Summary
Combinatorial CRISPR screening is useful for investigating synthetic lethality (SL) gene pairs. Here, we detail the steps for dual-gRNA library construction, with the introduction of two backbones, LentiGuide_DKO and LentiCRISPR_DKO. We describe steps for in vitro screening with 22Rv1-Cas9 and SaOS2-Cas9 cells followed by sequencing and data analysis. By introducing two backbones, we optimized the library construction process, facilitated standard pair-end sequencing, and provided options of screening on cells with or without modification of Cas9 expression.
Subject areas: High Throughput Screening, Molecular Biology, CRISPR, Biotechnology and bioengineering
Graphical abstract

Highlights
-
•
Optimized library construction process by introducing two backbones
-
•
In vitro screening gene-gene interaction w/wo Cas9
-
•
Full screening process from gene selection to data analysis
Publisher’s note: Undertaking any experimental protocol requires adherence to local institutional guidelines for laboratory safety and ethics.
Combinatorial CRISPR screening is useful for investigating synthetic lethality (SL) gene pairs. Here, we detail the steps for dual-gRNA library construction, with the introduction of two backbones, LentiGuide_DKO and LentiCRISPR_DKO. We describe steps for in vitro screening with 22Rv1-Cas9 and SaOS2-Cas9 cells followed by sequencing and data analysis. By introducing two backbones, we optimized the library construction process, facilitated standard pair-end sequencing, and provided options of screening on cells with or without modification of Cas9 expression.
Before you begin
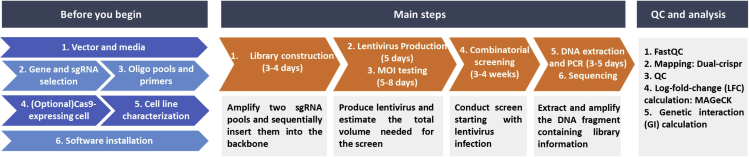
An overview of workflow chart can be found in Figure 1. Following this protocol, pooled library of 62,500 dual-gRNA combinations for targeting more than 1,000 gene pairs was successfully generated. This library was then applied to high-throughput combinatorial CRISPR screen on prostate cancer cell line 22Rv1-Cas9 and osteosarcoma cell line SaOS2-Cas9, to identify synergistic gene pairs that inhibited cancer cell growth when they were disrupted simultaneously.
Figure 1.
Protocol flow chart
In this study, we generated two backbone vectors, LentiGuide_DKO and LentiCRISPR_DKO (DKO refers to double knockout). The only difference between these two vectors is the additional Cas9-expressing sequence driven by EF-1-alpha promoter in LentiCRISPR_DKO. Cas9 is a bacterial protein and is not expressed by eukaryotic cells. Genetic screens in human cell lines with CRISPR technology often start with creation of cell line stably expressing Cas9, or using lentiviral backbone containing Cas9. With the built-in Cas9 expressing cassette, LentiCRISPR_DKO opens the option to perform the combinatorial CRISPR screen using cells that are not engineered to express Cas9.
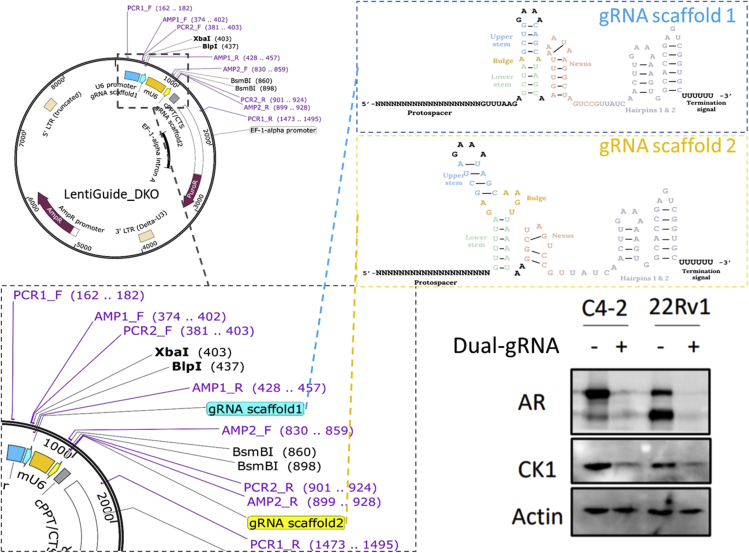
In LentiGuide_DKO (Figure 2), two different small RNA promoters (hU6 and mU6) and two versions of gRNA scaffolds were built into one plasmid. After sequentially inserting two sets of sgRNA, the simultaneously functional knockout (KO) of two genes can be achieved with the presence of Cas9 protein. The introduction of two different sgRNA scaffolds lifted the constraint of using customized Illumina sequencing that is required by currently available combinatorial CRISPR screening method ((Han et al., 2017)). The base sequence difference between two scaffolds allows the specific binding of PCR-2R to the beginning of second scaffold, which makes the second gRNA insertion within the sequencing length (Figure 3). Both versions of scaffold have been used extensively in single sgRNA CRISPR screens (Luo et al., 2008; Hart et al., 2017; Gilbert et al., 2014).
Figure 2.
Backbone design and KO efficiency validation
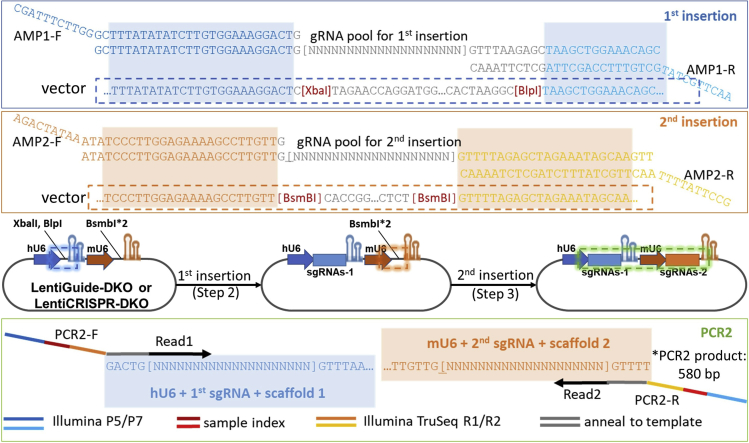
Figure 3.
Overall library construction process and gRNA pools/primer design
This backbone design has been validated in two cell lines (C4-2-Cas9 and 22Rv1-Cas9) with dual-target of AR and CK1. The western blot result showed that both the expression of AR and CK1 were inhibited after functional KO of AR and CK1 using the LentiGuide_DKO plasmid. This result also validated the same performance of the two scaffolds in the newly-built backbone regardless of their differences in the sequence and structure.
Preparation one: Vector and media
Timing: 1–3 days
-
1.
Purchase vector from Addgene: Addgene: LentiGuide_DKO (Plasmid #183193), or LentiCRISPR_DKO (Plasmid #183192). 2 μg of the backbone vector is needed in enzyme digestion to start the library construction (step 2).
-
2.
Ensure all the media are prepared. A detailed list can be found in the sections of key resources table, materials and equipment.
Preparation two: Gene and sgRNA selection
A guideline for choosing gene and sgRNA to prepare for the library construction is provided here. Please refer to Table 1 for the summary of the experiment details of the previous combinational CRISPR screens from several groups.
-
3.
Estimate library size and cell culture scale: The size of library is constrained by the cell culture limit; therefore, estimating the library size is critical before the experiment. For example, under the setting of library with 62,500 dual-gRNA constructions, ∼0.3 MOI, triplicates for each time point and at least 200 cell coverages for each gRNA-gRNA construction during screening, the corresponding pooled dual-gRNA library size is 62,500 and at least 2.5 × 10ˆ8 (62,500∗200∗6/0.3) cells will be needed at the time of infection.
Note: There are basically two scenarios when “coverage” is used in this protocol. One is the coverage when constructing the dual-gRNA pool library (refer as library coverage); another is the coverage when conducting screening (refer as cell coverage). For a library with 62,500 constructions, a total number of 6.25 × 10ˆ6 colonies are required to reach 100 library coverage. Later in the screen, at least 1.25 × 10ˆ7 cells each sample are needed for 200 cell coverage.
-
4.Gene selection: The gene selection is mainly based on experiment objective and library size. Several commonly used selection criteria are listed below:
-
a.Exclude the genes that its loss of function alone has detrimental effect on cell growth. The gene essentiality information can be obtained from previous screen or DepMap database (Broad, 2021). Note that gene essentiality is cancer type and cell-line specific.
-
b.Prioritize the genes that have been validated as drug targets.
-
c.Focus the genes from pathways of interest.
-
d.Include 2–4 gene pairs with known synthetic lethality as positive controls.
-
a.
Table 1.
Current SL research using combinatorial CRISPR technology
| Studies | Gene & sgRNA selection criteria | # of genes, control (ctrl)/safe gRNA and gRNA pairs | Coverages |
Screen details (Cell line, MOI, etc.) | Harvest timea | |
|---|---|---|---|---|---|---|
| Library | Cell | |||||
| Wong et al., 2016 | Epigenetics regulation | 50 genes (3 gRNAs per gene), 3 ctrl gRNAs, 23,409 gRNA pairs | NA | 300 | OVCAR8-ADR-Cas9; MOI 0.3-0.5 | Day 15, 20 |
| (Han et al., 2017) | Non-essential and expressed druggable gene targets (previous screen and DrugBank) | 207 genes (3 gRNAs per gene), 79 safe gRNAs, 21,321gRNA pairs | NA | 1000 | K562-Cas9; 2 replicates |
Day 0, 14 post-selection |
| (Shen et al., 2017) | validated oncogenes, tumor-suppressor genes and cancer-relevant drug targets | 73 genes (3 gRNAs per gene), 12 non-target gRNAs,141,912 gRNA pairs | 100 (in LB medium) | 200 | HeLa-Cas9, 293T-Cas9, A549-Cas9; 2 replicates; MOI 0.1-0.4 | Days 3, 14, 21, 28 post- infection |
| (Horlbeck et al., 2018) | Non-essential genes (CRISPRi v1 growth screen) | 472 genes (1-3 gRNAs per gene), 18 ctrl gRNAs , 1,044,484 gRNA pairs | 25 | 500 (initial 250) | K562-Cas9, Jurkat-Cas9; 2 replicates; | Day 0, ∼10 doubling |
| (Zhao et al., 2018) | Genes represent glycolysis and the pentose phosphate pathway (PPP) | 51 genes,11,475 gRNA pairs | 50 | 80 | HeLa-Cas9, A549-Cas9; 2 replicates; MOI 0.1-0.3 | Days 3, 14, 21, 28 post-infection |
| (Diehl et al., 2021) | Autophagy pathway | 160 genes, 247,032 gRNA pairs | NA | 20, 200 | RPE1-Cas9; triplicates; MOI 0.5 | Days 2, 14 post- infection, |
| (Ito et al., 2021) | Paralog families | 3,284 genes(6 gRNAs per gene), 110,874 gRNA pairs | NA | 750-1000 | 11 cancer cell lines with Cas9; triplicates; MOI 0.3-0.5 | NA |
| (Parrish et al., 2021) | Paralog and singleton essentiality | 2,060 gene pairs, 33,170 gRNA pairs | 1000 | 500 | PC9-Cas9; triplicates; MOI 0.3 | Plasmid pool, ∼12 doubling |
| (Thompson et al., 2021) | Paralog families and SL partners from previous studies | 1,191 gene pairs (3-5 gRNAs per gene), 41,838 gRNA pairs | NA | 1000 | RPE1-Cas9, A375-Cas9, MeWo-Cas9; triplicates; MOI 0.3 | Days 7, 14, 28 |
| This study | Genes from cell death pathways specified for prostate cancer; targetable genes for osteosarcoma | 1,225 gene pairs (4 gRNAs per gene, 62,500 gRNA pairs) | 10 | 200 | 22RV1-Cas9; SaOS2-Cas9; MOI 0.3 |
Day 0, 12 or 14 post-selection |
The sample harvest time is measured differently among different studies. For example, Han et al. harvested the samples at post-infection and puromycin selection day 0 and day 14. No info is presented if the time measurement is not specified in the original paper (such as the research done by Wong et al., 2016).
In this study, we selected 56 genes that can be targeted by available drugs (Drug Bank (Wishart et al., 2018), DepMap) and have low essentiality scores (DepMap). Among them, HDAC1-WEE1, HDAC2-WEE1 and KDR-KDM6B serve as the positive controls (Yu et al., 2015; Wang et al., 2020).
-
5.
sgRNA selection: typically 3 sgRNAs are required for each gene to ensure the quality of the screening. The tools for CRISPR/Cas9 sgRNA design were summarized by several groups (Cui et al., 2018; Doench et al., 2016; Liu et al., 2020).
The genes, sgRNAs and corresponding sources for generating our library are listed in the Table S1. In the project GitHub page, the sgRNA information and R code for information integration are provided under the sgRNA_info folder. In our library, 4 corresponding sgRNAs were selected for each gene with reference to available sgRNA sequence from current commonly-used CRISPR libraries, including TKOv3 (Mair et al., 2019), hGECKOv2 (Sanjana et al., 2014) and KinomeKO /Brunello(Doench et al., 2016), as well as their VBC (Vienna Bioactivity CRISPR) scores (Michlits et al., 2020). We prioritize the gRNAs that have overlapping on the abovementioned libraries. Additionally, another 26 safe gRNAs that target non-functional parts of genome were added into the pooled library as negative controls.
CRITICAL: Make sure to EXCLUDE the gRNAs contain the three restriction sites (XbaI, BlpI, BsmBI) used in the library construction.
Preparation three: Design oligo pools and primers
During the library construction process, the DNA fragments of digested vector and amplified sgRNA pools with 20-bp overlap at both ends can be ligated by HiFi DNA assembly. Two sequential HiFi DNA assemblies generate the library containing two sgRNAs. The designing for oligo pools and primers can be found in Figure 3.
Note: An extra G is added to the 5′-end of gRNA, since human promoter (hU6) prefers a 'G' at the transcription start site to have high expression, though it is possible for the plasmid to still express without the G (Wang et al., 2019).
Note: We recommend using IDT UltramerTM DNA oligonucleotides for PCR2 primer.
-
6.
Order oligo pools from commercial site with added 5′-flank and 3′-flank sequence to the gRNA insertion. For reference, we bought the oligo pools from oPools Oligo Pools (10 pmol/oligo) from IDT.
| 5′-flank | 3′-flank | |
|---|---|---|
| gRNA pool for 1st insertion | AAGTATCCCTTGGAGAACCACCTTG | GTTTAAGAGCTAAGCTGGAAACAGC |
| gRNA pool for 2nd insertion | TATCTTGTGGAAAGGACGAAACACC | GTTTTAGAGCTAGAAATAGCAAGTT |
-
7.
The primers are listed in the table below:
| Primers | Sequence | Note |
|---|---|---|
| AMP1 _F | CGATTTCTTGGCTTTATATATCTTGTGGAAAGGACTG | Amplification of 1st set of gRNA insertion (step 1) |
| AMP1 _R | AACTTGCTATGCTGTTTCCAGCTTAGCTCTTAAAC | |
| AMP2 _F | AGACTATAAATATCCCTTGGAGAAAAGCCTTGTT | Amplification of 2nd set of gRNA insertion (step 1) |
| AMP2 _R | GCCTTATTTTAACTTGCTATTTCTAGCTCTAAAAC | |
| PCR1_F∗ | GAGGGCCTATTTCCCATGATT | 1st round of the PCR after DNA extraction (step 20) |
| PCR1_R | GTTGCGAAAAAGAACGTTCACGG |
∗ PCR1_F can be used in Sanger sequencing in step2 – optional.
PCR2_F:
AATGATACGGCGACCACCGAGATCTACACNNNNNNNNACACTCTTTCCCTACACGACGCTCTTCCGATCTTTATATATCTTGTGGAAAGGACTG.
PCR2_R:
CAAGCAGAAGACGGCATACGAGATNNNNNNNNGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCTTAACTTGCTATTTCTAGCTCTAAA.
Annotation: Illumina P5/P7, sample index (i5/i7 index), Illumina TruSeq R1/R2, anneal to template (22 bp). The i5/i7 indexes are listed in the table below:
| i7 index name | i7 sequence | i7 index for sample sheet | i5 index name | i5 sequence | -i5 index for sample sheet |
|---|---|---|---|---|---|
| D701 | CGAGTAAT | ATTACTCG | D501 | TATAGCCT | TATAGCCT |
| D702 | TCTCCGGA | TCCGGAGA | D502 | ATAGAGGC | ATAGAGGC |
| D704 | GGAATCTC | GAGATTCC | D503 | CCTATCCT | CCTATCCT |
| D705 | TTCTGAAT | ATTCAGAA | D504 | GGCTCTGA | GGCTCTGA |
| D706 | ACGAATTC | GAATTCGT | D505 | AGGCGAAG | AGGCGAAG |
| D707 | AGCTTCAG | CTGAAGCT | D506 | TAATCTTA | TAATCTTA |
Note: The number of PCR2 primer pairs should be equal to the total number of screen samples, and each sample should have a different i5/i7 index combination.
Preparation four: Cas9-expressing cell line generation
Timing: 1–2 weeks
Note: You may skip this step if you choose to use lentiCRISPR-DKO vector.
-
8.
Please refer to Sanjana et al. (Sanjana et al., 2014) for generating the Cas9-expressing cell lines.
Preparation five: Cell line characterization
Timing: 1–2 weeks
-
9.
Ensure cell line is mycoplasma free before starting screen.
-
10.
Determine optimal cell plating density and passage number, typically in 15 cm plates. Measure and record the approximate cell doubling time.
-
11.
Determine puromycin and polybrene (up to 8 μg/mL)/protamine sulfate (up to 5 μg/mL) sensitivity of cell lines. Concentration of puromycin to be used in screen should be 0.5–1 μg/mL higher than that required to kill all uninfected cells in at least 48 h. The antibiotic selection time in the main step 14 should be the same here.
Preparation six: Software installation
Timing: 0.5 h
-
12.
Mali-dual-crispr-pipeline: This pipeline is provided through conda and can be installed on any linux-64 or osx-64 platform (Windows installation is not available at this time). A detailed installation process can be found at https://github.com/ucsd-ccbb/mali-dual-crispr-pipeline.
-
13.
MAGeCK: MAGeCK and associated software offer a range of functions and can be installed by conda/bioconda, Docker Image, or source code. We used MAGeCK RRA in the log-fold-change calculation. A detailed introduction and installation may be found at https://sourceforge.net/projects/mageck/.
-
14.
R and R-Studio.
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Bacterial and virus strains | ||
| psPAX2 | Didier Trono | AddGene #12260 |
| pMD2.G | Didier Trono | AddGene #12259 |
| Endura Electro Competent Cells | Thermo Fisher Scientific/Lucigen | 501047945 |
| Chemicals, peptides, and recombinant proteins | ||
| Bovine Serum Albumin | Sigma-Aldrich | A9418 |
| Polybrene Infection / Transfection Reagent | Sigma-Aldrich | TR-1003-G |
| Puromycin | Thermo Fisher Scientific | A1113803 |
| Dulbecco’s Modified Eagle Medium (DMEM) | Thermo Fisher Scientific | 11965118 |
| GlutaMAX Supplement | Thermo Fisher Scientific | 35050061 |
| Sodium Pyruvate (100 mM) | Thermo Fisher Scientific | 11360070 |
| Pen-Strep(5,000 U/mL) | Thermo Fisher Scientific | 15070063 |
| HEPES (1 M) | Thermo Fisher Scientific | 15630080 |
| Opti-MEM™ I Reduced Serum Medium | Thermo Fisher Scientific | 31985070 |
| Fetal Bovine Serum (FBS) | VWR/Avantor | 97068-091 |
| EZ-VISION Blue-Light DNA Dye | VWR | 10791-798 |
| XbaI | New England Biolabs | R0145S |
| BlpI | New England Biolabs | R0585S |
| BsmBI-v2 | New England Biolabs | R0739S |
| X-tremeGENE™ 9 DNA Transfection Reagent | Roche/Sigma | 6365787001 |
| Critical commercial assays | ||
| NEBNext Ultra II Q5 Master Mix | New England Biolabs | M0544L |
| NEBuilder HiFi DNA Assembly Cloning Kit | New England Biolabs | E5520S |
| QIAquick Gel Extraction Kit (50) | QIAGEN | 28704 |
| HiSpeed Plasmid Maxi Kit | QIAGEN | 12663 |
| QIAprep Spin Miniprep Kit | QIAGEN | 27106X4 |
| JetQuick Blood and Cell Culture DNA Maxiprep Kit | Thermo Fisher Scientific | A30706 |
| Deposited data | ||
| Sampler data and data analysis | Lang Li Lab (this paper) | https://github.com/Shantang3/Combinatorial-CRISPR-Screen-Anaylysis-Pipeline |
| Experimental models: Cell lines | ||
| 22RV1 with Cas9 expression | Xiaoqi Liu Lab | |
| SaOS2 with Cas9 expression | Lang Li Lab | |
| Oligonucleotides | ||
| oPools Oligo Pools | This paper | IDT |
| PCR Primers | This paper | Thermo Fisher, A15612 |
| UltramerTM DNA oligonucleotides | This paper | IDT |
| Recombinant DNA | ||
| LentiGuide_DKO | Lang Li Lab (this paper) | Addgene #183193 |
| LentiCRISPR_DKO | Lang Li Lab (this paper) | Addgene #183192 |
| LentiGuide_DKO based library | Lang Li Lab (this paper) | |
| Software and algorithms | ||
| dual_crispr | Amanda Birmingham, CCBB, UCSD | https://github.com/ucsd-ccbb/mali-dual-crispr-pipeline |
| MAGeCK | Wei Li, et al. | https://sourceforge.net/projects/mageck/ |
| R (version at least 4.0.3) | https://www.r-project.org/ | |
| R Studio | https://www.rstudio.com/ | |
| QC and Hit Identification | This paper | https://github.com/Shantang3/Combinatorial-CRISPR-Screen-Anaylysis-Pipeline |
| Other | ||
| Gene Pulser/MicroPulser Electroporation Cuvettes, 0.2 cm gap | Bio-Rad | 1652086 |
| Petri Dishes with Clear Lid, 150 × 15 MM,100/CS | Thermo Fisher Scientific | FB0875714 |
| Cell Culture Dish | VWR | 10062-882 |
| Plate Tissue Culture 6 Wells ST | VWR | 10062-892 |
Materials and equipment
Cell growth media
| Reagent | Final concentration | Amount |
|---|---|---|
| DMEM | N/A | 500 mL |
| FBS | N/A | 50 mL |
| GlutaMAX Supplement | N/A | 5 mL |
| Sodium Pyruvate (100 mM) | ∼1 mM | 5 mL |
| Pen-Strep(5,000 U/mL) | N/A | 5 mL |
| HEPES (1 M) | ∼ 10 nM | 5 mL |
| Total | N/A | 570 mL |
Store at 4°C up to two months.
None-antibiotics media
| Reagent | Final concentration | Amount |
|---|---|---|
| DMEM | N/A | 500 mL |
| FBS | 10% | 50 mL |
| GlutaMAX Supplement | N/A | 5 mL |
| Sodium Pyruvate (100 mM) | ∼1 mM | 5 mL |
| HEPES (1 M) | ∼ 10 nM | 5 mL |
| Total | N/A | 565 mL |
Store at 4°C up to two months.
Viral harvest media- Serum-Free, High-BSA 293T growth media
| Reagent | Final concentration | Amount |
|---|---|---|
| DMEM | N/A | 500 mL |
| BSA | 1.1 g/100 mL | 5.5 g |
| GlutaMAX Supplement | N/A | 5 mL |
| Sodium Pyruvate (100 mM) | ∼1 mM | 5 mL |
| Pen-Strep (5,000 U/mL) | 50 U/mL | 5 mL |
| Total | N/A | 515 mL |
Store at 4°C up to two months.
Step-by-step method details
Part 1: Library construction & amplification
Timing: 3–4 days
This part generates the customized dual-gRNA screening library that contains 2 promoters (hU6 and mU6), 2 gRNAs, and 2 scaffolds of different versions within one vector. Briefly, the amplified gRNA pool inserts and digested vectors were ligated via their compatible ends.
Note: Refer to troubleshooting 1 for reducing recombination and troubleshooting 2 to improve efficiency.
-
1.
(Day 1) Oligo pool amplification: assemble two PCR reactions for amplifying two gRNA pools separately on ice using corresponding primers, then follow the cycling conditions for each PCR reaction.
PCR reaction master mix: amplification of gRNA pool for 1st insertion
| Reagent | Amount |
|---|---|
| gRNA pool for 1st insertion (100 nM) | 5 μL |
| Primer AMP1 _F (10 μM) | 2.5 μL |
| Primer AMP1 _R (10 μM) | 2.5 μL |
| 2× NEBNext Μltra II Q5 Master Mix | 25 μL |
| ddH2O | 15 μL |
PCR reaction master mix: amplification of gRNA pool for 2nd insertion
| Reagent | Amount |
|---|---|
| gRNA pool for 2nd insertion (100 nM) | 5 μL |
| Primer AMP2 _F (10 μM) | 2.5 μL |
| Primer AMP2 _R (10 μM) | 2.5 μL |
| 2× NEBNext Μltra II Q5 Master Mix | 25 μL |
| ddH2O | 15 μL |
PCR cycling conditions
| Steps | Temperature | Time | Cycles |
|---|---|---|---|
| Initial Denaturation | 98°C | 30 s | 1 |
| Denaturation | 98°C | 10 s | 15 cycles |
| Annealing | 66°C | 30 s | |
| Extension | 72°C | 10 min | |
| Final extension | 72°C | 4 min | 1 |
| Hold | 4°C | Forever | |
After PCR amplification, run out the reaction on an EZ-VISION Blue-Light DNA Dye-stained 1% agarose gel. Visualize and cut the DNA fragment at ∼90 bp, then extract DNA using the QIAGEN Gel Extraction Kit according to the manufacturer’s instructions, elute in 20 μL ddH2O.
CRITICAL: Elute the amplified gRNA and digested vector in ddH2O to avoid introduction of the excess salt, which might influence the electroporation in later step.
Note: A guide line about the agarose gel concentration selection and optimization could be found at https://www.qiagen.com/us/knowledge-and-support/knowledge-hub/bench-guide/dna/analysing-dna/dna-analysis-using-analytical-gels.
-
2.(Day 1–2) Insert first set of gRNAs. Troubleshooting 1 and troubleshooting 2.
-
a.Vector linearization (could be done in parallel to step 1): assemble the following reaction on ice and incubate at 37°C for 1 h.Enzyme digestion reaction master mix
Reagent Amount Vector: LentiGuide_DKO 2 μg XbaI 1 μL BlpI 1 μL 10× NEBuffer CutSmart 5 μL ddH2O To 50 μL Then run out the sample on an EZ-VISION Blue-Light DNA Dye -stained 1% agarose gel. Extract linearized vector using QIAGEN Gel Extraction Kit, elute with 30 μL ddH2O. -
b.HiFi assembling: the molar ratio between the fragments of linearized vector: insertion (gRNA pools) is optimized to 1:100 with 100 ng of linearized vector. NEBioCacμlator helps to convert dsDNA mass to moles of dsDNA (or vice versa): https://nebiocalculator.neb.com/#!/dsdnaamt.Assemble the reaction on ice, incubate at 50°C for 60 min (use thermocycler), then proceed to transformation (step 1c) directly.HiFi reaction master mix
Reagent Length Mass Molar Amount Linearized LentiGuide_DKO 8750 bp 100 ng 18.5 fmol (Based on concentration) 1st gRNA pool insertion 90 bp 103 ng 1850 fmol (Based on concentration) 2× HiFi DNA Assembly Master Mix – – – 10 μL ddH2O – – – To 20 μL -
c.(Afternoon) Transformation by electroporation:
-
i.Calculate the number of reactions needed based on the library size and coverage. For example, we usually get around 25,000–30, 000 colonies per reaction. For a gRNA pool with 250 gRNA, 1–2 electroporation reactions are required to reach at least 100 coverages (250∗100 = 25,000) for each gRNA.
-
ii.Prepare MicroPulser Cuvettes (2 mm) on ice for at least 10 min. Set up the electroporator with conditions of voltage 3,000 V, Capacitance 25 μF, resistance 200 Ω, cuvette 2 mm.
-
iii.Thaw Endura Electrocompetent cells on ice for 15 min.
-
iv.For each reaction: add 1 μL of the HiFi reaction product to Endura Electrocompetent cells, mix them gently. Then transfer the mixture into MicroPulser Cuvettes, electroporate at the setting listed in step ii, add 975 of Recovery Medium ASAP.
 CRITICAL: Use fresh HiFi product for the transformation each time.
CRITICAL: Use fresh HiFi product for the transformation each time. CRITICAL: Quickly adding the Recovery Medium into the cuvette ASAP after electroporation is one key step. The electroporation efficiency will dramatically decrease if this step is delayed.Note: One trial run is suggested to test how many colonies can grow for each transformation reaction. Then calculate the coverage and corresponding number of reactions needed.
CRITICAL: Quickly adding the Recovery Medium into the cuvette ASAP after electroporation is one key step. The electroporation efficiency will dramatically decrease if this step is delayed.Note: One trial run is suggested to test how many colonies can grow for each transformation reaction. Then calculate the coverage and corresponding number of reactions needed. -
v.Transfer cells mixture to a bacterial culture tube with additional 1 mL recovery medium (or SOC medium). Shake the tubes for 1 h at 37°C in a shaking incubator.
-
i.
-
d.Titer the library: Set up dilutions for counting colonies: set up serial dilution (1:10, 1:100, 1: 1000,…) of the culture and plate onto 10 cm ampicillin LB agar plate. Count the colonies next morning to calculate the library coverage. The diluted plate is used to calculate the total colony number to ensure library coverage.
-
e.Plate the library: Spread the original culture onto pre-warmed 15 cm ampicillin LB agar plate (0.5 mL/plate), incubate at 30°C for approximate 18 h.
 CRITICAL: 30°C incubation is intended to minimize recombination of the lentiviral plasmid library.
CRITICAL: 30°C incubation is intended to minimize recombination of the lentiviral plasmid library. -
f.Harvest colonies and maxi-prep: Transfer 6 mL of LB+ ampicillin medium on one 15 cm plate, scrape the colonies off with a cell spreader. Then use another 6 mL of LB+ ampicillin medium to wash the plate. Collect all colonies and proceed to max-prep according to QIAGEN Plasmid Kit procedures.Optional: Sanger Sequencing. To ensure the success of gRNA pool’s insertion, pick several colonies from the plate (2d or 2e) to 4 mL LB culture with ampicillin. Shake the tubes for at least 8 h at 37°C. Use QIAGEN Mini Prep Kit to extract the plasmid and send it to Sanger Sequencing with primer of PCR1_F.
-
a.
-
3.(Day 2–3) Insert second set of gRNAs: The process is similar to step 2 of inserting first set of gRNA pools. The only difference is the enzyme digestion setting.
-
a.Vector linearization: assemble the following reaction on ice and incubate at 55°C for 1 h.Enzyme digestion reaction master mix
Reagent Amount Vector: LentiGuide_DKO+1st gRNA 2 μg BsmbI_v2 1 μL 10× N NEBuffer r3.1 5 μL ddH2O To 50 μL Then run out the sample on an EZ-VISION Blue-Light DNA Dye -stained 1% agarose gel. Extract linearized DNA using QIAGEN Gel Extraction Kit, elute with 30 μL ddH2O. PCR purification kit could be used on the enzyme digestion product to save time. -
b.Repeat HiFi assembling as steps 2b to 2f. Harvest colonies and max prep are the same as step2.Note: Step 3 usually requires much more reactions to reach appropriate library coverage, considering the combination of 2 sgRNAs. For example, for two sets of gRNA pools with 250 different gRNAs, there will be 62,500 combinations. To reach 10 coverages for each gRNA-gRNA construction, 20 electroporate reactions (62500∗10/30,000) are needed.
-
a.
-
4.
Library QC: Directly proceed to steps 20–24 and data analysis part. Sequencing the whole library at least 50× coverage (refer to step 23 for how to choose sequencing service) for the QC purpose. Pay attention to mapping ratio, number/percentage of missed gRNA pairs, the distribution and evenness of the gRNA-gRNA constructions.
-
5.
Library amplification: depending on the experiment designs, such as cell coverage and number of replicates, the amount of library generated from steps 1–3 might be insufficient. In this case, amplify the pooled library by electroporation as described in steps 2c–2e. Instead of using 1 μL of HiFi reaction product, dilute the pooled library to 50 ng/μL and then use 2 μL 50 ng/μL pooled library for each reaction.
Part 2: Large scale lentivirus production
Timing: 5 days
This part generates lentivirus that will be used to integrate dual-gRNA cassette into cellular genome during the screening steps. In this protocol we transfected 293T cells with psPAX2, pMD2.G and the dual-gRNA plasmid library for lentiviral production. psPAX2 and pMD2.G are the second generation lentiviral packaging plasmid and VSV-G envelope expressing plasmid, respectively.
CRITICAL: Biosafety precautions: Double gloves should be worn, and all virus waste requires incubation in 20% bleach for 30 min before disposal. No vacuum pumps can be used.
-
6.
Day 1. Seed 8–9 × 106/15 cm plate 293T packaging cells (recommended: passage number < 15) in 20 mL none-antibiotic growth media. For 1 L virus production prepare ∼60 plates. Incubate cells for 24 h (37°C, 5% CO2), or until the following afternoon. The cells should be ∼70% confluent at the moment of transfection.
-
7.Day 2 afternoon. Transfect 293T cells:
-
a.Prepare the mixture of 3 transfection plasmid (∼1:1:1 molar ratio) in OptiMEM:Transfection plasmid mix (per 15 cm plate)
Plasmid Mass Amount per plate Total amount psPAX2 6.5 μg per plate pMD2.G 3.6 μg per plate Library 5.4 μg per plate OptiMEM – To 200 μL -
b.For each plate, prepare one 1× OptiMEM and X-tremeGENE transfection reagent (i.e., 60 tubes OptiMEM and X-tremeGENE mix will be needed for 60 plates): directly add 48 μL X-tremeGENE into the tubes with 600 μL OptiMEM, and gently mix them by flicking the tubes.
 CRITICAL: Do not mix by pipetting or vertex here.
CRITICAL: Do not mix by pipetting or vertex here. -
c.After the incubation, add 200 μL OptiMEM with transfection plasmids drop-wise to each tube with OptiMEM and X-tremeGENE dilutions (1:3 ratio), then mix gently.
-
d.Carefully transfer the transfection mix to 293T packaging cells by drop-wise adding in circular motion.
-
e.Incubate cells for 18 h (37°C, 5% CO2).
-
a.
-
8.
Day 3 morning. Replace the media containing transfection components with high-BSA growth media for viral production (∼20 mL/15 cm plate).
-
9.
Day 4 morning. Harvest media containing lentivirus at ∼40 h post-transfection. Transfer media to 50 mL plastic tube. Spin the media containing lentivirus at 180 × g for 3 min, then collect and aliquot the supernatant into 15/50 mL sterile polypropylene storage tube. Lentivirus should be frozen at −80°C for long-term storage but can be stored temporarily at 4°C for hrs to days.
Note: Aliquot the lentivirus to smaller tubes according to the following steps’ usage to avoid freeze/thaw cycles.
Optional: After harvesting the lentivirus at day 4 morning, add high-BSA growth media again for continuing viral production. Then harvest lentivirus at day 5 morning. Note that the lentivirus amount might drop a lot compared to day-4 batch.
Part 3: Multiplicity of infection (MOI) determine
Timing: 5–8 days
This part tests the titer of lentivirus produced from Part 2 and determine the volume of virus needed to reach the MOI ∼0.3 in the lentivirus transduction of the pooled library in Part 4.
CRITICAL: The MOI of lentivirus must be determined for each batch and under the exact same cell culture conditions, such as media and cell density, used in the screening. Measurements from different conditions cannot be scaled to determine the MOI.
CRITICAL: Biosafety precautions for virus waste. In our institution, all virus waste requires incubation in 20% bleach for 30 min before disposal. No vacuum pumps can be used.
Note: To test MOI, two sets of samples with lentivirus infection are needed. The MOI can be calculated by measuring the ratio of living cells number in the lentivirus + antibiotics group over living cell number in the lentivirus only (no antibiotics selection, 100% survival control) group. Besides the samples with virus, another set of sample without virus but selected by antibiotics is needed for negative control purpose (0% survival control).
Note: The cells are transduced at a concentrated seeding density in 12-well plates while centrifuging (‘spinfection’), to achieve higher transduction efficiency. For normal transduction without spinning, refer to Hart et al. (Hart et al., 2015). If the cells can reach MOI ∼0.3 with sensible volume of lentivirus (<50% medium), normal transduction would be an easier option.
Note: The ideal spinfection conditions can be highly variable across cell type and should be optimized. A detailed protocol for the optimization of lentiviral transduction spinfection can be found at https://portals.broadinstitute.org/gpp/public/dir/download?dirpath=protocols/production&filename=Optimization_of_Lentiviral_Spinfection_Oct2018.pdf.
Note: The antibiotics selection time may vary from 2 to 5 days.
-
10.Day 1. Lentivirus spinfection.
-
a.Thaw a fresh aliquot of pooled lentivirus on ice.
-
b.Harvest cells for spinfection test and measure the number of cells/mL. Suspend the cells in small volume of median since spinfection requires concentrated seeding density.
-
c.Design dilution series of virus for MOI determination between 0–1 mL (e.g., 0.1 mL, 0.2 mL, 0.5 mL). For each dilution series, prepare two sets (one for testing MOI, one for positive control). Add 1 mL of cell suspension and 1 mL of media/polybrene/lentivirus solution to the corresponding labeled wells (total 2 mL each well). Prepare another one set of samples without lentivirus for negative control purpose (i.e., one set of 6 × 10ˆ6 SaOS2 in 8 mL media/polybrene solution ready for 4 wells).Plating conditions
Solution Amount per well in 6-well plate SaOS2-Cas9 cells 1.5 × 10ˆ6 in 1 mL media Polybrene 1.6 μL (final conc. 8 μg/mL) Pooled lentivirus Median To 2 mL -
d.Centrifuge the plate: spin the plates at 930 × g for 1.5 h at 30°C.
-
e.After 1.5 h spinning, add 2 mL of media without polybrene dropwise on top of each well.
-
f.Incubate cells for 24 h (37°C, 5% CO2).
-
a.
-
11.Day 2. Transfer the cells into 15 cm plate and start antibiotics selection:
-
a.Inspect cells visually, take note of the following things: cell conditions, the confluence of the wells, how adhered the cells are, approximate number of dead cells.
-
b.Collect the cells from the 6-well plate.
-
i.Remove the median, gently wash the cells with 1 mL pre-warmed PBS.
-
ii.Add 250 μL of trypsin each well and let the cells detach from the plate.
-
iii.Add 750 μL of media to each well and transfer the cell suspension to corresponding labeled tubes.
-
iv.Spin down and then re-suspend the cells.
-
i.
-
c.Reseed the cells to 15 cm plate (4 corresponding wells for one 15 cm plate for SaOS2) with either media containing puromycin at required concentration (2.5 μL/mL for SaOS2) or media without puromycin. Add media containing puromycin to the negative control (cells + polybrene, no virus) set.
-
a.
Note: If your incubator is not level, you can let plates sit level in hood for up to 1 h.
-
12.Day 4. MOI determination.
-
a.After 48 h, all uninfected cells should be dead. There should be no living cells in the negative control group.
-
b.Remove media, gently wash the cells with pre-warmed PBS to remove all the remaining dead cells. Trypsinize and collect cells from each plate into 15 mL tubes.
-
c.Count the cells from all plates and two settings of with or without puromycin selection.
-
d.Determine the lentivirus volume that yields to ∼30% survival with puromycin selection vs. without puromycin selection, which gives MOI ∼0.3 and ensure each cell is infected by no more than one virus.
-
a.
Part 4: Screening with customized library pool
Timing: 3–5 weeks
Part 4 performs pooled combinatorial CRISPR screen in human cells. The initial- and end- time point samples were harvested.
CRITICAL: Carefully consider the number of replicates and coverage you are going to use. Based on the previous MOI testing, cell coverage and replicate number, calculate the total number of cells, number of plates, and corresponding lentivirus volume (refer to step 13b).
CRITICAL: Biosafety precautions for virus waste (steps 12 and 13). In our institution, all virus waste requires incubation in 20% bleach for 30 min before disposal. No vacuum pumps can be used.
Note: Please refer to troubleshooting 3 to address the low infection efficiency issue.
Note: 1.25 × 10ˆ7 cells give ∼200-fold representation for each gRNA-gRNA construct in the 62,500 pool. If desired, you may use higher representation. In the previous combinatorial CRISPR screen, 80-fold is the minimum coverage ((Zhao et al., 2018)).
Note: Previous CRISPR/Cas9 screens have shown high genomic modification efficiencies after 6–12 days post-expression of Cas9 and gRNA. However, this genomic modification efficiency might be varied due to variations among gRNA performance and cell types. To confirm the ability of the CRISPR/Cas9 system to edit genes in specified cell lines, surveyor assay can be used (Wong et al., 2016). Nevertheless, the culture time for CRISPR/Cas9 screen post antibiotics selection typically ranges from 10 to 30 days (refer to Table1).
Note: A transient slowing of cell growth may be noticed. The cells should be able to return to grow normally after 5–15 days after infection, depending on the cell lines.
Note: Collect more cells if possible, considering the material loss during downstream processes such as DNA extraction.
-
13.Day 1. Spinfection. Troubleshooting 3.
-
a.The following extra plates with same cell number each plate are needed for negative and positive control purpose: one negative control plate: no virus + puromycin (0% survival control); one positive control plate: virus + no puromycin (100% survival control).
-
b.Calculate the plate number and total number of cells plated for infection with MOI ∼0.3. As discussed in Preparation4, for the SaOS2-Cas9 with triplicate at each time point, at least 2.5 × 10ˆ8 cells are needed at the time of infection, which requires ∼30 6-wheel plates for spinfection.
-
c.Thaw pooled lentivirus on ice.
-
d.Harvest cells for spinfection with corresponding virus volume to achieve ∼0.3 MOI. Follow the spinfection process as step 9.
-
a.
-
14.
Day 2. Transfer the cells into 15 cm plate and start antibiotics selection: same as step 10.
-
15.Day 4. T0 sample collection and MOI checking.
-
a.After 48 h, all uninfected cells should be dead. There should be no living cells in the negative control group.
-
b.Remove media, gently wash the cells with pre-warmed PBS to remove all the remaining dead cells. Trypsinize and collect cells from each plate into 15 mL tubes.
-
c.Count the cells from all plates and two settings of with or without puromycin selection. Ensure the MOI in the formal run is around ∼0.3.
-
d.Collect T0 sample: collect 3 replicates of 1.25 × 10ˆ7 cells by centrifuge (500 × g for 5 min). Wash the T0 samples with PBS and centrifuge again. Label the tube and freeze the cell pellets at −80°C.
-
e.Re-seed the rest of cells into three replicate groups. Ensure each replicate contains at least in total 1.25 × 10ˆ7 cells to maintain the 200 cell coverage. Do NOT use puromycin in this or subsequent plating steps.
-
a.
-
16.
Onward. (T1, T2, …, end point): The pool-infected cells should be cultured and passaged as the same way that you normally use. Each instance that you passage the cells out to the endpoint, ensure at least 1.5 × 10ˆ7 cells in total for each replicate group.
Note: Monitor the cell growth carefully, you may need to consider harvesting the Tend sample earlier if cells started dying fast.
-
17.
End time point. Collect Tend samples as step 14d.
Part 5: Genomic DNA extraction and PCR
Timing: 3–5 days
Part 5 performs two rounds of PCR after genomic DNA (gDNA) extraction. PCR1 enriches dual-gRNA cassette regions from the whole genome. PCR2 adds Illumina adapters and sample indices to the amplified dual-gRNA to facilitate Illumina sequencing in part 6.
Note: IDT UltramerTM DNA oligonucleotide is suggested for the long PCR2 primers (with adapter to flow cell, sample index, sequencing primer), which is more expensive than PCR1 primers. Thus two-step PCR is performed to reduce cost as well as PCR-induced error.
Note: Assuming a diploid genome is around 6.5 pg and one dual-gRNA construction per genome, 80 μg (62,500∗200∗6.5pg) of genomic DNA yields ∼200-fold coverage of a library with size of 62,500. Note that the average 6.5 pg genome weight might not fit every cell line.
Note: Please refer to troubleshooting 4 for PCR failure.
-
18.
gDNA Extraction: extract gDNA following the protocol of JetQuick Blood and Cell Culture DNA Maxiprep Kit.
-
19.
Precipitate and purify gDNA, and then quantify gDNA using Qubit dsDNA BR Assay.
-
20.
PCR1 (extract and enrich gRNA regions from the genome): add 2.5 μg of gDNA per 50 μL reaction. Set up identical 50 μL reactions to achieve the desired coverage.
Note: Use the lowest amount of the gDNA among all samples to calculation the number of PCR1 reaction for each sample. For example, if 80 μg is the lowest yield from all gDNA samples, then set up 32 PCR1 reactions for all samples.
PCR1 reaction master mix
| Reagent | Amount |
|---|---|
| gDNA | 2.5 μg |
| Primer PCR1_F (10 μM) | 2.5 μL |
| Primer PCR1_R (10 μM) | 2.5 μL |
| 2× NEBNext Μltra II Q5 Master Mix | 25 μL |
| ddH2O | To 50 μL |
PCR1 cycling conditions
| Steps | Temperature | Time | Cycles |
|---|---|---|---|
| Initial Denaturation | 98°C | 30 s | 1 |
| Denaturation | 98°C | 10 s | 25 cycles |
| Annealing | 66°C | 30 s | |
| Extension | 72°C | 15 s | |
| Final extension | 72°C | 2 min | 1 |
| Hold | 4°C | forever | |
Pool all individual 50 μL PRC1 reaction products, mix by vertex. Then directly proceed to PCR2.
Optional: Run 2 μL of PCR1 product on 1% gel. Visualize the PCR product on a gel imager. PCR1 yields a product of ∼1,300 bp.
-
21.
PCR2 (amplify gRNA with Illumina TruSeq adapters with i5/i7 indices and barcodes):
PCR2 reaction master mix
| Reagent | Amount |
|---|---|
| PCR1 product | 5 μL |
| Primer PCR2_F (10 μM) | 2.5 μL |
| Primer PCR2_R (10 μM) | 2.5 μL |
| 2× NEBNext Μltra II Q5 Master Mix | 25 μL |
| ddH2O | To 50 μL |
PCR2 cycling conditions
| Steps | Temperature | Time | Cycles |
|---|---|---|---|
| Initial Denaturation | 98°C | 30 s | 1 |
| Denaturation | 98°C | 10 s | 10 cycles |
| Annealing | 66°C | 30 s | |
| Extension | 72°C | 10 s | |
| Final extension | 72°C | 4 min | 1 |
| Hold | 4°C | forever | |
-
22.Gel extraction and QC:
-
a.After PCR amplification, run out the reaction on an EZ-VISION Blue-Light DNA Dye-stained 1% agarose gel. Visualize and cut the DNA fragment at ∼580 bp, then extract DNA using the QIAGEN Gel Extraction Kit according to the manufacturer’s instructions, elute in 20 μL ddH2O.
-
b.Quantify the fragment concentration using Qubit dsDNA BR Assay.
-
c.Run the sample in TapeStation DNA1000 or Bioanalyzer High Sensitivity DNA chip for QC check.
-
a.
Part 6: Sequencing
Timing: depending on the sequencing core
Part 6 provides a guideline for selecting the sequencing service.
Note: Please refer to troubleshooting 5 for addressing the low diversity issue.
-
23.Choose appropriate sequencing service.
-
a.Calculate the total reads: the total reads needed can be calculated as: coverages∗library size∗number of samples / mapping ratio. For example, to reach 200 coverage for a library with 62.5k different gRNA-gRNA constructions with triplicate in the screening (T0 and Tend in triplicates, 6 samples in total), ∼ 9.5 × 10ˆ7 reads (200∗62,500∗6/0.8) will be needed. Ideally the mapping ratio should be greater than 0.8, this number can be reduced in read estimation if you were not confident about the library quality.
-
b.Standard Pair-End (PE) sequencing with dual-index is required to sequence the two gRNAs on the both ends of the DNA fragment. At least 2 × 50 bp read cycle is suggested to cover the 20-bp gRNA sequence at both ends of the DNA fragment.
-
c.Given the total reads and PE sequencing, you may choose the sequencing service. The library samples can be sequenced on NovaSeq, NextSeq and MiSeq, which we have all used previously.
-
a.
Expected outcomes
During the library construction, ∼30,000 colonies per electroporation are expected. In our study, less than 0.5% construction is missed in the final construction using 10 coverages in step 3. For the references, 300 μL lentivirus (LentiGuide_DKO backbone, spin-infection) is needed to reach ∼0.3 MOI for 6 × 106 SaOS2 cells per 15 cm plate. In the gDNA extraction part, ∼50 μg gDNA is the average yield for 20–25 × 106 cells. After two rounds of PCR and gel extraction (step 22), ∼15 ng/μL DNA (for 20 μL elution, Qubit) is expected. It should be only one peak in the report of TapeStation DNA1000 Bioanalyzer. Note that the fragment size reported by TapeStation might be slightly larger than actual size.
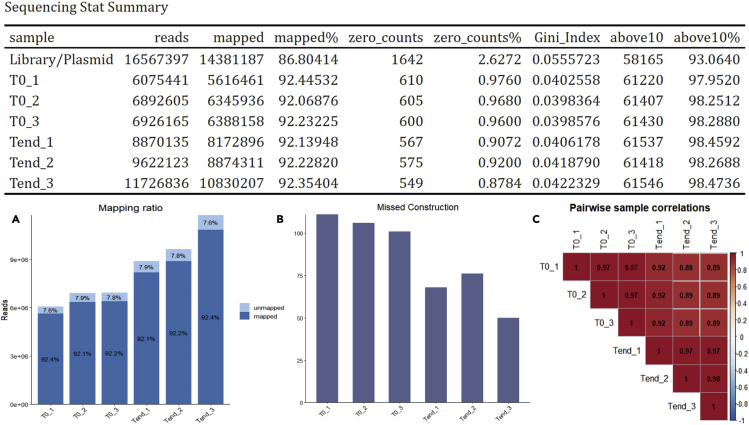
Following this protocol, pooled library of 62,500 dual-gRNA combinations for targeting more than 1,000 gene pairs was generated. Then combinatorial CRISPR screen on prostate cancer cell line 22Rv1 and osteosarcoma cell line SaOS2 (Cas9-expressing) was conducted to identify gene pairs that inhibited cancer cell growth when they were disrupted simultaneously. The dual-gRNA construction in initial- and end-time point samples were successfully sequenced and quantified. The diversity of the large dual-gRNA library was maintained throughout library generation, delivery, and screening, as more than 99% of the expected dual-gRNA constructions were detected in plasmid, T0 and Tend samples (Figure 4B). Additionally, the high mapping rate (above 90%, Figure 4A) and correlation between biological replicates (above 0.95, Figure 4C) further demonstrated the success of library construction and large-scale screening. Note that the difference in sequencing statistics between plasmid and screen samples is caused by the sequencing batch effect (MiSeq for single plasmid sample and NextSeq for multiple screen samples). Validation of growth-inhibiting effects of specific gene combinations that are identified by this study is in progress.
Figure 4.
Expected sequencing outcomes
Expected sequencing outcomes: sequencing statistic summary with (A) mapping ratio and mapped reads; (B) number of missed constructions; (C) pairwise sample correlation.
In general, a successful library construction should have a mapping rate greater than 80%, and the percentage of missed gRNA-gRNA construction should be less than 1%, or at most 5%. If any positive control is included, a substantial drop in the reads of the positive control constructs is expected.
Quantification and statistical analysis
Note: A small processed sampler dataset and all code for QC, hit identification etc. are available at GitHub: https://github.com/Shantang3/Combinatorial-CRISPR-Screen-Anaylysis-Pipeline.
-
1.
FastQC: FastQC provides an overview of basic quality control metrics for raw next generation sequencing data. A detailed demonstration of FastQC can be found at https://www.bioinformatics.babraham.ac.uk/projects/fastqc/Help/3%20Analysis%20Modμles/.
-
2.
Mapping and counting for gRNA-gRNA construction: Dual-crispr.
-
3.Quality control:
-
a.Mapping rate: should be above 80%.
-
b.Missed construct or zero count: should be less than 1%, or at most less than 5%.
-
c.Count distribution.
-
d.Sample correlation: should be greater than 0.8 between biological replicates.
-
e.gRNA distribution evenness: In MAGeCK (Li et al., 2014), Gini index, a coefficient measuring the degree of income distribution inequality in economy, is used to measure the evenness of the gRNA distribution in different samples. Smaller Gini index represents more evenly distributed sgRNAs and thus is expected in the library and baseline samples.
-
a.
-
4.
Log fold change (LFC) for the abundance for each gene pair using MAGeCK RRA (Li et al., 2014).
MAGeCK RRA uses a modified robust ranking aggregation (RRA) algorithm and prioritizes genes whose sgRNA rankings are consistently higher than expected. In this step, the gene pairs with construct of A-B and B-A are considered to be the same. All dual-gRNA constructions targeting the same gene pair were pooled together when calculating the gene-level LFC.
-
5.
Genetic Interaction (GI) calculation and hit identification on gene pairs:
The output file of “gene_summary.txt”, specifically the LFC values, from MAGeCK RRA is used for the following analysis:-
a.Observed LFC for gene-gene pairs: by subtracting the LFC with median LFC for pairs in which both sgRNAs were safe controls. This step set the median LFC of safe-safe constructions to zero.
-
b.Individual gene LFC: the median LFCs of all pairs compromising specified gene and other safe controls.
-
c.Expected LFC for gRNA-gRNA pairs: summing of LFCs of two individual genes from 5a.
-
d.GI calculation: the LFC deviation between observed and expected gene-gene pairs divided by the standard deviation of the individual genes’ LFCs from 5a. The more negative GI score is, the higher possibility of synergistic lethality exists between the two genes.
-
a.
Limitations
The capability of combinational CRISPR screen is mainly constrained by the library size and cell culture limit, especially for adherent cell lines. During the library construction, growing colony on plate from electroporation using HiFi assembly product has been optimized to reach the yield of 25,000–30,000 colonies per reaction in this protocol. This number yet needs to be improved for libraries with larger size or higher coverage. Instead of plating, some studies directly used LB median culture in the library construction process to reach higher library coverage. However, the problem of recombination using liquid median culture may compromise the library quality (Wang et al., 2016).
Moreover, the performance of CRISPR/Cas9 system relies on well-designed gRNAs. Despite that many models and computational tools have been developed to design gRNA, there is no golden-standard to quantify or compare the efficacy and specificity of gRNAs. Even though we selected multiple gRNAs targeting specified genes from validated merchandise or the commonly used CRISPR library, their performance might differ one from another.
Lastly, unlike single-gene CRISPR screen which has well-established analysis pipelines, limited analysis pipelines are available for the combinatorial CRISPR screen. The calculation of genetic interaction (GI) score and definition of SL gene pairs also vary study to study. With growing interest in combinatorial CRISPR screen, more bioinformatics tools are expected to facilitate the research community.
Nevertheless, this protocol, covering the whole combinatorial CRISPR screen process from gene selection to data analysis, provides the researchers with a framework for conducting experiment, analyzing data, and nominating potential combinational targets from scratch.
Troubleshooting
Problem 1
Plasmid library is recombined.
Potential solution
The gel electrophoresis can be used to assess the recombination issues. The use of Endura cells, growing bacteria on agar plate instead of in liquid culture, and incubation at 30°C are all intended to minimize recombination of the lentiviral plasmid library. Additionally, using the vector without Cas9 helps to reduce recombination during plasmid amplification.
Problem 2
Insufficient library coverage due to low transformation efficiency.
Potential solution
There are two common causes of this issue. Firstly, electrocompetent cells are no longer good to use. A test electroporation with control plasmid can be performed to check the condition of electrocompetent cells. Secondly, the salt in the HiFi product might be too much. To avoid introducing excess salt, we use ddH2O to elute DNA in step1, step2a and step3a. Diluting the HiFi product with ddH2O before transforming also helps.
Additionally, the transformation efficiency can be improved by adjusting the ratio of digested plasmid and insertion during HiFi assembly, the amount of HiFi product used, and electroporation setting. Notably, we noticed that using more HiFi product in electroporation does NOT guarantee higher efficiency. Furthermore, unlike directly using plasmid DNA, the number of the colony each electroporation using HiFi product is restrained by the input of HiFi Assembly. For example, less than 20 fmol of the 8k linearized vector is suggested. Finally, adding recovery media to the competent cells right after electroporation is critical. For more guidelines for using HiFi DNA assembly, please refer to: https://www.neb.com/tools-and-resources/usage-guidelines/guidelines-for-using-nebuilder-hifi-dna-assembly.
Problem 3
Low MOI, or low transduction efficiency.
Potential solution
One factor that influences viral titers is the size of the vector. Higher viral titers (typically 20 to 100-fold) during viral packaging can be achieved using the vector without Cas9. Wang et al. suggested only who plant to conduct screens across multiple cell lines to clone libraries into the Cas9-containing vector (Wang et al., 2016).
The spin-infection is designed to achieve higher efficient transduction already. Additionally, using products like ViralBoost Reagent during packaging can increase viral titers.
Problem 4
PCR failure.
Potential solution
The most likely reason is the low quality of the gDNA. Check the DNA quality by A260/A280, or Bioanalyzer. More details about the troubleshooting of gDNA preparation can be found at: https://www.sigmaaldrich.com/US/en/technical-documents/technical-article/genomics/dna-and-rna-purification/problems-during-genomic-dna-preparation.
Problem 5
Low diversity libraries.
Potential solution
Low diversity libraries can suffer from Illumina focusing issues, a problem not found with random fragment libraries. Traditionally this problem can be solved in two ways: 1) ‘Spiking in’ a higher diversity sample such as PhiX, or RNAseq samples; 2) Use amplicon primers with a series of random ‘N’ (25%A, 25%T, 25%G, 25%C) bases upstream of their gene target. Note that Illuminia sequencer prefers shorter reads; therefore, if you want to sequence the libraries with RNAseq samples, make sure the RNAseq samples’ size are ∼600 bp. Additionally, the mean insert size of the PhiX v3 library is approximately 400-bp, which is smaller than the 580-bp library. In one of our test runs using Illumina MiSeq with 40% PhiX spike-in, 75% reads went to PhiX in the end. Thus, consider lowering the PhiX spike-in ratio when loading together with the library samples.
Problem 6
False positive hits.
Potential solution
In our screen, the single genes that are already essential for cell growth tend to have a greater chance appearing in the SL gene pairs. Thus, excluding the single essential genes from the initial gene list (prep 4a) is critical for screen and hit identification. Alternatively, removing the gene combinations with single essential gene from the SL gene list is suggested after data analysis. In particular, following validation by means of transcriptional repression, protein inhibition or KO generation is needed to avoid false positive results.
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Lang Li (lang.li@osumc.edu).
Materials availability
Plasmids generated in this study have been deposited to Addgene: LentiGuide_DKO (Plasmid #183193), or LentiCRISPR_DKO (Plasmid #183192).
Cas9-expressing cell lines (22RV1-Cas9, SaOS2-Cas9) generated in relevant studies are available upon request.
Acknowledgments
The work related to cell culture is done by Dr. Liu’s lab and was supported by NIH R01 CA157429 (X. Liu), R01 CA196634 (X. Liu), R01 CA264652 (X. Liu), R01 CA256893 (X. Liu).
Author contributions
Conceptualization: S.T., X.W., and L.L.; Gene selection: B.G., K.F., F.L., and L.C.; Library constriction: S.T., X.W., and S.S.; Screening in vitro: J.L., Q.Z., and X.W.; Data analysis: S.T.; Writing – original draft: S.T.; Writing – review and editing: X.W. and L.L.; Supervision: L.L. and X.L.
Declaration of interests
The authors declare no competing interests.
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.xpro.2022.101556.
Contributor Information
Shan Tang, Email: shan.tang@osumc.edu.
Lang Li, Email: lang.li@osumc.edu.
Supplemental information
Data and code availability
The original sequencing files (fastq) of the dual-gRNA library generated by this protocol are stored at Sequence Read Archive (SRA) under PRJNA835479 (BioSample accession SAMN28103246).
A sampler processed dataset and the code generated during this study is available at GitHub: https://github.com/Shantang3/Combinatorial-CRISPR-Screen-Anaylysis-Pipeline, with Zenodo https://doi.org/10.5281/zenodo.6592122.
References
- Broad DepMap. 2021. https://depmap.org/portal/
- Cui Y., Xu J., Cheng M., Liao X., Peng S. Review of CRISPR/Cas9 sgRNA design tools. Interdiscip. Sci. 2018;10:455–465. doi: 10.1007/s12539-018-0298-z. [DOI] [PubMed] [Google Scholar]
- Diehl V., Wegner M., Grumati P., Husnjak K., Schaubeck S., Gubas A., Shah V.J., Polat I.H., Langschied F., Prieto-Garcia C. Minimized combinatorial CRISPR screens identify genetic interactions in autophagy. Nucleic Acids Res. 2021;49:5684–5704. doi: 10.1093/nar/gkab309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doench J.G., Fusi N., Sullender M., Hegde M., Vaimberg E.W., Donovan K.F., Smith I., Tothova Z., Wilen C., Orchard R., et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 2016;34:184–191. doi: 10.1038/nbt.3437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilbert L.A., Horlbeck M.A., Adamson B., Villalta J.E., Chen Y., Whitehead E.H., Guimaraes C., Panning B., Ploegh H.L., Bassik M.C., et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell. 2014;159:647–661. doi: 10.1016/j.cell.2014.09.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han K., Jeng E.E., Hess G.T., Morgens D.W., Li A., Bassik M.C. Synergistic drug combinations for cancer identified in a CRISPR screen for pairwise genetic interactions. Nature Biotechnology. 2017;35:463. doi: 10.1038/nbt.3834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hart T., Chandrashekhar M., Aregger M., Steinhart Z., Brown K.R., Macleod G., Mis M., Zimmermann M., Fradet-Turcotte A., Sun S., et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell. 2015;163:1515–1526. doi: 10.1016/j.cell.2015.11.015. [DOI] [PubMed] [Google Scholar]
- Hart T., Tong A.H.Y., Chan K., Van Leeuwen J., Seetharaman A., Aregger M., Chandrashekhar M., Hustedt N., Seth S., Noonan A., et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. G3. 2017;7:2719–2727. doi: 10.1534/g3.117.041277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horlbeck M.A., Xu A., Wang M., Bennett N.K., Park C.Y., Bogdanoff D., Adamson B., Chow E.D., Kampmann M., Peterson T.R. Mapping the genetic landscape of human cells. Cell. 2018;174:953–967. doi: 10.1016/j.cell.2018.06.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito T., Young M.J., Li R., Jain S., Wernitznig A., Krill-Burger J.M., Lemke C.T., Monducci D., Rodriguez D.J., Chang L., et al. Paralog knockout profiling identifies DUSP4 and DUSP6 as a digenic dependence in MAPK pathway-driven cancers. Nat Genet. 2021;53:1664–1672. doi: 10.1038/s41588-021-00967-z. [DOI] [PubMed] [Google Scholar]
- Li W., Xu H., Xiao T., Cong L., Love M.I., Zhang F., Irizarry R.A., Liu J.S., Brown M., Liu X.S. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 2014;15:554. doi: 10.1186/s13059-014-0554-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu G., Zhang Y., Zhang T. Computational approaches for effective CRISPR guide RNA design and evaluation. Comput. Struct. Biotechnol. J. 2020;18:35–44. doi: 10.1016/j.csbj.2019.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo B., Cheung H.W., Subramanian A., Sharifnia T., Okamoto M., Yang X., Hinkle G., Boehm J.S., Beroukhim R., Weir B.A., et al. Highly parallel identification of essential genes in cancer cells. Proc. Natl. Acad. Sci. USA. 2008;105:20380–20385. doi: 10.1073/pnas.0810485105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mair B., Tomic J., Masud S.N., Tonge P., Weiss A., Usaj M., Tong A.H.Y., Kwan J.J., Brown K.R., Titus E., et al. Essential gene profiles for human pluripotent stem cells identify uncharacterized genes and substrate dependencies. Cell Rep. 2019;27:599–615.e12. doi: 10.1016/j.celrep.2019.02.041. [DOI] [PubMed] [Google Scholar]
- Michlits G., Jude J., Hinterndorfer M., De Almeida M., Vainorius G., Hubmann M., Neumann T., Schleiffer A., Burkard T.R., Fellner M., et al. Multilayered VBC score predicts sgRNAs that efficiently generate loss-of-function alleles. Nat. Methods. 2020;17:708–716. doi: 10.1038/s41592-020-0850-8. [DOI] [PubMed] [Google Scholar]
- Parrish P.C.R., Thomas J.D., Gabel A.M., Kamlapurkar S., Bradley R.K., Berger A.H. Discovery of synthetic lethal and tumor suppressor paralog pairs in the human genome. Cell Rep. 2021;36 doi: 10.1016/j.celrep.2021.109597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanjana N.E., Shalem O., Zhang F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods. 2014;11:783–784. doi: 10.1038/nmeth.3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang D., Zhang C., Wang B., Li B., Wang Q., Liu D., Wang H., Zhou Y., Shi L., Lan F., Wang Y. Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning. Nat. Commun. 2019;10:4284. doi: 10.1038/s41467-019-12281-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen J.P., Zhao D., Sasik R., Luebeck J., Birmingham A., Bojorquez-Gomez A., Licon K., Klepper K., Pekin D., Beckett A.N. Combinatorial CRISPR–Cas9 screens for de novo mapping of genetic interactions. Nat. Methods. 2017;14:573–576. doi: 10.1038/nmeth.4225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson N.A., Ranzani M., Van Der Weyden L., Iyer V., Offord V., Droop A., Behan F., Gonçalves E., Speak A., Iorio F. Combinatorial CRISPR screen identifies fitness effects of gene paralogues. Nat. Commun. 2021;12:1–11. doi: 10.1038/s41467-021-21478-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L., Huang X., You X., Yi T., Lu B., Liu J., Lu G., Ma M., Zou C., Wu J., Zhao W. Nanoparticle enhanced combination therapy for stem-like progenitors defined by single-cell transcriptomics in chemotherapy-resistant osteosarcoma. Signal Transduct. Target Ther. 2020;5:196. doi: 10.1038/s41392-020-00248-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang T., Lander E.S., Sabatini D.M. Single guide RNA library design and construction. Cold Spring Harbor Protoc. 2016;2016 doi: 10.1101/pdb.prot090803. pdb.prot090803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wishart D.S., Feunang Y.D., Guo A.C., Lo E.J., Marcu A., Grant J.R., Sajed T., Johnson D., Li C., Sayeeda Z., et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 2018;46:D1074–D1082. doi: 10.1093/nar/gkx1037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong A.S.L., Choi G.C.G., Cui C.H., Pregernig G., Milani P., Adam M., Perli S.D., Kazer S.W., Gaillard A., Hermann M., et al. Multiplexed barcoded CRISPR-Cas9 screening enabled by CombiGEM. Proc. Natl. Acad. Sci. USA. 2016;113:2544–2549. doi: 10.1073/pnas.1517883113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu D., Kahen E., Cubitt C.L., Mcguire J., Kreahling J., Lee J., Altiok S., Lynch C.C., Sullivan D.M., Reed D.R. Identification of synergistic, clinically achievable, combination therapies for osteosarcoma. Sci. Rep. 2015;5:16991. doi: 10.1038/srep16991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao D., Badur M.G., Luebeck J., Magaña J.H., Birmingham A., Sasik R., Ahn C.S., Ideker T., Metallo C.M., Mali P. Combinatorial CRISPR-Cas9 metabolic screens reveal critical redox control points dependent on the KEAP1-NRF2 regulatory axis. Mo. Cell. 2018;69:699–708. doi: 10.1016/j.molcel.2018.01.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The original sequencing files (fastq) of the dual-gRNA library generated by this protocol are stored at Sequence Read Archive (SRA) under PRJNA835479 (BioSample accession SAMN28103246).
A sampler processed dataset and the code generated during this study is available at GitHub: https://github.com/Shantang3/Combinatorial-CRISPR-Screen-Anaylysis-Pipeline, with Zenodo https://doi.org/10.5281/zenodo.6592122.