Abstract
The Bacillus subtilis 168 derivative JH642 produces a bacteriocin, subtilosin, which possesses activity against Listeria monocytogenes. Inspection of the amino acid sequence of the presubtilosin polypeptide encoded by the gene sboA and sequence data from analysis of mature subtilosin indicate that the precursor subtilosin peptide undergoes several unique and unusual chemical modifications during its maturation process. The genes of the sbo-alb operon are believed to function in the synthesis and maturation of subtilosin. Nonpolar mutations introduced into each of the alb genes resulted in loss or reduction of subtilosin production. sboA, albA, and albF mutants showed no antilisterial activity, indicating that the products of these genes are critical for the production of active subtilosin. Mutations in albB, -C, and -D resulted in reduction of antilisterial activity and decreased immunity to subtilosin, particularly under anaerobic conditions. A new gene, sboX, encoding another bacteriocin-like product was discovered residing in a sequence overlapping the coding region of sboA. Construction of an sboX-lacZ translational fusion and analysis of its expression indicate that sboX is induced in stationary phase of anaerobic cultures of JH642. An in-frame deletion of the sboX coding sequence did not affect the antilisterial activity or production of or immunity to subtilosin. The results of this investigation show that the sbo-alb genes are required for the mechanisms of subtilosin synthesis and immunity.
The bacteriocins are a group of antimicrobial peptides that are produced by microorganisms inhabiting diverse environments (10, 12, 14). Typically, these small, gene-encoded polypeptides are first made as unprocessed, unmodified precursors that often undergo interesting and unique chemical modifications during their maturation. They possess hydrophobic N termini that mediate secretion and are subsequently removed proteolytically, yielding the active bacteriocins. The lantibiotic class of bacteriocins, the so-called group I bacteriocins (7, 10, 12, 44), include modified polypeptides that contain dehydrated threonine (T) or serine (S) residues, which condense with cysteine (C) to yield methyllanthionines and lanthionines. The lanthionines form intrachain thioether bridges that impart a cyclic character to the group I bacteriocins. Lantibiotics are produced by gram-positive bacteria such as the Bacillus species, the lactic acid bacteria (Lactococcus, Lactobacillus, and Carnobacterium spp.), and other gram-positive cocci (7, 10, 12, 34, 51). Since many of these bacteria are common components of fermented food, they have found use in food preservation and the inhibition of food-borne pathogens (18, 19). The lantibiotics, such as nisin, and the unmodified, or group II, bacteriocin pediocin PA-1 are used to reduce bacterial contamination in dairy products and meats (19).
The spore-forming, nonpathogenic soil bacterium Bacillus subtilis is capable of growth under aerobic and anaerobic conditions (24, 39) on a variety of substrates and is a common component of traditional fermented foods (2), particularly in Asian and African cultures. It also is known to produce an abundance of antimicrobial compounds, including the bacteriocin subtilosin (1, 49). While many lantibiotics and group II bacteriocins have been discovered and characterized (19), subtilosin is unique in that it appears to undergo unusual chemical modifications during its maturation that are unlike those of lantibiotics. Although codons for Phe are present in the nucleotide sequence of the gene (sboA) encoding subtilosin, no Phe residues are detected by amino acid analysis or by sequencing of peptides released from purified subtilosin through partial hydrolysis or proteolysis (1, 49). Other unusual properties can be discerned by inspection of the amino acid sequence, particularly the very short leader peptide of the subtilosin precursor and the covalent linkage of the N-terminal Asn and the C-terminal Gly.
We had reported that the operon was composed of eight genes (sboA and albABCDEFG). It is likely that some of the products of the alb genes function in carrying out the unusual modifications and processing of the subtilosin prepeptide. A search of proteins showing homology with the amino acid sequence of the alb products reveals similarities to proteins that are known to be involved with peptide processing, some of which are unlike those that carry out the processing of lantibiotics. Knowledge of how modifications are carried out during subtilosin maturation may provide new strategies for chemically modifying peptides.
The transcriptional start site of sbo-alb was localized to a sequence at position −45 with respect to the ATG translational start codon of sboA. It is preceded by a −35 and −10 region resembling a ςA-utilized promoter. A stem-loop structure located at the end of the sboA gene and upstream of the albA coding sequence may reduce transcriptional readthrough into the alb genes, thereby ensuring that the alb products are produced in smaller amounts with respect to the SboA substrate peptide. The sbo-alb genes are transcriptionally regulated by factors, such as Spo0A (8) and AbrB (41), which control the nutritional stress response and processes of cellular differentiation in B. subtilis. In a previous report (T. Stein, S. Düsterhus, A. Stroh, and K.-D. Entian, Abstr. 10th Int. Conf. Bacilli, abstr. P103, p. 65, 1999) and in the accompanying paper (23), we show that sbo-alb is induced under anaerobic conditions and controlled by the ResDE signal transduction system that regulates gene expression in response to limiting oxygen (24, 26, 42).
In this report, the phenotypes caused by nonpolar insertion mutations in each of the alb genes are described along with the identification of a new gene, sboX, the coding sequence of which overlaps the sboA gene. Genes required for immunity to subtilosin are also identified.
MATERIALS AND METHODS
Bacterial strains, plasmids, and oligonucleotide primers.
The bacterial strains used in this study are described in Table 1. All are derivatives of B. subtilis strain JH642. The indicator bacterium Listeria monocytogenes F4244, which was used to detect production of subtilosin, was obtained from M. Slavik (48). Strains ORB3148 and ORB3149 bear neo gene cassette insertions in the sboA gene (49). In ORB3148, the neo gene is oriented in the direction of sbo-alb transcription, and in ORB3149, the neo gene is oriented in the opposite direction.
TABLE 1.
Strain used in this study
| Strain | Genotype | Reference or source |
|---|---|---|
| JH642 | trpC2 pheA1 | J. A. Hoch |
| ORB3148 | trpC2 pheA1 sbo::neo-1 | 49 |
| ORB3149 | trpC2 pheA1 sbo::neo-2 | 49 |
| ORB3351 | trpC2 pheA1 albA::pAG-albA (Cmr Phler) | This study |
| ORB3400 | trpC2 pheA1 albB::cat (Cmr) | This study |
| ORB3353 | trpC2 pheA1 albC::pAG-albC (Cmr Phler) | This study |
| ORB3354 | trpC2 pheA1 albD::pAG-albD (Cmr Phler) | This study |
| ORB3355 | trpC2 pheA1 albE::pAG-albE (Cmr Phler) | This study |
| ORB3356 | trpC2 pheA1 albF::pAG-albF (Cmr Phler) | This study |
| ORB3354 | trpC2 pheA1 albG::pAG-albG (Cmr Phler) | This study |
| ORB3412 | trpC2 pheA1 amyE::pTRP-sboX (Cmr) | This study |
| ORB3413 | trpC2 pheA1 amyE::pTRP-sboX sboA::neo (Cmr Neor) | This study |
| ORB3441 | trpC2 pheA1 amyE::pTRP-sboX abrB::neo (Cmr Neor) | This study |
| ORB3442 | trpC2 pheA1 sboX::neo-1 (Neor) | This study |
| ORB3445 | trpC2 pheA1 sboX::neo-2 (Neor) | This study |
| ORB3470 | trpC2 pheA1 amyE::pDR-sboAXalbABC′ (Cmr) | This study |
| ORB3471 | trpC2 pheA1 amyE::pDR-sboAXalbABC′::spc (Specr) | This study |
| ORB3472 | trpC2 pheA1 amyE::pDR-sboAXalbABC′::spc albB::cat (Specr Cmr) | This study |
| ORB2552 | pheA1 sboX::neo-1 | This study |
| ORB3568 | pheA1 sboXΔ89-139 | This study |
The oligonucleotides used in this study are described in Table 2. Plasmids pAG-albA, pAG-albC, pAG-albD, pAG-albE, pAG-albF, and pAG-albG are derivatives of pAG58-ble-1 (47) and were used to create nonpolar disruptions of the alb genes. To construct these plasmids, a HindIII-XbaI fragment containing an internal segment of an alb gene (except albB) was generated by PCR using a pair of oligonucleotides as detailed below. Each individual fragment was then inserted into HindIII-XbaI-cleaved pAG58-ble-1. The resulting plasmids, pAG-albA, pAG-albC, pAG-albD, pAG-albE, pAG-albF, and pAG-albG, were used to transform competent cells of JH642 with selection for chloramphenicol resistance (Cmr) to create alb gene disruption mutants. To construct pAG-albA, oligonucleotides oywiA-U and oywiA-L were used in a PCR to obtain a fragment extending from 9 to 681 bp downstream of the TTG translation start site of albA. For pAG-albC, oligonucleotides oywiQ-U and oywiQ-L were used to generate a PCR fragment extending from 43 to 604 bp downstream of the ATG start site of albC. In the construction of pAG-albD, oligonucleotides oywhP-U and oywhP-L were used to generate a PCR fragment extending from 155 to 786 bp downstream of the ATG start site of albD. To construct pAG-albE, oligonucleotides oywhO-U and oywhO-L were used to generate a PCR product containing sequences from 49 to 1,035 bp downstream of the TTG start site of albE. In the construction of pAG-albF, oligonucleotides oywhN-U and oywhN-L were used to generate a PCR fragment extending from 219 to 725 bp downstream of the ATG start site of albF. To create pAG-albG, oligonucleotides oywhM-U and oywhM-L2 were used to generate a PCR fragment extending from 10 to 546 bp downstream of the ATG start site of albG.
TABLE 2.
Oligonucleotides used in this study
| Oligonucleotide | Sequence |
|---|---|
| oGZ1 | CAGCTCAAGCTTACCAAATGACATTTT |
| sboP1 | CCTCATGACCAGGACTTCGCCTTCGCTTACTTT |
| sboP2 | CGGTGCCGAGCGCTTCAGGTAAGCTTTCCAAA |
| sboP3 | TGCTGGATCCGAGCCGCTTGTCTAGTGGACGGTCCTAT |
| sboP4 | CTCGGATCCAGCATGTTGCACAACCTTTGTTTTCTA |
| oywiA-U | GAGCAGAAGCTTCCATTTATTAATGAAA |
| oywiA-L | CACTATCTAGAGAAATGCCGACGACGATGATTT |
| oywiQ-U | GGGACAACGTAAGCTTAGAACAAGTGGATT |
| oywiQ-L | GCACGACATCTAGAAGATGAGAGAAGAAAA |
| oywhP-U | GCGCAAAGCTTATATCATACTCGCGCTTCTT |
| oywhP-L | GGTGTTCATCATTCTAGAACGAAAGAA |
| oywhO-U | GCCGCAAGCTTTCTGAAACCGATCGAAA |
| oywhO-L | CCCCATCTAGACCAATAGACCTTCGGGAGAAT |
| oywhN-U | GGCCGCAAGCTTTACAGTGATTTTTTT |
| oywhN-L | GAGAGGTCTAGATTTCATGCCGGGATGATT |
| oywhM-U | GCACTGTCTTTAAGCTTTTATTGCTGCTTA |
| oywhM-L2 | CACTTTTCTAGACAGGCTGCCGTGCCGGAAAA |
| oywhR-U1 | GTGGGAATTCCACGATATGGCCCAAAA |
| oywhR-L1 | CCAAGGAAAGCTTCGAAATTCTCCTATT |
| oorfX-U | GCGGAATTCGATGACTTCTTGGTTT |
| oorfX-L | GGGAGAAGCTTACCCCCATAGACCGAATA |
| oorfX-U1 | CGCGGATCCCCCAAAAGGGCATAGTCATT |
| oorfX-L1 | CGCGGATCCTTACCCCCATAGACCGAATA |
| osboXd-3 | TTAAGGATCCCGTTTTTCATTTGAATCATAT |
| osboXd-5 | AACGGGATCCTTAAAAAGGGCATAGTA |
To construct pUC-albB::CAT, an EcoRI-HindIII fragment containing the albB gene along with the sequence 547 bp upstream and 577 bp downstream of albB was generated by PCR using the oligonucleotides oywihR-U1 and oywihR-L1. The EcoRI-HindIII fragment was inserted into EcoRI-HindIII-cleaved pUC18 to create pUC-albB. Plasmid pUC-albB was cut with BssHII at the two BssHII sites within the albB gene and treated with T4 DNA polymerase (28) to create blunt ends. A SmaI fragment bearing a cat (Cmr) cassette from pMMN7 (25) was inserted into the blunt ends of pUC-albB to create pUC-albB::CAT. The orientation of the cat gene in the plasmid was determined by restriction analysis. The resulting pUC-albB::CAT plasmid was used to transform competent cells of JH642 to obtain the albB::CAT insertion mutant, the structure of which was confirmed by PCR using oligonucleotides oywhR-U1 and oywhR-L1.
The sboX translational lacZ fusion was constructed by using plasmid ptrpBGI (37), which bears a promoterless lacZ gene. An EcoRI-HindIII fragment extending from 477 bp upstream to the 14th codon downstream of the putative TTG start site of sboX was generated by PCR using primers oorfX-U and oorfX-L. The PCR product was inserted into EcoRI-HindIII-cleaved ptrpBGI, which placed the sboX coding sequence in frame with the lacZ coding sequence. The resulting plasmid, pTRP-sboX, was used to transform competent cells of JH642 and ORB3148 to generate ORB3412 and ORB3413, respectively. The amylase-defective phenotype of ORB3412 and ORB3413 was confirmed by the iodine staining method (28). The chromosomal DNA of LAB2332 was used to transform competent cells of ORB3412 with selection for Cmr and Neor to obtain ORB3441.
Plasmid pUC-sboXBH is a derivative of pUC18 (47). A BamHI-HindIII fragment extending from 45 to 490 bp downstream of the TTG start codon of sboX was generated by PCR using oligonucleotides oorfX-U1 and osboP2 and inserted into BamHI-HindIII-cleaved pUC18. An EcoRI-BamHI fragment extending from 477 bp upstream to 45 bp downstream of the TTG start site of sboX was generated by PCR using primers oorfX-U and oorfX-L1 and cloned into the EcoRI-BamHI sites of pUC-sboXBH. The resulted plasmid was pUC-sboXEBH. To construct an sboX::neo strain, a BglII-BamHI fragment bearing the neo cassette from pDG782 (6) was inserted into the BamHI site of pUC-sboXEBH and the orientation of the neo cassette was determined by restriction analysis. The pUC-sboXEBH derivative containing the neo cassette was used to transform competent cells of JH642 to obtain strain ORB3442. The sboX::neo insertion mutation was confirmed by PCR using osboP2 and osboP3.
To make pDR-sboAXalbABC′, a HindIII-ClaI fragment extending from 490 bp upstream of the start codon of sboA to 510 bp downstream of the start codon of albC was made by PCR using the primers oGZ1 and oywiQ-L. The fragment was then inserted into HindIII-ClaI-cleaved pDR67, thus creating plasmid pDR-sboAxalbABC′. pDR-sboAxalbABC′ was used to transform cells of JH642 with selection for Cmr and screening for amyE (amylase negative) to obtain ORB3470. The Cmr marker of ORB3470 was replaced with Specr by using plasmid pCm::Sp (40) to obtain strain ORB3471. The chromosomal DNA of ORB3471 was then used to transform cells of albB mutant strain ORB3400 to obtain ORB3472.
Plasmid pUC-sboXEBH-D was constructed to create an in-frame deletion within the sboX coding sequence. A BamHI-HindIII fragment containing 138 to 490 bp downstream of the TTG start site of sboX was generated by PCR using oligonucleotides osboXd-3 and osboP-2 and inserted into BamHI-HindIII-cleaved pUC18 to generate pUC-sboXBH-D. EcoRI-BamHI-cleaved pUC-sboXBH-D was ligated to an EcoRI-BamHI fragment extending from 477 bp upstream to 89 bp downstream of the TTG start site of sboX which had previously been obtained by PCR using primers osboXd-5 and osboP-1. The resulting plasmid, pUC-sboXEBH-D, and chromosomal DNA of ORB3552 were then used to transform cells of strain ORB3442 with selection for Trp+ and screening for neomycin sensitivity. The sboXΔ89-138 in-frame deletion in ORB3568 thus created was confirmed by PCR and DNA sequence analysis.
Culture media.
2×YT broth (22) was used for the routine growth of Escherichia coli and B. subtilis. A yeast extract-glucose (YG) agar previously described (49) was used to grow JH642 or ORB3148 for analysis of the sensitivity of alb mutants toward subtilosin A. YG broth with addition of 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) (or chloramphenicol at 5 μg/ml for the albB mutant) was used for examination of the subtilosin A production of alb mutants. 2×YT broth supplemented with 1% glucose and 0.2% KNO3 was used for the growth of B. subtilis in the β-galactosidase activity assays and the immunity analysis using partially purified subtilosin A. Brain heart infusion medium was used for growth and maintenance of L. monocytogenes F4244.
Transformation and transduction.
The preparation of competent E. coli and B. subtilis cells and genetic transformation were carried out as previously described (4, 22).
β-Galactosidase assay.
B. subtilis cells were cultured in 2×YT broth supplemented with 1% glucose and 0.2% KNO3 under aerobic or anaerobic conditions as previously described (20). Samples were withdrawn at 1-h intervals for measurement of β-galactosidase activity (50).
Bioautography of subtilosin by SDS-PAGE.
The purity and activity of subtilosin samples from the wild type and alb mutants were analyzed by sodium dodecyl sulfate (SDS)–16.5% polyacrylamide gel electrophoresis (PAGE) with Tricine running buffer (36). Each sample (2 × 10 μl) was applied to duplicate gels. After electrophoresis at 100 V for 2 h, each lane was cut vertically from the gels. For each sample, one gel slice was stained by Coomassie blue staining solution and the other was assayed for inhibitory activity by bioautography using the indicator strain L. monocytogenes F4244 as described by Zheng and Slavik (48).
Partial purification of subtilosin A.
Subtilosin A purification was carried out as previously described (49), with the following modifications. Subtilosin A was precipitated from culture fluid by ammonium sulfate to 80% saturation, extracted for 1 h by 1/20 of the culture volume of methanol, concentrated by evaporation at 55°C, and then resolved by LH-20 chromatography. Subtilosin A fractions were dried by evaporation and dissolved in 20 mM Tris-HCl buffer (pH 7.0). This subtilosin A preparation was electrophoretically pure as demonstrated by Tricine SDS-PAGE. The subtilosin A concentration was determined spectrophotometrically by using the Protein Assay (Bio-Rad, Hercules, Calif.) with bovine serum albumin as the standard.
Immunity assay.
Immunity of alb mutants was examined by two methods. Cells of JH642 and the alb mutants were grown in 2 ml of 2×YT broth (for alb mutants chloramphenicol at 5 μg/ml was added). Cells were collected at exponential phase (optical density at 600 nm [OD600], 0.6 to 0.9), and the OD600 was adjusted to 0.5 by adding 2×YT broth. In the first immunity assay, 50-μl volumes of these suspensions were mixed with 5 ml of soft 2×YT (0.8% agar) and then poured onto YG plates onto which JH642 and ORB3148 had previously been stabbed and incubated at 37°C for 24 h. The overlaid plates were incubated for another 18 to 24 h at 37°C under aerobic and anaerobic conditions to observe the appearance of inhibition zones around JH642 or ORB3148 colonies. In the second immunity analysis, 50-μl volumes of the 2×YT suspensions were blended with 5-ml volumes of soft 2×YT (0.8% agar supplemented with 1% glucose and 0.2% KNO3) and then poured onto 2×YT agar (supplemented with 1% glucose and 0.2% KNO3). For the albB mutant, the soft agar contained chloramphenicol at 5 μg/ml. For all other alb mutants, the soft agar included 1 mM IPTG. Duplicate plates were made for each mutant and dried for 30 min in a 37°C incubator. Ten-microliter volumes of serial twofold dilutions of subtilosin (>90% pure) were spotted onto the surfaces of the plates, which were then incubated aerobically or anaerobically at 37°C overnight to observe the appearance of inhibition zones. The minimum concentration of subtilosin A that resulted in a clear inhibition zone on each plate was determined and reported as the MIC (see Table 3).
TABLE 3.
The sbo and alb genes and the phenotypes conferred by their mutant alleles
| Gene | Product name or length (molecular mass)a | Homolog(s) (function) | Mutant phenotype
|
|
|---|---|---|---|---|
| Antilisterial activity (%)b | Immunity (MIC [μg/ml])c | |||
| sboA | Subtilosin | Bacteriocin | − (<0.5) | + (280)d |
| albA | 449 (51,493) | NirJ, PqqE, MoeA (cofactor synthesis) | − (<0.5) | + (280) |
| albB | 53 (6,116) | ? (immunity) | ± (6.5) | − (63) |
| albC | 240 (27,232) | ABC transporter, ATP-binding protein | ± (12.5) | − (96) |
| albD | 436 (49,516) | ? | ± (6.5) | − (150) |
| albE | 399 (44,611) | Processing protease | ± (12.5) | + (280) |
| albF | 427 (48,979) | Ubiquinol-cytochrome c reductase complex, Zn endoproteinase (N terminal only) | − (<0.5) | + (280) |
| albG | 234 (26,333) | ? (membrane protein) | ± (25) | + (280) |
Length is in amino acids, and molecular mass is in daltons.
Percentages of wild-type activity, based on arbitrary units per microgram of cellular protein, are shown in parentheses.
Defect in immunity observed in anaerobic incubation. MICs were measured under anaerobic conditions.
Value is for the sboA::neo-2 mutant.
RESULTS
The albA and albF products are critical for the production of subtilosin.
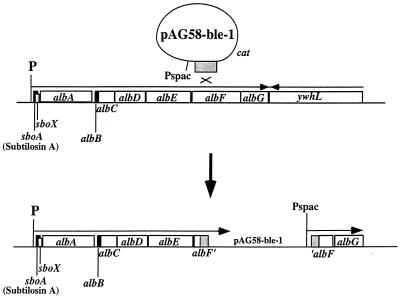
The sbo-alb genes are organized in an operon of 7 kb and are transcribed from a promoter residing upstream of the subtilosin structural gene sboA (Fig. 1). Nonpolar insertion mutations were created in each of the seven alb genes to determine if each functions in the production of and immunity to subtilosin. For albA, -C, -D, -E, -F, and -G, an internal fragment of the N-terminal coding half of each gene was amplified by PCR. Each fragment was inserted into the plasmid pAG58-bleo-1. Integration of the resulting recombinant plasmids into the alb locus by homologous recombination yielded insertion mutations that disrupt the alb open reading frames (Fig. 1). The Pspac promoter of pAG58-bleo-1 is directed downstream of the plasmid insertion so as to eliminate potential polarity effects exerted by the integrated plasmid. The albB gene is too small to create a gene disruption using an internal fragment of the albB coding sequence. In this case, a cat gene cassette was inserted into the BssHII sites residing within the albB gene and the resulting construct was introduced by double recombination into the sbo-alb operon. We had observed previously that the cat gene, if oriented in the same direction as the transcription of the operon into which it is inserted, directs transcription through the genes downstream of the insertion (21).
FIG. 1.
Organization of the sbo-alb operon and the recombination between the integrative plasmid and alb DNA that gives rise to the nonpolar insertion mutation. The example shown is albF. An internal region of the albF gene is amplified by PCR and inserted into the integration vector pAG58-bleo-1. After recombination, a copy of the plasmid is integrated into the alb locus. albF is disrupted, and transcription from the sbo promoter is blocked by the plasmid DNA. Expression of the downstream genes is driven by the IPTG-inducible Pspac promoter.
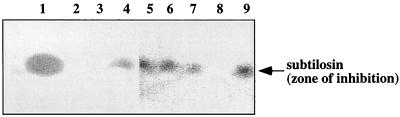
Each of the mutations thus generated was examined for effects on subtilosin production and immunity (Table 3). Subtilosin production was tested using the critical-dilution assay and bioautography (Fig. 2) of supernatants of YG cultures incubated for 36 h. The wild-type parent JH642 showed antilisterial activity and produced subtilosin, but each of the alb insertion mutants produced either none or reduced amounts of the bacteriocin. Only the sboA::neo-1, albA, and albF mutants failed to produce any detectable subtilosin. The albB, -C, -D, -E, and -G mutants exhibited antilisterial activity and produced small amounts of subtilosin, as judged by SDS-PAGE and bioautography (Fig. 2).
FIG. 2.
Bioautography of extracts from culture fluid of B. subtilis strain JH642 and the sboA and alb mutants. Supernatant fluid from YG liquid batch cultures collected at 36 h was subjected to (NH4)SO4 precipitation, followed by methanol extraction. Methanol extracts were evaporated, and the residue was dissolved in 20 mM Tris-HCl buffer, pH 7.0 (see Materials and Methods). Samples from extracts of wild-type and mutant culture supernatants were applied to Tricine SDS-polyacrylamide gels, and bioautography was performed. A zone of inhibition is observed in the area of the L. monocytogenes overlay corresponding to the position of bacteriocin in the gel (indicated by arrow). Lanes: 1, JH642; 2, sboA::neo; 3, albA; 4, albB; 5, albC; 6, albD; 7, albE; 8, albF; 9, albG.
The albB, albC, and albD products function in subtilosin immunity.
A bacteriocin-producing bacterium is immune to the specific bacteriocin that it produces (13, 35). The genes encoding immunity proteins are usually situated within the operon containing the genes required for bacteriocin biosynthesis. Products that function in immunity have been identified as small membrane-associated peptides, some having a lipid modification that is thought to anchor the peptide to the cytoplasmic membrane (33, 35). The sbo-alb operon contained one gene, albB, that encodes a small hydrophobic protein that we felt might function in subtilosin immunity. We tested all of the alb insertion mutants for defects in immunity by overlaying a lawn of alb mutant cells on colonies of JH642 or the sboA::neo-1 mutant and observing the zone of inhibition created (data not shown) or by determining the MIC of subtilosin required to create an inhibition zone on lawns of alb mutant cells embedded in a soft agar overlay (Table 3). Only the albB, -C, and -D mutants showed a reduction in subtilosin immunity, with the latter exhibiting the mildest defect.
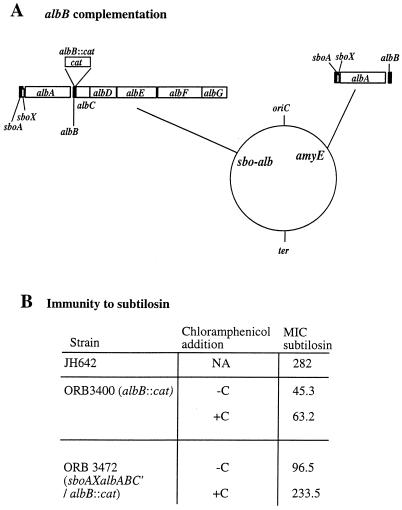
Although the cat gene used to create the albB insertion mutation had been shown to exert little polarity, a complementation experiment was designed to further show that the product of albB is necessary for optimal immunity to subtilosin. A DNA fragment containing the sbo-alb promoter, sboA, sboX, albA, and albB was inserted into the amyE integration plasmid pDR67. A partial diploid strain was constructed that contained the albB::cat insertion and the integrated sboAXalbABC′ construct (Fig. 3A). In the absence of added chloramphenicol, the ectopic albB allele only partially complemented the albB mutation (Fig. 3B). Addition of inducing levels of chloramphenicol, which activates expression of the cat gene insertion that drives expression of downstream alb genes, results in nearly complete complementation of the albB mutation. This confirms that the albB gene product is required for immunity and also indicates that the expression of albC and -D is needed for complete immunity.
FIG. 3.
Complementation of the albB mutation by the amyE::sboAXalbABC′ construct. (A) Genomic organization of albB/amyE::sboAXalbABC′. The circle indicates the B. subtilis genomic map, and the locations of the replication origin (oriC) and termination site (ter) are shown. The amyE and sbo-alb loci are labeled on the genomic map, and the organization of the sbo-alb operon bearing the albB::cat mutation and the sbo-alb DNA of the amyE::sboAXalbABC′ locus is shown. (B) MICs of subtilosin for the wild-type parent, JH642, the albB::cat mutant ORB3400, and the albB::cat/albB diploid ORB3472. An immunity assay was performed on anaerobically grown lawns of cells on 2×YT medium supplemented with glucose and KNO3 as described in Materials and Methods. The assay was performed in the presence (+C) or absence (−C) of chloramphenicol. NA, not applicable.
The sboA::neo-1 insertion mutant contains a neo gene cassette within the sboA coding sequence. The neo gene is transcriptionally oriented in the same direction as the sbo-alb operon (49) and has been shown to direct high-level expression of the downstream alb genes. Accordingly, the sboA::neo-1 mutant shows levels of subtilosin immunity higher than that exhibited by wild-type cells (MIC, >560 μg/ml). This higher level of self-protection can be attributed to the enhanced expression of albBCD in the sboA::neo-1 mutant. (Note that the value in Table 3 is that of the sboA::neo-2 mutant, in which the neo gene is oriented oppositely to the sbo-alb direction of transcription.)
Identification of the sboX gene and detection of its expression in anaerobic cultures.
Inspection of the sboA and sboA-albA intergenic region revealed an open reading frame that could potentially encode a bacteriocin-like product (Fig. 4). The coding sequence of this putative gene, sboX, begins at the 30th codon of sboA and extends to 28 nucleotides upstream from the TTG start of albA. A putative ribosome-binding site lies 16 to 20 bp from the TTG start codon. The product of sboX resembles a precursor of a type II bacteriocin in that it possesses a putative GG cleavage processing site, is cationic, and bears a hydropathy profile similar to that of some carnobacteriocins (32). To determine if the putative product of sboX functions in the production of antilisterial activity, an insertion mutation was introduced into the sboX open reading frame. Two restriction sites, HindIII and BamHI, were constructed in the sequence corresponding to the 16th and 17th codons, respectively, of the sboX coding sequence. The HindIII site was used to create a translation lacZ fusion as described below. Into the BamHI site was inserted a neomycin resistance cassette that would result in the insertional inactivation of the sboX gene. Introduction of the insertion into the sboX gene of the JH642 chromosome resulted in reduction of subtilosin activity, as shown by incubation of sboX::neo mutant colonies in a lawn of L. monocytogenes cells (data not shown). The neo cassette used was the same as that used to create sboA::neo-1, which was shown to direct the transcription of the downstream alb genes (49). It is unlikely that the insertion exerts a negative polar effect on alb gene expression.
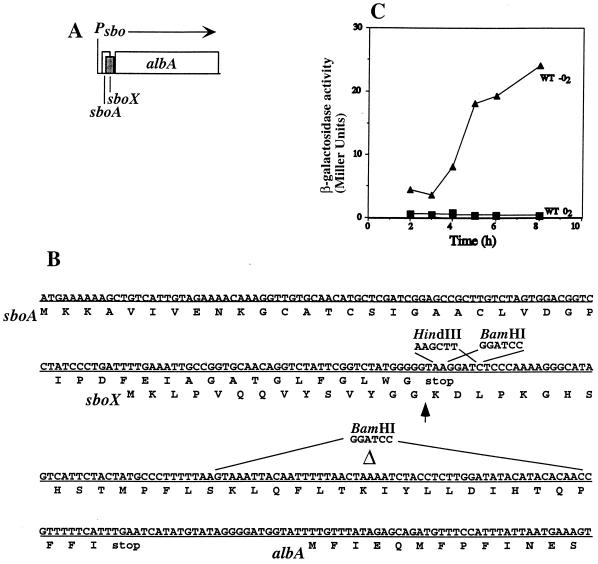
FIG. 4.
Organization of the sboAX locus. (A) Diagram of the sboA, sboX, and albA region of the sbo-alb gene cluster. Psbo indicates the location of the sbo-alb operon promoter, and the arrow marks the direction of transcription. (B) Nucleotide sequence of the sboAX genes and amino acid sequence (below the horizontal line) of the products. The restriction enzyme sites introduced into the sboX coding sequence (HindIII and BamHI) are indicated. The segment of the sboX gene that is deleted in the sboXΔ89-139 allele is labeled Δ. The BamHI sequence that replaced this segment is indicated. (C) Expression of sboX::lacZ under anaerobic (−O2) and aerobic (O2) conditions. An sboX-lacZ translational fusion was introduced into the amyE locus of B. subtilis JH642 cells. WT, wild type.
An in-frame deletion mutation was created that removed a segment of the sboX coding sequence extending from bp 89 to bp 138, replacing the fragment with a BamHI site (Fig. 4B). The resulting allele, sboXΔ89-139, did not confer a subtilosin-negative phenotype, nor did it detectably affect subtilosin production or immunity. It also did not have an effect on the expression of an ectopically expressed sboA-lacZ fusion (49). We conclude that the putative sboX product does not function in subtilosin production and is not required for antilisterial activity. The sboX::neo insertion mutations resides within the stem-loop structure located at the end of the sboA gene. It is possible that the insertion results in lower subtilosin production because it confers instability on the sboA mRNA.
To determine if the sboX coding sequence was translated in B. subtilis, the HindIII site was used to fuse sboX in frame with the truncated lacZ gene of plasmid ptrpGB1. The construct was introduced into the amyE locus, and lacZ activity was measured in samples collected from anaerobic and aerobic cultures. No sboX-lacZ expression could be detected in cells of aerobic cultures (Fig. 4C), but induction of expression was observed as anaerobic cultures entered the stationary phase of growth.
DISCUSSION
Eight of the nine genes of the sbo-alb operon function in the production of the antilisterial bacteriocin subtilosin, as shown by the phenotype produced by insertion mutations. Of the mutations created, those in sboA, albA, and albF are the most critical for the formation of active subtilosin. Mutations in sboX, albB, albC, albD, albE, and albG do not abolish subtilosin production but impair antilisterial activity. We do not know the primary functions of the alb gene products. The polypeptides encoded by albA, -C, -E, and -F show primary structural similarity to known proteins. AlbC very likely functions in the export of subtilosin, which it performs, perhaps, in conjunction with other alb products. We propose that AlbA, very likely a member of the MoeA/NifB/PqqE family (16), and AlbF, a member of a family of zinc endoproteinases, perform critical modifications of the presubtilosin peptide. AlbB, -C, and -D are required for immunity to subtilosin.
The newly discovered sboX gene encodes a bacteriocin-like precursor peptide bearing a GG motif resembling type II prebacteriocin cleavage sites (27). A neo cassette insertion mutation in sboX interrupts its coding sequence but also damages the sequence encoding the stem-loop structure residing at the 3′ end of sboA. The insertion would eliminate the sboX product but might also render sboA mRNA unstable and susceptible to 3′ exonucleolytic activity, which could explain the reduced subtilosin production of the sboX::neo mutant. The in-frame deletion allele sboXΔ89-139 has no detectable effect on subtilosin production or immunity or regulation of sboA expression. We do not know the function of the sboX gene.
AlbA and other members of the MoaA/NifB/PqqE family (16) possess two Cys clusters, one in the N-terminal half of the protein and the other at the C terminus. These are thought to be the locations of Fe-S centers that serve as the active sites in reactions involving hydration or dehydration of substrate compounds (38, 45). MoaA catalyzes a step of molybdenum cofactor synthesis in which precursor Z is produced from the phosphorylated guanosine precursor prior to MoeB-dependent sulfur addition (30, 45). Other members function in the synthesis of enzyme cofactors such as pyrroloquinoline quinone (5, 17, 43) and siroheme (11). A homolog of AlbA is encoded by the ybcQ gene of B. subtilis, which appears to reside in an operon that contains a small gene, ybcO, encoding another bacteriocin-like product (15).
The amino acid sequence of the N-terminal half of AlbF is very similar to those of known beta chain Zn metalloproteases of mitochondria that are associated with the cytochrome Bc1 reductase complex (3, 9, 29, 31) and the product of pqqF, which functions in cofactor (pyrroloquinoline quinone) synthesis (17, 44). The C-terminal half of AlbF shows no homology to known proteins but exhibits some similarity to the peptide binding protein encoded by oppA of Myxococcus xanthus. We suspect that AlbF catalyzes some step in the processing of presubtilosin.
Mutations in albB, -C, -D, and -G do not abolish subtilosin production but appear to reduce the amount of active peptide produced. In most cases, we do not know if partially active processing intermediates are produced and secreted or if the mutants simply produce less wild-type product. In the case of the albG mutant, nearly wild-type levels of a peptide are found in the culture supernatant, as judged from LH-20 chromatography and Tricine SDS-PAGE, but the activity of the product is significantly reduced (data not shown). This could be due to the accumulation of a processing intermediate that is efficiently secreted into the medium. The structure of this peptide product is under investigation.
Mutations in albB, -C, and, to a lesser extent, -D reduced the cell's immunity to subtilosin. The product of albC, a member of the ABC family of transport proteins, very likely participates in export of subtilosin. In several bacteriocin and lantibiotic production systems, export proteins have been shown to be required for complete immunity to the specific peptide produced (33, 35). The small 59-amino-acid hydrophobic peptide encoded by albB appears to play a critical role in immunity, since the albB mutant shows the most severe defect in subtilosin self-protection. AlbB may serve the same function as the other small, hydrophobic immunity peptides encoded by lantibiotic biosynthesis operons. Like other genes that confer bacteriocin immunity, albB is required for maximal bacteriocin production. How these products function in conferring self-protection and in bacteriocin production is not known.
In a previous report, we presented evidence that one or more products of the alb genes may function in the positive autoregulation of sbo-alb expression (49). This was based on the observation that high constitutive expression of the alb genes results in accelerated expression of an ectopically expressed sboA-lacZ fusion. We examined the effect of each alb insertion mutation on sboA-lacZ expression but found no significant changes. The high-level expression of the alb genes probably does not play a direct role in sbo-alb transcriptional regulation.
ACKNOWLEDGMENTS
We acknowledge the gift of high-pressure liquid chromatography-purified subtilosin from J. C. Vederas.
Support was provided by grant GM45898 from the National Institutes of Health and a grant from the Oregon Research Foundation. R.H. was supported by a grant from the Partnerships in Science Program that is funded through Research Corporation and the Murdock Charitable Trust. P.Z. gratefully acknowledges support from E. I. du Pont de Nemours, Inc.
REFERENCES
- 1.Babasaki K, Takao T, Shimonishi Y, Kurahashi K. Subtilosin A, a new antibiotic peptide produced by Bacillus subtilis 168: isolation, structural analysis, and biogenesis. J Biochem. 1985;98:583–603. doi: 10.1093/oxfordjournals.jbchem.a135315. [DOI] [PubMed] [Google Scholar]
- 2.Beuchat L R. Traditional fermented foods. In: Doyle M P, Beuchat L R, Montville T J, editors. Food microbiology: fundamentals and frontiers. Washington, D.C.: American Society for Microbiology; 1997. pp. 629–648. [Google Scholar]
- 3.Clary D O, Wahleithner J A, Wolstenholme D R. Sequence and arrangement of the genes for cytochrome b, URF1, URF4L, URF4, URF5, URF6 and five tRNAs in Drosophila mitochondrial DNA. Nucleic Acids Res. 1984;12:3747–3762. doi: 10.1093/nar/12.9.3747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Dubnau D, Davidoff-Abelson R. Fate of transforming DNA following uptake by competent Bacillus subtilis. I. formation and properties of the donor-recipient complex. J Mol Biol. 1971;56:209–221. doi: 10.1016/0022-2836(71)90460-8. [DOI] [PubMed] [Google Scholar]
- 5.Goodwin P M, Anthony C. The biochemistry, physiology and genetics of PQQ and PQQ-containing enzymes. Adv Microb Physiol. 1998;40:1–80. doi: 10.1016/s0065-2911(08)60129-0. [DOI] [PubMed] [Google Scholar]
- 6.Guerout-Fleury A M, Shazand K, Frandsen N, Stragier P. Antibiotic-resistance cassettes for Bacillus subtilis. Gene. 1995;167:335–336. doi: 10.1016/0378-1119(95)00652-4. [DOI] [PubMed] [Google Scholar]
- 7.Hansen J N. Nisin and related antimicrobial peptides. In: Strohl W R, editor. Biotechnology of antibiotics. New York, N.Y: Marcel Dekker Inc.; 1997. pp. 437–470. [Google Scholar]
- 8.Hoch J A. spo0 genes, the phosphorelay, and the initiation of sporulation. In: Sonenshein A L, Hoch J A, Losick R, editors. Bacillus subtilis and other gram-positive bacteria: biochemistry, physiology, and molecular genetics. Washington, D.C.: American Society for Microbiology; 1993. pp. 747–755. [Google Scholar]
- 9.Iwata S, Lee J W, Okada K, Lee J K, Iwata M, Rasmussen B, Link T A, Ramaswamy S, Jap B K. Complete structure of the 11-subunit bovine mitochondrial cytochrome bc1 complex. Science. 1998;281:64–71. doi: 10.1126/science.281.5373.64. [DOI] [PubMed] [Google Scholar]
- 10.Jack R W, Tagg F R, Ray B. Bacteriocins of gram-positive bacteria. Microbiol Rev. 1995;59:171–200. doi: 10.1128/mr.59.2.171-200.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kawasaki S, Arai H, Kodama T, Igarashi Y. Gene cluster for dissimilatory nitrite reductase (nir) from Pseudomonas aeruginosa: sequencing and identification of a locus for heme d1 biosynthesis. J Bacteriol. 1997;179:235–242. doi: 10.1128/jb.179.1.235-242.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Klaenhammer T R. Bacteriocins of lactic acid bacteria. Biochimie. 1988;70:337–349. doi: 10.1016/0300-9084(88)90206-4. [DOI] [PubMed] [Google Scholar]
- 13.Kleanthous C, Hemmings A M, Moore G R, James R. Immunity proteins and their specificity for endonuclease colicins: telling right from wrong in protein-protein recognition. Mol Microbiol. 1998;28:227–233. doi: 10.1046/j.1365-2958.1998.00811.x. [DOI] [PubMed] [Google Scholar]
- 14.Kolter R, Moreno F. Genetics of ribosomally synthesized peptide antibiotics. Annu Rev Microbiol. 1992;46:141–164. doi: 10.1146/annurev.mi.46.100192.001041. [DOI] [PubMed] [Google Scholar]
- 15.Kunst F, Ogasawara N, Moszer I, et al. The complete genome sequence of the gram-positive bacterium Bacillus subtilis. Nature. 1997;390:249–256. doi: 10.1038/36786. [DOI] [PubMed] [Google Scholar]
- 16.Menendez C, Igloi G, Henninger H, Brandsch R. A pAO1-encoded molybdopterin cofactor gene (moaA) of Arthrobacter nicotinovorans: characterization and site-directed mutagenesis of the encoded protein. Arch Microbiol. 1995;164:142–151. doi: 10.1007/BF02525320. [DOI] [PubMed] [Google Scholar]
- 17.Meulenberg J J, Sellink E, Riegman N H, Postma P W. Nucleotide sequence and structure of the Klebsiella pneumoniae pqq operon. Mol Gen Genet. 1992;232:284–294. doi: 10.1007/BF00280008. [DOI] [PubMed] [Google Scholar]
- 18.Montville T. The evolving impact of biotechnology of food microbiology. J Food Safety. 1989;10:87–97. [Google Scholar]
- 19.Montville T J, Winkowski K. Biologically based preservation systems and probiotic bacteria. In: Doyle M P, Beuchat L R, Montville T J, editors. Food microbiology: fundamentals and frontiers. Washington, D.C.: American Society for Microbiology; 1997. pp. 557–577. [Google Scholar]
- 20.Nakano M M, Hoffmann T, Zhu Y, Jahn D. Nitrogen and oxygen regulation of Bacillus subtilis nasDEF encoding NADH-dependent nitrite reductase by TnrA and ResDE. J Bacteriol. 1998;180:5344–5350. doi: 10.1128/jb.180.20.5344-5350.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Nakano M M, Magnuson R, Myers A, Curry J, Grossman A D, Zuber P. srfA is an operon required for surfactin production, competence development, and efficient sporulation in Bacillus subtilis. J Bacteriol. 1991;173:1770–1778. doi: 10.1128/jb.173.5.1770-1778.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Nakano M M, Marahiel M A, Zuber P. Identification of a genetic locus required for biosynthesis of the lipopeptide antibiotic surfactin in Bacillus subtilis. J Bacteriol. 1988;170:5662–5668. doi: 10.1128/jb.170.12.5662-5668.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nakano M M, Zheng G, Zuber P. Dual control of sbo-alb operon expression by the Spo0 and ResDE systems of signal transduction under anaerobic conditions in Bacillus subtilis. J Bacteriol. 2000;182:3274–3277. doi: 10.1128/jb.182.11.3274-3277.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Nakano M M, Zuber P. Anaerobic growth of a “strict aerobe.”. Annu Rev Microbiol. 1998;52:165–190. doi: 10.1146/annurev.micro.52.1.165. [DOI] [PubMed] [Google Scholar]
- 25.Nakano M M, Zuber P. Cloning and characterization of srfB, a regulatory gene involved in surfactin production and competence in Bacillus subtilis. J Bacteriol. 1989;171:5347–5353. doi: 10.1128/jb.171.10.5347-5353.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Nakano M M, Zuber P, Glaser P, Danchin A, Hulett F M. Two-component regulatory proteins ResD-ResE are required for transcriptional activation of fnr upon oxygen limitation in Bacillus subtilis. J Bacteriol. 1996;178:3796–3802. doi: 10.1128/jb.178.13.3796-3802.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Nes I F, Diep D B, Havarstein L S, Brurberg M B, Eijsink V, Holo H. Biosynthesis of bacteriocins in lactic acid bacteria. Antonie Van Leeuwenhoek. 1996;70:113–128. doi: 10.1007/BF00395929. [DOI] [PubMed] [Google Scholar]
- 28.Ogura M, Liu L, LaCelle M, Nakano M, Zuber P. Mutational analysis of ComS: evidence for the interaction of ComS and MecA in the regulation of competence development in Bacillus subtilis. Mol Microbiol. 1999;32:799–812. doi: 10.1046/j.1365-2958.1999.01399.x. [DOI] [PubMed] [Google Scholar]
- 29.Paces V, Rosenberg L E, Fenton W A, Kalousek F. The beta subunit of the mitochondrial processing peptidase from rat liver: cloning and sequencing of a cDNA and comparison with a proposed family of metallopeptidases. Proc Natl Acad Sci USA. 1993;90:5355–5358. doi: 10.1073/pnas.90.11.5355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pitterle D M, Johnson J L, Rajagopalan K V. The biosynthesis of molybdopterin. Purification and characterization of the converting factor. J Biol Chem. 1993;268:13506–13509. [PubMed] [Google Scholar]
- 31.Pollock R A, Hartl F U, Cheng M Y, Ostermann J, Horwich A, Neupert W. The processing peptidase of yeast mitochondria: the two co-operating components MPP and PEP are structurally related. EMBO J. 1988;7:3493–3500. doi: 10.1002/j.1460-2075.1988.tb03225.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Quadri L E, Sailer M, Roy K L, Vederas J C, Stiles M E. Chemical and genetic characterization of bacteriocins produced by Carnobacterium piscicola LV17B. J Biol Chem. 1994;269:12204–12211. [PubMed] [Google Scholar]
- 33.Ra S R, Wiao M, Immonen T, Pujana I, Saris E J. Genes responsible for nisin synthesis, regulation and immunity form a regulon of two operons and are induced by nisin in Lactococcus lactis N8. Microbiology. 1996;142:1281–1288. doi: 10.1099/13500872-142-5-1281. [DOI] [PubMed] [Google Scholar]
- 34.Sahl H-G, Dierbaum G. Lantibiotics: biosynthesis and biological activities or uniquely modified peptides from gram-positive bacteria. Annu Rev Microbiol. 1998;52:41–79. doi: 10.1146/annurev.micro.52.1.41. [DOI] [PubMed] [Google Scholar]
- 35.Saris P E, Immonen T, Reis M, Sahl H G. Immunity to lantibiotics. Antonie Van Leeuwenhoek. 1996;69:151–159. doi: 10.1007/BF00399420. [DOI] [PubMed] [Google Scholar]
- 36.Schagger H, von Jagow G. Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal Biochem. 1987;166:368–379. doi: 10.1016/0003-2697(87)90587-2. [DOI] [PubMed] [Google Scholar]
- 37.Shimotsu H, Henner D J. Construction of a single-copy integration vector and its use in analysis of regulation of the trp operon of Bacillus subtilis. Gene. 1986;43:85–94. doi: 10.1016/0378-1119(86)90011-9. [DOI] [PubMed] [Google Scholar]
- 38.Solomon P S, Shaw A L, Lane I, Hanson G R, Palmer T, McEwan A G. Characterization of a molybdenum cofactor biosynthetic gene cluster in Rhodobacter capsulatus which is specific for the biogenesis of dimethyl sulfoxide reductase. Microbiology. 1999;145:1421–1429. doi: 10.1099/13500872-145-6-1421. [DOI] [PubMed] [Google Scholar]
- 39.Sonenshein A L. Introduction to metabolic pathways. In: Sonenshein A L, Hoch J A, Losick R, editors. Bacillus subtilis and other gram-positive bacteria: physiology, biochemistry, and molecular biology. Washington, D.C.: American Society for Microbiology; 1993. pp. 127–132. [Google Scholar]
- 40.Steinmetz M, Richter R. Plasmids designed to alter the antibiotic resistance expressed by insertion mutations in Bacillus subtilis, through in vivo recombination. Gene. 1994;142:79–83. doi: 10.1016/0378-1119(94)90358-1. [DOI] [PubMed] [Google Scholar]
- 41.Strauch M A. AbrB, a transition state regulator. In: Sonenshein A L, Hoch J A, Losick R, editors. Bacillus subtilis and other gram-positive bacteria: physiology, biochemistry, and molecular biology. Washington, D.C.: American Society for Microbiology; 1993. pp. 757–764. [Google Scholar]
- 42.Sun G, Sharkova E, Chesnut R, Birkey S, Duggan M F, Sorokin A, Pujic P, Ehrlich S D, Hulett F M. Regulators of aerobic and anaerobic respiration in Bacillus subtilis. J Bacteriol. 1996;178:1374–1385. doi: 10.1128/jb.178.5.1374-1385.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Toyama H, Chistoserdova L, Lidstrom M E. Sequence analysis of pqq genes required for biosynthesis of pyrroloquinoline quinone in Methylobacterium extorquens AMI and the purification of a biosynthetic intermediate. Microbiology. 1997;143:595–602. doi: 10.1099/00221287-143-2-595. [DOI] [PubMed] [Google Scholar]
- 44.van Kraaij C, de Vos W M, Siezen R J, Kuipers O P. Lantibiotics: biosynthesis, mode of action and applications. Nat Prod Rep. 1999;16:575–587. doi: 10.1039/a804531c. [DOI] [PubMed] [Google Scholar]
- 45.Wuebbens M M, Rajagopalan K V. Investigation of the early steps of molybdopterin biosynthesis in Escherichia coli through the use of in vivo labeling studies. J Biol Chem. 1995;270:1082–1087. doi: 10.1074/jbc.270.3.1082. [DOI] [PubMed] [Google Scholar]
- 46.Yanisch-Perron C, Vieira J, Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33:103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- 47.Youngman P, Poth H, Green B, York K, Olmedo G, Smith K. Methods for genetic manipulation, cloning, and functional analysis of sporulation genes in Bacillus subtilis. In: Smith I, Slepecky R A, Setlow P, editors. Regulation of procaryotic development. Washington, D.C.: American Society for Microbiology; 1989. pp. 65–87. [Google Scholar]
- 48.Zheng G, Slavik M F. Isolation, partial purification and characterization of a bacteriocin produced by a newly isolated Bacillus subtilis strain. Lett Appl Microbiol. 1999;28:363–367. doi: 10.1046/j.1365-2672.1999.00545.x. [DOI] [PubMed] [Google Scholar]
- 49.Zheng G, Yan L Z, Vederas J C, Zuber P. Genes of the sbo-alb locus of Bacillus subtilis are required for production of the antilisterial bacteriocin subtilosin. J Bacteriol. 1999;181:7346–7355. doi: 10.1128/jb.181.23.7346-7355.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zuber P, Losick R. Use of a lacZ fusion to study the role of the spo0 genes of Bacillus subtilis in developmental regulation. Cell. 1983;35:275–283. doi: 10.1016/0092-8674(83)90230-1. [DOI] [PubMed] [Google Scholar]
- 51.Zuber P, Nakano M M, Marahiel M A. Peptide antibiotics. In: Sonenshein A L, Hoch J A, Losick R, editors. Bacillus subtilis and other gram-positive bacteria: physiology, biochemistry, and molecular biology. Washington, D.C.: American Society for Microbiology; 1993. pp. 897–916. [Google Scholar]