Abstract
Adipocyte browning is one of the potential strategies for the prevention of obesity-related metabolic syndromes, but it is a complex process. Although previous studies make it increasingly clear that several transcription factors and enzymes are essential to induce browning, it is unclear what dynamic and metabolic changes occur in induction of browning. Here, we analyzed the effect of a beta-adrenergic receptor agonist (CL316243, accelerator of browning) on metabolic change in mice adipose tissue and plasma using metabolome analysis and speculated that browning is regulated partly by inosine 5′-monophosphate (IMP) metabolism. To test this hypothesis, we investigated whether Ucp-1, a functional marker of browning, mRNA expression is influenced by IMP metabolism using immortalized adipocytes. Our study showed that mycophenolic acid, an IMP dehydrogenase inhibitor, increases the mRNA expression of Ucp-1 in immortalized adipocytes. Furthermore, we performed a single administration of mycophenolate mofetil, a prodrug of mycophenolic acid, to mice and demonstrated that mycophenolate mofetil induces adipocyte browning and miniaturization of adipocyte size, leading to adipose tissue weight loss. These findings showed that IMP metabolism has a significant effect on adipocyte browning, suggesting that the regulator of IMP metabolism has the potential to prevent obesity.
Keywords: metabolome analysis, adipocytes, browning, purine metabolism
Abbreviations: BAT, brown adipose tissue; BCAA, branched chain amino acid; EWAT, epididymal adipose tissue; IMP, inosine 5′-monophosphate; IWAT, inguinal adipose tissue; KEGG, Kyoto Encyclopedia of Genes and Genomes; MMF, mycophenolate mofetil; MPA, mycophenolic acid; MS, mass spectrometry; NEFA, nonesterified fatty acid; TG, triglyceride; WAT, white adipose tissue
Brown adipocyte and beige adipocyte contribute to an enhancement of thermogenesis and prevention of obesity. Recent studies have elucidated the functional and morphological characterization of these adipocytes partially. It is well known that there are three types of adipocytes (white adipocyte, brown adipocyte, and beige adipocyte). Although white adipocyte is unilocular cell and has an ability to store energy as triglyceride, brown adipocyte is multilocular cell and capable of thermogenesis to maintain body temperature under the condition of cold exposure (1, 2). The morphological feature of beige adipocyte is similar to that of brown adipocyte (3). The differentiation of beige adipocyte is induced by stimulation of cold exposure or adrenergic agonist treatment (known as ‘browning’) (4, 5, 6) and abolished by stop of these stimulations (7). This reversible phenomenon is characteristic of beige adipocytes. Previous studies have shown that brown adipose tissue (BAT) is protective against body fat accumulation in humans (8, 9). Therefore, the ability of brown adipocyte and beige adipocyte to induce thermogenesis paves the way for effective therapy of obesity-related diseases.
Uncoupling protein-1 (UCP-1) plays a critical role in thermogenesis of brown adipocyte and beige adipocyte and induces miniaturization of adipocytes. It is well known that UCP-1 is activated by the stimulation of beta-adrenergic receptor with its agonists (e.g., CL316243) and/or the activation of sympathetic nerve (e.g., cold exposure) and induces thermogenesis via uncoupling of oxidative phosphorylation in mitochondria of these adipocytes (5, 6). CL316243 is an adrenergic-β3 receptor agonist, which plays a critical role in the regulation of UCP-1 expression in vivo. The stimulation of beta3-adrenergic receptor in adipocyte induces lipolysis and UCP-1 upregulation via activation of adenylyl cyclase, protein kinase A, and hormone-sensitive lipase (6). It has been also reported that administration of acetic acid to mice increases mRNA expression level of Ucp-1 and induced miniaturization of adipocyte (10).
Although previous studies revealed the presence of several transcription factors and enzymes contributing to differentiation of brown adipocyte and beige adipocyte, it is largely unclear what metabolic pathway plays a key role in induction of brown adipocyte and beige adipocyte via UCP-1 upregulation. In the previous studies, it has been reported that PR domain containing protein 16, one of the transcription factors, is essential to differentiate into brown adipocyte (11) and that differentiation into beige adipocyte is inhibited by knockdown or KO of PR domain containing protein 16 (12). Furthermore, proteome analysis revealed that UCP-1 upregulation and browning of white fat is induced by knockdown of casein kinase 2 α1 (13). Recently, it has been reported that branched chain amino acid catabolism in brown adipocyte contributes to thermogenesis (14). Furthermore, it has been demonstrated that cinnamon extract has a browning activity in adipocytes via UCP-1 upregulation (15). These findings suggest that several primary metabolites and phytochemical contribute to activation of brown adipocyte and beige adipocyte. On the other hand, the relationship between browning and metabolic changes in adipocyte remains poorly understood.
Metabolome analysis is essential for overview analysis of metabolic variance. Over the past decade, metabolome analysis based on liquid chromatography coupled with ultra-accurate mass spectrometer is applied to investigation of animals (16, 17, 18), plants (19, 20, 21), and human (22, 23) metabolism. Metabolome analysis makes it possible to determine molecular formula provided by exact mass data and speculate on compound structure by metabolite database. Therefore, metabolome analysis is effective approach for comprehensive analysis and monitoring of phytochemical or animal metabolites. In the recently, mass spectrometry (MS)–based lipidomes of adipocytes showed that white adipocytes differed from beige and brown adipocytes in their lipid class and species profile, and this study identified that beta-adrenergic stimulation induces lysophosphatidylcholine in brown adipocytes leading to alteration of mitochondrial bioenergetics (24). Furthermore, several previous studies using metabolome analysis revealed that the exposure to cold or treatment with CL316243 induces the change of metabolomic profiling in brown and beige adipose tissue (25, 26). These findings showed that metabolome analysis is needed to better understand how metabolic change is reflected in adipocyte browning.
Our study aimed to find out metabolic pathway contributing induction of brown adipocyte and beige adipocyte. We performed metabolome analysis of mice adipose tissue and plasma metabolites treated with or without beta-adrenergic stimulation. In this analysis, we estimate that adipocyte browning is regulated by inosine 5′-monophosphate (IMP) metabolism. We showed that an administration of IMP dehydrogenase inhibitor to mice contributes to the upregulation of Ucp-1 mRNA expression and miniaturization adipocyte size, leading to adipose tissue weight loss. Our findings showed that IMP metabolic pathway contributes to regulation of adipocyte browning.
Results
The effect of CL316243 on body and adipose tissues weight, cell size, and browning of murine adipocyte
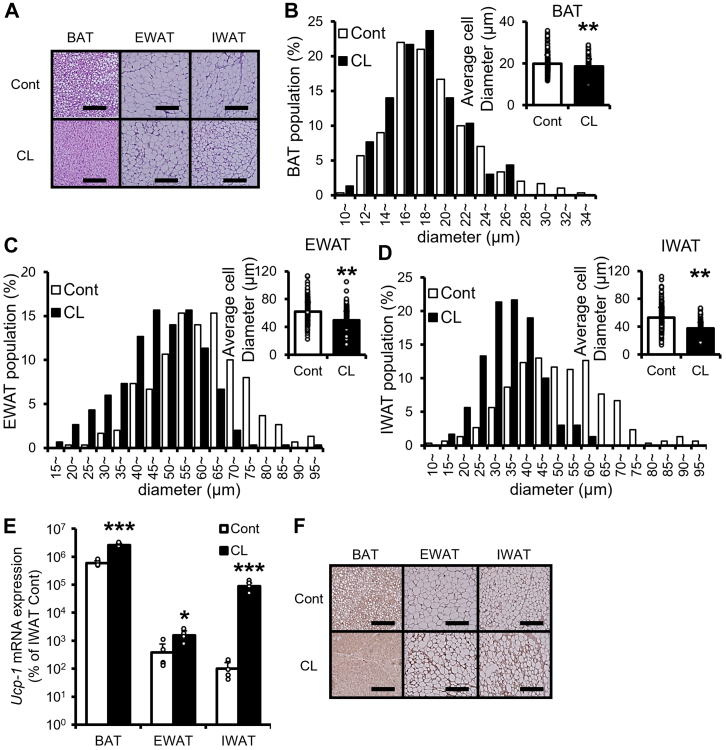
We hypothesized that endogenous metabolite that regulates browning is identified by the analysis of mice treated with or without adrenergic agonist. To test this hypothesis, we investigated the effect of CL316243, a selective β3-adrenoceptor agonist, on browning and metabolism. We analyzed the effect of CL316243 on body and adipose tissues weight, cell size, and browning of murine adipocyte. There were no significant differences in body weight among control and CL316243, an adrenergic receptor agonist, treatment group animals (Table 1). In the individual adipose tissues weight, although BAT was increased, epididymal adipose tissue (EWAT) was decreased by CL316243 treatment (Table 1). CL316243 had a tendency to decrease inguinal adipose tissue (IWAT) weight (Table 1). CL316243 had an ability to decrease adipocyte size (Fig. 1, A–D) and increase the mRNA expression of Ucp-1 in the individual adipose tissues (Fig. 1E). IWAT treated with CL316243 was the maximum change ratio of Ucp-1 mRNA expression among IWAT, EWAT, and BAT (Fig. 1E). Histological analysis suggested that CL316243 leads to a substantial increase in UCP-1 expression in the individual adipose tissues (Fig. 1F).
Table 1.
Effect of CL316243 on body and adipose tissue weight
| Weight | Control | CL316243 |
|---|---|---|
| Body (g) | 24.1 ± 0.8 | 23.7 ± 0.9 |
| BAT (mg) | 39.2 ± 6.0 | 46.9 ± 4.5∗ |
| IWAT (mg) | 280.3 ± 45.4 | 232.6 ± 25.5 |
| EWAT (mg) | 292.3 ± 47.3 | 172.0 ± 38.9∗ |
Data is expressed as means ± SD (n = 6–7/group). ∗p < 0.05 versus control group.
Figure 1.
Effect of CL316243 on adipocyte size, Ucp-1 mRNA, and UCP-1 protein expression level in mice.A, histological sections of BAT, EWAT, and IWAT. Adipocyte size population and average cell diameter (right panel, n = 300 cells/group) in (B) BAT, (C) EWAT, and (D) IWAT. E, Ucp-1 mRNA expression level of BAT, EWAT, and IWAT. F, UCP-1 immunostaining of BAT, EWAT, and IWAT (n = 6–7/group). Data are expressed as means ± SD. ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001 versus Cont. Cont: control group; CL: CL316243 treatment group; the scale bar represents 200 μm. BAT, brown adipose tissue; EWAT, epididymal adipose tissue; IWAT, inguinal adipose tissue.
Metabolomic analysis of mice adipose tissue and plasma treated with CL316243
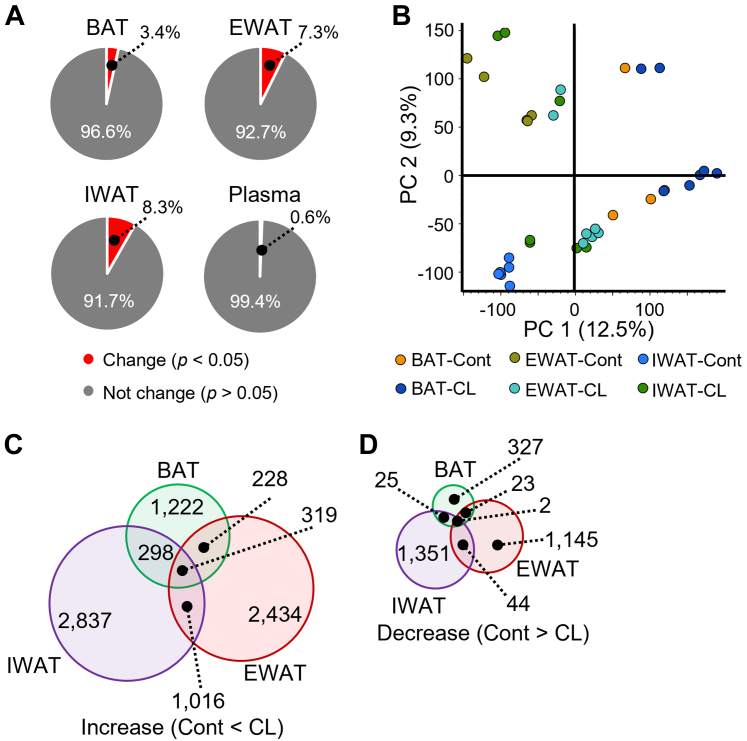
We performed metabolomic analysis of mice adipose tissue and plasma treated with CL316243 in order to search for metabolites associated with browning. At first, we analyzed the influence of CL316243 treatment on detected peaks in adipose tissue and plasma (Dataset S1). Among them, the change ratio of detected peaks was highest in IWAT (Fig. 2A). Approximately 3.4%, 7.3%, 8.3%, and 0.6% of detected peaks were changed in BAT, EWAT, IWAT, and plasma, respectively (Fig. 2A). In principal component analysis of metabolic profiling, it is difficult to be distinguished between control and CL316243 treatment group in each adipose tissue (Fig. 2B). On the other hand, the number of IWAT-specific changed peaks was maximum in significant different peak between control and CL316243 treatment group (Fig. 2, C and D).
Figure 2.
Effect of CL316243 on the variance of metabolic profile.A, the proportion of peaks altered by CL316243 treatment significantly. B, principal component analysis of adipose tissue in mice treated with or without CL316243. C and D, classification of peaks altered by CL316243 treatment significantly. Individual number of Venn diagrams show the number of peaks altered significantly by CL316243 treatment in individual region, respectively. Cont. Cont: control group; CL: CL316243 treatment group. The peak data is provided in Dataset S1.
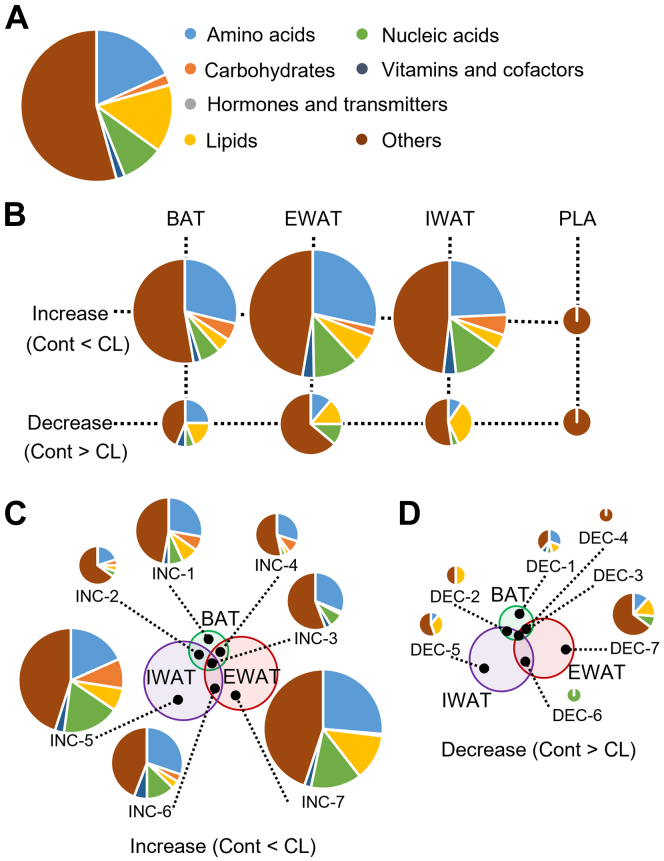
Next, compound name of detected peaks was estimated by Kyoto Encyclopedia of Genes and Genomes (KEGG) database based on exact mass data. About 1624 compounds were estimated (Dataset S2) and classified as ‘Amino acids group’, ‘Carbohydrates groups’, ‘Hormones and transmitters group’, ‘Lipids group’, ‘Nucleic acids group’, ‘Vitamins and cofactors group’, and ‘Others group’ (Fig. 3A). Among metabolite groups except for others group, metabolites of amino acids group including branched chain amino acids (BCAAs) were especially increased by CL316243 treatment in the individual adipose tissue (Fig. 3B and Dataset S3). Remarkably, metabolites of nucleic acids group in IWAT and EWAT were increased by CL316243 treatment (INC-5, INC-6, and INC-7 group, Fig. 3, C and D). On the other hand, lipid changes took significant proportion in the decreased metabolites (Fig. 3, B and D).
Figure 3.
Estimated categorization of metabolites in individual adipose tissue and plasma. Estimated metabolic categorization of (A) 1624 peaks among detected peaks, (B) peaks altered by CL316243 treatment significantly, and (C and D) 14 regions in Venn diagrams (INC1-7 and DEC1-7) using KEGG database. There was no estimated metabolic categorization in DEC3 region. Cont. Cont: control group; CL: CL316243 treatment group. The data of categorization information is provided in Dataset S2. KEGG, Kyoto Encyclopedia of Genes and Genomes.
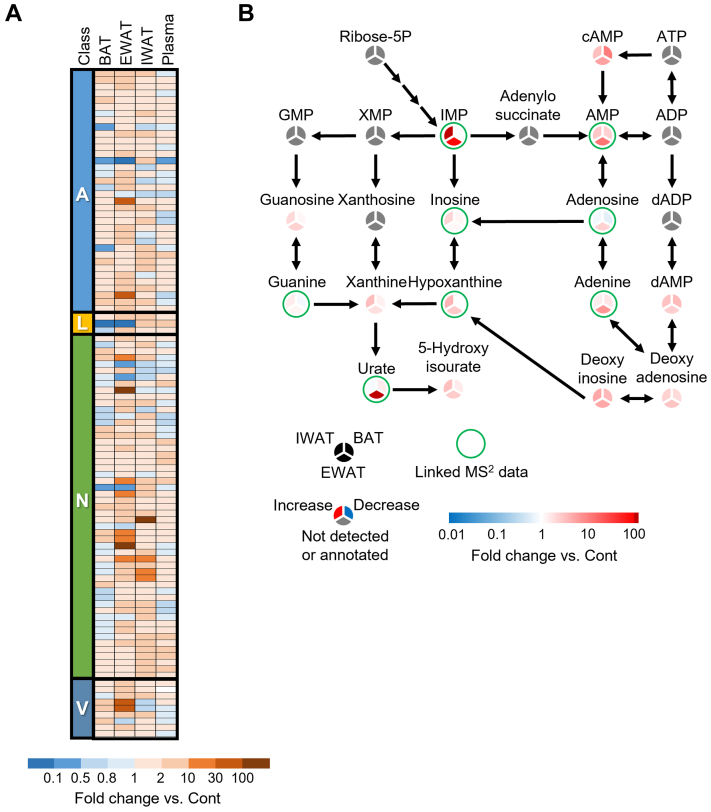
In order to better understand the information of compound structure, MS2 spectrums were analyzed by mzCloud database. The 36, 3, 53, and 9 peaks’ name of amino acids group, lipids group, nucleic acids group, and vitamins and cofactors group, respectively, were estimated by MS2 spectrums data (Dataset S2, Figs. S1, and 4A). We analyzed the effect of CL316243 treatment on purine metabolism (Dataset S4 and Fig. 4B). KEGG and mzCloud database analysis suggested that IMP metabolism in IWAT and EWAT is strongly influenced by CL316243 (Dataset S4 and Fig. 4B).
Figure 4.
Estimated metabolic pathway analysis using MS2spectrum data.A, heat map of metabolites estimated by mzCloud data. A: amino acids group; L: lipids group; N: nucleic acids group; V: vitamins and cofactors group. The peak data and MS2 data in detail are provided in the Dataset S2 and Fig. S1, respectively. B, the estimated effect of CL316243 on purine metabolic pathway. The peak data of the pathway in detail are provided in the Dataset S4. MS, mass spectrometry.
The effect of isoproterenol and mycophenolic acid on Ucp1 mRNA expression level, cAMP, and IMP metabolism in immortalized adipocytes
We tested the hypothesis that adrenergic agonist activates IMP metabolism in vitro. Isoproterenol (Iso), an artificial catecholamine and an adrenergic receptor agonist used as a positive regulator of Ucp-1 in cultured cells in general, induced not only cAMP (Fig. 5A) but also IMP (Fig. 5B) in immortalized adipocytes. We confirmed that the mRNA expression of Ucp-1 is increased by Iso treatment (Fig. 5C). Furthermore, we demonstrated that mycophenolic acid (MPA), an IMP dehydrogenase inhibitor, not only induces IMP (Fig. 5E) but also increases cAMP (Fig. 5D) and the mRNA expression of Ucp-1 (Fig. 5F) in immortalized adipocytes.
Figure 5.
Effect of isoproterenol and MPA on Ucp1 mRNA expression level, cAMP, and IMP metabolism in immortalized adipocytes. Effect of isoproterenol (Iso, 1 μM) on (A) cAMP and (B) IMP concentration and (C) Ucp-1 mRNA expression level. Effect of mycophenolic acid (MPA, 10 μM) on (D) cAMP and (E) IMP concentration and (F) Ucp-1 mRNA expression level. Data are expressed as means ± SD (n = 5–6/group). ∗p < 0.05, ∗∗p < 0.01 versus Cont. Cont: control group; Iso: isoproterenol treatment group; MPA: mycophenolic acid treatment group. IMP, inosine 5′-monophosphate; MPA, mycophenolic acid.
The effect of mycophenolate mofetil on body and adipose tissues weight, cell size, and browning of murine adipocyte
To test whether IMP metabolism regulates browning, we performed an administration of mycophenolate mofetil (MMF), a prodrug of MPA, to mice. There were no significant differences in body and BAT weight among control and MMF treatment group animals (Table 2). MMF had no effect on food intake (control group: 4.40 ± 0.52 g/day; MMF group: 4.03 ± 0.91 g/day; p = 0.66). On the other hand, IWAT and EWAT weight were decreased by MMF treatment (Table 2). We investigated the effect of MMF on plasma glucose, triglyceride (TG), and nonesterified fatty acid (NEFA) level and showed that plasma glucose level tends to be decreased by MMF treatment (Table S1). MMF had no effect on plasma TG and NEFA level (Table S1). In addition, we also showed that MMF has no effect on plasma glutamic oxaloacetic transaminase (GOT) and glutamic pyruvic transaminase (GPT) level (Table S1). We showed that the mRNA expression of Ucp-1 in the individual adipose tissues is increased by MMF treatment (Fig. 6A). We investigated the effect of MMF on other markers of thermogenesis (Pgc1a), brown/beige adipocyte identity (Cidea and Prdm16), and general adipogenesis (Pparγ, aP2, and Adiponectin). Although Cidea expression level tended to be increased by MMF treatment, the mRNA expressions of these markers were not influenced by MMF treatment statistically (Fig. S2). Histological analysis suggested that MMF leads to a substantial increase in UCP-1 expression especially in BAT and IWAT (Fig. 6B). Furthermore, we demonstrated that MMF has an ability to decrease adipocyte size in the individual adipose tissues (Fig. 6, C–F). MMF had no effect on adenosine, the known browning activator, level in adipose tissue (Table S2).
Table 2.
Effect of MMF on body and adipose tissue weight
| Weight | Control | MMF |
|---|---|---|
| Body (g) | 24.9 ± 0.2 | 24.2 ± 1.1 |
| BAT (mg) | 59.3 ± 33.5 | 57.0 ± 3.4 |
| IWAT (mg) | 300.7 ± 41.4 | 243.0 ± 16.9∗ |
| EWAT (mg) | 373.3 ± 35.9 | 266.0 ± 53.9∗ |
Data is expressed as means ± SD (n = 3–4/group). ∗p < 0.05 versus Control group.
Figure 6.
Effect of MMF on adipocyte size, Ucp-1 mRNA, and UCP-1 protein expression level in mice.A, Ucp-1 mRNA expression level (n = 3–4/group) and (B) UCP-1 immunostaining of BAT, EWAT, and IWAT. C, histological sections of BAT, EWAT, and IWAT. Adipocytes size population and average cell diameter (right panel, n = 300 cells/group) in (D) BAT, (E) EWAT, and (F) IWAT. Data are expressed as means ± SD. ∗p < 0.05, ∗∗p < 0.01 versus Cont. Cont: control group; MMF: MMF treatment group; LQ: low limited of quantification. The scale bar represents 200 μm. BAT, brown adipose tissue; EWAT, epididymal adipose tissue; IWAT, inguinal adipose tissue; MMF, mycophenolate mofetil.
The relation between IMP dehydrogenase (Impdh) and Ucp-1 mRNA expression levels
We investigated the direct effect of Impdh-1 and Impdh-2 gene on Ucp-1 mRNA expression levels using HB2 adipocytes, which are brown adipocyte originated from p53 KO mice. The mRNA expression of Ucp-1 in HB2 adipocytes was also decreased by Impdh-1 and Impdh-2 double-knockdown using siRNA (Fig. S3). We also investigated the effect of cold exposure on Impdh mRNA expression level in mice IWAT. We confirmed that Ucp-1 expression level is increased by cold exposure (Fig. 7A). In addition, we also demonstrated that Impdh-1 expression level is decreased by cold exposure condition (Fig. 7B). The cold exposure had no effect on the mRNA expression level of Impdh-2 (Fig. 7C).
Figure 7.
The effect of cold exposure on Ucp-1 and Impdh mRNA expression level in mice IWAT. The relative mRNA expression level of (A) Ucp-1, (B) Impdh-1, and (C) Impdh-2 in mice IWAT treated with RT or CE. Data are expressed as means ± SD (n = 6–7/group). ∗p < 0.05, ∗∗p < 0.01 versus RT. RT: room temperature (control) group; CE: cold exposure group. IWAT, inguinal adipose tissue.
Discussion
In the principal component analysis of control group, PC1 axis distinguished between BAT and white adipose tissue (WAT) (IWAT and EWAT). PC2 axis distinguished between IWAT and EWAT. On the other hand, the adipose tissues treated with CL316243 were not distinguished by PC1 and PC2 axes, suggesting that metabolic profile of WAT is changed by browning. IWAT in mice treated with CL316243 was not only the maximum change ratio of Ucp-1 mRNA expression but also maximum change of metabolites among IWAT, EWAT, and BAT. In the previous study, it has been reported that an administration of CL316243 on mice reduces WAT weight and adipocyte size, and these phenomena were partly abolished by UCP-1 KO (27), supporting our findings. Furthermore, we analyzed the effect of CL316243 on metabolism in adipose tissue and revealed that the maximum number of significant different peak between control and CL316243 treatment group is IWAT in each adipose tissue and plasma. Our data suggested that the effect of CL316243 on nucleic acids metabolism of IWAT and EWAT is larger than that of BAT. Further investigations are needed in order to reveal the difference of the effect of CL316243 on these adipose tissues. We showed that BCAAs are increased by CL316243 treatment. It has been reported that BCAAs contributes to thermogenesis via UCP-1 (14), supporting our data. CL316243 is an adrenergic-β3 receptor agonist and contributes to enhancement of lipid metabolism. Our metabolome data showed that lipid changes take significant proportion in the decreased metabolites, suggesting the effect of CL316243 on lipid metabolism.
It was demonstrated that adrenergic agonist induces not only upregulation of Ucp-1 mRNA but also that of IMP in adipocyte. Previous study has been reported that adrenergic agonist increases Ucp-1 mRNA expression via upregulation of cAMP concentration in adipocyte (6, 28, 29), supporting our data. Our study showed that IMP in adipocytes is also increased by adrenergic agonist, suggesting the participation of IMP in regulation of UCP-1 expression. To test this hypothesis, MPA was used to evaluate the effect of IMP dehydrogenase (IMPDH) on Ucp-1 mRNA expression in adipocyte. IMPDH catalyzes the oxidation of IMP to xanthosine monophosphate, which is the pivotal step in the biosynthesis of guanine nucleotides (30, 31, 32, 33). It is well known that MPA is a potent and specific inhibitor of IMPDH (34, 35). Our study revealed that MPA contributes to upregulation of cAMP and Ucp-1 mRNA expression in murine adipocytes. These data supported our hypothesis. Furthermore, we demonstrated that the mRNA expression level of Impdh-1 in IWAT is decreased by cold exposure, supporting the hypothesis that the inhibition of IMPDH activity increases UCP-1 expression. This finding suggests the existence of a regulation mechanism of UCP-1 expression in adipose tissue via IMPDH under physiological condition.
On the other hand, it remains unknown whether IMP has a direct and/or indirect action of Ucp-1 mRNA upregulation. At first, we tried to decrease the mRNA expression level of Impdh using the immortalized adipocytes and siRNA knockdown approach. However, the knockdown efficiency was very low using the immortalized adipocytes. Next, we investigated the direct effect of Impdh-1 and Impdh-2 gene on adipocyte browning using HB2 cells and siRNA knockdown approach. HB2 cells are pre–brown adipocyte originated from p53 KO mice and useful for in vitro studies of BAT and UCP functions (36). HB2 cells differentiate into brown adipocytes after induction of differentiation. Although this alternative method solved the problem of knockdown efficiency, the mRNA expression of Ucp-1 in HB2 adipocytes was decreased by Impdh-1 and Impdh-2 double knockdown, suggesting that Ucp-1 expression level is influenced by IMPDH activity. However, these data does not support our data including the immortalized adipocytes and administration of MMF to mice. The results using the immortalized adipocytes provide support for the results of in vivo experiments. The immortalized adipocytes obtained from WAT differentiate into beige/brite cells (37). On the other hand, HB2 cells were obtained from BAT (36). Therefore, the difference between HB2 adipocyte’s data and the immortalized adipocyte’s data might be attributed to differences in the cell type. In the previous study, it has been reported that adenosine activates BAT and beige adipocytes via adenosine receptor (38). Adenosine is one of metabolites of IMP. Therefore, we investigated the effect of MMF administration on adenosine level in adipose tissue in order to elucidate the molecular mechanisms underlying IMPDH-regulated adipocyte browning. MMF had no effect on adenosine level in adipose tissue, suggesting that MMF induces the expression of Ucp-1 via adenosine-independent pathway. T1R families and adenylosuccinate synthetic pathway may be the possible involvement in the molecular mechanisms underlying IMPDH-regulated adipocyte browning. T1R1/T1R3, the members of the T1R families of G protein–coupled receptors and known as taste receptor, responds to the umami taste stimulus L-glutamate, and this response is enhanced by IMP (39). In addition, it has been reported that T1R2 and T1R3 are expressed throughout adipogenesis and in adipose tissues (40). The findings of the previous studies and our study leave open the possibility of the involvement of T1R families in browning. Furthermore, IMP is metabolized to xanthosine monophosphate or adenylosuccinate via IMPDH or adenylosuccinate synthetase, respectively. Adenylosuccinate is substrate for AMP, which is indirect source of ATP and cAMP. Therefore, it may be also assumed that IMPDH inhibition promotes the influx of IMP to adenylosuccinate synthetic pathway, which induces the regulation of UCP-1 expression.
We showed that an administration of MMF to mice induces upregulation of Ucp-1 mRNA and inhibition of cell size and tissue weight in white/beige adipose tissue. We showed that the mRNA expressions of thermogenesis and brown/beige adipocyte markers are not influenced by MMF treatment. The sensitivity of Ucp-1 to browning is higher than that of other markers (41). Therefore, it is suggested that the short-term MMF administration in this study has a significant effect on only Ucp-1 and has no effect on other markers. In addition, we also showed that the mRNA expressions of general adipogenesis markers were not influenced by MMF treatment, suggesting that the short-term MMF administration in this study has no effect on general adipogenesis. The result of food intake and plasma chemical analysis suggested that the short-term MMF administration in this study has little effect on liver and plasma metabolism and there are no major safety concerns. MPA is the active form of the ester prodrug MMF. Although several previous studies have been reported that MPA has antiviral and anticancer activity in cell culture models (42, 43), the efficacy of MPA in vivo appears to be limited (35, 36). MPA eventually reached the clinic as an immunosuppressive drug for the prevention of transplant rejection in the form of MMF (35, 44, 45). In this study, we revealed that MMF has abilities to induce Ucp-1 mRNA expression, miniaturize adipocyte size, and reduce adipose tissue weight in mice. These findings are new effect of MMF, to our knowledge, and suggest that IMPDH inhibitor including MMF opens the possibility of new effective therapy of obesity-related diseases. Although MMF is a prodrug of MPA and a specific inhibitor of IMPDH (34, 35), MMF might increase UCP-1 expression via not only IMPDH inhibition but also other mechanism of action. In order to understand the effect of MMF on antiobesity in human, further investigations are needed.
It is likely that other metabolic pathways, except for purine metabolism, regulate Ucp-1 mRNA expression, cell size, and tissue weight in adipose tissue. In the present study, we analyzed the metabolites that were estimated by KEGG and mzCloud database. Other metabolites may contribute to regulation of UCP-1 expression level in adipocytes. Further investigations are needed in order to reveal the effect of other metabolic pathway, except for purine metabolism, on the regulation of UCP-1 expression and cell size in adipose tissue.
In conclusion, our study demonstrated the involvement of IMP metabolism in the regulation of UCP-1 expression and cell size in adipose tissue and provided new insight, to our knowledge, into the effect of MMF on browning in adipose tissue. These findings suggest that IMP metabolism is a potential therapeutic target for antiobesity.
Experimental procedures
Material and chemicals
All chemicals were obtained from Wako, Nacalai tesque, and Sigma. Buffers used were of HPLC or LC-MS grade.
Animal experiment
The animal experiments were performed as described in previous studies (46, 47) with some modifications. Male C57BL/6J mice (10 weeks) were purchased from CLEA Japan and maintained for 7 days on a standard diet. These mice were kept in individual cages in a temperature-controlled room at 23 °C ± 1 deg. C and maintained under a constant 12 h light/dark cycle. All the animal experiments were approved by the Kyoto University Animal Care Committee (NO. 31-62). In the experiment of administration of CL316243 to mice, the mice were divided into two groups of similar average body weights and then each group was administered of saline solution (control group) or CL316243 (1 mg/kg body weight/day, CL316243 treatment group) for 1 week. In the feeding experiment of MMF, the mice were divided into two groups and then each group was maintained on a standard diet (control group) and standard diet containing 0.2% (w/w) MMF for 3 days. In the experiment of cold exposure, male C57BL/6J mice (6 weeks) were divided into two groups of similar average body weights and then each group were kept in individual cages in a temperature-controlled room at 23 °C ± 1 deg. C (RT group) or 4 °C (CE group) for 24 h.
Histological analysis
Histological analysis was performed as described in previous studies (48, 49) with some modifications. Briefly, tissues were excised from each mouse and fixed in 4% (v/v) paraformaldehyde/PBS. After ethanol dehydration, the fixed samples were embedded in paraffin, cut into 5 μm sections with a microtome, and mounted on microscope slides (Matsunami Glass). The sections were stained with modified Mayer’s hematoxylin (Merck) and eosin Y (Wako Pure Chemical Industries Ltd). For immunohistochemistry analysis, the sections were incubated in 1% (v/v) hydrogen peroxide in methanol and treated with 10% (v/v) normal goat serum, rabbit anti-UCP1 (U6382; 1:200; Sigma–Aldrich), goat anti-rabbit IgG (Nichirei), and avidin-biotin-peroxidase complex (Nichirei).
Plasma chemical analysis
All plasma chemical indexes were measured by kit according to the manufacturer’s instructions. Plasma glucose, TG, and NEFA level were determined with glucose CII-test Kit (Wako), triglyceride E-test Kit (Wako), and NEFA C-test Kit (Wako), respectively. Plasma GOT and GPT were determined with transaminase CII-test Kit (Wako).
Metabolomic analysis
LC-MS–based metabolome analysis and sample extracts were performed as described in a previous study (17, 18, 50) with some modifications. Briefly, plasma and adipose tissue were extracted by 80% methanol (n = 4–7/group). LC-MS was performed using a HPLC system coupled to an LTQ Orbitrap XL-MS system (Thermo Fisher Scientific) equipped with an electrospray source operating in the positive ion mode. An aliquot of the extracted sample was injected into an Inertsil ODS-4 reversed-phase column (column size, 3.0 × 250 mm; particle size, 3.0 μm; GL Sciences Inc). Mobile phases A (0.1% formic acid) and B (acetonitrile including 0.1% formic acid) were used. The buffer gradient consisted of 5.0% B for 0.0 to 5.0 min, 5.0 to 95.0% B for 5.0 to 30.0 min, 95.0% B for 30.0 to 45.0 min, 95.0 to 5.0% B for 45.0 to 46.0 min, and 5.0% B for 14.0 min before the next injection, at a flow rate of 400 μl/min. These data were acquired using Compound Discoverer 2.1 (Thermo Fisher Scientific) linked to mzCloud database (Thermo Fisher Scientific) and KEGG database. Compound Discoverer 2.1 is a software package for detection and annotation of peaks obtained using LC-MS. The value of peak area was used to calculate the rate of change between different groups. In the processing of predictive composition and KEGG search, the mass tolerance was set at 5 ppm. In the processing of mzCloud search, the precursor mass tolerance and the fragment mass tolerance were set at 10 ppm and 0.4 Da, respectively.
Cell culture
Immortalized adipocytes were kindly provided by Shingo Kajimura, PhD (37). Immortalized adipocytes were cultured in a growth medium comprising Dulbecco's modified Eagle's medium (DMEM), supplemented with fetal bovine serum (10% v/v), penicillin (100 units/ml), and streptomycin (100 μg/ml), at 37 °C under CO2 (5%). The cells were cultured in a differentiation medium containing insulin (5 μg/ml), triiodothyronine (1 nM), indomethacin (0.125 mM), dexamethasone (2 μg/ml), rosiglitazone (0.5 μM), and IBMX (0.5 mM) in growth medium and incubated for 48 h (day 0–2). On the second day (day 2), the medium was replaced with a fresh regular growth medium containing insulin (5 μg/ml) and triiodothyronine (1 nM). On the fourth and sixth day, the medium was replaced with fresh regular growth medium. Iso (1 μM) or MPA (10 μM) were used for treatment for 4 h or 24 h, respectively, before total RNA and IMP extraction. Total RNA and IMP in immortalized adipocytes were extracted at day 8. Iso or MPA were treatment for 15 min or 5 min, respectively, before the measurement of cAMP in immortalized adipocytes at day 8.
HB2 cells were kindly provided by Masayuki Saito, PhD (36). HB2 cells which are brown adipocyte cell lines induced to develop into brown adipocytes by treating with 1 μM dexamethasone and 0.5 mM IBMX for 48 h (day 0–2) (induction medium). The cells were maintained in DMEM containing 10 μg/ml insulin and 50 nM triiodothyronine (maintenance medium). On the fourth day, the medium was replaced with fresh maintenance medium.
RNA preparation and quantification of gene expression
The experiment of gene expression was performed as described in previous studies (51, 52) with some modifications. Total RNA was extracted from adipose tissue and immortalized adipocytes using Sepasol-RNA I Super reagent (Nacalai Tesque), in accordance with the manufacturer’s protocol. Total RNA was reverse transcribed using M-MLV reverse transcriptase (Promega Corporation). To quantify the mRNA expression, real-time PCR was performed using a Light cycler system (Roche Diagnostics) and SYBR Green (TOYOBO CO, LTD). The oligonucleotide primer sets of genes were designed as follows: mouse Ucp-1 (forward [Fwd]: 5′- CAAAGTCCGCCTTCAGATCC-3′; reverse [Rev]: 5′- AGCCGGCTGAGATCTTGTTT-3′), mouse Impdh-1 (Fwd: 5′-GTGTCTCCGGTTCCATCCAG-3′; Rev: 5′-GCCGCTTCTCGTAAGAGTGT-3′), mouse Impdh-2 (Fwd: 5′-AGGTCATTGGAGGCAATGTAGTC-3′; Rev: 5′-ATAGCAGCCCGAGACAGTAG-3′), mouse Pparγ (Fwd: 5′-GGAGATCTCCAGTGATATCGACCA-3′; Rev: 5′-ACGGCTTCTACGGATCGAAACT-3′), mouse aP2 (Fwd: 5′-AAGACAGCTCCTCCTCGAAGGTT-3′; Rev: 5′-TGACCAAATCCCCATTTACGC-3′), mouse Adiponectin (Fwd: 5′-TACAACCAACAGAATCATTATG-3′; Rev: 5′-GAAAGCCAGTAAATGTAGAGTCGTTGA-3′), mouse Pgc1α (Fwd: 5′-CCCTGCCATTGTTAAGACC-3′; Rev: 5′-TGCTGCTGTTCCTGTTTTC-3′), mouse Cidea (Fwd: 5′-ATCACAACTGGCCTGGTTACG-3′; Rev: 5′-TACTACCCGGTGTCCATTTCT-3′), mouse Prdm16 (Fwd: 5′-CAGCACGGTGAAGCCATTC-3′; Rev: 5′-GCGTGCATCCGCTTGTG-3′), and mouse 36B4 as an internal control (Fwd: 5′-TCCTTCTTCCAGGCTTTGGG-3′; Rev: 5′-GACACCCTCCAGAAAGCGAG-3′). All data indicating mRNA expression levels are presented as a ratio relative to a control in each experiment.
RNAi
Six days after the differentiation induction, HB2 cells were transiently transfected using the transfection reagent, Lipofectamine RNAiMAX (Invitrogen). Impdh-1 (Silencer Select siImpdh1, s76613, Thermo Fisher), Impdh-2 (Silencer Select siImpdh2, s76617, Thermo Fisher), and a nontargeted control siRNA (MISSION siRNA universal negative control, Sigma–Aldrich) were used. The siRNA molecules (4 pmol/well) and 1.2 μl/well of Lipofectamine were first diluted in 100 μl/well of reduced serum medium (Opti-MEM; Invitrogen) and then mixed in the 24-well cell culture plate (IWAKI). The mixture was incubated for 20 min at room temperature and 300 μl/well of cell suspension containing 1.2 × 105 cells/well of differentiated HB2 were added to the siRNA mixture. After 48 h of transfection, media containing siRNA were removed and fresh DMEM was added. The next day, cells were harvested for later analysis.
Quantification of IMP, adenosine, and cAMP in immortalized adipocytes
About 80% methanol was used for extraction of IMP and adenosine. LC-MS for IMP and adenosine quantification analysis was performed using a Acquity UPLC system coupled to a Xevo QTOF-MS system (Waters), equipped with an electrospray source operating. The capillary, sampling cone, and extraction cone voltages were set at 1000, 10, and 1.0 V, respectively. The source and desolvation temperatures were 120 °C and 450 °C, respectively. The cone and desolvation gas flow rates were set at 50 and 800 l/h, respectively. An aliquot of the extracted sample (3 μl) was injected into an Acquity UPLC BEH-C18 reversed phase column (column size, 2.1 × 100 mm; particle size, 1.7 μm). Mobile phases A and B were used. The column temperature was set at 40 °C. The buffer gradient consisted of 0.1% to 30.0% B for 0 to 7 min, 30.0% to 99.0% B for 7 to 7.1 min, 99.0% B for 7.1 to 10.75 min, 99.0% to 0.1% B for 10.75 to 11 min, and 0.1% B for 11 to 15 min before the next injection, at a flow rate of 250 μl/min. These data were acquired with the MassLynx software (Waters).
The cAMP content was enzymatically determined with a cAMP-Glo max assay kit (Promega) according to the manufacturer’s instructions.
Statistical analysis
The data are presented as the mean ± SD. Data were analyzed by Student’s t test. Differences were considered significant at p < 0.05.
Data availability
All data that support the findings of this study are included in this article and Supplemental Information.
Supporting information
This article contains supporting information.
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Acknowledgments
The authors thank Ms R. Takahashi for secretarial support. The authors also thank Mr M. Komori for technical support. This work was supported in part by Japan Society for the Promotion of Science (JSPS) Grants-in-Aid for Scientific Research (Grant Number 21K05486, 18K14420, 19H02910, 20K21755, and 16H02551) and Japan Foundation for Applied Enzymology (Front Runner of Future Diabetes Research).
Author contributions
H. T. conceptualization; H. T. methodology; H. T., M. T., S. K., H. N., M. I., K. N., H. O., S. M., T. I., T. A., H.-F. J., W. N., K. I., and T. G. validation; H. T., M. T., S. K., H. N., M. I., K. N., and H. O. formal analysis; H. T., M. T., S. K., H. N., M. I., K. N., H. O., S. M., T. I., T. A., H.-F. J., and W. N. investigation; H. T., M. T., S. K., H. N., M. I., K. N., and H. O. resources; H. T., M. T., S. K., H. N., M. I., K. N., and H. O. data curation; H. T. writing–original draft; T. G. writing–review & editing; H. T. visualization; T. K., K. I., and T. G. supervision; T. G. project administration; H. T., T. K., and T. G. funding acquisition.
Edited by Qi-Qun Tang
Supporting information
References
- 1.Cannon B., Nedergaard J. Brown adipose tissue: function and physiological significance. Physiol. Rev. 2004;84:277–359. doi: 10.1152/physrev.00015.2003. [DOI] [PubMed] [Google Scholar]
- 2.Nedergaard J., Bengtsson T., Cannon B. Unexpected evidence for active brown adipose tissue in adult humans. Am. J. Physiol. Endocrinol. Metab. 2007;293:E444–E452. doi: 10.1152/ajpendo.00691.2006. [DOI] [PubMed] [Google Scholar]
- 3.Sidossis L., Kajimura S. Brown and beige fat in humans: thermogenic adipocytes that control energy and glucose homeostasis. J. Clin. Invest. 2015;125:478–486. doi: 10.1172/JCI78362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Nagase I., Yoshida T., Kumamoto K., Umekawa T., Sakane N., Nikami H., et al. Expression of uncoupling protein in skeletal muscle and white fat of obese mice treated with thermogenic beta 3-adrenergic agonist. J. Clin. Invest. 1996;97:2898–2904. doi: 10.1172/JCI118748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kajimura S., Saito M. A new era in brown adipose tissue biology: molecular control of brown fat development and energy homeostasis. Annu. Rev. Physiol. 2014;76:225–249. doi: 10.1146/annurev-physiol-021113-170252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Harms M., Seale P. Brown and beige fat: development, function and therapeutic potential. Nat. Med. 2013;19:1252–1263. doi: 10.1038/nm.3361. [DOI] [PubMed] [Google Scholar]
- 7.Altshuler-Keylin S., Shinoda K., Hasegawa Y., Ikeda K., Hong H., Kang Q., et al. Beige adipocyte maintenance is regulated by autophagy-induced mitochondrial clearance. Cell Metab. 2016;24:402–419. doi: 10.1016/j.cmet.2016.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yoneshiro T., Saito M. Activation and recruitment of brown adipose tissue as anti-obesity regimens in humans. Ann. Med. 2015;47:133–141. doi: 10.3109/07853890.2014.911595. [DOI] [PubMed] [Google Scholar]
- 9.Yoneshiro T., Aita S., Matsushita M., Kayahara T., Kameya T., Kawai Y., et al. Recruited brown adipose tissue as an antiobesity agent in humans. J. Clin. Invest. 2013;123:3404–3408. doi: 10.1172/JCI67803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hanatani S., Motoshima H., Takaki Y., Kawasaki S., Igata M., Matsumura T., et al. Acetate alters expression of genes involved in beige adipogenesis in 3T3-L1 cells and obese KK-Ay mice. J. Clin. Biochem. Nutr. 2016;59:207–214. doi: 10.3164/jcbn.16-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kajimura S., Seale P., Kubota K., Lunsford E., Frangioni J.V., Gygi S.P., et al. Initiation of myoblast to brown fat switch by a PRDM16-C/EBP-beta transcriptional complex. Nature. 2009;460:1154–1158. doi: 10.1038/nature08262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ohno H., Shinoda K., Spiegelman B.M., Kajimura S. PPARγ agonists induce a white-to-brown fat conversion through stabilization of PRDM16 protein. Cell Metab. 2012;15:395–404. doi: 10.1016/j.cmet.2012.01.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Shinoda K., Ohyama K., Hasegawa Y., Chang H.-Y., Ogura M., Sato A., et al. Phosphoproteomics identifies CK2 as a negative regulator of beige adipocyte thermogenesis and energy expenditure. Cell Metab. 2015;22:997–1008. doi: 10.1016/j.cmet.2015.09.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Yoneshiro T., Wang Q., Tajima K., Matsushita M., Maki H., Igarashi K., et al. BCAA catabolism in brown fat controls energy homeostasis through SLC25A44. Nature. 2019;572:614–619. doi: 10.1038/s41586-019-1503-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kwan H.Y., Wu J., Su T., Chao X.J., Liu B., Fu X., et al. Cinnamon induces browning in subcutaneous adipocytes. Sci. Rep. 2017;7:2447. doi: 10.1038/s41598-017-02263-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Soga T., Baran R., Suematsu M., Ueno Y., Ikeda S., Sakurakawa T., et al. Differential metabolomics reveals ophthalmic acid as an oxidative stress biomarker indicating hepatic glutathione consumption. J. Biol. Chem. 2006;281:16768–16776. doi: 10.1074/jbc.M601876200. [DOI] [PubMed] [Google Scholar]
- 17.Takahashi H., Goto T., Yamazaki Y., Kamakari K., Hirata M., Suzuki H., et al. Metabolomics reveal 1-palmitoyl lysophosphatidylcholine production by peroxisome proliferator-activated receptor α. J. Lipid Res. 2015;56:254–265. doi: 10.1194/jlr.M052464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Takahashi H., Sanada K., Nagai H., Li Y., Aoki Y., Ara T., et al. Over-expression of PPARα in obese mice adipose tissue improves insulin sensitivity. Biochem. Biophys. Res. Commun. 2017;493:108–114. doi: 10.1016/j.bbrc.2017.09.067. [DOI] [PubMed] [Google Scholar]
- 19.Iijima Y., Nakamura Y., Ogata Y., Tanaka K., Sakurai N., Suda K., et al. Metabolite annotations based on the integration of mass spectral information. Plant J. 2008;54:949–962. doi: 10.1111/j.1365-313X.2008.03434.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Akimoto N., Ara T., Nakajima D., Suda K., Ikeda C., Takahashi S., et al. FlavonoidSearch: a system for comprehensive flavonoid annotation by mass spectrometry. Sci. Rep. 2017;7:1243. doi: 10.1038/s41598-017-01390-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sakurai N., Mardani-Korrani H., Nakayasu M., Matsuda K., Ochiai K., Kobayashi M., et al. Metabolome analysis identified okaramines in the soybean rhizosphere as a legacy of hairy vetch. Front. Genet. 2020;11:114. doi: 10.3389/fgene.2020.00114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Bijlsma S., Bobeldijk I., Verheij E.R., Ramaker R., Kochhar S., Macdonald I.A., et al. Large-scale human metabolomics studies: a strategy for data (pre-) processing and validation. Anal. Chem. 2006;78:567–574. doi: 10.1021/ac051495j. [DOI] [PubMed] [Google Scholar]
- 23.Crews B., Wikoff W.R., Patti G.J., Woo H.-K., Kalisiak E., Heideker J., et al. Variability analysis of human plasma and cerebral spinal fluid reveals statistical significance of changes in mass spectrometry-based metabolomics data. Anal. Chem. 2009;81:8538–8544. doi: 10.1021/ac9014947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Schweizer S., Liebisch G., Oeckl J., Hoering M., Seeliger C., Schiebel C., et al. The lipidome of primary murine white, brite, and brown adipocytes-impact of beta-adrenergic stimulation. PLoS Biol. 2019;17 doi: 10.1371/journal.pbio.3000412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Mills E.L., Pierce K.A., Jedrychowski M.P., Garrity R., Winther S., Vidoni S., et al. Accumulation of succinate controls activation of adipose tissue thermogenesis. Nature. 2018;560:102–106. doi: 10.1038/s41586-018-0353-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fujimoto Y., Hashimoto O., Shindo D., Sugiyama M., Tomonaga S., Murakami M., et al. Metabolic changes in adipose tissues in response to β3 -adrenergic receptor activation in mice. J. Cell. Biochem. 2019;120:821–835. doi: 10.1002/jcb.27443. [DOI] [PubMed] [Google Scholar]
- 27.Inokuma K., Okamatsu-Ogura Y., Omachi A., Matsushita Y., Kimura K., Yamashita H., et al. Indispensable role of mitochondrial UCP1 for antiobesity effect of beta3-adrenergic stimulation. Am. J. Physiol. Endocrinol. Metab. 2006;290:E1014–E1021. doi: 10.1152/ajpendo.00105.2005. [DOI] [PubMed] [Google Scholar]
- 28.Nguyen K.D., Qiu Y., Cui X., Sharon Goh Y.P., Mwangi J., David T., et al. Alternatively activated macrophages produce catecholamines to sustain adaptive thermogenesis. Nature. 2011;480:104–108. doi: 10.1038/nature10653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Collins S. β-Adrenoceptor signaling networks in adipocytes for recruiting stored fat and energy expenditure. Front. Endocrinol. (Lausanne) 2012;2:102. doi: 10.3389/fendo.2011.00102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Crabtree G.W., Henderson J.F. Rate-limiting steps in the interconversion of purine ribonucleotides in Ehrlich ascites tumor cells in vitro. Cancer Res. 1971;31:985–991. [PubMed] [Google Scholar]
- 31.Snyder F.F., Henderson J.F., Cook D.A. Inhibition of purine metabolism--computer-assisted analysis of drug effects. Biochem. Pharmacol. 1972;21:2351–2357. doi: 10.1016/0006-2952(72)90386-3. [DOI] [PubMed] [Google Scholar]
- 32.Jackson R.C., Weber G., Morris H.P. IMP dehydrogenase, an enzyme linked with proliferation and malignancy. Nature. 1975;256:331–333. doi: 10.1038/256331a0. [DOI] [PubMed] [Google Scholar]
- 33.Weber G. Biochemical strategy of cancer cells and the design of chemotherapy: G. H. A. Clowes Memorial Lecture. Cancer Res. 1983;43:3466–3492. [PubMed] [Google Scholar]
- 34.Farazi T., Leichman J., Harris T., Cahoon M., Hedstrom L. Isolation and characterization of mycophenolic acid-resistant mutants of inosine-5'-monophosphate dehydrogenase. J. Biol. Chem. 1997;272:961–965. doi: 10.1074/jbc.272.2.961. [DOI] [PubMed] [Google Scholar]
- 35.Hedstrom L. IMP dehydrogenase: structure, mechanism, and inhibition. Chem. Rev. 2009;109:2903–2928. doi: 10.1021/cr900021w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Irie Y., Asano A., Cañas X., Nikami H., Aizawa S., Saito M. Immortal brown adipocytes from p53-knockout mice: differentiation and expression of uncoupling proteins. Biochem. Biophys. Res. Commun. 1999;255:221–225. doi: 10.1006/bbrc.1998.9999. [DOI] [PubMed] [Google Scholar]
- 37.Aune U.L., Ruiz L., Kajimura S. Isolation and differentiation of stromal vascular cells to beige/brite cells. J. Vis. Exp. 2013 doi: 10.3791/50191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Gnad T., Scheibler S., von Kügelgen I., Scheele C., Kilić A., Glöde A., et al. Adenosine activates brown adipose tissue and recruits beige adipocytes via A2A receptors. Nature. 2014;516:395–399. doi: 10.1038/nature13816. [DOI] [PubMed] [Google Scholar]
- 39.Li X., Staszewski L., Xu H., Durick K., Zoller M., Adler E. Human receptors for sweet and umami taste. Proc. Natl. Acad. Sci. U. S. A. 2002;99:4692–4696. doi: 10.1073/pnas.072090199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Simon B.R., Parlee S.D., Learman B.S., Mori H., Scheller E.L., Cawthorn W.P., et al. Artificial sweeteners stimulate adipogenesis and suppress lipolysis independently of sweet taste receptors. J. Biol. Chem. 2013;288:32475–32489. doi: 10.1074/jbc.M113.514034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.An J.-Y., Jheng H.-F., Nagai H., Sanada K., Takahashi H., Iwase M., et al. A phytol-enriched diet activates PPAR-α in the liver and brown adipose tissue to ameliorate obesity-induced metabolic abnormalities. Mol. Nutr. Food Res. 2018;62 doi: 10.1002/mnfr.201700688. [DOI] [PubMed] [Google Scholar]
- 42.Franklin T.J., Jacobs V., Jones G., Plé P., Bruneau P. Glucuronidation associated with intrinsic resistance to mycophenolic acid in human colorectal carcinoma cells. Cancer Res. 1996;56:984–987. [PubMed] [Google Scholar]
- 43.Papageorgiou C. Enterohepatic recirculation: a powerful incentive for drug discovery in the inosine monophosphate dehydrogenase field. Mini Rev. Med. Chem. 2001;1:71–77. doi: 10.2174/1389557013407269. [DOI] [PubMed] [Google Scholar]
- 44.Sollinger H.W. Mycophenolate mofetil for the prevention of acute rejection in primary cadaveric renal allograft recipients. U.S. Renal Transplant Mycophenolate Mofetil Study Group. Transplantation. 1995;60:225–232. doi: 10.1097/00007890-199508000-00003. [DOI] [PubMed] [Google Scholar]
- 45.Demirkiran A., Sewgobind V.D.K.D., van der Weijde J., Kok A., Baan C.C., Kwekkeboom J., et al. Conversion from calcineurin inhibitor to mycophenolate mofetil-based immunosuppression changes the frequency and phenotype of CD4+FOXP3+ regulatory T cells. Transplantation. 2009;87:1062–1068. doi: 10.1097/TP.0b013e31819d2032. [DOI] [PubMed] [Google Scholar]
- 46.Evans B.A., Papaioannou M., Anastasopoulos F., Summers R.J. Differential regulation of beta3-adrenoceptors in gut and adipose tissue of genetically obese (ob/ob) C57BL/6J-mice. Br. J. Pharmacol. 1998;124:763–771. doi: 10.1038/sj.bjp.0701867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.von Vietinghoff S., Ouyang H., Ley K. Mycophenolic acid suppresses granulopoiesis by inhibition of interleukin-17 production. Kidney Int. 2010;78:79–88. doi: 10.1038/ki.2010.84. [DOI] [PubMed] [Google Scholar]
- 48.Iwase M., Tokiwa S., Seno S., Mukai T., Yeh Y.S., Takahashi H., et al. Glycerol kinase stimulates uncoupling protein 1 expression by regulating fatty acid metabolism in beige adipocytes. J. Biol. Chem. 2020;295:7033–7045. doi: 10.1074/jbc.RA119.011658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Kawarasaki S., Kuwata H., Sawazaki H., Sakamoto T., Nitta T., Kim C.S., et al. A new mouse model for noninvasive fluorescence-based monitoring of mitochondrial UCP1 expression. FEBS Lett. 2019;593:1201–1212. doi: 10.1002/1873-3468.13430. [DOI] [PubMed] [Google Scholar]
- 50.Takahashi H., Ochiai K., Sasaki K., Izumi A., Shinyama Y., Mohri S., et al. Metabolome analysis revealed that soybean-Aspergillus oryzae interaction induced dynamic metabolic and daidzein prenylation changes. PLoS One. 2021;16 doi: 10.1371/journal.pone.0254190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hara H., Takahashi H., Mohri S., Murakami H., Kawarasaki S., Iwase M., et al. β-Cryptoxanthin induces UCP-1 expression via a RAR pathway in adipose tissue. J. Agric. Food Chem. 2019;67:10595–10603. doi: 10.1021/acs.jafc.9b01930. [DOI] [PubMed] [Google Scholar]
- 52.Takahashi H., Hara H., Goto T., Kamakari K., Wataru N., Mohri S., et al. 13-Oxo-9(Z),11(E),15(Z)-octadecatrienoic acid activates peroxisome proliferator-activated receptor γ in adipocytes. Lipids. 2015;50:3–12. doi: 10.1007/s11745-014-3972-x. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data that support the findings of this study are included in this article and Supplemental Information.