Abstract
Serine protease subtilase, found widely in both eukaryotes and prokaryotes, participates in various biological processes. However, how fungal subtilase regulates plant immunity is a major concern. Here, we identified a secreted fungal subtilase, UvPr1a, from the rice false smut (RFS) fungus Ustilaginoidea virens. We characterized UvPr1a as a virulence effector localized to the plant cytoplasm that inhibits plant cell death induced by Bax. Heterologous expression of UvPr1a in rice (Oryza sativa) enhanced plant susceptibility to rice pathogens. UvPr1a interacted with the important rice protein SUPPRESSOR OF G2 ALLELE OF skp1 (OsSGT1), a positive regulator of innate immunity against multiple rice pathogens, degrading OsSGT1 in a protease activity-dependent manner. Furthermore, host-induced gene silencing of UvPr1a compromised disease resistance of rice plants. Our work reveals a previously uncharacterized fungal virulence strategy in which a fungal pathogen secretes a subtilase to interfere with rice immunity through degradation of OsSGT1, thereby promoting infection. These genetic resources provide tools for introducing RFS resistance and further our understanding of plant–pathogen interactions.
A secreted fungal subtilase from Ustilaginoidea virens interacts with the positive regulator of innate immunity SUPPRESSOR OF G2 ALLELE OF skp1 to enhance susceptibility to multiple rice pathogens.
Introduction
Plants have evolved a complicated and precise innate immune system to resist the attack of microbial pathogens (Jones and Dangl, 2006). The plant immune system includes PAMP-triggered immunity (PTI) triggered by pathogen-associated molecular patterns (PAMPs) and effector-triggered immunity (ETI) triggered by effector proteins (Jones and Dangl, 2006; Dangl et al., 2013). PTI is an early defense response of plants against microbial pathogens. It is activated by pattern recognition receptors (PRR) located on plant cell membranes that specifically recognize conserved PAMPs (Couto and Zipfel, 2016). PTI can effectively prevent infection by a variety of phytopathogens (Dodds and Rathjen, 2010). However, in order to survive, phytopathogens secrete virulence effectors into host cells, block PRR’s recognition of PAMPs, interfere with host PTI, and inhibit basal defense, which result in successful infection of the host, via effector-triggered susceptibility (Jones and Dangl, 2006; Dou and Zhou, 2012; Dangl et al., 2013).
Subtilase as a type of serine protease is widely found in plants, fungi, bacteria, and parasites (Figueiredo et al., 2018). Most subtilases are composed of three conserved domains: peptidase S8 domain, PA domain, and inhibitor I9 domain. The peptidase S8 domain is a typical Asp/Ser/His catalytic domain which is present in all subtilases (Dodson and Wlodawer, 1998). The majority of subtilases contain an inhibitor I9 domain, which is autoinhibitory maintaining the inactive state of the zymogen and preventing access of the substrate to the active site, and also works as an intramolecular chaperone that is transiently required to assist in catalytic domain folding (Zhu et al., 1989; Baker et al., 1993; Li and Inouye, 1994; Huang et al., 1997; Bryan, 2002). The majority of subtilases also contain a protease-associated (PA) domain, which is responsible for protein–protein interactions and substrate recognition (Rautengarten et al., 2005; Schaller et al., 2012). Subtilase has a wide range of physiological functions, from nonspecific protein degradation functions to specific participation in the formation of polypeptide hormones, and it also participates in the degradation of proprotein to form an active mature protein (Figueiredo et al., 2018). Subtilases have been detected in the secretome of various pathogens and in infection situations (Figueiredo et al., 2018). A gene expression profiling study has shown that subtilases are involved in the wheat (Triticum aestivum)–Fusarium graminearum interaction (Pekkarinen and Jones, 2003; Walter et al., 2010). In the biocontrol entomopathogenic fungus Verticillium lecanii, a subtilase VlPr1, was found to be important in the biological control activity of some insect pests by aiding in degradation of insect cuticles (Yu et al., 2012). An extracellular subtilase, AsES, which acts as a plant defense response elicitor, can trigger a strong defense response in strawberry (Fragaria × ananassa) plants, manifested by an enhanced resistance against hemibiotrophic and necrotrophic fungal pathogens (Chalfoun et al., 2013). Recently, the conserved subtilisin-like Pr1 proteases, functioning in insect cuticle degradation, were found to possibly play important roles in the virulence of Metarhizium anisoplia (Gao et al., 2020). A fungal subtilase from Fusarium eumartii was identified as capable of degrading potato (Solanum tuberosum) pathogenesis-related (PR) proteins in vitro (Olivieri et al., 2002). However, the mechanisms by which pathogen subtilases target host defense proteins remain unexplored.
Rice false smut (RFS), caused by the ascomycete pathogen Ustilaginoidea virens, is one of the most devastating diseases affecting rice (Oryza sativa) worldwide (Sun et al., 2020). RFS not only reduces yields, but also threatens animal and human health by the production of cyclic peptide mycotoxins (Koiso et al., 1994; Sun et al., 2020). Genome analysis of U. virens showed that there are at least 628 secreted proteins, of which 193 are putative effectors (Zhang et al., 2014). Among them, a few virulence effectors which inhibit or trigger plant cell death (PCD) have been identified, including UvSCRE1, UvSCRE2 (UV_1261), and UvSGP1 (Fang et al., 2016, 2019; Zhang et al., 2020; Song et al., 2021). However, host targets of these effectors in U. virens are still unknown.
In this study, we identified a fungal virulence effector UvPr1a from U. virens, and uncovered a virulence strategy in which UvPr1a directly targets and degrades rice protein OsSGT1 (SUPPRESSOR OF G2 ALLELE OF skp1) to interfere host immunity and promote U. virens infection. This revealed a previously undescribed counter-defense mechanism evolved in a plant pathogen that can inactivate host defense responses.
Results
Identification of UvPr1a in U. virens
In our transcriptome data of U. virens infecting rice (Tang et al., 2021), we found that Uv8b_1567 which was annotated as a subtilisin-like protease was substantially upregulated in different infection processes. Functional domain analysis revealed that Uv8b_1567 contained signal peptide (SP), Inhibitor I9, and Peptidase S8 (Figure 1A). Blastp against NCBI NR database revealed Uv8b_1567 homologs to be widely distributed in Ascomycetes. Phylogenetic analysis revealed that Uv8b_1567 is closest to the subtilisin-like protease Pr1a in Metarhizium acridium, and thus Uv8b_1567 was named UvPr1a in this study (Supplemental Figure S1). We then examined the expression profile of UvPr1a at different developmental and infection stages of U. virens by real-time quantitative PCR (RT-qPCR), and found that UvPr1a expression was markedly increased during infection (Figure 1B), suggesting that UvPr1a might function in virulence of U. virens.
Figure 1.
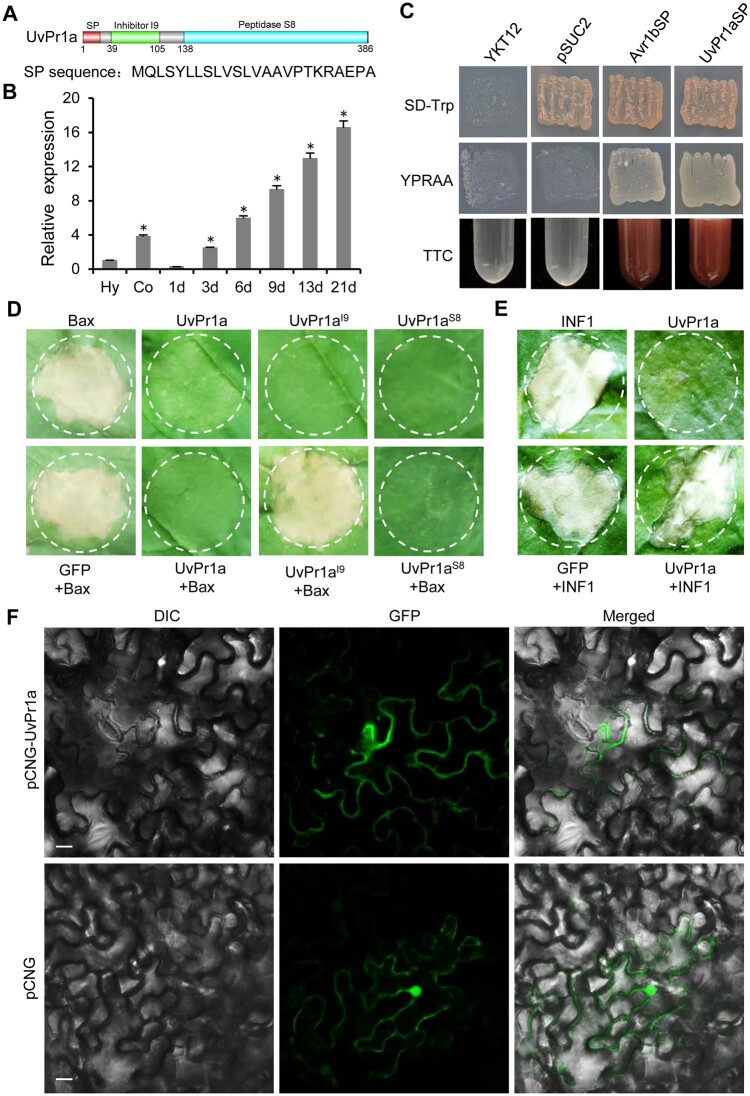
UvPr1a is a cytoplasmic effector. A, Predicted Pfam domain of UvPr1a. B, RT-qPCR detects the transcription level of UvPr1a relative to β-tubulin in conidia (Co) and hyphae (Hy) in PSB and at different infection stages (1–21 dpi) of U. virens. Error bars represent the standard deviation. Asterisks represent significant differences from Hy using the test of LSD at P < 0.05. C, Yeast secretion assay of UvPr1a signal peptide. Yeast strain YTK12 was transformed with the empty vector pSUC2 (negative control), pSUC2-UvPr1aSP, or pSUC2-Avr1SP (positive control) and tested for growth on SD–Trp or YPRAA medium and for invertase activity with a colorimetric TTC assay. D, Transient expression of UvPr1a, UvPr1aI9, and UvPr1aS8 suppresses PCD triggered by Bax in N. benthamiana leaves. Representative leaves were photographed 5 days after infiltration. E, Transient expression of UvPr1a did not suppresses PCD triggered INF1 in N. benthamiana leaves. Representative leaves were photographed 5 days after infiltration. F, UvPr1a-GFP localization in N. benthamiana leaves. DIC, differential interference contrast; GFP, green fluorescent protein. Scale bar = 20 μm.
UvPr1a is a cytosolic effector that suppresses Bax-induced cell death in the plant
To determine the functionality of the SP of UvPr1a, we conducted a secretion assay in yeast (Saccharomyces cerevisiae). The resulting transformants accumulating UvPr1aSP or the positive control Avr1bSP grew on YPRAA medium containing raffinose as the sole carbon source (Figure 1C). The transformants also secreted invertase, as detected by formation of the red product 1,3,5-triphenylformazan (TPF) (Figure 1C). In contrast, transformants harboring the empty vector failed to grow on YPRAA medium and did not produce the characteristic change in color during the 2,3,5-triphenyltetrazolium chloride (TTC) assay (Figure 1C), and these results indicated that the SP of UvPr1a acted as a functional secretion signal, confirming that UvPr1a is a secreted protein.
To evaluate the role of UvPr1a as an effector in the plant immune response, we tested the effects of expressing UvPr1a on PCD induced by Bax (a mouse proapoptotic protein) and INF1 (a PAMP from Phytophthora infestans). Co-expressing UvPr1a and Bax suppressed the PCD normally induced by Bax in Nicotiana benthamiana leaves (Figure 1D). However, UvPr1a did not prevent INF1-induced cell death (Figure 1E). To further determine the region of UvPr1a that mediates the inhibition of Bax-induced cell death, we generated constructs expressing two UvPr1a truncations, UvPr1aI9 (Inhibitor I9 domain) or UvPr1aS8 (Peptidase S8 domain), which were co-infiltrated them with Bax in N. benthamiana. UvPr1aS8 showed a comparable ability to suppress Bax-induced cell death as the full-length UvPr1a (Figure 1D). These results suggest that the Peptidase S8 domain of UvPr1a can suppress the PCD induced by Bax in the plant.
To examine UvPr1a localization in plant cells, we examined the subcellular localization of an UvPr1a-GFP (green fluorescent protein) fusion protein whose sequence was driven by the cauliflower mosaic virus 35S promoter and expressed in N. benthamiana leaves. Through fluorescence microscopy, GFP fluorescence was detected in the cytoplasm of plant cells, whereas the control vector encoding untagged GFP showed GFP signals in the nucleus and plasma membrane (Figure 1F), suggesting that UvPr1a is localized to the cytoplasm in plants.
UvPr1a is required for development and virulence of U. virens
From the UvPr1a knockout library, two independent deletion mutants (ΔUvPr1a-9 and ΔUvPr1a-24) identified by PCR analyses (Supplemental Figure S2) and showing similar phenotypes were obtained. For complementation of the UvPr1a deletion mutant, the complementary vector p3300neoIII-UvPr1a was transformed into ΔUvPr1a-24, and the complementation transformant CΔUvPr1a-24 was confirmed by PCR analyses (Supplemental Figure S2). In the CΔUvPr1a-24 mutant, GFP signals were enriched in the cytoplasm of vegetative hyphae or conidia under confocal microscopy (Supplemental Figure S3).
To determine whether the function of UvPr1a was associated with the development of U. virens, we first compared the mycelial growth rate between the mutants and wild-type on potato sucrose agar (PSA). The results showed that the mycelial growth rate of ΔUvPr1a mutants was slightly reduced compared to that of HWD-2 and CΔUvPr1a-24 (Figure 2, A and B). We then compared conidial production among HWD-2, ΔUvPr1a mutants, and complementation strains. Deletion of UvPr1a significantly increased the conidial production by U. virens (Figure 2C). Pathogenicity assays were performed with the HWD-2, ΔUvPr1a mutants, and complementation strains inoculated onto a susceptible rice cultivar, Wanxian-98. At 21-day postinoculation (dpi), ΔUvPr1a mutants exhibited significantly reduced virulence compared to the wild-type strain HWD-2 or the complementation strain (Figure 2, D and E). These results suggested that UvPr1a was required for full virulence of U. virens.
Figure 2.
UvPr1a is a key virulence effector. A, Colony morphology of U. virens wild-type HWD-2, ΔUvPr1a-9, ΔUvPr1a-24, CΔUvPr1a, CΔUvPr1aΔS8, and CΔUvPr1aΔI9 strains on PSA after 14 days of darkness at 28°C. B, Colony diameters of mutant strains on PSA after 14 days at 28°C. C, Conidial production of mutant strains grown in PSB medium at 180 rpm for 7 days. D, Virulence assays of mutant strains on rice spikelets at 21 dpi. E, Mean number of rice smut balls per panicle. Data were collected from three independent experiments for each treatment. Error bars represent standard deviation, and asterisks represent significant difference from HWD-2 using the test of LSD at P = 0.05.
To evaluate the domain roles of Inhibitor I9 and Peptidase S8 in development and virulence of U. virens, truncated sequences UvPr1aΔI9 (lack of Inhibitor I9) or UvPr1aΔS8 (lack of Peptidase S8) were then transformed separately into ΔUvPr1a-24. Phenotype analysis revealed that Inhibitor I9 domain (UvPr1aI9) of UvPr1a was essential for mycelial growth, conidiation, and virulence of U. virens (Figure 2, A–E).
Heterologous expression of UvPr1a in rice increases susceptibility to rice pathogens
We generated transgenic rice plants overexpressing UvPr1a (Supplemental Figure S4), and 35S-UvPr1a transgenic lines grew similarly to the wild-type Nipponbare (Nip) rice plants (Figure 3A). In pathogenicity tests, 35S-UvPr1a transgenic rice plants inoculated with U. virens produced significantly more rice smut balls than Nip, suggesting increased susceptibility to U. virens (Figure 3B). We then evaluated the 35S-UvPr1a transgenic rice plants for resistance against other three rice pathogens, the blast fungus Magnaporthe oryzae, the bacterial blight pathogen Xanthomonas oryzae pv. oryzae (Xoo), and the sheath blight pathogen Rhizoctonia solani. Twenty-one days after inoculation with Xoo PXO99A, 35S-UvPr1a transgenic lines showed more disease, and lesion lengths on 35S-UvPr1a transgenic rice plants were more than 7 times longer than those on Nip (Figure 3C). When spot-inoculated with M. oryzae Guy11, larger disease lesions were found on 35S-UvPr1a transgenic rice plants at 5 dpi (Figure 3D). Likewise, leaf lesion areas caused by R. solani HG81 on 35S-UvPr1a transgenic rice plants were larger than those on Nip (Figure 3E). Overexpression of UvPr1a in rice plants enhanced host susceptibility to the four rice pathogens, suggesting that the UvPr1a protein can suppress the immune response.
Figure 3.
Heterologous expression of UvPr1a in rice increases susceptibility to rice pathogens. A, Representative images of 35S-UvPr1a transgenic rice plants and the wild-type Nip rice plants at the mature stage following growth in field conditions. B, Left: Resistance assays of 35S-UvPr1a transgenic rice plants and the wild-type Nip rice plants to U. virens strain HWD-2 infection at 25 dpi. Right: Mean number of rice smut balls measured in resistance assays. C, Left: Disease symptoms at 21 dpi of 35S-UvPr1a transgenic rice plants and the wild-type Nip rice plants after inoculation with Xoo PXO99A. Right: Mean lesion lengths at 21 dpi on the leaves of 35S-UvPr1a transgenic rice plants and the wild-type Nip rice plants after inoculation with Xoo PXO99A. D, Left: Leaves of 35S-UvPr1a transgenic rice plants and the wild-type Nip rice plants at 5 dpi following spot-inoculation with a spore suspension of M. oryzae Guy11. Right: Relative fungal biomass was determined using qPCR for the M. oryzae Pot2 gene normalized to rice OsUBQ1. E, Left: Disease symptoms at 3 dpi of 35S-UvPr1a transgenic rice plants and the wild-type Nip rice plants after inoculation with R. solani HG81. Right: Leaf lesion area of 35S-UvPr1a transgenic rice plants and the wild-type Nip rice plants after inoculation with R. solani HG81. All inoculation experiments were repeated 3 times. Asterisks represent significant differences using the test of LSD at P < 0.05 between Nip and 35S-UvPr1a transgenic rice plants, and standard errors are shown in bar graphs.
UvPr1a interacts with rice protein OsSGT1
To further characterize the function of UvPr1a during U. virens infection, we performed a yeast two-hybrid (Y2H) screen with UvPr1a as bait against a cDNA library constructed from RNA extracted from U. virens-infected rice spikelets. After screening of this library, 47 putative UvPr1a-interacting proteins were identified (Supplemental Table S1). We found that the rice protein OsSGT1 (LOC_Os01g43540) was a frequently screened target of UvPr1a and confirmed the interaction between UvPr1a and OsSGT1 by Y2H assay (Figure 4A). We then attempted to delineate the interaction interface by testing two UvPr1a fragments (UvPr1aI9 and UvPr1aS8) for interaction with OsSGT1. UvPr1aI9 and UvPr1aS8 were both found to interact with OsSGT1 (Figure 4A). We attempted to delineate the interaction interface by testing three OsSGT1 fragments (OsSGT1TPR, OsSGT1CS, and OsSGT1SGS) for interaction with UvPr1a, and all three OsSGT1 fragments were found to interact with UvPr1a (Figure 4A). We performed in vitro GST pull-down assays with recombinant OsSGT1-His and UvPr1a-GST proteins purified from Escherichia coli. OsSGT1-His was detected in protein samples pulled down with UvPr1a-GST loaded onto glutathione beads (Figure 4B), but was not found in GST pull-down fractions (Supplemental Figure S5A), suggesting that UvPr1a-GST and OsSGT1-His interacted in vitro. To validate the interaction in vivo, we next performed co-immunoprecipitation (Co-IP) assays on N. benthamiana leaves transiently co-expressing with OsSGT1-Flag and UvPr1a-GFP, and found that UvPr1a was immunoprecipitated by OsSGT1 (Figure 4C), whereas GFP was not immunoprecipitated by OsSGT1-Flag (Supplemental Figure S5B), suggesting that UvPr1a-GFP interacted with OsSGT1-Flag. Finally, we transiently co-infiltrated N. benthamiana leaves with UvPr1a-nYFP and OsSGT1-cYFP fusion constructs for bimolecular fluorescence complementation (BiFC) analysis, and observed YFP fluorescence in the nucleus and cytoplasm of N. benthamiana leaf epidermal cells (Figure 4D). These results demonstrated that UvPr1a interacts with OsSGT1 in the cytoplasm of plant cells.
Figure 4.
UvPr1a physically interacts with rice OsSGT1 in vitro and in vivo. A, Y2H analysis of the interaction between UvPr1a and OsSGT1. SD-3, SD–Trp–Leu–His; BD, pGBKT7; AD, pGADT7. BD-53 + AD-T served as the positive control. BD-UvPr1a + AD or BD + AD-OsSGT1 served as the negative control. B, GST pull-down assay showing the interaction between OsSGT1-His and UvPr1a-GST. Recombinant UvPr1a-GST bound resin was incubated with E. coli crude extracts containing OsSGT1-His and analyzed by immunoblotting. C, Co-IP assay showing that UvPr1a interacts with OsSGT1 in vivo. Co-IP assay was performed on protein extracts from N. benthamiana leaves co-expressed with OsSGT1-Flag and UvPr1a-GFP. D, BiFC assays of the interaction between UvPr1a and OsSGT1. Nicotiana benthamiana leaves were co-infiltrated with UvPr1a-nYFP and OsSGT1-cYFP constructs. YFP signals were observed 2 days after infiltration. Co-infiltration with UvPr1a-nYFP and cYFP or nYFP and OsSGT1-cYFP constructs were used as negative control. Scale bar = 20 µm.
OsSGT1 positively regulates resistance to multiple diseases in rice
To explore the function of OsSGT1 in resistance against rice pathogens, we generated overexpressing OsSGT1 transgenic rice plants (OsSGT1-OE) and OsSGT1 knockout mutants (ossgt1) rice plants by Clustered Regularly Interspaced Short Palindromic Repeats-associated clause Cas9 (CRISPR/Cas9) (Supplemental Figure S6, A–D). Morphology of these OsSGT1-OE and ossgt1 rice plants was similar to that of the Nip rice plants (Figure 5A;Supplemental Table S2). After inoculation with mycelial/spore suspension of U. virens HWD-2, OsSGT1-OE transgenic rice plants produced fewer smut balls than the Nip, whereas the ossgt1 rice produced more smut balls than the Nip (Figure 5B). After inoculation with Xoo PXO99A using the scissor-clipping method, lesions were ∼80% shorter on the OsSGT1-OE rice plants than the Nip rice plants at 21 dpi, and lesion lengths on ossgt1 rice plants were much longer than on the Nip plants (Figure 5C). Following spot inoculation with compatible M. oryzae Guy11, as expected, weaker symptoms were observed on OsSGT1-OE rice plants at 5 dpi (Figure 5D). Likewise, ossgt1 rice plants were more susceptible to infection by R. solani HG81, whereas OsSGT1-OE rice plants were more resistant to infection by R. solani as observed at 3 dpi (Figure 5E). These results demonstrated that OsSGT1 positively regulates resistance to multiple diseases in rice.
Figure 5.
OsSGT1 positively regulates rice disease resistance against multiple rice pathogens. A, Representative images of Nip, OsSGT1-OE, and ossgt1 rice plants at the mature stage following growth under field conditions. B, Left: Resistance assays of Nip, OsSGT1-OE and ossgt1 rice plants against infection by U. virens strain HWD-2 at 25 dpi. Right: Mean number of rice smut balls seen in resistance assays. C, Left: Leaf lesions on Nip, OsSGT1-OE, and ossgt1 rice plants after inoculated with Xoo PXO99 at 21 dpi. Right: Lesion lengths on rice leaves at 21 dpi after inoculation with Xoo PXO99A. D, Left: Disease symptoms on the leaves of Nip, OsSGT1-OE, and ossgt1 rice plants. Leaves were spot inoculated with a spore suspension of M. oryzae Guy11 and photographed at 5 dpi. Right: Relative fungal biomass was determined using qPCR for M. oryzae Pot2 and normalized to rice OsUBQ1. E, Left: Disease symptoms at 3 dpi of Nip, OsSGT1-OE, and ossgt1 rice plants after inoculation with R. solani HG81. Right: Leaf lesions on Nip, OsSGT1-OE and ossgt1 rice plants after inoculation with R. solani HG81 at 3 dpi. All inoculation experiments were repeated 3 times. Asterisks represent significant differences using the test of LSD at P < 0.05, and standard errors are shown in bar graphs.
UvPr1a mediates protein degradation of OsSGT1
We performed experiments in vitro and in vivo to examine whether UvPr1a suppresses OsSGT1-mediated plant immunity by degradation of OsSGT1 protein. Extensive expression of OsSGT1 could induce PCD, and when OsSGT1 was co-expressed with UvPr1a, PCD was substantially attenuated (Figure 6A). As a negative control, expression of the GFP (empty vector pVX) did not abolish the induction of PCD by OsSGT1 (Figure 6A), suggesting that UvPr1a have a specific role in suppressing OsSGT1-triggered cell death. To explore whether UvPr1a directly degrades OsSGT1, we performed an in vitro protein degradation experiment. After co-incubation of OsSGT1 protein with different concentrations of UvPr1a, we found that the degradation level of OsSGT1 was increased along with increase abundance of UvPr1a (Figure 6B). The rice mitogen-activated protein kinase OsMAPK33 (LOC_Os02g05480) which was set as a negative control was confirmed to be an interaction protein of UvPr1a by Y2H, Co-IP, and GST pull-down assays (Supplemental Figure S7, A–C). To determine whether UvPr1a degrades OsMAPK33, we performed an in vitro protein degradation experiment and the result showed that UvPr1a did not degrade OsMAPK33 (Supplemental Figure S7D), suggesting that the degradation of OsSGT1 by UvPr1a is specific. To examine whether UvPr1a degrades OsSGT1 in a protease activity-dependent manner, we performed an in vitro degradation experiment with co-incubation of OsSGT1 and UvPr1a after denaturation. The results revealed that OsSGT1 was not degraded (Supplemental Figure S8A). We have performed an in vitro degradation experiment with co-incubation of OsSGT1 and UvPr1aI9 or UvPr1aS8. The results revealed that UvPr1aS8 could degrade OsSGT1, whereas UvPr1aI9 did not degrade OsSGT1 (Supplemental Figure S8B). To further confirm that UvPr1a can degrade OsSGT1 in plants, we first co-expressed UvPr1a-GFP and OsSGT1-Flag in N. benthamiana leaves. Compared with the co-expression of OsSGT1 and GFP, the protein amount of OsSGT1-Flag in the co-expression of UvPr1a-GFP and OsSGT1-Flag was considerably reduced, indicating that UvPr1a could degrade OsSGT1 in N. benthamiana leaves (Figure 6C). We also used an SGT1-specific antibody to detect the amount of OsSGT1 protein in Nip and in 35S-UvPr1a transgenic rice plants, and the results showed that the content of OsSGT1 protein in 35S-UvPr1a transgenic rice was markedly lower than that in Nip rice plants (Figure 6D). Taken together, these results showed that the serine protease UvPr1a degrades OsSGT1 in vivo and in vitro in a protease activity-dependent manner.
Figure 6.
UvPr1a mediates protein degradation of OsSGT1. A, Transient expression of UvPr1a suppresses PCD triggered by OsSGT1 in N. benthamiana leaves. pVX-UvPr1 and pVX-OsSGT1 fusion constructs were co-infiltrated in N. benthamiana leaves. Representative leaves were photographed 5 days after infiltration. B, UvPr1a degrades OsSGT1 protein in vitro, as determined by immunoblotting using 5 µL of purified OsSGT1-His, with 1, 2, 5, and 10 µL of recombinant UvPr1a-GST or GST incubated at 30°C for 15 min. The reaction products were analyzed by immunoblotting with anti-His and anti-GST antibodies. C, OsSGT1-Flag co-expression with UvPr1a-GFP or GFP in N. benthamiana leaves. The relative protein level of OsSGT1-Flag was detected by Western blotting. D, Relative protein levels of OsSGT1 in Nip and 35S-UvPr1a transgenic rice plants as detected with anti-SGT1.
Engineering RFS resistance rice via HIGS of UvPr1a
We employed host-induced gene silencing (HIGS) to generate transgenic Nip plants expressing an RNA interference (RNAi) construct against the UvPr1a transcript (Supplemental Figure S9, A–C). Notably, UvPr1aHIGS transgenic rice plants maintained agronomic traits that were indistinguishable from those of the untransformed Nip plants (Supplemental Table S3). After inoculation with U. virens at 25 dpi, an average of 11 smut balls per panicle was scored on the spikelets of Nip plants. In contrast, two independent UvPr1aHIGS transformants displayed a strong resistance to RFS, with an average of one to three smut balls per panicle (Figure 7, A and B).
Figure 7.
HIGS of UvPr1a increases rice resistance to RFS. A, Resistance assays of T2 and T3 HIGS transgenic rice plants harboring the UvPr1aHIGS construct against U. virens at 25 dpi. B, Mean number of rice smut balls in T2 and T3 transgenic rice plants from resistance assays. The experiment was repeated 3 times, and 30 rice panicles were used per inoculation. Error bars represent the standard deviation. C, Relative UvPr1a transcript levels of T3 transgenic lines at 6 dpi. Levels of the U. virens β-tubulin gene were used for normalization in RT-qPCR. The experiment was repeated 3 times and error bars represent the standard deviation. D, Visualization of siRNAs targeting UvPr1a in infected transgenic rice spikelets at 6 dpi by FISH using a specific probe. An, anther; hy, U. virens hyphae. Scale bar = 20 μm. Asterisks indicate statistically significant differences compared to Nip using the test of LSD at P < 0.05.
To confirm that the resistance to RFS seen in infected UvPr1aHIGS transgenic rice plants was caused by in planta-mediated silencing of UvPr1a, we collected infected spikelets from T3 transgenic rice plants at 6 dpi and quantified UvPr1a transcript levels by RT-qPCR. Indeed, relative UvPr1a transcript levels were much lower in the UvPr1aHIGS transgenic rice plants compared to Nip (Figure 7C).
To ascertain that silencing of UvPr1a in infected U. virens was mediated by small interfering RNAs (siRNAs) generated by UvPr1aHIGS transgenic rice plants, we sequenced small RNAs in the T3 UvPr1aHIGS transgenic rice plants. The sequencing data indicated that siRNAs mapping to UvPr1a were highly abundant in UvPr1aHIGS transgenic rice plants, accounting for 0.42% (L2 line) or 0.51% (L4 line) of all small RNAs detected in these transgenic rice plants. The siRNAs matching the UvPr1a transcript had a size distribution of 18–30 nucleotides (nts), with 21-nt siRNAs being the most abundant (Supplemental Figure S9D).
In fluorescence in situ hybridization (FISH) assays, fluorescence signal was observed both in rice flower organ tissue and infection hyphae of U. virens in the infected UvPr1aHIGS transgenic rice plants at 6 dpi, whereas no fluorescence signal was detected in the Nip plants (Figure 7D). This supported the notion that the UvPr1aHIGS vector produced effective siRNAs against UvPr1a in the UvPr1aHIGS transgenic rice plants that were then translocated to fungal cells during infection to reduce UvPr1a transcript levels in invading U. virens hyphae. These results demonstrated that silencing of UvPr1a in transgenic rice plants could promote rice resistance to U. virens.
Discussion
Previous studies revealed that fungal subtilases play an essential role in pathogenicity during pathogenic infection. In M. oryzae, knockout of the subtilisin gene SPM1 substantially reduces conidiation, pathogenicity, and autophagy (Donofrio et al., 2006). The insect parasitic fungus, M. anisoplia, contains 21 subtilisin genes, and Pr1C, Pr1G, Pr1A2, Pr1B1, and Pr1B2 were involved in pathogenicity of the pathogen to insects (Gao et al., 2020). In F. graminearum, subtilisin gene FgPrb1 regulates pathogenicity, stress response, toxin synthesis, autophagy, and lipid metabolism (Xu et al., 2020). However, subtilisin proteins have not been reported as virulence effectors during pathogenic infection, and the molecular mechanisms of pathogen subtilases in the infection process are still unknown. In this study, we found that a subtilisin protein UvPr1a which was secreted by wild-type HWD-2 of U. virens and identified as a virulence effector, interacted with the immune regulator OsSGT1, which positively regulated resistance to multiple diseases in rice. The UvPr1a directly degraded OsSGT1, resulting in inhibition of plant immunity, thereby promoting pathogen infection. However, after infection by the ΔUvPr1a mutants, the OsSGT1 protein in rice was expressed normally, and the plants showed immune responses and resistance to pathogens (Figure 8). The RAR1–SGT1–Heat shock protein 90 (HSP90) chaperone complex is involved in the stabilization of NLR (Nucleotide-binding domain and leucine-rich repeat containing receptor) protein (Shirasu, 2009). NLR proteins such as MLA1, MLA6, and Rx, as well as RPS5, were shown to require RAR1 for steady-state accumulation. Similarly, OsSGT1 interacts with OsRAR1 and OsHSP90 in rice, and they positively regulate basal disease resistance against Xoo and M. oryzae (Wang et al., 2008; Yan et al., 2012). Overexpression of OsSGT1 might enhance the stabilization of RAR1–SGT1–HSP90 chaperone complex in rice. Targeting of the major host immune regulator SGT1 for degradation, thereby interfering with the plant immune response, represents a previously uncharacterized mechanism for pathogen effectors.
Figure 8.
A working model illustrating how UvPr1a degrades OsSGT1 to suppress rice immunity during U. virens infection. During infection, the U. virens effector UvPr1a is secreted by HWD-2 and translocated into host cells, and then physically interacts with OsSGT1. UvPr1a directly degraded OsSGT1, thereby inhibiting plant immunity and showed susceptibility to pathogens. However, in the ΔUvPr1a mutants, the OsSGT1 protein was expressed normally, and the rice plants had normal immune responses and showed resistance to pathogens.
In general, many effectors are highly expressed in the early stages during pathogen infection. However, in this study, we found that the expression of UvPr1a was increased gradually and continuously after infection, and this is consistent with the results of Sreedhar et al. (1999), who reported that expression of a secreted subtilisin-like proteinase Mp1 of Magnaporthe poae in host infected roots corresponded with increasing severity of disease symptoms. Since UvPr1a was also found to be involved in hyphal growth and conidiation of U. virens, it is speculated that the high expression level of UvPr1a in late infection stages might be related to fungal development, smut ball formation, or toxin synthesis by U. virens.
The plant apoplast is the extracellular compartment serving as a major battlefield for plant–pathogen interactions (Du et al., 2016). The plant apoplast is a harsh environment which is full of digestive enzymes including proteases. Several plant proteases are highly upregulated during pathogen infection and have long been tied to plant immunity (Ramírez et al., 2013; Duan et al., 2016; Fan et al., 2016). For example, the subtilase P69 was located in the vacuole and in the intercellular spaces of leaf parenchyma cells of tomato. The P69 subtilase complex consists of a gene family with six closely related members, named P69A to P69F. P69A, P69D, and P69E are associated with developmental processes, whereas P69B and P69C are shown to behave as PR PR-P69 protease, exhibit a high similarity to each other and are involved in protein degradation against pathogen infection (Tornero et al., 1997; Jordá et al., 1999; Tian et al., 2004). By analyzing the subtilase of pathogens and plants, we found that they are highly conserved in these domains (peptidase S8 domain, PA domain, and inhibitor I9 domain). In the process of pathogenic infection, an extremely complex and precise “attack vs defense” arms race is staged. We speculate that the pathogen secretes subtilases as virulence factors, and because of their homology with plant subtilases, plants do not recognize them as foreign. Hence pathogen subtilases are not degraded, and lead to increased infection.
Previous studies showed that the Inhibitor I9 domain is involved in regulating the folding and activity of the zymogen, and that the Peptidase S8 domain is responsible for the peptidase function. The Inhibitor I9 domain keeps peptidase inactive by preventing the substrate from entering the active center of the enzyme and binding to the substrate-binding site (Figueiredo et al., 2018). Complementation of the Inhibitor I9 domain alone could restore the defects in mycelial growth, conidiation and pathogenicity of UvPr1a knockout mutants, indicating that the Inhibitor I9 domain is indispensable in the pathogenesis of U. virens. It was found that Inhibitor I9 domain did not inhibit Bax-induced PCD, whereas the Peptidase S8 domain could inhibit PCD, suggesting that Inhibitor I9 domain might not directly regulate Bax-related immune pathways. In this study, eight subtilisin proteins were found in the genome of U. virens, six of which contained SPs (Supplemental Figure S10A). Among them, we found that deletion of Uv8b_44, Uv8b_1567 (UvPr1a) and Uv8b_8155 affected the mycelial growth, conidiation, and pathogenicity of U. virens (Supplemental Figure S10, B–E). Interestingly, these three subtilisin proteins all contained an Inhibitor I9 domain whereas the other five did not. These suggest that every Inhibitor I9 domain from the three subtilisin proteins is important for U. virens–rice interaction. In previous studies, chaperoning function of the family of I9 inhibitors was confirmed for subtilisin proteins in mammals and plants (Shinde and Thomas, 2011; Meyer et al., 2016). Deletion of I9 inhibitor in tomato subtilase SBT3 led to defect in secretion of the active protease to the apoplast (Meyer et al., 2016). We speculate that the Inhibitor I9 domain plays an important role in the process of effector secretion into plant cells, while the Peptidase S8 domain functions in regulating plant immunity after effector entry into the host. In this study, we found that lack of Peptidase S8 in UvPr1a did not cause reduced virulence of U. virens. In the genome of U. virens, there are many Peptidase S8-containing proteins. Deletion of each one of these proteins might not affect plant immunity since some Peptidase S8-containing proteins might have similar functions. Hence, we hypothesized that the Inhibitor I9 domain is required for degradation function in planta of the Peptidase S8 domain in UvPr1a or other Peptidase S8-containing proteins.
In plant defense, many functional plant resistance (R)-genes have been identified and cloned as well as components of the downstream defense signaling network underlying R-protein, basal, and nonhost resistance and those of importance to the induction of systemic responses (Hammond-Kosack and Parker, 2003; Jones and Dangl, 2006). Key defense signaling components include RAR1 (required for Mla12 resistance) and SGT1 (suppressor of G2 allele of skp1). These proteins are involved in triggering various responses including production of reactive oxygen species, localized PCD manifesting as a hypersensitive response, and the accumulation of salicylic acid and antimicrobial PR proteins (Hammond-Kosack and Parker, 2003). RAR1 and SGT1 are required for the function of many R-proteins in both monocotyledonous and dicotyledonous plants (Liu et al., 2002; Muskett et al., 2002; Azevedo et al., 2002; Tornero et al., 2002; Tor et al., 2002; Shirasu and Schulze-Lefert, 2003). In Arabidopsis, SGT1b has been shown to have a role in defense (Azevedo et al., 2006). HSP90 is able to interact with SGT1 and RAR1 in barley and N. benthamiana as well as SGT1a, SGT1b, and RAR1 in Arabidopsis to regulate R-protein stability (Hubert et al., 2003; Liu et al., 2004). Upstream of the SGT1/RAR1/HSP90 complex is the enhanced disease susceptibility 1 (EDS1) protein which has some identity to lipases. In Arabidopsis, EDS1 is required for the function of several R-proteins (Aarts et al., 1998), basal resistance to virulent isolates of several pathogens (Parker et al., 1996; Aarts et al., 1998), and nonhost resistance to two biotrophic pathogens of Brassica oleracea (Parker et al., 1996). OsSGT1 interacts with OsRAR1 and OsHSP90 in rice, and positively regulates disease resistance against Xoo and M. oryzae (Wang et al., 2008; Yan et al., 2012). Nucleotide-binding domain and leucine-rich repeat-containing proteins (NLRs/NB-LRR) function as sensors that perceive pathogen molecules and activate immunity. In plants, the accumulation and activation of NLRs is regulated by SGT1 (Azevedo et al., 2006). In this study, we found that UvPr1a interacted with OsSGT1, and UvPr1a directly cleaved OsSGT1 to interfere plant immunity and promote infection. How OsSGT1 regulates rice resistance against various pathogens, and whether OsSGT1 also functions as a chaperone involved in the stabilization of NLR protein in rice need further study.
The development of durable and broad-spectrum resistance is an economical and sustainable approach to control crop diseases for agricultural production. Recently, HIGS, an RNAi-based approach, was developed as an alternative strategy to control fungal diseases, where siRNAs targeting selected genes of the invading pathogen are produced by transgenic host plants to silence fungal genes during infection (Nowara et al., 2010). For example, engineering Fusarium head blight resistance wheat via HIGS of SGE1, FGP1, and STE12 genes of F. graminearum (Wang et al., 2020). In recent years, HIGS technology has been widely used in cotton, wheat, rice, and other crops for cultivating green, durable, and stable resistant materials because of high silencing efficacy (Xu et al., 2018; Dou et al., 2020; Li et al., 2022; Chen et al., 2022). Our study now adds HIGS against UvPr1a as an effective strategy to reduce severity of U. virens in rice, and provides a start for the development of stable transgenic plants using a HIGS-based strategy to enhance rice resistance to U. virens.
In summary, we identified an important secreted virulence effector, UvPr1a, which interacts with rice OsSGT1. UvPr1a directly cleaves OsSGT1 to interfere with plant immunity and promote infection. More importantly, stable rice transgenic lines expressing UvPr1a-silencing RNAs showed increased resistance to U. virens, pointing to another strategy for protecting rice crops.
Materials and methods
Fungal strains and inoculation
The fungal strains U. virens HWD-2, M. oryzae Guy11, X. oryzae pv. oryzae (Xoo) PXO99A, and Rhizoctonia solani HG81 were used for rice (O. sativa) inoculations. The U. virens HWD-2 and transformed strains were cultured in potato sucrose broth (PSB) at 28°C for 7 days with shaking at 180 rpm. The PSB cultures were then homogenized with a blender to prepare mycelial/spore suspensions. Rice plants were inoculated with U. virens HWD-2 at booting stage with 2 mL mycelial/spore suspensions (1 × 106 conidia mL−1) using a syringe, and then placed in a greenhouse with a relative humidity of 95% ± 5% and a temperature of 28°C ± 2°C. The number of false smut balls was counted at 25 dpi (Chen et al., 2020). For spot-inoculations, 4-week-old leaves were punctured with a needle, and a droplet of 10-μL M. oryzae Guy11 conidial suspensions (5 × 104 conidia mL−1) in 0.25% (v/v) Tween-20 was placed at each wound site. Rice plants were inoculated with Xoo PXO99A at the booting stage by the leaf clipping method, and lesion lengths were scored at 21 dpi (Chen et al., 2021a). Four-week-old rice leaves were inoculated with R. solani HG81 and incubated at 25°C for 3 days. The length and width of lesions were measured using ImageJ software (http://rsbweb.nih.gov/ij/) and lesions areas were calculated. All inoculation experiments were repeated 3 times.
RNA extraction and RT-qPCR
Total RNA was extracted using TRIzol reagent (Vazyme Biotech, Nanjing, China). First-strand cDNA synthesis was carried out with a cDNA Synthesis SuperMix (TransGen Biotech, China). RT-qPCR was performed with TransStart Tip Green qPCR SuperMix (TransGen Biotech, Beijing, China). Transcript levels were normalized using the U. virens β-tubulin gene (Uv8b_900) or rice ubiquitin gene (OsUBQ1). The experiment was repeated 3 times. For each replicate, at least three spikelets were used per treatment.
UvPr1a secretion assay
The yeast invertase secretion assay was conducted, using the SP trap vector pSUC2 as previously described (Jacobs et al., 1997). Briefly, the SP sequence of UvPr1a was cloned into the vector pSUC2. The pSUC2, pSUC2-UvPr1aSP, and pSUC2-Avr1bSP (positive control) fusion vectors were transformed into yeast (S. cerevisiae) strain YTK12. The transformants were grown on SD-Trp (Clontech, Mountain View, CA, USA) or YPRAA medium (1% [w/v] yeast extract, 2% [w/v] peptone, 2% [w/v] raffinose, 2 μg L−1 antimycin A, 2% [w/v] agar) for invertase secretion assays. Invertase enzymatic activity was detected by the reduction of TTC to the insoluble red precipitate TPF.
Agroinfiltration assays
For agroinfiltration assays, Agrobacterium tumefaciens strain EHA105 cells were re-suspended in infiltration buffer (10-mM MgCl2, 0.5-mM MES, 0.2-mM acetylacetone) to a final optical density at 600 nm (OD600) of 0.5 and infiltrated into 4-week-old N. benthamiana leaves. To assay suppression of Bax- or INF1-induced programmed cell death (PCD), N. benthamiana leaves were first infiltrated with A. tumefaciens cells carrying the vector pVX-UvPr1a or pVX followed 1 d later by infiltration of the same areas with A. tumefaciens cells carrying the pVX-Bax or pVX-INF1 vector. PCD was assessed at 5 dpi. For subcellular localization of UvPr1a-GFP in N. benthamiana, the coding sequences of UvPr1a was cloned into the vector pCN-GFP. pCN-UvPr1a-GFP and pCN-GFP vectors were then introduced into Agrobacterium strain EHA105. Agrobacterium tumefaciens cells harboring each construct were resuspended in infiltration buffer to a final OD600 of 1.0 and infiltrated into N. benthamiana leaves. After 2 days, the fluorescence signal of the infiltrated areas was examined with a Zeiss LSM 510 Meta confocal microscope (Carl Zeiss, Jena, Germany). GFP expression was detected after excitation with a 488-nm wavelength laser, and emissions were collected between 500 and 540 nm.
Gene deletion and complementation
To generate UvPr1a gene deletion mutants, ∼1.5 kb of upstream and downstream flanking sequences of the gene were ligated with pGKO to generate the final deletion vector pGKO-UvPr1a. The construct was inserted into A. tumefaciens strain EHA105, and then transformed into conidia of HWD-2 (Chen et al., 2021b). Hygromycin-resistant transformants were isolated and subsequently screened by PCR with primers P1/P2 listed in Supplemental Table S4. For complementation assays, a native promoter region and a full-length UvPr1a, segmental complementary region sequence UvPr1aI9, and UvPr1aS8 were cloned into vector pNeo3300III. The EHA105 strain with the complementary vectors was transformed through A. tumefaciens-mediated transformation by co-culture with conidia of the ΔUvPr1a-24 mutant. Transformants were selected on PSA supplemented with 500 μg mL−1 antibiotic G418, and screened by PCR with the primers P1/P2.
Generation of transgenic rice plants
To generate genome-edited ossgt1 mutants by CRISPR/Cas9, two simple guide RNAs targeting the single exon were ligated into the pRGEB32 vector. To generate OsSGT1 overexpression or UvPr1a heterologous transgenic rice plants, the coding sequences of UvPr1a or OsSGT1 were cloned into the vector pU1301. To generate UvPr1aHIGS transgenic rice plants, sense and antisense gene fragments of UvPr1a were ligated into the pDS1301 vector, resulting in the UvPr1aHIGS vector. The vectors pDS1301-UvPr1a, pU1301-OsSGT1, and pU1301-UvPr1a were used in A. tumefaciens-mediated transformation with japonica rice cv. Nip.
Y2H assay
The coding sequence of UvPr1a (without the SP) was cloned into pGBKT7 to generate the bait vector pGBKT7-UvPr1a. The mRNA of rice spikelets that had been inoculated 6 days earlier with U. virens was used to construct a cDNA library in the prey vector pGADT7. Library screening for UvPr1a interactors was performed as previously described (Chen et al., 2020). For interaction analysis of UvPr1a and OsSGT1 in yeast, the vectors pGADT7-OsSGT1, pGADT7-OsSGT1TPR, pGADT7-OsSGT1CS, pGADT7-OsSGT1SGS, pGBKT7-UvPr1a, pGBKT7-UvPr1aI9, and pGBKT7-UvPr1aS8 were co-transformed into yeast strain Y2HGold (Clontech, USA). Positive clones were selected on SD–Trp–Leu–His (SD-3) agar (Clontech, USA) containing Aureobasidin A (Clontech, USA), and further confirmed on SD-3 agar (Clontech, USA) containing X-α-Gal (Clontech, USA). The interaction between pGBKT7-53 and pGADT7-T was used as the positive control, and that of pGBKT7-Lam and pGADT7-T, as the negative control.
Co-IP assays
The coding sequences of UvPr1a and OsSGT1 were cloned into the vectors pCN-GFP and pCN-Flag, respectively. pCN-UvPr1a-GFP and pCN-OsSGT1-Flag vectors were introduced into Agrobacterium strain EHA105. Agrobacterium cultures harboring each construct were resuspended in infiltration buffer to a final OD600 of 1.6 and co-infiltrated into N. benthamiana leaves. Total proteins were extracted from N. benthamiana leaves 2 days after infiltration and incubated with Anti-Flag M2 affinity gel (Yeasen Biotech, China). Proteins eluted from gels were analyzed by immunoblotting with anti-GFP (Thermo Fisher Scientific, Waltham, MA, USA) or anti-Flag antibodies (Thermo Fisher Scientific, USA).
GST pull-down assays
The coding sequences of UvPr1a (without the SP) and OsSGT1 were cloned into vectors pGEX-4T-2 and pET28a, respectively. The constructs OsSGT1-pET28a and UvPr1a-pGEX4T-2 were introduced individually into E. coli BL21(DE3) cells for production and purification. The UvPr1a-GST fusion protein was extracted from E. coli cells and incubated with 100 µL of glutathione–agarose beads (Yeasen Biotech, China) at 4°C for 4 h with shaking. After centrifugation at 6000 rpm for 5 min at 4°C, the beads were collected and washed with phosphate-buffered saline (PBS) 3 times. Beads were then incubated with recombinant OsSGT1-His protein at 4°C for 2 h with shaking and then washed with PBS 3 times. Beads were boiled for 5 min at 100°C in 40-µL sodium dodecyl sulfate (SDS) sample loading buffer, and the proteins were analyzed by immunoblotting with anti-His and anti-GST antibodies (Thermo Fisher Scientific, USA).
Western blotting assay
Proteins were separated by 12% (w/v) sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS–PAGE) and transferred onto a polyvinylidene fluoride membrane (Merck Millipore, Burlington, MA, USA) using wet transfer at 80 V for 90 min with a BioRad electroblotting apparatus. Membranes were blocked in Tris-buffered saline 0.1% (v/v) Tween-20 (TBST) containing 10% (w/v) nonfat dry milk at room temperature for 2 h. Primary antibodies were: anti-SGT1 (1:2,000, ABclonal, Wuhan, China), anti-GFP (1:5,000, ABclonal, China), anti-Flag (1:5,000, ABclonal, China), anti-His (1:5,000, ABclonal, China), or anti-GST (1:5,000, ABclonal, China). The membrane was incubated with primary antibodies in TBST with 5% (w/v) nonfat dry milk at room temperature for 2 h with shaking, and washed 6 times (5 min each) with TBST. Next, the membrane was incubated with goat anti-mouse (1:10,000, Thermo Fisher Scientific, USA) secondary antibody in TBST with 5% (w/v) nonfat dry milk at room temperature for 1.5 h with shaking. The membrane was washed 6 times (5 min each) with TBST, and the signals were detected using Pierce ECL Western blotting substrate (Thermo Fisher Scientific, USA) in a ChemiDoc XRS+ system (Bio-Rad, Hercules, CA, USA).
BiFC assays
The coding sequences of OsSGT1 and UvPr1a were cloned into the BiFC vectors pCAMBIA1301-nYFP and pCAMBIA1301-cYFP, respectively. The resulting constructs encoding the OsSGT1-cYFP and UvPr1a-nYFP fusion proteins were co-infiltrated into N. benthamiana leaves. After 2 days, the YFP fluorescence signal of the infiltrated areas was examined with A Zeiss LSM 510 Meta confocal microscope (Carl Zeiss). YFP expression was detected after excitation with a 488-nm wavelength laser, and emissions were collected between 500 and 540 nm.
Protein purification and in vitro degradation assay
OsSGT1-His, OsMAPK33-His, and UvPr1a-GST fusion proteins were produced in E. coil BL21 (DE3) cells by induction with 0.1-M isopropyl β-d-1-thiogalactopyranoside. The cells were then harvested by centrifugation at 6000 rpm for 5 min at 4°C and subjected to sonication. Recombinant OsSGT1-His and UvPr1a-GST proteins were purified with HisPur Ni-NTA Resin and GSTrap Resin (Thermo Scientific, USA), respectively. For in vitro degradation assay, 5 µL of purified OsSGT1-His or OsMAPK33-His, with 1, 2, 5, and 10 µL of recombinant UvPr1a-GST or GST were incubated at 30°C for 15 min. The reaction products were analyzed by immunoblotting with anti-His and anti-GST antibodies.
Small RNA sequencing
Total RNA extracted from spikelets of UvPr1aHIGS transgenic rice lines at the booting stage was used for small RNA sequencing. RNA libraries were constructed with the NEB Next Multiplex Small RNA Library Prep Set for Illumina (New England Biolabs, Inc., Ipswich, MA, USA) and sequenced on an Illumina HiSeq platform by Wuhan IGENEBOOK Biotechnology Co., Ltd (Wuhan, China). Unique reads were mapped to the corresponding target fragment of UvPr1a.
FISH assays
For FISH assays, a 21-bp oligonucleotide probe (GGCCCCTACACCAAGTGTATC), which was the largest siRNA sequence from small RNA-seq, was labeled using Alexa Fluor 594 Tyramide SuperBoost Kit (ThermoFisher Scientific, USA). Infected spikelets at 6 dpi were placed in fixative solution (Servicebio, Wuhan, China) for 6 h. To preserve tissue morphology, samples were then dehydrated in 75% (v/v) ethanol, 50% (v/v) ethanol/PBS and 25% (v/v) ethanol/PBS each for 10 min. The dehydrated spikelets were embedded in paraffin and cut into slices and stored at −20°C. After dewaxing and rehydration, Proteinase K (20 mg L−1) was used for protein digestion. Samples were then incubated in Hybridization Mix at 55°C for 16 h, and then placed in wash solution. 4, 6-diamidino-2-phenylindole (DAPI) was used for nuclear staining, and samples were analyzed by laser confocal microscopy. Fluorescence of DAPI was excited at 405 nm and detected at 415–450 nm, while fluorescence of Alexa Fluor 594 was excited at 591 nm and detected at 600–640 nm.
Domain and phylogenetic analyses
Protein domains were predicted with SMART online software (http://smart.embl.de/smart/set_mode.cgi?NORMAL=1). A phylogenetic tree was constructed with MEGA software (version 7.0, http://www.megasoftware.net/index.php) using the neighbor-joining (NJ) algorithm.
Statistical analyses
Statistical analyses of each treatment were performed using SPSS version 14.0 software (SPSS, Chicago, IL, USA), and when significant treatment effects (P < 0.05) were found, the means were compared by the test of least significant difference at P < 0.05.
Accession numbers
Sequence data from this article can be found in the GenBank/EMBL data libraries under the following accession numbers: _Uv8b_1567, KDB17683.1; Uv8b_44, KDB18653.1; Uv8b_8155, KDB11027.1; Uv8b_6002, KDB13242.1; Uv8b_7084, KDB12091.1; Uv8b_4545, KDB14689.1; Uv8b_7700, KDB11477.1; Uv8b_6469, KDB12623.1; OsSGT1, LOC_Os01g43540.
Supplemental data
The following materials are available in the online version of this article.
Supplemental Figure S1. NJ tree of putative UvPr1a homologs from 13 fungal genomes generated using MEGA version 7.0.
Supplemental Figure S2. Gene deletion and complementation of UvPr1a in U. virens.
Supplemental Figure S3. Subcellular localization of UvPr1a-GFP in U. virens.
Supplemental Figure S4. Expression of UvPr1a in Nip and 35S-UvPr1a transgenic lines detected by PCR and immunoblotting.
Supplemental Figure S5. Controls in GST pull-down and Co-IP assays to prove that UvPr1a physically interacts with OsSGT1.
Supplemental Figure S6. Verification of OsSGT1 knockout and overexpression.
Supplemental Figure S7. UvPr1a does not degrade its interacting protein OsMAPK33.
Supplemental Figure S8. UvPr1a mediated protein degradation of OsSGT1 depends on its protease activity.
Supplemental Figure S9. Construction of UvPr1aHIGS transgenic rice lines.
Supplemental Figure S10. Subtilase family genes Uv8b_44, Uv8b_1567 (UvPr1a), and Uv8b_8155 are important for the development and virulence of U. virens.
Supplemental Table S1. Putative UvPr1a-interacting proteins identified by Y2H assays.
Supplemental Table S2. Agronomic traits of Nip, ossgt1, and OsSGT1-OE transgenic rice plants.
Supplemental Table S3. Agronomic traits of Nip and UvPr1aHIGS transgenic rice plants.
Supplemental Table S4. Primers used in this study.
Supplementary Material
Acknowledgments
We are grateful to Prof. Wenxian Sun (China Agricultural University) for providing the gene deletion vector pGKO.
Funding
This work was financially supported by the National Natural Science Foundation of China (32172372), the Key Research & Development Program of Hubei Province (2021BBA236), and the Fundamental Research Funds for the Central Universities of China (2021ZKPY016).
Conflict of interest statement. The authors declare that they have no conflicts of interest.
Contributor Information
Xiaoyang Chen, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Xiabing Li, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Yuhang Duan, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Zhangxin Pei, Wuhan Institute of Landscape Architecture, Wuhan 430081, China.
Hao Liu, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Weixiao Yin, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Junbin Huang, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Chaoxi Luo, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Xiaolin Chen, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Guotian Li, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
Kabin Xie, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China; National Key Laboratory of Crop Genetic Improvement, Huazhong Agricultural University, Wuhan 430070, China.
Tom Hsiang, School of Environmental Sciences, University of Guelph, Guelph N1G 2W1, Canada.
Lu Zheng, State Key Laboratory of Agricultural Microbiology/Hubei Key Laboratory of Plant Pathology, Huazhong Agricultural University, Wuhan 430070, China.
L.Z. conceived the project. L.Z. and X.Y.C. designed the experiments. X.Y.C. and X.B.L conducted most of the experiments. Y.H.D., Z.X.P., W.X.Y., X.L.C., and H.L. conducted parts of the research. J.B.H., G.T.L., K.B.X., and C.X.L. gave critical suggestions for the structure and writing of the manuscript. The manuscript was prepared by X.Y.C., T.H., and L.Z. All authors read and approved the contents of this paper.
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (https://academic.oup.com/plphys/pages/general-instructions) is Lu Zheng (luzheng@mail.hzau.edu.cn).
References
- Aarts N, Metz M, Holub E, Staskawicz BJ, Daniels MJ, Parker JE (1998) Different requirements for EDS1 and NDR1 by disease resistance genes define at least two R gene-mediated signaling pathways in Arabidopsis. Proc Natl Acad Sci USA 95: 10306–10311 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azevedo C, Sadanandom A, Kitagawa K, Freialdenhoven A, Shirasu K, Schulze-Lefert P (2002) The RAR1 interactor SGT1, an essential component of R gene-triggered disease resistance. Science 295: 2073–2076 [DOI] [PubMed] [Google Scholar]
- Azevedo C, Betsuyaku S, Peart J, Takahashi A, Noel L, Sadanandom A, Casais C, Parker J, Shirasu K (2006) Role of SGT1 in resistance protein accumulation in plant immunity. EMBO J 25: 2007–2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker D, Shiau AK, Agard DA (1993) The role of pro regions in protein folding. Curr Opin Cell Biol 5: 966–970 [DOI] [PubMed] [Google Scholar]
- Bryan PN (2002) Prodomains and protein folding catalysis. Chem Rev 102: 4805–4816 [DOI] [PubMed] [Google Scholar]
- Chalfoun NR, Grellet-Bournonville CF, Martínez-Zamora MG, Díaz-Perales A, Castagnaro AP, Díaz-Ricci JC (2013) Purification and characterization of AsES protein: a subtilisin secreted by Acremonium strictum is a novel plant defense elicitor. J Biol Chem 288: 14098–14113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen XY, Tang JT, Pei ZX, Liu H, Huang JB, Luo CX, Hsiang T, Zheng L (2020) The “pears and lemons” protein UvPal1 regulates development and virulence of Ustilaginoidea virens. Environ Microbiol 22: 5414–5432 [DOI] [PubMed] [Google Scholar]
- Chen XY, Xu QT, Duan YH, Liu H, Chen XL, Huang JB, Luo CX, Zhou DX, Zheng L (2021a) Ustilaginoidea virens modulates lysine 2‐hydroxyisobutyrylation in rice flowers during infection. J Integr Plant Biol 63: 1801–1814 [DOI] [PubMed] [Google Scholar]
- Chen XY, Li XB, Li PP, Chen XL, Liu H, Huang JB, Luo CX, Hsiang T, Zheng L (2021b) Comprehensive identification of lysine 2‐hydroxyisobutyrylated proteins in Ustilaginoidea virens reveals the involvement of lysine 2-hydroxyisobutyrylation in fungal virulence. J Integr Plant Biol 63: 409–425 [DOI] [PubMed] [Google Scholar]
- Chen XY, Pei ZX, Liu H, Huang JB, Chen XL, Luo CX, Hsiang T, Zheng L (2022) Host-induced gene silencing of fungal-specific genes of Ustilaginoidea virens confers effective resistance to rice false smut. Plant Biotechnol J 20: 253–255 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Couto D, Zipfel C (2016) Regulation of pattern recognition receptor signaling in plants. Nat Rev Immunol 16: 537–552 [DOI] [PubMed] [Google Scholar]
- Dangl JL, Horvath DM, Staskawicz BJ (2013) Pivoting the plant immune system from dissection to deployment. Science 341: 746–751 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant-pathogen interactions. Nat Rev Gene. 11: 539–548 [DOI] [PubMed] [Google Scholar]
- Dodson G, Wlodawer A (1998) Catalytic triads and their relatives. Trends Biochem Sci 23: 347–352 [DOI] [PubMed] [Google Scholar]
- Donofrio NM, Oh Y, Lundy R, Pan H, Brown DE, Jeong JS, Coughlan S, Mitchell TK, Dean RA (2006) Global gene expression during nitrogen starvation in the rice blast fungus, Magnaporthe grisea. Fungal Genet Biol 43: 605–617 [DOI] [PubMed] [Google Scholar]
- Dou DL, Zhou JM (2012) Phytopathogen effectors subverting host immunity: different foes, similar battleground. Cell Host Microbe 12: 484–495 [DOI] [PubMed] [Google Scholar]
- Dou T, Shao XH, Hu CH, Liu SW, Sheng O, Bi F, Deng G, Ding L, Li C, Dong T, et al. (2020) Host-induced gene silencing of Foc TR 4 ERG 6/11 genes exhibits superior resistance to Fusarium wilt of banana. Plant Biotechnol J 18: 11–13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du Y, Stegmann M, Misas VJC (2016) The apoplast as battleground for plant-microbe interactions. New Phytol 209: 34–38 [DOI] [PubMed] [Google Scholar]
- Duan X, Zhang Z, Wang J, Zuo K (2016) Characterization of a novel cotton subtilase gene GbSBT1 in response to extracellular stimulations and its role in Verticillium resistance. PLoS One 11: e0153988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan T, Bykova NV, Rampitsch C, Xing T (2016) Identification and characterization of a serine protease from wheat leaves. Eur J Plant Pathol 146: 293–304 [Google Scholar]
- Fang AF, Han YQ, Zhang N, Zhang M, Liu LJ, Li S, Lu F, Sun WX (2016) Identification and characterization of plant cell death-inducing secreted proteins from Ustilaginoidea virens. Mol Plant-Microbe Interact 29: 405–16 [DOI] [PubMed] [Google Scholar]
- Fang AF, Gao H, Zhang N, Zhang XH, Qiu SS, Li YJ, Zhou S, Cui FH, Sun WX (2019) A novel effector gene SCRE2 contributes to full virulence of Ustilaginoidea virens to rice. Front Microbiol 10: 845–860 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Figueiredo J, Marta SS, Andreia F (2018) Subtilisin‐like proteases in plant defence: the past, the present and beyond. Mol Plant Pathol 19: 1017–1028 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao BJ, Mou YN, Tong SM, Ying SH, Feng MG (2020) Subtilisin-like Pr1 proteases marking the evolution of pathogenicity in a wide-spectrum insect-pathogenic fungus. Virulence 11: 365–380 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hammond-Kosack KE, Parker JE (2003) Deciphering plant-pathogen communication: fresh perspectives for molecular resistance breeding. Curr Opin Biotech 14: 177–193 [DOI] [PubMed] [Google Scholar]
- Huang HW, Chen WC, Wu CY, Yu HC, Lin WY, Chen ST, Wang KT (1997) Kinetic studies of the inhibitory effects of propeptides subtilisin BPN 0 and Carlsberg to bacterial serine proteases. Protein Eng 10: 1227–1233 [DOI] [PubMed] [Google Scholar]
- Hubert DA, Tornero P, Belkhadir Y, Krishna P, Takahashi A, Shirasu K, Dangl JL (2003) Cytosolic HSP90 associates with and modulates the Arabidopsis RPM1 disease resistance protein. EMBO J 22: 5679–5689 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobs KA, Collins-Racie LA, Colbert M, Duckett M, Golden-Fleet M, Kelleher K, Kriz R, LaVallie ER, Merberg D, Spaulding V, et al. (1997) A genetic selection for isolating cDNAs encoding secreted proteins. Gene 198: 289–296 [DOI] [PubMed] [Google Scholar]
- Jones JD, Dangl JL (2006) The plant immune system. Nature 44: 323–329 [DOI] [PubMed] [Google Scholar]
- Jordá L, , CoegoA, , ConejeroV, , Vera P (1999) A genomic cluster containing four differentially regulated subtilisin-like processing protease genes is in tomato plants. J Biol Chem 274: 2360–2365 [DOI] [PubMed] [Google Scholar]
- Koiso Y, Li YIN, Iwasaki S, Hanaoka K, Kobayashi T, Sonoda R, Fujita Y, Yaegashi H, Sato Z, et al. (1994) Ustiloxins, antimitotic cyclic peptides from false smut balls on rice panicles caused by Ustilaginoidea virens. J Antibiot 47: 765–773 [DOI] [PubMed] [Google Scholar]
- Li X, Huang R, Liu J, Xu G, Yuan M (2022) Engineering false smut resistance rice via host‐induced gene silencing of two chitin synthase genes of Ustilaginoidea virens. Plant Biotechnol J 1: 2386–2388 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y, Inouye M (1994) Autoprocessing of prothiolsubtilisin E in which active-site serine 221 is altered to cysteine. J Biol Chem 269: 4169–4174 [PubMed] [Google Scholar]
- Liu YL, Schiff M, Serino G, Deng XW, Dinesh-Kumar SP (2002) Role of SCF ubiquitin-ligase and the COP9 signalosome in the N gene-mediated resistance response to Tobacco mosaic virus. Plant Cell 14: 1483–1496 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu YL, Burch-Smith T, Schiff M, Feng SH, Dinesh-Kumar SP (2004) Molecular chaperone HSP90 associates with resistance protein N and its signaling proteins SGT1 and RAR1 to modulate an innate immune response in plants. J Biol Chem 279: 2101–2108 [DOI] [PubMed] [Google Scholar]
- Meyer M, Leptihn S, Welz M, Schaller A (2016) Functional characterization of propeptides in plant subtilases as intramolecular chaperones and inhibitors of the mature protease. J Biol Chem 291: 19449–19461 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muskett PR, Kahn K, Austin MJ, Moisan LJ, Sadanandom A, Shirasu K, Jones JDG, Parker JE (2002) Arabidopsis RAR1 exerts rate-limiting control of R gene-mediated defenses against multiple pathogens. Plant Cell 14: 979–992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nowara D, Gay A, Lacomme C, Shaw J, Ridout C, Douchkov D, Hensel G, Kumlehn J, Schweizer P (2010) HIGS: host-induced gene silencing in the obligate biotrophic fungal pathogen Blumeria graminis. Plant Cell 22: 3130–3141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olivieri F, Zanetti ME, Oliva CR, Covarrubias AA, Casalongué CA (2002) Characterization of an extracellular serine protease of Fusarium eumartii and its action on pathogenesis related proteins. Eur J Plant Pathol 108: 63–72 [Google Scholar]
- Parker JE, Holub EB, Frost LN, Falk A, Gunn ND, Daniels MJ (1996) Characterization of eds1, a mutation in Arabidopsis suppressing resistance to Peronospora parasitica specified by several different RPP genes. Plant Cell 8: 2033–2046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pekkarinen AI, Jones BL (2003) Purification and identification of barley (Hordeum vulgare L.) proteins that inhibit the alkaline serine proteinases of Fusarium culmorum. J Agric Food Chem 51: 1710–1717 [DOI] [PubMed] [Google Scholar]
- Rautengarten C, Steinhauser D, Büssis D, Stintzi A, Schaller A, Kopka J, Altmann T (2005) Inferring hypotheses on functional relationships of genes: analysis of the Arabidopsis thaliana subtilase gene family. PLoS Comput Biol 1: e40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramírez V, López A, Mauch-Mani B, Gil MJ, Vera P (2013) An extracellular subtilase switch for immune priming in Arabidopsis. PLoS Pathog 9: e1003445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaller A, Stintzi A, Graff L (2012) Subtilases-versatile tools for protein turnover, plant development, and interactions with the environment. Physiol Plant 145: 52–66 [DOI] [PubMed] [Google Scholar]
- Shinde U, Thomas G (2011) Insights from bacterial subtilases into the mechanisms of intramolecular chaperone-mediated activation of furin. Methods Mol Biol 768: 59–106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shirasu K, Schulze-Lefert P (2003) Complex formation, promiscuity and multi-functionality: protein interactions in disease-resistance pathways. Trends Plant Sci 8: 252–258 [DOI] [PubMed] [Google Scholar]
- Shirasu K (2009) The HSP90-SGT1 chaperone complex for NLR immune sensors. Annu Rev Plant Biol 60: 139–64 [DOI] [PubMed] [Google Scholar]
- Song TQ, Zhang Y, Zhang Q, Zhang X, Shen D, Yu J, Yu M, Pan X, Cao H, Yong M, et al. (2021) The N-terminus of an Ustilaginoidea virens Ser-Thr-rich glycosylphosphatidylinositol-anchored protein elicits plant immunity as a MAMP. Nat Commun 12: 1–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sreedhar L, Kobayashi DY, Bunting TE, Hillman BI, Belanger FC (1999) Fungal proteinase expression in the interaction of the plant pathogen Magnaporthe poae with its host. Gene 235: 121–129 [DOI] [PubMed] [Google Scholar]
- Sun WX, Fan J, Fang AF, Li YJ, Tariqjaveed M, Li DY, Hu DW, Wang WM (2020) Ustilaginoidea virens: Insights into an emerging rice pathogen. Annu Rev Phytopathol 58: 363–385 [DOI] [PubMed] [Google Scholar]
- Tang JT, Chen XY, Yan YQ, Huang JB, Luo CX, Hsiang T, Zheng L (2021) Comprehensive transcriptome profiling reveals abundant long non-coding RNAs associated with development of the rice false smut fungus, Ustilaginoidea virens. Environ Microbiol 23: 4998–5013 [DOI] [PubMed] [Google Scholar]
- Tian M,, Huitema E,, Da Cunha L,, Torto-Alalibo T,, Kamoun S (2004) A Kazal-like extracellular serine protease inhibitor from Phytophthora infestans targets the tomato pathogenesis-related protease P69B. J Biol Chem 279: 26370–26377 [DOI] [PubMed] [Google Scholar]
- Tor M, Gordon P, Cuzick A, Eulgem T, Sinapidou E, Mert-Turk F, Can C, Dangl JL, Holub EB (2002) Arabidopsis SGT1b is required for defense signaling conferred by several downy mildew resistance genes. Plant Cell 14: 993–1003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tornero P, , ConejeroV, , Vera P (1997) Identification of a new pathogen-induced member of the subtilisin-like processing protease family from plants. J Biol Chem 272: 14412–14419 [DOI] [PubMed] [Google Scholar]
- Tornero P, Merritt P, Sadanandom A, Shirasu K, Innes RW, Dangl JL (2002) RAR1 and NDR1 contribute quantitatively to disease resistance in Arabidopsis, and their relative contributions are dependent on the R gene assayed. Plant Cell 14: 1005–1015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walter S, Nicholson P, Doohan FM (2010) Action and reaction of host and pathogen during Fusarium head blight disease. New Phytol 185: 54–66 [DOI] [PubMed] [Google Scholar]
- Wang MH, Wu L, Mei YZ, Zhao YF, Ma ZH, Zhang X, Chen Y (2020) Host-induced gene silencing of multiple genes of Fusarium graminearum enhances resistance to Fusarium head blight in wheat. Plant Biotechnol J 18: 2373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Gao M, Li Q, Wang L, Wang J, Jeon JS, Qu N, Zhang Y, He Z (2008) OsRAR1 and OsSGT1 physically interact and function in rice basal disease resistance. Mol Plant Microbe In 21: 294–303 [DOI] [PubMed] [Google Scholar]
- Xu J, Wang X, Li Y, Zeng J, Wang G, Deng C, Guo W (2018) Host-induced gene silencing of a regulator of G protein signaling gene (VdRGS1) confers resistance to Verticillium wilt in cotton. Plant Biotechnol J 16: 1629–1643 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu L, Wang HK, Zhang CQ, Wang JL, Chen A, Chen Y, Ma ZH (2020) System-wide characterization of subtilases reveals that subtilisin-like protease FgPrb1 of Fusarium graminearum regulates fungal development and virulence. Fungal Genet Biol 144: 103449. [DOI] [PubMed] [Google Scholar]
- Yan JJ, Zhang YB, Ding Y (2012) Binding mechanism between Hsp90 and Sgt1 explored by homology modeling and molecular dynamics simulations in rice. J Mol Model 18: 4665–4673 [DOI] [PubMed] [Google Scholar]
- Yu G, Liu JL, Xie LQ, Wang XL, Zhang SH, Pan HY (2012) Characterization, cloning, and heterologous expression of a subtilisin-like serine protease gene VlPr1 from Verticillium lecanii. J Microbiol 50: 939–946 [DOI] [PubMed] [Google Scholar]
- Zhang N, Yang J, Fang AF, Wang J, Li D, Li Y, Wang S, Cui F, Yu J, Liu Y, et al. (2020) The essential effector SCRE1 in Ustilaginoidea virens suppresses rice immunity via a small peptide region. Mol Plant Pathol 21: 445–459 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Zhang K, Fang AF, Han Y, Yang J, Xue M, Bao J, Hu D, Zhou B, Sun X, et al. (2014) Specific adaptation of Ustilaginoidea virens in occupying host florets revealed by comparative and functional genomics. Nat Commun 5: 3849. [DOI] [PubMed] [Google Scholar]
- Zhu X, Ohta Y, Jordan F, Inouye M (1989) Pro-sequence of subtilisin can guide the refolding of denatured subtilisin in an intermolecular process. Nature 339: 483–484 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.