Abstract
The fifth edition WHO classification of Tumors of the Central nervous system (WHO-CNS5) integrated new molecular parameters to refine CNS tumor classification. This study aimed to reclassify a retrospective cohort of adult glioma patients according to WHO-CNS5, and assess if overall survival (OS) correlated with the revised diagnosis. Further, the diagnostic impact of methylation profiling (MP) was evaluated. Adult gliomas diagnosed according to 2016 WHO-CNS (n = 226) were evaluated according to WHO-CNS5 criteria. All patients had diagnostic NGS performed. 29 patients had 850k MP performed due to challenging tumor cases. OS was analyzed using Kaplan-Meier plots and log-rank test. 19 patients were reclassified. Specifically, diffuse astrocytic glioma, IDH-wildtype, with molecular features of glioblastoma (DAG-G) were reclassified as glioblastoma (n = 15). Shifts to glioblastoma were because of TERT promoter (TERTp) mutation (n = 9), EGFR amplification (n = 2), EGFR amplification and TERTp mutation (n = 1), and TERTp mutation with gain of chromosome 7, but uncertain chromosome 10 status due to lack of NGS coverage (n = 3). Lower grade IDH-mutant astrocytomas were reclassified as astrocytoma IDH-mutant, WHO grade 4 due to CDKN2A/B homozygous deletion (n = 4). No significant difference in OS was found for reclassified DAG-G in whole group (p = 0.59) and for TERTp mutation only (p = 0.44), compared to glioblastoma. MP resulted in revised diagnosis (n = 2), confirmed diagnosis (n = 15) and no match (n = 12). Our study showed similar overall survival for glioblastoma and DAG patients, supporting that isolated TERTp mutation may have a prognostic role in IDH-wildtype gliomas. Further, our study suggests MP is useful for confirming the diagnoses in challenging tumors.
Keywords: cIMPACT-NOW, Next-generation sequencing, Retrospective reclassification, TERT promoter mutation, WHO-CNS5, DNA methylation profiling
Highlights
-
•
Retrospective cohort of adult glioma reclassified using WHO-CNS5 molecular criteria.
-
•
8.4% of the cohort received a new diagnosis and often a higher WHO grade.
-
•
TERT promoter mutation suggested as a prognostic factor in IDH wildtype gliomas.
-
•
DNA methylation profiling useful for diagnostically difficult cases.
1. Introduction
Gliomas represent a heterogenic group of primary tumors that develop from glial cells in the brain and central nervous system, where they make up approximately 30% of all diagnosed tumors, and 80% of all malignant brain tumors (Goodenberger and Jenkins, 2012). Glioblastomas are the most common malignant brain tumor in adults with a median survival of around 14 months even with treatment (Stupp et al., 2005, Wen et al., 2020).
In 2016, the WHO Classification of Tumors of the Central Nervous system (WHO-CNS) introduced the principle of integrated diagnostics for brain tumors. Diagnosis of diffuse gliomas was defined as a result of not only histopathological findings, but also three additional molecular genetic alterations: mutations in the isocitrate dehydrogenase gene 1 and 2 (IDH1 and IDH2) and co-deletion of chromosomal arms 1p and 19q (Louis et al., 2016). This resulted in glioblastoma being split into glioblastoma IDH-wildtype and glioblastoma IDH-mutant, among many others (Louis et al., 2016).
Previously, the malignant potential of a tumor was graded based on histopathological evaluation of morphologic characteristics, such as nuclear atypia, necrosis, microvascular proliferation, and cellular proliferation (Louis et al., 2007, Cohen et al., 2013). In 2018 the Consortium to Inform Molecular and Practical Approaches to CNS Tumor Taxonomy (cIMPACT-NOW) released their third update, wherein they recommended IDH-wildtype diffuse astrocytic tumors with either TERT promoter mutation, EGFR gene amplification or a combination of gain of entire chromosome 7 and loss of entire chromosome 10 (+7/−10) should be considered WHO grade 4 due to much more aggressive behavior, despite lacking the typical histological features (Brat et al., 2018). This entity was named diffuse astrocytic glioma, IDH wildtype, with molecular features of glioblastoma, WHO grade 4 (DAG-G). In cIMPACT-NOW update 6, with sufficient clinical data confirming that survival for DAG-G is similar to glioblastomas IDH-wildtype, the following diagnostic criteria were suggested for glioblastoma: microvascular proliferation, or necrosis, or one (or more); TERT promoter mutation, EGFR amplification or + 7/− 10. These suggestions were incorporated into the fifth edition of WHO-CNS (WHO-CNS5) (Louis et al., 2021). Similarly, in WHO-CNS5, the grading of IDH-mutant astrocytomas was expanded by CDKN2A/B homozygous deletion. Specifically, the diagnosis of astrocytoma IDH mutant, WHO grade 4 (previously named glioblastoma, IDH-mutant) is based on necrosis or microvascular proliferation or CDKN2A/B homozygous deletion (Louis et al., 2021).
DNA methylation profiling is another method of classifying tumors. While more commonly used in pediatric settings, for adults, it is mostly used in cases with ambiguous histological and genetic profiling, or when there are contradictory results (Priesterbach-Ackley et al., 2020). DNA methylation is an epigenetic modification that affects gene activity. The methylation status of a wide range of CPG sites can be assessed using 850k methylation array (Moran et al., 2016). The results are then compared to the brain tumor methylation classifier developed by Heidelberg University, which outputs a diagnosis based on the methylation profile (Capper et al., 2018a).
Classification of CNS tumors is advancing rapidly, with genetic profiling allowing us to more narrowly define diagnostic entities, better predict patients’ outcome and optimize treatment strategies. In this study we aimed to reclassify a retrospective cohort according to the WHO-CNS5 criteria in order to evaluate the prognostic impact of the new criteria. Further, we evaluated the diagnostic impact and benefit of DNA methylation profiling in a subset of our cohort using 850k methylation array.
2. Methods and materials
2.1. Study population
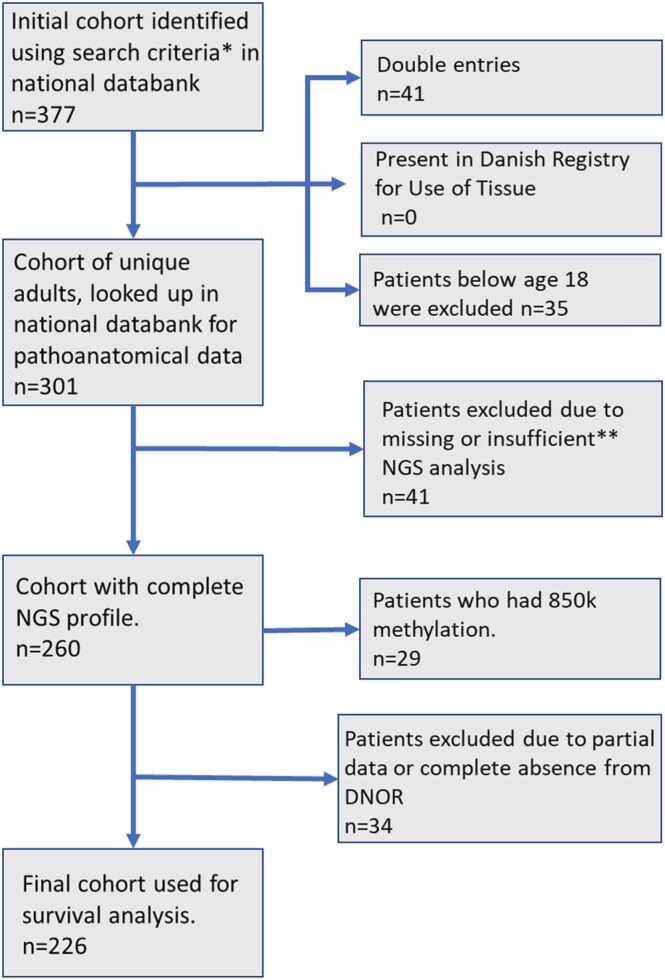
This study investigated adult glioma patients, who had been examined and diagnosed according to the 2016 WHO-CNS and cIMPACT-NOW update 3 at the Department of Pathology, Aarhus University Hospital, during the period of October 1, 2017 (the date a NGS panel was introduced into the routine examination of glioma patients) to December 31, 2019. The study population was identified using Patobank, a Danish national database containing data from every pathoanatomic examination performed in any of the Departments of Pathology in Denmark. The study was approved by the Central Denmark Region Committees on Health Research Ethics. A defined set of a glioma subtypes were included in this study: astrocytomas (including pilocytic astrocytoma, diffuse astrocytoma and anaplastic astrocytoma), oligodendrogliomas (including oligodendroglioma and anaplastic oligodendroglioma), glioblastomas (including glioblastoma, gliosarcoma, and diffuse astrocytic gliomas, IDH wildtype, with molecular features of glioblastoma), ependymomas, diffuse midline gliomas, and xanthoastrocytomas. Several other rare subtypes were included, but no cases were found for the given time period. A search was done in Patobank using the SNOMED codes corresponding to the abovementioned glioma subtypes, and for the given time period. A total of 377 cases were found, but after sorting for patients with double entries (n = 41), presence in the Danish Registry for Use of Tissue (n = 0), and patients below the age of 18 (n = 35), a total of 301 unique adults were found and included in the study. Forty-one patients were excluded due to NGS not being performed, not having tissue suitable for NGS analysis, or due to inadequate NGS analysis (poor tissue quality or low tumor percentage in the examined biopsy, making duplication and amplification assessment unreliable). Further, 34 patients were excluded due to not being identified in the Danish Neuro-oncology Registry (DNOR), which was used for obtaining various radiological and clinical data, including vital status. Thus, 226 patients were evaluated according to the WHO-CNS5 diagnostic criteria, reclassified if the criteria were met and subsequently used for survival analysis.
A subgroup of patients were also used for evaluation of the diagnostic impact of DNA methylation profiling in adult gliomas. During the diagnostic workup, some patients (n = 29) had tumor tissue submitted for DNA methylation profiling due to challenging tumor cases. A full NGS dataset was required to be included for analysis. All 29 patients had full NGS data and were included.
The above is illustrated in Fig. 1.
Fig. 1.
Illustration of how the study population was acquired. DNOR; Danish Neuro-Oncology Registry. NGS; Next-generation sequencing. *Patients examined and diagnosed at the Department of Pathology, Aarhus University Hospital, between October 1, 2017 and December 31, 2019, with SNOMED codes corresponding to astrocytomas, oligodendrogliomas, glioblastomas, ependymomas, diffuse midline gliomas and xanthoastrocytomas and their subtypes (if any). ** patients with tissue unsuitable for NGS, or inadequate NGS analysis due to poor tissue quality or low tumor percentage. DNOR was used for various radiological and clinical data, including vital status.
2.2. Histological, genetic and clinical data
Histological and genetic data was available on Patobank, which included data from the microscopic examination of the tissue by neuropathologists at Aarhus University Hospital, and genetic assessment by molecular biologists using a NGS panel spanning 28 genes, at Aarhus University Hospital. Study data were collected and managed using REDCap electronic data capture tools hosted at Aarhus University Hospital (Harris et al., 2009, Harris et al., 2019). This database was used to collect data from the microscopic examination, this included the presence of mitosis, necrosis, microvascular proliferation or nuclear pleomorphism in the biopsy, the rate of proliferation, methylation assessment of the promoter region of the MGMT gene, tumor percentage, and the outcome of DNA methylation profiling, if performed. The genetic examination consisted of a custom-made NGS panel. This panel is based on studies that have examined several of the genes, whose diagnostic and prognostic role in gliomas is most well-known (Zacher et al., 2017, Dubbink et al., 2016). The panel looks for mutations in three genes: ATRX, IDH1 (R132), IDH2 (R140 and R172), and for co-deletion of 1p19q. Furthermore, it examines the following genes in full (mutations, deletions, amplifications): CDKN2A, CDKN2B, CDKNC, CIC, EGFR, FUBP1, NF1, NF2, NOTCH 1, PTEN, RB1 and TP53 and it examines for hotspot mutations in BRAF, FGFR1, H3F3A, Hist1H3B, Hist1H3C, PIK3CA, PIK3R1, MET, NRAS, SMO and in the TERT promoter. The NGS panel did not cover entire chromosome 7, and evaluation of chromosome 7 gain was based on simultaneous gain of EGFR and MET, which were considered surrogate markers. The NGS panel also covers the following RNA fusions, but were only examined in selected cases: ZFTA-RELA, ZFTA-YAP1, YAP1-MAMLD1, PTPRZ1-MET, MYB-QKI, SRGAP3-RAF1, MYBL1 inversion, FGFR1-TACC1, QKI-NTRK2, CDK5RAP2-PDGFRA, CEP85L-ROS1, EGFR-SEPT14, NFASC-NTRK1, KIAA1549-BRAF, FGFR3-TACC3, NTRK3-ETV6 and ETV6-NTRK3. Of note, the NGS panel used in the diagnostic process of these patients did not include analysis of chromosome 10 loss. The same NGS panel was used for all patients.
An overview of the NGS findings for the different diagnostic groups are depicted in Supplementary file, Fig. 1.
Evaluation of the NGS data and gene statuses was done by an experienced clinical molecular biologist using the bioinformatics software from Thermo Fisher’s Ion reporter (version 5.6 and 5.10). For evaluation of mutations this included manual inspection of raw NGS data of all identified variants and in known problematic areas, such as the TERT promoter region – areas that were identified during the in-house clinical validation of the panel. When possible, this was combined with known tumor percentages (evaluated by a neuropathologist) and allelic frequencies of common polymorphisms. Evaluation of gene amplifications and deletions were evaluated using visual inspection of baseline-normalized coverage data in Thermo Fisher Scientific’s Ion reporter genome viewer.
The Danish Neuro-oncology Registry was used to gather clinical and radiological data for each patient. Clinical data included radiotherapy and chemotherapy treatment information, pre-operative Karnofsky performance status, gender, age at diagnosis and vital status. Radio-chemotherapy treatment was divided into completed long-course radio-chemotherapy (60 Gy and concomitant temozolomide), short-course radio-chemotherapy (40 Gy and concomitant temozolomide), radiotherapy alone, and no treatment. Data was not available on adjuvant chemotherapy treatment. Radiological data included if pre-operative MRI was performed, tumor location and the presence of contrast enhancement on pre-operative MRI. Vital status was evaluated on April 20 2021 for each patient, where the date of death (if applicable) was registered.
2.3. Diagnostic distribution
Two hundred and twenty six patients were included in the study. According to the 2016 WHO classification, the cohort was distributed as the following: Glioblastoma IDH wildtype (n = 141), DAG-G (n = 15), Anaplastic astrocytoma IDH mutant (n = 13), Diffuse astrocytoma IDH mutant (n = 11), Anaplastic oligodendroglioma (n = 10), Oligodendroglioma (n = 10), Anaplastic astrocytoma IDH wildtype (n = 8), Glioblastoma IDH mutant (n = 6), Gliosarcoma (n = 4), Giant-cell glioblastoma (n = 3), diffuse midline glioma (n = 3), Pilocytic astrocytoma (n = 1), Pleomorphic xanthoastrocytoma (n = 1). Due to lack of NGS coverage of chromosome 10, the diagnosis of DAG-G was based on either TERT promoter mutation or EGFR amplification. 1p/19q was evaluated in every patient, and 1p/19q co-deletion was not present in any of the IDH mutant astrocytomas. This is shown in Supplementary file, Fig. 1.
2.4. DNA methylation profiling
The DNA methylation data was generated using Illumina EPIC 850k methylation array (Illumina, San Diego, USA), followed by algorithmic classification by the brain tumor methylation classifier developed by Heidelberg University (from now on referred to as “The Classifier”). The Classifier outputs a classification score between 0 and 1, that indicates the degree of resemblance to known CNS tumor classes. In order to achieve valid classification using methylation profiling, patients where The Classifier returned with a calibrated score lower than 0.84, were considered as having no match, as recommended (Capper et al., 2018b).
3. Statistical analysis
Categorial variables were reported by frequencies and proportions and continuous data as medians and ranges. Select characteristics were compared using Fisher’s exact test (categorial data) or Wilcoxon rank-sum test (continuous data). Overall survival was defined as the time from primary surgery until death of any cause or date of censoring in April 2021. Overall survival was estimated by the Kaplan–Meier estimator and using the log-rank test for comparison (GraphPad software, Inc., California, USA). Results were judged significant if the p value was less than 0.05.
4. Results
4.1. Reclassification
A total of 19 patients were reclassified. Diffuse astrocytomas, IDH-mutant (n = 1) and anaplastic astrocytomas, IDH-mutant (n = 3) with CDKN2A/B homozygous deletion were reclassified as Astrocytoma, IDH mutant, WHO grade 4. A total of 24 patients with either Anaplastic Astrocytoma, IDH-mutant and Diffuse Astrocytoma, IDH-mutant were identified in the cohort. Reclassification resulted in 16.7% of IDH-mutant WHO grade 2–3 astrocytomas receiving WHO grade 4. IDH-mutant glioblastomas have been renamed Astrocytoma, IDH mutant, WHO grade 4 in the newest WHO-CNS edition.
All DAG-G patients (n = 15) were reclassified as glioblastoma IDH-wildtype due to either isolated TERT promoter mutation (n = 9), isolated EGFR amplification (n = 2), TERT promoter mutation and EGFR amplification (n = 1), or TERT promoter mutation with gain of chromosome 7, but unknown chromosome 10 status (n = 3). The NGS panel used in the diagnostic process for the included patients did not cover chromosome 10 loss. Therefore, lack of PTEN deletion, a gene which is located on chromosome 10 (Abdulkareem and Blair, 2013), was considered a surrogate marker for intact chromosome 10. However, the presence of PTEN deletion was not considered a surrogate marker for chromosome 10 loss. Two of the patients in the isolated TERT promoter mutation group had chromosome 7 gain, but no PTEN deletion, and were therefore considered to have an isolated TERT promoter mutation. The three patients with TERT promoter mutation, gain of chromosome 7, but unknown chromosome 10 status, had PTEN deletion, and were classified separately. The above is depicted in Fig. 2.
Fig. 2.
Illustration of the patients reclassified according to the WHO-CNS5 criteria. DAG-G: Diffuse astrocytic glioma, IDH wildtype, with molecular features of glioblastoma. IDH-wildtype gliomas were reclassified due to TERT promoter mutation, EGFR amplification and +7/−10. The NGS panel used in this study did not cover loss of entire chromosome 10. Intact PTEN gene was used a surrogate marker for intact chromosome 10. Patients with TERT promoter mutation, +7 and PTEN deletion were classified as having uncertain +7/−10 status. No anaplastic astrocytoma, IDH wildtype were reclassified. IDH-mutant gliomas were reclassified according to CDKN2A/B homozygous deletion status.
4.2. Survival
Due to lack of proper chromosome 10 coverage, and too few cases with isolated EGFR amplification (n = 2), only the entire DAG-G group (without stratification according to individual genetic markers), and DAG-G cases with isolated TERT promoter mutation were used for survival analysis and compared to glioblastoma, IDH wildtype. The median survival for the overall DAG-G group (n = 15) was 18.3 months, while the median survival for patients with isolated TERT promoter mutations (n = 9) was 15.8 months. Glioblastoma, IDH wildtype (n = 141) had a median survival of 13.6 months. No significant difference in overall survival was found for all DAG-G patients (p = 0.59) or DAG-G patients with isolated TERT promoter mutations (p = 0.44) compared to patients with glioblastoma, IDH wildtype. Kaplan-Meier analysis are shown in Fig. 3.
Fig. 3.
Kaplan-Meier overall survival curves. IDHwt = IDH wildtype. DAG-G = diffuse astrocytic glioma, IDH wildtype, with molecular features of glioblastoma, WHO grade 4. All DAG-G patients were reclassified as glioblastoma, IDH-wildtype, according to the WHO-CNS5 criteria. Log-rank P-values are shown in each subfigure. A) compares glioblastoma, IDH-wildtype with all DAG-G patients, without stratification according to genetic alteration. No significant difference was found (p = 0.59). B) compares glioblastoma, IDH-wildtype with DAG-G patients with isolated TERT promoter mutation. No significant difference was found in overall survival (p = 0.44).
Reclassified IDH-mutant astrocytomas only ascribed to 4 patients, which was considered too few to assess the impact of CDKN2A/B homozygous deletion on the survival for IDH-mutant astrocytomas. Therefore, no survival analysis was performed in this group.
Oligodendrogliomas, Diffuse midline glioma, Giant-cell glioblastoma, Gliosarcoma, Pleomorphic xanthoastrocytoma and pilocytic astrocytomas were not evaluated for overall survival as they were not affected by the reclassification.
Patient characteristics, including possible confounding factors, such as MGMT methylation status and CDKN2A/B homozygous deletion status is presented for the diagnostic groups included in the survival analysis in Table 1.
Table 1.
Clinical and demographic patient characteristics.
| Glioblastoma IDH wildtype | DAG-G (all) | p | DAG-G (TERT) | P | |
|---|---|---|---|---|---|
| Patients, n | 141 | 15 | 9 | ||
| Age in years (median, range) | 65 (25–88) | 67 (31–80) | 0.99 | 67 (31–80) | 0.94 |
| Gender, n (%) | |||||
| Male | 91 (64.5) | 9 (60) | 7 (77.8) | ||
| Female | 50 (35.5) | 6 (40) | 2 (22.2) | ||
| Performance status, n (%) | |||||
| 0–1 | 114 (80.9) | 11 (73.3) | 6 (66.7) | ||
| 2–4 | 23 (16.3) | 4 (26.7) | 3 (33.3) | ||
| Unknown | 4 (2.8) | 0 | 0 | ||
| MGMT methylation, n (%) | |||||
| Methylated | 58 (41.1) | 7 (46.7) | 0.41 | 3 (33.3) | 0.73 |
| Non-methylated | 77 (54.6) | 8 (53.3) | 6 (66.7) | ||
| Missing | 6 (4.3) | 0 | 0 | ||
| CDKN2A/B homozygous deletion, n (%) | 86 (61) | 6 (40) | 0.17 | 3 (33.3) | 0.16 |
| Treatment, n (%) | |||||
| Long course RCT | 86 (60.9) | 10 (66.7) | 5 (55.5) | ||
| Short course RCT | 15 (10.6) | 1 (6.6) | 1 (11.1) | ||
| RT alone | 22 (15.6) | 3 (20) | 2 (22.2) | ||
| No treatment | 18 (12.7) | 1 (6.7) | 1 (11.1) | ||
| Contrast enhancement on MRI, n (%) | |||||
| Yes | 136 (96.5) | 10 (66.6) | 6 (66.6) | ||
| No | 5 (3.5) | 5 (33.3) | 3 (33.3) | ||
| Location, n (%) | |||||
| Frontal Lobe | 28 (19.9) | 5 (33.3) | 4 (44.4) | ||
| Parietal lobe | 36 (25.5) | 3 (20) | 1 (11.1) | ||
| Occipital lobe | 8 (5.7) | 0 | 0 | ||
| Temporal lobe | 57 (40.4) | 3 (20) | 2 (22.2) | ||
| Thalamus | 2 (1.4) | 2 (13.3) | 1 (11.1) | ||
| Brainstem | 1 (0.7) | 1 (6.7) | 0 | ||
| Cerebellum | 2 (1.4) | 0 | 0 | ||
| Missing | 7 (5.0) | 1 (6.67) | 1 (11.1) | ||
| Vital status, n (%) | |||||
| Alive | 30 (21.3) | 5 (33.3) | 1 (11.1) | ||
| Dead | 111 (78.7) | 10 (66.6) | 8 (88.8) | ||
| Survival (median, months) | 13.6 | 18.3 | 0.59 | 15.8 | 0.44 |
DAG-G; Diffuse Astrocytic Glioma, with molecular features of glioblastoma, WHO grade 4.
RCT; radio-chemotherapy. RT; radiotherapy.
4.3. 850k methylation analysis
For each patient, the diagnosis provided by histopathological and genetic examination was compared to the final pathological diagnosis after methylation profiling. Evaluation was done according to WHO-CNS5 criteria, where cases with a histopathological diagnosis of DAG-G would be considered as glioblastoma, IDH-wildtype.
Using a cut-off value of 0.84 for the calibrated score, The Classifier found no methylation class match in 41.4% (n = 12) of cases, and as a result, methylation profiling did not substantially add to the diagnostic workup in these patients. In 51.7% (n = 15) of the analyzed cases, the methylation class suggested by The Classifier corresponded to and supported the original histopathological diagnosis. in 6.9% (n = 2) of cases, methylation profiling resulted in a new final pathological diagnosis. An overview for each patient displaying the histopathological diagnosis, the diagnosis suggested by 850k methylation and the Heidelberg Classifier and the impact on diagnosis is shown in Table 2.
Table 2.
Summary of diagnostic findings for patients who had 850k methylation performed.
| Patient ID | Histopathological diagnosis | 850k methylation diagnosis | Impact on diagnosis |
|---|---|---|---|
| 62 | Ganglioglioma | Anaplastic pleomorphic xanthoastrocytoma | New diagnosis |
| 73 | Unspecified low grade glial tumor | Pilocytic astrocytoma | New diagnosis |
| 7 | Diffuse astrocytoma, IDHmut | Diffuse astrocytoma, IDHmut | Concordance |
| 57 | Diffuse astrocytoma, IDHmut | Diffuse astrocytoma, IDHmut | Concordance |
| 79 | Diffuse astrocytoma, IDHmut | Diffuse astrocytoma, IDHmut | Concordance |
| 214 | Diffuse astrocytoma, IDHmut | Diffuse astrocytoma, IDHmut | Concordance |
| 152 | Pilocytic astrocytoma | Pilocytic astrocytoma | Concordance |
| 188 | Ependymoma | Ependymoma | Concordance |
| 27 | Glioblastoma, IDHwt | Glioblastoma, IDHwt | Concordance |
| 106 | Glioblastoma, IDHwt | Glioblastoma, IDHwt | Concordance |
| 123 | Glioblastoma, IDHmut | Glioblastoma, IDHmut | Concordance |
| 164 | Glioblastoma, IDHwt | Glioblastoma, IDHwt | Concordance |
| 192 | Glioblastoma, IDHwt | Glioblastoma, IDHwt | Concordance |
| 200 | Glioblastoma, IDHwt | Glioblastoma, IDHwt | Concordance |
| 226 | Glioblastoma, IDHwt | Glioblastoma, IDHwt | Concordance |
| 256 | Glioblastoma, IDHwt | Glioblastoma, IDHwt | Concordance |
| 262 | Diffuse midline glioma | Diffuse midline glioma | Concordance |
| 5 | Anaplastic astrocytoma, IDHwt | No match | None |
| 49 | Anaplastic astrocytoma, IDHwt | No match | None |
| 66 | Anaplastic Astrocytoma, IDHwt | No match | None |
| 112 | Anaplastic astrocytoma, IDHwt | No match | None |
| 228 | Anaplastic astrocytoma, IDHwt | No match | None |
| 202 | Anaplastic astrocytoma, IDHwt | No match | None |
| 28 | Glioblastoma, IDHwt | No match | None |
| 48 | Glioblastoma, IDHwt | No match | None |
| 122 | Glioblastoma, IDHwt | No match | None |
| 133 | Glioblastoma, IDHwt | No match | None |
| 210 | Glioblastoma, IDHwt | No match | None |
| 34 | Diffuse astrocytoma, IDHmut | No match | None |
Summary for each patient, showing the histopathological diagnosis and the diagnosis provided by 850k methylation and the Heidelberg Classifier. Patients where the Classifier returned with a calibrated score lower than 0.84, were considered as having no match. If the histopathological diagnosis corresponded with the Classifier diagnosis, the impact of methylation profiling was considered as “concordance”. IDHwt; IDH wildtype. IDHmut; IDH mutant.
5. Discussion
In this study we evaluated a retrospective cohort of 226 adult glioma patients according the new molecular criteria implemented in the fifth edition of WHO-CNS. Nineteen patients were reclassified. Reclassification resulted in 16.7% (n = 4) of lower grade IDH-mutant astrocytomas (WHO grade 2–3) receiving WHO grade 4, and DAG-G patients (n = 15) being reclassified as glioblastoma, IDH-wildtype. While DAG-G patients were already considered WHO grade 4 tumors, reclassifying them as glioblastoma, IDH-wildtype adds to simplify tumor classification of gliomas, better predict patient prognosis and optimize treatment strategies for individual patients. While based on a limited number of patients, our results support the prognostic impact of isolated TERT promoter mutations in DAG-G patients, showing no significant difference in overall survival compared to glioblastoma, IDH-wildtype. Similarly, no significant difference was found for the whole DAG-G group (without stratification according to genetic alteration). Further, this study showed that 6.9% (n = 2) of patients submitted for methylation profiling received a new diagnosis, while 51.7% (n = 15) had their original diagnosis supported/confirmed. In adult glioma patients, methylation profiling is mostly performed in complicated diagnostic cases, so while only a few patients received a new diagnosis, having the original diagnosis supported is also of great value, acting as double confirmation and helps prevent misdiagnoses. While methylation profiling may not be part of routine diagnostic workup in adult gliomas, it remains a valuable tool in difficult cases.
Despite being officially implemented in the WHO-CNS5, the role of TERT promoter mutation as an independent prognostic factor in gliomas remain controversial. A large number of studies have supported it as a negative independent prognostic factor, while others have not, instead, some have correlated this impact to confounding molecular and clinical factors such as age, MGMT status, EGFR amplification and IDH status (Olympios et al., 2021, Giannini and Giangaspero, 2021, Berzero et al., 2021). While the cohort of DAG-G patients was too small to perform multivariate analysis and thereby identify possible confounders, we accounted for several known and possible prognostic factors in Table 1, such as CDKN2A/B homozygous deletion, MGMT promoter methylation and performance status (Ma et al., 2020, Hegi et al., 2005, Matsuda et al., 2011). The cohort was originally diagnosed according to the 2016 WHO-CNS, and thereby already stratified according to IDH status. The glioblastoma patients used for comparison therefore did not include any IDH-mutant specimens, likewise, all DAG-G patients were IDH-wildtype. EGFR amplification was covered by the NGS panel used during the diagnostic workup for all patients in the cohort, and DAG-G patients were stratified according to their molecular alterations, including EGFR amplification, thus, no patients in the DAG-G group with isolated TERT promoter mutation had EGFR amplification. The median age for glioblastoma, IDH-wildtype and DAG-G (TERT) patients was 65 years and 67 years, respectively, with no significant difference found (p = 0.94). MGMT promoter methylation is an epigenetic modification that silences a DNA-repair gene, which in turn compromises the normal DNA repair function. The presence of MGMT methylation confers a better prognosis in glioblastoma patients regardless of treatment, and is a predictor of the tumor’s response to temozolomide (Hegi et al., 2005, Esteller et al., 2000). In our cohort, MGMT promoter methylation was present in 41% and 33% of cases for glioblastoma IDH-wildtype and DAG-G (TERT), respectively, but no significant difference was found (p = 0.73). Therefore, it is unlikely for MGMT promoter methylation to explain the similar overall survival seen in our study. Similarly, the proportion with CDKN2A/B homozygous deletion, the presence of which might confer to a worse prognosis (Ma et al., 2020), was lower among DAG-G patients (33%) compared to glioblastoma (61%), however, no significant difference was found (p = 0.16). A possible confounding factor may have been introduced by surgical sampling not representing the highest grade of the lesion due to intratumoral heterogeneity (Paulus and Peiffer, 1989), which could result in glioblastoma, IDH wildtype erroneously being classified as DAG-G. While not possible to rule out possible confounders due to the low number of patients, the groups used for survival analysis appear comparable concerning most clinical and genetic characteristics. Therefore, and despite the small cohort, the results of the study are suggestive of TERT promoter mutations having a prognostic role in diffuse astrocytic IDH-wildtype gliomas as suggested by WHO-CNS5, however, with the large number of conflicting reports, caution should be taken before assigning cases as molecular glioblastoma.
Methylation array profiling is an up and coming method of classifying CNS tumors, with several larger studies documenting its clinical significance (Priesterbach-Ackley et al., 2020, Wu et al., 2022, Jaunmuktane et al., 2019). Here, the reported number of patients receiving a new diagnosis, out of the total number of patients included, range between 9.8% and 17.9%. Similarly, the reported number of patients returning with a calibrated score above the cutoff of 0.84 or 0.9 range between 54.4% and 66.4%. In this study, the Heidelberg Classifier returned with a calibrated score above the cutoff of 0.84 in 58.6% of patients submitted for 850k methylation profiling. The patients submitted for methylation profiling were diagnostically difficult cases with atypical histological and genetic findings, and as seen in Table 2, these patients included several anaplastic astrocytoma, IDH wildtype. Anaplastic astrocytoma, IDH wildtype, is a broadly and poorly defined diagnostic entity, which in many cases can be allocated to other tumor entities based on molecular profiles, several of which are rare (Grimm and Chamberlain, 2016). This, along with factors such as low tumor percentages in the reviewed specimen, may contribute to the high incidence of no match. However, our findings are still in accordance with reports from other studies (Priesterbach-Ackley et al., 2020, Wu et al., 2022, Jaunmuktane et al., 2019). While the classifier returns a calibrated score below the recommended cutoff of 0.84 in many cases, values below the cutoff may still provide informative results that can help guide clinicians (Capper et al., 2018b). This study found that 6.9% of patients submitted for methylation profiling received a new diagnosis. This is slightly lower than other reports, and may be explained by the small study size and the statistical uncertainties associated with this.
While time and budget constraints along with further demands to laboratory facilities may prohibit DNA methylation from being used in routine diagnostics for adult brain tumors, it remains a valuable tool in cases with ambiguous histological and genetic findings. Our understanding of DNA methylation is, however, still expanding. Recently, a small group of IDH-mutant gliomas with a prognosis comparable to glioblastoma showed reduced hypermethylation (Aoki and Natsume, 2019). In the future, DNA methylation patterns may be incorporated into tumor grading systems.
This study was limited by the small number of patients affected by the WHO-CNS5 reclassification, especially lower grade IDH-mutant astrocytomas with CDKN2A/B homozygous deletion were rare, and the prognostic impact of this criteria could not be assessed. It would have been of interest to examine each of the criteria for IDH-wildtype diffuse astrocytomas (TERT promoter mutation, EGFR amplification and +7/−10) separately. Due to relatively few DAG-G cases, it was not possible to evaluate the prognostic impact of isolated EGFR amplification, nor perform multivariate analysis and thereby identify possible confounders. Further, the NGS panel used for every patient did not include a total coverage of chromosome 10 loss, and the true + 7/− 10 status for the patients could not be evaluated. Instead, an intact PTEN gene was considered a surrogate marker for intact chromosome 10. While not impossible, it would be very unlikely that loss of entire chromosome 10 would occur along with an unaffected PTEN gene assessed by NGS. For DAG-G patients with isolated TERT promoter mutations, calculations were also completed, where the patients, who had gain of entire chromosome 7, were excluded, and similarly, no significant difference in overall survival was found.
Introducing integrated diagnostics for brain tumor has allowed us to more narrowly define tumors and better predict patient outcome. Histopathological examinations and diagnostics have long suffered from intra and inter-observation variation (Jaunmuktane et al., 2019, van den Bent, 2010), and issues with lack of clinical-pathological correlation, as is evident from our study with DAG-G patients, where the histological findings suggest a lower malignant potential, but clinically, the tumors behave as glioblastoma. Introducing genetic data offers a degree of objectivity that has been missing in brain tumor diagnostics. Glioblastoma still retains a poor prognosis, and more research to further our understanding of gliomas and their treatment options is needed to improve patient care.
Ethical statement
1) This material is the authors' own original work, which has not been previously published elsewhere.
2) The paper is not currently being considered for publication elsewhere.
3) The paper reflects the authors' own research and analysis in a truthful and complete manner.
4) The paper properly credits the meaningful contributions of co-authors and co-researchers.
5) The results are appropriately placed in the context of prior and existing research.
6) All sources used are properly disclosed (correct citation). Literally copying of text must be indicated as such by using quotation marks and giving proper reference.
7) All authors have been personally and actively involved in substantial work leading to the paper, and will take public responsibility for its content.
CRediT authorship contribution statement
Danny Mortensen: Conceptualization, Methodology, Investigation, Writing – original draft. Benedicte Parm Ulhøi: Validation, Resources. Slávka Lukacova: Writing – review & editing. Jan Alsner: Formal analysis, Writing – review & editing. Magnus Stougaard: Formal analysis, Validation. Jens Randel Nyengaard: Supervision, Resources, Conceptualization.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
Thanks to the administrative staff and especially Tina Tofthøj at the Department of Pathology, Aarhus University Hospital, for the assistance in identifying the cohort in Patobank and retrieving the results of the histological and genetic examinations for each patient. The authors have no conflicts of interest to declare. This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors. The data that support the findings of this study are available from the corresponding author upon reasonable request.
An abstract on the main results was accepted for oral presentation at the 12th European Congress of Neuropathology in Odense, DK, in May 2021 and for poster presentation at the Danish Pathology Society (DPAS) annual meeting in Kolding, DK, March 2022.
Footnotes
Supplementary data associated with this article can be found in the online version at doi:10.1016/j.ibneur.2022.09.005.
Appendix A. Supplementary material
Supplementary material
.
References
- Abdulkareem I.H., Blair M. Phosphatase and tensin homologue deleted on chromosome 10. Niger. Med. J. J. Niger. Med. Assoc. 2013;54(2):79–86. doi: 10.4103/0300-1652.110033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aoki K., Natsume A. Overview of DNA methylation in adult diffuse gliomas. Brain Tumor Pathol. 2019;36(2):84–91. doi: 10.1007/s10014-019-00339-w. [DOI] [PubMed] [Google Scholar]
- van den Bent M.J. Interobserver variation of the histopathological diagnosis in clinical trials on glioma: a clinician's perspective. Acta Neuropathol. 2010;120(3):297–304. doi: 10.1007/s00401-010-0725-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berzero G., Di Stefano A.L., Ronchi S., Bielle F., Villa C., Guillerm E., et al. IDH-wildtype lower-grade diffuse gliomas: the importance of histological grade and molecular assessment for prognostic stratification. Neuro-Oncology. 2021;23(6):955–966. doi: 10.1093/neuonc/noaa258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brat D.J., Aldape K., Colman H., Holland E.C., Louis D.N., Jenkins R.B., et al. cIMPACT-NOW update 3: recommended diagnostic criteria for "Diffuse astrocytic glioma, IDH-wildtype, with molecular features of glioblastoma, WHO grade IV". Acta Neuropathol. 2018;136(5):805–810. doi: 10.1007/s00401-018-1913-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capper D., Jones D.T.W., Sill M., Hovestadt V., Schrimpf D., Sturm D., et al. DNA methylation-based classification of central nervous system tumours. Nature. 2018;555(7697):469–474. doi: 10.1038/nature26000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capper D., Stichel D., Sahm F., Jones D.T.W., Schrimpf D., Sill M., et al. Practical implementation of DNA methylation and copy-number-based CNS tumor diagnostics: the Heidelberg experience. Acta Neuropathol. 2018;136(2):181–210. doi: 10.1007/s00401-018-1879-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohen A.L., Holmen S.L., Colman H. IDH1 and IDH2 mutations in gliomas. Curr. Neurol. Neurosci. Rep. 2013;13(5):345. doi: 10.1007/s11910-013-0345-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubbink H.J., Atmodimedjo P.N., van Marion R., Krol N.M.G., Riegman P.H.J., Kros J.M., et al. Diagnostic detection of allelic losses and imbalances by next-generation sequencing: 1p/19q co-deletion analysis of gliomas. J. Mol. Diagn. JMD. 2016;18(5):775–786. doi: 10.1016/j.jmoldx.2016.06.002. [DOI] [PubMed] [Google Scholar]
- Esteller M., Garcia-Foncillas J., Andion E., Goodman S.N., Hidalgo O.F., Vanaclocha V., et al. Inactivation of the DNA-repair gene MGMT and the clinical response of gliomas to alkylating agents. N. Engl. J. Med. 2000;343(19):1350–1354. doi: 10.1056/NEJM200011093431901. [DOI] [PubMed] [Google Scholar]
- Giannini C., Giangaspero F. TERT promoter mutation: is it enough to call a WHO grade II astrocytoma IDH wild-type glioblastoma? Neuro-Oncology. 2021;23(6):865–866. doi: 10.1093/neuonc/noab052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodenberger M.L., Jenkins R.B. Genetics of adult glioma. Cancer Genet. 2012;205(12):613–621. doi: 10.1016/j.cancergen.2012.10.009. [DOI] [PubMed] [Google Scholar]
- Grimm S.A., Chamberlain M.C. Anaplastic astrocytoma. CNS Oncol. 2016;5(3):145–157. doi: 10.2217/cns-2016-0002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris P.A., Taylor R., Thielke R., Payne J., Gonzalez N., Conde J.G. Research electronic data capture (REDCap)--a metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inform. 2009;42(2):377–381. doi: 10.1016/j.jbi.2008.08.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris P.A., Taylor R., Minor B.L., Elliott V., Fernandez M., O'Neal L., et al. The REDCap consortium: Building an international community of software platform partners. J. Biomed. Inform. 2019;95 doi: 10.1016/j.jbi.2019.103208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hegi M.E., Diserens A.C., Gorlia T., Hamou M.F., de Tribolet N., Weller M., et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N. Engl. J. Med. 2005;352(10):997–1003. doi: 10.1056/NEJMoa043331. [DOI] [PubMed] [Google Scholar]
- Jaunmuktane Z., Capper D., Jones D.T.W., Schrimpf D., Sill M., Dutt M., et al. Methylation array profiling of adult brain tumours: diagnostic outcomes in a large, single centre. Acta Neuropathol. Commun. 2019;7(1):24. doi: 10.1186/s40478-019-0668-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Louis D.N., Ohgaki H., Wiestler O.D., Cavenee W.K., Burger P.C., Jouvet A., et al. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007;114(2):97–109. doi: 10.1007/s00401-007-0243-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Louis D.N., Perry A., Reifenberger G., von Deimling A., Figarella-Branger D., Cavenee W.K., et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 2016;131(6):803–820. doi: 10.1007/s00401-016-1545-1. [DOI] [PubMed] [Google Scholar]
- Louis D.N., Perry A., Wesseling P., Brat D.J., Cree I.A., Figarella-Branger D., et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro-Oncology. 2021;23(8):1231–1251. doi: 10.1093/neuonc/noab106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma S., Rudra S., Campian J.L., Dahiya S., Dunn G.P., Johanns T., et al. Prognostic impact of CDKN2A/B deletion, TERT mutation, and EGFR amplification on histological and molecular IDH-wildtype glioblastoma. Neuro-Oncology Adv. 2020;2(1) doi: 10.1093/noajnl/vdaa126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuda M., Yamamoto T., Ishikawa E., Nakai K., Zaboronok A., Takano S., et al. Prognostic factors in glioblastoma multiforme patients receiving high-dose particle radiotherapy or conventional radiotherapy. Br. J. Radiol. 2011;84 (Spec No. 1)(Spec Issue 1):S54–S60. doi: 10.1259/bjr/29022270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moran S., Arribas C., Esteller M. Validation of a DNA methylation microarray for 850,000 CpG sites of the human genome enriched in enhancer sequences. Epigenomics. 2016;8(3):389–399. doi: 10.2217/epi.15.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olympios N., Gilard V., Marguet F., Clatot F., Di Fiore F., Fontanilles M. TERT promoter alterations in glioblastoma: a systematic review. Cancers. 2021;13(5) doi: 10.3390/cancers13051147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulus W., Peiffer J. Intratumoral histologic heterogeneity of gliomas. A quantitative study. Cancer. 1989;64(2):442–447. doi: 10.1002/1097-0142(19890715)64:2<442::aid-cncr2820640217>3.0.co;2-s. [DOI] [PubMed] [Google Scholar]
- Priesterbach-Ackley L.P., Boldt H.B., Petersen J.K., Bervoets N., Scheie D., Ulhøi B.P., et al. Brain tumour diagnostics using a DNA methylation-based classifier as a diagnostic support tool. Neuropathol. Appl. Neurobiol. 2020;46(5):478–492. doi: 10.1111/nan.12610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stupp R., Mason W.P., van den Bent M.J., Weller M., Fisher B., Taphoorn M.J., et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 2005;352(10):987–996. doi: 10.1056/NEJMoa043330. [DOI] [PubMed] [Google Scholar]
- Wen P.Y., Weller M., Lee E.Q., Alexander B.M., Barnholtz-Sloan J.S., Barthel F.P., et al. Glioblastoma in adults: a Society for Neuro-Oncology (SNO) and European Society of Neuro-Oncology (EANO) consensus review on current management and future directions. Neuro-Oncology. 2020;22(8) doi: 10.1093/neuonc/noaa106. 1073-113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Z., Abdullaev Z., Pratt D., Chung H.J., Skarshaug S., Zgonc V., et al. Impact of the methylation classifier and ancillary methods on CNS tumor diagnostics. Neuro-Oncology. 2022;24(4):571–581. doi: 10.1093/neuonc/noab227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zacher A., Kaulich K., Stepanow S., Wolter M., Köhrer K., Felsberg J., et al. Molecular diagnostics of gliomas using next generation sequencing of a glioma-tailored gene panel. Brain Pathol. 2017;27(2):146–159. doi: 10.1111/bpa.12367. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary material