. (2010), De Beer & Wingfield (2013), Dreaden et al. (2014), Musvuugwa et al. (2015), De Beer et al. (2016a, b), Vanderpool et al. (2018), Wingfield et al. (2017), Li et al. (2018), Saucedo-Carabez et al. (2018).
Notes: This species forms part of Harringtonia (Fig. 5), previously the R. lauricola species complex (De Beer & Wingfield 2013). This species should not be confused with Monilia brunnea, a soil inhabiting species.
Hawksworthiomyces (Lineage XXII)
Hawksworthiomyces Z.W. de Beer et al., Fungal Biol. 120: 1329. 2016. MycoBank MB 815685. Fig. 8G, H.
Etymology: ‘Named for Dr David Hawksworth, in recognition of the leading role that he has played in guiding the global mycological community through the controversial and often challenging transition from a dual nomenclature to a One Fungus-One Name-based system’ (De Beer et al. 2016b).
Sexual morph: Unknown.
Asexual morph: Conidiophores hyaline, micronematous to macronematous, mononematous, simple or branched. Conidiogenous cells hyaline, polyblastic, denticulate. Conidia hyaline, 1-celled, broadly ellipsoidal to cylindrical, producing secondary cells.
Type species: Hawksworthiomyces lignivorus (De Mey. et al.) Z.W. de Beer et al., Fungal Biol. 120: 1329. 2016. MycoBank MB 815686.
Other species: Listed in Table 1.
Notes: Hawksworthiomyces was erected to accommodate Sporothrix lignivora (now Haw. lignivora) and three other species not previously associated with the Ophiostomatales. De Beer & Wingfield (2013) listed S. lignivora as “incertae sedis”, as this species formed a lineage on its own, separate from other species in Ophiostoma and Sporothrix, even though its morphology resembled species of Sporothrix (De Meyer et al. 2008). BLAST searches on GenBank later revealed sequences from environmental studies and isolates from diversity studies with strong similarity to S. lignivora. Consequently, De Beer et al. (2016) proceeded to erect the new genus, Hawksworthiomyces, to incorporate S. lignivora and these “new” species. In the analyses of our data, Haw. lignivora, Haw. taylorii, Haw. hibbettii and Haw. crousii formed a well-supported monophyletic lineage, supporting their treatment in a distinct genus. An ENAS (Environmental Nucleic Acid Sequence) species, named Haw. sequentia ENAS, was also included in Hawksworthiomyces by De Beer et al. (2016). Hawksworthiomyces species have been isolated from diverse sources (e.g. Eucalyptus wood poles, insect fungal gardens, rhizospheres of plants) in South Africa, North and Central America, South Korea and Europe (De Beer et al. 2016). In the analyses of our datasets based on protein coding genes (Figs S3, S4) Hawksworthiomyces spp. formed two separate lineages. However, in the concatenated (Fig. 5) and LSU (Fig. S1) datasets, a monophyletic lineage emerged.
Heinzbutinia (Lineage XXIV)
Heinzbutinia Z.W. de Beer & M. Procter, gen. nov. MycoBank MB 840404. Fig. 9A, B.
Fig. 9.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Heinzbutinia. C, D. Intubia. E, F. Jamesreidia. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Etymology: Named for Prof. Heinz Butin (1928–2021), a German forest pathologist who described 10 new species in the Ophiostomatales between 1968 and 1990, including the type species of this novel genus.
Sexual morph: Ascomatal bases black, subglobose to globose; necks black, cylindrical, curved. Ostiolar hyphae absent. Asci elongate ovoid or clavate, evanescent. Ascospores hyaline, 1-celled, orange segment-like in side view, ellipsoidal in face view.
Asexual morph: Sporothrix-like; conidiophores micronematous, macronematous, mononematous, simple or branched, hyaline; conidiogenous cells sympodial, denticulate; conidia hyaline, 1-celled, ellipsoid, often curved, reniform, secondary conidia originating from swollen ellipsoidal conidia, oblong, straight or curved.
Type species: Heinzbutinia grandicarpa (Kowalski & Butin) Z.W. de Beer & M. Procter, comb. nov. MycoBank MB 840405.
Basionym: Ceratocystis grandicarpa Kowalski & Butin, J. Phytopathol. 124: 243. 1989. MycoBank MB 134497.
Synonym: Ophiostoma grandicarpum [as ‘grandicarpa’] (Kowalski & Butin) Rulamort, Bull. Soc. Bot. Centre-Ouest 21: 511. 1990. MycoBank MB 130251.
Phylogenetic data: Villarreal et al. (2005), De Beer & Wingfield (2013), Reid & Hausner (2015), De Beer et al. (2016a, b), De Errasti et al. (2018).
Notes: The O. grandicarpum complex was described by De Beer et al. (2016b) for the monophyletic lineage formed by O. grandicarpum and O. microsporum among other genera in the Ophiostomatales (De Beer & Wingfield 2013, Reid & Hausner 2015, De Beer et al. 2016a, De Errasti et al. 2018). In our expanded phylogenies, the two species again consistently formed a well-supported lineage distinct from all other genera in the Ophiostomatales (Figs 5, S2–S4). Both species are characterised by very long (more than 1 mm) ascomatal necks (Kowalski & Butin 1989, Hunt 1956), sporothrix-like asexual morphs and reniform ascospores (Fig. 3F; Type A as categorised by De Beer & Wingfield 2013). Both species were isolated from hardwoods. Based on the distinct phylogenetic placement and supported by a unique morphology and ecology, we have designated the O. grandicarpum complex as the new genus Heinzbutinia.
Other new combinations:
1) Heinzbutinia microspora (Arx) Z.W. de Beer & M. Procter, comb. nov. MycoBank MB 840406.
Basionym: Ophiostoma microsporum Arx, Antonie van Leeuwenhoek 18: 211. 1952. MycoBank MB 302079.
Synonyms: Ceratostomella microspora R.W. Davidson, Mycologia 34: 650. 1942. MycoBank MB 284860. nom. illegit., Art. 53.1, later homonym for Cs. microspora Ellis & Everh., see De Beer et al. 2013, Section C.1.
Ceratocystis perparvispora J. Hunt, Lloydia 19: 46. 1956. MycoBank MB 294227. Art. 52.1, superfluous nom. nov.
Ceratocystis microspora (R.W. Davidson) R.W. Davidson & Aoshima, Ph.D. thesis, University of Tokyo: 20. 1965. nom. inval., Arts 29.1, 39.1 or 39.2.
Ceratocystis microspora (Arx) R.W. Davidson, J. Colorado-Wyoming Acad. Sci. 6: 16. 1969. MycoBank MB 453851.
Descriptions: Hunt (1956: 46–47), Griffin (1968: 710), De Hoog (1974: 63–64, fig. 25), Olchowecki & Reid (1974: 1709), Upadhyay (1981: 50, figs 104–108), Maekawa et al. (1987: 8, 10, figs 1–6).
Phylogenetic data: Hausner et al. (1993b), Mullineux et al. (2011), De Beer & Wingfield (2013), Musvuugwa et al. (2015), De Beer et al. (2016a, b).
Notes: This species grouped with He. grandicarpa and distinct from other Ophiostoma spp. (De Beer et al. 2016a, b), in the O. grandicarpum complex as designated by De Beer et al. (2016b). See note under He. grandicarpa. The name He. microspora should not be confused with Leptographium microsporum or Ceratostomella microspora (see De Beer et al. 2013, section C.2).
2) Heinzbutinia solheimii (Strzałka & Jankowiak) Z.W. de Beer & M. Procter, comb. nov. MycoBank MB 840407.
Basionym: Ophiostoma solheimii Strzałka & Jankowiak, Antonie van Leeuwenhoek 112: 1517. 2019. MycoBank MB 830198.
Description: Jankowiak et al. (2019: 1516–1517, fig. 8).
Phylogenetic data: Jankowiak et al. (2019).
Notes: This species is closely related to and formed a well-supported lineage with He. grandicarpa and He. microspora (Jandowiak et al. 2019, figs 1, S4).
Intubia
Intubia W.J. Nel et al., Mycologia 113: 1206. 2021. MycoBank MB 837565. Fig. 9C, D.
Etymology: ‘From the Xhosa language, Intubi for termite, recognising the source where the fungus was found’ (Nel et al. 2021).
Sexual morph: Ascomatal bases dark brown, globose; necks uniformly dark, often slightly curved, tapering towards apex, extremely long. Ostiolar hyphae absent. Asci not seen. Ascospores produced in slimy droplet at apex of neck, hyaline, 1-celled, cylindrical, sometimes slightly curved, no sheath.
Asexual morph: Conidiophores micronematous, hyaline, arising singly. Hyalorhinocladiella-like; conidiogenous cells smooth; conidia hyaline, 1-celled, bacilliform tapering toward one end, occasionally producing secondary spores. Sporothrix-like; conidiogenous cells denticulate; conidia of two types, first type formed on vegetative hyphae, hyaline, 1-celled, round to obovoid, second type formed on conidiophores, hyaline, 1-celled, bacilliform, secondary conidia present.
Type species: Intubia macrotermitinarum W.J. Nel et al., Mycologia 113: 1208. 2021. MycoBank MB 837567.
Other species: Listed in Table 1.
Notes: Distinguished from other phylogenetically related genera in the Ophiostomatales for their unique habitat with dark coloured ascomata embedded in the substrate of abandoned Termitomyces combs.
Jamesreidia (Lineage XII)
Jamesreidia Z.W. de Beer & M. Procter, gen. nov. MycoBank MB 840408. Fig. 9E, F.
Etymology: Named for the Canadian mycologist, Dr James Reid, who described 28 novel ophiostomatoid species, including the type species of this new genus. He was also involved in producing some of the first phylogenetic data based on ribosomal sequences distinguishing between the Ophiostomatales and Microascales (Olchowecki & Reid 1974, Hausner et al. 1992).
Synonym: Ophiostoma tenellum complex sensu Z.W. de Beer & M.J. Wingf., In The Ophiostomatoid Fungi: Expanding Frontiers: 34. 2013.
Sexual morph: Ascomatal bases black, globose; necks absent or present, black, straight or bent. Ostiolar hyphae divergent. Asci evanescent, clavate to subglobose. Ascospores hyaline, 1-celled, orange segment-like in side view, oblong in face view, broadly ellipsoidal to globose in end view.
Asexual morph: Conidiophores macronematous, semimacronema-tous, mononematous, hyaline, simple or branched. Conidiogenous cells polyblastic, denticulate. Conidia hyaline, 1-celled, broadly ovoid to nearly cylindrical, slightly curved.
Type species: Jamesreidia tenella (R.W. Davidson) Z.W. de Beer & M. Procter, comb. nov. MycoBank MB 840409.
Basionym: Ceratocystis tenella R.W. Davidson, Mycologia 50: 666. 1958. MycoBank MB 294240.
Synonyms: Ophiostoma tenellum (R.W. Davidson) M. Villarreal, Mycotaxon 92: 263. 2005. MycoBank MB 346181.
Ceratocystis capitata H.D. Griffin, Canad. J. Bot. 46: 699. 1968. MycoBank MB 327623.
Descriptions: Griffin (1968: 713, 715, fig. 93, pI. Ill), Olchowecki & Reid (1974: 1708, pI. XVI figs 307, 308, 311, 312), Upadhyay (1981: 114, figs 408–412), Maekawa et al. (1987: 10–11, figs 19, 20), Hutchison & Reid (1988a: 68).
Phylogenetic data: Villarreal et al. (2005), Linnakoski et al. (2010, 2016), De Beer & Wingfield (2013), De Errasti et al. (2018), De Beer et al. (2016a), De Errasti et al. (2018), Wang et al. (2018), Chang et al. (2019).
Notes on the type species: Jamesreidia tenella groups with J. coronata, J. nigricarpa (Fig. 5; Linnakoski et al. 2010, De Beer & Wingfield 2013) and J. rostrocoronata (in the phylogenies of De Beer et al. 2016a) to form Jamesreidia, previously designated as the O. tenellum species complex (De Beer & Wingfield 2013). This species serves as the type species of the new genus, due to the fact that it is the oldest described species in the genus.
Notes on the genus: The O. tenellum complex was defined in Ophiostoma s.l. by De Beer & Wingfield (2013), who noted that members of the complex shared a similar morphology with species in the S. schenckii-O. stenoceras complex (also treated in Ophiostoma s.l. at the time, but now recognised as the genus Sporothrix). The asexual morphs of all species can be described as sporothrix-like and sexual morphs form Type A ascospores (Fig. 3F; as categorised by De Beer & Wingfield 2013). All species in this complex colonise conifer wood in North America (Linnakoski et al. 2010).
De Beer et al. (2016a) noted that the generic status of the O. tenellum complex should be further investigated, because it grouped distinctly from both Sporothrix and Ophiostoma s.s. in their phylogenies. In the present study, this complex grouped separately from what we have designated as Ophiostoma and Sporothrix in our phylogeny (Fig. 5) as well as in the phylogenies of other authors (Villarreal et al. 2005, Linnakoski et al. 2010, De Beer & Wingfield 2013, De Beer et al. 2016a). We have thus elevated the status of the O. tenellum complex to that of a genus, for which we have provided the name Jamesreidia.
Other new combinations:
1) Jamesreidia coronata (Olchow. & J. Reid) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840410.
Basionym: Ceratocystis coronata Olchow. & J. Reid, Canad. J. Bot. 52: 1705. 1974. MycoBank MB 310490.
Synonym: Ophiostoma coronatum (Olchow. & J. Reid) M. Villarreal, Mycotaxon 92: 263. 2005. MycoBank MB 346368.
Description: Hutchison & Reid (1988a: 66, 68).
Phylogenetic data: Hausner et al. (1993b), Thwaites et al. (2005), Villarreal et al. (2005), Linnakoski et al. (2010), Mullineux et al. (2011), De Beer & Wingfield (2013), Linnakoski et al. (2016), De Beer et al. (2016a), De Errasti et al. (2018), Wang et al. (2018), Chang et al. (2019).
Notes: This species grouped in Jamesreidia (Fig. 5), previously designated as the O. tenellum species complex (De Beer & Wingfield 2013).
2) Jamesreidia nigrocarpa (R.W. Davidson) M. Procter & Z.W. de Beer, comb. nov. [MycoBank MB 840411]
Basionym: Ceratocystis nigrocarpa R.W. Davidson, Mycopathol. Mycol. Appl. 28: 276. 1966. MycoBank MB 327637.
Synonym: Ophiostoma nigrocarpum (R.W. Davidson) de Hoog, Stud. Mycol. 7: 62. 1974. MycoBank MB 319024.
Descriptions: De Hoog (1974: 62–63, fig. 24), Olchowecki & Reid (1974: 1709), Upadhyay (1981: 104, figs 378–381), Benade et al. (1997: 1110–1111, figs 6–11).
Phylogenetic data: Aghayeva et al. (2004), Zhou et al. (2004a, 2006), Roets et al. (2006, 2008, 2010), Zipfel et al. (2006), De Meyer et al. (2008), Linnakoski et al. (2010), Madrid et al. (2010), Romón et al. (2014b), De Errasti et al. (2016, 2018), Linnakoski et al. (2016), De Beer et al. (2016a), Giraldo et al. (2017), Chang et al. (2019).
Notes: This species forms part of Jamesreidia (Fig. 5), previously the O. tenellum species complex (De Beer & Wingfield 2013).
3) Jamesreidia rostrocoronata (R.W. Davidson & Eslyn) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840412.
Basionym: Ceratocystis rostrocoronata R.W Davidson & Eslyn, Mem. N.Y. Bot. Gard. 28: 50. 1976. MycoBank MB 310518.
Synonym: Ophiostoma rostrocoronatum (R.W. Davidson & Eslyn) de Hoog & R.J. Scheff., Mycologia 76: 297. 1984. MycoBank MB 107077.
Descriptions: Upadhyay (1981: 112), Hutchison & Reid (1988a: 76–78).
Phylogenetic data: Hausner et al. (1993b), Jacobs et al. (2003), Villarreal et al. (2005), Linnakoski et al. (2010), De Beer & Wingfield (2013), Reid & Hausner (2015), De Beer et al. (2016b), Osorio et al. (2016), Jankowiak et al. (2017b), Wang et al. (2018).
Notes: A LSU sequence generated by Jacobs et al. (2003) was shown to group in Sporothrix (De Beer & Wingfield 2013, Osorio et al. 2016, Jankowiak et al. 2017, Wang et al. 2018). However, sequence data generated by De Beer et al. (2016b) for the same isolate, placed O. rostrocoronatum with the O. tenellum complex. Based on analyses of sequences from four gene regions and morphological characteristics, De Beer et al. (2016b) proceeded to treat O. rostrocoronatum in the O. tenellum complex. We have consequently treated this species as part of the lineage described here as Jamesreidia.
Leptographium (Lineage I)
Leptographium Lagerb. & Melin, Svenska Skogsv.-Fören. Tidskr. 25: 257. 1927. MycoBank MB 8749, emend. Z.W. de Beer & M. Procter. Figs 10, 11.
Fig. 10.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Leptographium clavigerum complex. C, D. Leptographium galeiforme complex. E, F. Leptographium lundbergii complex. G, H. Leptographium olivaceum complex. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Fig. 11.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Leptographium piceiperdum complex. C, D. Leptographium procerum complex. E, F. Leptographium serpens complex. G, H. Leptographium wageneri complex. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Etymology: ‘A thin, small brush. From the Greek adjective, λɛπτóζ (thin) and the Greek noun γραϕισν (a small brush). The generic name refers to the conidiophores that resemble small brushes’ (Jacobs & Wingfield 2001).
Synonyms: Scopularia Preuss, Linnaea 24: 133. 1851. MycoBank MB 22345, nom. illegit., Art. 53.1. (Type species Sc. venusta Preuss).
Phialographium H.P. Upadhyay & W.B. Kendr., Mycologia 66: 183. 1974. MycoBank MB 9340. (Type species Ph. sagmatosporae H.P. Upadhyay & W.B. Kendr.).
Graphiocladiella H.P. Upadhyay, A Monograph of Ceratocystis and Ceratocystiopsis: 138. 1981. MycoBank MB 8394. (Type species G. clavigera H.P. Upadhyay).
?= Europhium A.K. Parker, Canad. J. Bot. 35: 175. 1957. MycoBank MB 1938. [Type species E. trinacriforme A.K. Parker; Placement uncertain, see De Beer et al. (2013b)].
Sexual morph: Ascomata cleitothecial or perithecial; bases black, subglobose to globose; necks absent or present, very short, elongated, cylindrical, straight or flexuous. Ostiolar hyphae absent. Asci evanescent, clavate to broadly clavate. Ascospores hyaline, 1-celled, enclosed in sheath, reniform or cucullate in side view, oblong or rectangular in face view, ovoid, globose or triangular in end view.
Asexual morph: Conidiophores macronematous, micronematous, mononematous, singly or in groups, upright or prone, branches brown, becoming paler towards upper branches, bases simple or rhizoid-like. Conidiogenous cells hyaline, cylindrical. Conidia hyaline, 1-celled, oblong, pyriform, broadly ellipsoidal to obovoid, with truncate base. Hyalorhinocladiella-like; conidiophores simple or branched, upright; conidia hyaline, 1-celled, oblong to ellipsoid. Pesotum-like; conidiophores pale to dark brown, hyaline to light grey, reddish brown at apex; conidia hyaline, 0–4-celled, oblong, cylindrical to clavate, broadly fusiform, or subglobose to ellipsoid with truncate base. Ramoconidia hyaline, clavate to ellipsoid, 1-celled spores. Serprentine hyphae present in L. lundbergii, L. serpens complexes and L. douglasii in L. wageneri complex.
Type species: Leptographium lundbergii Lagerb. & Melin, Svenska Skogsv.-Fören. Tidskr. 25: 248. 1927. MycoBank MB 269891.
Synonym: Scopularia lundbergii (Lagerb. & Melin) Goid., Ann. Mycol. 31: 138. 1933. MycoBank MB 253103.
Notes on the genus: Leptographium was originally described as an asexual genus with L. lundbergii as type species (Lagerberg et al. 1927). Soon afterwards, the sexual genus Grosmannia was described to accommodate the sexual morph of another species, Leptographium penicillatum (Goidànich 1935). During the course of the next 65 years, an additional 28 asexual species were described in Leptographium and 17 species with known sexual morphs that were treated in different genera (Harrington 1988, Wingfield 1993). This prompted a revision of the genus in a comprehensive monograph by Jacobs & Wingfield (2001), who treated the sexual morphs of these species in Ophiostoma, of which Grosmannia was listed as one of several synonyms (Jacobs & Wingfield 2001). As mentioned in the introduction to this study, Zipfel et al. (2006) reinstated Grosmannia for all sexual species with Leptographium asexual morphs. However, they included only one asexual species, L. lundbergii, in their phylogeny.
De Beer & Wingfield (2013) were the first authors to apply the One Fungus-One Name principles to the Ophiostomatales, and listed Grosmannia as synonym of the older name, Leptographium. However, they recognised that the type species for these two genera grouped in different lineages but concluded that their LSU and ITS phylogenies were not sufficiently robust to resolve the generic status of these sub-lineages within what they referred to as ‘Leptographium sensu lato’. They listed 94 valid species in this broad concept but refrained from making new combinations for species that did not have names in Leptographium, recommending that phylogenies based on a greater number of gene regions should be used to resolve the genera.
Prior to the study of De Beer & Wingfield (2013), several authors defined phylogenetic species complexes in ‘Leptographium sensu lato’ (Massoumi Alamouti et al. 2011, Six et al. 2011, Duong et al. 2012, Linnakoski et al. 2012). The results of the present study clearly show that ‘Leptographium sensu lato’ represents two distinct genera. These include Leptographium (Lineage I) with L. lundbergii as type species and including eight species complexes defined below and Grosmannia (Lineage II) defined by G. penicillata including two species complexes defined above. A third complex could emerge as new species are described in Grosmannia.
The last formal morphological description of Leptographium was provided by Jacobs & Wingfield (2001) for asexual fungi forming mononematous conidiophores and branched conidiogenous apparatuses. In view of the One Fungus-One Name principles, we have emended the description of Leptographium to accommodate the morphology of the sexual morphs, treated previously in Grosmannia, Ophiostoma, Ceratocystis or Europhium (Upadhyay 1981, Jacobs & Wingfield 2001, De Beer & Wingfield. 2013). Morphology of synnematous asexual morphs previously excluded from Leptographium and treated variously as species of Graphium, Pesotum, Phialographium or Graphiocladiella (Upadhyay & Kendrick 1974, Upadhyay 1981, Seifert & Okada 1993, Okada et al. 1998) have also been incorporated in the revised circumscription of the genus.
Species complexes:
The L. clavigerum complex
The L. clavigerum complex was first described by Massoumi Alamouti et al. (2011) as the G. clavigera complex. Grosmannia clavigera is also the type species of Graphiocladiella, a genus originally described for the asexual morphs of ‘Ceratocystis’ species forming mononematous and synnematous conidiophores (Upadhyay 1981, Harrington 1988). Harrington (1988) proceeded to reduce Graphiocladiella to synonymy with Graphium (in which he also included species producing leptographium-like conidiophores). Species in the G. clavigera complex produce mostly synnematous aggregates of leptographium-like conidiophores, cleistothecal ascomata and reniform ascospores with hat-shaped sheaths (Linnakoski et al. 2012, De Beer & Wingfield 2013).
The first whole genome sequence produced for species in the Ophiostomatales was that for G. clavigera. This work was justified by the close association of G. clavigera with the mountain pine beetle (Dendroctonus ponderosae), a native but invasive pest causing devastating losses in the Northwestern USA and Canada (Kim et al. 2004, Massoumi Alamouti et al. 2011, De Beer & Wingfield 2013). Genome data were then used to show that what was previously thought to be a highly variable population of G. clavigera, actually represents a complex including multiple species (Massoumi Alamouti et al. 2011). Later studies contributed additional species to this new species complex (Six et al. 2011, De Beer & Wingfield 2013). In the present study, the G. clavigera complex formed a well-supported monophyletic lineage, and currently includes eight well-defined species, as well as one undescribed species (Massoumi Alamouti et al. 2011).
The L. galeiforme complex
The L. galeiforme complex was first described as the O. galeiformis complex when Zhou et al. (2004b) designated an epitype for the species (Bakshi 1951). Kim et al. (2005b) showed that G. radiaticola was part of the complex, and that this species was conspecific with the asexual species, Hyalopesotum pini. The latter species name was thus used for the asexual morph of G. radiaticola (Kim et al. 2005b, De Beer & Wingfield 2013). However, when the One Fungus–One Name principles are applied, the oldest epithet ‘pini’ (Hutchison & Reid 1988b) should take precedence over ‘radiaticola’ (Kim et al. 2005b). We have consequently provided a new combination, L. pini, to represent this species. It was previously believed that G. radiaticola and G. galeiformis were the same species (Thwaites et al. 2005). However, data from the studies of Kim et al. (2005b) and Linnakoski et al. (2012) showed that these are distinct species, and that in addition to these, the complex includes two as yet undescribed cryptic species. Chang et al. (2017) reported L. pini (as G. radiaticola) from China, as well as another possible new species in the complex.
All species in the L. galeiforme complex produce synnemata that appear to be loose aggregates of leptographium-like conidiophores (Zhou et al. 2004b, Linnakoski et al. 2012, De Beer & Wingfield 2013). The distinction between the two species is also supported by the ecology of these fungi, with L. galeiforme being associated with conifer-infesting bark beetles in Europe, and L. pini (as G. radiaticola) associated with pine-infesting bark beetles with a wider distribution (South Africa, Chile, Europe, China, Korea, and New Zealand; Linnakoski et al. 2012).
The L. lundbergii complex
The L. lundbergii complex was first defined by Jacobs et al. (2005). In a study emerging from a survey of ophiostomatoid fungi in Finland and Russia, Linnakoski et al. (2012) expanded the complex to include additional species that group close to L. lundbergii. Although these species always grouped close together, they never formed a well-supported monophyletic lineage, neither in the analyses of Linnakoski et al. (2012), various subsequent analyses (De Beer & Wingfield 2013, Jankowiak et al. 2017, 2018, Chang et al. 2017, Pan et al. 2020) nor those in the present study. However, for the sake of convenience, we have chosen to treat these species as a group. All the species produce relatively short conidiophores that have conidia with truncate bases, and the two sexually reproducing species produce cucullate ascospores (Linnakoski et al. 2012, De Beer & Wingfield 2013). Most of the species in this complex have been described from conifers in Asia and they have also been found in Europe, Canada, South Africa and New Zealand as the causal agents of sapstain in lumber (Linnakoski et al. 2012).
The L. olivaceum complex
The L. olivaceum complex accommodates species that are characterised by brownish synnematous asexual morphs and cucullate ascospores produced in ascomata with almost cylindrical necks and prominent ostiolar hyphae (Yin et al. 2019). The complex was first demarcated in a phylogeny by Six et al. (2011), more comprehensively defined by Linnakoski et al. (2012), and subsequently recognised by De Beer & Wingfield (2013) and Jankowiak et al. (2017, 2018). The complex includes L. sagmatosporum, the type species of the genus Phialographium (Upadhyay & Kendrick 1974), and its inclusion in the complex renders that name a synonym of Leptographium (De Beer & Wingfield 2013).
Most recently, Yin et al. (2019) added six new species to the complex. All species in this complex are associated with various bark beetles, mostly infesting spruce and pine trees in North America and Eurasia (Linnakoski et al. 2012, Jankowiak et al. 2017, 2018, Yin et al. 2019). The only exception is L. francke-grosmanniae that groups peripheral to the complex, has a morphology quite distinct from other species and was isolated from a bark beetle gallery on oak trees in Europe (Davidson 1971, Mouton et al. 1992, Yin et al. 2019). It is possible that this species represents a distinct species complex that cannot be distinguished at the present time.
The L. piceiperdum complex
Linnakoski et al. (2012) first defined this species complex when they showed that isolates described as G. piceiperda separated into five distinct species, two of which were unnamed. Ando et al. (2016) focused on isolates from different sources in Japan, revealing several additional cryptic species. Both Linnakoski et al. (2012) and Ando et al. (2016) concluded that epitypification is needed for L. piceiperdum and L. europhioides in order to confirm the placement of these species, because sequences for them were derived from herbarium type material. All members of this complex form typical leptographium-like asexual morphs, have cucullate ascospores (De Beer & Wingfield 2013, Ando et al. 2016), and have been found in association with conifer infesting bark beetles (mostly Ips species) in Europe, North America, Russia and Japan (Linnakoski et al. 2012, Ando et al. 2016).
The L. procerum complex
The L. procerum complex was first defined by De Beer & Wingfield (2013), and studied by Yin et al. (2015). These authors used seven gene regions to evaluate the species boundaries in the complex, and showed that it accommodates nine species. No sexual morph has been observed for any species in this complex, and asexual morphs can be described as leptographium-like (Jacobs et al. 2000b, 2006, Jacobs & Wingfield 2001, Lu et al. 2008, Paciura et al. 2010, De Beer & Wingfield 2013, Yin et al. 2015). Species in this complex are associated with conifer-infesting bark beetles in North America and Eurasia, with most described from East Asia. The exception is L. profanum that occurs on hardwood trees in the USA (Linnakoski et al. 2012, Yin et al. 2015). Most species in this complex are not widely distributed, with single species being found only in single countries. However, L. procerum has been isolated from Pinus spp. in China, Russia and the USA (Yin et al. 2015).
The L. procerum complex has received considerable attention by researchers. This is due to the fact that L. procerum, the type species of the complex has been associated with a root and root-collar disease of Pinus spp. in the USA known as white pine root decline (Kendrick 1962, Dochinger 1967, Wingfield 1983, Wingfield et aI. 1994). Furthermore, L. procerum is associated with the red turpentine beetle (Dendroctonus valens), which is native to North America and possibly western Europe and was introduced into China resulting in devastating losses of Chinese native pines (Yan et al. 2005, Lu et al. 2008, Taerum et al. 2012, 2013, 2017, Zhou et al. 2013, Sun et al. 2013, Yin et al. 2015).
The L. serpens complex
The L. serpens species complex was first recognised based on phylogenetic analyses by Six et al. (2011). In a focused study to resolve typification issues in the complex, Duong et al. (2012) showed that L. serpens, previously recognised as a single taxon represented five distinct species. All of these species produce typical leptographium-like conidiophores, with characteristic serpentine hyphae on agar, while ascospores produced by two of the species are sheathed (Wingfield & Marasas 1980, Jacobs & Wingfield 2001, Duong et al. 2012). Interestingly, most of the members of the complex have relatively narrow distributions, most being known from only a single country. For example, L. serpens occurs in Italy, L. gibbsii in the UK, L. yamaoake in Japan and L. castellanum in Spain and the Dominican Republic. The exception here is L. alacre that is more widely distributed and associated with conifer-root infesting bark beetles in South Africa and Italy (Duong et al. 2012), although its occurrence in South Africa is certainly due to an accidental introduction likely from Europe. Marincowitz et al. (2017) described a morphologically similar species, L. rhodanense from Switzerland that groups between the L. serpens and the L. wageneri complexes.
The L. wageneri complex
Six et al. (2011) first defined the L. wageneri species complex based on phylogenetic analyses. All species in this complex form typical leptographium-like asexual morphs. However, the sexual morph has been observed only for L. wageneri var. ponderosae, and this was described as G. wageneri (Goheen & Cobb 1978, Zipfel et al. 2006). This fungus produces allantoid ascospores, but unlike other Leptographium ascospores, no sheath was reported (Goheen & Cobb 1978, Jacobs & Wingfield 2001). Most of the species in the complex are associated with conifer-root infesting beetles in the USA (Cobb et al. 1974, 1984, Hansen et al. 1988, Witcosky & Hansen 1985, Witcosky et al. 1986). The only exception is L. reconditum, which was isolated from the roots of Triticum species in South Africa (Jooste 1978).
The L. wageneri complex includes three host-specific varieties that are important pathogens of conifers restricted to the western USA and causing black stain root disease (Cobb 1988, Hessburg & Hansen 2000). These three formally described varieties can be distinguished from each other based on their morphological characteristics and the host tree species that they infect (Harrington & Cobb 1984, 1987, Jacobs & Wingfield 2001). Leptographium wageneri var. pseudotsugae occurs on Douglas fir (Pseudotsuga menziesii), L. wageneri var. ponderosae on hard pines (Pinus ponderosa, P. jeffreyi and P. contorta), and L. wageneri var. wageneri on pinyon pines (P. monophylla and P. edulis). Phylogenetic analyses in this study support the lineage defined as the G. wageneri complex by De Beer & Wingfield (2013) and Six et al. (2011).
Groups A & B
Leptographium pineti grouped singly in Leptographium and this is consistent with phylogenies emerging from other studies (Paciura et al. 2010, Six et al. 2011, Duong et al. 2012, De Beer & Wingfield 2013). Chang et al. (2017) described a new species from China, L. ningerense (not included in our data), which grouped with L. pineti in their datasets.
Grosmannia cainii formed a lineage distinct from others in Leptographium in our analyses (Fig. 5) and this was also seen in the phylogenies of De Beer & Wingfield (2013).
New combinations:
1) Leptographium cainii (Olchow. & J. Reid) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840389.
Basionym: Ceratocystis cainii Olchow. & J. Reid, Canad. J. Bot. 52: 1697. 1974. MycoBank MB 310486.
Synonyms: Ophiostoma cainii (Olchow. & J. Reid) T.C. Harr., Mycotaxon 28: 41. 1987. MycoBank MB 128912.
Grosmannia cainii (Olchow. & J. Reid) Zipfel, Z.W de Beer & M.J. Wingf., Stud. Mycol. 55: 89. 2006. MycoBank MB 500812.
Descriptions: Upadhyay (1981: 39, figs 43–47), Seifert & Okada (1993: 32, fig. 3D).
Phylogenetic data: Hausner et al. (2000), Masuya et al. (2004), Kim et al. (2005b), Six et al. (2011), Duong et al. (2012), De Beer & Wingfield (2013), Jankowiak et al. (2017, 2018), Liu et al. (2017), De Errasti et al. (2018), Chang et al. (2019).
Notes: Leptographium cainii groups alone and distinct from other species complexes in Leptographium in our data (Fig. 5). This is consistent with the data presented by De Beer & Wingfield (2013).
2) Leptographium europhioides (E.F. Wright & Cain) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840392.
Basionym: Ceratocystis europhioides E.F. Wright & Cain, Canad. J. Bot. 39: 1222. 1961. MycoBank MB 327627.
Synonyms: Ophiostoma europhioides (E.F. Wright & Cain) H. Solheim, Nordic J. Bot. 6: 203. 1986. MycoBank MB 102979.
Grosmannia europhioides (E.F. Wright & Cain) Zipfel, Z.W. de Beer & M.J. Wingf., Stud. Mycol. 55: 90. 2006. MycoBank MB 500818.
Ceratocystis shikotsuensis Aoshima, Ph. D. thesis, University of Tokyo: 10. 1965. nom. inval., Arts 29.1, 39.1 or 39.2.
Descriptions: Davidson et al. (1967: 929–930), Griffin (1968: 709, 713), Olchowecki & Reid (1974: 1699, pI. XIII, figs 259–261), De Hoog & Scheffer (1984: 295, fig. 2), Yamaoka et al. (1997: 1221–1222), Jacobs et al. (1998: 290–291), Jacobs et al. (2000a: 239).
Phylogenetic data: Hausner et al. (1993b, 2000), Okada et al. (1998), Schroeder et al. (2001), Masuya et al. (2004), Greif et al. (2006), Mullineux & Hausner (2009), Matsuda et al. (2010), Paciura et al. (2010), Six et al. (2011), Linnakoski et al. (2012), De Beer & Wingfield (2013), Musvuugwa et al. (2015), Ando et al. (2016), Yamaoka (2017), De Errasti et al. (2018).
Notes: Leptographium europhiodes groups in the L. piceiperdum complex (Fig. 5). De Beer & Wingfield (2013) noted that species such as G. europhioides, previously treated as synonyms of G. piceiperda based on morphology (Jacobs et al. 2000a, Jacobs & Wingfield 2001), were possibly distinct species (Linnakoski et al. 2012). Ando et al. (2016) investigated the phylogenetic relationships of Japanese isolates assigned to the L. piceiperdum complex and recognised 13 lineages within the complex. These included lineages representing G. aenigmatica, G. laricis, G. piceiperda group D and eight representing distinct, but undescribed species (Ando et al. 2016, Yamaoka 2017). Ando et al. (2016) concluded that although L. piceiperdum (as G. piceiperda) and L. europhioides (as G. europhioides) are valid species, epitypification is needed for both species to resolve the identity of the remaining undescribed taxa.
3) Leptographium galeiforme (B.K. Bakshi) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840393.
Basionym: Ceratocystis galeiformis Bakshi, Mycol. Pap. 35: 13. 1951. MycoBank MB 294208.
Synonyms: Ophiostoma galeiforme (B.K. Bakshi) Math.-Käärik, Meddeland. Statens Skogs-Forstningsinst. 43: 47. 1953. MycoBank MB 302075 (as ‘galeiformis’).
Grosmannia galeiformis (B.K. Bakshi) Zipfel, Z.W. de Beer & M.J. Wingf., Stud. Mycol. 55: 90. 2006. MycoBank MB 500820.
Descriptions: Mathiesen-Käärik (1953: 47–50), Hunt (1956: 33), Wingfield (1993: 48, fig. 8), Zhou et al. (2004b: 1309–1311, fig. 2).
Phylogenetic data: Hausner et al. (2000), Zhou et al. (2004b), Kim et al. (2005b, 2011), Thwaites et al. (2005), Greif et al. (2006), Zipfel et al. (2006), Lu et al. (2009), Mullineux & Hausner (2009), Harrington et al. (2010), Matsuda et al. (2010), Paciura et al. (2010), Six et al. (2011), Duong et al. (2012), Linnakoski et al. (2012), De Beer & Wingfield (2013), Taerum et al. (2013), Masuya et al. (2013), Huang & Chen (2014), Wang et al. (2014), Musvuugwa et al. (2015), De Beer et al. (2016a, b), Chang et al. (2017, 2019), Wingfield et al. (2017), Jankowiak et al. (2017), Liu et al. (2017), De Errasti et al. (2018).
Notes: This species grouped with L. radiaticola and undescribed species (Linnakoski et al. 2012, De Beer & Wingfield 2013) to form the L. galeiforme species complex (Fig. 5). In the phylogenies of Chang et al. (2019), L. koraiensis grouped within this species complex. The latter species was not included in our analyses.
4) Leptographium pseudoeurophioides (Olchow. & J. Reid) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840413.
Basionym: Ceratocystis pseudoeurophioides Olchow. & J. Reid, Canad. J. Bot. 52: 1700. 1974. MycoBank MB 310514.
Synonyms: Ophiostoma pseudoeurophioides (Olchow. & J. Reid) Hausner et al., Canad. J. Bot. 71: 1264. 1993. MycoBank MB 362667.
Grosmannia pseudoeurophioides (Olchow. & J. Reid) Zipfel et al., Stud. Mycol. 55: 91. 2006. MycoBank MB 500826.
Descriptions: Olchowecki & Reid (1974: 1700, figs 219–229).
Phylogenetic data: Hausner et al. (1993b, 2000), Masuya et al. (2004), Mullineux & Hausner (2009), Mullineux et al. (2011), De Beer & Wingfield (2013), Musvuugwa et al. (2015), De Errasti et al. (2018).
Notes: Leptographium pseudoeurophioides was assigned to the G. penicillata complex (Hausner et al. 1993b, 2000). However, based on a short LSU sequence produced by Hausner et al. (1993b, 2000) and ascospore morphology, De Beer & Wingfield (2013) suggested that L. pseudoeurophioides should form part of the L. piceiperdum complex, and that additional collections and fresh material would be required to confirm its placement. For the present, we treat this species in Leptographium.
5) Leptographium radiaticola (J.J. Kim et al.) M. Procter & Z.W. de Beer comb. nov. [MycoBank MB 840396]
Basionym: Ophiostoma radiaticola J.J. Kim et al., Mycotaxon 91: 486. 2005. MycoBank MB 500832.
Synonyms: Grosmannia radiaticola (J.J. Kim et al.) Z.W. de Beer & M.J. Wingf., Stud. Mycol. 55: 91. 2006. MycoBank MB 500827.
Hyalopesotum pini L.J. Hutchison & J. Reid, New Zealand J. Bot. 26: 90. 1988. MycoBank MB 135442.
= Pesotum pini (L.J. Hutchison & J. Reid) G. Okada & Seifert, Canad. J. Bot. 76: 1504. 1998. MycoBank MB 446360.
Descriptions: Hutchison & Reid (1988b: 90–91, figs 32–35, of Hy. pini), Kim et al. (2005b: 486–489, figs 1–14).
Phylogenetic data: Masuya et al. (2004), Kim et al. (2005a, b), Thwaites et al. (2005), Zipfel et al. (2006), Lu et al. (2009), Mullineux & Hausner (2009), Paciura et al. (2010), Six et al. (2011), Duong et al. (2012), Linnakoski et al. (2012), De Beer & Wingfield (2013), Taerum et al. (2013), Huang & Chen (2014), Wang et al. (2014), Romón et al. (2014a), Musvuuwga et al. (2015), Chang et al. (2017, 2019), Liu et al. (2017), De Errasti et al. (2018).
Notes: This species grouped in the L. galeiforme species complex (Fig. 5; Linnakoski et al. 2012, De Beer & Wingfield 2013). Pesotum pini has been recognised as the asexual morph of L. radiaticola (Kim et al. 2005b). Following the One Fungus-One Name principles, the oldest name, irrespective of morph, should take precedence in the selection of the basionym. In this case, Hyalopesotum pini would serve that purpose. However, to avoid confusion with the name O. pini, currently treated as a synonym of O. minus, but possibly a distinct taxon (De Beer & Wingfield, 2013), we have designated O. radiaticola as basionym.
Other species: Listed in Table 1.
Leptographium and Grosmannia incertae sedis (Lineages III & VI)
Leptographium piriforme (Lineage III)
De Beer & Wingfield (2013) applied a relatively wide taxonomic concept for Leptographium s.l. and their phylogenies placed L. piriforme in a distinct lineage with G. crassivaginata. The data in the present study did not include G. crassivaginata. Greif et al. (2006) noted some morphological similarities between L. piriforme and G. crassivaginata when they described L. piriforme. Consequently, G. crassivaginata may form part of Lineage III if it were included in analyses.
Leptographium piriforme produces curved conidia and pear-shaped cells, distinguishing it from other Leptographium species (Greif et al. 2006, Jankowiak & Kolařik 2010). In the concatenated dataset utilised in the present study, L. piriforme grouped basal to Lineage I and II (Fig. 5). Although the phylogeny presented by Greif et al. (2006) was much smaller than that presented here, L. piriforme also grouped outside of the lineage containing Leptographium and Grosmannia. For the present, we have retained L. piriforme in Leptographium although its taxonomic placement clearly deserves further consideration.
Leptographium verrucosum (Lineage VI)
Gebhardt et al. (2002) was the first to describe L. verrucosum (as O. verrucosum) and their description was based solely on morphology. De Beer & Wingfield (2013) transferred O. verrucosum to Leptographium based on LSU sequences, and their recommendation was that new species grouping with either Leptographium or Grosmannia should be described in Leptographium. In our LSU dataset (Fig. S1), L. verrucosum grouped basal to Lineage I (Leptographium s.s.), but this grouping was not seen in the other gene trees. For the present and until additional material is available for further analyses, we have chosen to retain it in Leptographium.
Masuyamyces (Lineage XVII)
Masuyamyces Z.W. de Beer & M. Procter, gen. nov. MycoBank MB 840414. Fig. 12A, B.
Fig. 12.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Masuyamyces. C, D. Ophiostoma clavatum complex. E, F. Ophiostoma ips complex. G, H. Ophiostoma minus complex. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Etymology: Named for Dr Hayato Masuya, a Japanese mycologist who has described 17 species of ophiostomatoid fungi, including the type species of this novel genus.
Sexual morph: Ascomatal bases black, subglobose to globose; necks black, nearly cylindrical, curved or straight. Ostiolar hyphae absent or present. Asci evanescent, clavate to sub-globose when young. Ascospores hyaline, 1-celled, oblong or cylindrical in front view, allantoid in side view, globose in end view, enclosed in uniform hyaline sheath.
Asexual morph: Conidiophores macronematous, mononematous to synnematous. Hyalorhinocladiella-like; conidiophores simple or branched, hyaline; conidiogenous cells annellidic, cylindrical; conidia hyaline, obovoid to globose, oblong to ellipsoidal, 1-celled. Pesotum-like; conidiophores loosely compacted, stipes hyaline to pale brown; conidiogenous cells hyaline annellidic; conidia hyaline, oblong to ellipsoidal.
Type species: Masuyamyces botuliformis (Masuya) Z.W. de Beer & M. Procter, comb. nov. MycoBank MB 840415.
Basionym: Ophiostoma botuliforme Masuya, Mycoscience 44: 304. 2003. MycoBank MB 489292.
Description: Masuya et al. (2003: 304, figs 11–21).
Phylogenetic data: Fig. 5.
Notes on the type species: No DNA sequence data were previously available for this species (De Beer & Wingfield 2013). Although M. botuliformis resembles O. allantosporum morphologically (Masuya et al. 2003), it did not group with the latter species, which groups in Ophiostoma (Fig. 5). Recently, sequences of M. botuliformis (as O. botuliforme) have been deposited in GenBank, which are identical to our ITS and LSU sequences. See note under M. saponiodorus.
Notes on the genus: In our phylogenies, Lineage XVII consistently formed a monophyletic group, distinct from Ophiostoma. This group includes O. ambrosium, O. botuliforme, O. pallidulum (not included in our analyses) and O. saponiodorum. All described from conifers and all having reniform ascospores (Fig. 3F; Type A as categorised by De Beer & Wingfield 2013). Although O. pallidulum was not included in our analyses, it grouped with O. saponiodorum in the phylogenies of Linnakoski et al. (2010) and De Beer & Wingfield (2013).
The isolate of O. ambrosium included in our analyses is not the ex-type culture but was designated as an “isolate from type collection” by Hausner et al. (1993b). Our LSU sequence is almost identical to that of Hausner et al. (1993b), although theirs is very short. However, the morphology of this isolate does not match that of the type of O. ambrosium, and although this isolate is from the type collection, it may represent a different species.
Our isolate of O. saponiodorum was also not the ex-type isolate for the species, but our ITS sequence is identical to the one produced by Linnakoski et al. (2010). The latter species grouped with isolate CMW 30883 and was used in the cross to produce the holotype of the species. Therefore, although two species in this clade are not represented by ex-type isolates, we have sufficient confidence in the sequences to justify describing this lineage as a new genus.
Masuyamyces ambrosius was described from Betula albae infested by the beetle Trypodendron domesticum in Great Britain (Bakshi 1950). Masuyamyces botuliformis was isolated from Pinus densiflora in Japan (Masuya et al. 2003), M. pallidulus was from P. sylvestris infested by Hylastes brunneus in Finland (Linnakoski et al. 2010), and M. saponiodorus was from Picea abies infested with Ips typographus and Pityogenes chalcographus in Russia as well as Finland (Linnakoski et al. 2010). The asexual morph of M. botuliformis was described as pesotum-like (Masuya et al. 2003), that of M. ambrosium as raffaelea-like (De Beer & Wingfield 2013), that of M. pallidulus as hyalorhinocladiella-like, and that of M. saponiodorus as pesotum-like with a hyalorhinocladiella-like synasexual morph (Linnakoski et al. 2010).
Other new combinations:
1) Masuyamyces acarorum (R. Chang & Z.W. de Beer) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840416.
Basionym: Ophiostoma acarorum R. Chang & Z.W. de Beer, MycoKeys 28: 40. 2017. MycoBank MB 823693.
Description: Chang et al. (2017: 40–41, fig. 7).
Phylogenetic data: Chang et al. (2017, 2019).
Notes: Sexual morph unknown. This species resides in Masuyamyces and grouped with M. saponiodorus (as O. saponiodorum) in the phylogenies of Chang et al. (2017, 2019). See note under M. saponiodorus.
2) Masuyamyces ambrosius (B.K. Bakshi) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840526.
Basionym: Ceratocystis ambrosia B.K. Bakshi, Trans. Brit. Myc. Soc. 33: 116. 1950. MycoBank MB 294192.
Synonym: Ophiostoma ambrosium (B.K. Bakshi) Georg Hausner, J. Reid & Klassen, Canad. J. Bot. 71: 1264. 1993. MycoBank MB 362662.
Description: Bakshi (1950: 116–118, fig. 2)
Notes: Sexual morph unknown.This species grouped with M. saponiodorus and M. botuliformis (Fig. 5).
3) Masuyamyces jilinensis (R. Chang et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840417.
Basionym: Ophiostoma jilinense R. Chang et al., MycoKeys 28: 63. 2019. MycoBank MB 825086.
Description: Chang et al. (2019: 63–65, fig. 16).
Phylogenetic data: Chang et al. (2019).
Notes: Sexual morph unknown. This species grouped with M. saponiodorus (as O. saponiodorum) in the phylogeny of Chang et al. (2019), and is consequently included in the the genus. See note under M. saponiodorus.
4) Masuyamyces lotiformis (Z. Wang & Q. Lu) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840418.
Basionym: Ophiostoma lotiforme Z. Wang & Q. Lu, IMA Fungus 11: 17. 2020. MycoBank MB 830612.
Description: Wang et al. (2020: 17–18, fig. 14)
Phylogenetic data: Wang et al. (2020).
Notes: Sexual morph unknown. This species grouped with M. saponiodorus (as O. saponiodorum) in the phylogeny of Wang et al. (2020), and is consequently included in the genus. See note under M. saponiodorus.
5) Masuyamyces massonianae M. Procter & Z.W. de Beer, sp. nov. MycoBank MB 840419.
Synonym: Ophiostoma massonianae H.M. Wang & Q. Lu, MycoKeys 39: 15. 2018. nom. inval., Art. 40.7. MycoBank MB 827856.
Description: Wang et al. (2018: 15–17, fig. 4).
Holotype: PREM 63310.
Phylogenetic data: Wang et al. (2018)
Notes: Sexual morph unknown. This species grouped with M. saponiodorus (as O. saponiodorum) in the phylogenies of Wang et al. (2018), and is thus included in the genus. See note under M. saponiodorus.
6) Masuyamyces pallidulus (Linnak. et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840420.
Basionym: Ophiostoma pallidulum Linnak. et al., Persoonia 25: 86. 2010. MycoBank MB 518884.
Description: Linnakoski et al. (2010: 86, 88, fig. 9).
Phylogenetic data: Linnakoski et al. (2010, 2016), De Beer & Wingfield (2013), De Beer et al. (2016a, b), Chang et al. (2017, 2019), Wang et al. (2018).
Notes: Sexual morph unknown. Although not included in our analyses, this species was shown to consistently group with M. saponiodorus (as O. saponiodorum) and is thus treated in Masuyamyces (Linnakoski et al. 2010, 2016, De Beer & Wingfield 2013, Chang et al. 2017, 2019, Wang et al. 2018). See note under M. saponiodorus.
7) Masuyamyces saponiodorus (Linnak. et al.) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 840421.
Basionym: Ophiostoma saponiodorum Linnak. et al., Persoonia 25: 88. 2010. MycoBank MB 518885.
Description: Linnakoski et al. (2010: 88, fig. 10).
Phylogenetic data: Linnakoski et al. (2010, 2016), Six et al. (2011), De Beer & Wingfield (2013), Romón et al. (2014b), Chang et al. (2017, 2019), Wang et al. (2018).
Notes: Masuyamyces saponiodorus consistently grouped separately from what we have defined as Ophiostoma in our phylogenies (Fig. 5, S1–S4), and with the other species listed above in previously published phylogenies (Linnakoski et al. 2010, 2016, De Beer & Wingfield 2013, Chang et al. 2017, 2019, Wang et al. 2018).
Ophiostoma (Lineage XV)
Ophiostoma Syd. & P. Syd., Ann. Mycol. 17: 43. 1919. MycoBank MB 3614, emend. Z.W. de Beer & M. Procter. Figs 12C–13F.
Fig. 13.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Ophiostoma piceae complex. C, D. Ophiostoma pluriannulatum complex. E, F. Ophiostoma ulmi complex. G, H. Raffaelea. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Synonyms: Linostoma Höhn., Ann. Mycol. 16: 91. 1918. MycoBank MB 2885. nom. illegit. Art. 53.1, De Beer et al. 2013a. [Type species Linostoma piliferum (Fr.) Höhn.].
Pesotum J.L. Crane & Schokn., Amer. J. Bot. 60: 347. 1973. MycoBank MB 9270. [Type species Pesotum ulmi (M.B. Schwarz) J.L. Crane & Schokn.].
Hyalopesotum H.P. Upadhyay & W.B. Kendr., Mycologia 67: 801. 1975. MycoBank MB 8579. (Type species H. introcitrinum H.P. Upadhyay & W.B. Kendr.).
Pachnodium H.P. Upadhyay & W.B. Kendr., Mycologia 67: 802. 1975. Asexual synonym: MycoBank MB 9189 (Type species P. canum H.P. Upadhyay & W.B. Kendr.).
Etymology: Ophio- from the Greek for ‘snake-like’ and ‘stoma’ meaning mouth, referring to the long, tubular necks of the ascomata.
Sexual morph: Ascomatal bases black, subglobose to globose; necks black, cylindrical, straight or slightly curved. Ostiolar hyphae absent or present, divergent. Asci evanescent, clavate. Ascospores hyaline, 1-celled, enclosed in sheath, allantoid, reniform, cylindrical to ossiform in side view, ellipsoid in face view, globose in end view.
Asexual morph: Conidiophores macronematous, micronematous, mononematous, synnematous. Hyalorhinocladiella-like; conidiophores simple or branched; conidia hyaline, 1-celled, oblong to ellipsoid. Leptographium-like; conidiophores upright or prone, bases simple or rhizoid-like; conidiogenous cells hyaline, subcylindrical; conidia hyaline, 1-celled, oblong to obovoid with truncate base. Pesotum-like; conidiophores single or in groups, black becoming paler towards apex; stipes extending beyond conidiogenous cells, becoming seta-like structures (O. cupulatum); conidia hyaline, mostly 1-celled, oblong to ellipsoid, round apex, tapering towards base, clavate to obovoid; conidia elongated cylindrical or clavate with tapering towards base, 1–3-celled (O. nikkonse), giving rise to secondary conidia (O. nikkoense). Sporothrix-like; conidiogenous cells proliferating, sympodial, denticulate; conidia hyaline, 1-celled, oblong to ellipsoid, elongate or broadly cylindrical, round apex, tapering towards base, clavate or obovoid, obpyriform, giving rise to secondary spores (O. cupulatum, O. flexuosum).
Type species: Ophiostoma piliferum (Fr.) Syd. & P. Syd., Ann. Mycol. 17: 43. 1919. MycoBank MB 431882.
Other species: Listed in Table 1.
Notes on the genus: The definition of Ophiostoma emerging from this study relates to what was previously treated as Ophiostoma s.l. by De Beer & Wingfield (2013). In this case it excludes the S. schenckii-O. stenoceras complex (now Sporothrix) and the O. tenellum complex. Species in the newly defined Ophiostoma sensu stricto typically produce Type A or B ascospores (Fig. 3F, G; as categorised by De Beer & Wingfield 2013) and a variety of asexual morphs including those that are sporothrix-like, pesotum-like, hyalorhinocladiella-like; some having more than one morph and commonly a continuum between these morphs (De Beer & Wingfield 2013).
Species complexes:
The O. clavatum complex
The O. clavatum complex was defined by Linnakoski et al. (2016). Species in this complex generally have rectangular to cylindrical shaped ascospores (Fig. 3F; Type A as categorised by De Beer & Wingfield 2013), spirally coiled ostiolar hyphae that are brown in colour and they produce pesotum-like asexual morphs (Mathiesen-Käärik 1960, Linnakoski et al. 2016). These fungi are found in Eurasia and are mostly associated with conifer-infesting bark beetles residing in the genus Ips and some species cause blue stain in the sapwood of trees (Linnakoski et al. 2016).
Ophiostoma ainoae and O. tapionis grouped peripherally to other species in the O. clavatum complex in the datasets of Linnakoski et al. (2016). Ophiostoma ainoae has ascospores similar in shape to species in the O. ips complex, but groups within the O. clavatum complex (Linnakoski et al. 2016). Ophiostoma ainoae also has a hyphal morphology similar to that of O. clavatum but it grouped peripheral to that complex (Linnakoski et al. 2016). Due to its morphological similarity with other species in the complex, O. ainoae is best treated as part of the O. clavatum species complex. This grouping was also supported in our analyses (Fig. 5).
The O. ips complex
The O. ips complex was first defined by De Beer & Wingfield (2013). Species in the complex all have cylindrical ascospores in a pillow-shaped sheath (Fig. 3H; Type E as categorised by De Beer & Wingfield 2013) and this distinguishes them from other species in Ophiostoma. These species form pesotum- to hyalorhinocladiella-like asexual morphs (De Beer & Wingfield 2013). Members of this complex are associated with bark beetles in the genera Ips and Dendroctonus, infesting conifers mostly in North America and areas where pine trees have been introduced (Upadhyay 1981, Zhou et al. 2004a, Linnakoski et al. 2010).
The ex-type isolate of the asexual species Hyalorhinocladiella ips (CMW 14175) grouped in our analyses within the O. ips complex, similar to the results of De Beer & Wingfield (2013). Hyalorhinocladiella ips was originally described as Tuberculariella ips (Leach et al. 1934), later treated in Ambrosiella (Batra 1967) and most recently in Hyalorhinocladiella (Harrington et al. 2010). This fungus has been suggested to represent the asexual morph of O. montium, because the two species share morphological similarities and there are minimal differences in their sequence data (Massoumi Alamouti et al. 2009, De Beer & Wingfield 2013, De Beer et al. 2013b). In our analyses (Fig. 5) O. montium and “H. ips” grouped very close to each other. De Beer et al. (2013b) formally listed H. ips as synonym of O. montium (Rumbold 1941), even though H. ips is the older of the two names (Leach et al. 1934) and should thus be designated as basionym for the species. However, the epithet “ips” is already in use for O. ips, the name bearing species of this complex, that has Ceratostomella ips (Rumbold 1931) as basionym. To avoid the creation of a later homonym by using H. ips as basionym, we have chosen to select Ceratostomella montium (Rumbold 1941) as basionym for the species and for which O. montium (Von Arx 1952) is then the appropriate current name. The O. ips complex formed a well-supported lineage that might be excluded from Ophiostoma s.s. in future to be treated as a distinct genus. However, for the present and in the absence of more robust datasets, we have retained species in this complex in Ophiostoma s.s.
The O. minus complex
The O. minus complex was well supported in our phylogenies even though this was not true in the phylogenies of De Beer & Wingfield (2013). The O. minus complex was first defined by Jacobs & Kirisits (2003). Species in this complex produce hyalorhinocladiella-like asexual morphs (Fig. 3A; as categorised by De Beer & Wingfield 2013) and have been isolated from conifers in North America and Europe. Ophiostoma minus is a well-known causal agent of blue stain in conifers. It is associated with the southern pine beetle (Dendroctonus frontalis), a native but invasive pest in the USA (Jacobs & Kirisits 2003, Gorton et al. 2004).
It must be noted that our phylogenies included isolates of only the North American form of O. minus and O. pseudominus, both available in the CMW collection. This was due to the fact that there were no available cultures of other species included in the complex by previous authors (Jacobs & Kirisits 2003, Gorton et al. 2004, Linnakoski et al. 2010, De Beer & Wingfield 2013).
The O. piceae complex
Harrington et al. (2001) first defined the O. piceae complex, but the complex was not applied by De Beer & Wingfield (2013). Because the complex did not have any phylogenetic support in their trees, De Beer and Wingfield (2013) treated the species in the complex as part of Ophiostoma s.s. Yin et al. (2016) redefined the O. piceae complex, including the discripiton of three new species described from spruce-infesting bark beetles in China. The two sub-clades seen to be formed by the O. piceae complex (Harrington et al. 2001) are now known as the O. ulmi complex and the O. piceae complex. The O. ulmi complex includes what was previously referred to as the ‘hardwood clade’, and the newly redefined O. piceae complex includes what was previously referred to as the ‘conifer clade’ (Harrington et al. 2001, Grobbelaar et al. 2010, 2011, Kamgan Nkuekam et al. 2011, De Beer & Wingfield 2013, Yin et al. 2016). Species in this complex form ascomata with long necks of up to 500 μm and can form either sporothrix- or pesotum-like asexual morphs, or both, while some form hyalorhinocladiella-like asexual morphs (Fig. 3A, C, E; as categorised by De Beer & Wingfield 2013). Members of this complex have been isolated from conifers mostly in the Northern Hemisphere, and one species (O. cupulatum) was described from New Zealand (Yin et al. 2016). In our phylogenies, the O. piceae complex grouped distinct from the O. ulmi complex.
The O. pluriannulatum complex
The O. pluriannulatum complex was initially defined as the O. multiannulatum complex (Villarreal et al. 2005) and later renamed to the O. pluriannulatum complex (Kamgan Nkuekam et al. 2008, Zanzot et al. 2010, De Beer & Wingfield 2013). Ascospores of species in the complex are allantoid (Fig. 3F; Type A in De Beer & Wingfield 2013) (Hunt 1956, Kamgan Nkuekam et al. 2008, Zanzot et al. 2010, De Beer & Wingfield 2013). This is with the exception of O. carpenteri that has Type B (Fig. 3G) narrowly clavate ascospores (Hausner et al. 2003, De Beer & Wingfield 2013). Other than O. carpenteri, all members of the O. pluriannulatum complex have very long ascomatal necks relative to other Ophiostoma species (Zanzot et al. 2010, De Beer & Wingfield, 2013). Ophiostoma carpenteri has very short necks but does produce a sporothrix-like asexual morph as in all other members of the complex (Hausner et al. 2003, De Beer & Wingfield 2013). Species in this complex have been found in Eurasia, North America and Africa on conifers and hardwoods, and some associated with bark beetles in the genera Ips and Dendroctonus (Zhou et al. 2004a, Linnakoski et al. 2010, Pacuira et al. 2010). This complex may represent a discrete genus, but our analyses were not sufficiently robust to support such a distinction.
The O. ulmi complex
The O. ulmi complex was first known as the ‘hardwood clade’ in the O. piceae complex (Harrington et al. 2001, Linnakoski et al. 2010). It was later referred to as the O. quercus complex (Kamgan Nkuekam et al. 2011), until De Beer & Wingfield (2013) defined it as the O. ulmi species complex. Most of the members of the complex produce allantoid ascospores (Fig. 3F; Type A) that lack sheaths (Hunt 1956, Brasier 1991, De Beer & Wingfield, 2013). Species in this complex occur in Eurasia, Australia, Africa and North and South America and are associated with a variety of bark beetles (Upadhyay 1981, Brasier 1991, Brasier & Mehrotra 1995, Villarreal et al. 2005, Harrington et al. 2001, Kamgan Nkuekam et al. 2010, 2011, Linnakoski et al. 2008, 2009, Grobbelaar et al. 2010, Paciura et al. 2010). De Beer & Wingfield (2013) chose to have O. ulmi as the name bearing species for this complex, justified because it is the best-known species in this complex, as one of the causal agents of Dutch Elm Disease (DED). Ophiostoma novo-ulmi and O. himal-ulmi are also causal agents of DED (Brasier 1991, Harrington et al. 2001). DED is one of the most catastrophic tree disease pandemics in the Northern Hemisphere, leading to the death of large populations of Elm trees (Brasier 1979, 1991, Gibbs 1978). Another species in this complex, O. quercus, is associated with substantial economic losses due to sapstain in Quercus spp. (Harrington et al. 2001).
Ophiostoma bacillisporum, O. torulosum and O. undulatum have asexual morphs that are different in morphology to other members of the O. ulmi complex. Their asexual morphs are mycelial while other members of the complex form sporothrix- and pesotum-like asexual morphs. The ascospores of the species are, however, similar to those in the O. ulmi complex (De Beer & Wingfield, 2013). We have supported the decision of De Beer & Wingfield (2013) to include species residing in what was known as the O. quercus complex (Kamgan Nkuekam et al. 2011) in the O. ulmi complex. This decision is justified based on the fact that O. quercus groups within the O. ulmi complex. However, our analyses grouped O. torulosum outside the O. piceae complex (Fig. 5) and its position requires further consideration.
Group H
De Beer & Wingfield (2013) included O. triangulosporium in the O. ulmi complex but noted that its unique ascospore morphology (allantoid ascospores with triangular sheaths) suggested that its placement in the complex required further study, including inspection of the type material and a greater number of isolates. In our analyses (Fig. 5), O. triangulosporium grouped peripheral to other species in the O. ulmi complex.
Group I
Ophiostoma macrosporum and an undescribed Ophiostoma species (previously treated as a Hyalorhinocladiella sp.) formed a small lineage outside of any of the species complexes in Ophiostoma.
Group J
The type species of Ophiostoma, O. piliferum, grouped on its own within Ophiostoma. Although it is the type of the genus, it does not group in any species complex (De Beer & Wingfield 2013, Yin et al. 2016, De Beer et al. 2016a, b).
Group K
Ophiostoma tetropii grouped peripheral to the O. minus and O. piceae complexes (Fig. 5). Linnakoski et al. (2010) included O. tetropii in the O. minus complex. However, the ascospores of O. tetropii have a distinct morphology different to species in the O. minus complex (De Beer & Wingfield 2013). The phylogenies of De Beer & Wingfield (2013) also showed that O. tetropii grouping separately from the O. minus complex and it clearly requires further study.
Ophiostoma incertae sedis (Lineages XVIII, XX & XXIII)
Ophiostoma valdivianum (Lineage XVIII)
As mentioned above, O. valdivianum formed a lineage with S. fumea in the phylogenies of De Beer et al. (2016a), grouping basal to most of Sporothrix. However, in the phylogenies arising from the present study, these two species did not group together.
Ophiostoma angusticollis and O. denticulatum (Lineage XX)
Ophiostoma angusticollis and O. denticulatum consistently formed a lineage distinct from Ophiostoma. In the phylogenies of De Beer & Wingfield (2013), O. angusticollis formed a lineage with O. sejunctum (not included in the present study) close to the O. tenellum complex (Lineage XIII) but distinct from Ophiostoma. We have chosen not to describe this lineage as a new genus because we were not able to include the holotype of O. angusticollis.
Ophiostoma noisomeae (Lineage XXIII)
Ophiostoma noisomeae consistently grouped distinct from Ophiostoma in our datasets. In the phylogenies of De Beer et al. (2016a), O. noisomeae also grouped distinct from Ophiostoma.
Paleoambrosia (No sequence data available)
Paleoambrosia Poinar & F.E. Vega, Fungal Biol. 122: 1160. 2018. MycoBank MB 840457. nom. inval., Art. F.5.1.
Etymology: ‘From the Greek “palaios” = ancient, and the Greek “ambrosia” = immortal, referring to the fossilized ambrosial fungus discovered with a Platypodine beetle in ~100 million-year-old amber in Myanmar’ (Poinar & Vega 2018).
Sexual morph: Unknown.
Asexual morph: Conidiophores borne in sporodochia, simple, black. Conidia black, 1-celled, globose to obovoid, borne in slimy droplet terminally. Yeast-like cells present in mycangium.
Type species: Paleoambrosia entomophila Poinar & F.E. Vega, Fungal Biol. 122(12): 1160 (2018). MycoBank MB 840458. nom. inval., Art. F.5.1.
No other species known at present.
Notes: This genus was recently described from a specimen found in amber and was included in the current study for completeness, despite the fact that its placement in the Ophiostomatales (Poinar & Vega 2018) is open to debate. The fossil beetle associated with this fungus could have been misidentified and might not be a member of the Platipodinae. Likewise, whether the cavity shown on the beetle reflects a functional mycangium is unclear. The growth form of the fungus growing alongside the beetle is unusual for any beetle-related ophiostomatalean fungus, and mucilage-bound phialidic spores could be those of an asexual morph of an entomopathogenic or another fungus producing phialidica conidiophores.
Raffaelea (Lineage XII)
Raffaelea Arx & Hennebert, Mycopathol. Mycol. Appl. 25: 310. 1965. MycoBank MB 9685, emend. Z.W. de Beer & M. Procter. Fig 13G, H.
Etymology: Named for the Italian mycologist and plant pathologist, Prof. Raffaele Ciferri (1897–1964).
Sexual morph: Ascomatal bases black, subglobose to globose; necks black, straight or curved. Ostiolar hyphae present or absent. Asci evanescent. Ascospores hyaline, 1-celled or rarely 2-celled, cylindrical to oblong, enclosed in hyaline rectangular sheath.
Asexual morph: Colony confluent, mucilaginous. Conidiophores micronematous, macronematous, mononematous, solitary or aggregated in sporodochia, hyaline, simple or branched, monilioid. Conidiogenous cells flask-shaped, proliferating percurrently or sympodially, with denticles, inconspicuous scars or annellations. Conidia hyaline, ellipsoidal, obovoid, globose, pyriform, sometimes T- or Y-shaped, 1-celled, producing secondary cells through budding. Associated with ambrosia beetles.
Type species: R. ambrosiae Arx & Hennebert, Mycopathol. Mycol. Appl. 25: 310. 1965. MycoBank MB 338171.
Other species: Listed in Table 1.
Notes: Raffaelea was originally described to accommodate a number of fungal mutualists of wood-boring ambrosia beetles (Von Arx & Hennebert 1965). In the same paper, Von Arx & Hennebert (1965) validated a similar genus, Ambrosiella, with a mode of conidial development different to that in Raffaelea. However, confusion persisted regarding the generic placement of species in these two genera until DNA sequence data showed that Ambrosiella belonged in the Ceratocystidaceae (Microascales) and Raffaelea in the Ophiostomatales (Batra 1967, Cassar & Blackwell 1996, Massoumi Alamouti et al. 2009, Harrington et al. 2010, De Beer et al. 2013a, 2014). Species of Raffaelea have reduced conidiophore morphology, distinguishing them from other Ophiostomatalean genera (De Beer & Wingfield 2013). All species in the genus are mutualists of ambrosia beetles in the sub-families Scolytinae and Platypodinae (Curculionidae), respectively, in North America, South Africa, Europe and Asia (Harrington et al. 2010, Hulcr & Stelinski 2017).
Prior to the study of Musvuugwa et al. (2015), Raffaelea was considered as an exclusively asexual genus. Musvuugwa et al. (2015) described a Raffaelea species forming sexual structures and emended the generic description of Raffaelea to accommodate these structures. In our analyses, R. vaginata (labelled as Lineage X) did not consistently group within Raffaelea (see below). However, our analyses support the inclusion of R. seticollis in Raffaelea, which was one of the other two sexually reproducing species treated in Raffaelea by Musvuugwa et al. (2015). This species was originally described from ambrosial galleries in a hemlock (Tsuga canadensis) stump in New York State (Davidson 1966) but has not been recorded again. Musvuugwa et al. (2015) also transferred O. deltoideosporum to Raffaelea based on a short LSU sequence of the fungus produced by Hausner et al. (1993b), and morphological similarities to O. seticolle. We did not have access to material of this species and could therefore not include it in our analyses. For the present, we have followed the suggestion of Musvuugwa et al. (2015) and include it in Raffaelea.
Raffaelea incertae sedis (Lineage X)
Raffaelea vaginata
Notes: This species was described from a beetle in the genus Lanurgus collected from a native tree in the Southern Cape of South Africa (Musvuugwa et al. 2015). The fungus formed a sexual morph similar to O. deltoideosporum and O. seticolle that grouped in Raffaelea in the study of De Beer & Wingfield (2013). As stated above, analyses of our data supported the inclusion of R. seticollis (O. seticolle) in Raffaelea. Because R. vaginata did not consistently group within Raffaelea in our analyses, it is not included in our narrower definition of Raffaelea. While it might respresent a novel genus, there is insufficient information to make this decision and it is thus treated as incertae sedis.
Sporothrix (Lineage XIV)
Sporothrix Hektoen & C.F. Perkins, J. Exp. Med. 5: 80. 1900. MycoBank MB 10046, emend. Z.W. de Beer et al., Stud. Mycol. 83: 171. 2016. Figs 14, 15.
Fig. 14.
Genera of the Ophiostomatales redrawn from published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Sporothrix candida complex. C, D. Sporothrix gossypina complex. E, F. Sporothrix inflata complex. G, H. Sporothrix pallida complex. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Fig. 15.
Genera of the Ophiostomatales redrawn published images with sexual morphs (if known) on the left and asexual morphs on the right. A, B. Sporothrix pathogenic clade. C, D. Sporothrix stenoceras complex. (Pale grey shading reflects hyaline to subhyaline colouration, medium-tone grey brown to dark brown and dark grey reflects fuscous black to dark black colouration).
Synonyms: Spumatoria Massee & E.S. Salmon, Ann. Bot. (Oxford) 15: 350. 1901. MycoBank MB 5175. [Type species Spumatoria longicollis Massee & E.S. Salmon, Ann. Bot. (Oxford) 15: 351 (1901)].
Sporotrichopsis Guég., Arch. Parasitol. 15: 104. 1911. MycoBank MB 10048. [Type species S. beurmannii (Matr. & Ramond) Guég.; nom. inval., Art. 38.1].
Dolichoascus Thibaut & Ansel, Compt. Rend. Hebd. Seances Acad. Sci. 270: 2173. 1970. MycoBank MB 1684. (Type species D. schenckii Thibaut & Ansel; nom. inval., Art. 40.1).
Etymology: The name is derived from the Latin for ‘spore hair’ (De Hoog 1974).
Sexual morph: Ascomatal bases black, globose; necks black, becoming paler towards apex, straight or slightly curved. Ostiolar hyphae parallel to divergent. Asci evanescent. Ascospores hyaline, 1-celled, allantoid to reniform in side view, almost triangular-shaped, ovoid to oblong.
Asexual morph: Conidiophores micronematous, semi-macronematous, mononematous, simple. Conidiogenous cells showing sympodial growth, terminal or intercalary, cylindrical, denticulate or not denticulate, hyaline. Conidia hyaline to subhyaline, 1-celled, subglobose to oblong, obovoid, clavate to strongly curved, guttuliform to fusiform with round apices and pointed base that sometimes slightly curved, sometimes slightly curved at base; (pathogenic clade and S. inflata complex) borne directly on hyphae (sessile) brown to dark brown, subglobose to globose, narrowly obovoid, ellipsoid, 1-celled. Secondary conidia borne by yeast-like budding, absent or present.
Type species: Sporothrix schenckii Hektoen & C.F. Perkins, J. Exp. Med. 5: 77. 1900.
Other species: Listed in Table 1.
Notes: Sporothrix was described as part of a medical case study concerning a child who had contracted a fungal infection after injuring his finger with a hammer (Hektoen & Perkins 1900). The fungus isolated from the infection was described as Sporothrix schenckii. The epithet referred to B.R. Schenck who described, but did not name a similar fungus in an earlier study. The genus description of Sporothrix by Hektoen & Perkins (1900) was invalid, and the name was validated years later by Nicot & Mariat (1973). In the late 1960’s, S. schenckii was suggested to represent the asexual morph of O. stenoceras (Nicot & Mariat 1973), and subsequently the asexual morphs of many other Ophiostoma species were also treated in Sporothrix under the dual nomenclature system (De Hoog 1974, De Beer & Wingfield 2013).
The phylogenetic placement of S. schenckii in the Ophiostomatales based on ribosomal DNA sequences by Berbee & Taylor (1992) became the first example where asexual (anamorph) and sexual (teleomorph) genera were connected based on phylogenetic inference. This represented a first important step towards the abandonment of the the dual nomenclature system for the fungi, which culminated in the Amsterdam Declaration (Hawksworth et al. 2011) and subsequent changes to the Code where one fungus could only have one species name.
Following the adoption of the One Fungus-One Name convention, De Beer & Wingfield (2013) listed Sporothrix as a synonym of Ophiostoma, with the type species of Sporothrix (S. schenckii) forming part of the S. schenckii-O. stenoceras complex in a broad concept for Ophiostoma. De Beer & Wingfield (2013) noted that this species complex could represent a genus separate from other species in Ophiostoma s.l. However, the available data at the time were insufficient to formally separate the two genera. In a more comprehensive study including sequences of all Sporothrix spp. and all Ophiostoma spp. with Sporothrix asexual morphs, De Beer et al. (2016a) showed that the S. schenckii-O. stenoceras complex formed a well-supported monophyletic lineage, distinct from other species in Ophiostoma. Because this lineage accommodated the type species of Sporothrix, they reinstated the genus, transferring all Ophiostoma species in this lineage to Sporothrix. De Beer et al. (2016a) also emended the description of Sporothrix to include sexual morphs. Furthermore, they defined several species complexes within Sporothrix. The analyses emerging from the present study support the separation of Sporothrix and Ophiostoma based on strong monophyletic lineages. There are a few exceptions where species formed independent lineages and these are discussed below.
Species complexes:
See De Beer et al. (2016a) and below.
The S. candida complex
The S. candida complex accommodates four species that have been described from hardwoods in South Africa (Kamgan Nkuekam et al. 2012, Musvuugwa et al. 2016) and one species from hardwoods in Argentina (De Errasti et al. 2016). Sporothrix itsvo was previously included in this species complex, but it grouped within the S. gossypina/S. stenoceras complex in our phylogenetic analyses (Fig. 5).
The S. inflata complex
The S. inflata complex accommodates species that have been isolated from oak in Europe, soil in Europe, Canada and Malaysia (De Beer et al. 2016a). It also includes the epitype isolate of Spumatoria longicollis, a coprophilous fungus. In a recent study forming part of The Genera of Fungi project (http://www.mycobank.org), Giraldo et al. (2017) showed an isolate of the monotypic genus Spumatoria (Sp. longicollis) grouping within Sporothrix, based on ITS, LSU and BT sequence data. These authors considered this as the first report of the fungus after it was initially described. The type of this species has been lost (De Beer et al. 2014) and consequently Giraldo et al. (2017) designated an epitype, isolated from cow dung in the Netherlands, which was used in the present study. Although this species has a sporothrix-like asexual morph, its septate ascospores distinguished it from all other ophiostomatalean species. Furthermore, its coprophilous nature and pale coloured ascomata distinguish it from other Sporothrix species (De Beer et al. 2014, Giraldo et al. 2017). De Beer et al. (2013a) excluded Spumatoria from the Ophiostomatales, based on these atypical characteristics and characteristics shared with Kathistes, which is phylogeneticially distant from the Ophiostomatales. Our analyses included LSU and TEF-1α sequence data for the epitype isolate, which placed it within the S. inflata species complex (Figs 5, S1, S3).
The S. stenoceras and S. gossypina complexes
The S. stenoceras and S. gossypina species complexes formed an aggregated lineage in the analyses of our datasets (Figs 5, S1, S3). Interestingly, the lineage within the S. stenoceras complex including species isolated from Protea (De Beer et al. 2016a, Ngubane et al. 2018), and S. gemella (also from Protea), formed a lineage distinct from the other species in these complexes, none of which are associated with Protea.
The S. gossypina complex accommodates species that have diverse ecologies (De Beer et al. 2016a). Some species are associated with conifer-infesting bark beetles (Davidson 1971, Marmolejo & Butin 1990, Zhou et al. 2004a, 2006, Lu et al. 2009, Linnakoski et al. 2010, Six et al. 2011, Taerum et al. 2013), others specifically with pine-infesting bark beetles (Davidson 1971, Zhou et al. 2006, Romón et al. 2014a, b), and a few species that have been isolated from stained oak wood (Aghayeva et al. 2004). Interestingly, there is also a species isolated from cankers on chestnut caused by the aggressive tree pathogen Cryphonectria parasitica (Davidson 1978). Species are widely distributed in the USA, Europe, Asia and South Africa (De Beer et al. 2016a).
Sporothrix stenoceras has been isolated from soils and hardwoods on many continents. This could explain why it grouped with species in the S. gossypina complex.
Sporothrix gemella was previously included in the S. pallida complex, but it grouped within the complex in the analyses of our combined dataset (Fig. 5). This species is found on Protea species in South Africa and is vectored by mites, a niche similar to that of S. splendens (Roets et al. 2013, De Beer et al. 2016a).
Ophiostoma ponderosae grouped in this species complex in the analyses of our combined dataset (Fig. 5). The LSU sequence generated by De Beer et al. (2016a) for the same isolate placed this species in Ophiostoma s.s. Our sequence data for O. ponderosae differed from the sequence generated by De Beer et al. (2016a) in only a small number of bases.
The S. pallida complex
Three species forming part of the S. pallida complex (S. pallida, S. mexicana and S. chilensis) have been reported as rare and opportunistic pathogens of humans, but mostly occurring in soil along with other species in this complex. Two species are associated with Protea species occurring in South Africa. Species in this complex have been found in Japan, Mexico, Chile, England and South Africa (De Beer et al. 2016a).
The pathogenic clade
Most Sporothrix species are relatively harmless to humans, but there are four species (S. schenckii, S. luriei, S. brasiliensis and S. globosa) that are human and animal pathogens. These species are the causal agents of the disease known as sporotrichosis (Teixeira et al. 2014). All four species form pigmented blastoconidia, in addition to the more typical sporothrix-like morphology of the asexual morphs. This is possibly an adaptation to both the soil inhabiting and animal-pathogenic lifestyle of these fungi (De Beer et al. 2016a). Collectively, these pathogens have an almost global distribution. Sporothrix schenckii has been isolated in the Americas, Australia, Southern Africa and Europe, and S. globosa in Central America and parts of Eurasia. However, S. brasiliensis is known only from Brazil, where it is responsible for serious epidemic (Rodrigues et al. 2013, 2014). Sporothrix luriei is known based on only a single isolate from South Africa (Zhang et al. 2013, Teixeira et al. 2014).
Group D
Sporothrix polyporicola grouped alone in Sporothrix in our analyses. (Fig. 5). This is consistent with the phylogenies of Osorio et al. (2016) and De Beer et al. (2016a).
Group E
In a similar manner, S. polyporicola and S. phasma grouped alone in Sporothrix in our analyses (Fig. 5). Both species were closely related in the LSU dataset of De Beer et al. (2016a).
Group F
Group F accommodated S. nigrograna, S. curviconia, S. thermara, S. eucalyptigena and S. bragantina. Sporothrix thermara and S. bragantina grouped together as “Lineage H” and S. curviconia as “Lineage G” in the phylogenies of De Beer et al. (2016a).
Group G
Sporothrix dombeyi formed a single lineage basal to Sporothrix in our concatenated dataset (Fig. 5). Previously S. dombeyi (as O. nothofagi) was transferred to Sporothrix and renamed by De Beer et al. (2016a). This species also grouped basal to Sporothrix in their datasets (along with other species treated here as Lineage XVI, XVIII, XIX).
New combination:
1) Sporothrix longicollis (Massee & E.S. Salmon) M. Procter & Z.W. de Beer, comb. nov. MycoBank MB 841005.
Basionym: Spumatoria longicollis Massee & E.S. Salmon, Ann. Bot. 15: 351. 1901. MycoBank MB 171713.
Description: Massee & Salmon (1901: 350–351, fig. 27), Giraldo et al. (2017: 344–345, fig. 9).
Phylogenetic data: Giraldo et al. (2017).
Notes: Sporothrix longicollis is different from other Sporothrix spp. by having 1-septate ascospores, pale-coloured ascomata and coprophilous biology. The inclusion of this species in Sporothrix was also suggested by Giraldo et al. (2017).
Sporothrix incertae sedis (Lineages XVI & XIX)
Sporothrix fumea (Lineage XVI)
Sporothrix fumea grouped basal to all the above-mentioned lineages in the analyses of our combined dataset (Fig. 5). In the phylogenies of De Beer et al. (2016a), S. fumea formed a lineage with O. valdivianum (Lineage XVIII), but in our phylogenies these species grouped separately.
Sporothrix brunneoviolaceae (Lineage XIX)
In the phylogenies of De Beer & Wingfield (2013), S. brunneoviolaceae formed a distinct lineage in Ophiostoma s.l. with S. fumea and O. fasciatum but these were not available for the present study. However, this species grouped basal to Sporothrix in the phylogenies of De Beer et al. (2016a). In the analyses for the present study, S. brunneoviolaceae consistently grouped separately from Sporothrix.
CONCLUSIONS
The results of this study have provided support for the delineation of most of the genera that have been recognised during the course of the last decade in the Ophiostomatales. This study also revealed robust novel lineages described here in four new genera. In addition to phylogenetic support, most of these genera include coherent groups of species that are characterised by similar morphology, ecology and/or geographical origin. The study has also resolved remaining One Fungus-One Name issues that were not dealt with in the revision of De Beer & Wingfield (2013). This is especially by providing new combinations for several species in Grosmannia and Leptographium.
This study is the most extensive and most comprehensive ever undertaken on the taxonomy of the Ophiostomatales. The concatenated dataset included 264 taxa representing all major lineages in the Order. It was not possible to resolve the taxonomic status of some smaller lineages. These lineages may represent new genera or reside in existing genera that will emerge as new species are discovered. We have consequently chosen to retain species in these lineages in their current genera until such data can provide clarity on their taxonomic placement.
The phylogenies generated in this study will serve as a framework for future taxonomic studies on the Ophiostomatales. In the short term, they will facilitate the appropriate generic placement of novel species. But they will also provide the required lists of the appropriate species to include in smaller phylogenies of single genera and/or species complexes that will make it possible to confirm whether studied isolates represent novel taxa.
Acknowledgments
We thank Dr Hugh Glen for advice on Latinized names and Dr Konstanze Bensch of MycoBank for advice on nomenclatural elements. We acknowledge funding from members of the Tree Protection Cooperative Programme (TPCP), the DST/NRF Centre of Excellence in Plant Health Biotechnology (CPHB), South Africa, and the Fibre Processing & Manufacturing Sector Education and Training Authority (FP&M SETA) bursary programme. We acknowledge the support from the South African Research Chairs Initiative (SARChI), specifically Brenda Wingfield’s SARChI Chair in Fugal Genetics for providing early access to some of the fungal genome sequences during this study. Furthermore, thank the many herbaria and culture collections that provided us with material for this study. Reviewers of the originally submitted manuscript, and particularly in one case, provided extensive and important suggestions. These enabled us to re-think particular taxonomic decisions and thus to substantially refine the final product, for which we are most grateful.
DECLARATION ON CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
Supplementary Material: https://studiesinmycology.org/
Phylogenetic tree derived from maximum likelihood analysis of the LSU gene region. The dataset consisted of 233 isolates and 859 characters before and after Gblocks treatment. Bootstrap values above 60 % are shown. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks genera that we redefine here, and blue blocks indicate genera that we here re-instate and re-define.
Phylogenetic tree derived from maximum likelihood analysis of the ITS gene region. The dataset consisted of 235 isolates and 169 characters after Gblocks treatment (1 171 characters prior to Gblocks, including gaps). ML values above 60 % are shown. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks genera that we redefine here, and blue blocks indicate genera that we here re-instate and re-define.
Phylogenetic tree derived from maximum likelihood analysis of the TEF1-α gene region. The dataset consisted of 207 isolates and 380 characters after Gblocks treatment (655 characters prior to Gblocks, including gaps). ML values above 60 % are shown. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks genera that we redefine here, and blue blocks indicate genera that we here re-instate and re-define.
Phylogenetic tree derived from maximum likelihood analysis of the RPBII gene region. The dataset consisted of 171 isolates and 952 characters after Gblocks treatment (1 108 characters prior to Gblocks, including gaps). ML values above 60 % are shown. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks genera that we redefine here, and blue blocks indicate genera that we here re-instate and re-define.
Taxa included in the phylogenomic analyses and their genome sequence statistics
REFERENCES
- Aghayeva DN, Wingfield MJ, De Beer ZW, et al. (2004). Two new Ophiostoma species with Sporothrix anamorphs from Austria and Azerbaijan. Mycologia 96: 866–878. [DOI] [PubMed] [Google Scholar]
- Ando Y, Masuya H, Motohashi K, et al. (2016). Phylogenetic relationship of Japanese isolates belonging to the Grosmannia piceiperda complex (Ophiostomatales). Mycoscience 57: 123–135. [Google Scholar]
- Bakshi BK. (1950). Fungi associated with ambrosia beetles in Great Britain. Transactions of the British Mycological Society 33: 111–120. [Google Scholar]
- Bakshi BK. (1951). Studies on four species of Ceratocystis, with a discussion on fungi causing sap-stain in Britain. Mycological Paper 35: 1–16. [Google Scholar]
- Bateman C, Huang Y-T, Simmons DR, et al. (2017). Ambrosia beetle Premnobius cavipennis (Scolytinae: Ipini) carries highly divergent ascomycotan ambrosia fungus, Afroraffaelea ambrosiae gen. nov. et sp. nov. (Ophiostomatales). Fungal Ecology 25: 41–49. [Google Scholar]
- Batra LR. (1967). Ambrosia fungi: a taxonomic revision and nutritional studies of some species. Mycologia 59: 976–1017. [Google Scholar]
- Benade E, Wingfield MJ, Van Wyk PS. (1997). Conidium development in Sporothrix anamorphs of Ophiostoma. Mycological Research 101: 1108–1112. [Google Scholar]
- Benny GL, Kimbrough JW. (1980). A synopsis of the orders and families of Plectomycetes with keys to genera. Mycotaxon 12: 1–91. [Google Scholar]
- Berbee ML, Taylor JW. (1992). 18S Ribosomal RNA gene sequence characters place the human pathogen Sporothrix schenckii in the genus Ophiostoma. Experimental Mycology 16: 87–91. [Google Scholar]
- Blanco-Ulate B, Rolshausen P, Cantu D. (2013). Draft genome sequence of the ascomycete Phaeoacremonium aleophilum strain UCR-PA7, a causal agent of the esca disease complex in grapevines. Genome Announcements 3: e00390-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brasier CM. (1979). Dual origin of recent Dutch elm disease outbreaks in Europe. Nature 281: 78–80. [Google Scholar]
- Brasier CM. (1991). Ophiostoma novo-ulmi sp nov. – causative agent of current Dutch elm disease pandemics. Mycopathologia 115: 151–161. [Google Scholar]
- Brasier CM, Mehrotra MD. (1995) Ophiostoma himal-ulmi sp. nov., a new species of Dutch elm disease fungus endemic to the Himalayas. Mycological Research 99: 205–215. [Google Scholar]
- Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T. (2009). trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25: 1972–1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cassar S, Blackwell M. (1996). Convergent origins of ambrosia fungi. Mycologia 88: 596–601. [Google Scholar]
- Castresana J. (2000). Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Molecular Biology and Evolution 17: 540–552. [DOI] [PubMed] [Google Scholar]
- Chang R, Duong TA, Taerum SJ, et al. (2017). Ophiostomatoid fungi associated with conifer-infesting beetles and their phoretic mites in Yunnan, China. MycoKeys 28: 19–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang R, Duong TA, Taerum SJ, et al. (2019). Ophiostomatoid fungi associated with the spruce bark beetle Ips typographus, including 11 new species from China. Persoonia 42: 50–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chesters CGC. (1935). Studies on British Pyrenomycetes. I. The life histories of three species of Cephalotheca Fuck. Transactions of the British Mycological Society 19: 261–279. [Google Scholar]
- Cobb FW, Jr. (1988) Leptographium wageneri, cause of black-stain root disease: a review of its discovery, occurrence and biology with emphasis on pinyon and ponderosa pine. Leptographium root diseases on conifers. (Harrington TC, Cobb FW, ed.): 41–62. APS press, St. Paul, Minnesota. [Google Scholar]
- Cobb FW, Jr., Goheen DJ, Harrington TC. (1984). Black-stain root disease caused by Verticicladiella wageneri. In: Proceedings of the Fourth International Congress for Plant Pathology, Melbourne, Australia. p. 84. [Google Scholar]
- Cobb FW, Jr., Parmeter JR, Jr., Wood DL, et al. (1974). Root pathogens as agents predisposing ponderosa pine and white fir to bark beetles. In: Proceedings of the fourth international conference on Fomes annosus (Kuhlman E.G., ed.): 8–15. USDA Forestry Service, Washington, DC, USA. [Google Scholar]
- D’Alessandro E, Giosa D, Huang L, et al. (2016). Draft genome sequence of the dimorphic fungus. Genome Announcements 4: e00184-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davidson RW. (1942). Some additional species of Ceratostomella in the United States. Mycologia 34: 650–662. [Google Scholar]
- Davidson RW. (1958). Additional species of Ophiostomataceae from Colorado. Mycologia 50: 661–670. [Google Scholar]
- Davidson RW. (1966). New species of Ceratocystis from conifers. Mycopathologia et Mycologia Applicata 28: 273–286. [Google Scholar]
- Davidson RW. (1971). New species of Ceratocystis. Mycologia 63: 5–15. [Google Scholar]
- Davidson RW. (1978). A new species of Ceratocystis on Endothia parasitica canker of American chestnut. Mycologia 70: 856–858. [Google Scholar]
- Davidson RW, Käärik A, Francke-Grosmann H. (1967). A restudy of Ceratocystis penicillata and report of two American species of this genus from Europe. Mycologia 59: 928–932. [Google Scholar]
- De Beer ZW, Begerow D, Bauer R, et al. (2006). Phylogeny of the Quambalariaceae fam. nov., including important Eucalyptus pathogens in South Africa and Australia. Studies in Mycology 55: 289–298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Beer ZW, Duong TA, Barnes I, et al. (2014). Redefining Ceratocystis and allied genera. Studies in Mycology 79: 187–219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Beer ZW, Duong TA, Wingfield MJ. (2016a). The divorce of Sporothrix and Ophiostoma: solution to a problematic relationship. Studies in Mycology 83: 165–191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Beer ZW, Marincowitz S, Duong TA, et al. (2016b). Hawksworthiomyces gen. nov. (Ophiostomatales), illustrates the urgency for a decision on how to name novel taxa known only from environmental nucleic acid sequences (ENAS). Fungal Biology 120: 1323–1340. [DOI] [PubMed] [Google Scholar]
- De Beer ZW, Seifert K, Wingfield MJ. (2013a). The ophiostomatoid fungi: their dual position in the Sordariomycetes. In: The Ophiostomatoid Fungi: Expanding Frontiers, (Seifert KA, De Beer ZW, Wingfield MJ, ed.): 1–19. CBS-KNAW Fungal Biodiversity Centre, Utrecht, The Netherlands. [Google Scholar]
- De Beer ZW, Seifert KA, Wingfield MJ. (2013b). A nomenclator for ophiostomatoid genera and species in the Ophiostomatales and Microascales. In: The Ophiostomatoid Fungi: Expanding Frontiers, (Seifert KA, De Beer ZW, Wingfield MJ, ed.): 245–322. CBS-KNAW Fungal Biodiversity Centre, Utrecht, The Netherlands. [Google Scholar]
- De Beer ZW, Wingfield MJ. (2013). Emerging lineages in the Ophiostomatales. In: The Ophiostomatoid Fungi: Expanding Frontiers, (Seifert KA, De Beer ZW, Wingfield MJ, ed.): 21–46. CBS-KNAW Fungal Biodiversity Centre, Utrecht, The Netherlands. [Google Scholar]
- De Errasti A, De Beer ZW, Coetzee MPA, et al. (2016). Three new species of Ophiostomatales from Nothofagus in Patagonia. Mycological Progress 15: 17. [Google Scholar]
- De Errasti A, Pildain MB, Rajchenberg M. (2018). Ophiostomatoid fungi isolated from three different pine species in Argentinian Patagonia. Forest Pathology 48: e12393. [Google Scholar]
- De Hoog GS. (1974). The genera Blastobotrys, Sporothrix, Calcarisporium and Calcarisporiella gen. nov. Studies in Mycology 7: 1–84. [Google Scholar]
- De Hoog GS, Scheffer R. (1984). Ceratocystis versus Ophiostoma: a reappraisal. Mycologia 76: 292–299. [Google Scholar]
- De Meyer EM, De Beer ZW, Summerell RC, et al. (2008). Taxonomy and phylogeny of new wood- and soil-inhabiting Sporothrix species in the Ophiostoma stenoceras-Sporothrix schenckii complex. Mycologia 100: 647–661. [DOI] [PubMed] [Google Scholar]
- Dean RA, Talbot NJ, Ebbole DJ, et al. (2005). The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434: 980–986. [DOI] [PubMed] [Google Scholar]
- DiGuistini S, Wang Y, Liao NY, et al. (2011). Genome and transcriptome analyses of the mountain pine beetle-fungal symbiont Grosmannia clavigera, a lodgepole pine pathogen. Proceedings of the National Academy of Sciences 108: 2504–2509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dochinger LS. (1967). Leptographium root decline of eastern white pine. Phytopathology 57: 809. [Google Scholar]
- Dreaden TJ, Davis JM, De Beer ZW, et al. (2014). Phylogeny of ambrosia beetle symbionts in the genus Raffaelea. Fungal Biology 118: 970–978. [DOI] [PubMed] [Google Scholar]
- Duong TA, De Beer ZW, Wingfield BD, et al. (2012). Phylogeny and taxonomy of species in the Grosmannia serpens complex. Mycologia 104: 715–732. [DOI] [PubMed] [Google Scholar]
- Endoh R, Suzuki M, Okada G, et al. (2011). Fungus symbionts colonizing the galleries of the ambrosia beetle Platypus quercivorus. Microbial Ecology 62: 106–120. [DOI] [PubMed] [Google Scholar]
- Fraedrich SW, Harrington TC, Rabaglia RJ, et al. (2008). A fungal symbiont of the redbay ambrosia beetle causes a lethal wilt in redbay and other Lauraceae in the southeastern United States. Plant Disease 92: 215–224. [DOI] [PubMed] [Google Scholar]
- Forgetta V, Leveque G, Dias J, et al. (2013). Sequencing of the Dutch elm disease fungus genome using the Roche/454 GS-FLX titanium system in a comparison of multiple genomics core facilities. Journal of Biomolecular Techniques 24: 39–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gharabigloozare Y. (2015). Raffaelea spp. from five ambrosia beetles in the genera Xyleborinus and Cyclorhipidion (Coleoptera: Curcurlionidae: Scolytinae: Xyleborini): 11–47. Iowa State University, Graduate Theses and Dissertations. 14815. Ames, Iowa, USA. [Google Scholar]
- Gardes M, Bruns TD. (1993). ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Molecular Ecology 2: 113–118. [DOI] [PubMed] [Google Scholar]
- Gebhardt H, Kirschner R, Oberwinkler F. (2002). A new Ophiostoma species isolated from the ambrosia beetle Xyleborus dryographus (Coleoptera: Curculionidae, Scolytinae). Mycological Progress 1: 377–382. [Google Scholar]
- Gebhardt H, Weiss M, Oberwinkler F. (2005). Dryadomyces amasae: a nutritional fungus associated with ambrosia beetles of the genus Amasa (Coleoptera: Curculionidae, Scolytinae). Mycological Research 109: 687–696. [DOI] [PubMed] [Google Scholar]
- Gibbs JN. (1978). Development of the Dutch elm disease epidemic in southern England, 1971–6. Annals of Applied Biology 88: 219–228. [Google Scholar]
- Giraldo A, Crous PW, Schumacher RK, et al. (2017). The Genera of Fungi—G3: Aleurocystis, Blastacervulus, Clypeophysalospora, Licrostroma, Neohendersonia and Spumatoria. Mycological Progress 16: 325–348. [Google Scholar]
- Goheen DJ, Cobb F. (1978). Occurrence of Verticicladiella wagenerii and its perfect state, Ceratocystis wageneri sp. nov., in insect galleries. Phytopathology 68: 1192–1195. [Google Scholar]
- Goidànich G. (1935). A new species of Ophiostoma living on pear and some observations on the exact systematic position of the ascigerous form and the metagenetic forms of the genus. Bollettino della R Stazione di Patologia Vegetale 15: 122–168. [Google Scholar]
- Gorton C, Kim SH, Henricot B, et al. (2004). Phylogenetic analysis of the bluestain fungus Ophiostoma minus based on partial ITS rDNA and β-tubulin gene sequences. Mycological Research 108: 759–765. [DOI] [PubMed] [Google Scholar]
- Greif MD, Gibas CFC, Currah RS. (2006). Leptographium piriforme sp. nov., from a taxonomically diverse collection of arthropods collected in an aspen-dominated forest in western Canada. Mycologia 98: 771–780. [DOI] [PubMed] [Google Scholar]
- Griffin HD. (1968). The genus Ceratocystis in Ontario. Canadian Journal of Botany 46: 689–718. [Google Scholar]
- Grobbelaar JW, Bloomer P, Wingfield MJ, et al. (2011). Discovery of Ophiostoma tsotsi on Eucalyptus wood chips in China. Mycoscience 52: 111–118. [Google Scholar]
- Grobbelaar JW, De Beer ZW, Bloomer P, et al. (2010) Ophiostoma tsotsi sp. nov., a wound-infesting fungus of hardwood trees in Africa. Mycopathologia 169: 413–423. [DOI] [PubMed] [Google Scholar]
- Hansen EM, Goheen DJ, Hessburd PF, et al. (1988). Biology and management of black stain root disease in Douglas fir. In: root disease in conifers (Harrington TC, Cobb FW, eds): 63–80. American Phytopathological Society, St Paul, Minnesota, USA. [Google Scholar]
- Haridas S, Wang Y, Lim L, et al. (2013). The genome and transcriptome of the pine saprophyte Ophiostoma piceae, and a comparison with the bark beetle-associated pine pathogen Grosmannia clavigera. BMC Genomics 14: 373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrington TC. (1981). Cycloheximide sensitivity as a taxonomic character in Ceratocystis. Mycologia 73: 1123–1129. [Google Scholar]
- Harrington TC. (1988). Leptographium species, their distributions, hosts and insect vectors. In: Leptographium root diseases on conifers, (Harrington TC, Cobb FW, eds.): 1–40. American Phytopathological Society Press, St. Paul, Minnesota, USA. [Google Scholar]
- Harrington T, Cobb F., Jr (1984). Host specialization of three morphological variants of Verticicladiella wageneri. Phytopathology 74: 286–290. [Google Scholar]
- Harrington T, Cobb F., Jr (1987). Leptographium wageneri var. pseudotsugae, var. nov., cause of black stain root disease on Douglas fir. Mycotaxon 30: 501–507. [Google Scholar]
- Harrington TC, Aghayeva DN, Fraedrich SW. (2010). New combinations in Raffaelea, Ambrosiella, and Hyalorhinocladiella, and four new species from the redbay ambrosia beetle, Xyleborus glabratus. Mycotaxon 111: 337–361. [Google Scholar]
- Harrington T, Fraedrich S, Aghayeva D. (2008). Raffaelea lauricola, a new ambrosia beetle symbiont and pathogen on the Lauraceae. Mycotaxon 104: 399–404. [Google Scholar]
- Harrington TC, McNew D, Steimel J, et al. (2001). Phylogeny and taxonomy of the Ophiostoma piceae complex and the Dutch Elm Disease fungi. Mycologia 93: 111–136. [Google Scholar]
- Harrington TC, Yun HY, Lu S-S, et al. (2011). Isolations from the redbay ambrosia beetle, Xyleborus glabratus, confirm that the laurel wilt pathogen, Raffaelea lauricola, originated in Asia. Mycologia 103: 1028–1036. [DOI] [PubMed] [Google Scholar]
- Hausner G, Eyjolfsdottir GG, Reid J. (2003). Three new species of Ophiostoma and notes on Cornuvesica falcata. Canadian Journal of Botany 81: 40. [Google Scholar]
- Hausner G, Reid J, Klassen GR. (1992). Do galeate-ascospore members of the Cephaloascaceae, Endomycetaceae and Ophiostomataceae share a common phylogeny? Mycologia 84: 870–881. [Google Scholar]
- Hausner G, Reid J, Klassen GR. (1993a). Ceratocystiopsis: a reappraisal based on molecular criteria. Mycological Research 97: 625–633. [Google Scholar]
- Hausner G, Reid J, Klassen GR. (1993b). On the phylogeny of Ophiostoma, Ceratocystis s.s., and Microascus, and relationships within Ophiostoma based on partial ribosomal DNA sequences. Canadian Journal of Botany 71: 1249–1265. [Google Scholar]
- Hausner G, Reid J, Klassen GR. (1993c). On the subdivision of Ceratocystis s.l., based on partial ribosomal DNA sequences. Canadian Journal of Botany 71: 52–63. [Google Scholar]
- Hausner G, Reid J, Klassen GR. (2000). On the phylogeny of members of Ceratocystis s.s. and Ophiostoma that possess different anamorphic states, with emphasis on the anamorph genus Leptographium, based on partial ribosomal DNA sequences. Canadian Journal of Botany 78: 903–916. [Google Scholar]
- Hawksworth DL. (2012). Managing and coping with names of pleomorphic fungi in a period of transition. IMA Fungus 3: 15–24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawksworth DL, Crous PW, Redhead SA, et al. (2011). The Amsterdam Declaration on fungal nomenclature. IMA Fungus 2: 105–112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hektoen L, Perkins CF. (1900). Refractory subcutaneous abscesses caused by Sporothrix schenckii. A new pathogenic fungus. The Journal of Experimental Medicine 5(1): 77–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hessburg PF, Hansen EM. (2000). Infection of Douglas-fir by Leptographium wageneri. Canadian Journal of Botany 78: 1254–1261. [Google Scholar]
- Hoang DT, Chernomor O, Von Haeseler A, et al. (2018). UFBoot2: Improving the Ultrafast Bootstrap Approximation. Molecular Biology and Evolution 35: 518–522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang Y-T, Chen C-Y. (2014). Leptographium globosum sp. nov., a new species with globose conidia. Mycological Progress 13: 841–848. [Google Scholar]
- Huang L, Gao W, Giosa D, et al. (2016). Whole-genome sequencing and in silico analysis of two strains of Sporothrix globosa. Genome Biology and Evolution 8: 3292–3296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hulcr J, Stelinski LL. (2017). The ambrosia symbiosis: from evolutionary ecology to practical management. Annual Review of Entomology 62: 285–303. [DOI] [PubMed] [Google Scholar]
- Hunt J. (1956). Taxonomy of the genus Ceratocystis. Lloydia 19: 1–58. [Google Scholar]
- Hutchison LJ, Reid J. (1988a). Taxonomy of some potential wood-staining fungi from New Zealand 1. Ophiostomataceae. New Zealand Journal of Botany 26: 63–81. [Google Scholar]
- Hutchison LJ, Reid J. (1988b). Taxonomy of some potential wood-staining fungi from New Zealand. 2. Pyrenomycetes, Coelomycetes and Hyphomycetes. New Zealand Journal of Botany 26: 83–98. [Google Scholar]
- Jacobs K, Bergdahl DR, Wingfield MJ, et al. (2004). Leptographium wingfieldii introduced into North America and found associated with exotic Tomicus piniperda and native bark beetles. Mycological Research 108: 411–418. [DOI] [PubMed] [Google Scholar]
- Jacobs K, Eckhardt LG, Wingfield MJ. (2006). Leptographium profanum sp. nov., a new species from hardwood roots in North America. Canadian Journal of Botany 84: 759–766. [Google Scholar]
- Jacobs K, Kirisits T. (2003). Ophiostoma kryptum sp. nov. from Larix decidua and Picea abies in Europe, similar to O. minus. Mycological Research 107: 1231–1242. [DOI] [PubMed] [Google Scholar]
- Jacobs K, Seifert KA, Harrison KJ, et al. (2003). Identity and phylogenetic relationships of ophiostomatoid fungi associated with invasive and native Tetropium species (Coleoptera: Cerambycidae) in Atlantic Canada. Canadian Journal of Botany 81: 316–329. [Google Scholar]
- Jacobs K, Solheim H, Wingfield BD, et al. (2005). Taxonomic re-evaluation of Leptographium lundbergii based on DNA sequence comparisons and morphology. Mycological Research 109: 1149–1161. [DOI] [PubMed] [Google Scholar]
- Jacobs K, Wingfield MJ. (2001). Leptographium species: tree pathogens, insect associates, and agents of blue-stain. American Phytopathological Society Press, St. Paul, Minnesota, USA. [Google Scholar]
- Jacobs K, Wingfield MJ, Crous PW. (2000b). Ophiostoma europhioides and Ceratocystis pseudoeurophioides, synonyms of O. piceaperdum. Mycological Research 104: 238–243. [Google Scholar]
- Jacobs K, Wingfield MJ, Crous PW, et al. (1998). Leptographium engelmannii, a synonym of Leptographium abietinum, and description of Leptographium hughesii sp. nov. Canadian Journal of Botany 76: 1660–1667. [Google Scholar]
- Jacobs K, Wingfield MJ, Jacobs A, et al. (2001b). A taxonomic re-evaluation of Phialocephala phycomyces. Canadian Journal of Botany 79: 110–117. [Google Scholar]
- Jacobs K, Wingfield MJ, Pashenova NV, et al. (2000a). A new Leptographium species from Russia. Mycological Research 104: 1524–1529. [Google Scholar]
- Jacobs K, Wingfield MJ, Uzunovic A, et al. (2001c) Three new species of Leptographium from pine. Mycological Research 105: 490–499. [Google Scholar]
- Jacobs K, Wingfield MJ, Wingfield BD. (2001a). Phylogenetic relationships in Leptographium based on morphological and molecular characters. Canadian Journal of Botany 79: 719–732. [Google Scholar]
- Jankowiak R, Bilański P, Strzałka B, et al. (2019). Four new Ophiostoma species associated with conifer- and hardwood-infesting bark and ambrosia beetles from the Czech Repubic and Poland. Antonie van Leeuwenhoek 112: 1501–1521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jankowiak R, Ostafińska A, Aas T, et al. (2018). Three new Leptographium spp. (Ophiostomatales) infecting hardwood trees in Norway and Poland. Antonie van Leeuwenhoek 111: 2323–2347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jankowiak R, Solheim H, Bilański P, et al. (2020) Seven new species of Graphilbum from conifers in Norway, Poland, and Russia. Mycologia 112: 1240–1262. [DOI] [PubMed] [Google Scholar]
- Jankowiak R, Strzałka B, Bilański P, et al. (2017). Two new Leptographium spp. reveal an emerging complex of hardwood-infecting species in the Ophiostomatales. Antonie van Leeuwenhoek 110: 1537–1553. [DOI] [PubMed] [Google Scholar]
- Jeon J, Kim KT, Song H, et al. (2017). Draft Genome Sequence of the Fungus Associated with Oak Wilt Mortality in South Korea, Raffaelea quercus-mongolicae KACC44405.Genome Announcements 5: e00797-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jooste W. (1978). Leptographium reconditum sp. nov. and observations on conidiogenesis in Verticicladiella. Transactions of the British Mycological Society 70: 152–155. [Google Scholar]
- Junier T, Zdobnov EM. (2010). The Newick utilities: high-throughput phylogenetic tree processing in the UNIX shell. Bioinformatics 26: 1669–1670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamgan Nkuekam G, De Beer ZW, Wingfield MJ, et al. (2011). Ophiostoma species (Ophiostomatales, Ascomycota), including two new taxa on eucalypts in Australia. Australian Journal of Botany 59: 283–297. [Google Scholar]
- Kamgan Nkuekam G, De Beer ZW, Wingfield MJ, et al. (2012) A diverse assemblage of Ophiostoma species, including two new taxa on eucalypt trees in South Africa. Mycological Progress 11(2): 515–533. [Google Scholar]
- Kamgan Nkuekam G, Jacobs K, De Beer ZW, et al. (2008). Ceratocystis and Ophiostoma species, including three new taxa, associated with wounds on native South African trees. Fungal Diversity 29: 37–59. [Google Scholar]
- Kamgan Nkuekam G, Solheim H, De Beer ZW, et al. (2010) Ophiostoma species, including Ophiostoma borealis sp. nov., infecting wounds of native broad-leaved trees in Norway. Cryptogamie, Mycologie 31: 285–303. [Google Scholar]
- Katoh K, Standley DM. (2013). MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution 30: 772–780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kendrick WB. (1962). The Leptographium complex. Verticicladiella Hughes. Canadian Journal of Botany 40: 771–797. [Google Scholar]
- Khoshraftar S, Hung S, Khan S, et al. (2013). Sequencing and annotation of the Ophiostoma ulmi genome. BMC Genomics 14: 162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim J-J, Lim YW, Breuil C, et al. (2005a). A new Leptographium species associated with Tomicus piniperda infesting pine logs in Korea. Mycological Research 109: 275–284. [DOI] [PubMed] [Google Scholar]
- Kim J-J, Lim YW, Seifert K, et al. (2005b). Taxonomy of Ophiostoma radiaticola sp. nov. (Ophiostomatales, Ascomycetes), the teleomorph of Pesotum pini, isolated from logs of Pinus radiata. Mycotaxon 91: 481–496. [Google Scholar]
- Kim J-J, Lim YW, Wingfield MJ, et al. (2004). Leptographium bistatum sp. nov., a new species with a Sporothrix synanamorph from Pinus radiata in Korea. Mycological Research 108: 699–706. [DOI] [PubMed] [Google Scholar]
- Kim K-H, Choi Y-J, Seo S-T, et al. (2009). Raffaelea quercus-mongolicae sp. nov. associated with Platypus koryoensis on oak in Korea. Mycotaxon 110: 189–197. [Google Scholar]
- Kim S, Harrington TC, Lee JC, et al. (2011). Leptographium tereforme sp. nov. and other Ophiostomatales isolated from the root-feeding bark beetle Hylurgus ligniperda in California. Mycologia 103: 152–163. [DOI] [PubMed] [Google Scholar]
- Kowalski T, Butin H. (1989). Taxonomy of known and new species of Ceratocystis from oak (Quercus robur L.). Journal of Phytopathology 124: 236–248. [Google Scholar]
- Kubono T, Ito S. (2002). Raffaelea quercivora sp. nov. associated with mass mortality of Japanese oak, and the ambrosia beetle (Platypus quercivorus). Mycoscience 43: 255–260. [Google Scholar]
- Kück P, Meusemann K. (2010). FASconCAT: convenient handling of data matrices. Molecular Phylogenetics and Evolution 56: 1115–1118. [DOI] [PubMed] [Google Scholar]
- Lagerberg T, Lundberg G, Melin E. (1927). Biological and practical researches into blueing in pine and spruce. Svenska Skogsvårdsfören Tidskrift 25: 145–272. [Google Scholar]
- Lah L, Löber U, Hsiang T, et al. (2017). A genomic comparison of putative pathogenicity-related gene families in five members of the Ophiostomatales with different lifestyles. Fungal Biology 121: 234–252. [DOI] [PubMed] [Google Scholar]
- Lanfear R, Fandsen PB, Wright AM, et al. (2017). PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Molecular Biology and Evolution 34(3): 772–773. [DOI] [PubMed] [Google Scholar]
- Lartillot N, Rodrigue N, Stubbs D, et al. (2013). PhyloBayes MPI: Phylogenetic reconstruction with infinite mixtures of profiles in a parallel environment. Systematic Biology 62: 611–615. [DOI] [PubMed] [Google Scholar]
- Leach JG, Orr LW, Christiansen C. (1934). The interrelationship of bark beetles and blue staining fungi in felled Norway pine timber. Journal of Agricultural Research 49: 315–341. [Google Scholar]
- Li Y, Huang Y-T, Kasson MT, et al. (2018). Specific and promiscuous ophiostomatalean fungi associated with Platypodinae ambrosia beetles in the southeastern United States. Fungal Ecology 35: 42–50. [Google Scholar]
- Linnakoski R, De Beer ZW, Ahtiainen J, et al. (2010). Ophiostoma spp. associated with pine- and spruce-infesting bark beetles in Finland and Russia. Persoonia 25: 72–93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linnakoski R, De Beer ZW, Duong TA, et al. (2012). Grosmannia and Leptographium spp. associated with conifer-infesting bark beetles in Finland and Russia, including Leptographium taigense sp. nov. Antonie van Leeuwenhoek 102: 375–399. [DOI] [PubMed] [Google Scholar]
- Linnakoski R, De Beer ZW, Rousi M, et al. (2008) Fungi including Ophiostoma karelicum sp. nov., associated with Scolytus ratzeburgi infesting birch in Finland and Russia. Mycological Research 112: 1475–1488 [DOI] [PubMed] [Google Scholar]
- Linnakoski R, De Beer ZW, Rousi M, et al. (2009) Ophiostoma denticiliatum sp. nov. and other Ophiostoma species associated with the birch bark beetle in southern Norway. Persoonia 23:9–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linnakoski R, Jankowiak R, Villari C, et al. (2016). The Ophiostoma clavatum species complex: a newly defined group in the Ophiostomatales including three novel taxa. Antonie van Leeuwenhoek 109: 987–1018. [DOI] [PubMed] [Google Scholar]
- Liou JY, Shih JY, Tzean SS. (1999). Esteya, a new nematophagous genus from Taiwan, attacking the pinewood nematode (Bursaphelenchus xylophilus). Mycological Research 103: 242–248. [Google Scholar]
- Liu F, Chen S, Ferreira MA, et al. (2019). Draft genome sequences of five Calonectria species from Eucalyptus plantations in China, Celoporthe dispersa, Sporothrix phasma and Alectoria sarmentosa. IMA Fungus 10: 22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu X-W, Wang H-M, Lu Q, et al. (2017). Taxonomy and pathogenicity of Leptographium species associated with Ips subelongatus infestations of Larix spp. in northern China, including two new species. Mycological Progress 16: 1–13. [Google Scholar]
- Löytynoja A. (2014). Phylogeny-aware alignment with PRANK. Methods in Molecular Biology 1079: 155–170. [DOI] [PubMed] [Google Scholar]
- Lu Q, Decock C, Zhang XY, et al. (2008). Leptographium sinoprocerum sp. nov., an undescribed species associated with Pinus tabuliformis-Dendroctonus valens in northern China. Mycologia 100: 275–290. [DOI] [PubMed] [Google Scholar]
- Lu Q, Decock C, Zhang XY, et al. (2009). Ophiostomatoid fungi (Ascomycota) associated with Pinus tabuliformis infested by Dendroctonus valens (Coleoptera) in northern China and an assessment of their pathogenicity on mature trees. Antonie van Leeuwenhoek 96: 275–293. [DOI] [PubMed] [Google Scholar]
- Madrid H, Gené J, Cano J, et al. (2010). Sporothrix brunneoviolacea and Sporothrix dimorphospora, two new members of the Ophiostoma stenoceras-Sporothrix schenckii complex. Mycologia 102: 1193–1203. [DOI] [PubMed] [Google Scholar]
- Maekawa N, Tsuneda A, Arita I. (1987). Ceratocystis species occurring on the Lentinus edodes bedlogs. Reports of the Tottori Mycological Institute 25: 6–14. [Google Scholar]
- Mai U, Mirarab S. (2018). TreeShrink: fast and accurate detection of outlier long branches in collections of phylogenetic trees. BMC Genomics 19: 272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malloch D, Blackwell M. (1993). Dispersal biology of the ophiostomatoid fungi. In: Ceratocystis and ophiostoma: taxonomy, ecology, and pathogenicity, (Wingfield MJ, Seifert KA, Webber JF, ed.): 195–206. American Phytopathological Society Press., St. Paul, Minnesota. [Google Scholar]
- Malloch D, Cain RF. (1970). Five new genera in the new family Pseudeurotiaceae. Canadian Journal of Botany 48: 1815–1825. [Google Scholar]
- Marincowitz S, Duong TA, De Beer ZW, et al. (2015). Cornuvesica: A little known mycophilic genus with a unique biology and unexpected new species. Fungal Biology 119: 615–630. [DOI] [PubMed] [Google Scholar]
- Marincowitz S, Duong TA, Heiniger U, et al. (2017). A new Leptographium species from the roots of declining Pinus sylvestris in Switzerland. Forest Pathology 47: e12346. [Google Scholar]
- Marincowitz S, Duong TA, Stephen SJ, et al. (2020) Fungal associates of an invasive pine-infesting bark beetle, Dendroctonus valens including seven new Ophiostomatalean fungi. Persoonia 45: 177–195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marmolejo JG, Butin H. (1990). New conifer-inhabiting species of Ophiostoma and Ceratocystiopsis (Ascomycetes, Microascales) from Mexico. Sydowia 422: 193–199. [Google Scholar]
- Massee G, Salmon ES. (1901) Researches on coprophilous fungi. Annals of Botany 15: 313–357. [Google Scholar]
- Massoumi Alamouti S, Kim J-J, Breuil C. (2006). A new Leptographium species associated with the northern spruce engraver, Ips perturbatus, in western Canada. Mycologia 98: 149–160. [DOI] [PubMed] [Google Scholar]
- Massoumi Alamouti S, Tsui CKM, Breuil C. (2009). Multigene phylogeny of filamentous ambrosia fungi associated with ambrosia and bark beetles. Mycological Research 113: 822–835. [DOI] [PubMed] [Google Scholar]
- Massoumi Alamouti S, Wang V, DiGuistini S, et al. (2011). Gene genealogies reveal cryptic species and host preferences for the pine fungal pathogen Grosmannia clavigera. Molecular Ecology 20: 2581–2602. [DOI] [PubMed] [Google Scholar]
- Masuya H, Kaneko S, Yamaoka Y. (2003). Three new Ophiostoma species isolated from Japanese red pine. Mycoscience 44: 301–310. [Google Scholar]
- Masuya H, Manabe RI, Ohkuma M, et al. (2016). Draft genome sequence of Raffaelea quercivora JCM 11526, a Japanese oak wilt pathogen associated with the platypodid beetle, Platypus quercivorus. Genome Announcements 4: e00755-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masuya H, Wingfield M, Kubono T, et al. (2004). Leptographium pruni, sp. nov. from bark beetle-infested Prunus jamasakura in Japan. Mycologia 96: 548–557. [PubMed] [Google Scholar]
- Masuya H, Yamaoka Y, Wingfield MJ. (2013). Ophiostomatoid fungi and their associations with bark beetles in Japan. In: Ophiostomatoid Fungi: Expanding Frontiers, (Seifert KA, De Beer ZW, Wingfield MJ, ed.): 77–90. CBS-KNAW Fungal Biodiversity Centre, Utrecht, the Netherlands. [Google Scholar]
- Mathiesen-Käärik A. (1953). Eine Übersicht über die gewöhnlichsten mit Borkenkäfern assoziierten Bläuepilze in Schweden und einige für Schweden neue Bläuepilze. Meddelanden från Statens Skogsforskningsinstitut 43: 1–74. [Google Scholar]
- Mathiesen-Käärik A. (1960). Studies on the ecology, taxonomy and physiology of Swedish insect-associated blue stain fungi, especially the genus Ceratocystis. Oikos 11: 1–25. [Google Scholar]
- Matsuda Y, Kimura K, Ito S-I. (2010). Genetic characterization of Raffaelea quercivora isolates collected from areas of oak wilt in Japan. Mycoscience 51: 310–316. [Google Scholar]
- Mayers CG, Harrington TC, Masuya H, et al. (2020). Patterns of coevolution between ambrosia beetle mycangia and the Ceratocystidaceae, with five new fungal genera and seven new species. Persoonia 44: 41–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayers CG, McNew DL, Harrington TC, et al. (2015). Three genera in the Ceratocystidaceae are the respective symbionts of three independent lineages of ambrosia beetles with large, complex mycangia. Fungal Biology 119: 1075–1092. [DOI] [PubMed] [Google Scholar]
- Melin E, Nannfeldt JA. (1934). Researches into the blueing of ground wood-pulp. Svensk Skogsvardsforeningens Tidskrift 32: 397–616. [Google Scholar]
- McNeill J, Barrie FR, Buck WR, et al. (2012). International Code of Nomenclature for algae, fungi, and plants (Melbourne Code). Regnum Vegetabile 154. ARG Gantner Verlag KG, Germany. [Google Scholar]
- Minh BQ, Schmidt HA, Chernomor O, et al. (2020). IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Molecular Biology and Evolution 37: 1530–1534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirarab S, Reaz R, Bayzid MS, et al. (2014). ASTRAL: genome-scale coalescent-based species tree estimation. Bioinformatics 30: i541–i548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morales-Cruz A, Amrine KC, Blanco-Ulate B, et al. (2015). Distinctive expansion of gene families associated with plant cell wall degradation, secondary metabolism, and nutrient uptake in the genomes of grapevine trunk pathogens. BMC Genomics 16: 469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morelet M. (1998). Une espèce nouvelle de Raffaelea, isolée de Platypus cylindrus, coléoptère xylomycétophage des chênes. Annales de la Société des Sciences Naturelles et d’Archéologie de Toulon et du Var. 50: 185–193. [Google Scholar]
- Mouton M, Wingfield MJ, Van Wyk PS. (1992). The anamorph of Ophiostoma frankae-grosmanniae is a Leptographium. Mycologia 84: 857–862. [Google Scholar]
- Mullineux T, Hausner G. (2009). Evolution of rDNA ITS1 and ITS2 sequences and RNA secondary structures within members of the fungal genera Grosmannia and Leptographium. Fungal Genetics and Biology 46: 855–867. [DOI] [PubMed] [Google Scholar]
- Mullineux T, Willows K, Hausner G. (2011). Evolutionary dynamics of the mS952 Intron: A novel Mitochondrial Group II Intron encoding a LAGLIDADG homing endonuclease gene. Journal of Molecular Evolution 72: 433–449. [DOI] [PubMed] [Google Scholar]
- Musvuugwa T, De Beer ZW, Duong TA, et al. (2015). New species of Ophiostomatales from Scolytinae and Platypodinae beetles in the Cape Floristic Region, including the discovery of the sexual state of Raffaelea. Antonie van Leeuwenhoek 108: 933–950. [DOI] [PubMed] [Google Scholar]
- Musvuugwa T, De Beer ZW, Duong TA, et al. (2016). Wounds on Rapanea melanophloeos provide habitat for a large diversity of Ophiostomatales including four new species. Antonie van Leeuwenhoek 109: 877–894. [DOI] [PubMed] [Google Scholar]
- Nannfeldt JA. (1932). Studien uber die Morphologie und Systematik der nicht-lichenisierten inoperculaten Discomyceten. Nova Acta Regiae Societatis Scientiarum Upsaliensis IV 8: 1–368. [Google Scholar]
- Nel WJ, De Beer ZW, Wingfield MJ, et al. (2021). Phylogenetic and phylogenomic analyses reveal two new genera and three new species of ophiostomatalean fungi from termite-fungus combs. Mycologia. 113: 1199–1217. [DOI] [PubMed] [Google Scholar]
- Nel WJ, Duong TA, Wingfield BD, et al. (2018). A new genus and species for the globally important, multi-host root pathogen Thielaviopsis basicola. Plant Pathology 67: 871–882. [Google Scholar]
- Ngubane NP, Dreyer LL, Oberlander KC, et al. (2018). Two new Sporothrix species from Protea flower heads in South African Grassland and Savanna. Antonie van Leeuwenhoek 111: 965–979. [DOI] [PubMed] [Google Scholar]
- Nicot J, Mariat F. (1973). Caractères morphologiques et position systématique de Sporothrix schenckii, agent de la sporotrichose humaine. Mycopathologia et Mycologia Applicata 49: 53–65. [DOI] [PubMed] [Google Scholar]
- Okada G, Seifert KA, Takematsu A, et al. (1998). A molecular phylogenetic reappraisal of the Graphium complex based on 18S rDNA sequences. Canadian Journal of Botany 76: 1495–1506. [Google Scholar]
- Olchowecki A, Reid J. (1974) Taxonomy of the genus Ceratocystis in Manitoba. Canadian Journal of Botany 52: 1675–1711. [Google Scholar]
- Osorio JA, De Beer ZW, Wingfield MJ, et al. (2016). Ophiostomatoid fungi associated with mangroves in South Africa, including Ophiostoma palustre sp. nov. Antonie van Leeuwenhoek 109: 1555–1571. [DOI] [PubMed] [Google Scholar]
- Paciura D, De Beer ZW, Jacobs K, et al. (2010). Eight new Leptographium species associated with tree-infesting bark beetles in China. Persoonia 25: 94–108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan Y, Lu J, Zhou X-D, et al. (2020). Leptographium wushanense sp. nov., associated with Tomicus armandii on Pinus armandii in Southwestern China. Mycoscience 61: 43–48. [Google Scholar]
- Plattner A, Kim J-J, Reid J, et al. (2009). Resolving taxonomic and phylogenetic incongruence within species Ceratocystiopsis minuta. Mycologia 101: 878–887. [DOI] [PubMed] [Google Scholar]
- Poinar GO, Vega FE. (2018). A mid-Cretaceous ambrosia fungus, Paleoambrosia entomophila gen. nov. et sp. nov. (Ascomycota: Ophiostomatales) in Burmese (Myanmar) amber, and evidence for a femoral mycangium. Fungal Biology 122: 1159–1162. [DOI] [PubMed] [Google Scholar]
- Réblová M, Gams W, Seifert KA. (2011). Monilochaetes and allied genera of the Glomerellales, and a reconsideration of families in the Microascales. Studies in Mycology 68: 163–191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reid J, Hausner G. (2015). A new Graphilbum species from western hemlock (Tsuga heterophylla) in Canada. Mycotaxon 130: 399–419. [Google Scholar]
- Rodrigues AM, De Hoog GS, De Camargo ZP. (2013). Emergence of pathogenicity in the Sporothrix schenckii complex. Medical Mycology 51: 405–412. [DOI] [PubMed] [Google Scholar]
- Rodrigues AM, De Hoog GS, De Cássia Pires D, et al. (2014). Genetic diversity and antifungal susceptibility profiles in causative agents of sporotrichosis. BMC Infectious Diseases 14: 219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roets F, De Beer ZW, Dreyer LL, et al. (2006). Multi-gene phylogeny for Ophiostoma spp. reveals two new species from Protea infructescences. Studies in Mycology 55: 199–212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roets F, De Beer ZW, Wingfield MJ, et al. (2008). Ophiostoma gemellus and Sporothrix variecibatus from mites infesting Protea infructescences in South Africa. Mycologia 100: 496–510. [DOI] [PubMed] [Google Scholar]
- Roets F, Wingfield MJ, Crous PW, et al. (2013). Taxonomy and ecology of ophiostomatoid fungi associated with Protea infructescences. In: The Ophiostomatoid Fungi: Expanding Frontiers, (Seifert KA, De Beer ZW, Wingfield MJ, ed.): 177–187. CBS-KNAW Fungal Biodiversity Centre, Utrecht, The Netherlands. [Google Scholar]
- Roets F, Wingfield BD, De Beer ZW, et al. (2010). Two new Ophiostoma species from Protea caffra in Zambia. Persoonia: Molecular Phylogeny and Evolution of Fungi 24: 18–28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rollins F, Jones KG, Krokene P, et al. (2001). Phylogeny of asexual fungi associated with bark and ambrosia beetles. Mycologia 93: 991–996. [Google Scholar]
- Romón P, De Beer ZW, Fernández M, et al. (2014a). Ophiostomatoid fungi including two new fungal species associated with pine root-feeding beetles in northern Spain. Antonie van Leeuwenhoek 106: 1167–1184. [DOI] [PubMed] [Google Scholar]
- Romón P, De Beer ZW, Zhou X, et al. (2014b). Multigene phylogenies of Ophiostomataceae associated with Monterey pine bark beetles in Spain reveal three new fungal species. Mycologia 106: 119–132. [DOI] [PubMed] [Google Scholar]
- Rumbold CT. (1931) Two blue-stain fungi associated with bark-beetle infestation of pines. Journal of Agricultural Research 43: 847–873. [Google Scholar]
- Rumbold CT. (1941). A blue stain fungus, Ceratostomella montium n. sp., and some yeasts associated with two species of Dendroctonus. Journal of Agricultural Research 62: 589–601. [Google Scholar]
- Saccardo PA. (1881). Fungi Gallici lecti a cl. viris P. Brunaud, C.C. Gillet, Abb. Letendre, A. Malbranche, J. Therry vel editi in Mycotheca Gallica C. Roumeguèri. Series III. Michelia 2: 302–371. [Google Scholar]
- Saucedo-Carabez JR, Ploetz RC, Konkol JL, et al. (2018). Partnerships between ambrosia beetles and fungi: Lineage-specific promiscuity among vectors of the Laurel wilt pathogen, Raffaelea lauricola. Microbial Ecology 76: 925–940. [DOI] [PubMed] [Google Scholar]
- Schoch CL, Robbertse B, Robert V, et al. (2014). Finding needles in haystacks: linking scientific names, reference specimens and molecular data. Database: 1–21, doi: 10.1093/dababase/bau061 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schoch CL, Seifert KA, Huhndorf S, et al. (2012). Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proceedings of the National Academy of Sciences 109: 6241–6246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schroeder S, Kim SH, Cheung WT, et al. (2001). Phylogenetic relationship of Ophiostoma piliferum to other sapstain fungi based on the nuclear rRNA gene. FEMS Microbiology Letters 195: 163–167. [DOI] [PubMed] [Google Scholar]
- Seifert KA, De Beer ZW, Wingfield MJ. (2013). The ophiostomatoid fungi: expanding frontiers. CBS Biodiversity Series 12. CBS-KNAW Biodiversity Centre, Utrecht, The Netherlands. [Google Scholar]
- Seifert KA, Okada G. (1993). Graphium anamorphs of Ophiostoma species and similar anamorphs of other Ascomycetes. In: Ceratocystis and Ophiostoma: Taxonomy, Ecology and Pathogenicity, (Wingfield MJ, Seifert KA, Webber J, eds.): 27–41. American Phytopathological Society Press, St. Paul, Minnesota. [Google Scholar]
- Seo ST, Kim KH, Lee SH, et al. (2010). Genotypic characterization of Oak Wilt pathogen Raffaelea quercus-mongolicae and R. quercivora strains. Research in Plant Disease 16: 219–223. [Google Scholar]
- Seppey M, Manni M, Zdobnov EM. (2019). BUSCO: assessing genome assembly and annotation completeness. Methods in Molecular Biology 1962: 227–245. [DOI] [PubMed] [Google Scholar]
- Shear CL. (1923). Life Histories and undescribed genera and species of fungi. Mycologia 15: 120–131. [Google Scholar]
- Siemaszko W. (1939). Zespoly grzybów towarzysza¸cych kornikom polskim. Planta Polonica 7: 1–54. [Google Scholar]
- Silvestro D, Michalak I. (2012). raxmlGUI: a graphical front-end for RAxML. Organisms Diversity & Evolution 12: 335–337. [Google Scholar]
- Simmons DR, De Beer ZW, Huang Y-T, et al. (2016). New Raffaelea species (Ophiostomatales) from the USA and Taiwan associated with ambrosia beetles and plant hosts. IMA Fungus 7: 265–273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Six D, De Beer ZW, Duong TA, et al. (2011). Fungal associates of the lodgepole pine beetle, Dendroctonus murrayanae. Antonie van Leeuwenhoek 100: 231–244. [DOI] [PubMed] [Google Scholar]
- Spatafora JW, Blackwell M. (1994). The polyphyletic origins of ophiostomatoid fungi. Mycological Research 98: 1–9. [Google Scholar]
- Stamatakis A. (2014). RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30: 1312–1313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stielow JB, Lévesque CA, Seifert KA, et al. (2015). One fungus, which genes? Development and assessment of universal primers for potential secondary fungal DNA barcodes. Persoonia: Molecular Phylogeny and Evolution of Fungi 35: 242–263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suh S-O, Blackwell M. (1999). Molecular phylogeny of the cleistothecial fungi placed in Cephalothecaceae and Pseudeurotiaceae. Mycologia 91: 836–848. [Google Scholar]
- Sun J, Lu M, Gillette NE, et al. (2013). Red Turpentine beetle: innocuous native becomes invasive tree killer in China. Annual Review of Entomology 58: 293–311. [DOI] [PubMed] [Google Scholar]
- Swofford DL. (2003). PAUP* 4.0: phylogenetic analysis using parsimony (*and other methods). Sunderland, Massachusetts: Sinauer Associates. [Google Scholar]
- Taerum S, De Beer ZW, Duong TA, et al. (2012). Fungal symbionts suggest an alternative origin for the red turpentine beetle (Dendroctonus valens) invasion in China. Mycological Society of America 2012 Meeting. Yale University, New Haven, Connecticut. [Google Scholar]
- Taerum SJ, Duong TA, De Beer ZW, et al. (2013). Large shift in symbiont assemblage in the invasive red turpentine beetle. PLoS ONE 8: e78126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taerum SJ, Hoareau TB, Duong TA, et al. (2017). Putative origins of the fungus Leptographium procerum. Fungal Biology 121: 82–94. [DOI] [PubMed] [Google Scholar]
- Teixeira M, De Almeida L, Kubitschek-Barreira P, et al. (2014). Comparative genomics of the major fungal agents of human and animal Sporotrichosis: Sporothrix schenckii and Sporothrix brasiliensis. BMC Genomics 15: 943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thwaites JM, Farrell RL, Duncan SM, et al. (2005). Survey of potential sapstain fungi on Pinus radiata in New Zealand. New Zealand Journal of Botany 43: 653–663. [Google Scholar]
- Upadhyay HP. (1981). A monograph of Ceratocystis and Ceratocystiopsis. University of Georgia Press. [Google Scholar]
- Upadhyay HP, Kendrick WB. (1975). Prodromus for a revision of Ceratocystis (Microascales, Ascomycetes) and its conidial states. Mycologia 67: 798–805. [Google Scholar]
- Van der Linde J, Six D, De Beer Z, et al. (2016). Novel ophiostomatalean fungi from galleries of Cyrtogenius africus (Scolytinae) infesting dying Euphorbia ingens. Antonie van Leeuwenhoek 109: 589–601. [DOI] [PubMed] [Google Scholar]
- Van der Nest MA, Bihon W, De Vos L, et al. (2014). Draft genome sequences of Diplodia sapinea, Ceratocystis manginecans, and Ceratocystis moniliformis. IMA Fungus 5: 135–140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vanderpool D, Bracewell RR, McCutcheon JP. (2018). Know your farmer: ancient origins and multiple independent domestications of ambrosia beetle fungal cultivars. Molecular Ecology 27: 2077–2094. [DOI] [PubMed] [Google Scholar]
- Verrall AF. (1943). Fungi associated with certain ambrosia beetles. Journal of Agricultural Research 66: 135–144. [Google Scholar]
- Vilgalys R, Hester M. (1990). Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. Journal of Bacteriology 172: 4238–4246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villarreal M, Rubio V, De Troya M, et al. (2005). New Ophiostoma species isolated from Pinus pinaster in the Iberian Peninsula. Mycotaxon 92: 259–268. [Google Scholar]
- Von Arx JA. (1952). Ueber die Ascomycetengattungen Ceratostomella Sacc., Ophiostoma Syd. und Rostrella Zimmermann. Antonie van Leeuwenhoek 18: 13–213. [DOI] [PubMed] [Google Scholar]
- Von Arx JA, Hennebert GL. (1965) Deux champignons ambrosia. Mycopathologia et Mycologia Applicata 25: 309–315. [Google Scholar]
- Wang CY, Fang ZM, Sun BS, et al. (2008). High infectivity of an endoparasitic fungus strain, Esteya vermicola, against nematodes. Journal of Microbiology 46: 380–389. [DOI] [PubMed] [Google Scholar]
- Wang H, Lun Y, Lu Q, et al. (2018). Ophiostomatoid fungi associated with pines infected by Bursaphelenchusxylophilus and Monochamusalternatus in China, including three new species. MycoKeys 39: 1–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, Wang T, Wang J, et al. (2014). Morphological, molecular and biological characterization of Esteya vermicola, a nematophagous fungus isolated from intercepted wood packing materials exported from Brazil. Mycoscience 55: 367–377. [Google Scholar]
- Wang Y-B, Pang W-X, Yv X-N, et al. (2015). The effects of fluctuating culture temperature on stress tolerance and antioxidase expression in Esteya vermicola. Journal of Microbiology 53: 122–126. [DOI] [PubMed] [Google Scholar]
- Wang Z, Liu Y, Wang H, et al. (2020). Ophiostomatoid fungi associated with Ips subelongatus, including eight new species from northeastern China. IMA Fungus 11:1–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weijman AC, De Hoog GS. (1975). On the subdivision of the genus Ceratocystis. Antonie van Leeuwenhoek 41: 353–360. [DOI] [PubMed] [Google Scholar]
- White TJ, Bruns T, Lee S, et al. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR protocols: a guide to methods and applications 18: 315–322. [Google Scholar]
- Wingfield BD, Ades PK, Al-Naemi FA, et al. (2015a). Draft genome sequences of Chrysoporthe austroafricana, Diplodia scrobiculata, Fusarium nygamai, Leptographium lundbergii, Limonomyces culmigenus, Stagonosporopsis tanaceti, and Thielaviopsis punctulata. IMA Fungus 6: 233–248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wingfield BD, Ambler JM, Coetzee MPA, et al. (2016a) Draft genome sequences of Armillaria fuscipes, Ceratocystiopsis minuta, Ceratocystis adiposa, Endoconidiophora laricicola, E. polonica and Penicillium freii DAOMC 242723. IMA Fungus 7: 217–227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wingfield BD, Barnes I, De Beer ZW, et al. (2015). Draft genome sequences of Ceratocystis eucalypticola, Chrysoporthe cubensis, C. deuterocubensis, Davidsoniella virescens, Fusarium temperatum, Graphilbum fragrans, Penicillium nordicum, and Thielaviopsis musarum. IMA Fungus 6: 493–506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wingfield BD, Berger DK, Steenkamp ET, et al. (2017). Draft genome of Cercospora zeina, Fusarium pininemorale, Hawksworthiomyces lignivorus, Huntiella decipiens and Ophiostoma ips. IMA Fungus 8: 385–396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wingfield BD, Duong TA, Hammerbacher A, et al. (2016b) Draft genome sequences for Ceratocystis fagacearum, C. harringtonii, Grosmannia penicillata, and Huntiella bhutanensis. IMA Fungus 7: 317–323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wingfield BD, Liu M, Nguyen HDT, et al. (2018). Nine draft genome sequences of Claviceps purpurea s.lat., including C. arundinis, C. humidiphila, and C. cf. spartinae, pseudomolecules for the pitch canker pathogen Fusarium circinatum, draft genome of Davidsoniella eucalypti, Grosmannia galeiformis, Quambalaria eucalypti, and Teratosphaeria destructans. IMA Fungus 9: 401–418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wingfield MJ. (1983). Association of Verticicladiella procera and Leptographium terebrantis with insects in the Lake States. Canadian Journal of Forest Research 13: 1238–1245. [Google Scholar]
- Wingfield MJ. (1993). Leptographium species as anamorphs of Ophiostoma. In: Ceratocystis and Ophiostoma: Taxonomy, Ecology and Pathogenicity, (Wingfield MJ, Seifert KA, Webber J, eds.): 43–51. American Phytopathological Society Press, St. Paul, Minnesota. [Google Scholar]
- Wingfield MJ, Barnes I, De Beer ZW, et al. (2017). Novel associations between ophiostomatoid fungi, insects and tree hosts: current status—future prospects. Biological Invasions 19: 3215–3228. [Google Scholar]
- Wingfield MJ, Harrington TC, Crous PW. (1994). Three new Leptographium species associated with conifer roots in the United States. Canadian Journal of Botany 72: 227–238. [Google Scholar]
- Wingfield MJ, Marasas WFO. (1980). Verticicladiella alacris sp. nov., associated with a root disease of pines in South Africa. Transactions of the British Mycological Society 75: 21–28. [Google Scholar]
- Wingfield MJ, Seifert KA, Webber JF. (1993). Ceratocystis and Ophiostoma: taxonomy, ecology, and pathogenicity. American Phytopathological Society, St. Paul, Minnesota, USA. [Google Scholar]
- Witcosky JJ, Hansen EM. (1985). Root-colonizing insects recovered from Douglas-fir in various stages of decline due to black-stain root disease. Phytopathology 75: 399–402. [Google Scholar]
- Witcosky JJ, Schowalter TD, Hansen EM. (1986). Hylastes nigrinus (Coleoptera: Scolytidae), Pissodes fasciatus, and Steremnius carinatus (Coleoptera: Curculionidae) as Vectors of Black-stain Root Disease of Douglas-fir. Environmental Entomology 15: 1090–1095. [Google Scholar]
- Yaguchi T, Sano A, Yarita K, et al. (2006). A new species of Cephalotheca isolated from a Korean patient. Mycotaxon 96: 309–322. [Google Scholar]
- Yamaoka Y. (2017). Taxonomy and pathogenicity of ophiostomatoid fungi associated with bark beetles infesting conifers in Japan, with special reference to those related to subalpine conifers. Mycoscience 58: 221–235. [Google Scholar]
- Yamaoka Y, Wingfield MJ, Takahashi I, et al. (1997). Ophiostomatoid fungi associated with the spruce bark beetle Ips typographus f. aponicus in Japan. Mycological Research 101: 1215–1227. [Google Scholar]
- Yan Z, Sun J, Don O, et al. (2005). The red turpentine beetle, Dendroctonus valens LeConte (Scolytidae): an exotic invasive pest of pine in China. Biodiversity and Conservation 14: 1735–1760. [Google Scholar]
- Yin M, Duong TA, Wingfield MJ, et al. (2015). Taxonomy and phylogeny of the Leptographium procerum complex, including Leptographium sinense sp. nov. and Leptographium longiconidiophorum sp. nov. Antonie van Leeuwenhoek 107: 547–563. [DOI] [PubMed] [Google Scholar]
- Yin M, Wingfield MJ, Zhou X, et al. (2016). Multigene phylogenies and morphological characterization of five new Ophiostoma spp. associated with spruce-infesting bark beetles in China. Fungal Biology 120: 454–470. [DOI] [PubMed] [Google Scholar]
- Yin M, Wingfield MJ, Zhou X, et al. (2019). Taxonomy and phylogeny of the Leptographium olivaceum complex (Ophiostomatales, Ascomycota), including descriptions of six new species from China and Europe. MycoKeys 60: 93–123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yin ML, Wingfield MJ, Zhou XD, et al. (2020). Phylogenetic re-evaluation of the Grosmania penicillata complex (Ascomycota, Ophiostomatales), with the description of five new species from China and USA. Fungal Biology 124: 110–124. [DOI] [PubMed] [Google Scholar]
- Zanzot JW, De Beer ZW, Eckhardt LG, et al. (2010). A new Ophiostoma species from loblolly pine roots in the southeastern United States. Mycological Progress 9: 447–457. [Google Scholar]
- Zhang Y, Hagen F, Stielow B, et al. (2013). Global evolutionary patterns in pathogenic Sporothrix species. 1st International Meeting on Sporothrix and Sporotrichosis. Rio de Janeiro, Brazil. [Google Scholar]
- Zhou X, De Beer ZW, Cibrian D, et al. (2004a). Characterisation of Ophiostoma species associated with pine bark beetles from Mexico, including O. pulvinisporum sp. nov. Mycological Research 108: 690–698. [DOI] [PubMed] [Google Scholar]
- Zhou X, De Beer ZW, Harrington TC, et al. (2004b). Epitypification of Ophiostoma galeiforme and phylogeny of species in the O. galeiforme complex. Mycologia 96: 1306–1315. [PubMed] [Google Scholar]
- Zhou XD, De Beer ZW, Wingfield MJ. (2006). DNA sequence comparisons of Ophiostoma spp., including Ophiostoma aurorae sp. nov., associated with pine bark beetles in South Africa. Studies in Mycology 55: 269–277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou X, De Beer ZW, Wingfield M. (2013). Ophiostomatoid fungi associated with conifer-infesting bark beetles in China. In: The Ophiostomatoid Fungi: Expanding Frontiers, (Seifert KA, De Beer ZW, Wingfield MJ, ed.): 91–98. CBS-KNAW Fungal Biodiversity Centre, Utrecht, The Netherlands. [Google Scholar]
- Zipfel RD, De Beer ZW, Jacobs K, et al. (2006). Multi-gene phylogenies define Ceratocystiopsis and Grosmannia distinct from Ophiostoma. Studies in Mycology 55: 75–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Phylogenetic tree derived from maximum likelihood analysis of the LSU gene region. The dataset consisted of 233 isolates and 859 characters before and after Gblocks treatment. Bootstrap values above 60 % are shown. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks genera that we redefine here, and blue blocks indicate genera that we here re-instate and re-define.
Phylogenetic tree derived from maximum likelihood analysis of the ITS gene region. The dataset consisted of 235 isolates and 169 characters after Gblocks treatment (1 171 characters prior to Gblocks, including gaps). ML values above 60 % are shown. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks genera that we redefine here, and blue blocks indicate genera that we here re-instate and re-define.
Phylogenetic tree derived from maximum likelihood analysis of the TEF1-α gene region. The dataset consisted of 207 isolates and 380 characters after Gblocks treatment (655 characters prior to Gblocks, including gaps). ML values above 60 % are shown. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks genera that we redefine here, and blue blocks indicate genera that we here re-instate and re-define.
Phylogenetic tree derived from maximum likelihood analysis of the RPBII gene region. The dataset consisted of 171 isolates and 952 characters after Gblocks treatment (1 108 characters prior to Gblocks, including gaps). ML values above 60 % are shown. Purple blocks indicate existing genera, yellow blocks new genera described in this study, green blocks genera that we redefine here, and blue blocks indicate genera that we here re-instate and re-define.
Taxa included in the phylogenomic analyses and their genome sequence statistics