ABSTRACT
Enterococcus faecalis encodes two proteins, AcpA and AcpB, having the characteristics of acyl carrier proteins (ACPs). We report that the acpA gene located in the fatty acid synthesis operon is essential for fatty acid synthesis and the ΔacpA strain requires unsaturated fatty acids for growth. The ΔacpA strain could be complemented by a plasmid carrying a wild-type acpA gene, but not by a plasmid carrying a wild-type acpB gene. Substitution of four AcpA residues for those of AcpB resulted in a protein that modestly complemented the ΔacpA strain and restored fatty acid synthesis, although the acyl chains synthesized were unusually short.
IMPORTANCE Enterococcus faecalis, as well as related species, has two genes—acpA and acpB—encoding putative acyl carrier proteins (ACPs). It has been assumed that AcpA is essential for fatty acid synthesis whereas AcpB is involved utilization of environmental fatty acids. We report here the first experimental test of the essentiality of acpA and show that it is indeed an essential gene that cannot be replaced by acpB.
KEYWORDS: acyl carrier proteins, fatty acid synthesis, phospholipids
INTRODUCTION
Enterococcus faecalis and related streptococci are atypical in that their genomes encode two acyl carrier proteins (ACPs), whereas most bacteria have only a single ACP gene (1). One of these genes, called acpA, is located at the beginning of an operon that encodes virtually all of the genes required for saturated fatty acid synthesis (Fig. 1). The other gene, acpB, is located in a separate operon with plsX, a gene involved in producing activated long-chain acyl donors of phospholipid synthesis (Fig. 1). Both proteins are modified with the 4′-phosphopantetheine moiety (2). In previous work, we deleted acpB and found that it has an important role in the uptake of exogenous fatty acids (2, 3). Since ΔacpB strains grow normally, AcpB could not play an essential role in fatty acid synthesis, although acyl-AcpB was a substrate for the enoyl-ACP reductases of E. faecalis (2). However, several bacterial enoyl-ACP reductases have low substrate specificity (see below).
FIG 1.
Strategy for construction of the E. faecalis ΔacpA strain and its characterization by PCR. The genomic locations of the fatty acid synthesis genes are given above the scheme.
Based on the location of acpA in the fatty acid synthesis operon, AcpA has been designated as the carrier protein essential for fatty acid synthesis (4). However, there are no published data to support this designation, and no acpA mutant strains have been reported. We deleted acpA from the chromosome and found that that AcpA is required for fatty acid synthesis and that AcpB cannot replace AcpA. However, by substitution of four residues in the putative helix II of AcpB for those found in helix II of AcpA, we obtained an AcpB protein that could weakly complement loss of AcpA. The AcpP of Escherichia coli also weakly complemented the ΔacpA strain.
RESULTS AND DISCUSSION
Construction and properties of the E. faecalis ΔacpA strain.
Construction of the E. faecalis ΔacpA strain was difficult. Many attempts were made before success. We first tried to replace acpA with a chloramphenicol resistance cassette and to screen for the mutant strain using chloramphenicol resistance. However, an ΔacpA-Cm strain could not be obtained in liquid medium supplemented with 100 μM oleic acid. Subsequently, an ΔacpA-Cm strain was obtained by increasing the concentration of oleic acid to 5 mM and adding serum to 20%. However, complementation of this strain with a plasmid carrying the wild-type acpA gene was poor, indicating a polar effect on the downstream genes probably due to the chloramphenicol cassette. Finally, the ΔacpA strain was constructed by in-frame deletion, as shown in Fig. 1, using 5 mM oleic acid and 20% serum for the initial isolation. Additional details are given in Materials and Methods.
The ΔacpA strain is an unsaturated fatty acid auxotroph (Fig. 2) and completely lacks fatty acid synthetic ability as assayed by incorporation of [14C]acetate into phospholipid acyl chains (Fig. 3). Only unsaturated fatty acids (UFAs) allowed growth of the ΔacpA strain; saturated fatty acids (SFAs) fail to support growth. Serum which contains a mixture of UFA species also supported growth (Fig. 2). We tested complementation of the ΔacpA strain using a high-copy-number plasmid with wild-type acpA driven by a strong promoter, and this construct restored growth (Fig. 2). However, the growth was less robust than that of the wild-type strain, and the strain incorporated 30% less [14C]acetate into fatty acids relative to the wild-type strain (Fig. 3). This result is consistent with either a weak polar effect on the downstream genes or to toxicity of AcpA overproduction. To test these possibilities, the inducible expression system developed by Linares et al. (5) based on the regulatory system for agmatine degradation was used. Exogenous agmatine controls expression of agmatine degradation genes. We used low-copy-number (pTRK L2) or high-copy-number (pBM02) plasmids carrying the aguR gene (EF0731, which encodes a LuxR family transcriptional regulator for agmatine degradation), together with the aguB promoter that drives the agmatine degradation operon, to induce a range of AcpA levels in the ΔacpA strain. The low-copy-number acpA construct complemented growth of the ΔacpA strain but only upon addition of agmatine, whereas the high-copy-number acpA construct complemented growth without addition of agmatine (Fig. 4). Labeling with [14C]acetate showed that the low-copy-number acpA construct required agmatine for fatty acid synthesis, whereas the high-copy-number acpA construct did not (Fig. 5). However, low concentrations of agmatine (0.5 to 5 mM) progressively stimulated synthesis to wild-type levels. In contrast, higher agmatine concentrations inhibited fatty acid synthesis by about 80%. These results support our hypothesis that AcpA overexpression inhibits fatty acid synthesis. The fact that wild-type levels of fatty acid synthesis were obtained shows that the ΔacpA construct is nonpolar.
FIG 2.
Growth phenotypes of the ΔacpA strain with either 5% serum or different fatty acids.
FIG 3.
Growth phenotypes and fatty acid synthesis by [14C]acetate labeling of the ΔacpA strain and the complemented derivative strains. The wild-type acpA gene was expressed from the strong P32 promoter (1) in the high-copy-number plasmid pBM02. The number above each lane is the incorporation relative to the wild-type strain. Note that the label at the origin is from lipoproteins that extract into chloroform. One of the lipoprotein fatty acids is in amide linkage which is not cleaved under the basic conditions we use. Others have estimated 60 to 80 different lipoproteins in E. faecalis. The ester-linked lipoprotein fatty acids are derived from phospholipids.
FIG 4.
Growth phenotypes of the ΔacpA strain complemented with the wild-type acpA gene under agmatine control. Growth with agmatine induction at various concentrations was tested.
FIG 5.
Fatty acid synthesis of the agmatine-induced complemented strains of Fig. 4 by [14C]acetate labeling. The number above each lane is the incorporation relative to the wild-type strain.
We constructed a high-copy-number plasmid with E. coli acpP driven by a strong promoter. This construct gave very weak complementation of the ΔacpA strain and labeling with [14C]acetate demonstrated that this plasmid supported saturated fatty acid synthesis, but only traces of UFA were synthesized (Fig. 3). This suggests incompatibility E. coli ACP with the E. faecalis UFA synthesis enzymes (FabO and FabN). We also tested the AcpA of Lactococcus lactis II1403 and found that it fully complemented the ΔacpA strain (data not shown). This was expected since the AcpA proteins are 68% identical, and the bacteria are both members of the Lactobacillales.
Relationship between AcpA and AcpB.
Deletion of the E. faecalis acpB gene had no ffect on fatty acid synthesis consistent with the role of AcpB in fatty acid uptake (3, 6). Expression of acpB is not regulated by FabT, the repressor of the fatty acid synthesis operon which includes acpA (3) (Fig. 1). Expression of AcpB failed to complement either the growth or fatty acid synthesis defects of the ΔacpA strain despite prior data showing activity with the E. faecalis enoyl-ACP reductases (6). This discrepancy can be attributed to the low substrate specificity of enoyl-ACP reductases (7) and the fact that AcpB has a 4′-phosphopantetheine moiety (6). Coenzyme A thioesters are known substrates for enoyl-ACP reductases of the diverse bacteria listed in Table S3 in the supplemental material, so ACP substrates are not required in vitro. An extreme example is Bacillus subtilis in which both enoyl-ACP reductases are active with acyl-N-acetylcysteamine substrates. N-Acetylcysteamine is the thiol proximal half of 4′-phosphopantetheine.
The most striking differences between AcpA and AcpB are in helix II (6), the helix involved in most ACP-enzyme interactions (8) (Fig. 6). By comparison of the ACP sequences of several diverse bacteria, we first chose two acpB residues for substitution with the analogous AcpA residues (Fig. 6). This construct was unable to complement growth of the ΔacpA strain and labeling with [14C]acetate showed only traces of saturated fatty acids (Fig. 7). We therefore chose two additional residues for substitution (Fig. 6). The AcpB derivative having four substitutions with AcpA residues complemented the ΔacpA strain although growth was modest (Fig. 7). This construct showed fatty acid synthesis activity, and both saturated and unsaturated acyl chains were produced (Fig. 8). However, the phospholipid acyl chains were shorter than those seen in the wild-type strain. Fourteen carbon acids, both saturated and unsaturated, were synthesized, and only traces of cis-vaccenic acid (C18:1), the major chain in wild-type phospholipids, were present (Table 1). The accumulation of short-chain acids in the phospholipids argues that fatty acid synthesis is abnormally slow in the strain expressing the AcpB having four substituted AcpA residues, and this slow synthesis competes poorly with the phospholipid acyltransferases. The liquid chromatography-mass spectroscopy data for all the strains except the AcpB with two substituted AcpA residues are presented in Table 1. Note that the AcpB having four substitutions with AcpA residues retained much of the AcpB function in incorporation of exogenous oleic acid into phospholipids (Fig. 9).
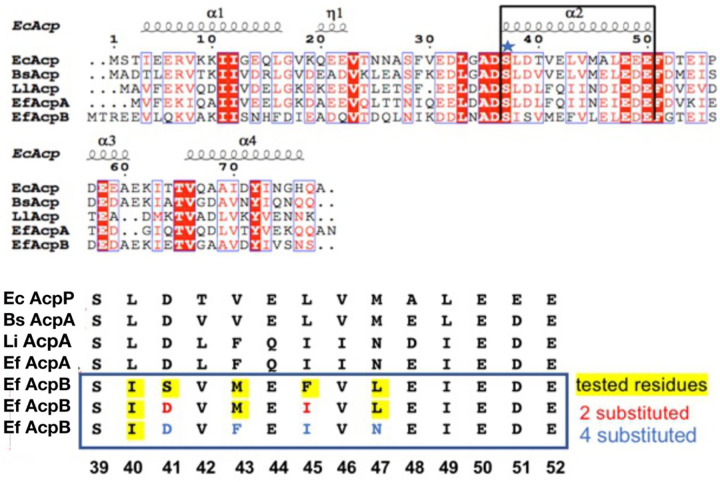
FIG 6.
(Upper panel) Sequence alignments of ACPs of various bacteria with the helix II sequences in a bracket. (Lower panel) Helix II sequences from the upper panel and sites where AcpA residues were substituted in place of AcpB residues. The ACPs are from various organisms as follows: Ec, E. coli; Bs, Bacillus subtilis; Ll, Lactococcus lactis; and Ef, E. faecalis.
FIG 7.
Complementation of the ΔacpA strain by acpB and substitution derivatives in complementation of the ΔacpA strain. 2S and 4S denote the residue substitutions given in Fig. 6. The acpB genes were expressed from the strong P32 promoter (10) in the high-copy-number plasmid pBM02.
FIG 8.

Fatty acid synthesis by [14C]acetate labeling of the ΔacpA strain complemented by plasmids encoding the substituted acpB derivatives. The two unsaturates in the right-hand lane are 14:1 (probably cis-7-tetradecenoic acid) and palmitoleic acid (cis-9-hexadecenoic acid) (Table 1). The number above each lane is the incorporation relative to the wild-type strain.
TABLE 1.
Fatty acid compositions of wild-type, ΔacpA, and complemented strainsa
| Fatty acid (%) | Mean fatty acid composition (% of total fatty acid esters) ± SD |
|||||
|---|---|---|---|---|---|---|
| Wild type | ΔacpA mutant + 5% serum | ΔacpA mutant + oleic acid | ΔacpA/acpA mutant | ΔacpA/Ecacp mutant | ΔacpA/acpB 4 Sub mutant | |
| C12:0 | 10.3 ± 1.1 | |||||
| C14:0 | 1.7 ± 0.3 | 0.8 ± 0.7 | 2.2 ± 0.6 | 20.9 ± 1.4 | 51.4 ± 2.9 | |
| C14:1 (7) | 1.3 ± 0.1 | 7.8 ± 3.4 | 11.0 ± 0.4 | |||
| C16:0 | 29.6 ± 2.7 | 31.0 ± 2.2 | 32.1 ± 0.4 | 43.1 ± 0.7 | 10.1 ± 0.2 | |
| C16:1 (9) | 3.3 ± 0.3 | 5.1 ± 0.2 | 4.1 ± 0.3 | 15.8 ± 2.3 | ||
| C18:0 | 5.1 ± 0.6 | 10.6 ± 1.6 | 0.7 ± 0.1 | 7.5 ± 0.4 | 7.4 ± 2.8 | 1.0 ± 0.2 |
| C18:1 (9) | 51.1 ± 4.8 | 99.4 ± 0.2 | ||||
| C18:1 (11) | 59.0 ± 3.5 | 52.6 ± 1.4 | 20.9 ± 3.3 | 0.5 ± 0.1 | ||
| C19:0-cyclo | 1.1 ± 0.2 | 2.0 ± 0 | 0.6 ± 0.2 | |||
| UFA/SFA | 1.73 | 1.39 | 142 | 1.42 | 0.40 | 0.38 |
UFAs and UFA/SFA ratios are indicated in boldface type. The entry on the right side of the shill denotes a plasmid carried gene.
FIG 9.

Fatty acid uptake by [14C]oleic acid labeling of the ΔacpA strain complemented by plasmids encoding the substituted acpB derivatives. The compounds running at the top of the plate are cyclopropane fatty acid derived from [14C]oleic acid. The number above each lane gives the incorporation relative to the wild-type strain.
MATERIALS AND METHODS
Materials.
The fatty acids and antibiotics were purchased from Sigma-Aldrich. The DNA polymerases, Gibson assembly kit, restriction endonucleases, and T4 ligase were obtained from NEB. Sodium [1-14C]acetate (specific activity, 58.6 mCi/mmol) was provided by Moravek, Inc., while the [1-14C]oleic acid (specific activity, 55 mCi/mmol) was purchased from American Radiolabeled Chemicals.
Bacterial strains, plasmids, and incubation.
The bacterial strains and plasmids used in this study are listed in Table S1, and the primers used for this study are listed in Table S2. E. coli cells were incubated at 37°C in Luria-Bertani medium (tryptone, 10 g/L; yeast extract, 5 g/L; NaCl, 10 g/L), whereas the E. faecalis cells were cultured at 37°C in M17 medium (Sigma). In a few cases, we used AC medium (Sigma), which contains low levels of fatty acids. M17 medium plates were solidified with agarose to avoid the fatty acids in agar. Antibiotics were added at the following concentrations (in mg/L): sodium ampicillin, 100 (for E. coli); kanamycin sulfate, 50 (for E. coli); chloramphenicol, 30 (for E. coli) and 10 (for E. faecalis); and erythromycin, 250 (for E. coli) and 10 (for E. faecalis). Fatty acids were added at 0.1 mM unless otherwise stipulated.
Construction of E. faecalis acpA deletion strains.
The E. faecalis ΔacpA deletion strain was constructed using methods described earlier (9) and the PCR primers listed Table S2. A cassette for deletion of the gene was constructed by overlap PCR. Upstream and downstream ~500-bp DNA fragments were assembled by overlap PCR and then inserted into temperature-sensitive vector pBVGh using Gibson assembly to produce pHJ493. Plasmid pHJ493 was transformed into competent E. faecalis FA2-2 cells by electroporation. One blue transformant colony was streaked on M17 agar medium containing 10 μg/mL erythromycin and 0.1 mg/mL 5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside (X-Gal) and then incubated overnight at 42°C to confirm the blue colony, indicating chromosomal integration of the plasmid. One such single-crossover strain was cultured in AC liquid medium with 5 mM oleic acid and 20% serum for 4 h at 30°C and then shifted to 42°C overnight. This step was repeated several times, and the final culture was diluted and plated on AC agar containing X-Gal and 200 μM oleic acid, followed by incubation for 24 to 48 h at 42°C. Genomic DNAs from white colonies, which represent double-crossover events, were extracted, and screened by PCR analysis to give the ΔacpA strain (named HJ484).
The acpA and acpB expression plasmids were constructed by inserting the acpA or acpB genes together with a P32 promoter or agmatine induced promoter into a shuttle plasmid vector either pBM02 or pTRK L2 through Gibson assembly. Transformation of E. faecalis ΔacpA or ΔacpB strains with these plasmids produced the complementation strains.
TLC analysis of radioactive labeled fatty acid methyl esters from membrane phospholipids.
To test de novo fatty acid biosynthesis, E. faecalis strains were started at an optical density (OD) of 0.5 in M17 medium, followed by incubation at 37°C in the presence of 1 mCi/L sodium [1-14C]acetate with or without an exogenous fatty acid at 0.1 mM. The ΔacpA strain was cultured in M17 liquid medium with 100 μM oleic acid, and the cells were collected and washed three times with M17 liquid medium before labeling using a 14C-labeled compound. After labeling, the cells were lysed with methanol-chloroform (2:1) solution, and the phospholipids were extracted in chloroform and then dried under nitrogen. The fatty acyl groups were methylated by 25% (wt/vol) sodium methoxide, extracted by hexanes, and processed for thin-layer chromatography (TLC) analysis on Analtech silica gel containing 20% silver nitrate in toluene at −20°C. The plates containing the 14C-labeled fatty acid methyl esters were exposed on the phosphorimager GE Typhoon FLA700 Scanner and then analyzed using ImageQuant TL software.
Monounsaturated methyl esters separate from saturated esters because the double bonds interact with the silver nitrate in the plate (π-π interactions). Under the right conditions (toluene as solvent and run at −20°C), monounsaturated methyl esters separate according to the position of the double bond. Hence, Δ9 C16 runs slower than Δ11 C18.
To test incorporation of exogenous unsaturated fatty acids, E. faecalis strains were inoculated at an OD of 0.3 in M17 medium, followed by incubation at 37°C to OD600 values of about 1.0% ± 10%, in the presence of 0.1 mCi/L [1-14C]oleic acid with 0.1 mM nonradioactive oleate. The cells were washed with water, and the phospholipids were extracted, methylated, and processed for TLC analysis as described above.
Gas chromatography-mass spectrometry analysis of the fatty acids of cell membrane phospholipids.
E. faecalis strains were inoculated at logarithmic phase in M17 medium. The cells were lysed by methanol-chloroform (2:1) solution, and the phospholipids were extracted into chloroform and dried under a nitrogen stream. The fatty acyl groups were methylated by transesterification using 25% (wt/vol) sodium methoxide in methanol, extracted with hexanes, and sent to the Carver Metabolomics Center of the University of Illinois at Urbana-Champaign for gas chromatography-mass spectrometry analysis. The same procedure was used in the presence of exogenous oleic acid since sodium methoxide in methanol cannot methylate free fatty acids.
ACKNOWLEDGMENTS
We thank Alexander Vladimirovich Ulanov for the gas chromatography-mass spectrometry facility.
This study was supported by National Institutes of Health grant AI15650 from the National Institute of Allergy and Infectious Diseases.
Footnotes
Supplemental material is available online only.
Contributor Information
John E. Cronan, Email: jecronan@illinois.edu.
Michael J. Federle, University of Illinois at Chicago
REFERENCES
- 1.Dehal PS, Joachimiak MP, Price MN, Bates JT, Baumohl JK, Chivian D, Friedland GD, Huang KH, Keller K, Novichkov PS, Dubchak IL, Alm EJ, Arkin AP. 2010. MicrobesOnline: an integrated portal for comparative and functional genomics. Nucleic Acids Res 38:D396–D400. 10.1093/nar/gkp919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Zhu L, Bi H, Ma J, Hu Z, Zhang W, Cronan JE, Wang H. 2013. The two functional enoyl-acyl carrier protein reductases of Enterococcus faecalis do not mediate triclosan resistance. mBio 4:e00613-13–e00613. 10.1128/mBio.00613-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Zou Q, Dong H, Zhu L, Cronan JE. 2022. The Enterococcus faecalis FabT transcription factor regulates fatty acid biosynthesis in response to exogeneous fatty acids. Front Microbiol 13:877582. 10.3389/fmicb.2022.877582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Jerga A, Rock CO. 2009. Acyl-Acyl carrier protein regulates transcription of fatty acid biosynthetic genes via the FabT repressor in Streptococcus pneumoniae. J Biol Chem 284:15364–15368. 10.1074/jbc.C109.002410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Linares DM, Perez M, Ladero V, Del Rio B, Redruello B, Martin MC, Fernandez M, Alvarez MA. 2014. An agmatine-inducible system for the expression of recombinant proteins in Enterococcus faecalis. Microb Cell Fact 13:169. 10.1186/s12934-014-0169-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhu L, Zou Q, Cao X, Cronan JE. 2019. Enterococcus faecalis encodes an atypical auxiliary acyl carrier protein required for efficient regulation of fatty acid synthesis by exogenous fatty acids. mBio 10:e00577-19. 10.1128/mBio.00577-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Massengo-Tiassé RP, Cronan JE. 2009. Diversity in enoyl-acyl carrier protein reductases. Cell Mol Life Sci 66:1507–1517. 10.1007/s00018-009-8704-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cronan JE. 2014. The chain-flipping mechanism of ACP (acyl carrier protein)-dependent enzymes appears universal. Biochem J 460:157–163. 10.1042/BJ20140239. [DOI] [PubMed] [Google Scholar]
- 9.Blancato VS, Magni C. 2010. A chimeric vector for efficient chromosomal modification in Enterococcus faecalis and other lactic acid bacteria. Lett Appl Microbiol 50:542–546. 10.1111/j.1472-765X.2010.02815.x. [DOI] [PubMed] [Google Scholar]
- 10.Zhu D, Liu F, Xu H, Bai Y, Zhang X, Saris PE, Qiao M. 2015. Isolation of strong constitutive promoters from Lactococcus lactis subsp. lactis N8. FEMS Microbiol Lett 362:fnv107. 10.1093/femsle/fnv107. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Tables S1 to S3. Download jb.00202-22-s0001.pdf, PDF file, 0.1 MB (137.4KB, pdf)