Abstract
A rearrangement between the symbiotic plasmid (pRleVF39d) and a nonsymbiotic plasmid (pRleVF39b) in Rhizobium leguminosarum bv. viciae VF39 was observed. The rearranged derivative showed the same plasmid profile as its parent strain, but hybridization to nod, fix, and nif genes indicated that most of the symbiotic genes were now present on a plasmid corresponding in size to pRleVF39b instead of pRleVF39d. On the other hand, some DNA fragments originating from pRleVF39b now hybridized to the plasmid band at the position of pRleVF39d. These results suggest that a reciprocal but unequal DNA exchange between the two plasmids had occurred.
Rhizobium leguminosarum bv. viciae is a native soil bacterium that can enter nitrogen-fixing symbioses (root nodules) with several legumes such as Pisum, Vicia, and Lens spp. It generally contains 1 to 10 plasmids which vary in size from 30 kb to more than 800 kb (11, 12, 14). Most of the genes required for nodule formation (nod) and nitrogen fixation (nif and fix) are carried on a plasmid that is traditionally called the symbiotic plasmid or pSym (6, 11).
Plasmids are important genetic components for the divergence and adaptation of microbial populations because they contribute to genomic plasticity. Although plasmid profiles in rhizobia can be considered as a comparatively stable character, strains can lose some of their traits due to loss or partial deletion of a plasmid (3, 27). Recombination and rearrangement events between plasmids is also frequently observed. For example, for R. leguminosarum bv. viciae it was found that a transmissible plasmid could pick up symbiotic determinants from the nontransmissible pSym (1), and pSym rearrangements have been described and studied in detail in R. etli and R. tropici (2, 4, 21, 22). One of these rearrangements is a deletion of the symbiotic gene region, accompanied by a concomitant amplification of other sequences, so that the overall size of the plasmid is not significantly changed (21).
In this study, we present direct evidence of a DNA rearrangement between two indigenous plasmids of R. leguminosarum bv. viciae VF39SM, a model strain that is routinely used in our laboratory (Table 1). It contains six plasmids, ranging in size from 130 kb to approximately 600 kb (9). They were designated as pRleVF39a to pRleVF39f in order of increasing molecular size. The symbiotic plasmid pRleVF39d carries the nod, nif, and most of the fix genes (9, 10). Some fix genes, such as fixKL and a second copy of the fixNOQP operon, reside on plasmid pRleVF39c, which also carries genes involved in lipopolysaccharide synthesis (10, 15, 24). Little is known about pRleVF39b except that it carries a restriction-modification system (20) and that it is self-transmissible (9).
TABLE 1.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Relevant characteristicsa | Source or reference |
|---|---|---|
| R. leguminosarum bv. viciae | ||
| VF39SM | Streptomycin-resistant derivative of wild-type strain VF39 | 17 |
| VF39SM′ | A derivative of VF39SM with genomic rearrangement | This work |
| VF39SM" | A derivative of VF39SM′ with deletion in plasmid pRleVF39b′ | This work |
| Plasmids | ||
| pXT | A derivative of plasmid pGEM-7Zf(+); Apr | 7 |
| pXTZ11 to pXTZ14 | pXT derivatives containing XhoI/HindIII fragments of pRleVF39b | This work |
| pJQ177 | pK19 derivative, 6.6-kb EcoRI/BamHI fragment carrying VF39 fixLKNO genes; Kmr | 24 |
| pTP100 | pUC19 derivative, 4.3-kb fragment carrying VF39 fixBCX and nifA genes; Apr | 15 |
| Cosmid 220 | 30-kb DNA region from pRleVF39b cloned in vector pRK7813 | 20 |
Apr, ampicillin resistant.
Hybridization analyses reveal a pSym rearrangement.
The rearrangement described here was found when single colonies of the original VF39SM stock culture, stored in 20% glycerol at −20°C since 1993, were checked by plasmid profiling using a modified Eckhardt procedure (8, 16) and subsequent Southern hybridization analysis with nodABC genes as a probe. The digoxigenin-labeled probe was prepared via PCR by using primers pnodA (5′-AGT GCG ATG GAA AAT ATG CTG-3′) and pnodC (5′-GGA AGC GCA AGC AAA GTA TC-3′), which correspond to positions 4521 to 4541 and positions 2376 to 2395, respectively, in the pRl1JI nodABC sequence (accession number X99520). Although all isolates showed the plasmid pattern characteristic for VF39SM (Fig. 1A), 1 out of 100 colonies was identified in which hybridization to nodABC occurred to a plasmid corresponding in size to pRleVF39b (240 kb), whereas in all the others the hybridization was as expected to pRleVF39d (pSym, 330 kb) (Fig. 1B). This result suggested the event of a DNA rearrangement in this isolate. Therefore, it was renamed VF39SM′ (Table 1), and its plasmids were accordingly renamed pRleVF39a′ to pRleVF39f′.
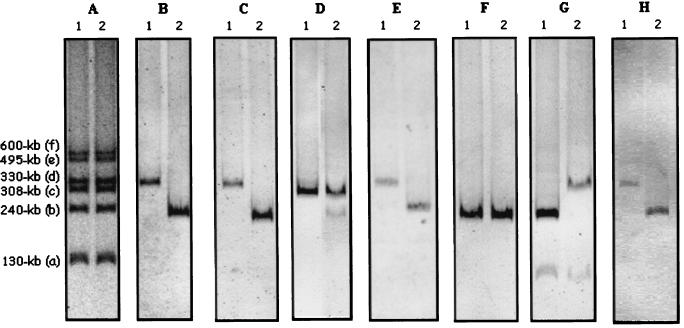
FIG. 1.
Profiles and hybridization results of the plasmids in R. leguminosarum bv. viciae VF39SM (lane 1) and its rearranged derivative VF39SM′ (lane 2). (A) Eckhardt gel showing the banding pattern of the six plasmids (negative image); their designation (for the parental VF39SM strain) and the estimated sizes of the plasmid bands are given on the left. (B to H) Hybridization results of the plasmid gels with nodABC (B), nodO (C), fixLKNO (D), fixBCX-nifA (E), pXTZ11 (F), pXTZ12 (G), and repC (H).
To estimate the extent of this rearrangement, the localization of other symbiotic genes was analyzed. Southern hybridization with a nodO probe, amplified and labeled using primers pnodO1 (5′-CGT AAG CGA GGT AAG ATC CG-3′) and pnodO2 (5′-AGA ACG ACT GGA TTG ATG CC-3′), which were designed on the basis of the pRL1JI sequence (accession number X17285), indicated that in VF39SM′ this gene was now also carried on the 240-kb plasmid instead of pRleVF39d (Fig. 1C). When the fixLKNO gene region was used as a probe, a strong signal with plasmid pRleVF39c, which carries one copy of the fixNOQP operon upstream of the fixKL region (15, 24), was observed for both strains; the second copy of fixNOQP, however, which normally is carried on pRleVF39d in VF39SM (24) was detected at the position of pRleVF39b′ in the rearranged strain (Fig. 1D; the hybridization signals of this copy are weak because it does not contain the fixLK genes). Likewise, hybridization to the fixBCX-nifA genes had changed from pRleVF39d to pRleVF39b′ (Fig. 1E). These results suggest that in the rearranged strain VF39SM′ almost the entire symbiotic region originally present on plasmid pRleVF39d is now carried on a plasmid banding at position pRleVF39b′.
Plant tests with Vicia hirsuta confirmed that the two strains had similar symbiotic properties with respect to nodule formation, acetylene reduction activity, and plant shoot dry weights (data not shown).
The rearrangement occurred by reciprocal but unequal exchange of DNA.
To test whether there was also a shift in the hybridization pattern of sequences originally carried by plasmid pRleVF39b, DNA of the parental pRleVF39b plasmid was isolated from the Eckhardt gel, digested completely with XhoI and HindIII, and cloned into vector pXT (7). Four clones (pXTZ11 to pXTZ14 [Table 1]), which contained 600-bp to 2.8-kb inserts, were randomly chosen and used to probe the plasmids of VF39SM and VF39SM′. The 2.5-kb DNA fragment cloned in pXTZ11 hybridized to the 240-kb plasmid in both strains (Fig. 1F); pXTZ12, pXTZ13, and pXTZ14 hybridized to plasmid pRleVF39b in the parental strain but to the pRleVF39d′ band in the rearranged derivative (exemplified for pXTZ12 in Fig. 1G; in both strains, pXTZ12 also gave a weak hybridization to pRleVF39a). These data suggest that the two plasmids had exchanged genetic information.
Since the DNA arrangement did not visibly change the overall plasmid profile of the strain, it could have occurred in two possible ways. The first is that the two plasmids have exchanged a DNA region of exactly the same size, which would mean that plasmid pRleVF39d′ is a rearranged version of the original pSym (now without the sym region and containing part of pRleVF39b instead) and that the plasmid pRleVF39b′ represents plasmid pRleVF39b, which has acquired the sym gene region from plasmid pRleVF39d in exchange for its own DNA of the same size. The alternative possibility is that the DNA region obtained by plasmid pRleVF39b exceeds the region transferred to plasmid pRleVF39d by 90 kb, thus shifting the size of plasmid pRleVF39b from 240 to 330 kb (i.e., to the gel position of pRleVF39d) and decreasing the size of pRleVF39d accordingly. This would mean that in VF39SM′ the plasmid band with a size of 240 kb (pRleVF39b′) is actually a rearranged version of pSym (pRleVF39d) still containing the sym region.
To determine which possibility had occurred, the replication regions of the two plasmids were analyzed. By using a pair of conserved PCR primers, RC1 and RC3, developed to amplify repC genes of R. leguminosarum (19, 26), a product with the expected size of 750 bp was amplified from total DNA of VF39SM, and it was used as a probe to hybridize the plasmid gels. As shown in Fig. 1H, the repC probe was found to hybridize to pRleVF39d in VF39SM and to pRleVF39b′ in VF39SM′. This means that the symbiotic genes are still contained on the original replicon, suggesting that the rearrangement had occurred by the second way described above.
Since several copies of a putative insertion sequence (ISRle39) were found to be present on plasmid pRleVF39b (20), we tested whether these elements were involved in the DNA rearrangement. ISRle39 was amplified from total VF39SM DNA and used as a hybridization probe. In VF39SM, a strong signal was found to plasmid pRleVF39b (and also to plasmid pRleVF39a), whereas in VF39SM′ the signal had shifted to the position of pRleVF39d′ (data not shown). Provided that the rearrangement had occurred by the second mechanism proposed (i.e., that the plasmid band designated pRleVF39d′ corresponds to the rearranged pRleVF39b plasmid), this would mean that these IS elements have not been part of the DNA exchange.
It was also checked whether further rearrangements could occur. For this purpose, 50 single colonies of strain VF39SM′ were studied by plasmid profile and nodABC Southern hybridization analyses. A deletion in plasmid pRleVF39b′ of about 45 kb was detected in one colony (VF39SM"). No hybridization signal to nodABC was found in this deletion derivative and it failed to nodulate V. hirsuta, indicating that the deleted DNA region included the nodulation genes. All other colonies had normal plasmid profiles, and their nod genes were still located on pRleVF39b′.
Delineation of one rearrangement site.
By using cosmid 220 (kindly provided by M. F. Hynes), which contains a 30-kb insert from pRleVF39b, and appropriate subclones thereof, a 2.2-kb BamHI/HindIII fragment was identified that hybridized only to pRleVF39b in the original strain but to both pRleVF39b′ and pRleVF39d′ in VF39SM′ (data not shown), suggesting that this region was involved in the generation of the rearrangement. DNA sequence analysis (accession number AJ292764) revealed an open reading frame (ORF1) with 73% homology, at both the DNA and amino acid levels, to Y4JO, a hypothetical 36.1-kDa protein encoded by pNGR234a from Sinorhizobium sp. strain NGR234 (accession number AE000080, positions 3600 to 4646). Y4JO may be a fragment of a larger protein of as-yet-unknown function (5). It is notable that the C-terminal part of ORF1 (positions 324 to 359) shows 69% homology at protein level with the C-terminal end of a putative transposase of Vibrio cholerae (accession number AB012957), suggesting that the ORF1 gene product may have some function in the observed DNA rearrangement, which will be studied in the future. Considering that the other sequences upstream and downstream of ORF1 show no homology with the neighboring sequences of Y4JO, it seems that this DNA region has undergone recombination several times.
Although at present, the molecular mechanisms leading to the rearrangement described here are not understood, the data presented imply that it is caused by a reciprocal but unequal exchange of DNA, resulting in a decrease in the size of pSym and a simultaneous increase in the size of pRleVF39b. Symbiotic plasmid rearrangement have been previously described in field isolates of Sinorhizobium meliloti (18) and R. leguminosarum (23, 25). Mazurier and Laguerre (13) recently reported an unusual location of symbiotic genes in some R. leguminosarum bv. viciae field isolates, which could be the result of genome rearrangements between the pSym and another replicon. Our own work on the pSym diversity among R. leguminosarum field isolates indicated that identical sym gene genotypes can be carried by plasmids of different sizes, ranging from 190 to 630 kb (unpublished data). These findings suggest that plasmid DNA rearrangements similar to that described here may also occur in the natural environment.
Acknowledgments
We thank M. F. Hynes (University of Calgary, Calgary, Ontario, Canada) for fruitful discussions and for providing cosmid clone 220. We also thank A. Seibold for helpful technical advice and assistance and B. Boesten for constructive criticism.
X.-X. Zhang was supported by an Alexander-von-Humboldt fellowship.
REFERENCES
- 1.Brewin N J, Wood E A, Johnston A W B, Dibb N J, Hombrecher G. Recombinant nodulation plasmids in Rhizobium leguminosarum. J Gen Microbiol. 1982;128:1817–1827. [Google Scholar]
- 2.Brom S, García de los Santos A, Girard M L, Dávila G, Palacios R, Romero D. High-frequency rearrangements in Rhizobium leguminosarum bv. phaseoli plasmids. J Bacteriol. 1991;173:1344–1346. doi: 10.1128/jb.173.3.1344-1346.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Djordjevic M A, Zurkowski W, Rolfe B G. Plasmids and stability of symbiotic properties of Rhizobium trifolii. J Bacteriol. 1982;151:560–568. doi: 10.1128/jb.151.2.560-568.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Flores M, González V, Pardo M A, Leija A, Martínez E, Romero D, Pinero D, Dávila G, Palacios R. Genomic instability in Rhizobium phaseoli. J Bacteriol. 1988;170:1191–1196. doi: 10.1128/jb.170.3.1191-1196.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Freiberg C, Fellay R, Bairoch A, Broughton W J, Rosenthal A, Perret X. Molecular basis of symbiosis between Rhizobium and legumes. Nature. 1997;387:394–401. doi: 10.1038/387394a0. [DOI] [PubMed] [Google Scholar]
- 6.García de los Santos A, Brom S, Romero D. Rhizobium plasmids in bacteria-legume interactions. World J Microbiol Biotechnol. 1996;12:119–125. doi: 10.1007/BF00364676. [DOI] [PubMed] [Google Scholar]
- 7.Harrison J, Molloy P L, Clark S J. Direct cloning of polymerase chain reaction products in an XcmI T-vector. Anal Biochem. 1994;216:235–236. doi: 10.1006/abio.1994.1033. [DOI] [PubMed] [Google Scholar]
- 8.Hynes M F, Simon R, Pühler A. The development of plasmid-free strains of Agrobacterium tumefaciens by using incompatibility with a Rhizobium meliloti plasmid to eliminate pAEC58. Plasmid. 1985;13:99–105. doi: 10.1016/0147-619x(85)90062-9. [DOI] [PubMed] [Google Scholar]
- 9.Hynes M F, Brucksch K, Priefer U B. Melanin production encoded by a cryptic plasmid in a Rhizobium leguminosarum strain. Arch Microbiol. 1988;150:326–332. [Google Scholar]
- 10.Hynes M F, McGregor N F. Two plasmids other than the nodulation plasmid are necessary for formation of nitrogen-fixing nodules by Rhizobium leguminosarum. Mol Microbiol. 1990;4:567–574. doi: 10.1111/j.1365-2958.1990.tb00625.x. [DOI] [PubMed] [Google Scholar]
- 11.Martínez-Romero E, Palacios R. The Rhizobium genome. Crit Rev Plant Sci. 1990;9:59–93. [Google Scholar]
- 12.Martínez-Romero E, Caballero-Mellado J. Rhizobium phylogenies and bacterial genetic diversity. Crit Rev Plant Sci. 1996;15:113–140. [Google Scholar]
- 13.Mazurier S I, Laguerre G. Unusual location of nod and nif genes in Rhizobium leguminosarum bv. viciae. Can J Microbiol. 1997;43:399–402. [Google Scholar]
- 14.Mercado-Blanco J, Toro N. Plasmids in rhizobia: the role of nonsymbiotic plasmids. Mol Plant-Microbe Interact. 1996;9:535–545. [Google Scholar]
- 15.Patschkowski T, Schlüter A, Priefer U B. Rhizobium leguminosarum bv. viciae contains a second fnr/fixK-like gene and an unusual fixL homologue. Mol Microbiol. 1996;21:267–280. doi: 10.1046/j.1365-2958.1996.6321348.x. [DOI] [PubMed] [Google Scholar]
- 16.Priefer U B. Isolation of plasmid DNA. In: Pühler A, Timmis K N, editors. Advanced molecular genetics. Berlin, Germany: Springer-Verlag; 1984. pp. 14–24. [Google Scholar]
- 17.Priefer U B. Genes involved in lipopolysaccharide production and symbiosis are clustered on the chromosome of Rhizobium leguminosarum bv. viciae VF39. J Bacteriol. 1989;171:6161–6168. doi: 10.1128/jb.171.11.6161-6168.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Rastogi V K, Bromfield E S P, Whitwill S T, Barran L R. A cryptic plasmid of indigenous Rhizobium meliloti possesses reiterated nodC and nifE genes and undergoes DNA arrangement. Can J Microbiol. 1992;38:563–568. [Google Scholar]
- 19.Rigottier-Gois L, Turner S L, Young J P W, Amarger N. Distribution of repC plasmid-replication sequences among plasmids and isolates of Rhizobium leguminosarum bv. viciae from field populations. Microbiology. 1998;144:771–780. doi: 10.1099/00221287-144-3-771. [DOI] [PubMed] [Google Scholar]
- 20.Rochepeau P, Selinger L B, Hynes M F. Transposon-like structure of a new plasmid-encoded restriction-modification system in Rhizobium leguminosarum VF39SM. Mol Gen Genet. 1997;256:387–396. doi: 10.1007/s004380050582. [DOI] [PubMed] [Google Scholar]
- 21.Romero D, Brom S, Martínez-Salazar J, Girard M L, Palacios R, Dávila G. Amplification and deletion of a nod-nif region in the symbiotic plasmid of Rhizobium phaseoli. J Bacteriol. 1991;173:2435–2441. doi: 10.1128/jb.173.8.2435-2441.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Romero D, Martínez-Salazar J, Girard L, Brom S, Dávila G, Palacios R, Flores M, Rodriguez C. Discrete amplifiable regions (amplicons) in the symbiotic plasmid of Rhizobium etli CFN42. J Bacteriol. 1995;177:973–980. doi: 10.1128/jb.177.4.973-980.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Schofield P R, Gibson A H, Dudman W F, Watson J M. Evidence for genetic exchange and recombination of Rhizobium symbiotic plasmids in a soil population. Appl Environ Microbiol. 1987;53:2942–2947. doi: 10.1128/aem.53.12.2942-2947.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Schlüter A, Patschkowski T, Quandt J, Selinger L B, Weidner S, Krämer M, Zhou L M, Hynes M F, Priefer U B. Functional and regulatory analysis of the two copies of the fixNOQP operon of Rhizobium leguminosarum strain VF39. MPMI. 1997;10:605–616. doi: 10.1094/MPMI.1997.10.5.605. [DOI] [PubMed] [Google Scholar]
- 25.Selbitscka W, Lotz W. Instability of cryptic plasmids affects the symbiotic efficiency of Rhizobium leguminosarum bv. viciae strains. Mol Plant-Microbe Interact. 1991;4:608–618. [Google Scholar]
- 26.Turner S L, Rigottier-Gois L, Power R S, Amarger N, Young J P W. Diversity of recC plasmid-replication sequences in Rhizobium leguminosarum. Microbiology. 1996;142:1705–1713. doi: 10.1099/13500872-142-7-1705. [DOI] [PubMed] [Google Scholar]
- 27.Weaver R W, Wei G R, Berryhill D L. Stability of plasmids in R. phaseoli during culture. Soil Biol Biochem. 1990;22:465–469. [Google Scholar]