Abstract
The Pharmacogene Variation Consortium (PharmVar) catalogs star (*) allele nomenclature for the polymorphic human CYP3A5 gene. Genetic variation within the CYP3A5 gene locus impacts the metabolism of several clinically important drugs, including the immunosuppressants tacrolimus, sirolimus, cyclosporine, and the benzodiazepine midazolam. Variable CYP3A5 activity is of clinical importance regarding tacrolimus metabolism. This GeneFocus provides a CYP3A5 gene summary with a focus on aspects regarding standardized nomenclature. In addition, this review also summarizes recent changes and updates including the retirement of several allelic variants and provides an overview of how PharmVar CYP3A5 star allele nomenclature is utilized by the Pharmacogenomics Knowledgebase (PharmGKB) and the Clinical Pharmacogenetics Implementation Consortium (CPIC).
A Brief History of CYP3A5 and its Nomenclature
In 1989, a cytochrome P450 enzyme related to CYP3A4 was purified from human fetal liver samples (1). In that same year, the full-length cDNA sequence for this enzyme (the gene now known as CYP3A5) was published (2). These studies revealed that this newly described enzyme was only expressed in 10–20% of livers derived from European ancestry samples. In 1996, a study investigating the genetics underlying the variable expression of CYP3A5 suggested that a nonsynonymous variant in exon 11 (defined at the time as CYP3A5*2) segregated with the absence of the protein (3); however, the role of this variant remained inconclusive. Finally, in 2001, two studies by Kuehl et al. and Hustert et al. (4, 5) uncovered the presence of a variant in intron 3 of CYP3A5 (rs776746) (5), which creates a cryptic splice site resulting in aberrant splicing and the creation of a premature stop codon that results in transcript degradation. This allele, now known as CYP3A5*3, explained the highly variable expression of CYP3A5 in the liver. Based on homozygosity for the CYP3A5*3 allele, individuals were divided into CYP3A5 non-expressors (CYP3A5*3/*3) and CYP3A5 expressors (CYP3A5*1/*3 and CYP3A5*1/*1). CYP3A5*3 is the most common allele in European and Asian populations, but it is the minor allele in people of African ancestry. In addition to CYP3A5*3, two additional variants abolishing the function of CYP3A5 were identified and subsequently designated as CYP3A5*6 (5) and CYP3A5*7 (4). Differences in CYP3A5*3, *6 and *7 across populations explained why CYP3A5 is only expressed in 10–30% of Europeans and Asians but approximately 70% in people of African ancestry (5).
CYP3A5 star (*) allele nomenclature was first devised by the Human Cytochrome P450 Allele Nomenclature Database and has since been widely utilized including clinical genetic testing laboratories, the Pharmacogenomics Knowledgebase (PharmGKB), and the Clinical Pharmacogenetics Implementation Consortium (CPIC). CYP3A5 was transitioned to the Pharmacogene Variation (PharmVar) Consortium in 2017 (6) and introduced into the interactive PharmVar database in August 2020.
Nine major CYP3A5 haplotypes, CYP3A5*1 through *9, and several so-called suballeles were cataloged before the gene was transitioned to PharmVar (the CYP3A5 original content before it transitioned to PharmVar remains accessible through the ‘Archive’ link on the PharmVar homepage (7)). While the common CYP3A5*3 allele has been well defined, CYP3A5*2 and *4 through *9 were based on ambiguous genetic information that raised concerns of whether their respective allele defining variants indeed occur by themselves or rather exist in cis (on the same haplotype) with the intron 3 variant that defines the nonfunctional CYP3A5*3 allele. Furthermore, 5′ and 3′ flanking regions were not consistently included when a haplotype was first reported or inferred.
Resources cited throughout this review are summarized in Table 1.
Table 1.
Online CYP3A5 Resources - Links to Sites and Online Resources Referenced Throughout the Review
| Sources | References |
|---|---|
| PharmVar | |
|
CYP3A5 Gene Page Read Me Document Change Log Document Structural Variation Document Other Documents (Allele Frequency/Genotype Reporting Templates) |
(73) |
| Standards | (94) |
| Allele Designation and Evidence Level Criteria Document | (78) |
| CYP3A Gene Expert Panel Roster | (83) |
| P450 Nomenclature Site – Archive | (7) |
| PharmGKB | |
| CYP3A5 Gene Page | (59) |
| Gene-Specific Information Tables for CYP3A5 Allele Definition Table Allele Functionality Table Frequency Table Diplotype-Phenotype Table Gene Resource Mappings |
(60) |
| CYP3A5 Drug Label Annotations | (95) |
| PGx Publication Tips | (63) |
| CPIC | |
| Guidelines Standard Operating Procedure (SOP) for Assigning Allele Function Gene/Drug Pairs Process for assigning CPIC levels Levels for gene/drug pairs Process for prioritizing CPIC guidelines |
(64) (82) (27) |
| Other Resources | |
| FDA Pharmacogenomic Biomarkers in Drug Labeling and FDA Table of Pharmacogenetic Associations |
(96, 97) |
| Drug Interactions Flockhart Table™ | (98) |
| GTR:Genetic Testing Registry | (72) |
| Human Genome Variation Society (HGVS) Nomenclature | (74) |
| NCBI Reference Sequences Database | (99) |
| Locus Reference Genomic (LRG) Project | (80) |
| Database of Genomic Variants – catalogue of human genomic structural variation | (75) |
| Pharmacogenomics Clinical Annotation Tool (PharmCAT) | (61) |
Clinical Relevance
CYP3A5 and its paralog CYP3A4 have 83% amino acid sequence similarity and overlapping substrate specificity, e.g., in the metabolism of steroid hormones, nifedipine (103), cyclosporine (8), and fentanyl (9, 10). The main differences between CYP3A4 and CYP3A5 are in their active centers and substrate access channels (11), which may explain the observed differences in the kinetic parameters of catalysis and inhibition by various compounds (12, 13) and the formation of alternative CYP3A5 metabolites for some substrates (14, 15). The CYP3A5 enzyme contributes to the metabolism of diverse clinical drugs, including tacrolimus (16), cyclosporine (17), sirolimus (18), saquinavir (19), maraviroc (20), midazolam (21), vincristine (22), and statins (23). CYP3A5 was also suggested to confer resistance to tyrosine kinase inhibitors and paclitaxel (24); however, these associations have not been confirmed (25). The most robust evidence supporting a clinical impact of CYP3A5 genetic variation is for tacrolimus metabolism (CPIC guideline (26); PharmGKB level of evidence 1A). Clinical implications of CYP3A5 genetic variation for other drugs include the immunosuppressants cyclosporin and sirolimus (CPIC level C; PharmGKB level of evidence 3) and the sedative midazolam (CPIC level C/D; PharmGKB level of evidence 3) (27). Genetic variation has also been suggested to play a role in the metabolism of vinca-alkaloids, such as the anti-cancer drug vincristine (22, 28, 29).
Tacrolimus, usually given in combination with mycophenolate mofetil and steroids to improve graft survival (30), is the first-line immunosuppressant used to prevent graft rejection in solid organ transplantation. Due to its narrow therapeutic index and the large variability in patient drug plasma levels, therapeutic drug monitoring is routinely performed to individualize tacrolimus dose (31). Optimal dosing is crucial to decrease the risk of graft rejection, when underexposed, or nephrotoxicity in cases of overexposure. CYP3A5 genotyping can be used to optimize tacrolimus dosing to achieve target therapeutic blood concentrations.
Tacrolimus is extensively metabolized in the small intestine and liver predominantly by CYP3A5, with CYP3A4 and POR playing minor roles (16, 32, 33). Thus, genetic variation of CYP3A5 is highly relevant for tacrolimus dose requirements. Associations between CYP3A5 genotype and tacrolimus plasma levels have been observed in adult and pediatric kidney, liver, heart, and lung transplant recipients, although CYP3A5 variation has not been proven to cause acute rejection (34). A randomized controlled clinical trial comparing the efficacy of CYP3A5 genotype-based tacrolimus dosing against standard care demonstrated that dosing based on pre-emptive genotyping reduced the time required to reach optimal drug exposure (35). Since tacrolimus is extensively metabolized in the liver, it has been suggested that both donor and recipient CYP3A5 genotype may contribute to tacrolimus pharmacokinetics (36, 37), although there is not enough evidence to support this hypothesis fully.
There is conflicting evidence regarding the association between CYP3A5 variation and drug plasma levels of the immunosuppressive drug cyclosporine (38–43). Renal CYP3A5-mediated cyclosporine metabolism may have clinical importance, as CYP3A5 expressors locally generate more nephrotoxic metabolites than non-expressors, which may lead to an increased risk of nephrotoxicity (44). Another study demonstrated significantly longer survival for kidney transplant patients with a CYP3A5*1 allele receiving cyclosporine treatment (45). Despite these observations, the clinical relevance of CYP3A5 pharmacogenetics for cyclosporine metabolism remains unclear.
CYP3A5 genotype was also reported to have a significant influence on sirolimus metabolism; however, this drug is metabolized by both CYP3A4 and CYP3A5, with CYP3A4 being the major metabolizing enzyme (18).
Factors Influencing CYP3A5 activity
Similar to CYP3A4, CYP3A5 expression levels are inherently higher in women than men when assessed in liver tissue samples (46). This sex dimorphism has also been observed via in vivo assessment of plasma metabolite levels (47). It is believed that these differences are due to sex-dependent control of the transcription networks responsible for the hormone-induced expression of liver enzymes. Regarding developmental expression of CYP3A5, protein is detectable in liver samples at early gestational ages (48, 49) and expression after that seems to remain constant and independent of age (from early gestation to 18 years) (48).
Cytochrome P450 oxidoreductase serves as the only electron donor for CYP3A5 (50). Monooxygenase catalysis by CYP3A5 depends on this electron donor (51), thus the cellular level of oxidoreductase may impact CYP3A5 activity. Otherwise, the catalytic activity of CYP3A5 largely depends on the presence of the microsomal form of cytochrome b (CYB5A) (52), which is substrate dependent. There is also evidence that CYP3A5 forms protein-protein complexes with other CYPs such as CYP3A4 and CYP2E1 (53), which may impact interindividual variability.
CYP3A5 activity can be impacted by inhibitors, leading to phenoconversion (54), or inducers, that may increase CYP3A5 expression in individuals with a functional allele. Decreased activity may also be caused by inflammation (55). Thus, patients with a CYP3A5*1 allele and chronic kidney disease may require dose adjustments for drugs metabolized by CYP3A5 (56). Furthermore, the expression of CYP3A5 appears to be regulated, at least to some extent, by a long non-coding RNA, AC069294.1. This regulatory mechanism may account for additional variability in CYP3A4 and CYP3A5 expression (57). Of note, the expression of AC069294.1 has been shown to be impacted by a CYP3A4 intronic variant, rs2242480, that is part of the CYP3A4*1G haplotype.
CYP3A5 in PharmGKB and CPIC
PharmGKB collects, curates, and disseminates knowledge about the impact of human genetic variation on drug response (58). The PharmGKB CYP3A5 gene page (59) allows structured access to gene-specific pharmacogenomic knowledge. Information is presented in sections, including prescribing information, drug label annotations, clinical annotations, variant annotations, and curated pathways. As of January 2022, the PharmGKB CYP3A5 gene page includes 3 clinical guideline annotations for tacrolimus (CPIC (18), Dutch Pharmacogenetics Working Group (DPWG) and French National Network of Pharmacogenetics (RNPGx)), and 5 drug label annotations for three drugs: Maraviroc (Swiss Agency of Therapeutic Products (Swissmedic)), Prasugrel (European Medicines Agency (EMA), FDA, and Swissmedic), and Dolutegravir (Swissmedic). PharmGKB contains 67 CYP3A5-related clinical annotations, which are evidence-rated genotype-level summaries for specific variant/allele–drug combinations based on curated literature (variant annotations). Pharmacokinetic pathways depicting CYP3A5 in drug metabolism are available for 40 drugs, although the significance for CYP3A5 involvement varies by drug. PharmGKB and CPIC work collaboratively to develop gene-specific resources that accompany each CPIC guideline, including allele definition mapping, allele functionality, allele frequency, and diplotype to phenotype mapping files in a standardized format. Gene-specific information tables for CYP3A5 are available from PharmGKB (60). In addition, the Pharmacogenomics Clinical Annotation Tool (PharmCAT) facilitates the interpretation and reporting of pharmacogenomic-based dosing recommendations, including those for CYP3A5 (61, 62).
PharmVar and PharmGKB collaborated to develop templates to facilitate more consistent and transparent reporting of genotype details and how genotype is translated into phenotype. The two template files that have been developed may be adapted to individual needs and are available as supplemental materials (available through the PharmVar CYP3A5 gene page under ‘More Documents’ and at PharmGKB under ‘PGx Publication Tips’ (63)). The first template collects information, including methods or platforms used for genotyping and which genetic variants are interrogated; the template also provides a standardized set-up for reporting genotype results for individual subjects, as well as allele frequencies. The second template facilitates the reporting of how genotype is translated into phenotype and genotype frequencies. Publication of this information in a structured form greatly facilitates access to data for subsequent curation by PharmGKB and other groups.
CPIC develops structured, evidence-based clinical practice guidelines for drugs affected by pharmacogenetic variation (64). To date, one CYP3A5 CPIC guideline has been published that includes CYP3A5 and tacrolimus (26). The guideline has multiple components, including CYP3A5 phenotype-specific therapeutic recommendations, a systematic evidence review, and implementation resources to support the translation of the guideline into electronic health records (EHRs) with examples of clinical decision support text.
For CYP3A5 phenotypic classification, individuals are categorized into the following CPIC-recommended phenotype categories: poor (PM), intermediate (IM), and normal (NM) (formerly EM, extensive) metabolizers. Individuals are, however, often also described as ‘expressors’ (those having one or two CYP3A5*1 alleles) and ‘non-expressors’ (those having two nonfunctional alleles) in the literature. The Diplotype-Phenotype-Table provided by the PharmGKB and CPIC serves as a reference for translating CYP3A5 genotype to phenotype (60).
Need for standardized genetic variation definitions and reporting of functional/clinical impact
To guide drug therapy, it is imperative to understand how CYP3A5 allelic variation can impact CYP3A5 function and utilize standardized reporting and data representation. This effort aligns with recent reports emphasizing that clinically actionable pharmacogenetic information must be accurately represented in EHRs using a harmonized system for genotype and phenotype information (65, 66). Clinical testing for CYP3A5 can be performed on a variety of platforms using different methodologies, and although genotyping data can be reported in different ways, such as chromosomal or genomic position on reference sequence (RefSeq), amino acid change, Single Nucleotide Polymorphism Database (dbSNP) rsID, and/or using star allele nomenclature (6, 66), many laboratories use PharmVar star allele nomenclature and the CPIC-recommended system to translate genotype to phenotype.
A study performed by the Genetic Testing Reference Material Program (GeT-RM) concluded that many pharmacogenetic variants were not interrogated consistently across commercial and laboratory platforms (67), which was substantiated by Moyer et al. surveying reporting pharmacogenetic test results for CYP2D6 and CYP2C19 (68). The use of nomenclature as provided by PharmVar will minimize inconsistent interpretations of pharmacogenetic test results. To guide pharmacogenetic testing, the Association of Molecular Pathology (AMP) and the College of American Pathologists (CAP) are publishing recommendations for clinical genotyping allele selection using PharmVar nomenclature, exemplifying its clinical utility. These recommendations also include information on test platforms. AMP/CAP recommendations have been published for CYP2C9 (69), CYP2C19 (70), and most recently, CYP2D6 (71) allele testing. Recommendations are planned for CYP3A5, which will include many commercially available pharmacogenetic test panels (72).
Throughout this review, variants are denoted according to their relative position in the CYP3A5 NM_000777.5 reference transcript sequence with the ‘A’ of the ATG translation start codon being +1 (as shown on the PharmVar gene page (73)). For example, the CYP3A5*3 allele-defining variant (rs776746) is referred to as c.219–237A>G (splice defect), indicating its intronic position. The CYP3A5*6 defining variant (rs10264272) is located within the coding region and is thus shown as c.624G>A (splice defect). The CYP3A5*7 allele-defining variant (rs41303343) has a 1-nucleotide insertion which is annotated by PharmVar as c.1035_1036insT (p.T346fs), denoting a frameshift at amino acid position 346. Per the Human Genome Variation Society (HGVS) (74), this insertion is described as NM_000777.5:c.1035dup.
CYP3A5 reference materials
The Centers for Disease Control’s GeT-RM Coordination program is a combined effort with the Coriell Institute for Medical Research and members of the pharmacogenetic testing community. Established sets of well-characterized reference materials are needed for assay development, validation, quality control, and proficiency testing. The increasing need for reference materials based on the growing use of pharmacogenetic testing prompted the establishment of a set of 137 genomic DNA samples characterized for 28 pharmacogenes, including CYP3A5 and “consensus” genotypes (67). Although several platforms included testing of variants identifying the rare CYP3A5*8, and *9 alleles, only samples with CYP3A5*3, *6, and *7 alleles were identified among the samples tested. Testing and research laboratories can acquire these materials from the Coriell Institute (Camden, NJ, USA), as they are publicly available.
The CYP3A5 Gene Locus
The CYP3A5 gene is a member of the CYP3A family; it has 13 exons and is translated into a protein of 502 amino acids. The gene is located on the minus strand of chromosome 7q22.1 spanning a region of approximately 32 kilobases (kb). The CYP3A locus, in addition to CYP3A5, also harbors CYP3A4, CYP3A7, and CYP3A43, spanning 231 kb of genomic region (Figure 1). Genotyping assays need to employ CYP3A5-specific regions for primer design (e.g., using intronic sequences) to avoid amplifying any of the other genes in the locus. Numerous genotyping assays and tests are available commercially for CYP3A5 testing (72). Per the Database of Genomic Variants, a curated catalog of human genomic structural variation (75), copy number variation appears to be exceedingly rare at the CYP3A5 gene locus and thus is not routinely tested.
Figure 1. Overview of the gene locus and allelic variation.
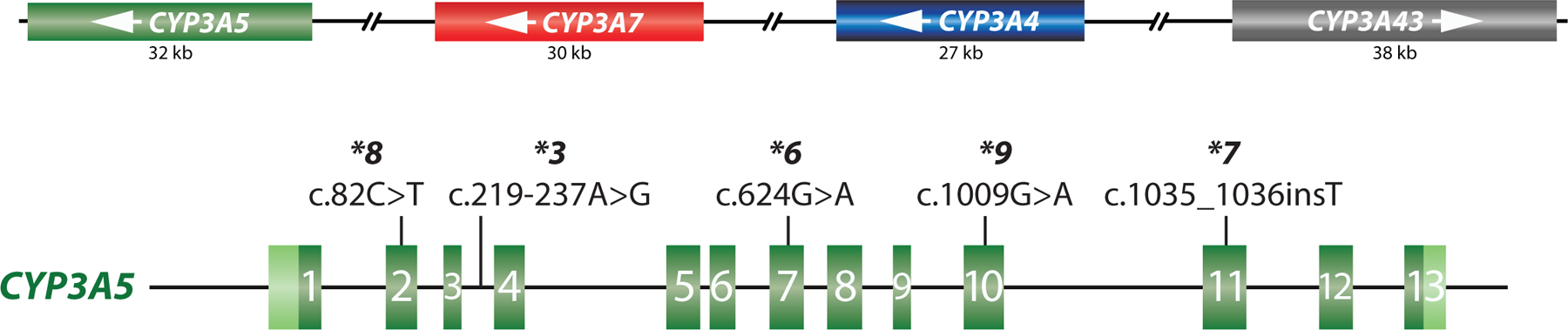
The top panel provides a graphical overview of the CYP3A gene locus containing the CYP3A5, CYP3A7, CYP3A4 and CYP3A43 genes and their approximate length in kilobase pairs (kb). CYP3A43 is encoded on the forward strand, while the other three genes are encoded on the minus strand as indicated by the white arrows. The bottom panel shows the CYP3A5 gene in the forward orientation (5’ to 3’). CYP3A5 comprises 13 exons as indicated by the green boxes; 5’ and 3’ untranslated regions (UTRs) are highlighted in light green. The core variants defining the CYP3A5*3, *6, *7, *8 and *9 alleles are as shown.
CYP3A5 allele, genotype, and phenotype frequencies across populations
The CYP3A5 frequency table available at PharmGKB (60) summarizes population-based allele frequencies reported in the literature. Studies were considered for inclusion if 1) the population’s ethnicity was clearly indicated, 2) either allele frequencies or genotype frequencies were reported, 3) the methodology by which the genes were genotyped was indicated, and 4) the study represented an original publication. The ethnicities/locations reported in the articles were mapped into seven geographically defined groups (American, Central/South Asian, East Asian, European, Near Eastern, Oceanian, and Sub-Saharan African) and two admixed groups (African American/Afro-Caribbean and Latino), using the biogeographical grouping system developed by PharmGKB (76). The CYP3A5 frequency table is periodically updated and contains multiple tabs summarizing ‘allele frequencies by biogeographical group’, ‘diplotype frequencies by biogeographical group’, ‘phenotype frequency’, and ‘references’; the latter describes allele frequencies for each publication included in the listing, which also allows the user to customize allele frequencies as needed. There are, however, limitations regarding the accuracy of allele frequencies as follows: 1) frequencies are based on published allele data (limited or unavailable for some populations and alleles), 2) most studies test for a limited number of allelic variants that may lead to an underestimation of certain alleles. For example, most tests only interrogate the c.219–237A>G splice defect and default to a CYP3A5*1 assignment in its absence. This limitation may inflate the frequency of the CYP3A5*1 allele and cause underreporting of other alleles. Therefore, all calculations based on allele frequencies are estimates.
There is considerable variation among the estimated frequencies for individual alleles across and within the biogeographical groups. The CYP3A5*3 allele frequency is highest in Europeans and lowest in African American/Afro-Caribbeans and Sub-Saharan Africans, averaging 92%, 32%, and 24%, respectively. Its frequency is also high in Asians (67%–75%), Latinos (77%), and Near-Eastern populations (84%). In contrast, CYP3A5*6 and *7 are more readily observed in individuals with African ancestry (11%–19% and 9%–12%, respectively), while these alleles are less than 4% or rarely found in other ethnic groups (60).
Considering the relatively small number of CYP3A5 haplotypes and the low frequencies of most of the alleles, the actual number of genotypes found in a population or patient cohort is quite small compared to those seen for other CYP genes. Phenotype frequencies calculated from the averaged allele frequencies across populations are provided in the ‘Phenotype frequency’ tab of the CYP3A5 Frequency Table (60). Calculated phenotype frequencies should be viewed with caution owing to the limitations mentioned above regarding the accuracy of allele frequencies, inconsistencies in the classification of ‘population’, ‘ethnicity’, or ‘race’ (77), as well as the recent retirement of CYP3A5*2, *4 and *5 discussed in more detail below.
PharmVar Nomenclature and CYP3A5 allele designation
PharmVar stores and displays allelic data consistently across genes, relying on public standards and data sources wherever possible. The “Allele Designation and Evidence Level Criteria” document describes the nomenclature system (78). A new star number is only issued if a haplotype contains a sequence variant that: 1) results in an amino acid change (example: CYP3A5*8 was defined based on having c.82C>T (rs55817950) causing a p.R28C change; 2) contains a nonsense variant (example: CYP3A5*7 contains a 1-nucleotide insertion, c.1035_1036insT, causing a frameshift at amino acid position p.T346fs), or 3) changes expression levels or affects splicing (example: CYP3A5*3 contains an intronic variant, c.219–237A>G, which causes a splice defect). Significantly, new haplotypes that contain previously characterized variants that lead to a nonfunctional protein are cataloged under the original star allele number as a suballele. For example, any haplotype having a novel variant in addition to c.219–237A>G will be designated as a CYP3A5*3 suballele and considered to have no function, regardless of whether the novel variant impacts function on its own or not.
The PharmVar CYP3A5 gene page
The PharmVar CYP3A5 gene page (73) details all currently defined star alleles. Each allele is listed in sequential order on the CYP3A5 gene page and cross-referenced with its legacy name, if existing. Each allele contains information on variants, including core variants (see Core Allele section below), haplotype evidence level, and CPIC clinical allele function assignment. A ‘Compare View’ allows the viewer to toggle between the standard allele table and the Comparative Allele ViewEr (CAVE) tool. The CYP3A5 gene page also includes ‘Read Me’, ‘Change Log’, and ‘More’ documents providing additional relevant information and resources, including examples and links to other websites, such as a link to PharmGKB’s gene information. Also, each characterized haplotype receives a PharmVar ID (PVID), i.e., a unique numeric identifier analogous to dbSNP rsIDs (79). PVIDs and their haplotype definitions can be tracked in the database via the PVID Lookup function.
Variant Mapping:
Variants are mapped to the genomic (NG_007938.2) and transcript (NM_000777.5) reference sequences as well as to the genome builds GRCh37 (NC_000007.13) and GRCh38 (NC_000007.14). Of note, LRG_1431, the CYP3A5 Locus Reference Genomic record (80) matches 100% with the RefSeq identifier NG_007938.2, and NM_000777.5 represents the Matched Annotation from NCBI and EMBL-EBI (MANE) select transcript which is representative of biology, is well-supported, expressed, and highly conserved (81). This transcript matches GRCh38 and is 100% identical with its RefSeq for 5’UTR, CDS, splicing, and 3’UTR.
One rather crucial difference between GRCh37 and GRCh38 is that GRCh37 represents a CYP3A5*3 allele (has a ‘G’ at c.219–237 which corresponds to g.99270539C on GRCh37) while GRCh38 matches the CYP3A5*1 allele (has an ‘A’ at c.219–237 which corresponds to g.99270539T). Consequently, c.219–237A>G is not reported as a ‘variant’ when sequences are compared to GRCh37. Therefore, when choosing GRCh37 as the reference setting on the CYP3A5 gene page, the CYP3A5*3 core allele does not have any variants displayed while all other core alleles including their respective suballeles, have g.99270539C>T. Lastly, g.99270539C>T is not annotated as ‘splice defect’ because this change represents the nucleotide allowing normal splicing. Also see illustration in the Read Me document available through the CYP3A5 gene page on PharmVar.
On the PharmVar CYP3A5 gene page, the user can easily cross-reference genomic coordinates and cDNA positions by choosing the respective reference sequence or genome build of interest; there is also the option of two count modes, i.e., counting from the first nucleotide in the reference sequence or the ATG translation start codon being +1. Variant annotations are also provided according to the Human Genome Variation Society (HGVS) (74) and the more traditional PharmVar display format. Figure 2 provides an excerpt of the page illustrating CYP3A5*3, *6, and *7.
Figure 2. Overview of core allele and suballele categorization.
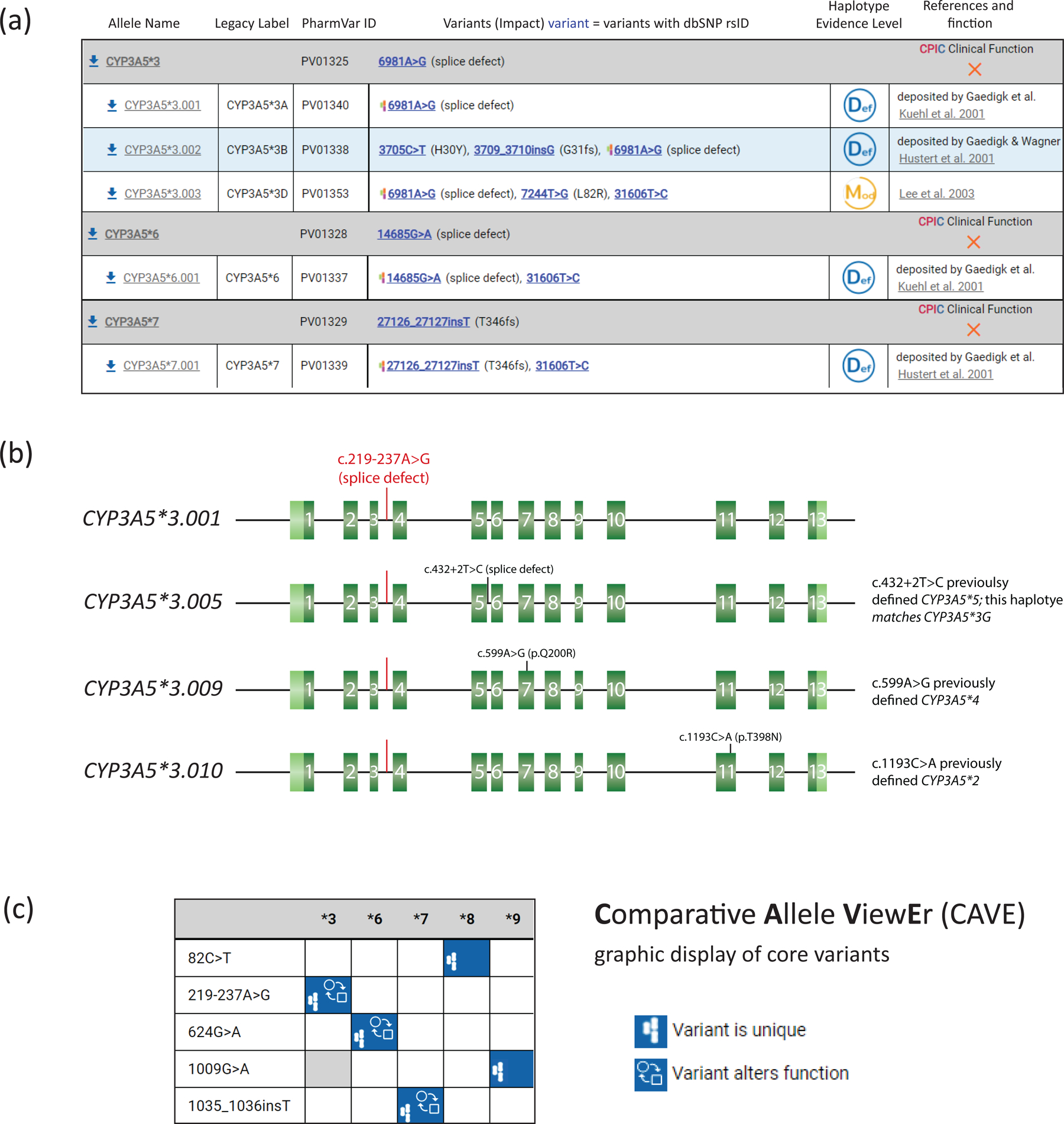
Panel (a) is an excerpt of the CYP3A5 gene page showing *3, *6 and *7 allele definitions with NM_000777.5 as the reference sequence; their respective core allele definitions are depicted by gray bars. Core variants, PharmVar ID (PVID), and haplotype evidence levels are shown for each allele. Selected CYP3A5*3 suballeles are displayed underneath the core allele bar; currently, CYP3A5*6 and *7 have only one suballele. Legacy allele designations are cross-referenced (e.g., CYP3A5*3.001 corresponds to *3A and *3.003 corresponds *3D). Panel (b) is a graphical representation of the CYP3A5*3 suballeles containing the variants that previously defined the CYP3A5*2, *4 and *5 alleles; these are now cataloged as CYP3A5*3 suballeles due to having the intron 3 c.219–237A>G splice variant (highlighted in red). Green boxes represent exons; the 5’ and 3’ untranslated regions are shown in light green. Panel (c) represents the graphical output of the Comparative Allele ViewEr (CAVE) when comparing the five currently existing star alleles. The PharmVar symbols indicate that the variant is unique to the allele and the function symbol signifies that the variant alters function.
CYP3A5 Allele Function:
PharmVar displays allele clinical function as determined by CPIC using their standardized protocol detailing the criteria for assigning clinical function to alleles to harmonize the process across guidelines (82). It is important to realize that CPIC’s primary focus is to assign allele function based on clinical actionability, not solely on molecular or biochemical function. The expert consensus for allele functions can be accessed on PharmGKB (60). The table includes allele clinical functional status (displayed on PharmVar), and references reviewed during the assignment process. The filter option on the PharmVar CYP3A5 gene page allows the user to sort alleles by functional status.
CYP3A5 haplotype evidence levels:
PharmVar designates the “Haplotype Evidence Level” for each star allele reported on the CYP3A5 gene page. Evidence levels are displayed as symbols indicating ‘definitive’ (Def), ‘moderate’ (Mod), or ‘limited’ (Lim) levels of support for a given haplotype. This three-tiered system represents a modified ClinVar classification system; more detailed information is provided in the ‘Allele Designation and Evidence Criteria Level’ document (78). This type of information (e.g., whether an allele was sequenced across the gene, how haplotype was determined) was not systematically captured prior to PharmVar. For existing haplotype definitions, a literature review was conducted to assign evidence levels. Several CYP3A5*3 suballeles are currently labeled as ‘Lim’ and ‘Mod’ because their definitions are not based on current PharmVar requirements, or the phase of the variants was computationally inferred. The value of evidence levels is centered on providing as much information on haplotype reliability as possible, enabling users to quickly parse haplotypes based on robust, high evidence as required for ‘Def’ versus other haplotypes with ‘Lim’ or ‘Mod’ evidence levels. PharmVar solicits submissions for all alleles labeled ‘Lim’ and ‘Mod’, but especially CYP3A5*8 and *9, to substantiate their current definitions and raise their evidence levels to ‘Def’. PharmVar encourages encore submissions for alleles with single citations and shown as ‘Def’ to further corroborate a haplotype definition.
Core allele definitions:
While many CYP genes have star alleles that share one or more ‘core’ defining sequence variant(s), all CYP3A5 haplotypes defined to date are characterized by a single unique variant. Suballele information may be valuable for designing assays or test platforms (sequence or genotype-based) and the interpretation of genotyping test results. There is no need to distinguish suballeles for phenotype prediction because all alleles under a star number are presumed to be functionally equivalent. Thus, even if a test can distinguish suballeles, these can be reported simply using their respective core allele definition.
A core allele is defined only by sequence variations that cause an amino acid change, or impact function by changing expression levels, or interfere with splicing, and are present in all suballeles within an allele group. This rule-based system allows all suballeles categorized under one star number to be collapsed into a single ‘core’ definition. For example, all CYP3A5*3 suballeles share the c.219–237A>G variant, causing a splice defect rendering this haplotype nonfunctional. Therefore, this variant constitutes the CYP3A5*3 core allele definition (Figure 2).
The core alleles are the basis of the CYP3A5 allele definition table used in CPIC guidelines and PharmGKB (Table 1). The CYP3A5 core allele definitions are also utilized for clinical annotations in PharmGKB.
The PharmVar Comparative Allele ViewEr:
PharmVar developed the Comparative Allele ViewEr (CAVE) tool to compare core alleles easily. This tool can be accessed using the “Compare View” button on the CYP3A5 gene page. Figure 2c visualizes the utility of this tool by comparing the CYP3A5*3, *6, *7, *8 and *9 alleles. In this display mode, it is easy to see that no core variants are shared among the currently defined star alleles and which genetic variants are known to alter function.
Curation efforts
CYP3A5 nomenclature was curated by a panel of international experts representing research, clinical testing, implementation interests, and PharmGKB/CPIC representatives to ensure that the nomenclature is consistent with CPIC guidelines and to facilitate dissemination to a greater audience through PharmGKB and other resources, such as ClinGen. The composition of the panel can be found on the PharmVar website (83).
Gene region mapped/required for allele definition:
CYP3A5 allele definitions include variants within the coding region, 1000 bp upstream of the ATG translation start codon, and 250 bp of the 3’ untranslated region. Intronic variants are not considered for allele definitions unless they affect enzyme activity. Consequently, the CYP3A5*3E, *3H, and *3I suballeles were not transitioned into the PharmVar database. New submissions require covering the regions mentioned above and information for the intronic c.219–237A>G splice variant. Some of the alleles defined before PharmVar are based on limited sequencing data (e.g., lack information for up- and downstream regions) or were computationally inferred.
Revisions and corrections:
Extensive curation efforts were part of the content transfer from the P450 nomenclature webpage into the PharmVar database to standardize the annotations to the above-mentioned conventions. Revisions made are detailed in the ‘Change Log’ document on the CYP3A5 gene page (73) and are summarized in Table 2; this document also records submissions received by PharmVar. The following sections highlight selected efforts undertaken.
Table 2.
Novel suballeles, confirmed alleles and alleles not transitioned into the PharmVar database
| Core Allele Designation | Novel Suballeles | Confirmed Existing Allele Definitions (Legacy Name) | Other Revisions |
|---|---|---|---|
| *1 | *1.003, *1.004, *1.005 | *1.002 (*1D) |
*1B, *1C were not transitioned due to uncertainty of having c.-86G>A and c.-74C>T with c.219–237A as originally published *1E not transitioned due to uncertainty of having c.1523T>C |
| *3 | *3.012 - *3.017 |
*3.001 (*3A),
*3.002 (*3B), *3.005 (*3G) *3.009 *3.010 |
*3C was not transitioned due to uncertainty of having c.1523T>C *3E, *3H and *3I were not transitioned; these matched *3.001 after removing intronic SNVs |
| *6, *7, and *8 | none | *6.001 (*6), *7.001 (*7), and *8.001 (*8) | none |
All novel suballeles received a haplotype evidence level of ‘definitive’ (Def); the haplotype evidence level of those confirmed with new data was raised from ‘limited’ (Lim) to ‘definitive’ (Def).
During the transition process into the PharmVar database, comments and footnotes were removed and errors corrected. References that support allele definitions were updated and those solely describing function removed (note that references describing function are summarized in the PharmGKB/CPIC CYP3A5 Allele Functionality table (60)). As detailed in Table 2, several suballeles (CYP3A5*1B, *1C, *1E, *3C) were not transitioned due to insufficient evidence supporting the haplotypes as initially defined. Finally, a variant in the 3’ untranslated region (rs15524, c.1523T>C) was annotated as “C>T” on the legacy page (7) but is now shown as “T>C”. Lifting allele definitions to the current RefSeq NG_007938 caused a switch in annotations for this variant, i.e., haplotypes shown on the legacy page as having this variant no longer do, while all others gained c.1523T>C (note that this variant is annotated by HGVS as c.*14T>C). CYP3A5*1E and *3C were not transitioned as it was uncertain whether these haplotypes have c.1523T>C or not.
Allele retirements and novel haplotypes:
PharmVar has retired three star alleles, namely CYP3A5*2 (defined before the CYP3A5*3 allele was discovered (3)), *4, and *5. As detailed in Table 3, it was uncertain whether the variants defining the latter indeed occurred on their own or are part of CYP3A5*3 haplotypes. To address these concerns, the PharmVar team utilized data available through the 1000 Genomes Project to show that c.1193C>A (CYP3A5*2 core variant), c.599A>G (CYP3A5*4 core variant), and c.432+2T>C (CYP3A5*5 core variant) do not occur by themselves, as projected by their original star allele definitions, but are part of CYP3A5*3 haplotypes. Several samples heterozygous for each, c.1193C>A, c.599A>G, or c.432+2T>C, were homozygous for the CYP3A5*3 c.219–237A>G splice variant unequivocally informing both haplotypes (Figure 3a). Unpublished genotype data further corroborated this observation as c.1193C>A, c.599A>G, or c.432+2T>C, when tested, were only found in samples that were either homozygous or heterozygous for the CYP3A5*3 splice variant (84). The c.1193C>A and c.599A>G variants are now part of the newly created CYP3A5*3.010 and CYP3A5*3.009 suballeles; regarding c.432+2T>C, there already was a suballele, CYP3A5*005 (formerly *3G) accurately representing this haplotype. Of note, CYP3A5*2, *4, and *5 are reported as CYP3A5*3 if a pharmacogenetic test does not interrogate their respective identifying variants. For example, a patient who tested homozygous for the CYP3A5*3 c.219–237A>G splice variant and heterozygous for the CYP3A5*2 c.1193C>A variant may previously have been reported as “CYP3A5*2+*3/*3” or “CYP3A5*3/*3 +*2”. Moving forward, these should now simply be reported as CYP3A5*3/*3.
Table 3.
Retirement of CYP3A5*2, *4, and *5. Variants that defined these star alleles are now part of CYP3A5 suballeles.
| Allele | Notes |
|---|---|
| CYP3A5*2 | Jounaïdi et al. 1996 (3) found c.1193C>A (p.T398N) in the cDNA of two subjects; the *3 intronic variant c.219–237A>G was unknown at the time this variant was first described. Data available through the 1000 Genomes Project showed unequivocally that c.1193C>A is part of a CYP3A5*3 haplotype and does not occur on its own. Therefore, the CYP3A5*2 allele was retired; c.1193C>A is now part of the *3.010 suballele. |
| CYP3A5*4 | Chou et al 2001 (100) first identified c.599A>G (p.Q200R) but did not provide information regarding the subject’s *3 genotype (c.219–237A>G). Shih et al. (101) identified a subject heterozygous for c.599A>G and homozygous for c.219–237A>G. Data available through the 1000 Genomes Project showed unequivocally that c.599A>G is part of a CYP3A5*3 haplotype and does not occur on its own. Therefore, the CYP3A5*4 allele was retired; c.599A>G is now part of the *3.009 suballele. |
| CYP3A5*5 | Chou et al 2001 (100) first identified c.432+2T>C (splice defect) but did not provide information regarding the subject’s *3 genotype (c.219–237A>G). Saeki et al. 2003 (102) identified a subject who was heterozygous for c.432+2T>C and homozygous for c.219–237A>G; this allele was defined as CYP3A5*3G (now *3.005). Data available through the 1000 Genomes Project showed unequivocally that c.432+2T>C is part of the CYP3A5*3.005 haplotype and does not occur on its own. Therefore, the CYP3A5*5 allele was retired; c.599A>G was already part of the *3.005 suballele. |
| CYP3A5*3.011 | Lee et al 2003 (85) found c.1009G>A (p.A337T) in a subject who was heterozygous for *3; it was uncertain, however, whether c.1009G>A is on the *3 allele or indeed on the newly defined *9 allele. *3.011 was created to reflect this uncertainty. New data supports the current *9 definition and thus, *3.011 was retracted. |
Figure 3. Methods and approaches to characterize CYP3A5 allelic variants.
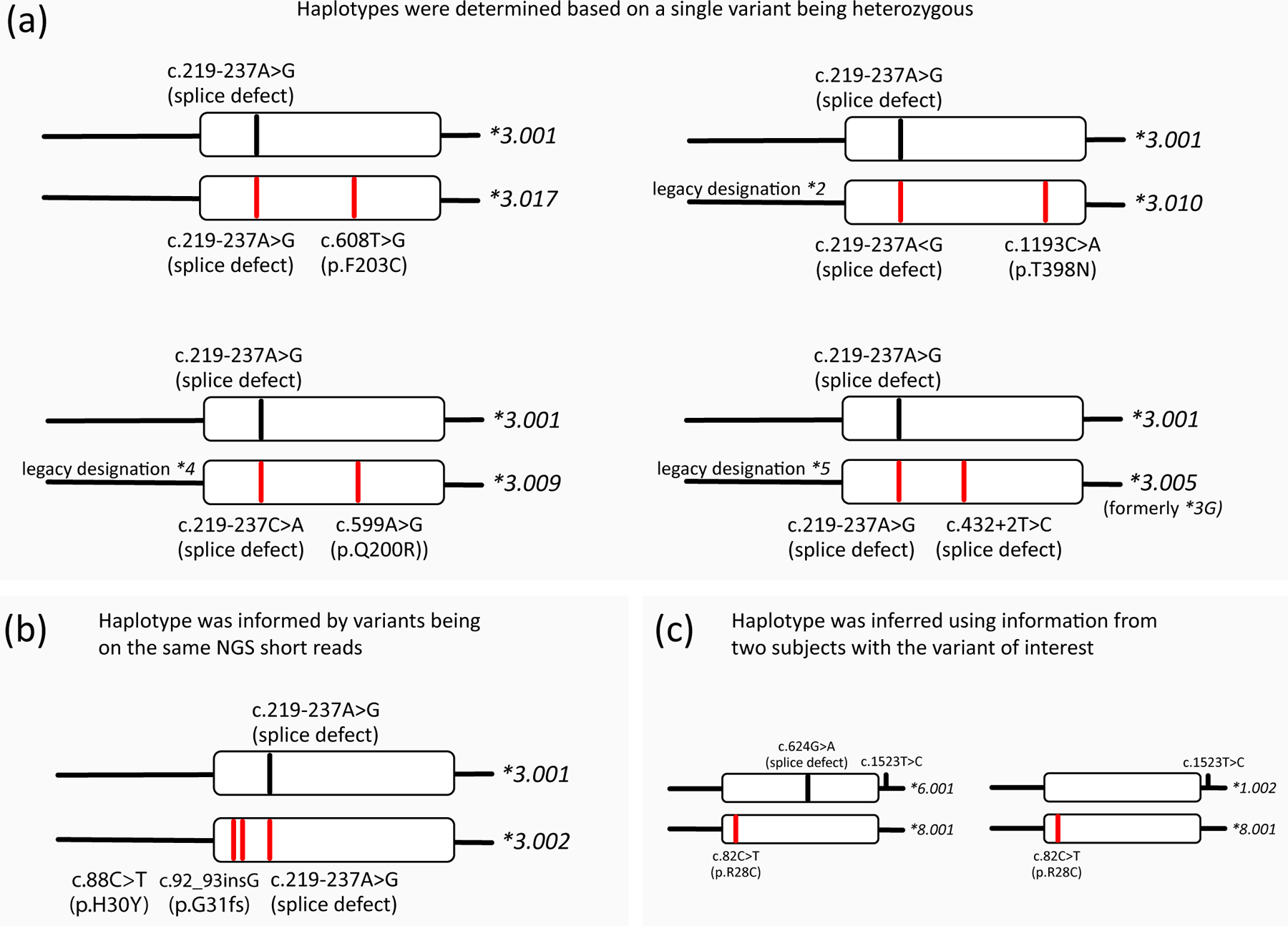
Panels (a-c) provide examples of alleles submitted to PharmVar to name or confirm existing allele definitions. Red lines highlight variants found on the submitted alleles. Haplotypes shown in panels a and b utilized WGS data either confirmed by WES or targeted NGS-based sequencing panels; haplotypes in panel c represent two subjects of the Simons Genome Diversity Project (93) who underwent WGS. Haplotypes are inferred; variants were not independently confirmed.
Panel (a) provides examples of haplotypes that can unequivocally be deduced as only a single variant is heterozygous. These include selected submissions supporting the reclassification of CYP3A5*2, *4 and *5 as CYP3A5*3 suballeles as shown. Each subject was homozygous for the CYP3A5*3 splice variant c.219–237A>G and heterozygous for c.1193C>A, c.599A>G or c.432+2T>C. Thus, the latter variants are part of CYP3A5*3 haplotypes and do not occur independently. Panel (b) shows a subject whose three heterozygous variants were placed on the same chromosome because c.88C>T and c.92_93insG were found on the same short reads. Thus, variants in close proximity may be deduced directly from short-read data. Panel (c) illustrates two samples with the CYP3A5*8 core variant c.82C>T. Although the phase of their respective haplotypes could not be inferred with certainty, the data support that c.82C>T is not on a CYP3A5*3 haplotype.
CYP3A5*8 was initially found in a subject that did not have the CYP3A5*3 splice variant, suggesting that its core variant (c.82C>T, p.R28C) does indeed occur by itself as initially published (85). Diplotypes of two subjects submitted by Twesigomwe & Hazelhurst (Figure 3c) further substantiate the initial findings and current CYP3A5*8 allele definition. Uncertainty remains, however, regarding CYP3A5*9. The sentinel subject was heterozygous for c.1009G>A (p.A337T) and c.219–237A>G, and thus it remained uncertain whether c.1009G>A indeed occurs on its own (85). One sample in the St Jude database (86) heterozygous for c.1009G>A and negative for CYP3A5*3 supports the current allele definition and thus, the CYP3A5*3.011 suballele that was created to reflect this uncertainty, was retracted (Table 3). Since c.1009G>A is exceedingly rare, it may be difficult to identify additional subjects for further analysis.
Lastly, while no novel haplotypes have been identified, PharmVar has expanded its catalog of CYP3A5*3 suballeles (Table 2) and raised evidence levels for several alleles to ‘Def’.
CYP3A5 allele characterization: Methods and approaches
While of considerably lower genetic complexity than many other pharmacogenetically relevant CYP genes, the determination of variant linkage is nevertheless essential for the complete characterization of CYP3A5 haplotypes. This section offers examples from CYP3A5 submissions to PharmVar to illustrate how alleles can be characterized. Additional information regarding this topic can also be found in previous GeneFocus articles (87, 88).
Haplotypes can unequivocally be determined using deductive logic. Homozygosity of any number of variants indicates that all define the haplotype; the haplotype is also evident if all variants but one are homozygous, as shown in Figure 3a. Homozygosity of c.219–237A>G and heterozygosity of c.608T>G informs that the new suballele, designated as CYP3A5*3.017, has both variants. Likewise, subjects homozygous for the CYP3A5*3 splice variant (c.219–237A>G) and heterozygous for c.1193C>A, c.599A>G or c.432+2T>C, the core variants that were initially used to define CYP3A5*2, *4 and *5, respectively, classifies these as CYP3A5*3 suballeles. As described above in the section on curation, all available diplotypes containing c.1193C>A, c.599A>G or c.432+2T>C were either homozygous or heterozygous for c.219–237A>G and thus warranted their retirement as individual star alleles and reclassification as CYP3A5*3.010, *3.009 and *3.005, respectively as shown in Figure 3a.
Variant phases can, in some instances, be informed by short-read NGS data. As shown in Figure 3b, a subject homozygous for the splice variant has two additional variants, c.88C>T and c.92_93insG, which were found on the same NGS short reads indicating that both are on the same allele. This haplotype confirmed CYP3A5*3.002 as it was originally defined. Phasing variants via short reads is, however, limited to variants that are in close proximity. Other approaches such as single molecule sequencing (e.g., PacBio single-molecule real-time (SMRT) sequencing and Oxford Nanopore sequencing) can be employed to resolve the phase of distant variants (exceeding the length of short reads) (89, 90). While there are no examples to demonstrate the utility of these methods for CYP3A5 allele characterization to date, these methods have successfully been utilized for other highly polymorphic pharmacogenomic loci, including CYP2C9, CYP2C19 and CYP2B6.
Variants may also be phased using statistical approaches that leverage population-specific reference genomes. However, the reliability of such computationally inferred haplotypes critically depends on the quality of the reference genome in question and the number of samples in the data set. Thus, this method may only provide limited or moderate evidence of whether a set of variants occurs in cis or trans.
In some cases, such as that illustrated in Figure 3c, the phase of the variants found in subjects of interest cannot be determined with certainty (the shown diplotypes were manually inferred). While these data on their own would not be sufficient to define a novel haplotype, they are supportive of the current CYP3A5*8 allele definition.
Lastly, haplotypes may also be determined using inheritance. This approach has been successfully utilized to infer haplotypes of other CYP genes including CYP2B6, CYP2C8, CYPC9 and CYP2C19 and CYP2C19 (87, 88, 91, 92).
Conclusions
This PharmVar GeneFocus on CYP3A5 provides essential information for understanding this important pharmacogene, complementing clinically relevant information provided by clinical guidelines and other pharmacogenetic resources. We summarize PharmVar’s efforts systematically cataloging CYP3A5 allelic variation, including the retirement of three star alleles: CYP3A5*2, *4, and *5. This review also offers examples of submissions highlighting different approaches to characterize novel haplotypes fully. In addition, we stress our collaborative efforts to make the information useful and easily accessible to the entire pharmacogenetics community through the PharmGKB.
Acknowledgments
We would like to thank Dr Yoshiro Saito of the National Institute of Health Sciences, Kawasaki, Japan, for his expertise and serving on the expert panel and Drs Cyrine Haidar (St Jude Children’s Research Hospital, Memphis, TN, USA), Ann Moyer (Mayo Clinic, Rochester, MN, USA) and Charity Nofziger (Pharmgenetix GmBH, Salzburg, Austria) for sharing genotype data and Drs David Twesigomwe and David Hazelhurst (University of the Witwatersrand, Johannesburg, SA) for collaborating on CYP3A5*8 and submitting their data to PharmVar.
Funding
PharmVar is supported by the Children’s Mercy Research Institute. PharmGKB is supported by the National Institutes of Health (U24 HG010615; PI, T.E.K.) and D.W. by R35GM140845. V.M.L. acknowledges support from the Swedish Research Council [grant agreement numbers: 2016-01153, 2016-01154 and 2019-01837], the EU/EFPIA/OICR/McGill/KTH/Diamond Innovative Medicines Initiative 2 Joint Undertaking (EUbOPEN grant number 875510) and the Robert Bosch Foundation, Stuttgart, Germany. B.I.D. is supported by a Canadian Institute of Health Research Tier 2 Canada Research Chair in Pharmacogenomics and Precision Medicine. G.A.A. acknowledges support from the Russian Federation Fundamental Research Program for the long-term period for 2021–2030. V.M.P. acknowledges partial support from National Human Genome Research Institute (U01 HG010245).
Footnotes
Conflicts of Interest:
Indiana University School of Medicine Pharmacogenomics Laboratory is a fee-for-service clinical laboratory that offers clinical pharmacogenetic testing. V.M.L. is co-founder and shareholder of PersoMedix AB, CEO and shareholder of HepaPredict AB and discloses consultancy work for Enginzyme AB. Sema4 is a fee-for-service clinical laboratory that offers clinical pharmacogenetic testing. B.E.R. is a paid employee of Sema4 and the founder and C.E.O. of Phoenix Laboratory Consulting, LLC.
All other authors declared no competing interests for this work.
References
- (1).Aoyama T et al. Cytochrome P-450 hPCN3, a novel cytochrome P-450 IIIA gene product that is differentially expressed in adult human liver. cDNA and deduced amino acid sequence and distinct specificities of cDNA-expressed hPCN1 and hPCN3 for the metabolism of steroid hormones and cyclosporine. J Biol Chem 264, 10388–95 (1989). [PubMed] [Google Scholar]
- (2).Schuetz JD, Molowa DT & Guzelian PS Characterization of a cDNA encoding a new member of the glucocorticoid-responsive cytochromes P450 in human liver. Arch Biochem Biophys 274, 355–65 (1989). [DOI] [PubMed] [Google Scholar]
- (3).Jounaidi Y, Hyrailles V, Gervot L & Maurel P Detection of CYP3A5 allelic variant: a candidate for the polymorphic expression of the protein? Biochem Biophys Res Commun 221, 466–70 (1996). [DOI] [PubMed] [Google Scholar]
- (4).Hustert E et al. The genetic determinants of the CYP3A5 polymorphism. Pharmacogenetics 11, 773–9 (2001). [DOI] [PubMed] [Google Scholar]
- (5).Kuehl P et al. Sequence diversity in CYP3A promoters and characterization of the genetic basis of polymorphic CYP3A5 expression. Nat Genet 27, 383–91 (2001). [DOI] [PubMed] [Google Scholar]
- (6).Gaedigk A et al. The Pharmacogene Variation (PharmVar) Consortium: Incorporation of the Human Cytochrome P450 (CYP) Allele Nomenclature Database. Clin Pharmacol Ther 103, 399–401 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (7).PharmVar. P450 Nomenclature site – Archive; previoulsy at cypalleles.ki/se <https://www.pharmvar.org/htdocs/archive/index_original.htm;
- (8).Dai et al. , In vitro metabolism of cyclosporine A by human kidney CYP3A5. Biochem Pharmacol. 2004. Nov 1;68(9):1889–902. doi: 10.1016/j.bcp.2004.07.012. [DOI] [PubMed] [Google Scholar]
- (9).Saiz-Rodriguez M et al. Polymorphisms associated with fentanyl pharmacokinetics, pharmacodynamics and adverse effects. Basic Clin Pharmacol Toxicol 124, 321–9 (2019). [DOI] [PubMed] [Google Scholar]
- (10).Williams ML, Kannankeril PJ, Breeyear JH, Edwards TL, Van Driest SL & Choi L Effect of CYP3A5 and CYP3A4 Genetic Variants on Fentanyl Pharmacokinetics in a Pediatric Population. Clin Pharmacol Ther, (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (11).Hsu MH & Johnson EF Active-site differences between substrate-free and ritonavir-bound cytochrome P450 (CYP) 3A5 reveal plasticity differences between CYP3A5 and CYP3A4. J Biol Chem 294, 8015–22 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (12).Wu JJ et al. A Naturally Occurring Isoform-Specific Probe for Highly Selective and Sensitive Detection of Human Cytochrome P450 3A5. J Med Chem 60, 3804–13 (2017). [DOI] [PubMed] [Google Scholar]
- (13).Niwa T, Murayama N, Emoto C & Yamazaki H Comparison of kinetic parameters for drug oxidation rates and substrate inhibition potential mediated by cytochrome P450 3A4 and 3A5. Curr Drug Metab 9, 20–33 (2008). [DOI] [PubMed] [Google Scholar]
- (14).Lu Y, Hendrix CW & Bumpus NN Cytochrome P450 3A5 plays a prominent role in the oxidative metabolism of the anti-human immunodeficiency virus drug maraviroc. Drug Metab Dispos 40, 2221–30 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (15).Li X et al. Characterization of T-5 N-oxide formation as the first highly selective measure of CYP3A5 activity. Drug Metab Dispos 42, 334–42 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (16).Hesselink DA, Bouamar R, Elens L, van Schaik RH & van Gelder T The role of pharmacogenetics in the disposition of and response to tacrolimus in solid organ transplantation. Clin Pharmacokinet 53, 123–39 (2014). [DOI] [PubMed] [Google Scholar]
- (17).Dai Y et al. In vitro metabolism of cyclosporine A by human kidney CYP3A5. Biochem Pharmacol 68, 1889–902 (2004). [DOI] [PubMed] [Google Scholar]
- (18).Le Meur Y et al. CYP3A5*3 influences sirolimus oral clearance in de novo and stable renal transplant recipients. Clin Pharmacol Ther 80, 51–60 (2006). [DOI] [PubMed] [Google Scholar]
- (19).Josephson F et al. CYP3A5 genotype has an impact on the metabolism of the HIV protease inhibitor saquinavir. Clin Pharmacol Ther 81, 708–12 (2007). [DOI] [PubMed] [Google Scholar]
- (20).Lu Y, Fuchs EJ, Hendrix CW & Bumpus NN CYP3A5 genotype impacts maraviroc concentrations in healthy volunteers. Drug Metab Dispos 42, 1796–802 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (21).Wong M, Balleine RL, Collins M, Liddle C, Clarke CL & Gurney H CYP3A5 genotype and midazolam clearance in Australian patients receiving chemotherapy. Clin Pharmacol Ther 75, 529–38 (2004). [DOI] [PubMed] [Google Scholar]
- (22).Dennison JB, Jones DR, Renbarger JL & Hall SD Effect of CYP3A5 expression on vincristine metabolism with human liver microsomes. J Pharmacol Exp Ther 321, 553–63 (2007). [DOI] [PubMed] [Google Scholar]
- (23).Kim KA, Park PW, Lee OJ, Kang DK & Park JY Effect of polymorphic CYP3A5 genotype on the single-dose simvastatin pharmacokinetics in healthy subjects. J Clin Pharmacol 47, 87–93 (2007). [DOI] [PubMed] [Google Scholar]
- (24).Noll EM et al. CYP3A5 mediates basal and acquired therapy resistance in different subtypes of pancreatic ductal adenocarcinoma. Nat Med 22, 278–87 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (25).Ingelman-Sundberg M & Lauschke VM Can CYP Inhibition Overcome Chemotherapy Resistance? Trends Pharmacol Sci 41, 503–6 (2020). [DOI] [PubMed] [Google Scholar]
- (26).Birdwell KA et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guidelines for CYP3A5 Genotype and Tacrolimus Dosing. Clin Pharmacol Ther 98, 19–24 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (27).CPIC. Gene/drug pairs <https://cpicpgx.org/genes-drugs/>.
- (28).Dennison JB, Mohutsky MA, Barbuch RJ, Wrighton SA & Hall SD Apparent high CYP3A5 expression is required for significant metabolism of vincristine by human cryopreserved hepatocytes. J Pharmacol Exp Ther 327, 248–57 (2008). [DOI] [PubMed] [Google Scholar]
- (29).Dennison JB, Kulanthaivel P, Barbuch RJ, Renbarger JL, Ehlhardt WJ & Hall SD Selective metabolism of vincristine in vitro by CYP3A5. Drug Metab Dispos 34, 1317–27 (2006). [DOI] [PubMed] [Google Scholar]
- (30).Shapiro R et al. A prospective, randomized trial of tacrolimus/prednisone versus tacrolimus/prednisone/mycophenolate mofetil in renal transplant recipients. Transplantation 67, 411–5 (1999). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (31).Wallemacq P et al. Opportunities to optimize tacrolimus therapy in solid organ transplantation: report of the European consensus conference. Ther Drug Monit 31, 139–52 (2009). [DOI] [PubMed] [Google Scholar]
- (32).Dai Y et al. Effect of CYP3A5 polymorphism on tacrolimus metabolic clearance in vitro. Drug Metab Dispos 34, 836–47 (2006). [DOI] [PubMed] [Google Scholar]
- (33).Kamdem LK et al. Contribution of CYP3A5 to the in vitro hepatic clearance of tacrolimus. Clin Chem 51, 1374–81 (2005). [DOI] [PubMed] [Google Scholar]
- (34).Hesselink DA et al. CYP3A5 genotype is not associated with a higher risk of acute rejection in tacrolimus-treated renal transplant recipients. Pharmacogenet Genomics 18, 339–48 (2008). [DOI] [PubMed] [Google Scholar]
- (35).Thervet E et al. Optimization of initial tacrolimus dose using pharmacogenetic testing. Clin Pharmacol Ther 87, 721–6 (2010). [DOI] [PubMed] [Google Scholar]
- (36).Moes DJ et al. Population pharmacokinetics and pharmacogenetics of once daily tacrolimus formulation in stable liver transplant recipients. Eur J Clin Pharmacol 72, 163–74 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (37).Li D, Lu W, Zhu JY, Gao J, Lou YQ & Zhang GL Population pharmacokinetics of tacrolimus and CYP3A5, MDR1 and IL-10 polymorphisms in adult liver transplant patients. J Clin Pharm Ther 32, 505–15 (2007). [DOI] [PubMed] [Google Scholar]
- (38).Zhu HJ, Yuan SH, Fang Y, Sun XZ, Kong H & Ge WH The effect of CYP3A5 polymorphism on dose-adjusted cyclosporine concentration in renal transplant recipients: a meta-analysis. Pharmacogenomics J 11, 237–46 (2011). [DOI] [PubMed] [Google Scholar]
- (39).Sun B et al. Influence of CYP3A and ABCB1 polymorphisms on cyclosporine concentrations in renal transplant recipients. Pharmacogenomics 18, 1503–13 (2017). [DOI] [PubMed] [Google Scholar]
- (40).Tang HL, Ma LL, Xie HG, Zhang T & Hu YF Effects of the CYP3A5*3 variant on cyclosporine exposure and acute rejection rate in renal transplant patients: a meta-analysis. Pharmacogenet Genomics 20, 525–31 (2010). [DOI] [PubMed] [Google Scholar]
- (41).Hesselink DA et al. Genetic polymorphisms of the CYP3A4, CYP3A5, and MDR-1 genes and pharmacokinetics of the calcineurin inhibitors cyclosporine and tacrolimus. Clin Pharmacol Ther 74, 245–54 (2003). [DOI] [PubMed] [Google Scholar]
- (42).Kreutz R, Zurcher H, Kain S, Martus P, Offermann G & Beige J The effect of variable CYP3A5 expression on cyclosporine dosing, blood pressure and long-term graft survival in renal transplant patients. Pharmacogenetics 14, 665–71 (2004). [DOI] [PubMed] [Google Scholar]
- (43).Debette-Gratien M et al. Influence of Donor and Recipient CYP3A4, CYP3A5, and ABCB1 Genotypes on Clinical Outcomes and Nephrotoxicity in Liver Transplant Recipients. Transplantation 100, 2129–37 (2016). [DOI] [PubMed] [Google Scholar]
- (44).Zheng S et al. CYP3A5 gene variation influences cyclosporine A metabolite formation and renal cyclosporine disposition. Transplantation 95, 821–7 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (45).Kreutz R et al. CYP3A5 genotype is associated with longer patient survival after kidney transplantation and long-term treatment with cyclosporine. Pharmacogenomics J 8, 416–22 (2008). [DOI] [PubMed] [Google Scholar]
- (46).Thangavel C, Boopathi E & Shapiro BH Inherent sex-dependent regulation of human hepatic CYP3A5. Br J Pharmacol 168, 988–1000 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (47).Ito A et al. Sex Differences in the Blood Concentration of Tacrolimus in Systemic Lupus Erythematosus and Rheumatoid Arthritis Patients with CYP3A5*3/*3. Biochem Genet 55, 268–77 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (48).Stevens JC et al. Developmental expression of the major human hepatic CYP3A enzymes. J Pharmacol Exp Ther 307, 573–82 (2003). [DOI] [PubMed] [Google Scholar]
- (49).Hakkola J et al. Cytochrome P450 3A expression in the human fetal liver: evidence that CYP3A5 is expressed in only a limited number of fetal livers. Biol Neonate 80, 193–201 (2001). [DOI] [PubMed] [Google Scholar]
- (50).Gillam EM et al. Expression of cytochrome P450 3A5 in Escherichia coli: effects of 5’ modification, purification, spectral characterization, reconstitution conditions, and catalytic activities. Arch Biochem Biophys 317, 374–84 (1995). [DOI] [PubMed] [Google Scholar]
- (51).Yablokov EO et al. A large-scale comparative analysis of affinity, thermodynamics and functional characteristics of interactions of twelve cytochrome P450 isoforms and their redox partners. Biochimie 162, 156–66 (2019). [DOI] [PubMed] [Google Scholar]
- (52).Yamaori S et al. Effects of cytochrome b(5) on drug oxidation activities of human cytochrome P450 (CYP) 3As: similarity of CYP3A5 with CYP3A4 but not CYP3A7. Biochem Pharmacol 66, 2333–40 (2003). [DOI] [PubMed] [Google Scholar]
- (53).Davydov DR, Davydova NY, Sineva EV & Halpert JR Interactions among cytochromes P450 in microsomal membranes: oligomerization of cytochromes P450 3A4, 3A5, and 2E1 and its functional consequences. J Biol Chem 290, 3850–64 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (54).Klomp SD, Manson ML, Guchelaar HJ & Swen JJ Phenoconversion of Cytochrome P450 Metabolism: A Systematic Review. J Clin Med 9, (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (55).Dunvald AD, Jarvinen E, Mortensen C & Stage TB Clinical and Molecular Perspectives on Inflammation-Mediated Regulation of Drug Metabolism and Transport. Clin Pharmacol Ther, (2021). [DOI] [PubMed] [Google Scholar]
- (56).Suzuki Y et al. Factors involved in phenoconversion of CYP3A using 4beta-hydroxycholesterol in stable kidney transplant recipients. Pharmacol Rep 71, 276–81 (2019). [DOI] [PubMed] [Google Scholar]
- (57).Collins JM & Wang D Regulation of CYP3A4 and CYP3A5 by a lncRNA: a potential underlying mechanism explaining the association between CYP3A4*1G and CYP3A metabolism. Pharmacogenet Genomics 32, 16–23 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (58).Whirl-Carrillo M et al. An Evidence-Based Framework for Evaluating Pharmacogenomics Knowledge for Personalized Medicine. Clin Pharmacol Ther 110, 563–72 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (59).PharmGKB. CYP3A5 gene page. <https://www.pharmgkb.org/gene/PA131>.
- (60).PharmGKB. Gene-specific Information Tables for CYP3A5. <https://www.pharmgkb.org/page/cyp3a5RefMaterials>.
- (61).PharmCAT. Pharmacogenomics Clinical Annotation Tool. <https://pharmcat.org/>. [Google Scholar]
- (62).Sangkuhl K et al. Pharmacogenomics Clinical Annotation Tool (PharmCAT). Clin Pharmacol Ther 107, 203–10 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (63).PharmGKB. PGx Publication Tips. https://www.pharmgkb.org/page/curationTips>
- (64).CPIC. Guidelines. <https://cpicpgx.org/guidelines/>.
- (65).Caraballo PJ, Bielinski SJ, St Sauver JL & Weinshilboum RM Electronic Medical Record-Integrated Pharmacogenomics and Related Clinical Decision Support Concepts. Clin Pharmacol Ther 102, 254–64 (2017). [DOI] [PubMed] [Google Scholar]
- (66).Hicks JK, Dunnenberger HM, Gumpper KF, Haidar CE & Hoffman JM Integrating pharmacogenomics into electronic health records with clinical decision support. Am J Health Syst Pharm 73, 1967–76 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (67).Pratt VM et al. Characterization of 137 Genomic DNA Reference Materials for 28 Pharmacogenetic Genes: A GeT-RM Collaborative Project. J Mol Diagn 18, 109–23 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (68).Moyer AM, Rohrer Vitek CR, Giri J & Caraballo PJ Challenges in Ordering and Interpreting Pharmacogenomic Tests in Clinical Practice. Am J Med 130, 1342–4 (2017). [DOI] [PubMed] [Google Scholar]
- (69).Pratt VM et al. Recommendations for Clinical CYP2C9 Genotyping Allele Selection: A Joint Recommendation of the Association for Molecular Pathology and College of American Pathologists. J Mol Diagn 21, 746–55 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (70).Pratt VM et al. Recommendations for Clinical CYP2C19 Genotyping Allele Selection: A Report of the Association for Molecular Pathology. J Mol Diagn 20, 269–76 (2018). [DOI] [PubMed] [Google Scholar]
- (71).Pratt VM et al. Recommendations for Clinical CYP2D6 Genotyping Allele Selection: A Joint Consensus Recommendation of the Association for Molecular Pathology, College of American Pathologists, Dutch Pharmacogenetics Working Group of the Royal Dutch Pharmacists Association, and the European Society for Pharmacogenomics and Personalized Therapy. J Mol Diagn 23, 1047–64 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (72).Genetic Testing Registry (GTR). Central location for voluntary submission of genetic test information by providers. <https://www.ncbi.nlm.nih.gov/gtr/>.
- (73).PharmVar. CYP3A5 gene page. <https://www.pharmvar.org/gene/CYP3A5>.
- (74).Human Genome Variation Society (HGVS); Nomenclature for the description of sequence variations <http://www.hgvs.org/content/guidelines>.
- (75).Database of Genomic Variants (DGV). Database of Genomic Variants. A curated catalogue of human genomic structural variation. <http://dgv.tcag.ca/dgv/app/home>.
- (76).Huddart R et al. Standardized Biogeographic Grouping System for Annotating Populations in Pharmacogenetic Research. Clin Pharmacol Ther 105, 1256–62 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (77).Zhang F & Finkelstein J Inconsistency in race and ethnic classification in pharmacogenetics studies and its potential clinical implications. Pharmgenomics Pers Med 12, 107–23 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (78).PharmVar. Allele Designation and Evidence Level Criteria document. <www.pharmvar.org/documents/Allele_Designation_and_Evidence_Level_Criteria_V2.1.pdf >.
- (79).Single Nucleotide Polymorphism Database (dbSNP). <https://www.ncbi.nlm.nih.gov/snp/>.
- (80).Locus Reference Genomic (LRG) Records. <https://www.lrg-sequence.org/>.
- (81).NCBI. Matched Annotation from NCBI and EMBL-EBI (MANE). <https://www.ncbi.nlm.nih.gov/refseq/MANE/>.
- (82).CPIC. Standard Operating Protocol for Assigning CPIC Allele Clinical Function. <https://cpicpgx.org/resources/cpic-draft-allele-function-sop/>.
- (83).PharmVar. CYP3A Gene Expert Panel. <https://www.pharmvar.org/expert-panels >.
- (84).Personal communication with Cyrine Haidar, St Jude Children’s Research Hospital, Ann Moyer, Mayo Clinic and Charity Nofziger, PharmGenetix GmBH. [Google Scholar]
- (85).Lee SJ et al. Genetic findings and functional studies of human CYP3A5 single nucleotide polymorphisms in different ethnic groups. Pharmacogenetics 13, 461–72 (2003). [DOI] [PubMed] [Google Scholar]
- (86).Personal communication with Cyrine Haidar, St Jude Children’s Research Hospital. [Google Scholar]
- (87).Desta Z et al. PharmVar GeneFocus: CYP2B6. Clin Pharmacol Ther 110, 82–97 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (88).Sangkuhl K et al. PharmVar GeneFocus: CYP2C9. Clin Pharmacol Ther 110, 662–76 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (89).Lauschke VM, Milani L & Ingelman-Sundberg M Pharmacogenomic Biomarkers for Improved Drug Therapy-Recent Progress and Future Developments. AAPS J 20, 4 (2017). [DOI] [PubMed] [Google Scholar]
- (90).Russell LE, Zhou Y, Almousa AA, Sodhi JK, Nwabufo CK & Lauschke VM Pharmacogenomics in the era of next generation sequencing - from byte to bedside. Drug Metab Rev 53, 253–78 (2021). [DOI] [PubMed] [Google Scholar]
- (91).Botton MR et al. PharmVar GeneFocus: CYP2C19. Clin Pharmacol Ther 109, 352–66 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- (92).Gaedigk A et al. CYP2C8, CYP2C9 and CYP2C19 characterization using Next Generation Sequencing and Haplotype Analysis: A GeT-RM Collaborative Project. J Mol Diagn, online ahead of print Feb 5:S1525–1578 (2022). doi: 10.1016/j.jmoldx.2021.12.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- (93).Simons Diversity Project. <https://www.simonsfoundation.org/simons-genome-diversity-project/>.
- (94).PharmVar. Standards. <https://www.pharmvar.org/genes>.
- (95).PharmGKB. CYP3A5 Drug Label Annotations. <https://www.pharmgkb.org/gene/PA131/labelAnnotation>.
- (96).FDA. Pharmacogenomic Biomarkers in Drug Labeling <https://www.fda.gov/drugs/science-research-drugs/table-pharmacogenomic-biomarkers-drug-labeling >.
- (97).FDA. Table of Pharmacogenetics Associations. <https://www.fda.gov/medical-devices/precision-medicine/table-pharmacogenetic-associations>.
- (98).Drug Interactions Flockhart Table™. <https://drug-interactions.medicine.iu.edu/Main-Table.aspx >.
- (99).NCBI. NCBI Reference Sequences database <https://www.ncbi.nlm.nih.gov/refseq/>.
- (100).Chou FC, Tzeng SJ & Huang JD Genetic polymorphism of cytochrome P450 3A5 in Chinese. Drug Metab Dispos 29, 1205–9 (2001). [PubMed] [Google Scholar]
- (101).Shih PS & Huang JD Pharmacokinetics of midazolam and 1’-hydroxymidazolam in Chinese with different CYP3A5 genotypes. Drug Metab Dispos 30, 1491–6 (2002). [DOI] [PubMed] [Google Scholar]
- (102).Saeki M et al. Single nucleotide polymorphisms and haplotype frequencies of CYP3A5 in a Japanese population. Hum Mutat 21, 653 (2003). [DOI] [PubMed] [Google Scholar]
- (103).Patki et al. In vitro metabolism of midazolam, triazolam, nifedipine, and testosterone by human liver microsomes and recombinant cytochromes p450: role of cyp3a4 and cyp3a5) after nifedipine. Drug Metab Dispos. 2003. Jul;31(7):938–44. doi: 10.1124/dmd.31.7.938. [DOI] [PubMed] [Google Scholar]


