Summary
The cyclin-dependent kinase CDK12 has garnered interest as a cancer therapeutic target as DNA damage response genes are particularly suppressed by loss of CDK12 activity. In this study, we assessed the acute effects of CDK12 inhibition on transcription and RNA processing using nascent RNA Bru-seq and BruChase-seq. Acute transcriptional changes were overall small after CDK12 inhibition but over 600 genes showed intragenic premature termination, including DNA repair and cell cycle genes. Furthermore, many genes showed reduced transcriptional readthrough past the end of genes in the absence of CDK12 activity. RNA turnover was dramatically affected by CDK12 inhibition and importantly, caused increased degradation of many transcripts from DNA damage response genes. We also show that co-transcriptional splicing was suppressed by CDK12 inhibition. Taken together, these studies reveal the roles of CDK12 in regulating transcription elongation, transcription termination, co-transcriptional splicing, and RNA turnover.
Subject areas: Biological sciences, molecular biology, cell biology
Graphical abstract

Highlights
-
•
Over 600 genes showed prematurely terminated transcription when CDK12 was inhibited
-
•
CDK12 promotes transcriptional readthrough past transcription end sites (TESs)
-
•
CDK12 promotes splicing and affects transcript stability
Biological sciences; Molecular biology; Cell biology.
Introduction
CDK12 is a kinase that phosphorylates the carboxy-terminal domain (CTD) of the largest subunit of RNA polymerase II. It interacts with cyclin K and localizes to active genes (Bartkowiak et al., 2010). The human CDK12 gene was first cloned under the name CrkRS (Ko et al., 2001) but was later re-named CDK12 (Chen et al., 2006). As a CTD-kinase, CDK12 acts downstream of CDK7 (Bosken et al., 2014) and CDK9 (Bartkowiak et al., 2010) during transcription elongation and phosphorylates Ser2, Ser5, and Thr4 of the CTD of RNAPII. Such phosphorylation is stimulated by the pre-phosphorylation of the Ser-7 site by CDK7 (Bosken et al., 2014; Glover-Cutter et al., 2009; Greenleaf, 2019; Liang et al., 2020; Tellier et al., 2020). ChIP-seq experiments have shown that CDK12-binding is more prominent in gene bodies and toward the 3′-ends of genes compared to the 5′-ends of genes. In addition, CDK12 binding is highly enriched at enhancer elements (Greenleaf, 2019; Zhang et al., 2016). CDK12 phosphorylation of RNPII CTD at Ser2 and Thr4 plays important roles in transcription elongation (Bayles et al., 2019; Chirackal Manavalan et al., 2019; Tellier et al., 2020; Zhang et al., 2016) as well as for 3′-end complex formation and termination (Davidson et al., 2014; Eifler et al., 2015). Depletion or inhibition of CDK12 activity in cells leads to premature polyadenylation and termination at intronic polyadenylation sites (IPAs) (Dubbury et al., 2018; Krajewska et al., 2019). Genes encoding DNA damage response genes are especially prone to undergo premature termination following CDK12 inactivation owing to their above-average gene length and high number of IPAs (Dubbury et al., 2018; Krajewska et al., 2019). In addition to reduced Ser-2 phosphorylation in the bodies and 3′-ends of genes, loss of CDK12 indirectly leads to reduced H3K36me3 in the chromatin of active genes (Bowman et al., 2013). Loss of H3K36me3 in active genes has been shown to lead to increased sensitivity to ionizing radiation by reduced homologous recombination repair (HRR) (Aymard et al., 2014; Greenleaf, 2019; Krajewska et al., 2019; Pfister et al., 2014).
CDK12 proteins co-localize with hyperphosphorylated RNAPII to splicing speckles (Ko et al., 2001; Liang et al., 2015), suggesting that CDK12 may coordinate the splicing of pre-mRNAs (Emadi et al., 2020; Liang et al., 2020; Tien et al., 2017). In further support of its link to splicing are findings that CDK12 directly interacts with and phosphorylates many components of the spliceosome (Bartkowiak and Greenleaf, 2015; Krajewska et al., 2019; Liang et al., 2015) and components of the exon junction complex (EJC) (Eifler et al., 2015). Mutation of Ser2 to alanine in the CTD of RNAPII results in the inhibition of co-transcriptional splicing (Gu et al., 2013) and depletion of CDK12 results in alternative splicing patterns (Chen et al., 2006; Tien et al., 2017). Other protein complexes that CDK12 interacts with are Integrator, the RNA exosome, condensin, elongin, hnRNPs, 5′Cap proteins, 3′-cleavage complex, and the DNA repair proteins ERCC6, XPC, MRE11, and RAD50 (Bartkowiak and Greenleaf, 2015; Bartkowiak et al., 2019; Krajewska et al., 2019). CDK12 expression peaks in the early G1-phase of the cell cycle and then drops to its lowest level in the late S-phase (Chirackal Manavalan et al., 2019).
CDK12 has been found to be recurrently mutated in certain cancers such as ovarian (Joshi et al., 2014; Popova et al., 2016) and prostate cancer (Chou et al., 2020; Rescigno et al., 2021; Wu et al., 2018). Many of these tumors show genomic instability leading to hundreds of tandem DNA duplications (TD). Although the mechanism responsible for the formation of TDs is not fully understood, the fact that these duplications are enriched in regions of active genes (Wu et al., 2018) suggests that the loss of CDK12 functions regulating transcription elongation and splicing may be involved in the formation of TDs. DNA damage response (DDR) genes are typically long and contain an above-average number of IPAs, and consequently have been found to be down-regulated following the acute loss of CDK12 activity (Blazek et al., 2011; Dubbury et al., 2018; Ekumi et al., 2015; Joshi et al., 2014; Krajewska et al., 2019; Wang et al., 2020; Zhang et al., 2016). This downregulation of DDR genes can be exploited therapeutically using DNA-damaging chemotherapy or PARP1 inhibitors (Bajrami et al., 2014; Chila et al., 2016; Chou et al., 2020; Joshi et al., 2014; Paculova and Kohoutek, 2017). However, CDK12-defective tumors with a TD phenotype show no suppression of expression of DDR genes (Greenleaf, 2019; Popova et al., 2016; Wu et al., 2018). These tumors may instead qualify for immunotherapy because of the frequent formation of fusion genes giving rise to new neoantigens (Chou et al., 2020). Several small molecule inhibitors of CDK12 have been developed (Dieter et al., 2021; Gao et al., 2018; Zhang et al., 2016) and they appear to be especially effective against Ewing’s sarcoma and osteosarcoma (Chou et al., 2020), triple-negative breast cancer (Quereda et al., 2019), hepatocellular carcinoma (Wang et al., 2020), and a subset of colorectal cancers (Dieter et al., 2021). The compound THZ531 selectively inhibits CDK12 and CDK13 and leads to a significantly reduced output of RNA within 6h (Zhang et al., 2016). Recently, a selective CDK12 degrader BSJ-4-116 was developed with effects on gene expression very similar to THZ531 according to poly(A) RNA-seq analysis following an 8-h treatment (Jiang et al., 2021a). This inhibition has been ascribed to premature transcriptional termination at intronic polyadenylation sites (IPAs) and significant loss of expression of DDR genes and super enhancer-associated genes (Zhang et al., 2016). Furthermore, the compound NCT02, which acts as a molecular glue inducing ubiquitylation of CDK12, showed efficacy in in vivo studies of metastatic colorectal cancer (Dieter et al., 2021). Owing to its role in regulating DDR genes, CDK12 has been considered a potential therapeutic target in different cancers especially when combined with PARP inhibitors (Chou et al., 2020; Paculova and Kohoutek, 2017). Furthermore, the diminished levels of H3K36me3 in active genes suppressing HRR (Aymard et al., 2014; Greenleaf, 2019; Krajewska et al., 2019; Pfister et al., 2014) suggest that CDK12 inhibitors may act as sensitizers for radiation therapy and chemotherapy drugs invoking HRR.
In this study, we used Bru-seq and BruChase-seq (Paulsen et al., 2013, 2014) to investigate the transcriptional and post-transcriptional effects of CDK12 inhibition by the CDK12/CDK13 inhibitor THZ531, and by using an analog-sensitive HeLa-AS cell line treated with the adenine analog 1-NM-PP1 (Bartkowiak and Greenleaf, 2015; Bartkowiak et al., 2015). The results show a reduced transcription signal toward the 3′ ends of long genes and reduced transcriptional readthrough past transcription end sites (TESs). We identified 688 genes showing premature termination and these genes were predominantly associated with DNA damage and repair and cell cycle functions. Using BruChase-seq to assess the turnover of newly synthesized transcripts revealed a dramatic effect of CDK12 inhibition on the stabilization of RNA transcripts representing gene sets such as “TNFA-signaling”, “G2/M checkpoint”, “p53 pathway” and “hypoxia” and KEGG pathway gene sets such as “homologous recombination”, “base excision repair”, “nucleotide excision repair” and “mismatch repair”. In addition, the inhibition of CDK12 showed a slight, but significant, reduction of overall co-transcriptional splicing. Our results show that CDK12 inhibition causes defects in transcription elongation as previously reported but also an acute induction of transcriptional programs involving MYC and E2F as well as dramatic changes in transcript stability. Transcripts involved in DNA repair pathways were both prematurely terminated and de-stabilized by CDK12 inhibition, suggesting that transcript turnover may contribute to the DNA repair defect reported following CDK12 inhibition.
Results
Acute inhibition of CDK12 alters the nascent transcriptional profile
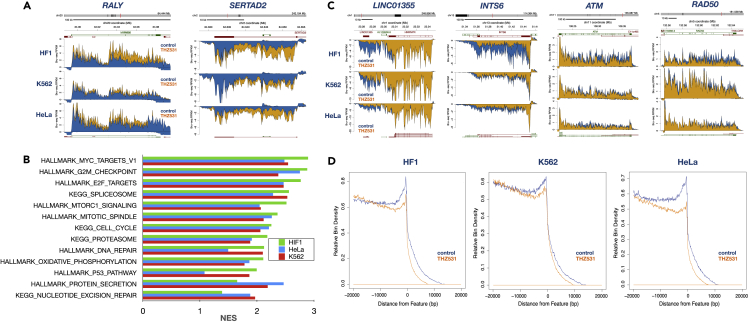
To explore the acute effects of CDK12 inhibition on transcription, we treated HF1, K562, and HeLa cells for 1 h with the CDK12/CDK13 inhibitor THZ531 and measured nascent RNA synthesis using Bru-seq. We observed several genes upregulated in response to THZ531 treatment across the three cell lines such as RALY and SERTAD2 (Figure 1A). RALY is an RNA-binding protein that regulates pre-mRNA splicing and miRNA metabolism and may have oncogenic properties (Song et al., 2020; Sun et al., 2021). SERTAD2 is a transcriptional co-activator of E2F-regulated genes and promotes diet-induced obesity (Liew et al., 2013). Performing gene set enrichment analysis (GSEA) on THZ531-treated cells, we observed very similar patterns of upregulated gene sets across the three cell lines (Figures 1B and S1). The top upregulated gene sets included “MYC targets”, “E2F targets”, “G2/M checkpoint” and “spliceosome”. Surprisingly, the GSEA analysis did not reveal any gene sets that were downregulated by THZ531 although there were indications of reduced transcription toward the 3′-ends of many genes such as Integrator complex subunit gene INTS6, and the DNA damage response genes ATM and RAD50 (Figure 1C). The non-coding RNA gene LINC01355 was suppressed following CDK12 inhibition, and this may be owing to a reduced transcriptional readthrough from the upstream gene HNRNPR (Figure 1C). Inspecting aggregate plots of transcription reads 20 kb upstream and downstream of the TESs, we observed that THZ531 suppressed transcription toward the 3′end of genes in all three cell lines (Figure 1D). Furthermore, THZ531 also suppresses transcription past the TES in all cell lines, implying that CDK12 activity plays a role in extending transcriptional readthrough or alternatively, reducing the turnover of the readthrough RNA.
Figure 1.
Effects of THZ531 on nascent transcription
(A) HF1, K562, and HeLa cells were treated with 400 nM THZ531 for 1 h at 37°C with the addition of bromouridine for the last 30 min and Bru-labeled RNA was then prepared for Bru-seq. Nascent RNA signal from Bru-seq libraries is represented as reads per kilobase per million (RPKM) and is shown for the control (blue) and THZ531 (yellow) for each cell line. The RPKM sign indicates the transcribed strand (positive values for plus strand signal; negative values for minus strand signal). Gene models are above the signal tracks and transcript isoform models are below (green for plus strand; red for minus strand). The nascent RNA reads were mapped to Hg38. Examples of genes showing upregulated transcription following THZ531 treatment in all three cell lines are RALY and SERTAD2.
(B) Gene set enrichment analysis (GSEA) of Bru-seq data showing gene sets upregulated across all three cell lines following a 1-h THZ531 treatment. Gene sets in the Hallmark and KEGG pathways are shown and the data is expressed as normalized enrichment scores (NES).
(C) Nascent transcription of genes, represented as in (A), showing reduced signal toward their 3′-ends following THZ531 treatment.
(D) Aggregated profile of gene transcription end sites (TESs) showing that THZ531 reduces transcription toward the 3′-ends and past the TES. For the aggregate graphs, 2637 genes >20kb in length and expressed at >0.5 RPKM in the HF1 control sample were used.
Specific inhibition of CDK12 using 1-NM-PP1 in HeLa-AS cells
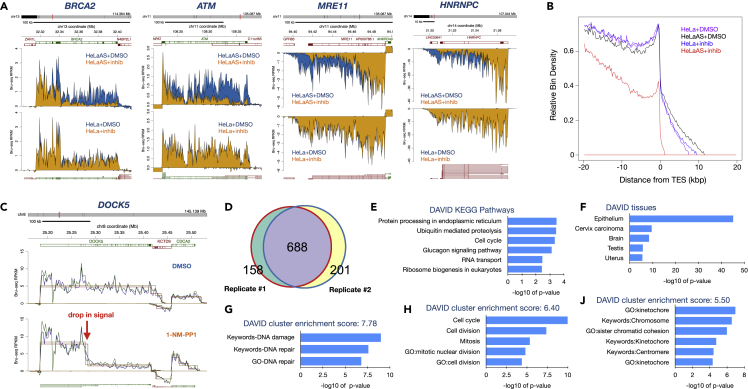
THZ531 inhibits both CDK12 and CDK13 and therefore our results above may reflect the inhibition of CDK12, CDK13, or both. To specifically target CDK12 in cells, we utilized HeLa-AS (analog-sensitive) cells in which CDK12 has been mutated in such a way as to retain normal activity but render it susceptible to inhibition by the bulky adenine analog 1-NM-PP1 (Bartkowiak and Greenleaf, 2015; Bartkowiak et al., 2015). Similarly to treatments with THZ531, HeLa-AS cells treated with 1-NM-PP1 showed reduced nascent RNA reads toward the 3′-end of many genes such as the DNA damage response genes BRCA2, ATM, and MRE11 (Figure 2A). Furthermore, many genes showed reduced transcription that appears to be owing to diminished transcriptional readthrough from upstream genes such as HNRNPC leading to loss of expression of downstream lncRNA LINC00641 (Figures 2A and S2). Assessing genome-wide nascent transcription, we found many significantly downregulated genes following CDK12 inhibition with fewer significantly upregulated genes (Figure S3A). Surprisingly, GSEA analyses did not show any gene sets significantly enriched for the downregulated genes while several gene sets were enriched for genes upregulated following CDK12 inhibition (Figure S3B). Although some of these upregulated gene sets were unique for 1-NM-PP1 treatment of HeLa-AS cells compared to gene sets induced by THZ531, such as “TNF-signaling via NFkB”, many of the gene sets were common, suggesting that it was the loss of CDK12 activity, and not CDK13 activity, that resulted in the upregulation of the genes driving these pathways. Examples of genes where the inhibition of CDK12 led to increased transcription are shown in Figure S3C. We next asked whether gene length may influence the degree to which CDK12 inactivation may affect transcription. Comparing the average size of significantly up-regulated genes (padj<0.01, n = 472) with the average gene size of down-regulated genes (padj<0.01, n = 1766) we observed a marked difference in that upregulated genes were much smaller (33 kb) than the downregulated genes (121 kb) supporting the hypothesis that CDK12 activity plays a role promoting the completion of transcription of long genes (Figure S3D). Finally, when we aggregated the data, we observed a similar reduction of transcription reads toward the 3′-end of genes as seen for THZ531 treated cells (Figure 1D) and suppression of transcription elongation past the TES in the CDK12 inactivated cells (Figures 2B and S2).
Figure 2.
Effects of the adenine analog 1-NM-PP1 on nascent transcription in HeLa-AS cells
(A) HeLa-AS cells were treated in two biological experiments with DMSO or 1-NM-PP1 for 30 min in the presence of bromouridine and Bru-labeled RNA was then prepared for Bru-seq. Inhibition of CDK12 resulted in the rapid reduction of transcription reads at the 3′-ends of the indicated genes (represented as in Figure 1A). No transcriptional effects of 1-NM-PP1 treatment were observed in parental cells.
(B) Aggregate view of gene transcription end sites (TESs) from Bru-seq data showing that the inhibition of CDK12 results in reduced transcription toward the 3′-end of genes and past the TES. For the aggregate graphs, 2619 genes >20kb in length and expressed at >0.5 RPKM in the HeLa + DMSO sample are represented.
(C) Contiguous regions of transcription at different expression levels (shown as rectangular overlays; one overlay per replicate) for the DOCK5 gene. A single, long segment overlays the gene body in the control (top), but is partitioned into two segments (high and low expression) after CDK12 inhibition, indicating the region containing a presumptive intergenic termination site.
(D) Genes with intragenic termination events identified by transcription segment partitioning. 688 genes were found to be common to both biological replicates.
(E–H) DAVID analyses using the 688 common gene list show enrichment for gene sets. E) KEGG pathways, F) tissues and cluster enrichment, G) DNA damage responses, H) cell cycle and J) kinetochore and chromosomes. Enrichment is expressed as -log10 p value.
To identify genes showing intragenic premature termination, a transcription segmentation algorithm was used to first identify de novo transcribed regions (Paulsen et al., 2014) and then to find adjacent segments that indicate a drop in expression within gene bodies. Using regions of consistent transcription (represented by a single contiguous segment) in the control condition as a reference, we searched for divided segments in the CDK12-inhibited condition with a high-expressing upstream segment and a low-expressing downstream segment. Applying this approach to the HeLa-AS control and 1-NM-PP1 treated cells, we identified 846 genes in replicate 1 and 889 in replicate 2, with a substantial overlap of 688 genes in common (Figures 2D and Table S1). One example of a gene showing intragenic termination is DOCK5 where following treatment with 1-NM-PP1, we observe two segments with different levels of transcription (Figure 2C). Performing DAVID (https://david.ncifcrf.gov/home.jsp, version 6.8) analyses using the 688 gene set, we found enrichment of genes in the KEGG pathways “protein processing in the ER”, “ubiquitin-mediated proteolysis” and “cell cycle” (Figure 2E). Furthermore, this gene set was found to be highly enriched for the tissue term “epithelium” (Figure 2F). Exploring DAVID cluster enrichment scores, we found that the top clusters involved “DNA damage” and “DNA repair” (Figure 2G) followed by “cell cycle” (Figure 2H) and mitosis-related gene sets such as “kinetochore”, “chromosome” and “chromatin cohesion” (Figure 2J).
Dramatic alterations in RNA turnover by CDK12 inhibition
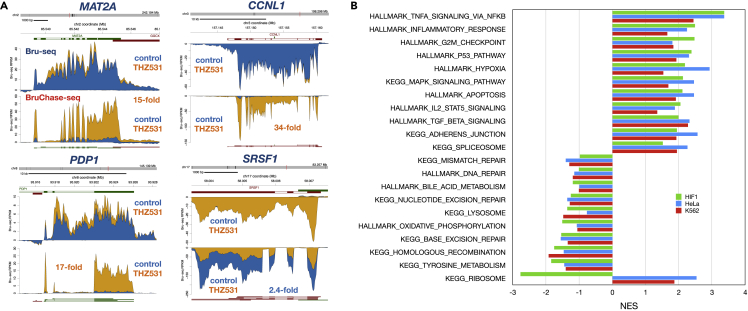
To investigate the effects of CDK12 inhibition on RNA turnover, we treated the three human cell lines with THZ531 for 1 h with Bru-labeling for the last 30 min followed by a 2-h (HeLa) or a 6-h chase (HF1 and K562) in the presence of THZ531. The reason for the shorter chase period for the HeLa cells is that we have found that post-transcriptional processing is very fast in these cells. The effects of THZ531 on RNA turnover were assessed by comparing the ratio of normalized gene counts obtained after the chase with the normalized gene counts without the chase for the treated and control conditions. Transcripts stabilized by THZ531 treatment include MAT2A (15-fold), which encodes a subunit of methionine adenosyltransferase (Halim et al., 2001) which can be stabilized by METLL16-induced splicing (Pendleton et al., 2017; Shima et al., 2017), CCNL1 (34-fold) encoding cyclin L1 that has been shown to interact with CDK12 (Chen et al., 2006) and has a role in pre-mRNA splicing (Dickinson et al., 2002) and PDP1 (17-fold), coding for pyruvate dehydrogenase phosphatase that may play a role in circadian rhythm regulation (Cyran et al., 2003) (Figure 3A). Among transcripts that were de-stabilized following THZ531 treatment was the transcript for the splicing factor SRSF1 (2.4-fold) which has been previously shown to be stabilized in a CDK12-dependent manner (Liang et al., 2015). (Figure 3A).
Figure 3.
Effects of THZ351 treatment on the turnover of RNAs
(A) Treatment with THZ531 stabilized several transcripts such as MAT2A, CCNL1, and PDP1 while de-stabilized transcripts such as SRSF1. HF1 cells were either mock-treated or treated for 1h with 400 nM THZ531 with labeling with bromouridine for the last 30 min. Cells were then chased in 20 mM uridine in the presence or absence of THZ531 for 6 h and prepared for BruChase-seq. Bru-seq and BruChase-seq signal is presented in Figure 1A.
(B) Enriched gene sets from BruChase-seq differential expression in THZ531-treated HF1, K562, and HeLa cells and presented as normalized expression scores (NES) from GSEA.
Performing GSEA on gene stability values from HF1 as well as HeLa and K562 cells showed that the Hallmark gene sets “TNFA signaling via NFkB”, “inflammatory response”, “G2/M checkpoint”, “p53 pathway” and “hypoxia” were stabilized in all cell lines following THZ531 treatment (Figure 3B). In contrast, the KEGG pathway gene sets “homologous recombination”, “base excision repair”, “nucleotide excision repair” and “mismatch repair” showed transcripts de-stabilized in all cell lines. Thus, rapid turnover of DNA repair gene transcripts contributes to the loss of expression of DNA damage response genes. Interestingly, transcripts in the KEGG pathway “ribosome”, were highly de-stabilized by THZ531 in the normal fibroblasts while they were stabilized in both K562 and HeLa cells.
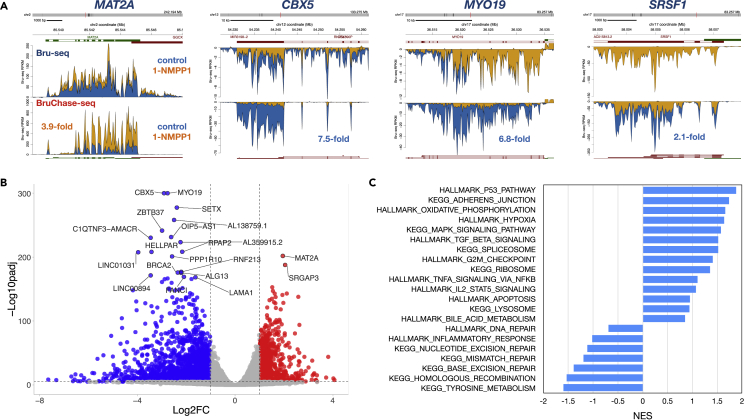
We next explored whether the inhibition of CDK12 was responsible for the dramatic changes in RNA stability observed with the THZ531 inhibitor by using the 1-NM-PP1 compound on the HeLa-AS cells. Again, we observed transcripts that became stabilized such as MAT2A (3.9-fold) and transcripts that became de-stabilized such as CBX5 (7.5-fold) which codes for the heterochromatin protein HP1 (Wreggett et al., 1994), MY O 19 (6.8-fold), which encodes a myosin associated with mitochondria (Quintero et al., 2009) and the splicing factor encoding transcript SRSF1 (2- to 1-fold)which is known to be stabilized by CDK12 (Liang et al., 2015) (Figure 4A). Plotting the transcripts with significantly altered stability (padj<0.01) and with a fold-change of at least 2, we observed 1130 genes that were stabilized and 2467 genes that were de-stabilized within 2h following CDK12 inactivation (Figure 4B and Table S2). Performing GSEA on the RNA stability data obtained from HeLa-AS cells following 1-NM-PP1 treatment and a 2-h chase, we detected many gene sets enriched in common with what we detected for THZ531 treatment (Figure 4C). The gene sets with the most highly stabilized transcripts were “p53 pathway”, “adherens junctions”, “oxidative phosphorylation” and “hypoxia”. De-stabilized transcripts were found in “tyrosine metabolism”, “homologous recombination”, “base excision repair”, “mismatch repair” and “nucleotide excision repair”.
Figure 4.
Effect of 1-NM-PP1 treatment of HeLa-AS cells on RNA turnover
(A) HeLa-AS cells were mock-treated with DMSO or 1-NM-PP1 for 30 min in the presence of bromouridine. The cells were then chased in uridine in the presence of either DMSO or 1-NM-PP1 for 2 h before preparing the Bru-RNA for BruChase-seq (n = 3). The top panels are displaying Bru-seq reads and the bottom panels are displaying BruChase-seq reads (2-h chase). For a transcript to be considered stabilized by 1-NM-PP1 treatment, the ratio of the treated sample (yellow) to control (blue) is higher in the BruChase-seq data than in Bru-seq data (e.g. MAT2A)and lower for de-stabilized transcripts (e.g. CBX5, MY O 19 and SRSF1).
(B) Volcano plot showing transcripts stabilized (red) and transcripts de-stabilized (blue) after CDK12 inhibition (adjusted p value < 0.01; log2 fold-change > 1 or < −1).
(C) Enrichment of Hallmark and KEGG pathway gene sets using BruChase-seq stability following CDK12 inhibition shown as normalized enrichment scores (NES) from GSEA.
Inhibition of CDK12 results in reduced co-transcriptional splicing
The splicing of introns occurs co-transcriptionally as the pre-mRNA is synthesized by RNA polymerase II (Ameur et al., 2011; Bauren and Wieslander, 1994; Beyer and Osheim, 1988; Bhatt et al., 2012; Darnell, 2013; Djebali et al., 2012; Naftelberg et al., 2015; Oesterreich et al., 2016; Tilgner et al., 2012). We recently used nascent Bru-seq data to estimate co-transcriptional splicing and showed that the splicing efficiencies differ within genes and between cell types (Bedi et al., 2021). Here we assessed whether the inhibition of CDK12/13 with THZ531 would affect the global splicing index (SI) of introns in the three human cell lines HF1, K562, and HeLa. Here we compared the distributions of SI values from DMSO-treated cells and THZ531-treated cells comparing splicing junction with sufficient reads to make the cut-off as previously described (Bedi et al., 2021). It was found using the Wilcoxon test that THZ531 treatment slightly, but significantly, reduced co-transcriptional splicing (Figure 5A). We also observed a similar shift in SI values in HeLa-AS cells treated with 1-NM-PP1 compared with DMSO-treated control cells (Figure 5B). We next assessed the SI values for these cells after a chase period (6 h for HF1 and K562; 2 h for HeLa) and found that most introns were fully spliced and that there was no difference globally in SI values between THZ531-treated cells and DMSO-treated control cells (Figure 5C). We found a slight, but significant, difference in the 1-NM-PP1-treated HeLa-AS cells compared to DMSO-treated controls (Figure 5D). Taken together, the inhibition of CDK12 with THZ531 or 1-NM-PP1 showed slight, but significant, inhibitory effects on co-transcriptional splicing.
Figure 6.
Model of the roles of CDK12 in transcription and post-transcriptional processes
Our results implement CDK12 playing roles in transcription both in suppressing premature termination at cryptic polyadenylation sites (PAS) as well as promoting transcriptional readthrough past transcription end sites (TES). CDK12 also was found to promote more efficient general splicing of transcripts. Finally, our study shows that CDK12 is involved in the regulation of transcript stability where transcripts encoding DNA repair enzymes are stabilized by CDK12 while transcripts encoding proteins in the p53, G2/M checkpoint, hypoxia pathways as well as the spliceosome are de-stabilized.
Figure 5.
Effect of CDK12 inhibition on co-transcriptional splicing
(A) THZ531 suppresses global co-transcriptional splicing in HF1, K562, and HeLa cells. Splicing index (SI) values were calculated from Bru-seq data using intron junctions with sufficient reads to make the cut-off as previously described (Bedi et al., 2021). The number of introns qualifying for the analyses is shown below the violin plots. The median SI values are shown in the white rectangles.
(B) Inhibition of CDK12 with 1NMPP1 in HeLa-AS cells shows slight but significantly reduced co-transcriptional splicing.
(C) SI-values for HF1, K562, and HeLa cells from BruChase-seq data show no significant difference between THZ531-treated and control cells for any of the cell lines.
(D) SI-values for 1NMPP1-treated HeLa-AS cells show a slight, but significant difference following a 2-h chase. ∗∗∗ = p value<10−15, ∗∗ = p value<0.0007.
Discussion
CDK12 plays important roles in transcription and RNA processing. By phosphorylating the CTD during elongation, CDK12 promotes transcription by passing through cryptic intronic polyadenylation sites (Bayles et al., 2019; Chirackal Manavalan et al., 2019; Tellier et al., 2020; Zhang et al., 2016) as well as being involved in 3′-end complex formation and termination (Davidson et al., 2014; Eifler et al., 2015). Furthermore, phosphorylating and associating with splicing factors it is thought to promote the splicing process (Bartkowiak and Greenleaf, 2015; Eifler et al., 2015; Emadi et al., 2020; Ko et al., 2001; Krajewska et al., 2019; Liang et al., 2015, 2020; Tien et al., 2017). Inhibition of CDK12 leads to loss of expression of DNA damage response genes sensitizing cells to DNA-damaging agents and PARP inhibitors (Blazek et al., 2011; Dubbury et al., 2018; Ekumi et al., 2015; Joshi et al., 2014; Krajewska et al., 2019; Wang et al., 2020; Zhang et al., 2016). Thus, inhibition of CDK12 in combination with DNA damaging agents or PARP inhibitors has attracted significant clinical interest with several small molecule inhibitors or degraders being developed (Gao et al., 2018; Jiang et al., 2021b; Zhang et al., 2016). To fully capitalize on CDK12 as a cancer therapeutic target, a more comprehensive understanding of its role in transcription and RNA processing is needed.
In this study, we inhibited CDK12/13 using THZ531 and used the adenine analog 1-NM-PP1 to specifically inhibit CDK12 in analog-sensitive HeLa-AS cells and assessed transcription and RNA processing using the nascent RNA sequencing methods Bru-seq and BruChase-seq. Our results confirm previous studies that the acute inhibition of CDK12 results in decreased transcription reads toward the 3′-ends of genes likely owing to premature transcription termination by the utilization of intronic poly(A) sites (Zhang et al., 2016) (Figures 1 and 2). We also observed reduced transcription signals past the transcriptional termination following inhibition, suggesting that CDK12 plays some role in extending transcription past gene termination sites. The purpose of such transcriptional readthrough is not clear but may lead to the expression of downstream genes such as miRNA or lncRNA genes. Alternatively, CDK12 inhibition may enhance the degradation of the RNA generated by RNA polymerase II as it traverses the TES. When comparing the average gene size of the top up-regulated genes versus the top down-regulated genes it shows that inhibited genes are on average substantially longer than induced genes (Figure S3D). Although no gene enrichments were found for acutely down-regulated genes, we observed enrichment of several gene sets acutely upregulated transcriptionally across all three cell lines by THZ531 and by 1-NM-PP1 in HeLa-AS cells (Figures 1, S1, and S2). These gene sets included “MYC targets”, “G2/M checkpoint”, “E2F targets”, “spliceosome” and surprisingly “DNA repair”. The mechanism by which CDK12 inhibition leads to the upregulation of these pathways is not clear.
Using BruChase-seq we found that inhibition of CDK12 had a dramatic effect on the stability of several transcripts (Figures 3 and 4). Gene sets such as “p53 pathway”, “G2/M checkpoint”, “hypoxia” and “spliceosome” showed transcript stabilization. Significantly, DNA damage repair pathway gene sets such as “homologous recombination”, “base excision repair”, “nucleotide excision repair”, “DNA repair” and “mismatch repair” showed transcript de-stabilization upon CDK12 inhibition. Thus, the well-documented downregulation of DNA repair pathways after CDK12 inhibition may not only be driven by premature transcription termination but also by increased degradation of transcripts of DNA repair genes. One transcript that was highly stabilized by CDK12 inhibition was MAT2A (Figures 3A and 4A). The splicing of the last intron of the MAT2A transcript is regulated by the cellular levels of S-adenosyl methionine (SAM) which is the product of the MAT2A SAM synthase. When SAM levels are high, the last intron is not spliced resulting in nuclear retention of the transcript and rapid degradation (Pendleton et al., 2017; Shima et al., 2017). Furthermore, when SAM levels are low, the N6-adenosine transferase METLL16 promotes efficient splicing of the last intron and the transcript is exported and translated. However, we do not observe a difference in the splicing pattern of MAT2A following CDK12 inhibition, suggesting that the mechanism of MAT2A transcript stabilization following CDK12 inhibition does not involve altered splicing.
While CDK12 has been shown to associate with and phosphorylate splicing factors (Bartkowiak and Greenleaf, 2015; Eifler et al., 2015; Emadi et al., 2020; Ko et al., 2001; Krajewska et al., 2019; Liang et al., 2015, 2020; Tien et al., 2017), the role of CDK12 in co-transcriptional splicing has not been previously assessed. We recently reported that co-transcriptional splicing can be determined from nascent RNA Bru-seq data and showed that the efficiency of splicing is regulated in both a gene and cell type-dependent manner (Bedi et al., 2021). Here we show that the inhibition of CDK12 caused a slight, but significant lowering of co-transcriptional splicing in HF1, K562, and HeLa cells treated with THZ531 (Figure 5A) as well as in HeLa-AS cells treated with 1-NM-PP1 (Figure 5B). When assessing SI values following a 2 or 6-h chase, the differences between control and treated cells were erased, suggesting that transcripts not fully spliced co-transcriptionally are either spliced later or purged by nuclear surveillance ribonucleases. As CDK12 is known to localize to nuclear speckles that also host many splicing factors (Ko et al., 2001; Liang et al., 2015) it is possible that the CDK12 phosphorylation of RNAPII CTD promotes the localization of the transcription machinery to speckles where efficient co-transcriptional splicing occurs.
Our study is the first to show that CDK12 plays important roles in co-transcriptional splicing, transcript stability, and promoting transcriptional elongation within genes and transcriptional readthrough past transcription end sites (Figure 6). One other important finding of our study is that increased transcript degradation may contribute to the well-known suppression of DNA repair gene expression following CDK12 inhibition. The mechanisms by which CDK12 regulates co-transcriptional splicing and transcript stability need further investigations and may lead to the development of better cancer therapeutic agents targeting CDK12.
Limitation of the study
This work was performed on cell lines in culture using pharmacological inhibitors and the results may, therefore, not fully represent the loss of function of CDK12 in vivo and off-target effects of these drugs are likely. Furthermore, the drug THZ531 is known to inhibit both CDK12 and CDK13 making it impossible to distinguish whether either or both kinases are affected by the drug. However, the results obtained using 1-NM-PP1 on the analog-sensitive HeLa-AS cell line should be CDK12-specific.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| anti-BrdU | BD Biosciences | 555627 (RRID:AB_10015222) |
| Chemicals, peptides, and recombinant proteins | ||
| MEM | Invitrogen | 11095-080 |
| IMDM | Invitrogen | 12440-053 |
| DMEM | Invitrogen | 11965-092 |
| FBS | Invitrogen | 10437-028 |
| Penicillin/Streptomycin | Invitrogen | 15140-122 |
| THZ531 | MedChem Express | HY-103618 |
| 1-NM-PP1 | Axon MedChem | 1892 |
| Bromouridine | MilliporeSigma | 850187 |
| Uridine | MilliporeSigma | U3750 |
| Trizol | Invitrogen | 15596-018 |
| Bru-seq RNA spike-ins | This manuscript | N/A |
| Critical commercial assays | ||
| Tru-seq Kit | Illumina | RS-122-2001 |
| Deposited data | ||
| Raw and processed data | This manuscript | GEO: GSE191222 |
| Experimental models: Cell lines | ||
| Human: HF1; hTERT immortalized human diploid foreskin fibroblasts | Dr. Mary Davis, Department of Radiation Oncology, University of Michigan, Ann Arbor | N/A |
| Human: K562; CML-derived lymphoblast | ATCC | CCL-243 (RRID CVCL_0004) |
| Human: HeLa; epitheloid cervix carcinoma | Millipore-SIGMA | 93021013 |
| Human: HeLaAS; HeLa stably expressing analog-sensitive (AS) CDK12 | Bartkowiak and Greenleaf (2015); Bartkowiak et al. (2015) | N/A |
| Software and algorithms | ||
| Casava v1.8.2 | Illumina | N/A |
| BB Tools v38.46 | Bushnell B. | http://sourceforge.net/projects/bbmap/ |
| Bowtie2 v2.3.3 | Langmead and Salzberg (2012) | http://bowtie-bio.sourceforge.net/bowtie2/index.shtml |
| STAR v2.5.3a | Dobin et al. (2013) | https://github.com/alexdobin/STAR/releases |
| DESeq2 v1.18.1 (R library) | Love et al. (2014) | https://bioconductor.org/packages/release/bioc/html/DESeq2.html |
| bedtools 2 v2.28.0 | Quinlan and Hall, 2010 | https://bedtools.readthedocs.io/en/latest/ |
| R v3.4.3 | R Core Team, 2020 | https://www.R-project.org/ |
| Other | ||
| Bru-seq RNA spike-ins (reference) | ENCODE Project | https://www.encodeproject.org/files/ENCFF267BFZ |
Resource availability
Lead contact
Further information and requests for resources should be directed to and will be fulfilled by the Lead Contact, Mats Ljungman (ljungman@umich.edu).
Materials availability
This study did not generate new unique reagents.
Experimental model and subject details
HF1 fibroblasts (male) are from primary source (Dr. Mary Davis, University of Michigan) and thus authentic. K562 CML lymphoblasts (ATCC: CCL-242) and HeLa cervix carcinoma (Millipore-Sigma: 93021013) are authenticated via their sources. HeLa with analog sensitive CDK12 (HeLaAS) is not authenticated.
Method details
Cell cultures and reagents
hTERT immortalized human diploid foreskin fibroblasts (HF1) (a gift from Dr. Mary Davis, Department of Radiation Oncology, University of Michigan, Ann Arbor) were grown in Minimal Essential Medium with 10% FBS (Invitrogen) (Paulsen et al., 2013, 2014) K562 cells (ATCC) were grown in suspension in IMDM with 10% FBS. HeLa cells (Millipore-SIGMA) DMEM, 10% FBS, 100 U/mL penicillin and 100 U/mL streptomycin. HeLa-AS cells were grown in DMEM as previously described (Bartkowiak and Greenleaf, 2015; Bartkowiak et al., 2015). THZ531 (MedChemExpress) and 1-NM-PP1 (Axon Medchem) were dissolved in DMSO.
Bru-seq and BruChase-seq
For the Bru-seq experiments, cells were either mock-treated with DMSO in conditioned media or treated with 400 nM THZ531 (HF1, K562 and HeLa) or 10 μM 1-NM-PP1 (HeLa-AS) for 60 min where bromouridine (Bru) (MilliporeSigma) was included during the last 30 min at a final concentration of 2 mM to label nascent RNA. Following Bru-labeling, cells were lysed directly in Trizol followed by isolation of total RNA. For BruChase-seq experiments, cells were first labeled for 30 min with 2 mM BrU in conditioned media after which the cells were washed in PBS and conditioned media containing 20 mM uridine was added back together with either 400 nM THZ531, 10 μM 1-NM-PP1 or no drug for 2 hours in HeLa and HeLa-AS cells, or 6 hours in HF1 and K562 cells.
Following the uridine chase, cells were lysed directly in TRIzol followed by isolation of total RNA. For experiments involving HF1, K562, and HeLa treated with THZ531 (including controls), the isolated total RNA was further treated with DNAse (TURBO DNA-free Kit; Invitrogen). For both Bru-seq and BruChase-seq, Bru-labeled RNA was immunocaptured using anti-BrdU antibodies (BD Biosciences). Size selection for all libraries was via agarose gel excision.
Strand-specific libraries for the HeLa-AS experiments were prepared from a modified protocol using the TruSeq Kit (Illumina) as previously described (Paulsen et al., 2013, 2014). This protocol used single-index barcoded ligation adapters and universal primers for PCR. These libraries were sequenced on a HiSeq 2000 (Illumina), yielding 50 bp, single-ended reads. For the experiments in HF1, K562, and HeLa cells involving THZ531 treatment, a spike-in cocktail of Bru-labeled and unlabeled RNA (described below) was mixed in with total RNA prior to Bru-RNA IP. Ribosomal RNA was reduced via QiaSeq FastSelect (Qiagen) prior to first strand synthesis. These strand-specific libraries were made using a universal ligation adapter and dual-index, barcoded primers for PCR. Sequencing was performed on a NovaSeq 6000 (Illumina), yielding 150 bp, paired-ended reads.
The RNA spike-in cocktail was designed in-house for the Bru-seq methods and consists of in vitro transcribed RNAs from the A. thaliana genes AGP23 (Bru-labeled), OBF5 (Bru-labeled), PDF1 (unlabeled), and AP2 (unlabeled), total RNA extracted from E. coli K-12 MG1655 strain (unlabeled), and total RNA extracted from D. melanogaster S3 cells (Bru-labeled). The spike-in cocktail was optimized such that final quantities as percentage by weight of total RNA were as follows: 0.001% each labeled IVT RNA, 0.1% each unlabeled IVT RNA, 4% unlabeled E. coli total RNA, and 4% Bru-labeled D. melanogaster S3 total RNA. For the 6-hour BruChase-seq assay, the spike-ins were added at 1/4 of these amounts. A reference sequence for the spike-ins is available on the ENCODE portal (https://www.encodeproject.org/files/ENCFF267BFZ/; FASTA format).
Bru-seq and BruChase-seq data analysis
Reads were pre-mapped to the ribosomal RNA (rRNA) repeating unit (GenBank U13369.1) and the mitochondrial and EBV genomes (from the hg38 analysis set) using Bowtie2 (2.3.3) (Langmead and Salzberg, 2012). Unaligned reads were subsequently mapped strand-specifically to human genome build hg38/GRCh38 using STAR (v 2.5.3a) (Dobin et al., 2013) and a STAR index created from GENCODE annotation version 27 (Frankish et al., 2019). Gene expression was quantified as previously described (Paulsen et al., 2013, 2014) and differential gene expression was performed using DESeq2 v1.18.1 (Love et al., 2014). DESeq data set objects were then normalized using the rlog function (parameter blind=TRUE) and rlog fold change calculated by subtracting rlog of the control group from rlog of the treated group. This rlog fold change was used to order genes for use in pre-ranked GSEA (v4.1.0) to identify enriched gene sets from mSigDB v7.4 Hallmark and KEGG collections (Broad Institute, Cambridge, MA).
In the analog-sensitive CDK12 experiment, genes with CDK12-dependent premature termination defects were identified using Bru-seq transcription segmentation. Segments were identified in individual samples by a hidden Markov model algorithm as previously described (Paulsen et al., 2014). Intronic regions from the AS-CDK12 control Bru-seq samples that overlapped only 1 transcription segment were defined as introns of constant expression. Of these introns, those in the 1-NM-PP1 treated sample were selected if they had >1 overlapping segments and the HMM score decreased in the downstream direction. Expressed genes (at least 1 RPKM) with such introns were selected from the two replicates to find the reproducible set of events.
Data are available at the Gene Expression Omnibus (GEO; accession GSE191222).
Quantification and statistical analysis
Bru-seq and BruChase-seq data analysis used software with statistical parameters described in the methods or in the software documentation.
Acknowledgments
We thank the personnel at the University of Michigan Advanced Genomics Core for professional technical assistance and Manhong Dai and Fan Meng for the administration and maintenance of the University of Michigan Molecular and Behavioral Neuroscience Institute (MBNI) computing cluster. This work was supported by grants from NCI, United States (R01 CA213214), NHGRI, United States (UM1 HG009382) and the Rogel Cancer Center Support Grant, United States (P30 CA046592).
Author contributions
B.M., B.B., A.G., and M.L. designed the experiments. B.B., H.B., and M.T.P. performed cell experiments and H.B. and M.T.P. prepared sequencing libraries. B.M., K.B., I.V.N., and M.L. performed data analyses. B.M. and M.L. wrote the article and prepared figures.
Declaration of interests
The authors declare no competing interest.
Published: September 16, 2022
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2022.105030.
Supplemental information
Data and code availability
Raw and processed sequencing data for Bru-seq and BruChase-seq have been deposited at the Gene Expression Omnibus (GEO), accession GSE191222.
This manuscript does not report original code.
Any additional information required to reanalyze the data reported herein is available upon request of the lead contact.
References
- Ameur A., Zaghlool A., Halvardson J., Wetterbom A., Gyllensten U., Cavelier L., Feuk L. Total RNA sequencing reveals nascent transcription and widespread co-transcriptional splicing in the human brain. Nat. Struct. Mol. Biol. 2011;18:1435–1440. doi: 10.1038/nsmb.2143. [DOI] [PubMed] [Google Scholar]
- Aymard F., Bugler B., Schmidt C.K., Guillou E., Caron P., Briois S., Iacovoni J.S., Daburon V., Miller K.M., Jackson S.P., Legube G. Transcriptionally active chromatin recruits homologous recombination at DNA double-strand breaks. Nat. Struct. Mol. Biol. 2014;21:366–374. doi: 10.1038/nsmb.2796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bajrami I., Frankum J.R., Konde A., Miller R.E., Rehman F.L., Brough R., Campbell J., Sims D., Rafiq R., Hooper S., et al. Genome-wide profiling of genetic synthetic lethality identifies CDK12 as a novel determinant of PARP1/2 inhibitor sensitivity. Cancer Res. 2014;74:287–297. doi: 10.1158/0008-5472.CAN-13-2541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartkowiak B., Greenleaf A.L. Expression, purification, and identification of associated proteins of the full-length hCDK12/CyclinK complex. J. Biol. Chem. 2015;290:1786–1795. doi: 10.1074/jbc.M114.612226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartkowiak B., Liu P., Phatnani H.P., Fuda N.J., Cooper J.J., Price D.H., Adelman K., Lis J.T., Greenleaf A.L. CDK12 is a transcription elongation-associated CTD kinase, the metazoan ortholog of yeast Ctk1. Genes Dev. 2010;24:2303–2316. doi: 10.1101/gad.1968210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartkowiak B., Yan C., Greenleaf A.L. Engineering an analog-sensitive CDK12 cell line using CRISPR/Cas. Biochim. Biophys. Acta. 2015;1849:1179–1187. doi: 10.1016/j.bbagrm.2015.07.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartkowiak B., Yan C.M., Soderblom E.J., Greenleaf A.L. CDK12 activity-dependent phosphorylation events in human cells. Biomolecules. 2019;9:6344–E714. doi: 10.3390/biom9100634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baurén G., Wieslander L. Splicing of Balbiani ring 1 gene pre-mRNA occurs simultaneously with transcription. Cell. 1994;76:183–192. doi: 10.1016/0092-8674(94)90182-1. [DOI] [PubMed] [Google Scholar]
- Bayles I., Krajewska M., Pontius W.D., Saiakhova A., Morrow J.J., Bartels C., Lu J., Faber Z.J., Fedorov Y., Hong E.S., et al. Ex vivo screen identifies CDK12 as a metastatic vulnerability in osteosarcoma. J. Clin. Invest. 2019;129:4377–4392. doi: 10.1172/JCI127718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bedi K., Magnuson B., Narayanan I.V., Paulsen M.T., Wilson T.E., Ljungman M. Co-transcriptional splicing efficiencies differ within genes and between cell types. RNA. 2021;27:829–840. doi: 10.1261/rna.078662.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beyer A.L., Osheim Y.N. Splice site selection, rate of splicing, and alternative splicing on nascent transcripts. Genes Dev. 1988;2:754–765. doi: 10.1101/gad.2.6.754. [DOI] [PubMed] [Google Scholar]
- Bhatt D.M., Pandya-Jones A., Tong A.J., Barozzi I., Lissner M.M., Natoli G., Black D.L., Smale S.T. Transcript dynamics of proinflammatory genes revealed by sequence analysis of subcellular RNA fractions. Cell. 2012;150:279–290. doi: 10.1016/j.cell.2012.05.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blazek D., Kohoutek J., Bartholomeeusen K., Johansen E., Hulinkova P., Luo Z., Cimermancic P., Ule J., Peterlin B.M. The Cyclin K/Cdk12 complex maintains genomic stability via regulation of expression of DNA damage response genes. Genes Dev. 2011;25:2158–2172. doi: 10.1101/gad.16962311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bösken C.A., Farnung L., Hintermair C., Merzel Schachter M., Vogel-Bachmayr K., Blazek D., Anand K., Fisher R.P., Eick D., Geyer M. The structure and substrate specificity of human Cdk12/Cyclin K. Nat. Commun. 2014;5:3505. doi: 10.1038/ncomms4505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowman E.A., Bowman C.R., Ahn J.H., Kelly W.G. Phosphorylation of RNA polymerase II is independent of P-TEFb in the C. elegans germline. Development. 2013;140:3703–3713. doi: 10.1242/dev.095778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen H.H., Wang Y.C., Fann M.J. Identification and characterization of the CDK12/cyclin L1 complex involved in alternative splicing regulation. Mol. Cell Biol. 2006;26:2736–2745. doi: 10.1128/MCB.26.7.2736-2745.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chilà R., Guffanti F., Damia G. Role and therapeutic potential of CDK12 in human cancers. Cancer Treat Rev. 2016;50:83–88. doi: 10.1016/j.ctrv.2016.09.003. [DOI] [PubMed] [Google Scholar]
- Chirackal Manavalan A.P., Pilarova K., Kluge M., Bartholomeeusen K., Rajecky M., Oppelt J., Khirsariya P., Paruch K., Krejci L., Friedel C.C., Blazek D. CDK12 controls G1/S progression by regulating RNAPII processivity at core DNA replication genes. EMBO Rep. 2019;20:e47592. doi: 10.15252/embr.201847592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chou J., Quigley D.A., Robinson T.M., Feng F.Y., Ashworth A. Transcription-associated cyclin-dependent kinases as targets and biomarkers for cancer therapy. Cancer Discov. 2020;10:351–370. doi: 10.1158/2159-8290.CD-19-0528. [DOI] [PubMed] [Google Scholar]
- Cyran S.A., Buchsbaum A.M., Reddy K.L., Lin M.C., Glossop N.R.J., Hardin P.E., Young M.W., Storti R.V., Blau J. vrille, Pdp1, and dClock form a second feedback loop in the Drosophila circadian clock. Cell. 2003;112:329–341. doi: 10.1016/s0092-8674(03)00074-6. [DOI] [PubMed] [Google Scholar]
- Darnell J.E., Jr. Reflections on the history of pre-mRNA processing and highlights of current knowledge: a unified picture. RNA. 2013;19:443–460. doi: 10.1261/rna.038596.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davidson L., Muniz L., West S. 3' end formation of pre-mRNA and phosphorylation of Ser2 on the RNA polymerase II CTD are reciprocally coupled in human cells. Genes Dev. 2014;28:342–356. doi: 10.1101/gad.231274.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickinson L.A., Edgar A.J., Ehley J., Gottesfeld J.M. Cyclin L is an RS domain protein involved in pre-mRNA splicing. J. Biol. Chem. 2002;277:25465–25473. doi: 10.1074/jbc.M202266200. [DOI] [PubMed] [Google Scholar]
- Dieter S.M., Siegl C., Codó P.L., Huerta M., Ostermann-Parucha A.L., Schulz E., Zowada M.K., Martin S., Laaber K., Nowrouzi A., et al. Degradation of CCNK/CDK12 is a druggable vulnerability of colorectal cancer. Cell Rep. 2021;36:109394. doi: 10.1016/j.celrep.2021.109394. [DOI] [PubMed] [Google Scholar]
- Djebali S., Davis C.A., Merkel A., Dobin A., Lassmann T., Mortazavi A., Tanzer A., Lagarde J., Lin W., Schlesinger F., et al. Landscape of transcription in human cells. Nature. 2012;489:101–108. doi: 10.1038/nature11233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dobin A., Davis C.A., Schlesinger F., Drenkow J., Zaleski C., Jha S., Batut P., Chaisson M., Gingeras T.R. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21. doi: 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubbury S.J., Boutz P.L., Sharp P.A. CDK12 regulates DNA repair genes by suppressing intronic polyadenylation. Nature. 2018;564:141–145. doi: 10.1038/s41586-018-0758-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eifler T.T., Shao W., Bartholomeeusen K., Fujinaga K., Jäger S., Johnson J.R., Luo Z., Krogan N.J., Peterlin B.M. Cyclin-dependent kinase 12 increases 3' end processing of growth factor-induced c-FOS transcripts. Mol. Cell Biol. 2015;35:468–478. doi: 10.1128/MCB.01157-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ekumi K.M., Paculova H., Lenasi T., Pospichalova V., Bösken C.A., Rybarikova J., Bryja V., Geyer M., Blazek D., Barboric M. Ovarian carcinoma CDK12 mutations misregulate expression of DNA repair genes via deficient formation and function of the Cdk12/CycK complex. Nucleic Acids Res. 2015;43:2575–2589. doi: 10.1093/nar/gkv101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emadi F., Teo T., Rahaman M.H., Wang S. CDK12: a potential therapeutic target in cancer. Drug Discov. Today. 2020;25:2257–2267. doi: 10.1016/j.drudis.2020.09.035. [DOI] [PubMed] [Google Scholar]
- Frankish A., Diekhans M., Ferreira A.M., Johnson R., Jungreis I., Loveland J., Mudge J.M., Sisu C., Wright J., Armstrong J., et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 2019;47:D766–D773. doi: 10.1093/nar/gky955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao Y., Zhang T., Terai H., Ficarro S.B., Kwiatkowski N., Hao M.F., Sharma B., Christensen C.L., Chipumuro E., Wong K.K., et al. Overcoming resistance to the THZ series of covalent transcriptional CDK inhibitors. Cell Chem. Biol. 2018;25:135–142.e5. doi: 10.1016/j.chembiol.2017.11.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glover-Cutter K., Larochelle S., Erickson B., Zhang C., Shokat K., Fisher R.P., Bentley D.L. TFIIH-associated Cdk7 kinase functions in phosphorylation of C-terminal domain Ser7 residues, promoter-proximal pausing, and termination by RNA polymerase II. Mol. Cell Biol. 2009;29:5455–5464. doi: 10.1128/MCB.00637-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greenleaf A.L. Human CDK12 and CDK13, multi-tasking CTD kinases for the new millenium. Transcription. 2019;10:91–110. doi: 10.1080/21541264.2018.1535211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gu B., Eick D., Bensaude O. CTD serine-2 plays a critical role in splicing and termination factor recruitment to RNA polymerase II in vivo. Nucleic Acids Res. 2013;41:1591–1603. doi: 10.1093/nar/gks1327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halim A.B., LeGros L., Chamberlin M.E., Geller A., Kotb M. Regulation of the human MAT2A gene encoding the catalytic alpha 2 subunit of methionine adenosyltransferase, MAT II: gene organization, promoter characterization, and identification of a site in the proximal promoter that is essential for its activity. J. Biol. Chem. 2001;276:9784–9791. doi: 10.1074/jbc.M002347200. [DOI] [PubMed] [Google Scholar]
- Jiang B., Gao Y., Che J., Lu W., Kaltheuner I.H., Dries R., Kalocsay M., Berberich M.J., Jiang J., You I., et al. Discovery and resistance mechanism of a selective CDK12 degrader. Nat. Chem. Biol. 2021;17:675–683. doi: 10.1038/s41589-021-00765-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang B., Jiang J., Kaltheuner I.H., Iniguez A.B., Anand K., Ferguson F.M., Ficarro S.B., Seong B.K.A., Greifenberg A.K., Dust S., et al. Structure-activity relationship study of THZ531 derivatives enables the discovery of BSJ-01-175 as a dual CDK12/13 covalent inhibitor with efficacy in Ewing sarcoma. Eur. J. Med. Chem. 2021;221:113481. doi: 10.1016/j.ejmech.2021.113481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joshi P.M., Sutor S.L., Huntoon C.J., Karnitz L.M. Ovarian cancer-associated mutations disable catalytic activity of CDK12, a kinase that promotes homologous recombination repair and resistance to cisplatin and poly(ADP-ribose) polymerase inhibitors. J. Biol. Chem. 2014;289:9247–9253. doi: 10.1074/jbc.M114.551143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ko T.K., Kelly E., Pines J. CrkRS: a novel conserved Cdc2-related protein kinase that colocalises with SC35 speckles. J. Cell Sci. 2001;114:2591–2603. doi: 10.1242/jcs.114.14.2591. [DOI] [PubMed] [Google Scholar]
- Krajewska M., Dries R., Grassetti A.V., Dust S., Gao Y., Huang H., Sharma B., Day D.S., Kwiatkowski N., Pomaville M., et al. CDK12 loss in cancer cells affects DNA damage response genes through premature cleavage and polyadenylation. Nat. Commun. 2019;10:1757. doi: 10.1038/s41467-019-09703-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B., Salzberg S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 2012;9:357–359. doi: 10.1038/nmeth.1923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang K., Gao X., Gilmore J.M., Florens L., Washburn M.P., Smith E., Shilatifard A. Characterization of human cyclin-dependent kinase 12 (CDK12) and CDK13 complexes in C-terminal domain phosphorylation, gene transcription, and RNA processing. Mol. Cell Biol. 2015;35:928–938. doi: 10.1128/MCB.01426-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang S., Hu L., Wu Z., Chen Z., Liu S., Xu X., Qian A. CDK12: a potent target and biomarker for human cancer therapy. Cells. 2020;9:1483. doi: 10.3390/cells9061483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liew C.W., Boucher J., Cheong J.K., Vernochet C., Koh H.J., Mallol C., Townsend K., Langin D., Kawamori D., Hu J., et al. Ablation of TRIP-Br2, a regulator of fat lipolysis, thermogenesis and oxidative metabolism, prevents diet-induced obesity and insulin resistance. Nat. Med. 2013;19:217–226. doi: 10.1038/nm.3056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love M.I., Huber W., Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naftelberg S., Schor I.E., Ast G., Kornblihtt A.R. Regulation of alternative splicing through coupling with transcription and chromatin structure. Annu. Rev. Biochem. 2015;84:165–198. doi: 10.1146/annurev-biochem-060614-034242. [DOI] [PubMed] [Google Scholar]
- Oesterreich F.C., Herzel L., Straube K., Hujer K., Howard J., Neugebauer K.M. Splicing of nascent RNA coincides with intron exit from RNA polymerase II. Cell. 2016;165:372–381. doi: 10.1016/j.cell.2016.02.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paculová H., Kohoutek J. The emerging roles of CDK12 in tumorigenesis. Cell Div. 2017;12:7. doi: 10.1186/s13008-017-0033-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulsen M.T., Veloso A., Prasad J., Bedi K., Ljungman E.A., Magnuson B., Wilson T.E., Ljungman M. Use of Bru-Seq and BruChase-Seq for genome-wide assessment of the synthesis and stability of RNA. Methods. 2014;67:45–54. doi: 10.1016/j.ymeth.2013.08.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulsen M.T., Veloso A., Prasad J., Bedi K., Ljungman E.A., Tsan Y.C., Chang C.W., Tarrier B., Washburn J.G., Lyons R., et al. Coordinated regulation of synthesis and stability of RNA during the acute TNF-induced proinflammatory response. Proc. Natl. Acad. Sci. USA. 2013;110:2240–2245. doi: 10.1073/pnas.1219192110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pendleton K.E., Chen B., Liu K., Hunter O.V., Xie Y., Tu B.P., Conrad N.K. The U6 snRNA m(6)A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell. 2017;169:824–835.e14. doi: 10.1016/j.cell.2017.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfister S.X., Ahrabi S., Zalmas L.P., Sarkar S., Aymard F., Bachrati C.Z., Helleday T., Legube G., La Thangue N.B., Porter A.C.G., Humphrey T.C. SETD2-dependent histone H3K36 trimethylation is required for homologous recombination repair and genome stability. Cell Rep. 2014;7:2006–2018. doi: 10.1016/j.celrep.2014.05.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Popova T., Manié E., Boeva V., Battistella A., Goundiam O., Smith N.K., Mueller C.R., Raynal V., Mariani O., Sastre-Garau X., Stern M.H. Ovarian cancers harboring inactivating mutations in CDK12 display a distinct genomic instability pattern characterized by large tandem duplications. Cancer Res. 2016;76:1882–1891. doi: 10.1158/0008-5472.CAN-15-2128. [DOI] [PubMed] [Google Scholar]
- Quereda V., Bayle S., Vena F., Frydman S.M., Monastyrskyi A., Roush W.R., Duckett D.R. Therapeutic targeting of CDK12/CDK13 in triple-negative breast cancer. Cancer Cell. 2019;36:545–558.e7. doi: 10.1016/j.ccell.2019.09.004. [DOI] [PubMed] [Google Scholar]
- Quintero O.A., DiVito M.M., Adikes R.C., Kortan M.B., Case L.B., Lier A.J., Panaretos N.S., Slater S.Q., Rengarajan M., Feliu M., Cheney R.E. Human Myo19 is a novel myosin that associates with mitochondria. Curr. Biol. 2009;19:2008–2013. doi: 10.1016/j.cub.2009.10.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rescigno P., Gurel B., Pereira R., Crespo M., Rekowski J., Rediti M., Barrero M., Mateo J., Bianchini D., Messina C., et al. Characterizing CDK12-mutated prostate cancers. Clin. Cancer Res. 2021;27:566–574. doi: 10.1158/1078-0432.CCR-20-2371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shima H., Matsumoto M., Ishigami Y., Ebina M., Muto A., Sato Y., Kumagai S., Ochiai K., Suzuki T., Igarashi K. S-adenosylmethionine synthesis is regulated by selective N(6)-adenosine methylation and mRNA degradation involving METTL16 and YTHDC1. Cell Rep. 2017;21:3354–3363. doi: 10.1016/j.celrep.2017.11.092. [DOI] [PubMed] [Google Scholar]
- Song G., Guo G., Du T., Li X., Wang J., Yan Y., Zhao Y. RALY may cause an aggressive biological behavior and a dismal prognosis in non-small-cell lung cancer. Exp. Cell Res. 2020;389:111884. doi: 10.1016/j.yexcr.2020.111884. [DOI] [PubMed] [Google Scholar]
- Sun L., Wan A., Zhou Z., Chen D., Liang H., Liu C., Yan S., Niu Y., Lin Z., Zhan S., et al. RNA-binding protein RALY reprogrammes mitochondrial metabolism via mediating miRNA processing in colorectal cancer. Gut. 2021;70:1698–1712. doi: 10.1136/gutjnl-2020-320652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tellier M., Zaborowska J., Caizzi L., Mohammad E., Velychko T., Schwalb B., Ferrer-Vicens I., Blears D., Nojima T., Cramer P., Murphy S. CDK12 globally stimulates RNA polymerase II transcription elongation and carboxyl-terminal domain phosphorylation. Nucleic Acids Res. 2020;48:7712–7727. doi: 10.1093/nar/gkaa514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tien J.F., Mazloomian A., Cheng S.W.G., Hughes C.S., Chow C.C.T., Canapi L.T., Oloumi A., Trigo-Gonzalez G., Bashashati A., Xu J., et al. CDK12 regulates alternative last exon mRNA splicing and promotes breast cancer cell invasion. Nucleic Acids Res. 2017;45:6698–6716. doi: 10.1093/nar/gkx187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tilgner H., Knowles D.G., Johnson R., Davis C.A., Chakrabortty S., Djebali S., Curado J., Snyder M., Gingeras T.R., Guigó R. Deep sequencing of subcellular RNA fractions shows splicing to be predominantly co-transcriptional in the human genome but inefficient for lncRNAs. Genome Res. 2012;22:1616–1625. doi: 10.1101/gr.134445.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang C., Wang H., Lieftink C., du Chatinier A., Gao D., Jin G., Jin H., Beijersbergen R.L., Qin W., Bernards R. CDK12 inhibition mediates DNA damage and is synergistic with sorafenib treatment in hepatocellular carcinoma. Gut. 2020;69:727–736. doi: 10.1136/gutjnl-2019-318506. [DOI] [PubMed] [Google Scholar]
- Wreggett K.A., Hill F., James P.S., Hutchings A., Butcher G.W., Singh P.B. A mammalian homologue of Drosophila heterochromatin protein 1 (HP1) is a component of constitutive heterochromatin. Cytogenet. Cell Genet. 1994;66:99–103. doi: 10.1159/000133676. [DOI] [PubMed] [Google Scholar]
- Wu Y.M., Cieślik M., Lonigro R.J., Vats P., Reimers M.A., Cao X., Ning Y., Wang L., Kunju L.P., de Sarkar N., et al. Inactivation of CDK12 delineates a distinct immunogenic class of advanced prostate cancer. Cell. 2018;173:1770–1782.e14. doi: 10.1016/j.cell.2018.04.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang T., Kwiatkowski N., Olson C.M., Dixon-Clarke S.E., Abraham B.J., Greifenberg A.K., Ficarro S.B., Elkins J.M., Liang Y., Hannett N.M., et al. Covalent targeting of remote cysteine residues to develop CDK12 and CDK13 inhibitors. Nat. Chem. Biol. 2016;12:876–884. doi: 10.1038/nchembio.2166. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Raw and processed sequencing data for Bru-seq and BruChase-seq have been deposited at the Gene Expression Omnibus (GEO), accession GSE191222.
This manuscript does not report original code.
Any additional information required to reanalyze the data reported herein is available upon request of the lead contact.