Abstract
FAT atypical cadherin 1 (FAT1) is among the most frequently mutated genes in many types of cancer. Its highest mutation rate is found in head and neck squamous cell carcinoma (HNSCC), in which FAT1 is the second most frequently mutated gene. Thus, FAT1 has great potential to serve as a target or prognostic biomarker in cancer treatment. FAT1 encodes a member of the cadherin-like protein family. Under normal physiological conditions, FAT1 serves as a molecular “brake” on mitochondrial respiration and acts as a receptor for a signaling pathway regulating cell–cell contact interaction and planar cell polarity. In many cancers, loss of FAT1 function promotes epithelial-mesenchymal transition (EMT) and the formation of cancer initiation/stem-like cells. However, in some types of cancer, overexpression of FAT1 leads to EMT. The roles of FAT1 in cancer progression, which seems to be cancer-type specific, have not been clarified. To further study the function of FAT1 in cancers, this review summarizes recent relevant literature regarding this protein. In addition to phenotypic alterations due to FAT1 mutations, several signaling pathways and tumor immune systems known or proposed to be regulated by this protein are presented. The potential impact of detecting or targeting FAT1 mutations on cancer treatment is also prospectively discussed.
Keywords: FAT1, Cancer progression, Gene mutations, Signaling regulatory network, Targeted treatment
Background
The challenges in effectively treating head and neck squamous cell carcinoma (HNSCC) are attributed to its extreme heterogeneity as far as anatomic locations and genetic aberrations [1–4]. These genetic alterations, especially gene mutations, which accumulate during the growth of cancer, create difficulties in understanding the biology of the disease and lead to ineffective and non-targeted approaches which can only go so far in altering the prognosis of patients. The Cancer Genome Atlas (TCGA) and other omics-based studies have provided the most comprehensive characterization to date of the genomic and proteomic landscape in many types of cancers. FAT atypical cadherin 1 (FAT1) is among the group of genes that is most frequently mutated in many cancers. For example, in TCGA dataset, the FAT1 mutation rate is around 10–18% in both lung adenocarcinoma (LUAD) and lung squamous cell carcinomas (LUSCC), esophageal cell carcinoma (ESCC), and cervical SCC [5]. The highest mutation rate was found to be ~ 23% in HNSCC, ranking as the second most mutated gene after TP53 in this disease and suggesting its important role in the biology of HNSCC [3, 4]. In HPV-negative (HPV-) HNSCC, the FAT1 mutation rate is as high as 28% with many truncation and nonsense mutations [6, 7], suggesting that wild-type FAT1 serves as a tumor suppressor gene in this disease [2]. These observations were also supported by some publications. Martin et al.reported that the FAT1 gene alteration rate is as high as 29.8% in HNSCC, which is the highest among solid tumors [5]. FAT1 mutation was reported to be more common in HPV-negative than in HPV-positive HNSCC (28% vs. 2.8%) [6]. In a study from Taiwan, 29% of HNSCC had FAT1 mutation [8]. Mann et al.examined 16 HNSCC cell lines and reported a FAT1 mutation rate of 43% [9]. Table 1 summaries FAT1 gene mutations in HNSCC cell lines from the CCLE Cancer Cell Line Encyclopedia (https//site.broadinstitute.org/ccle/datasets) [9–12].
Table 1.
FAT1 gene status in various HNSCC cell lines
| Cell line primary name | Variant Classification | Variant Type | Anato Class | Anatomy | Node Status | Gender | HPV |
|---|---|---|---|---|---|---|---|
| BHY | Nonsense_Mutation | SNP | OC | Alveolus | Negative | M | Negative |
| BICR 18 | Silent | SNP | LX | Larynx | Positive | M | N/A |
| Frame_Shift_Del | DEL | - | - | - | - | - | |
| Frame_Shift_Ins | INS | - | - | - | - | - | |
| BICR 31 | Nonsense_Mutation | SNP | OC | Tongue | Positive | M | N/A |
| BICR 56 | Nonsense_Mutation | SNP | OC | Tongue | Positive | F | N/A |
| BICR78 | Splice_Site | SNP | OC | Oral alveolus | N/A | M | N/A |
| CAL-33 | Nonsense_Mutation | SNP | OC | Tongue | Negative | M | Negative |
| FaDu | Frame_Shift_Del | DEL | OC | Hypopharynx | Positive | M | Negative |
| H357 | Frame_Shift_Del | DEL | OC | Tongue | N/A | M | N/A |
| Nonsense_Mutation | SNP | - | - | - | - | - | |
| H376 | Missense_Mutation | SNP | OC | Floor of mouth | N/A | F | N/A |
| HO1N1 | Nonsense_Mutation | SNP | OC | Baccul Muc | N/A | N/A | N/A |
| HO1U1 | Nonsense_Mutation | SNP | OC | Mouth Floor | N/A | N/A | N/A |
| HSC-2 | Missense_Mutation | SNP | OC | Floor of mouth | Negative | M | N/A |
| HSC-3 | Frame_Shift_Del | DEL | OC | Tongue | Positive | M | Negative |
| JHU022 | Missense_Mutation | SNP | LX | Larynx | Positive | M | Negative |
| MDA686TUa | Missense_Mutation | SNP | OP | Base of Tongue | Positive | M | Negative |
| OSC19 | Frame_Shift_Del | DEL | OC | Tongue | Positive | M | N/A |
| PE/CA-PJ41 | Missense_Mutation | SNP | OC | N/A | N/A | F | N/A |
| PE/CA-PJ49 | Nonsense_Mutation | SNP | OC | Tongue | M | N/A | |
| Missense_Mutation | SNP | - | - | - | - | - | |
| SCC-15 | Missense_Mutation | SNP | OC | Tongue | N/A | M | Negative |
| SNU-46 | Frame_Shift_Del | DEL | LX | Larynx | N/A | M | Negative |
| SNU-1041 | Missense_Mutation | SNP | OC | Pharynx | N/A | N/A | N/A |
| Nonsense_Mutation | SNP | - | - | - | - | - | |
| SqCCY1a | Frame-Shift-Ins | INS | OC | N/A | N/A | M | Negative |
| UMSCC1a | Missense_Mutation | SNP | OC | Floor of mouth | Negative | M | Negative |
| UMSCC9 | Missense_Mutation | SNP | OC | Tongue | Negative | F | Negative |
| UMSCC11A | Missense_Mutation | SNP | OC | Epiglottis | Positive | M | Negative |
| UMSCC25 | Frame-Shift-Del | DEL | LN | Lymph Node | Positive | M | Negative |
| UMSCC28 | Nonsense_Mutation | SNP | - | True cord | Negative | F | Negative |
| UMSCC41 | Missense_Mutation | SNP | - | Arytenoid | Positive | M | Negative |
| UMSCC74A | Missense_Mutation | SNP | OP | Base of Tongue | Negative | M | Negative |
| UMSCC74B | Missense_Mutation | SNP | Recurrence | Intraoral | Negative | M | Negative |
| UMSCC76 | Missense_Mutation | SNP | LN | Lymph Node | Positive | M | Negative |
| UMSCC81B | Missense_Mutation | SNP | - | True cord | Negative | M | Negative |
| UMSCC104 | Missense_Mutation | SNP | OC | Floor of mouth | Positive | M | Positive |
| UPCISCC116 | Frame_Shift_Del | DEL | OC | ALV Ridge | N/A | M | Nagative |
| YD-10B | Frame_Shift_Del | DEL | OC | Tongue | N/A | M | N/A |
Most of the information can be found in CCLE (https//site.broadinstitute.org/ccle/datasets)
a FAT1 status were determined by our group
The function of FAT1 in both normal and cancer tissues has been studied since FAT1 was discovered in Drosophila [13, 14]. However, FAT1 seems to be playing different roles in different tissues or cancer types based the finding that its expression level is upregulated in acute leukemia, hepatocellular carcinoma (HCC), glioblastoma (GMB), and gastric cancer, but downregulated in HNSCC, ESCC, breast cancer, and cervical cancer [15], though one recent study reported upregulation of FAT1 in oral cancer [16]. Particularly, the effect of FAT1 mutation on development of malignant phenotypes has not been extensively investigated, and little is known about its clinical implications. The objective of this review is to summarize the potential functions of FAT1 and its mutations in cancer progression to facilitate the development of treatments for patients harboring this specific mutated or deleted protein. HNSCC has been a focus in recent literature since this disease carries the highest FAT1 mutation rate among solid tumors [5].
Main text
Biology of normal FAT1 as an adhesion molecule
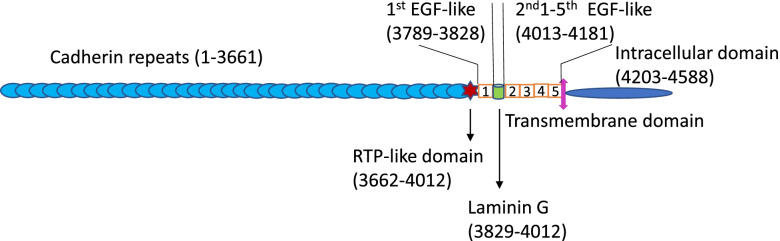
The human FAT1 gene was cloned from the human T-cell acute lymphoblastic leukemia (T-ALL) cell line in 1995 and is located on chromosome 4q34-35 and consists of 27 exons [17]. FAT1 is a cadherin-like protein family member, as a large type 1 transmembrane protein that encodes 4588 amino acid residues. It has 34 cadherin repeats, a laminin G domain, and five epidermal growth factor (EGF)-like repeats in the extracellular region, followed by a transmembrane region and a C-terminal cytoplasmic tail containing a PDZ-binding motif [18, 19] (Fig. 1). Early studies identified FAT1 as an ortholog of the Drosophila fat gene family. Under normal physiological conditions, FAT1 serves as a molecular “brake” on mitochondrial respiration [20], which regulates proliferation and migration of vascular smooth muscle cells in case of injury [21, 22]. It also acts as a receptor for a signaling pathway regulating cell–cell contact interaction and planar cell polarity [23, 24]. FAT1 has been found to be involved in the development of certain vertebrates and some hereditary diseases, such as eye abnormalities [15]. Loss of fat leads to cell cycle dysregulation and hyperproliferation in Drosophila larval imaginal discs [25]. Studies on regulation of FAT1 gene expression under normal physiological condition are limited, but two studies have identified transcriptional activator binding elements responding to NFκB and E2F1 in FAT1 gene promoter region [26, 27]. In addition to these functions in regulating normal cell activities, FAT1 plays roles in blocking or facilitating carcinogenesis and cancer progression depending on the cancer type.
Fig. 1.
Human FAT1 protein structure
FAT1 as a potential tumor suppressor
Since FAT1 is downregulated in many types of cancer and plays a role in controlling cell proliferation and migration, it has been considered a tumor suppressor. Strong evidence supporting this came from a transgenic mouse model. Pastushenko et al.performed conditional deletion of Fat1 in the skin epidermis using a Fat1-constitutive knockout (Fat1-cKO) mouse model [28]. After Fat1-cKO mice were born, DMBA/TPA was applied to the skin. The number of benign and malignant tumors per mouse was noted to increase in Fat1-cKO mice, suggesting that Fat1 acts as a tumor-suppressor gene in DMBA/TPA-induced skin SCCs. Immunohistochemistry analyses revealed that in Fat1-cKO mice, the polarity of the basal cells as well as the adherens and tight junctions were rapidly lost. In this study, the authors also confirmed their observations using combined deletion of Fat1 with p53 and kRASG12D expression in the lung epithelia by intratracheal installation of Cre-expressing adenovirus. It was found that Fat1 deletion considerably increased the number of tumors per lung tissue [28].
Tumor initiation is suggested to occur from cancer initiating cells that are also called stem-like cells. Pastushenko et al.identified stemness features in the Fat1-cKO cells, which was supported by an increased numbers of spheroids in Fat1-knockout as compared with FAT1 wild-type cells [28]. In another study, Li et al.reported that overexpression of wild-type FAT1 decreased stem-like cell markers and suppressed formation of spheroids in non-small cell lung cancer (NSCLC) cells [29]. They demonstrated that FAT1 might reduce the tumor-initiating ability in NSCLCs by promoting Yes-associated protein 1 (YAP1) nuclear-cytoplasmic translocation [29].
FAT1 as a potential cancer type-specific metastatic suppressor or promoter
In addition to the suppression of tumor initiation, FAT1 may also suppress metastasis. Multiple lines of evidence have suggested that inactivation of FAT1 results in epithelial-mesenchymal transition (EMT) through a variety of signaling pathways [28, 30–33]. In an ESCC study, overexpression of FAT1 led to inhibition of cell proliferation and colony formation, as well as cell migration and invasion. FAT1 knockdown led to a dramatic decrease in E-cadherin expression along with increased N-cadherin, vimentin, and snail mediated by MAPK/ERK signaling, while overexpression of FAT1 resulted in the opposite trends [32]. Jiang et al.reported that S100A14 suppressed proliferation and EMT in prostate cancer [30]. It turned out that S100 calcium binding protein A14 (S100A14) promoted the expression of FAT1 and activated the Hippo complex activity, which, therefore, suppressed prostate cancer progression. Depletion of FAT1 reversed the suppression of cell proliferation and EMT resulting from S100A14 overexpression in prostate cancer. These observations were confirmed in a PC3 prostate cancer xenograft mouse model [30]. More convincingly, in a Fat1-cKO mouse model, the proportion of spontaneous lymph node and lung metastases and the number of metastases per mouse were increased as compared with wild-type control. Also, intravenous injection of EPCAM+ Fat1-cKO tumor cells gave rise to a higher number of lung metastases as compared to tumor cells with wild-type FAT1, which clearly illustrated that deletion of FAT1 promotes metastasis in vivo [28].
In contrast, in some types of cancers, FAT1 may facilitate metastasis. GBM is characterized by the presence of hypoxia, stemness and local invasiveness. It was found that markers of EMT (Snail/LOX/Vimentin/N-cad), stemness (SOX2/OCT4/Nestin/REST) and hypoxia (HIF-1a/VEGF/PGK1/CA9) were upregulated in 39% of GBM tumors with significant positive correlation with the expression of FAT1, consistent with the data from TCGA. FAT1 knockdown in U87MG/A172 maintained in severe hypoxia primary glioma cultures led to significant reduction of EMT/stemness markers as compared to the controls, suggesting FAT1 is a regulator of EMT/stemness in hypoxic GBM [33]. A recent mechanistic study has identified an interaction between glypican-3 (GPC3), which is a surface heparan sulfate proteoglycan and FAT1 in HCC cells [31]. The GPC3 binding region on FAT1 was mapped to the C-terminal region (Q14517, residues 3662–4181), which covered a putative receptor tyrosine phosphatase (RTP)-like domain, a laminin G-like domain, and five EGF-like domains. Expression of both GPC3 and FAT1 in HCC were upregulated under hypoxia conditions, and thus were able to upregulate the expression of snail, vimentin, and downregulate E-cadherin, promoting HCC cell migration. However, these studies did not have supportive data from animal models of metastasis.
Function of circular FAT1
Several recent studies have identified circular FAT1 (circFAT1) in cancer cells. circFAT1 is a non-coding RNA with a cyclic structure which was initially reported in osteosarcoma in 2018 [34]. It is formed by the back-splicing of exon 2 of the FAT1 gene and head-to-tail binding. Like cell surface FAT1, circFAT1 has a dual-function, either inhibiting or promoting tumor progression in a cancer specific manner through sponging miRNAs and repressing their downstream pathways [35–40]. In ESCC, downregulation of circFAT1expression by siRNA promoted ESCC cell migration and invasive ability, but not proliferation. Consequently, the expression of miR-548 g was upregulated, which promoted ESCC cell migration and invasion [37].
In contrast, in HNSCC, Jia et al.screened 4573 circRNA in tumors and identified circFAT1 as one of 6 highly expressed circRNAs associated with shorter overall survival (OS) [35]. They further revealed that one of the mechanisms for circFAT1 to promote HNSCC progression was through binding with STAT3 and subsequently inhibiting this signaling pathway. Knockdown of circFAT1 by siRNA in HNSCC cell lines reduced tumorsphere formation in vitro and tumor growth in vivo. Interestingly, they also found that knockdown of circFAT1 significantly enhanced the efficacy of PD1 immunotherapy by enhancing CD8+ infiltration into tumor tissues. Additionally, circFAT1 expression is upregulated in HCC tissues and cells and positively correlated with TNM stage and tumor size [40]. Depletion of circFAT1 by siRNA repressed the proliferation and invasion of HCC cells in vitro and tumorigenesis in vivo. circFAT1 sponges miR-30a-5p to downregulate the expression of REEP3 and inhibits HCC proliferation and invasion, blocking hepatocarcinogenesis.
Potential contributions of FAT1 to immune regulation
Currently, there are limited publications regarding FAT1’s regulation of the cancer immune system. A notable publication is from Feng et al.who recently reported a high mutation rate of FAT1/2/3/4 (57.3%, 603/1052) in NSCLC patients [41]. They found that LUAD patients with FAT1 mutations showed significantly high infiltration of activated dendritic cells, while those with FAT2/3/4 mutations had high infiltration of CD8+T-cells, M1 macrophages, activated memory CD4+ T-cells, and helper follicular T-cells. They also found that FAT1/2/3 mutations were associated with longer progression-free survival in an immune checkpoint inhibitor (ICI)-treated NSCLC cohort. FAT1/4 mutations were related to better OS in pan-cancer patients treated with ICIs. Another study on NSCLC was consistent with their observation and further suggested that high FAT1 mutation rate is associated with high tumor mutation burden (TMB), which could be used to predict patient response to ICIs [42]. These observations were supported by a recent study on 631 melanoma and 109 NSCLC samples [43], which showed that patients with melanoma and NSCLC harboring FAT1 mutations had favorable outcomes from ICI therapy. Genomic and immunologic analysis showed that a high TMB, increased infiltration of immune-response cells, decreased infiltration of immune-suppressive cells, interferon and cell cycle-related pathways were enriched in patients with FAT1 mutations.
Investigations from fields unrelated to cancer have yielded some interesting clues regarding FAT1’s effects on the immune regulatory system. Studies from wild-type FAT1 gene transgenic mice showed downregulated gene expression of TNF-α, IL-6, NF-kB, and CCL2 as compared with non-transgenic wild-type mice, supporting that overexpression of FAT1 may downregulate these inflammatory cytokines/chemokines [44]. In contrast, mutated FAT1 may upregulate growth factors and proinflammatory cytokines, such as TGFB1 [45], IL-6 [46, 47], and FGF2 [48], consistent with our recent findings [49]. This mechanism may be mediated by YAP1, which is activated by FAT1 inactivation [5]. Activated YAP1 has been found to upregulate CCL2 in endothelial cells [50, 51].
However, a preliminary TIMER2.0 analysis of TCGA database revealed an inverse correlation of FAT1 expression with infiltration of tumor-inhibiting immune cells (e.g., monocytes and T cells) and a positive correlation with myeloid-derived suppressor cells (MDSCs) in GBM, HCC, cervical and pancreatic cancers, which is also positively correlated with TGFB1/2 expression and eventually results in immune suppression [52]. FAT1 knockdown in GBM primary cultures and cell lines led to a reduction in TGFB1/2 expression/secretion. These findings are consistent with the contribution of FAT1 to the progression in these cancers.
Abnormal signaling transduction of mutated FAT1
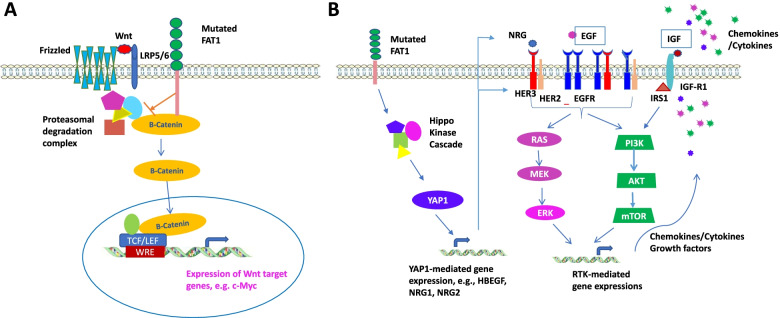
Due to its numerous biological activities in cell growth and cell–cell interaction, FAT1 is involved in the regulation of many signaling pathways. Mutation of FAT1 results in dysregulation of these signaling transductions, which potentially contributes to carcinogenesis and cancer progression (Fig. 2).
Fig. 2.
Major signaling pathways affected by mutated FAT1. A Wnt/β-catenin signaling pathway: FAT1 can bind to β-catenin. Mutated FAT1 releases β-catenin from proteasomal degradation complex, which enhances the nuclear translocation and transcriptional activity of β-catenin. B Hippo/YAP1 activation and receptor tyrosine kinase (RTK) signaling pathways: Mutated FAT1 releases YAP1 from Hippo complex and activates it as a transcription factor. Mutated FAT1 also enhances HER3 activation and IRS1 expression, which may contribute to activation of multiple RTK signaling pathways. A potential link between YAP1 and ERBB signaling may be due to an autocrine loop through their ligand EGF and NRGs
Wnt/β-catenin signaling pathways
Previous studies of the functional impact of FAT1 mutation were focused on the activation of Wnt signaling pathways. Activation of Wnt/β-catenin signaling pathway includes three steps: Wnt transducing signal at the membrane, stabilizing β-catenin in the cytoplasm, and activating Wnt/β-catenin target genes in the nucleus. FAT1 protein may affect Wnt signaling via enhancing β-catenin activity. Morris et al.reported that endogenous FAT1 bound to β-catenin in human cells [11]. In experiments using glioma cells and immortalized human brain astrocytes, they found that knockdown of FAT1 resulted in a decrease in plasma membrane β-catenin staining, and a significant increase in nuclear β-catenin translocation. Therefore, inactivated FAT1 expression could alter gene expression mediated by Wnt/β-catenin pathway components.
Hippo/YAP1 activation
The Hippo/YAP1 pathway is one of the key oncogenic regulatory pathways in multiple cancers including HNSCC [6]. FAT1 is one of the cell surface modulators involved in the regulation of YAP1 activation. A recent study by Martin et al.endorsed this observation [5]. They found that wild-type FAT1 assembled a multimeric Hippo signaling complex (signalome) which is required for activation of core Hippo kinases by TAOKs and consequent YAP1 phosphorylation. Phosphorylated YAP1 was inactive. When FAT1 was mutated, YAP1 was not restrained by phosphorylation, it then acted as an oncogenic driver in HNSCC and contributed to aggressiveness, suggesting that targeting YAP1 may serve as an attractive precision therapeutic option for cancers harboring genomic alterations in FAT1 tumor suppressor genes. In a study of ECC, Lu et al.revealed that FAT1 and PTPN14 regulated malignant progression and chemotherapy resistance through the Hippo/YAP1 signaling pathway [53]. In addition, there is reported evidence that YAP1 signaling, which was activated by FAT1 mutation [5], can activate EGFR family members though upregulation of their ligands in ovarian cancer [54], which suggests a potential linkage between YAP1 and EGFR signaling. Recently, Chen et al.reported that the YAP1/TAZ/TEAD transcriptional complex recruits BRD4 to promote an active chromatin state and regulate multiple oncogenic transcriptional programs in HNSCC [55]. Therefore, FAT1 mutated HNSCC exhibits selective and higher sensitivity to BRD4 inhibition.
EGFR family/MAPK/ERK signaling pathways
Whether the EGFR/MAPK signaling pathway is regulated by FAT1 through YAP1 has not been reported. Previous studies initially observed that in pituitary spindle cell tumor, FAT1 mutations were associated with increased ERK activity, highlighting an association between FAT1 and the ERK signaling pathway [56]. A later study in ECC also reported that knockout of FAT1 significantly increased the levels of p-ERK1/2, while overexpression of FAT1 decreased p-ERK1/2 levels [32].
In contrast, we recently reported that knocking out FAT1 in HNSCC cell lines significantly reduced p-ERK, which may be a consequence of EGFR inactivation [49]. These observations were consistent with FAT1 transgenic mouse studies [28]. Through a biostatistics/bioinformatics study using HNSCC TCGA proteomic database, we also observed increased cell surface proteins, such as HER3_pY1289, VEGFR2, and PDL1, plus IGFR signaling mediator IRS1 and cell cycle modulator CMYC in more than 90 HNSCC patient samples with FAT1 mutation. Specifically, our data showed that in both total and HPV(-) patients, HER3_pY1289 was upregulated in FAT1 mutated HNSCC. HER3 is one of the EGFR family members, with a high affinity of binding to the growth factor neuregulin, and HER3_pY1289 is the activated form that transduces signaling after partnering with other EGFR family members [57, 58]. In addition, IRS1 [59], a key regulator of IGF-1R, is also upregulated in FAT1 mutated HNSCC. Cross-talk between HER3 and IGF-R1 [60] signals could synergistically activate ERK/MAPK, PI3K/AKT, and RAS/RAF pathways and promote cell proliferation/survival, protein synthesis, and cell cycle through CMYC [61]. In contrast, similar downstream functions and biological categories are affected by different receptors and signaling molecules in tumors containing wild type FAT1. HER2_pY1248 [57] and RET_pY905 [62] are surface receptors upregulated through activated signaling molecules, including SRC_pY527 [63], SHC_pY317 [64], MTOR_pS2448 [65], and transcription factors CJUN_pS73 [66, 67], which could synergistically activate ERK/MAPK, PI3K/AKT, and RAS/RAF pathways, and promote cell proliferation/survival and protein synthesis. P16INK4A [68] and CYCLINE2 [69] regulate the cell cycle, while PAI1 [70] promotes cellular angiogenesis, migration, and invasion abilities. ASNS [71] and SCD1 [72] modulate protein and lipid metabolisms, affecting cell cycle, proliferation, and apoptosis.
Interaction between FAT1 and actin cytoskeletal dynamics
Actin dynamic is important in governing cell migration and cell–cell interaction. It has been reported that FAT1 cytoplasmic domain recruits components of actin such as Ena/VASP and Homer1/3 proteins that regulate the actin polymerization complex [13, 73, 74]. FAT1 knockdown decreased recruitment of endogenous VASP to the leading edge of the cell and resulted in weakening of lamellipodial dynamics, stopping of polarization, and controlling cell migration. Ena/VASP was found to be involved in migration of breast cancer cells [75]. Actin-mediated cellular cytoskeletal dynamics has been linked to cancer cell progression and metastasis [76, 77]. Most likely, inactivated FAT1 may regulate cancer progression and metastasis partially through the interaction with Ena/VASP and Homer-mediated cellular cytoskeletal dynamics [78].
Other cancer related proteins affected by FAT1
In our TCGA study, IPA analysis revealed the dominant upstream regulators, including TP53 and CMYC, as potentially activated transcription factors and cell surface molecules. In addition, Madan et al. revealed that under hypoxic conditions, depletion of endogenous FAT1 could reduce the expression of HIF1a and its downstream target genes such as CA9, GLUT1, VEGFA, MCT4, HK2, BNIP3 and REDD1. Consequently, a significant reduction in invasiveness was observed in GBM cells [79]. This observation is consistent with the study by Srivastava et al.who suggested FAT1 promoted EMT under hypoxia [33]. It is worth mentioning that Hayes et al.reported synergistic effects of FAT1 and CASP8 inactivation on migration and colony formation of oral cancer cell lines [80] since both are frequently mutated in this type of cancer [81]. The mechanism behind this observation remains to be determined.
Perspective: clinical significance of FAT1 as a prognostic marker or treatment target
As a frequently mutated protein in cancer, several studies have examined FAT1 as a cancer prognostic biomarker. In an earlier study, FAT1 was found to be overexpressed in paired diagnosis-relapse samples of precursor B-cell acute lymphoblastic leukemia. High FAT1 mRNA expression was correlated with shorter relapse-free and overall survival, and it was independent of other traditional prognostic markers, such as white blood cell count, sex and age in this disease [82]. In the last 10 years, several publications have reported a correlation between FAT1 mutation or expression with prognosis in different type of cancers, such as breast cancer [83], NSCLC [41], gastric cancer [84], and T-cell lymphoma [85]. Using HNSCC as an example, the Taiwanese study on HNSCC by Lin et al.showed significant correlations of FAT1 mutations with lymph node status and worse disease-free survival (DFS) [8]. Kim et a. examined 566 HNSCC patients who were classified into FAT1-associated low risk (FAT1-LR; n = 195) and FAT1-associated high risk (FAT1-HR; n = 371) subgroups. The five-year overall survival and recurrence-free survival rates were significantly lower in the FAT1-HR subgroup than in the FAT1-LR subgroup (P = 0.01 and 0.003, respectively). These results were validated using four independent cohorts [86]. It is worth mentioning that most oropharyngeal squamous cell carcinomas (OPSCCs) are HPV ( +), while FAT1 mutation occurs dominantly in HPV(-) SCC. Harbison et al.reported that metachronous recurrent OPSCCs share similar genomic features with HPV-unrelated HNSCC including FAT1 mutations [87], implicating a potential role of FAT1 mutation in recurrent HNSCC regardless of HPV status. Our recent study using TCGA proteomic database also demonstrated that HNSCC patients with FAT1 mutations had a shorter progression-free survival than those with wild-type FAT1 [49]. These studies support that FAT1 represents a promising prognostic biomarker. It is expected that a standard must be established for each cancer type based on additional studies with validations in multiple cohorts.
FAT1 expression or mutation has also been linked to cancer treatment sensitivity. Lepikhova et al.screened 45 HNSCC cell lines for sensitivity to EGFR, MEK, and mTOR inhibitors [88]. They found that cell lines harboring a stop-gain mutation in FAT1 showed a tendency for higher sensitivity to the mTOR inhibitor temsirolimus as compared with other cell lines. Pastushenko et al.examined the sensitivity of wild-type and isogenic FAT1-knockout human cancer cell lines to targeted inhibitors and found FAT1-knockout cells were significantly more resistant to the EGFR/HER2 inhibitor afatinib and MEK inhibitor trametinib as compared to FAT1 wild-type cells in vitro [28]. In contrast, FAT1-knockout tumor cells were significantly more sensitive to the SRC inhibitor dasatinib and SRC/Bcr-Abl inhibitor saracatinib and the CAMK2 inhibitor KN93 as compared to FAT1 wild-type cells. Administration of afatinib and dasatinib to mice transplanted with FAT1 wild-type and knockout human SCC cell lines showed that FAT1 wild-type tumor cells were more sensitive to afatinib and FAT1-knockout tumor cells were more sensitive to dasatinib, consistent with the differences in drug sensitivity observed in vitro. Li et al.performed genomic analysis of 348 estrogen receptor-positive breast cancer patients treated with CDK4/6 inhibitor and found that loss of function mutation of FAT1 led to resistance to the CDK4/6 inhibitor through the Hippo/YAP1 signaling pathway [89]. Interestingly, in a recently completed phase II clinical trial using a combination of HER3 inhibitor CDX-3379 and EGFR inhibitor cetuximab in recurrent/metastatic, HPV-negative, cetuximab-resistant HNSCC, the investigators analyzed tissues from 27 patients including one of two responders. They reported that the overall response rate was 1/10 (complete response; 10%; 95% CI 0.30–44.5) in the FAT1-mutated versus 0/17 (0%; 95% CI: 0–19.5) in the FAT1-wildtype cohorts, suggesting that FAT1 mutation may play a role in resistance to EGFR-targeting therapy through activation of HER3. Upon further follow up these correlations did not however hold upon completion of the trial [90]. FAT1 mutation may be an indicator for outcome in these specific combination strategies. A recent study by Zhai et al. showed that FAT1 downregulation in ESCC enhanced stemness and reduced patients’ sensitivity to cisplatin [91]. They found that knockdown of FAT1 could induce multi-drug resistant protein ABCC3 due to FAT1 mediated nuclear translocation of β-catenin. It seems that the effect of FAT1 on drug sensitivity is related to aggressive behaviors, such as stemness and EMT status of cancer cells. The contributions of FAT1 mutations to tumor and the tumor microenvironment interaction, particularly to the immune regulatory system, have not been clarified. These interactions may substantially affect tumor cell response to cancer therapy in the human body, which deserves further studies.
FAT1 is expected to define a new subclass of HNSCC. However, no agents are currently available to target FAT1 directly due to lack of fully understanding of FAT1 mutation sites and their functional alterations as compared with the wild-type protein. Our proteomic analysis has provided basic information that surface receptors and signaling molecules, such as HER3 phosphorylation, are associated with FAT1 mutation. On the bright side, some of these molecules are druggable with commercially available agents, such as HER3-DXd, a novel HER3 directed antibody drug conjugate, and Vertepofin, a YAP1 inhibitor interrupting the interaction between YAP1 and TEAD [92], which can be considered to suppress the oncogenic pathways mediated by FAT1. Kang et al.found that FAT1 was overexpressed in gastric cancer [84]. They demonstrated that verteporfin could suppress FAT1 expression, leading to decreased migration and invasion of gastric cancer cells. Recently, Gutkind’s group summarized genomic alterations in the Hippo pathway and persistent YAP/TAZ activation in HNSCC. Since the mutation frequency of FAT1 is high in this disease, blocking this pathway may provide novel multimodal precision therapies for HNSCC. We also suggest multiple strategies which can indirectly target this Hippo/YAP1 pathway [93], including use of EGFR inhibitor cetuximab, PI3K inhibitor alpelisib, MEK inhibitor trametinib, FAK inhibitor defactinib, tankyrase inhibitors, SRC inhibitor dasatinib, YAP1 inhibitor verteporfin, and smTEAD inhibitor. Most recently, Chen et al.performed comprehensive proteomic and drug-screening studies across pan-cancer models and confirmed that FAT1 mutated HNSCC had selective and higher sensitivity to BRD4 inhibition by JQ1 which has been also used to block YAP1 signaling [55]. Another approach is to disrupt the FAT1-associated protein complex by pre-designed stapled peptides. We have gained experience in developing this type of therapeutic peptide in the past [94–96] and are now working on identifying novel points of susceptibility for peptide intervention. No doubt, there is a long way to go. Still, it is imperative to further the knowledge regarding the impact of FAT1 mutations on cancer development at the molecular and therapeutic levels, as well as discover novel FAT1-targeted therapeutic strategies for personalized medicine.
Conclusion
The FAT1 mutation rate in HNSCC is the highest among major solid tumors, making its investigation of primary interest in this disease [49]. Although a high FAT1 mutation rate has been identified in many types of cancer, its functions and clinical significance remain to be further elucidated. Particularly, the current literature brings some conflict and contradiction regarding the function of this protein. Whether FAT1 serves a tumor suppressor or promoter seems to be cancer type-specific. Furthermore, since FAT1 is a gene with no identified mutation hot spots as seen for TP53, mutation-specific functional alterations of this protein remain to be discovered, which will be essential for targeting this protein in precision cancer therapy.
Acknowledgements
We would like to acknowledge Anthea Hammond for critical reading of this manuscript.
Abbreviations
- circFAT1
Circular FAT1
- DFS
Disease-free survival
- EGF
Epidermal growth factor
- EMT
Epithelial-mesenchymal transition
- ESCC
Esophageal cell carcinoma
- Fat1-cKO
Fat1-Constitutive knockout
- GMB
Glioblastoma
- GPC3
Glypican-3
- HCC
Hepatocellular carcinoma
- HNSCC
Head and neck squamous cell carcinoma
- ICI
Immune checkpoint inhibitor
- LUAD
Lung adenocarcinoma
- LUSCC
Lung squamous cell carcinomas
- NSCLC
Non-small cell lung cancer
- MDSCs
Myeloid-derived suppressor cells
- OPSCCs
Oropharyngeal squamous cell carcinomas
- OS
Overall survival
- RTK
Receptor tyrosine kinase
- RTP
Receptor tyrosine phosphatase
- S100A14
S100 calcium binding protein A14
- T-ALL
T-cell acute lymphoblastic leukemia
- TMB
Tumor mutation burden
- YAP1
Yes-associated protein 1
Authors’ contributions
ZGC and YT conceived the original idea and supervised the project. ZGC and YT wrote the manuscript. NFS provided critical feedback and helped shape the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work was supported in part by NIH/NIDCR grants R01DE028351, R03DE032084 and R03DE028387 (to YT), Winship Invest$ Team Science Award (to YT and NFS), Winship Invest$ Pilot Grant Award (to YT, ZGC and NFS), and I3 Nexus Research Award (to YT).
Availability of data and materials
Not applicable.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Cramer JD, Burtness B, Le QT, Ferris RL. The changing therapeutic landscape of head and neck cancer. Nat Rev Clin Oncol. 2019;16(11):669–683. doi: 10.1038/s41571-019-0227-z. [DOI] [PubMed] [Google Scholar]
- 2.Leemans CR, Snijders PJF, Brakenhoff RH. The molecular landscape of head and neck cancer. Nat Rev Cancer. 2018;18(5):269–282. doi: 10.1038/nrc.2018.11. [DOI] [PubMed] [Google Scholar]
- 3.Cancer Genome Atlas N Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015;517(7536):576–82. doi: 10.1038/nature14129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Campbell JD, Yau C, Bowlby R, Liu Y, Brennan K, Fan H, Taylor AM, Wang C, Walter V, Akbani R, Byers LA, Creighton CJ, Coarfa C, Shih J, Cherniack AD, Gevaert O, Prunello M, Shen H, Anur P, Chen J, Cheng H, Hayes DN, Bullman S, Pedamallu CS, Ojesina AI, Sadeghi S, Mungall KL, Robertson AG, Benz C, Schultz A, Kanchi RS, Gay CM, Hegde A, Diao L, Wang J, Ma W, Sumazin P, Chiu HS, Chen TW, Gunaratne P, Donehower L, Rader JS, Zuna R, Al-Ahmadie H, Lazar AJ, Flores ER, Tsai KY, Zhou JH, Rustgi AK, Drill E, Shen R, Wong CK, Cancer Genome Atlas Research N. Stuart JM, Laird PW, Hoadley KA, Weinstein JN, Peto M, Pickering CR, Chen Z, Van Waes C. Genomic, pathway network, and immunologic features distinguishing squamous carcinomas. Cell Rep. 2018;23(1):194–212. doi: 10.1016/j.celrep.2018.03.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Martin D, Degese MS, Vitale-Cross L, Iglesias-Bartolome R, Valera JLC, Wang Z, Feng X, Yeerna H, Vadmal V, Moroishi T, Thorne RF, Zaida M, Siegele B, Cheong SC, Molinolo AA, Samuels Y, Tamayo P, Guan KL, Lippman SM, Lyons JG, Gutkind JS. Assembly and activation of the Hippo signalome by FAT1 tumor suppressor. Nat Commun. 2018;9(1):2372. doi: 10.1038/s41467-018-04590-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Santos-de-Frutos K, Segrelles C, Lorz C. Hippo pathway and YAP signaling alterations in squamous cancer of the head and neck. J Clin Med. 2019;8(12):2131. doi: 10.3390/jcm8122131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kim KT, Kim BS, Kim JH. Association between FAT1 mutation and overall survival in patients with human papillomavirus-negative head and neck squamous cell carcinoma. Head Neck. 2016;38(Suppl 1):E2021–E2029. doi: 10.1002/hed.24372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lin SC, Lin LH, Yu SY, Kao SY, Chang KW, Cheng HW, Liu CJ. FAT1 somatic mutations in head and neck carcinoma are associated with tumor progression and survival. Carcinogenesis. 2018;39(11):1320–1330. doi: 10.1093/carcin/bgy107. [DOI] [PubMed] [Google Scholar]
- 9.Mann JE, Kulkarni A, Birkeland AC, Kafelghazal J, Eisenberg J, Jewell BM, Ludwig ML, Spector ME, Jiang H, Carey TE, Brenner JC. The molecular landscape of the University of Michigan laryngeal squamous cell carcinoma cell line panel. Head Neck. 2019;41(9):3114–3124. doi: 10.1002/hed.25803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cheng H, Yang X, Si H, Saleh AD, Xiao W, Coupar J, Gollin SM, Ferris RL, Issaeva N, Yarbrough WG, Prince ME, Carey TE, Van Waes C, Chen Z. Genomic and transcriptomic characterization links cell lines with aggressive head and neck cancers. Cell Rep. 2018;25(5):1332 e5–1345. doi: 10.1016/j.celrep.2018.10.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Morris LG, Kaufman AM, Gong Y, Ramaswami D, Walsh LA, Turcan S, Eng S, Kannan K, Zou Y, Peng L, Banuchi VE, Paty P, Zeng Z, Vakiani E, Solit D, Singh B, Ganly I, Liau L, Cloughesy TC, Mischel PS, Mellinghoff IK, Chan TA. Recurrent somatic mutation of FAT1 in multiple human cancers leads to aberrant Wnt activation. Nat Genet. 2013;45(3):253–261. doi: 10.1038/ng.2538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Brenner JC, Graham MP, Kumar B, Saunders LM, Kupfer R, Lyons RH, Bradford CR, Carey TE. Genotyping of 73 UM-SCC head and neck squamous cell carcinoma cell lines. Head Neck. 2010;32(4):417–426. doi: 10.1002/hed.21198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tanoue T, Takeichi M. New insights into Fat cadherins. J Cell Sci. 2005;118(Pt 11):2347–2353. doi: 10.1242/jcs.02398. [DOI] [PubMed] [Google Scholar]
- 14.Bryant PJ, Huettner B, Held LI, Jr, Ryerse J, Szidonya J. Mutations at the fat locus interfere with cell proliferation control and epithelial morphogenesis in Drosophila. Dev Biol. 1988;129(2):541–554. doi: 10.1016/0012-1606(88)90399-5. [DOI] [PubMed] [Google Scholar]
- 15.Peng Z, Gong Y, Liang X. Role of FAT1 in health and disease. Oncol Lett. 2021;21(5):398. doi: 10.3892/ol.2021.12659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lan T, Ge Q, Zheng K, Huang L, Yan Y, Zheng L, Lu Y, Zheng D. FAT1 upregulates in oral squamous cell carcinoma and promotes cell proliferation via cell cycle and DNA repair. Front Oncol. 2022;12:870055. doi: 10.3389/fonc.2022.870055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Dunne J, Hanby AM, Poulsom R, Jones TA, Sheer D, Chin WG, Da SM, Zhao Q, Beverley PC, Owen MJ. Molecular cloning and tissue expression of FAT, the human homologue of the Drosophila fat gene that is located on chromosome 4q34-q35 and encodes a putative adhesion molecule. Genomics. 1995;30(2):207–223. doi: 10.1006/geno.1995.9884. [DOI] [PubMed] [Google Scholar]
- 18.Katoh Y, Katoh M. Comparative integromics on FAT1, FAT2, FAT3 and FAT4. Int J Mol Med. 2006;18(3):523–8. doi: 10.3892/ijmm.18.3.523. [DOI] [PubMed] [Google Scholar]
- 19.Sadeqzadeh E, de Bock CE, Zhang XD, Shipman KL, Scott NM, Song C, Yeadon T, Oliveira CS, Jin B, Hersey P, Boyd AW, Burns GF, Thorne RF. Dual processing of FAT1 cadherin protein by human melanoma cells generates distinct protein products. J Biol Chem. 2011;286(32):28181–28191. doi: 10.1074/jbc.M111.234419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cao LL, Riascos-Bernal DF, Chinnasamy P, Dunaway CM, Hou R, Pujato MA, O'Rourke BP, Miskolci V, Guo L, Hodgson L, Fiser A, Sibinga NE. Control of mitochondrial function and cell growth by the atypical cadherin Fat1. Nature. 2016;539(7630):575–578. doi: 10.1038/nature20170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Riascos-Bernal DF, Maira A, Sibinga NES. The atypical cadherin FAT1 limits mitochondrial respiration and proliferation of vascular smooth muscle cells. Front Cardiovasc Med. 2022;9:905717. doi: 10.3389/fcvm.2022.905717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hou R, Liu L, Anees S, Hiroyasu S, Sibinga NE. The Fat1 cadherin integrates vascular smooth muscle cell growth and migration signals. J Cell Biol. 2006;173(3):417–429. doi: 10.1083/jcb.200508121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tanoue T, Takeichi M. Mammalian Fat1 cadherin regulates actin dynamics and cell-cell contact. J Cell Biol. 2004;165(4):517–528. doi: 10.1083/jcb.200403006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Saburi S, Hester I, Goodrich L, McNeill H. Functional interactions between Fat family cadherins in tissue morphogenesis and planar polarity. Development. 2012;139(10):1806–1820. doi: 10.1242/dev.077461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Mahoney PA, Weber U, Onofrechuk P, Biessmann H, Bryant PJ, Goodman CS. The fat tumor suppressor gene in Drosophila encodes a novel member of the cadherin gene superfamily. Cell. 1991;67(5):853–868. doi: 10.1016/0092-8674(91)90359-7. [DOI] [PubMed] [Google Scholar]
- 26.Srivastava C, Irshad K, Gupta Y, Sarkar C, Suri A, Chattopadhyay P, Sinha S, Chosdol K. NFsmall ka, CyrillicB is a critical transcriptional regulator of atypical cadherin FAT1 in glioma. BMC Cancer. 2020;20(1):62. doi: 10.1186/s12885-019-6435-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wang Y, Wang G, Ma Y, Teng J, Wang Y, Cui Y, Dong Y, Shao S, Zhan Q, Liu X. FAT1, a direct transcriptional target of E2F1, suppresses cell proliferation, migration and invasion in esophageal squamous cell carcinoma. Chin J Cancer Res. 2019;31(4):609–619. doi: 10.21147/j.issn.1000-9604.2019.04.05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Pastushenko I, Mauri F, Song Y, de Cock F, Meeusen B, Swedlund B, Impens F, Van Haver D, Opitz M, Thery M, Bareche Y, Lapouge G, Vermeersch M, Van Eycke YR, Balsat C, Decaestecker C, Sokolow Y, Hassid S, Perez-Bustillo A, Agreda-Moreno B, Rios-Buceta L, Jaen P, Redondo P, Sieira-Gil R, Millan-Cayetano JF, Sanmatrtin O, D'Haene N, Moers V, Rozzi M, Blondeau J, Lemaire S, Scozzaro S, Janssens V, De Troya M, Dubois C, Perez-Morga D, Salmon I, Sotiriou C, Helmbacher F, Blanpain C. Fat1 deletion promotes hybrid EMT state, tumour stemness and metastasis. Nature. 2021;589(7842):448–455. doi: 10.1038/s41586-020-03046-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Li M, Zhong Y, Wang M. Fat1 suppresses the tumor-initiating ability of nonsmall cell lung cancer cells by promoting Yes-associated protein 1 nuclear-cytoplasmic translocation. Environ Toxicol. 2021;36(11):2333–2341. doi: 10.1002/tox.23347. [DOI] [PubMed] [Google Scholar]
- 30.Jiang S, Zhu Y, Chen Z, Huang Z, Liu B, Xu Y, Li Z, Lin Z, Li M. S100A14 inhibits cell growth and epithelial-mesenchymal transition (EMT) in prostate cancer through FAT1-mediated Hippo signaling pathway. Hum Cell. 2021;34(4):1215–1226. doi: 10.1007/s13577-021-00538-8. [DOI] [PubMed] [Google Scholar]
- 31.Meng P, Zhang YF, Zhang W, Chen X, Xu T, Hu S, Liang X, Feng M, Yang X, Ho M. Identification of the atypical cadherin FAT1 as a novel glypican-3 interacting protein in liver cancer cells. Sci Rep. 2021;11(1):40. doi: 10.1038/s41598-020-79524-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hu X, Zhai Y, Kong P, Cui H, Yan T, Yang J, Qian Y, Ma Y, Wang F, Li H, Cheng C, Zhang L, Jia Z, Li Y, Yang B, Xu E, Wang J, Yang J, Bi Y, Chang L, Wang Y, Zhang Y, Song B, Li G, Shi R, Liu J, Zhang M, Cheng X, Cui Y. FAT1 prevents epithelial mesenchymal transition (EMT) via MAPK/ERK signaling pathway in esophageal squamous cell cancer. Cancer Lett. 2017;397:83–93. doi: 10.1016/j.canlet.2017.03.033. [DOI] [PubMed] [Google Scholar]
- 33.Srivastava C, Irshad K, Dikshit B, Chattopadhyay P, Sarkar C, Gupta DK, Sinha S, Chosdol K. FAT1 modulates EMT and stemness genes expression in hypoxic glioblastoma. Int J Cancer. 2018;142(4):805–812. doi: 10.1002/ijc.31092. [DOI] [PubMed] [Google Scholar]
- 34.Liu G, Huang K, Jie Z, Wu Y, Chen J, Chen Z, Fang X, Shen S. CircFAT1 sponges miR-375 to promote the expression of Yes-associated protein 1 in osteosarcoma cells. Mol Cancer. 2018;17(1):170. doi: 10.1186/s12943-018-0917-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jia L, Wang Y, Wang CY. circFAT1 Promotes Cancer Stemness and Immune Evasion by Promoting STAT3 Activation. Adv Sci (Weinh) 2021;8(13):2003376. doi: 10.1002/advs.202003376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ye Y, Ke Y, Liu L, Xiao T, Yu J. CircRNA FAT1 regulates osteoblastic differentiation of periodontal ligament stem cells via miR-4781-3p/SMAD5 pathway. Stem Cells Int. 2021;2021:5177488. doi: 10.1155/2021/5177488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Takaki W, Konishi H, Shoda K, Arita T, Kataoka S, Shibamoto J, Furuke H, Takabatake K, Shimizu H, Komatsu S, Shiozaki A, Fujiwara H, Masuda K, Otsuji E. Significance of circular FAT1 as a prognostic factor and tumor suppressor for esophageal squamous cell carcinoma. Ann Surg Oncol. 2021;28(13):8508–8518. doi: 10.1245/s10434-021-10089-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hu B, Xian Z, Zou Q, Zhang D, Su D, Yao J, Ren D. CircFAT1 suppresses colorectal cancer development through regulating miR-520b/UHRF1 Axis or miR-302c-3p/UHRF1 axis. Cancer Biother Radiopharm. 2021;36(1):45–57. doi: 10.1089/cbr.2019.3291. [DOI] [PubMed] [Google Scholar]
- 39.Gu H, Cheng X, Xu J, Zhou K, Bian C, Chen G, Yin X. Circular RNA circFAT1(e2) promotes osteosarcoma progression and metastasis by sponging miR-181b and regulating HK2 expression. Biomed Res Int. 2020;2020:3589871. doi: 10.1155/2020/3589871. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 40.Wei H, Yan S, Hui Y, Liu Y, Guo H, Li Q, Li J, Chang Z. CircFAT1 promotes hepatocellular carcinoma progression via miR-30a-5p/REEP3 pathway. J Cell Mol Med. 2020;24(24):14561–14570. doi: 10.1111/jcmm.16085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Feng Z, Yin Y, Liu B, Zheng Y, Shi D, Zhang H, Qin J. Prognostic and immunological role of FAT family genes in non-small cell lung cancer. Cancer Control. 2022;29:10732748221076682. doi: 10.1177/10732748221076682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Fang W, Ma Y, Yin JC, Hong S, Zhou H, Wang A, Wang F, Bao H, Wu X, Yang Y, Huang Y, Zhao H, Shao YW, Zhang L. Comprehensive genomic profiling identifies novel genetic predictors of response to Anti-PD-(L)1 therapies in non-small cell lung cancer. Clin Cancer Res. 2019;25(16):5015–5026. doi: 10.1158/1078-0432.CCR-19-0585. [DOI] [PubMed] [Google Scholar]
- 43.Zhang WT, Gao Y, Kong Y, Shi Y, Shen F, Wang C, Wang SQ. Favorable immune checkpoint inhibitor outcome of patients with melanoma and NSCLC harboring FAT1 mutations. NPJ Precis Oncol. 2022;6:46. doi: 10.1038/s41698-022-00292-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Fang X, Ge K, Song C, Ge Y, Zhang J. Effects of n-3PUFAs on autophagy and inflammation of hypothalamus and body weight in mice. Biochem Biophys Res Commun. 2018;501(4):927–932. doi: 10.1016/j.bbrc.2018.05.084. [DOI] [PubMed] [Google Scholar]
- 45.Cohen J, Chen Z, Lu SL, Yang XP, Arun P, Ehsanian R, Brown MS, Lu H, Yan B, Diallo O, Wang XJ, Van Waes C. Attenuated transforming growth factor beta signaling promotes nuclear factor-kappaB activation in head and neck cancer. Cancer Res. 2009;69(8):3415–3424. doi: 10.1158/0008-5472.CAN-08-3704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Johnson DE, O'Keefe RA, Grandis JR. Targeting the IL-6/JAK/STAT3 signalling axis in cancer. Nat Rev Clin Oncol. 2018;15(4):234–248. doi: 10.1038/nrclinonc.2018.8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Hong SH, Ondrey FG, Avis IM, Chen Z, Loukinova E, Cavanaugh PF, Jr, Van Waes C, Mulshine JL. Cyclooxygenase regulates human oropharyngeal carcinomas via the proinflammatory cytokine IL-6: a general role for inflammation? FASEB J. 2000;14(11):1499–1507. doi: 10.1096/fj.14.11.1499. [DOI] [PubMed] [Google Scholar]
- 48.Ornitz DM, Marie PJ. Fibroblast growth factor signaling in skeletal development and disease. Genes Dev. 2015;29(14):1463–1486. doi: 10.1101/gad.266551.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Chen Z, Zhang C, Chen J, Wang D, Tu J, Van Waes C, Saba NF, Chen ZG, Chen Z. The proteomic landscape of growth factor signaling networks associated with FAT1 mutations in head and neck cancers. Cancer Res. 2021;81(17):4402–4416. doi: 10.1158/0008-5472.CAN-20-3659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Young K, Tweedie E, Conley B, Ames J, FitzSimons M, Brooks P, Liaw L, Vary CP. BMP9 crosstalk with the hippo pathway regulates endothelial cell matricellular and chemokine responses. PLoS ONE. 2015;10(4):e0122892. doi: 10.1371/journal.pone.0122892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Xu S, Koroleva M, Yin M, Jin ZG. Atheroprotective laminar flow inhibits Hippo pathway effector YAP in endothelial cells. Transl Res. 2016;176(18–28):e2. doi: 10.1016/j.trsl.2016.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Irshad K, Srivastava C, Malik N, Arora M, Gupta Y, Goswami S, Sarkar C, Suri V, Mahajan S, Gupta DK, Suri A, Chattopadhyay P, Sinha S, Chosdol K. Upregulation of atypical cadherin FAT1 promotes an immunosuppressive tumor microenvironment via TGF-beta. Front Immunol. 2022;13:813888. doi: 10.3389/fimmu.2022.813888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Lu Y, Wang Z, Zhou L, Ma Z, Zhang J, Wu Y, Shao Y, Yang Y. FAT1 and PTPN14 regulate the malignant progression and chemotherapy resistance of esophageal cancer through the hippo signaling pathway. Anal Cell Pathol (Amst) 2021;2021:9290372. doi: 10.1155/2021/9290372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.He C, Lv X, Hua G, Lele SM, Remmenga S, Dong J, Davis JS, Wang C. YAP forms autocrine loops with the ERBB pathway to regulate ovarian cancer initiation and progression. Oncogene. 2015;34(50):6040–6054. doi: 10.1038/onc.2015.52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Chen N, Golczer G, Ghose S, Lin B, Langenbucher A, Webb J, Bhanot H, Abt NB, Lin D, Varvares M, Sattler M, Egloff AM, Joh R, Uppaluri R, Emerick KS, Lawrence MS, Saladi SV. YAP1 maintains active chromatin state in head and neck squamous cell carcinomas that promotes tumorigenesis through cooperation with BRD4. Cell Rep. 2022;39(11):110970. doi: 10.1016/j.celrep.2022.110970. [DOI] [PubMed] [Google Scholar]
- 56.Miller MB, Bi WL, Ramkissoon LA, Kang YJ, Abedalthagafi M, Knoff DS, Agarwalla PK, Wen PY, Reardon DA, Alexander BM, Laws ER, Jr, Dunn IF, Beroukhim R, Ligon KL, Ramkissoon SH. MAPK activation and HRAS mutation identified in pituitary spindle cell oncocytoma. Oncotarget. 2016;7(24):37054–37063. doi: 10.18632/oncotarget.9244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Yarden Y, Sliwkowski MX. Untangling the ErbB signalling network. Nat Rev Mol Cell Biol. 2001;2(2):127–137. doi: 10.1038/35052073. [DOI] [PubMed] [Google Scholar]
- 58.Jiang N, Saba NF, Chen ZG. Advances in targeting HER3 as an anticancer therapy. Chemother Res Pract. 2012;2012:817304. doi: 10.1155/2012/817304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Gual P, Le Marchand-Brustel Y, Tanti JF. Positive and negative regulation of insulin signaling through IRS-1 phosphorylation. Biochimie. 2005;87(1):99–109. doi: 10.1016/j.biochi.2004.10.019. [DOI] [PubMed] [Google Scholar]
- 60.Iams WT, Lovly CM. Molecular pathways: clinical applications and future direction of insulin-like growth factor-1 receptor pathway blockade. Clin Cancer Res. 2015;21(19):4270–4277. doi: 10.1158/1078-0432.CCR-14-2518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Robinson AM, Rathore R, Redlich NJ, Adkins DR, VanArsdale T, Van Tine BA, Michel LS. Cisplatin exposure causes c-Myc-dependent resistance to CDK4/6 inhibition in HPV-negative head and neck squamous cell carcinoma. Cell Death Dis. 2019;10(11):867. doi: 10.1038/s41419-019-2098-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Prazeres H, Torres J, Rodrigues F, Couto JP, Vinagre J, Sobrinho-Simoes M, Soares P. How to treat a signal? current basis for RET-genotype-oriented choice of kinase inhibitors for the treatment of medullary thyroid cancer. J Thyroid Res. 2011;2011:678357. doi: 10.4061/2011/678357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Parsons SJ, Parsons JT. Src family kinases, key regulators of signal transduction. Oncogene. 2004;23(48):7906–7909. doi: 10.1038/sj.onc.1208160. [DOI] [PubMed] [Google Scholar]
- 64.Ahmed SBM, Prigent SA. Insights into the shc family of adaptor proteins. J Mol Signal. 2017;12:2. doi: 10.5334/1750-2187-12-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Vander Broek R, Mohan S, Eytan DF, Chen Z, Van Waes C. The PI3K/Akt/mTOR axis in head and neck cancer: functions, aberrations, cross-talk, and therapies. Oral Dis. 2015;21(7):815–825. doi: 10.1111/odi.12206. [DOI] [PubMed] [Google Scholar]
- 66.Eferl R, Wagner EF. AP-1: a double-edged sword in tumorigenesis. Nat Rev Cancer. 2003;3(11):859–868. doi: 10.1038/nrc1209. [DOI] [PubMed] [Google Scholar]
- 67.Wagner K, Couillard-Despres S, Lehner B, Brockhoff G, Rivera FJ, Blume A, Neumann I, Aigner L. Prolactin induces MAPK signaling in neural progenitors without alleviating glucocorticoid-induced inhibition of in vitro neurogenesis. Cell Physiol Biochem. 2009;24(5–6):397–406. doi: 10.1159/000257432. [DOI] [PubMed] [Google Scholar]
- 68.Coordes A, Lenz K, Qian X, Lenarz M, Kaufmann AM, Albers AE. Meta-analysis of survival in patients with HNSCC discriminates risk depending on combined HPV and p16 status. Eur Arch Otorhinolaryngol. 2016;273(8):2157–2169. doi: 10.1007/s00405-015-3728-0. [DOI] [PubMed] [Google Scholar]
- 69.Ansari SS, Sharma AK, Zepp M, Ivanova E, Bergmann F, Konig R, Berger MR. Upregulation of cell cycle genes in head and neck cancer patients may be antagonized by erufosine's down regulation of cell cycle processes in OSCC cells. Oncotarget. 2018;9(5):5797–5810. doi: 10.18632/oncotarget.23537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Pavon MA, Arroyo-Solera I, Cespedes MV, Casanova I, Leon X, Mangues R. uPA/uPAR and SERPINE1 in head and neck cancer: role in tumor resistance, metastasis, prognosis and therapy. Oncotarget. 2016;7(35):57351–57366. doi: 10.18632/oncotarget.10344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Balasubramanian MN, Butterworth EA, Kilberg MS. Asparagine synthetase: regulation by cell stress and involvement in tumor biology. Am J Physiol Endocrinol Metab. 2013;304(8):E789–E799. doi: 10.1152/ajpendo.00015.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Dobrzyn P, Jazurek M, Dobrzyn A. Stearoyl-CoA desaturase and insulin signaling–what is the molecular switch? Biochim Biophys Acta. 2010;1797(6–7):1189–1194. doi: 10.1016/j.bbabio.2010.02.007. [DOI] [PubMed] [Google Scholar]
- 73.Moeller MJ, Soofi A, Braun GS, Li X, Watzl C, Kriz W, Holzman LB. Protocadherin FAT1 binds Ena/VASP proteins and is necessary for actin dynamics and cell polarization. EMBO J. 2004;23(19):3769–3779. doi: 10.1038/sj.emboj.7600380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Schreiner D, Muller K, Hofer HW. The intracellular domain of the human protocadherin hFat1 interacts with Homer signalling scaffolding proteins. FEBS Lett. 2006;580(22):5295–5300. doi: 10.1016/j.febslet.2006.08.079. [DOI] [PubMed] [Google Scholar]
- 75.Carmona G, Perera U, Gillett C, Naba A, Law AL, Sharma VP, Wang J, Wyckoff J, Balsamo M, Mosis F, De Piano M, Monypenny J, Woodman N, McConnell RE, Mouneimne G, Van Hemelrijck M, Cao Y, Condeelis J, Hynes RO, Gertler FB, Krause M. Lamellipodin promotes invasive 3D cancer cell migration via regulated interactions with Ena/VASP and SCAR/WAVE. Oncogene. 2016;35(39):5155–5169. doi: 10.1038/onc.2016.47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Biber G, Ben-Shmuel A, Sabag B, Barda-Saad M. Actin regulators in cancer progression and metastases: From structure and function to cytoskeletal dynamics. Int Rev Cell Mol Biol. 2020;356:131–196. doi: 10.1016/bs.ircmb.2020.05.006. [DOI] [PubMed] [Google Scholar]
- 77.Brandhagen BN, Tieszen CR, Ulmer TM, Tracy MS, Goyeneche AA, Telleria CM. Cytostasis and morphological changes induced by mifepristone in human metastatic cancer cells involve cytoskeletal filamentous actin reorganization and impairment of cell adhesion dynamics. BMC Cancer. 2013;13:35. doi: 10.1186/1471-2407-13-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Zhang X, Liu J, Liang X, Chen J, Hong J, Li L, He Q, Cai X. History and progression of Fat cadherins in health and disease. Onco Targets Ther. 2016;9:7337–7343. doi: 10.2147/OTT.S111176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Madan E, Dikshit B, Gowda SH, Srivastava C, Sarkar C, Chattopadhyay P, Sinha S, Chosdol K. FAT1 is a novel upstream regulator of HIF1alpha and invasion of high grade glioma. Int J Cancer. 2016;139(11):2570–2582. doi: 10.1002/ijc.30386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Hayes TF, Benaich N, Goldie SJ, Sipila K, Ames-Draycott A, Cai W, Yin G, Watt FM. Integrative genomic and functional analysis of human oral squamous cell carcinoma cell lines reveals synergistic effects of FAT1 and CASP8 inactivation. Cancer Lett. 2016;383(1):106–114. doi: 10.1016/j.canlet.2016.09.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Pickering CR, Zhang J, Yoo SY, Bengtsson L, Moorthy S, Neskey DM, Zhao M, Ortega Alves MV, Chang K, Drummond J, Cortez E, Xie TX, Zhang D, Chung W, Issa JP, Zweidler-McKay PA, Wu X, El-Naggar AK, Weinstein JN, Wang J, Muzny DM, Gibbs RA, Wheeler DA, Myers JN, Frederick MJ. Integrative genomic characterization of oral squamous cell carcinoma identifies frequent somatic drivers. Cancer Discov. 2013;3(7):770–781. doi: 10.1158/2159-8290.CD-12-0537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.de Bock CE, Ardjmand A, Molloy TJ, Bone SM, Johnstone D, Campbell DM, Shipman KL, Yeadon TM, Holst J, Spanevello MD, Nelmes G, Catchpoole DR, Lincz LF, Boyd AW, Burns GF, Thorne RF. The Fat1 cadherin is overexpressed and an independent prognostic factor for survival in paired diagnosis-relapse samples of precursor B-cell acute lymphoblastic leukemia. Leukemia. 2012;26(5):918–926. doi: 10.1038/leu.2011.319. [DOI] [PubMed] [Google Scholar]
- 83.Wang L, Lyu S, Wang S, Shen H, Niu F, Liu X, Liu J, Niu Y. Loss of FAT1 during the progression from DCIS to IDC and predict poor clinical outcome in breast cancer. Exp Mol Pathol. 2016;100(1):177–183. doi: 10.1016/j.yexmp.2015.12.012. [DOI] [PubMed] [Google Scholar]
- 84.Kang MH, Jeong GS, Smoot DT, Ashktorab H, Hwang CM, Kim BS, Kim HS, Park YY. Verteporfin inhibits gastric cancer cell growth by suppressing adhesion molecule FAT1. Oncotarget. 2017;8(58):98887–98897. doi: 10.18632/oncotarget.21946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Laginestra MA, Cascione L, Motta G, Fuligni F, Agostinelli C, Rossi M, Sapienza MR, Righi S, Broccoli A, Indio V, Melle F, Tabanelli V, Calleri A, Novero D, Facchetti F, Inghirami G, Sabattini E, Bertoni F, Pileri SA. Whole exome sequencing reveals mutations in FAT1 tumor suppressor gene clinically impacting on peripheral T-cell lymphoma not otherwise specified. Mod Pathol. 2020;33(2):179–187. doi: 10.1038/s41379-019-0279-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Kim SI, Woo SR, Noh JK, Lee MK, Lee YC, Lee JW, Ko SG, Eun YG. Clinical significance of FAT1 gene mutation and mRNA expression in patients with head and neck squamous cell carcinoma. Mol Oncol. 2021;16(8):1661–1679. doi: 10.1002/1878-0261.13171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Harbison RA, Kubik M, Konnick EQ, Zhang Q, Lee SG, Park H, Zhang J, Carlson CS, Chen C, Schwartz SM, Rodriguez CP, Duvvuri U, Mendez E. The mutational landscape of recurrent versus nonrecurrent human papillomavirus-related oropharyngeal cancer. JCI Insight. 2018;3(14):e99327. doi: 10.1172/jci.insight.99327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Lepikhova T, Karhemo PR, Louhimo R, Yadav B, Murumagi A, Kulesskiy E, Kivento M, Sihto H, Grenman R, Syrjanen SM, Kallioniemi O, Aittokallio T, Wennerberg K, Joensuu H, Monni O. Drug-sensitivity screening and genomic characterization of 45 HPV-negative head and neck carcinoma cell lines for novel biomarkers of drug efficacy. Mol Cancer Ther. 2018;17(9):2060–2071. doi: 10.1158/1535-7163.MCT-17-0733. [DOI] [PubMed] [Google Scholar]
- 89.Li Z, Razavi P, Li Q, Toy W, Liu B, Ping C, Hsieh W, Sanchez-Vega F, Brown DN, Da Cruz Paula AF, Morris L, Selenica P, Eichenberger E, Shen R, Schultz N, Rosen N, Scaltriti M, Brogi E, Baselga J, Reis-Filho JS, Chandarlapaty S. Loss of the FAT1 tumor suppressor promotes resistance to CDK4/6 Inhibitors via the hippo pathway. Cancer Cell. 2018;34(6):893–905 e8. doi: 10.1016/j.ccell.2018.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Bauman JE, Julian R, Saba NF, Wise-Draper TM, Adkins DR, O'Brien P, Fidler MJ, Gibson MK, Duvvuri U, Heath-Chiozzi M, Alvarado D, Gedrich R, Golden P, Cohen RB. Phase II trial of CDX-3379 and cetuximab in recurrent/metastatic, HPV-negative, cetuximab-resistant head and neck cancer. Cancers (Basel) 2022;14(10):2355. doi: 10.3390/cancers14102355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Zhai Y, Shan C, Zhang H, Kong P, Zhang L, Wang Y, Hu X, Cheng X. FAT1 downregulation enhances stemness and cisplatin resistance in esophageal squamous cell carcinoma. Mol Cell Biochem. 2022 doi: 10.1007/s11010-022-04475-4. [DOI] [PubMed] [Google Scholar]
- 92.Liu-Chittenden Y, Huang B, Shim JS, Chen Q, Lee SJ, Anders RA, Liu JO, Pan D. Genetic and pharmacological disruption of the TEAD-YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 2012;26(12):1300–1305. doi: 10.1101/gad.192856.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Faraji F, Ramirez SI, Anguiano Quiroz PY, Mendez-Molina AN, Gutkind JS. Genomic hippo pathway alterations and persistent YAP/TAZ Activation: new hallmarks in head and neck cancer. Cells. 2022;11(8):1370. doi: 10.3390/cells11081370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Teng Y, Bahassan A, Dong D, Hanold LE, Ren X, Kennedy EJ, Cowell JK. Targeting the WASF3-CYFIP1 complex using stapled peptides suppresses cancer cell invasion. Cancer Res. 2016;76(4):965–973. doi: 10.1158/0008-5472.CAN-15-1680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Teng Y, Qin H, Bahassan A, Bendzunas NG, Kennedy EJ, Cowell JK. The WASF3-NCKAP1-CYFIP1 complex is essential for breast cancer metastasis. Cancer Res. 2016;76(17):5133–5142. doi: 10.1158/0008-5472.CAN-16-0562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Xie X, Gao L, Shull AY, Teng Y. Stapled peptides: providing the best of both worlds in drug development. Future Med Chem. 2016;8(16):1969–1980. doi: 10.4155/fmc-2016-0102. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.