Abstract
The spacer A/T region is a positive cis-acting DNA element that was identified in the Chlamydia trachomatis rRNA promoter region. We have now demonstrated that similar sequences in other chlamydial promoters are important for transcription. Substitution of candidate spacer A/T regions in four chlamydial promoters decreased transcription by partially purified C. trachomatis RNA polymerase in an in vitro transcription assay. Addition of a spacer A/T region to the dnaK promoter, which does not contain an identifiable spacer A/T region, increased transcription 16-fold. Transcription of Escherichia coli promoters by C. trachomatis RNA polymerase also appeared to be dependent on the spacer A/T region. However, the effect of the spacer A/T region on transcription by E. coli RNA polymerase was small. In summary, the spacer A/T region is a novel DNA element that is required for high-level transcription of many promoters by chlamydial RNA polymerase.
Chlamydia trachomatis is a pathogenic bacterium with an unusual intracellular life cycle that involves sequential conversion between two morphologic forms (reviewed in references 13 and 16). Starting at 2 to 3 h postinfection (hpi), the infectious but metabolically inert elementary body (EB) begins a striking change, with decondensation of its DNA-containing nucleoid. By 6 to 8 hpi, this conversion into the larger and metabolically active reticulate body (RB) is complete. Later in the cycle, after proliferation of RBs by binary fission, individual RBs convert back to EBs.
Transcription is regulated during the chlamydial life cycle and appears to be generally restricted to RBs. Low levels of several early transcripts have been described within a few hours of infection (24). RNA is first measurable by Northern blots at the time when the EB-to-RB conversion is complete, with detection of rRNA and mRNA for the major outer membrane protein (MOMP) at 7 to 8 hpi. The mechanisms that regulate the transcription of rRNA and mRNA are not known, although it has been proposed that the condensed genome in EBs causes a global transcriptional silencing which is relieved upon conversion of EBs into RBs (2).
In previous work, we have shown that a cis-acting DNA element is required for high-level transcription of the C. trachomatis rRNA promoter (19, 20). We have called this element the spacer A/T region because of its location in the spacer region between the −35 and −10 promoter elements and because of the strong sequence preference for A and T residues. Substitution with a single C or G residue decreased rRNA promoter activity significantly. Candidate spacer A/T regions can be identified in many other chlamydial promoters as an AT-rich sequence immediately downstream of the predicted −35 promoter element (20). In the present study, we have examined additional promoters in order to determine whether the spacer A/T region is unique to the rRNA promoter or whether it has a more general role in chlamydial transcription. Our results demonstrate that in vitro transcription of chlamydial and Escherichia coli promoters by chlamydial RNA polymerase is stimulated by the presence of a spacer A/T region.
MATERIALS AND METHODS
Reagents.
The following products were obtained from the sources given and were used according to the manufacturer's specifications: restriction enzymes, calf alkaline phosphatase, T4 DNA ligase, rRNasin, and Thermus aquaticus DNA polymerase, Promega Biotech (Madison, Wis.); T4 polynucleotide kinase, New England Biolabs (Beverly, Mass.); T7 Sequenase DNA polymerase and dideoxynucleotide kit, United States Biochemical Corp. (Cleveland, Ohio); nucleoside triphosphates, 3′-O-methylguanosine 5′-triphosphate, and 32P-containing nucleoside triphosphates, Amersham Corp. (Arlington Heights, Ill.); SeaPlaque agarose, FMC Bioproducts (Rockland, Maine); ampicillin, Fisher Scientific (Pittsburgh, Pa.); and purified E. coli RNA polymerase, Epicentre (Madison, Wis.).
DNA manipulation.
Nucleic acid preparation and analysis were performed according to standard recombinant DNA protocols. DNA was amplified by PCR as described previously (20). The dideoxy-chain termination method for sequencing double-stranded plasmid DNA was performed with a Sequenase kit from United States Biochemical Corp.
Construction of transcription plasmids containing wild-type promoters.
The promoter regions of the following genes were amplified from C. trachomatis serovar L2 genomic DNA by PCR: omcB (−36 to +5), hctA (−135 to +5), ltuA (−135 to +5), ltuB (−132 to +5), and dnaK (−235 to +6). The E. coli lacUV5 promoter (−42 to +5) and rrnBp1 (−60 to +6, which includes the UP element [20]) were amplified from E. coli K-12 genomic DNA by PCR. Each promoter insert was cloned upstream of a promoterless G-less cassette transcription template in plasmid pMT504 (19). pMT504 also contains an internal control transcription template consisting of C. trachomatis rRNA P1 upstream of a shorter G-less cassette. Transcription of each plasmid produced a 158-nucleotide test transcript (159 nucleotides if the test promoter extended to +6 instead of +5) and a 130-nucleotide control transcript.
Construction of transcription plasmids containing substitutions in the spacer A/T regions.
Specific mutations were introduced into each of the wild-type promoters by PCR with an oligonucleotide primer containing the desired mutation in the predicted spacer A/T region. A 4-bp substitution was introduced at −28 to −25 of omcB, −26 to −23 of hctA, −28 to −25 of ltuA, and −29 to −26 of ltuB. In each case, A residues were replaced with C residues, and T residues were replaced with G residues. A sequence resembling a spacer A/T region was introduced into the dnaK promoter by substitution with A residues at −27, −25, and −24. Similarly, a spacer A/T region-like sequence was altered in E. coli lacUV5 with a 5-bp substitution (TTTAT to GGGCG at −29 to −25) and introduced into E. coli rrnBp1 (GGCCG to AAAAA at −30 to −26).
Purification of C. trachomatis RNA polymerase.
RNA polymerase was partially purified from C. trachomatis LGV serovar L2 at 20 hpi by heparin-agarose chromatography as previously described (19).
In vitro transcription.
The following reaction mixture was assembled: 50 mM potassium acetate, 8.1 mM magnesium acetate, 50 mM Tris acetate (pH 8.0), 27 mM ammonium acetate, 2 mM dithiothreitol (DTT), 400 μM ATP, 400 μM UTP, 1.2 μM CTP, 0.21 μM [α-32P]CTP (800 Ci/mmol), 100 μM 3′-O-methylguanosine 5′-triphosphate, Na salt, 18 U of rRNasin, 10% glycerol, and 0.5 μl of heparin-agarose-purified C. trachomatis RNA polymerase or 0.003 U of E. coli ς70 RNA polymerase. The supercoiled DNA template (final concentration, 25 nM) was added, and the reaction mixture was incubated at 37°C for 5 min. Heparin was added to a final concentration of 150 μg/ml, and the incubation was continued at 37°C for a further 10 min. The final reaction volume was 10 μl. The reaction was terminated by the addition of 10 μl of stop solution (95% formamide, 20 mM EDTA, 0.05% bromophenol blue, 0.05% xylene cyanol). A 7-μl portion of the sample was electrophoresed on an 8 M urea–6% polyacrylamide gel. Transcripts were visualized by autoradiography and quantified with a Molecular Dynamics (Sunnyvale, Calif.) PhosphorImager.
Calculation of promoter activity.
Promoter activity is expressed as the ratio of transcript produced by C. trachomatis RNA polymerase (or E. coli RNA polymerase) relative to transcription from the control promoter, C. trachomatis rRNA P1. Three measurements of promoter activity were obtained for each promoter, and a mean and a standard deviation were calculated.
RESULTS
Sequences resembling the rRNA P1 spacer A/T region are important for transcription of other chlamydial promoters.
To determine if candidate spacer A/T regions in other chlamydial promoters also function as positive cis-acting elements, we used a mutational approach to measure the effect of each spacer A/T region on transcription. For each of four C. trachomatis genes, omcB, hctA, ltuA, and ltuB (Fig. 1A), we constructed a mutant promoter of the spacer A/T region without alteration of the predicted −10 and −35 promoter elements. A 4-bp substitution was introduced into omcB at −28 to −25 (AATT→CCGG), into hctA at −26 to −23 (ATTT→CGGG), into ltuA at −28 to −25 (TTTA→GGGC), and into ltuB at −29 to −26 (AAAA→CCCC). Transcription of the mutant and wild-type promoters was then compared using in vitro transcription assays.
FIG. 1.
(A) Sequence alignment of selected C. trachomatis promoters (modified from reference 20) containing predicted spacer A/T regions. Predicted −35 and −10 promoter sequences are underlined, and predicted spacer A/T regions are double underlined. Dots above the sequence indicate in vivo transcription initiation sites. Spacer A/T regions that have been shown to be functional are indicated by plus signs. n/t, not tested. Substitutions which introduced a synthetic spacer A/T region in the dnaK promoter are shown in lowercase type. The consensus E. coli ς70 promoter sequence is shown for comparison. GenBank accession numbers are as follows: for rRNA P1 (mouse pneumonitis strain [MoPn]), M18268; for ompA (serovar L2 MOMP P2), M14738; for omcB (L2 60-kDa cysteine-rich protein), X54450; for hctA (L2), M60902; for ltuA (L2), L40822; for ltuB (L2), L40838; for rs1 (MoPn ribosomal protein S1), M23000; for ORF 8 (L2), X07547; for groE (MoPn), L12004; for ORF 7 (L2), X07547; and for dnaK (MoPn), M62819. (B) Sequences of the promoter regions of the E. coli lacUV5 promoter and E. coli rrnBp1. The sequence that resembles a chlamydial spacer A/T region in the wild-type (wt) lacUV5 promoter was replaced by a 5-bp substitution (A/T−). In rrnBp1, a 5-bp substitution was introduced to create a synthetic spacer A/T region (A/T+). Substitutions are in lowercase type.
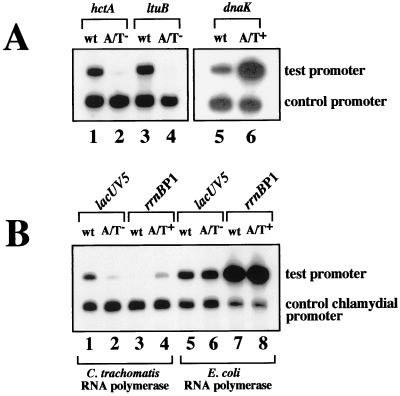
For all four promoters, sequence substitutions in the spacer A/T region significantly decreased transcription by partially purified C. trachomatis RNA polymerase. Examples of the assay results for the hctA promoter (lanes 1 and 2) and the ltuB promoter (lanes 3 and 4) are shown in Fig. 2A. Promoter activity was decreased 5-fold for the omcB promoter, 5-fold for the hctA promoter, 12-fold for the ltuA promoter, and 15-fold for the ltuB promoter. These results are shown in Table 1 as the fold activation of each promoter due to the spacer A/T region. Each wild-type and mutant promoter was also transcribed by E. coli RNA polymerase, but the effect of the substitutions was small. Substitutions in the spacer A/T region decreased transcription by E. coli RNA polymerase 1.6-fold for the omcB promoter, 0.9-fold for the hctA promoter, 2.2-fold for the ltuA promoter, and 1.6-fold for the ltuB promoter (Table 1).
FIG. 2.
(A) Effect of the spacer A/T region on in vitro transcription by C. trachomatis RNA polymerase. Lane 1, wild-type (wt) hctA promoter; lane 2, hctA promoter with a 4-bp substitution of the spacer A/T region (A/T−); lane 3, wild-type ltuB promoter; lane 4, ltuB promoter with a 4-bp substitution of the spacer A/T region; lane 5, wild-type dnaK promoter; lane 6, dnaK promoter containing a synthetic spacer A/T region (A/T+). C. trachomatis rRNA P1 (+53 to −5) was used as the control promoter. (B) Effect of the spacer A/T region on in vitro transcription of E. coli promoters by C. trachomatis RNA polymerase (lanes 1 to 4) or E. coli RNA polymerase (lanes 5 to 8). Lanes 1 and 5, wild-type lacUV5 promoter; lanes 2 and 6, lacUV5 promoter with a 5-bp substitution of the spacer A/T region; lanes 3 and 7, wild-type rrnBp1; lanes 4 and 8, rrnBp1 containing a synthetic spacer A/T region.
TABLE 1.
Effects of the spacer A/T region on promoter activity
| Gene | Fold activation ± SDa
|
|
|---|---|---|
| C. trachomatis RNA polymerase | E. coli RNA polymerase | |
| C. trachomatis | ||
| omcB | 5.4 ± 0.3 | 1.6 ± 0.0 |
| hctA | 4.8 ± 0.5 | 0.9 ± 0.2 |
| ltuA | 11.9 ± 2.2 | 2.2 ± 0.1 |
| ltuB | 15.3 ± 0.8 | 1.6 ± 0.1 |
| dnaK | 15.7 ± 0.9 | 0.7 ± 0.1 |
| E. coli lacUV5 | 7.0 ± 0.4 | 0.9 ± 0.0 |
| E. coli rrnBp1 | 13.3 ± 1.8 | 1.8 ± 0.1 |
Fold activation was determined as the ratio of transcription of each promoter in the presence and absence of the spacer A/T region. Each result is the mean of three separate experiments.
Addition of a spacer A/T region to the dnaK promoter increases transcription by C. trachomatis RNA polymerase.
Of the approximately one dozen chlamydial promoter sequences that have been predicted on the basis of a mapped transcription initiation site, only the promoters for dnaK and the plasmid ORF 7 do not contain AT-rich sequences in the location of the spacer A/T region (Fig. 1A). The dnaK promoter is transcribed by chlamydial RNA polymerase despite the lack of a predicted spacer A/T region (21). To determine if a synthetic spacer A/T region can stimulate transcription, we introduced a spacer A/T region into the dnaK promoter by substituting A residues at positions −27, −25 and −24 (Fig. 1A) and measured the effect on promoter activity. The synthetic spacer A/T region produced a 16-fold stimulation of transcription by C. trachomatis RNA polymerase (Fig. 2A, lanes 5 and 6), in contrast to a 1.5-fold decrease in transcription by E. coli RNA polymerase (Table 1).
Transcription of E. coli promoters by C. trachomatis RNA polymerase is dependent on the spacer A/T region.
To determine if the spacer A/T region is important for transcription of heterologous promoters by chlamydial RNA polymerase, we used partially purified C. trachomatis RNA polymerase to transcribe two E. coli promoters. We chose two strong E. coli promoters, the lacUV5 promoter, which contains a predicted spacer A/T region, and rrnBp1, which does not (Fig. 1B). While both promoters were transcribed by chlamydial RNA polymerase, rrnBp1 was transcribed at minimally detectable levels (Fig. 2B, lanes 1 and 3). A 5-bp substitution of the spacer A/T region in the lacUV5 promoter decreased transcription sevenfold (Fig. 2B, lane 2). Interestingly, chlamydial RNA polymerase was able to transcribe rrnBp1 at 13-fold-higher levels when a synthetic spacer A/T region was introduced into this E. coli promoter (Fig. 2B, lane 4). In contrast, the presence or absence of the spacer A/T region had little effect on transcription by E. coli RNA polymerase. Transcription of the lacUV5 promoter was increased 1.1-fold when the spacer A/T region was replaced, and transcription of rrnBp1 increased 1.8-fold with the addition of a spacer A/T region (Fig. 2B, lanes 5 to 8). The fold activation of each promoter due to the spacer A/T region is shown in Table 1.
DISCUSSION
In this report, we have demonstrated that the presence of the cis-acting spacer A/T region in multiple promoters stimulated transcription by partially purified chlamydial RNA polymerase but not by E. coli RNA polymerase. Transcription was stimulated by native spacer A/T regions and synthetic spacer A/T regions that were added to promoters which do not contain this cis-acting element. Interestingly, transcription of E. coli promoters by chlamydial RNA polymerase was also stimulated by the presence of a spacer A/T region. This novel cis-acting DNA element does not appear to be necessary for transcription of the wild-type dnaK promoter, although addition of a synthetic spacer A/T region to the dnaK promoter created the strongest promoter that we have tested in our in vitro C. trachomatis transcription system.
A functional spacer A/T region has now been identified in the native sequences of six chlamydial promoters. The four promoters presented here control genes that are only expressed late in the life cycle, at 20 to 30 hpi. These late genes include omcB, which encodes the 60-kDa cysteine-rich outer membrane protein (9), hctA, which encodes a histone-like protein (22), and ltuA and ltuB, whose gene products are of unknown function (6). The two promoters that have been previously shown to contain a spacer A/T region are rRNA P1 (19, 20) and the promoter for ompA, which encodes MOMP (4). Both of these genes are transcribed throughout the intracellular life cycle, starting from the time of conversion of EBs to RBs.
The spacer A/T region appears to be conserved among chlamydial species. Potential spacer A/T regions can be identified in Chlamydia pneumoniae and Chlamydia psittaci homologs of C. trachomatis genes containing functional spacer A/T regions (Fig. 3). Sequence inspection shows similarities between the predicted promoter sequence of homologs, suggesting a conserved chlamydial promoter specificity.
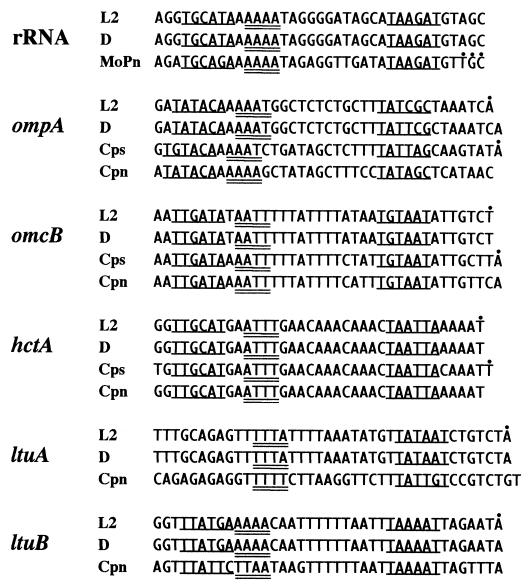
FIG. 3.
Sequence alignment of C. trachomatis promoters containing a functional spacer A/T region with predicted promoter sequences from other chlamydial species (C. trachomatis serovars L2 and D, and the mouse pneumonitis [MoPn] strain; C. psittaci [Cps]; and C. pneumoniae [Cpn]). Predicted −35 and −10 promoter elements are underlined, and predicted spacer A/T regions are double underlined. Dots above sequences indicate in vivo transcription initiation sites (for rRNA, in MoPn [5]; for ompA, in L2 [18] and C. psittaci strain GPIC [25]; for omcB, in L2 [6] and C. psittaci strain 6BC [11]; for hctA, in L2 [6] and the meningopneumonitis subspecies of C. psittaci [8]; and for ltuA and ltuB, in L2 [6]).
The optimal promoter sequence recognized by C. trachomatis RNA polymerase has not been completely determined. We have defined the optimal −10 promoter element recognized in the context of rRNA P1, and it strongly resembles the consensus E. coli ς70 −10 promoter element; however, the −35 promoter element of rRNA P1 appears to be suboptimal in function under in vitro transcription conditions (20). Sequences close to the consensus ς70 promoter structure alone do not appear to be sufficient for transcription, as E. coli rrnBp1 and the mutant lacUV5 promoter lacking a spacer A/T region were transcribed at minimally detectable levels.
While the spacer A/T region appears to function as a positive cis-acting element, its mechanism of action is not known. In E. coli, the spacer region is not known to be involved in a specific contact with RNA polymerase (23). Furthermore, E. coli promoters do not show sequence homology in this region (7). At least three distinct models can be proposed for the function of the spacer A/T region in Chlamydia. In the first two models, the spacer A/T region may serve as a binding site for a trans-acting factor, which could be either RNA polymerase or a transcriptional activator. In the third model, the DNA structure alone may affect transcription in a factor-independent manner.
In the first model, the spacer A/T region may function as a promoter element that makes contact with RNA polymerase. While the downstream promoter element at −10 appears to be conserved in Chlamydia, it is possible that the upstream promoter element may be located in the spacer A/T region rather than at −35. As noted, the chlamydial rRNA P1 −35 sequences are suboptimal for in vitro transcription (20). However, the −35 promoter recognition domain (sigma subunit subregion 4.2) of the major RNA polymerase is well conserved between C. trachomatis and other prokaryotes (10). Thus, chlamydial RNA polymerase would be predicted to recognize the −35 element. It is possible that the spacer A/T region, which is located immediately downstream of the −35 promoter region, might function as part of an extended −35 promoter element instead of as a replacement for the −35 sequences.
Activation by the spacer A/T region is dependent on ς66, the major sigma subunit of C. trachomatis RNA polymerase (19). While the spacer A/T region may serve as a binding site for ς66, it is also possible that another RNA polymerase subunit is involved in the contact. For example, the alpha subunit of prokaryotic RNA polymerase binds to the UP element, another AT-rich promoter element, to stimulate transcription (15). It may be possible to determine if ς66 is involved in a direct interaction with the spacer A/T region by using a hybrid RNA polymerase consisting of C. trachomatis ς66 and E. coli core enzyme (12).
In the second model, the spacer A/T region may function as a binding site for an activator of chlamydial transcription. This hypothesis predicts that the putative activator is present in the partially purified chlamydial RNA polymerase preparation used in our experiments. A transcriptional activator might also explain the “RNA polymerase sigma subunit paradox” propounded by Stephens (17). This observation noted the discordance between strong sequence conservation in the promoter recognition domains of C. trachomatis ς66 and E. coli ς70, which would predict a conserved promoter specificity, and the relative lack of conservation in the promoter sequences of the two bacteria. The observed sequence variation in the −35 regions of many chlamydial promoters may indicate a suboptimal structure that requires activation by a transcription factor. It should be noted that the spacer A/T region is located within the spacer region of the promoter, whereas the large majority of prokaryotic transcriptional activators bind to sites upstream of the promoter (14).
In the third model, the intrinsic sequence of the spacer A/T region may exert its effect on promoter activity through DNA structure without a specific contact with RNA polymerase. Experiments with E. coli have demonstrated that sequence substitutions in the spacer region can affect promoter activity, presumably by altering the relative orientations or local structures of the −10 and −35 promoter elements (1, 3). It is noteworthy that while substitution of the sequence of the spacer A/T region had a great effect on chlamydial transcription, there was a minimal effect on transcription by E. coli RNA polymerase.
The sequence of the spacer A/T region could contribute to DNA structure through its effect on DNA curvature or flexibility. Some of the chlamydial spacer A/T regions are part of a run of A residues or T residues (Fig. 1A), which are sequences known to form a stiff intrinsic DNA bend, although of opposite orientation (1). The spacer A/T region of chlamydial rRNA P1 is part of an oligo(A) tract, and we have shown that substitution of a T residue in this tract, which would disrupt the intrinsic DNA bend, did not alter promoter activity significantly (20). This result suggests that stimulation of transcription by the spacer A/T region is not mediated by a DNA bend.
The observed preference for an AT-rich sequence in the spacer A/T region could also suggest that energetic considerations, such as localized melting of DNA in this region, are important for promoter activity. Localized strand separation is associated with open complex formation during transcription initiation with ς70 RNA polymerase, but it occurs in the vicinity of the −10 promoter element and the transcription initiation site (23), and thus at a different location from the spacer A/T region.
The spacer A/T region appears to have a novel role in the transcription of promoters by chlamydial RNA polymerase. Its possible function as a promoter element or a DNA structural element would be unusual for transcription initiation by a ς70- type RNA polymerase. If, instead, chlamydial transcription is dependent on an activator that binds to the spacer A/T region, the implications for chlamydial biology are significant. Transcriptional activation via the spacer A/T region might provide a mechanism for the control of gene expression during the chlamydial life cycle. The putative activator would be predicted to stimulate transcription throughout much of the RB stage, as suggested by the presence of functional spacer A/T regions in the promoters of genes which are transcribed when EBs convert to RBs, as well as in late genes. In addition, transcriptional activation of rRNA synthesis may serve as a control point for the assembly of the translational machinery.
ACKNOWLEDGMENTS
We thank Jison Choi, Adam Wilson, and Hilda Hiu Yin Yu for support and suggestions and Bert Semler, Wes Hatfield, and Marian Waterman for critical review of the manuscript.
This work was supported by a grant from the NIH (AI 44198).
REFERENCES
- 1.Auble D T, Allen T L, deHaseth P L. Promoter recognition by Escherichia coli RNA polymerase. Effects of substitutions in the spacer DNA separating the −10 and −35 regions. J Biol Chem. 1986;261:11202–11206. [PubMed] [Google Scholar]
- 2.Barry C E, III, Hayes S, Hackstadt T. Nucleoid condensation in Escherichia coli that express a chlamydial histone homolog. Science. 1992;256:377–379. doi: 10.1126/science.256.5055.377. [DOI] [PubMed] [Google Scholar]
- 3.Collis C M, Molloy P L, Both G W, Drew H R. Influence of the sequence-dependent flexure of DNA on transcription in E. coli. Nucleic Acids Res. 1989;17:9447–9468. doi: 10.1093/nar/17.22.9447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Douglas A L, Hatch T P. Mutagenesis of the P2 promoter of the major outer membrane protein gene of Chlamydia trachomatis. J Bacteriol. 1996;178:5573–5578. doi: 10.1128/jb.178.19.5573-5578.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Engel J, Ganem D. Identification and comparison of putative chlamydial promoter elements. In: Van der Ploeg L, editor. Immune recognition and evasion: molecular aspects of host-parasite interaction. San Diego, Calif: Academic Press Inc.; 1990. pp. 245–260. [Google Scholar]
- 6.Fahr M J, Douglas A L, Xia W, Hatch T P. Characterization of late gene promoters of Chlamydia trachomatis. J Bacteriol. 1995;177:4252–4260. doi: 10.1128/jb.177.15.4252-4260.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hawley D K, McClure W R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983;11:2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kaul R, Tao S, Wenman W M. Interspecies structural diversity among chlamydial genes encoding histone H1. Gene. 1992;112:129–132. doi: 10.1016/0378-1119(92)90314-f. [DOI] [PubMed] [Google Scholar]
- 9.Lambden P R, Everson J S, Ward M E, Clarke I N. Sulfur-rich proteins of Chlamydia trachomatis: developmentally regulated transcription of polycistronic mRNA from tandem promoters. Gene. 1990;87:105–112. doi: 10.1016/0378-1119(90)90500-q. [DOI] [PubMed] [Google Scholar]
- 10.Lonetto M, Gribskov M, Gross C A. The ς70 family: sequence conservation and evolutionary relationships. J Bacteriol. 1992;174:3843–3849. doi: 10.1128/jb.174.12.3843-3849.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mathews S A, Douglas A, Sriprakash K S, Hatch T P. In vitro transcription in Chlamydia psittaci and Chlamydia trachomatis. Mol Microbiol. 1993;7:937–946. doi: 10.1111/j.1365-2958.1993.tb01185.x. [DOI] [PubMed] [Google Scholar]
- 12.Mathews S A, Stephens R S. DNA structure and novel amino and carboxyl termini of the Chlamydia sigma 70 analogue modulate promoter recognition. Microbiology. 1999;145:1671–1681. doi: 10.1099/13500872-145-7-1671. [DOI] [PubMed] [Google Scholar]
- 13.Moulder J W. Interaction of chlamydiae and host cells in vitro. Microbiol Rev. 1991;55:143–190. doi: 10.1128/mr.55.1.143-190.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Reznikoff W S, Siegele D A, Cowing D W, Gross C A. The regulation of transcription initiation in bacteria. Annu Rev Genet. 1985;19:355–387. doi: 10.1146/annurev.ge.19.120185.002035. [DOI] [PubMed] [Google Scholar]
- 15.Ross W, Gosink K K, Salomon J, Igarashi K, Zou C, Ishihama A, Severinov K, Gourse R L. A third recognition element in bacterial promoters: DNA binding by the α subunit of RNA polymerase. Science. 1993;262:1407–1413. doi: 10.1126/science.8248780. [DOI] [PubMed] [Google Scholar]
- 16.Schachter J. The intracellular life of Chlamydia. Curr Top Microbiol Immunol. 1988;138:109–139. [PubMed] [Google Scholar]
- 17.Stephens R S. Molecular genetics of Chlamydia. In: Bowie W R, et al., editors. Chlamydial infections 1990. Cambridge, United Kingdom: Cambridge University Press; 1990. pp. 63–72. [Google Scholar]
- 18.Stephens R S, Wagar E A, Edman U. Developmental regulation of tandem promoters for the major outer membrane protein of Chlamydia trachomatis. J Bacteriol. 1988;170:744–750. doi: 10.1128/jb.170.2.744-750.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Tan M, Engel J N. Identification of sequences necessary for transcription in vitro from the Chlamydia trachomatis rRNA P1 promoter. J Bacteriol. 1996;178:6975–6982. doi: 10.1128/jb.178.23.6975-6982.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tan M, Gaal T, Gourse R L, Engel J N. Mutational analysis of the Chlamydia trachomatis rRNA P1 promoter defines four regions important for transcription in vitro. J Bacteriol. 1998;180:2359–2366. doi: 10.1128/jb.180.9.2359-2366.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tan M, Wong B, Engel J N. Transcriptional organization and regulation of the dnaK and groE operons of Chlamydia trachomatis. J Bacteriol. 1996;178:6983–6990. doi: 10.1128/jb.178.23.6983-6990.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Tao S, Kaul R, Wenman W M. Identification and nucleotide sequence of a developmentally regulated gene encoding a eukaryotic histone H1-like protein from Chlamydia trachomatis. J Bacteriol. 1991;173:2818–2822. doi: 10.1128/jb.173.9.2818-2822.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.von Hippel P H, Bear D G, Morgan W D, McSwiggen J A. Protein-nucleic acid interactions in transcription: a molecular analysis. Annu Rev Biochem. 1984;53:389–446. doi: 10.1146/annurev.bi.53.070184.002133. [DOI] [PubMed] [Google Scholar]
- 24.Wichlan D G, Hatch T P. Identification of an early-stage gene of Chlamydia psittaci 6BC. J Bacteriol. 1993;175:2936–2942. doi: 10.1128/jb.175.10.2936-2942.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yuan Y, Zhang Y X, Manning D S, Caldwell H D. Multiple tandem promoters of the major outer membrane protein gene (omp1) of Chlamydia psittaci. Infect Immun. 1990;58:2850–2855. doi: 10.1128/iai.58.9.2850-2855.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]