Abstract
Mechanisms driving tumor growth and metastasis are complex, and involve the recruitment of many genes working in concert with each other. The tumor is characterized by the expression of specific sets of genes depending on its environment. Here we review the role of the carboxypeptidase E (CPE) gene which has been shown to be important in driving growth, survival and metastasis in many cancer types. CPE was first discovered as a prohormone processing enzyme, enriched in endocrine tumors, and later found to be expressed and secreted from many epithelial-derived tumors and cancer cell lines. Numerous studies have shown that besides wild-type CPE, a N-terminal truncated splice variant form of CPE (CPE-ΔN) has been cloned and found to be highly expressed in malignant tumors and cell lines derived from prostate, breast, liver and lung cancers and gliomas. The mechanisms of action of CPE and the splice variant in promoting tumor growth and metastasis in different cancer types are discussed. Mechanistically, secreted CPE activates the Erk/wnt pathways, while CPE-ΔN interacts with HDACs in a protein complex in the nucleus, to recruit various cell cycle genes and metastatic genes, respectively. Clinical studies suggest that CPE and CPE-ΔN mRNA and protein are potential diagnostic and prognostic biomarkers for multiple cancer types, assayed using solid tumors and secreted serum exosomes. CPE has been shown to be a therapeutic target for multiple cancer types. CPE/CPE-ΔN siRNA transported via exosomes and taken up by recipient high metastatic cancer cells, suppressed growth and proliferation of these cells. Thus future studies, delivering CPE/CPE-ΔN siRNA, perhaps via exosomes, to the tumor could be a novel treatment approach to suppress tumor growth and metastasis.
Keywords: Hepatocellular carcinoma, glioblastoma, lung adenocarcinoma, osteosarcoma, exosomes
1. Introduction
Cancer mortality may result from growth of the primary tumor, but more often from metastatic disease. Numerous mechanisms involving recruitment of genes and proteins that promote the pathogenesis of metastasis or local invasiveness have been described. Nevertheless, despite extensive studies, new molecules continue to emerge that play a role in this complex process. A molecule that has garnished attention is that of carboxypeptidase E (CPE). As the name suggests CPE was first identified as an enzyme that catalyzes the processing of prohormones and proneuropeptides to yield mature peptide hormones and neuropeptides within the secretory vesicles of endocrine and neuronal cells respectively [1]. Besides the soluble form, the membrane anchored form of CPE acts as a prohormone sorting receptor at the trans-Golgi network (TGN) to target prohormones to the regulated secretory pathway in (neuro)endocrine cells [2]. Several SNPs and mutations have been reported in human CPE gene, and shown to be associated with the risk of diabetes, obesity, infertility and Alzheimer’s disease [3, 4]. Two CPE deficient mouse models, CPE KO and Cpefat/fat mice exhibit severe diabetes, obesity, infertility and abnormalities in bone remodeling, as well as dysfunction in mood and emotional responses, sexual behavior, stress-induced neuroprotection, learning and memory [3, 5].
While it was known since 1984 that CPE is expressed in endocrine cells, neurons, and endocrine tumors, the first report of CPE expression in epithelial cancerous cells was in 1989, in human hepatoma (HepG2), but was absent in normal liver cells [6]. Subsequently, tumor cells of non-endocrine origin such as breast [7, 8], and cervical cancer cells [9] were found to express CPE, but not in their respective normal tissues. Microarray studies have shown that CPE mRNA expression is elevated in many types of tumors compared to their normal counterpart (Table 1). For example, while CPE mRNA expression in colon was minimal, during early onset of colorectal cancer (CRC) and lymph node metastasis, CPE mRNA expression was substantially elevated [10, 11]. Additionally, CPE mRNA expression was significantly elevated in neoplastic clear cell sarcoma of the kidney and in renal clear cell tumor compared to the normal control [12, 13] [GDS505/CPE]. Interestingly, CPE expression was down regulated in Wilms’ tumors [GDS1282/CPE]. Ewing sarcomas, which are malignant round-cell tumors in bones have exceptionally high levels of CPE expression, but less than rhabdomyosarcomas, malignant tumors derived from skeletal muscle [GDS971/CPE]. In brain, astrocytes and glial cells show CPE expression, but in the pathological state, as in human gliomas, elevated levels of CPE mRNA expression has been reported [14] [GDS1813/CPE]. Astrocytomas are CNS neoplasms, predominantly of astrocyte origin and they also show elevated CPE mRNA expression [GDS1813/CPE]. Table 1 shows analysis of microarray data, collected up to 2010, and examples of various cancers that exhibited elevated CPE levels compared to their counterpart normal tissue. These early clinical findings led to extensive studies to determine the function and mechanism of action of CPE in promoting growth and metastasis of various types of tumors. In this review, we summarize and discuss the current literature pertaining to the role and mechanism of action of CPE in tumorigenesis, its use as a potential diagnostic and prognostic biomarker and therapeutic target.
Table 1. CPE expression in various cancers.
Analysis of microarray data from the GEO profile database indicated that significantly elevated levels of CPE mRNA was found in many non-endocrine cancers: cervical, colorectal, renal cancers, Ewing sarcomas (bone cancer) and various types of astrocytomas and oligodendrogliomas (samples dated until 2010).
| Serial# | Cancer Type | Sample type and State | Number of samples | CPE expression fold change±SEM& | p value | Geo profile ID |
|---|---|---|---|---|---|---|
| a | Cervical cancer | Biopsies-Metastatic | 33 | 7.4±0.426 | <0.003-22 | GDS2416/CPE |
| b | Colorectal cancer | Biopsies-Metastatic | 24 | 9.5±0.623 | <0.0128 | GDS2609/CPE |
| c | Colorectal cancer | Primary cells Metastatic | 6 | 7±1.020 | <0.0007 | GDS1780/CPE |
| dΔ | Clear cell sarcoma | Biopsies-Metastatic | 14 | 2.49±0.260 | <0.0001 | GDS1282/CPE |
| eΔ | Wilms’ tumor | Biopsies-Metastatic | 15 | −0.5±0.260 | <0.001-4 | GDS1282/CPE |
| f$ | Renal Clear Cell Carcinoma | Biopsies-Metastatic | 8 | 16.3±1.894 | <0.0007 | GDS505/CPE |
| g | Oligodendroglioma | Biopsies-Benign | 8 | 1.75±0.726 | <0.0097 | GDS1813/CPE |
| h | Anaplastic Oligodendroglioma | Biopsies-Metastatic | 6 | 1.59±0.698 | <0.0112 | GDS1813/CPE |
| i | Glioblastoma | Biopsies-Metastatic | 30 | 1.48±0.425 | <0.0029 | GDS1813/CPE |
| j | Astrocytic Tumor | Biopsies-Metastatic | 5 | 1.65±0.325 | <0.0070 | GDS1813/CPE |
| k | Ewing’s sarcoma* | Biopsies-Metastatic | 11 | 8.5±0.650 | <0.031 | GDS971/CPE |
CPE expression fold changes in metastatic or benign tumors versus normal tissues;
CPE expression fold changes compared to normal human fetal kidney;
CPE expression fold changes compared to adjacent normal human kidney;
CPE expression fold changes in Ewing sarcomas vs rhabdomyosarcomas.
2. Human CPE mRNA variants and protein isoforms in tumors
The human CPE gene is located on chromosome 4q32.3, and is highly conserved among species [3, 15]. It contains nine exons and shares high homology with other members of the metallocarboxypeptidase gene family [16]. Recently a 2.4kb CPE mRNA has been identified from human hepatocarcinoma cell line HCC97H and found to have a shorter 5’ untranslated region than the normal human CPE mRNA (Fig. 1A). The coding region is identical to CPE mRNA in normal human cells and encodes a 476-amino acid pre-proprotein (~53kD,WT, wild type, Fig. 1) with different functional domains [17]. The N-terminus 25 amino acids comprise the signal peptide that directs the protein into the rough endoplasmic reticulum (RER) cisternae. The catalytic domain is in the middle and the C-terminus contains a transmembrane domain and a short cytoplasmic tail [18]. Upon entry of pre-pro CPE into the RER cisternae, the signal peptide is removed to generate the proprotein which is trafficked to the Golgi apparatus where it is further processed to the mature form (Fig. 1B). At the trans-Golgi network, (TGN), which is an acidic compartment, the C-terminus of CPE forms an amphipathic α-helix under acidic conditions and binds to the TGN membrane at lipid rafts. This membrane association facilitates its packaging into secretory vesicles. A sub-population of CPE assumes a transmembrane orientation, yielding a cytoplasmic tail which interacts with microtubule motors to transport the secretory vesicles to the cell plasma membrane for release [19]. Studies in mouse and bovine pituitary cells and secretory granules [3] reveal that a subpopulation of CPE is further processed within the secretory vesicle to remove the C-terminal region including the transmembrane domain to yield a soluble form of the enzyme that is secreted. This processing step probably occurs in human cancer cells as well, although the structure of secreted CPE from these cells has not been fully characterized. Nevertheless, CPE molecules that are loosely associated with the secretory vesicle membrane can be released upon exocytosis when exposed to neutral pH in the extracellular space.
Fig. 1. Schematic representation of CPE mRNA and protein:

A. CPE mRNA structure of wild type CPE, showing the position of the three putative start codons and the stop codon. UTR-untranslated regions. In cancer cells (HCC) and clinical tumors, the WT-CPE and CPE variants were found to use an alternate transcription start site at 133 nt. The 1.7kb CPE-ΔN variant has been cloned in HCC cells, and the corresponding deletions (yellow box) in the first exon and 589 nt deletions in the 3’-UTR, in comparison to WT-CPE are shown. The 1.7kb transcript encodes a 40kD CPE-ΔN variant was detected in HCC patient samples and various cancer cell lines.
Numbers refer to nt postions in homo sapiens CPE NM001873.2 sequence. nt- nucleotide. B. Structure of full-length CPE, mature CPE and 40kD CPE-ΔN protein. 40kD CPE-ΔN variant lacks 112 amino acids at the N’ end. Numbers refer to amino acid positions deduced from homo sapiens CPE NM001873.2. SP- signal protein, P- propeptide, TM- transmembrane protein.
More recently, variants of CPE have been found in cancer cells and mouse brain, although generated by different mechanisms in the two species [17, 20]. Protein variants are generated by alternative splicing. First a newly made precursor messenger RNA is spliced to remove introns (non-coding regions) and then the exons (coding regions) are spliced back together to produce a mature mRNA that is translated into a protein. Splicing of the introns occurs via several sequential reactions catalyzed by the spliceosome which is a complex of small nuclear ribonucleoproteins. In some instances, the splicing process can generate a variety of mRNAs containing different compositions of exons from the same primary precursor mRNA, thereby yielding different protein isoforms. This is known as alternative splicing and commonly occurs in most multiexon genes. During this process, exons can be extended or skipped and introns can be retained. Alternative splicing is regulated by trans-acting activator or repressor proteins which bind to cis-acting elements (enhancers or silencers) on the precursor mRNA to promote or reduce the usage of a specific splice site. Alternative splicing events are more common in cancer cells compared to normal cells, and contribute to cancer development and progression. An alternatively spliced 1.7 kb transcript named CPE-ΔN (Genebank # MT035840.1) was first cloned from hepatocellular carcinoma (HCC) cells, and was found in human hippocampus as well [17]. This CPE-ΔN transcript encodes a 40kD CPE protein lacking the N-terminal region due to alternative splicing, occurring at a noncanonical alternative splice site to remove 198 nucleotides of the GC-rich exon 1 (Fig.1). Such noncanonical splicing frequently occurs at a 5′ untranslated GC-rich region and is common in certain cancers.
Analysis of patient HCC tumors showed variability in the amount of the WT-CPE versus the 40kDa CPE-ΔN PCR products present [17]. Furthermore, sequence analysis from one of these HCC patient tumors indicate there may be another CPE splice variant with a deletion region at 288–385 nt in exon1. This CPE transcript variant has not been cloned yet; theoretically it is a 2.1kb transcript if its 3’UTR sequence is the same as the 2.4kb WT-CPE transcript, encoding a ~50kD CPE-ΔN protein. A 2.1kb human brain cDNA sequence (# AK090962.1) with identical splicing but shorter 3’-UTR has been deposited in the NCBI database. Additionally, such a full length variant transcript has been cloned in mouse brain, although its mechanism of generation is different from human and the encoded 47kD protein product has been found. Two PCR products with sizes: 336bp correlating with the 1.7kb CPE-ΔN transcript [17] and the 436bp product correlating with the other possible CPE-ΔN variant have been found in patient non-small cell lung adenocarcinomas [21]. The forms of CPE and CPE-ΔN proteins in other types of patient tumors are less clear, due to lack of molecular weight information in the Western blots reported.
Interestingly, the subcellular distribution of WT-CPE and CPE-ΔN isoforms differ. WT-CPE is packaged into secretory vesicles and appear to be rapidly secreted into the media where it can act extracellularly in an autocrine, paracrine fashion by binding to a putative membrane receptor [17]. Recently, WT-CPE has also been found in exosomes isolated from culture media of liver, prostate, colon and pancreatic cancer cell lines and glioblastoma [22]. The CPE mRNA levels are enriched in exosomes derived from high-metastatic cells compared to the low-metastatic cancer cells. CPE-ΔN, on the other hand, since it does not have a signal peptide, has only been found in the cytoplasm and nucleus of different cancer cell lines and interacts with nuclear components [17, 23, 24]. Studies have shown that CPE and CPE-ΔN regulate the expression of many genes involved in tumorigenesis in different cancers (see sections below).
3. CPE and CPE-ΔN promote tumor growth and metastasis in different cancer types
3.1. Hepatocellular Carcinoma
Hepatocellular carcinoma (HCC) is an aggressive and fatal cancer of the liver, with average survival rate of 6–20 months in the United States. Diagnosis at advanced stages and recurrence of tumor after surgical resection are the determining factors for high mortality in patients [25]. In a prospective study involving 120 HCC patients with early-stage disease who received curative resection, analysis of the ratio of CPE mRNA expression level which included the splice variants in the tumor tissue (T), versus paired non-tumor tissue (N) showed that CPE mRNA expression level is a significant prognostic factor for overall survival and recurrence-free survival. There was also significant correlation between tumor recurrence for stage I and stage II HCC patients and T/N ratio >2 [26]. Interestingly, there was a very small cohort of 8 patients that had a T/N <1 that showed poor recurrence-free survival. Analysis of forms of CPE transcripts in these tumors showed presence of CPE-WT and CPE-ΔN splice variant encoding the 40kD protein. The 436bp PCR product was also detected in some of the HCC tumors, suggesting the possible expression of another CPE-ΔN splice variant discussed above. The levels of each CPE form varied with different patient tumors [17]. This study highlighted the importance of CPE/CPE-ΔN mRNA as a potential biomarker for predicting tumor recurrence in early-stage HCC patients. Additionally, in another clinical study on a cohort of 205 HCC patients [24], higher expression of nuclear CPE-ΔN was associated with advanced disease stages and distant metastasis.
Cell biological studies have shown that in HCC cells, both WT-CPE and 40kD CPE-ΔN proteins are expressed. WT-CPE is secreted into the medium while 40kD CPE-ΔN is transported into the nucleus [17]. During metabolic stress such as nutrient deprivation and hypoxic conditions, recombinant WT-CPE added to the medium has been shown to promote survival of high metatstatic hepatocellular carcinoma cells (MHCC97H cell line) by up-regulating the expression of various genes related to cell survival, such as BCL-2, an anti-apoptotic protein, through activation of ERK1/2 signaling pathway. Other cell survival genes that increased in expression were TNF, NF-κB, IκB alpha, and IL-8 [27]. Additionally, there was an increase in expression of active β-catenin, a Wnt pathway gene, and phosphorylation of GSK3β(Ser9), a key regulator of nuclear factor NF-κB, which promotes proliferation and angiogenesis, and induces epithelial-mesenchymal transition that facilitates distant metastasis [28]. Indeed, β-catenin is also known to play an important role in cancer cell survival [29]. These findings show that WT-CPE acts extracellularly to increase transcription of genes that promote survival and proliferation under metabolic stress conditions in HCC cells.
Studies have shown that transfection of 40kD CPE-ΔN into low metastatic HCCL cells upregulated the expression of multiple metastasis-related genes [17]. Some of the key genes included CXCL12 and its receptor CXCR4, CXCL2 and MMP3, all of which promote aggressive transformation of tumors. In another study, a microarray analysis of differentially expressed genes after the transfection of MHCC97L cells with 40kD CPE-ΔN (Fig. 2 A, B) [20] was carried out. Comparing the genes that were up- and down-regulated in that study with those in MHCC97H versus MHCC97L cells analyzed in the same manner, there were 6 genes in common that were up-regulated (Fig. 2C) and 20 genes in common that were down-regulated (Fig. 2D). The 6 common genes that were up-regulated were associated with increased proliferation, invasion, metastasis and malignancy. Interestingly, many of the common down-regulated genes were also associated with tumor growth and metastasis, but some have been reported to have anti-tumorigenic activity. For example, NDRG1 has been characterized as a metastatic suppressor gene and also found to inhibit tumor progression [30]. DDIT4 has been reported to have a tumor suppressor role in cancer progression [31], while PPP1RC3 has been identified as a candidate tumor suppressor gene [32]. All the gene profiling data taken together indicate that 40kD CPE-ΔN up-regulates many genes that promote tumor growth and metastasis, yet down-regulate others. More importantly, 40kD CPE-ΔN down-regulated several tumor suppressor genes, thereby enhancing its tumorigenic effect. Ultimately, the level of 40kD CPE-ΔN expression in the tumor will significantly influence the gene profile and characteristics of the HCC tumor. With this insight, one might speculate that in the clinical study of HCC patients [26], where there were a few patients with T/N <1 that showed poor recurrence -free survival, perhaps the tumorigenic genes that were down-regulated in the highly malignant tumors (T/N>2), may be up-regulated in those (T/N<1) tumors, in the absence of CPE.
Fig. 2. CPE-ΔN differentially regulates tumor associated genes.
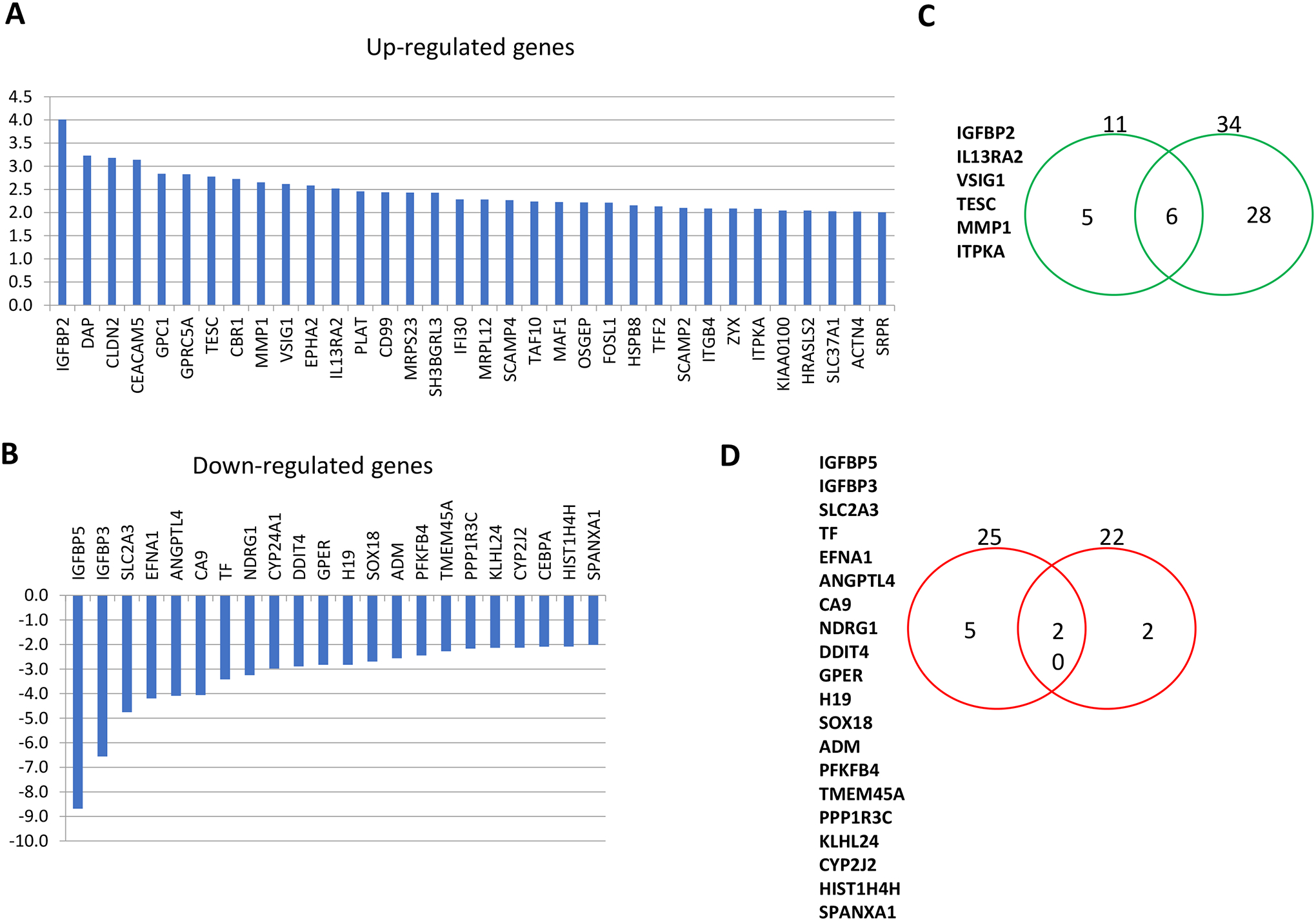
A. Bar graphs show genes that are increased ≥2 fold in MHCC97L-CPE-ΔN vs MHCC97L cells. RNA from MHCC97L cells (low-metastatic HCC cells) transfected with CPE-ΔN plasmid or un-transfected MHCC97L cells were subjected to genome-wide gene expression analysis. All genes that were increased at least two-fold were identified and plotted in a bar graph. Thirty four genes were up-regulated in CPE-ΔN transfected cells versus un-transfected control cells. B. Bar graphs show genes that are decreased ≥2 fold in MHCC97L-CPE-ΔN vs MHCC97L cells. All genes that were decreased at least two-fold were identified and plotted in a bar graph. Twenty two genes were down-regulated in CPE-ΔN transfected cells versus un-transfected control cells. C. RNA from MHCC97L cells transfected with CPE-ΔN plasmid, or MHCC97H (high-metastatic) and un-transfected MHCC97L cells were subjected to genome-wide gene expression level analysis. Venn diagram shows the number of genes up-regulated ≥2 fold in CPE-ΔN transfected cells (right green circle) and MHCC97H (left green circle) versus un-transfected MHCC97L cells. Six genes, listed, were found to overlap in these two conditions. D. RNA from MHCC97L cells transfected with CPE-ΔN plasmid, or MHCC97H (high metastatic) and un-transfected MHCC97L cells were subjected to genome-wide gene expression level analysis. Venn diagram shows the number of genes down-regulated ≥2 fold in CPE-ΔN transfected cells (right red circle) and MHCC97H (left red circle) versus un-transfected MHCC97L cells. Twenty genes, listed, were found to overlap in these two conditions.
HCC cells release exosomes or extracellular vesicles containing CPE mRNA and protein [22]. As exosomes are regarded as critical mediators of intercellular communication favoring tumorigenesis, this recent study investigated if exosomal CPE can contribute to tumor growth and metastasis. CPE is enriched within the exosomes isolated from highly malignant cells, across various cancer types, including HCC. When HCC97L cells were treated with HCC97H exosomes, their proliferation and invasion properties were significantly enhanced in a CPE-dependent manner. Preliminary clinical data showed that CPE transcript copy numbers are elevated in cancer patient sera as opposed to healthy controls, highlighting the utlity of exosome-based CPE to be developed as a cancer biomarker. Exosomes are ideal tools for delivering drugs and siRNA [33]. Modified exosomes from HEK293 cells carrying CPE-shRNA inhibited the proliferation of high-metastatic HCC cells, and down-regulated Cyclin D1 and c-MYC, two key genes associated with tumor growth and progression. Although this is an in vitro study, it lays the foundation for future research in understanding how CPE regulates HCC tumorigenesis and metastasis via exosomes, and in parallel, exploiting exosome-based platforms for targeting CPE in tumor.
3.2. Glioblastoma
In glioblastoma (GBM), the most common and aggressive brain tumor, CPE negatively regulates tumor migration but promotes cell proliferation, thereby supporting the ‘go or grow’ hypothesis. Primary GBM tissues express significantly lower levels of CPE mRNA than normal brain, and reduced CPE protein expression in GBM patients contributes to poor prognosis. Strongest loss of CPE in human GBM tissues was observed in the pseudopalisading perinecrotic areas, which are hypoxic and glucose-deprived, and are associated with enhanced tumor cell migration [34]. Overexpression of WT-CPE in GBM cell lines resulted in enhanced proliferation with a concomitant decrease in invasive potential, with the cells exhibiting altered morphology in the form of larger focal adhesions and actin stress fibers. Among the different forms of CPE, it is the secreted form (sCPE, ~51kD) that regulates the anti-migratory phenotype of glioma cells. Using phospho-proteomic analysis, it was found that the soluble CPE activates the mTOR pathway, signified by the phosphorylation of ribosomal protein S6 (RPS6), a downstream target of mTORC1, and this in turn blocked Rac-1 signaling, which is known to drive GBM cell migration. In addition, sCPE also causes a metabolic switch from aerobic glycolysis to tricarboxylic acid cycle, adversely affecting tumor cell motility [35], as shown in Fig. 3. At the transcriptional level, sCPE was shown to regulate the expression of several genes associated with cell motility and architecture, especially SLUG, acting down stream of ERK1/2 pathway and MMP-2, which is involved in ECM remodeling [36]. In summary, CPE plays a metabolic stress-based anti-migratory role in glioma, through activation of mTOR pathway and transcriptional regulation of cell structure and motility related genes.
Fig. 3. CPE inhibits glioma cell migration.

Soluble CPE possibly binds to a putative receptor (blue) to activate the mTOR pathway, causing phosphorylation of the downstream target, ribosomal protein S6 (RSP6), which inturn inhibits RAC1, and RAC1 induced cell migration. Lactate, produced by the metabolic switch from glycolysis to tricarboxylic acid cycle (TCA) also suppresses cell motility [35]. In addition, it is proposed that CPE regulates expression of SLUG, downstream of ERK1/2 and MMP-2, to modulate glioma cell migration [36]. Modified from Ilina I.E et al, 2017 [35].
3.3. Pancreatic cancer
From the studies investigating the association of CPE with various cancers, it is becoming clear that CPE could regulate tumorigenicity distinctly in different types of cancers. In pancreatic cancer, which is another malignant cancer with high mortality rates, elevated CPE expression was reported both in pancreatic cancer cell lines and tumor tissues of patients. Knockdown of endogenous CPE levels in BXPC-3, a pancreatic cancer cell line, suppressed the proliferation, migration and tumor forming ability of these cells while improving their sensitivity to the chemotherapeutic drug, cisplatin. The study identified NF- κB as the probable downstream target through which CPE exerted its effects on pancreatic cancer [37]. Similarly, down-regulation of CPE expression using several siRNAs that target both CPE and CPE-ΔN in PANC-1 cells, another pancreatic cancer cell line, decreased proliferation, migration and invasion, and cyclin D1 protein expression that is involved in cell cycle regulation [38]. Furthermore, silencing of CPE expression in these cells resulted in differential expression of multiple RNA species such as mRNA, miRNA, circRNA and IncRNA, related to cancer progression [39]. Our recent report suggested that the overexpression of 40kD CPE-ΔN in PANC-1 cells, increases proliferation and invasion, which was partly dependent on the expression of CXCR2, a metastasis driving gene [23]. These results suggest that in the same type of cancer, CPE full-length protein and CPE-ΔN variant could modulate tumor growth and metastasis using many different genes and/or signaling pathways.
3.4. Lung adenocarcinoma
A study has investigated if CPE-ΔN protein expression is valuable in early identification of recurring and metastatic lung adenocarcinoma, which accounts for more than 1 million mortalities worldwide every year. Out of the 95 patient samples that were analyzed for protein expression of CPE-WT and CPE-ΔN using Western Blot and immunohistochemistry (IHC), it was observed that while WT-CPE was expressed at similar levels in tumor tissue and peri-carcinoma tissues, CPE-ΔN was prominently expressed only in the tumor tissue. In this study, interpretation of CPE-ΔN expression is based on nuclear staining by IHC, as reported in the literature, but the form(s) of CPE-ΔN in these tumors was unclear due to lack of molecular weight information in the Western blot reported. Multivariate Cox regression analysis revealed that CPE-ΔN expression was associated with lymph node and distant metastasis. Patients having nuclear staining of CPE-ΔN exhibited higher 3- year postoperative recurrence and metastasis rates than patients lacking CPE-ΔN expression [40]. In another clinical study [21], quantifying CPE/CPE-ΔN mRNA copy numbers by qRT-PCR in lung adenocarcinoma (LADC) in 86 patients revealed that those with high CPE/CPE-ΔN copy numbers had a shorter time of disease-free survival and also shorter time to death, after 7 years follow-up, post-tumor resection, particularly in the early cancer stages. Analysis of the transcripts in 10 randomly selected LADC tumors in this study detected the expression of the 1.7kb transcript and potentially the second CPE-ΔN transcript as well, as discussed above (Fig. 1), but no WT-CPE transcript was detected in these tumors. Both these studies suggest that CPE transcripts or protein could potentially be powerful biomarkers for identifying LADC patients at high risk of recurrence especially at early stages I and II.
A recent study showed that CPE-ΔN enhances the metastatic properties of lung cancer cells both in vitro and in vivo in mouse xenograft experiments, by regulating the expression of E-cadherin, a protein important for epithelial tissue homeostasis. Loss of E-cadherin is a hallmark of epithelial-mesenchymal transition (EMT), a crucial process in malignant transformation. CPE-ΔN was found to interact and stabilize the Snail/HDAC1/HDAC3 complex in the promoter of CDH1, the E-cadherin encoding gene. Over expression of CPE-ΔN in H1299 and H1975 cells, human non-small cell carcinoma cell lines, resulted in alteration of the epigenetic marks which included reduction of histone H3K9 acetylation and an increase of H3K9 and H3K27 trimethylation in the CHD1 gene promoter, leading to the suppression of E-cadherin transcription. These studies indicate that CPE-ΔN plays a significant role in lung cancer metastasis via transcriptional regulation of EMT-related gene CDH1. Relating their findings to a clinical study, 195 patient samples were analyzed. The nuclear levels of CPE-ΔN in these tumor samples showed significant negative correlation with E-cadherin expression on the cell membrane [24]. Furthermore, higher expression of nuclear CPE-ΔN was associated with advanced disease stages and distant metastasis of lung adenocarcinoma. These findings further support the other two studies indicating that CPE-ΔN is a potential biomarker for LADC.
3.5. Colorectal cancer
Colorectal cancer is an aggressive tumor with high global prevalence, and interestingly enhanced CPE expression has been documented in both colorectal cancer (CRC) cell lines and patient tumor samples [10]. Using two CRC cell lines, it was shown that overexpression of full-length CPE enhances the proliferation and colony forming ability of these cells, by increasing the percentage of cells in the S phase of cell cycle, as determined by flow cytometry. In concurrence, CPE down-regulated the expression of CDK inhibitors p21 and p27, and increased cyclin D1 expression [41]. CPE has been shown to regulate canonical Wnt signaling pathway, which is implicated in the progression of several cancers including CRC [42]. CPE, through its N’-terminal domain forms an aggregate with Wnt3a, resulting in an insoluble complex. This complex causes ER stress and inhibition of the Wnt pathway [43]. We could hence speculate that CPE might modulate cancer cell survival and growth in either forward or reverse manner, depending on its expression levels within the cell and cues from the tumor microenvironment.
3.6. Osteosarcoma and Cervical Cancer
The contribution of cyclin D1 in CPE-induced tumorigenesis was also documented in another malignant tumor, osteosarcoma, which affects the bone. Silencing of CPE using CPE-shRNA inhibited proliferation, invasion and migration of MG-63, a human osteosarcoma cell line along with cell cycle arrest at G0/G1 and reduced tumor growth in nude mice [44]. Additionally, shRNA down-regulation of CPE expression in nude mice inhibited xenograft tumor growth [43]. On the other hand, stable overexpression of CPE-ΔN increased the proliferation, migration, invasion, and promoted EMT of Saos-2 osteosarcoma cells, 143B and U2OS. Concurrently, c-MYC and nuclear β-catenin were increased along with protein and activities of MMP2 and MMP9. However, E-cadherin was reduced, but vimentin and transcription factors Snail and Slug were up-regulated. These findings indicate that CPE-ΔN promotes osteosarcoma tumorigenesis via wnt-β-catenin signaling pathway [45].
Cervical cancer is ranked as the fourth common cancer affecting women on a worldwide scale (WHO). Elevated expression of CPE was reported in cervical cancer tissues from patients and cervical cancer cell lines. IHC analysis of cervical cancer samples revealed significant association of CPE levels and pelvic lymph node metastasis (PLNM), an important prognostic factor. Logistic regression analysis showed that high level of CPE expression is an independent risk factor for PLNM while using Cox-regression analyses and it was determined that upregulated CPE expression could serve as an independent determinant for overall survival in early-stage cervical cancer [46].
4. Summary of the mechanisms of action of CPE and CPE-ΔN in tumorigenesis
Studies on CPE and its splice variants in different cancer types indicate that they can regulate expression of many genes via various signaling pathways involved in tumor progression and metastasis. From all the studies reviewed, it is possible to summarize the mechanisms that are common to the various cancers. It has been demonstrated that WT-CPE is secreted from many types of cancer cells, such as liver, pancreatic, ovarian cancer cells and glioblastoma. Its action is presumably receptor-mediated, which then leads to activation of signal transduction pathways and gene transcription (Fig. 4). Recently, it was discovered that human HTR1E, a serotonin G-protein coupled receptor could bind CPE [47], and is expressed in many tumor cell lines (Fig. 5) that also express CPE [17, 23, 34, 47]. Moreover, HTR1E interacts with CPE to activate the ERK1/2 signaling pathway [47]. Hence, HTR1E may be the receptor involved since many genes up-regulated by WT-CPE in tumor progression are mediated through the ERK pathway as reviewed above. Studies from colorectal [41, 42] and pancreatic cancer [37] osteosarcoma [44] and HCC [27, 28] cell lines have demonstrated that other oncogenic pathways such as NF-κB and Wnt are also activated by WT-CPE. These signaling pathways then lead to up-regulation of the expression of cell survival genes (e.g., TNF, IL8), anti-apoptotic genes (e.g., Bcl2) and cell cycle regulatory genes (e.g. Cyclin D1). From the current studies on different cancer types, WT-CPE appears to be primarily mediating cancer cell proliferation and survival especially in hypoxic environments.
Fig. 4. Proposed mechanism for CPE-WT regulation of tumorigenesis.
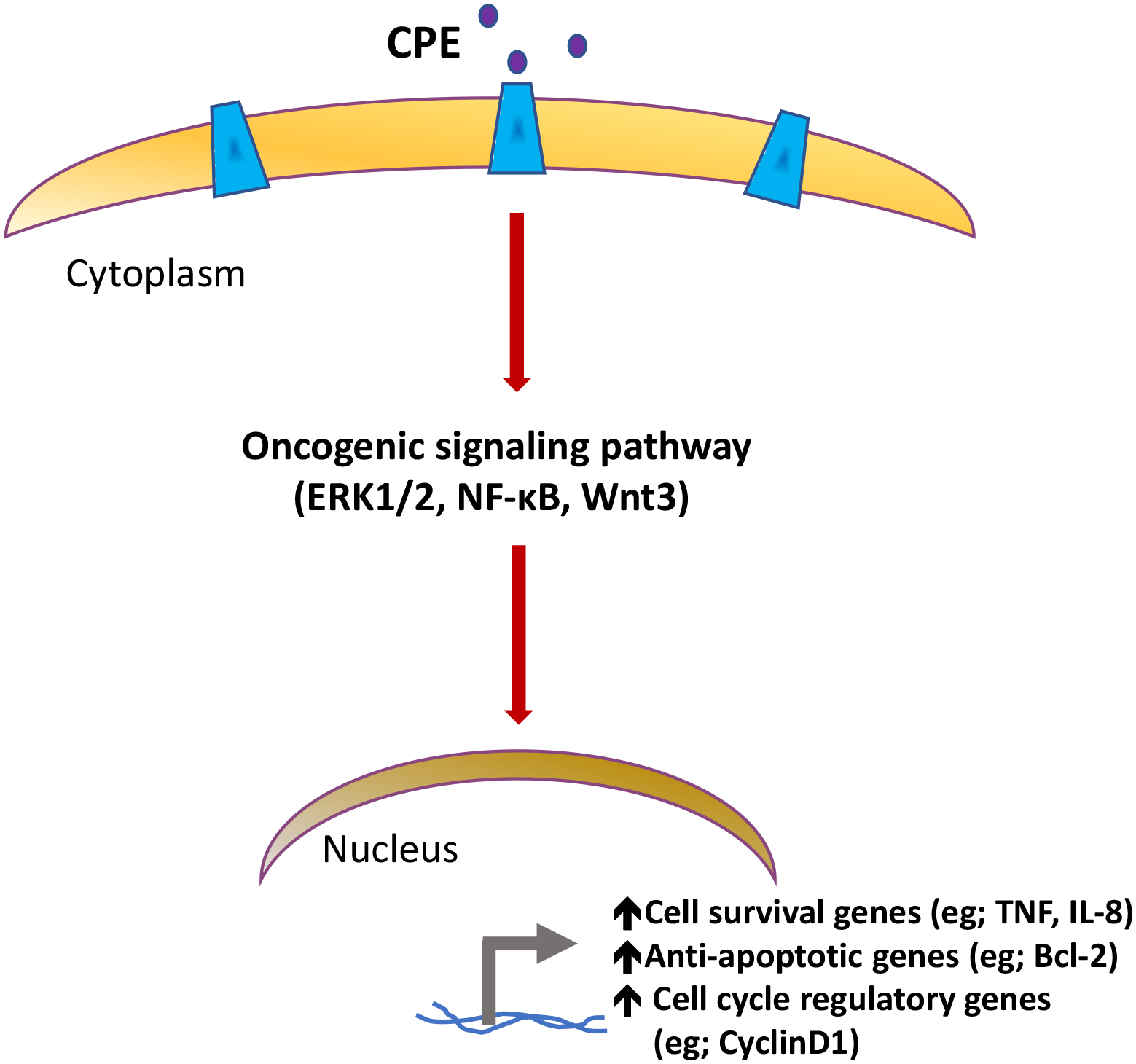
CPE present in the extracellular space binds to a putative receptor (HTR1E), to activate oncogenic signaling pathways such as ERK1/2, NF-κB or Wnt3a, to regulate tumor growth and metastasis, by inducing the expression of target genes involved in cell cycle regulation and survival/ anti-apoptosis.
Fig. 5. 5-HTR1E expression in cancer cells.

Western blot showing the 45kDa 5-HTR1E band in different cancer cells as indicated. Cells were lysed in RIPA lysis buffer and 30 μg lysate was loaded in each well of a 4–12% SDS-PAGE gel. Western blot membrane was incubated with 5-HTR1E, S31 rabbit ab (Cat. no. ab154813, dilution-1:2500) overnight at 4°C followed by one hour incubation with anti-rabbit secondary ab IRDye® 800CW (LI-COR- 925–32211). Blots were visualized on a LI-COR Odyssey system. Previous studies show that Panc-1, BxPC3 and ASPC1 [23], LN 18, LNT229 [34], HCC97H [17], U118 and SHSY5Y (our unpublished data) cells also express CPE.
CPE-ΔN, unlike WT-CPE is not secreted, but is present in the cytoplasm and translocated to the nucleus. Studies in HCC, lung adenocarcinoma and osteosarcoma cell lines indicate that CPE-ΔN up-regulates many important genes that are specifically involved in metastasis and anti-apoptosis (Fig. 2) [17, 20, 23, 24, 45]. Interestingly, CPE-ΔN also down-regulates some genes that promote cancer growth and metastasis, as well as tumor suppressor genes (see HCC section). A yeast two-hybrid screen reported that CPE interacted directly with HDAC1 and HDAC2 [48]. At the time of that publication, the existence CPE-ΔN was not known, but since CPE-ΔN was found to be translocated from the cytoplasm to the nucleus, it would be expected to be the form of CPE that would interact with nuclear HDACs. Our bioinformatic analysis identified a HDAC binding domain in CPE/CPE-ΔN (Fig. 6A). Co-immunoprecipitation experiments using HCC97L cells demonstrated interaction of CPE-ΔN with HDAC1 and HDAC2 [3], while Sun et al [24] showed interaction of CPE-ΔN with Snail/HADC1/HDAC3 complex in lung adenocarcinoma cells. Hence we propose a mechanism of action of CPE-ΔN as shown in the schematic in Fig. 6B. HDACs interact with the binding domain on CPE-ΔN stabilizing a protein complex that binds to the promoter of a specific gene to regulate transcription. Genes down-regulated by CPE-ΔN includes tumor suppressor genes such as E-Cadherin, NDGR1 and DDID4. CPE-ΔN also up-regulates many metastasis-related and anti-apoptotic genes. Besides the genetic and epigenetic regulation of E-cadherin expression in lung adenocarcinoma studied by Sun et al [24], the detail mechanisms for the different genes regulated by CPE-ΔN remain to be elucidated.
Fig. 6. Proposed mechanism of action of CPE-ΔN in tumorigenesis.

A. Data abstracted from Yeast-two-Hybrid analysis indicated that CPE interacts with HDAC1 and HDAC2 [48]. Bioinformatic analysis comparing HDAC interacting domain in other proteins identified a HDAC interacting domain in CPE, and CPE-ΔN (sequence in blue). B. Schematic of proposed mechanism involving CPE-ΔN interaction with HDACs in a protein complex on a promoter to up-regulate metastasis-related and anti-apoptotic gene, or down-regulate tumor suppressor gene expression.
5. CPE in exosomes: Regulator of tumorigenesis and potential biomarker for different cancers
CPE, in addition to being a potential biomarker for various cancers using resected tumor tissues, has recently been shown to be present in exosomes, and may be applicable in liquid biopsy. Exosomes are extracellular vesicles (EV) which range in size from 30–150 nm that are released from cells into the circulation [49]. The contents of EVs which include proteins, RNA and DNA, mirror that of the parent cell, and have been shown to be useful biomarkers for cancer [50–55]. Exosomes are also important in cell-cell communication and have been used as delivery vehicles for therapeutic agents. For example, KRAS si-RNA -loaded exosomes, when injected into orthotopic pancreatic cancer mouse models suppressed tumor growth and inhibited metastasis [33]. Recently it has been reported that WT-CPE protein and mRNA are packaged within exosomes and released from cancer cells, including highly malignant liver, prostate, pancreatic and glioblastoma cancer cells. Furthermore, there was a correlation between elevated CPE mRNA levels in exosomes derived from highly malignant versus low malignant cancer cells. A pilot clinical study documented significantly high CPE copy numbers in exosomes derived from sera of cancer patients versus healthy subjects, indicating that CPE transcripts within exosomes has the potential for being developed as a diagnostic biomarker for many types of cancer [22].
It was found that the phenocopying of metastatic behavior from highly metastatic HCC to low metastatic HCC cells via exosomes is a CPE-dependent process. Furthermore, exosomes loaded with CPE-shRNA when incubated with highly malignant HCC cells were able to effectively suppress the proliferation of these recipient cells. Concomitantly, the levels of Cyclin D1, a cell cycle regulatory protein and c-Myc, a proto-oncogene were down-regulated in the recipient cells [22]. This underscores the value of exosomes modified to carry CPE-shRNA as an anti-cancer therapeutic agent.
6. Conclusions and future prospective
This review has summarized evidence that CPE and CPE-ΔN can regulate many genes and signaling pathways involved in tumor progression and metastasis. More interestingly, distinct pathways such as ERK1/2, mTOR, NF-κB and Wnt3a are downstream effectors of secreted WT-CPE, which act extracellularly in different cancers to regulate expression of pro-survival and cell cycle-regulatory genes. In contrast CPE-ΔN acts in the nucleus as “a master gene regulator” for many down-stream metastasis-related and tumor suppressor genes. Analyses of CPE expression data from clinical samples of different cancer types such as liver cancer, pancreatic cancer, colorectal, cervical cancer and lung adenocarcinoma have consistently shown that elevated CPE levels are associated with poor prognosis. High levels of CPE/CPE-ΔN were identified as a risk factor for distant metastasis and tumor recurrence in patients who underwent curative resection, and was a determinant of their overall survival. Measuring CPE expression levels will be valuable to stratify patients with risk of disease progression and metastasis. However, there are some reports which suggest that low levels of CPE as in glioma [34], and a small sub-class of HCC (CPE mRNA levels T/N<1) [26] led to poor prognosis in these patients. Hence, it is possible that there is a homeostatic level at which CPE must be maintained to regulate normal physiological functions. At low levels, CPE could lose some of its regulatory functions such as inhibiting the Wnt pathway, which can drive tumorigenesis [43], while at abnormal high levels, CPE could induce expression of tumor and metastasis promoting genes [27, 41, 44]. Another possibility is the switch in the expression between the full-length protein versus the variant form(s) of CPE, which is likely to be more detrimental. This emphasizes the need for more research in the future to better understand the molecular mechanism underlying the regulation of CPE/ CPE-ΔN production and maintenance of its physiological levels inside the cell, such as promoter/ enhancer regulation and epigenetic control. It is equally important to verify the role of the HTR1E receptor which WT-CPE interacts with extracellularly in initiating the downstream signaling processes to mediate tumor progression. In conclusion, the studies reviewed here have provided cumulative evidence indicating that CPE/CPE-ΔN is a critical regulator of tumor growth and metastasis, a potential diagnostic/prognostic biomarker and an important therapeutic cancer target for many cancer types.
Highlights.
Carboxypeptidase E (CPE) and its splice variant CPE-ΔN are expressed in many cancers
CPE is secreted and promotes tumor growth by up-regulating anti-apoptotic genes via ERK/wnt signaling
CPE-ΔN is transported into nucleus and up-regulates metastasis-related genes
Clinical studies indicate CPE/CPE-ΔN is a potential diagnostic biomarker and therapeutic target for different cancers
Acknowledgements
This research was supported by the Intramural Research Program of the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD), National Institutes of Health, USA.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Conflicts of Interest: The authors declare no conflict of interest.
Declaration of interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- [1].Fricker LD, Snyder SH, Purification and characterization of enkephalin convertase, an enkephalin-synthesizing carboxypeptidase, J Biol Chem, 258 (1983) 10950–10955. [PubMed] [Google Scholar]
- [2].Cool DR, Normant E, Shen F, Chen HC, Pannell L, Zhang Y, Loh YP, Carboxypeptidase E is a regulated secretory pathway sorting receptor: genetic obliteration leads to endocrine disorders in Cpe(fat) mice, Cell, 88 (1997) 73–83. [DOI] [PubMed] [Google Scholar]
- [3].Cawley NX, Wetsel WC, Murthy SR, Park JJ, Pacak K, Loh YP, New roles of carboxypeptidase E in endocrine and neural function and cancer, Endocr Rev, 33 (2012) 216–253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [4].Cheng Y, Cawley NX, Yanik T, Murthy SR, Liu C, Kasikci F, Abebe D, Loh YP, A human carboxypeptidase E/NF-alpha1 gene mutation in an Alzheimer’s disease patient leads to dementia and depression in mice, Transl Psychiatry, 6 (2016) e973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [5].Chougule A, Kolli V, Baroi S, Ebraheim N, Czernik PJ, Loh YP, Lecka-Czernik B, Nonenzymatic and Trophic Activities of Carboxypeptidase E Regulate Bone Mass and Bioenergetics of Skeletal Stem Cells in Mice, JBMR Plus, 4 (2020) e10392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Grimwood BG, Plummer TH Jr., Tarentino AL, Carboxypeptidase H A regulatory peptide-processing enzyme produced by human hepatoma Hep G2 cells, J Biol Chem, 264 (1989) 15662–15667. [PubMed] [Google Scholar]
- [7].Du J, Keegan BP, North WG, Key peptide processing enzymes are expressed by breast cancer cells, Cancer Lett, 165 (2001) 211–218. [DOI] [PubMed] [Google Scholar]
- [8].Tang SS, Zhang JH, Liu HX, Li HZ, PC2/CPE-mediated pro-protein processing in tumor cells and its differentiated cells or tissues, Mol Cell Endocrinol, 303 (2009) 43–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Kontostathi G, Zoidakis J, Makridakis M, Lygirou V, Mermelekas G, Papadopoulos T, Vougas K, Vlamis-Gardikas A, Drakakis P, Loutradis D, Vlahou A, Anagnou NP, Pappa KI, Cervical Cancer Cell Line Secretome Highlights the Roles of Transforming Growth Factor-Beta-Induced Protein ig-h3, Peroxiredoxin-2, and NRF2 on Cervical Carcinogenesis, Biomed Res Int, 2017 (2017) 4180703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Zhou K, Liang H, Liu Y, Yang C, Liu P, Jiang X, Overexpression of CPE-DeltaN predicts poor prognosis in colorectal cancer patients, Tumour Biol, 34 (2013) 3691–3699. [DOI] [PubMed] [Google Scholar]
- [11].Supiot S, Gouraud W, Campion L, Jezequel P, Buecher B, Charrier J, Heymann MF, Mahe MA, Rio E, Cherel M, Early dynamic transcriptomic changes during preoperative radiotherapy in patients with rectal cancer: a feasibility study, World J Gastroenterol, 19 (2013) 3249–3254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [12].Lenburg ME, Liou LS, Gerry NP, Frampton GM, Cohen HT, Christman MF, Previously unidentified changes in renal cell carcinoma gene expression identified by parametric analysis of microarray data, BMC Cancer, 3 (2003) 31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].Cutcliffe C, Kersey D, Huang CC, Zeng Y, Walterhouse D, Perlman EJ, Renal G Tumor Committee of the Children’s Oncology, Clear cell sarcoma of the kidney: up-regulation of neural markers with activation of the sonic hedgehog and Akt pathways, Clin Cancer Res, 11 (2005) 7986–7994. [DOI] [PubMed] [Google Scholar]
- [14].Bredel M, Bredel C, Juric D, Harsh GR, Vogel H, Recht LD, Sikic BI, High-resolution genome-wide mapping of genetic alterations in human glial brain tumors, Cancer Res, 65 (2005) 4088–4096. [DOI] [PubMed] [Google Scholar]
- [15].Hall C, Manser E, Spurr NK, Lim L, Assignment of the human carboxypeptidase E (CPE) gene to chromosome 4, Genomics, 15 (1993) 461–463. [DOI] [PubMed] [Google Scholar]
- [16].Reznik SE, Fricker LD, Carboxypeptidases from A to z: implications in embryonic development and Wnt binding, Cell Mol Life Sci, 58 (2001) 1790–1804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [17].Yang X, Lou H, Chen YT, Huang SF, Loh YP, A novel 40kDa N-terminal truncated carboxypeptidase E splice variant: cloning, cDNA sequence analysis and role in regulation of metastatic genes in human cancers, Genes Cancer, 10 (2019) 160–170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Dhanvantari S, Arnaoutova I, Snell CR, Steinbach PJ, Hammond K, Caputo GA, London E, Loh YP, Carboxypeptidase E, a prohormone sorting receptor, is anchored to secretory granules via a C-terminal transmembrane insertion, Biochemistry, 41 (2002) 52–60. [DOI] [PubMed] [Google Scholar]
- [19].Park JJ, Loh YP, How peptide hormone vesicles are transported to the secretion site for exocytosis, Mol Endocrinol, 22 (2008) 2583–2595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Xiao L, Yang X, Sharma VK, Loh YP, Cloning, gene regulation, and neuronal proliferation functions of novel N-terminal-truncated carboxypeptidase E/neurotrophic factor-alphal variants in embryonic mouse brain, FASEB J, 33 (2019) 808–820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Kuo IY, Liu D, Lai WW, Wang YC, Loh YP, Carboxypeptidase E mRNA: Overexpression predicts recurrence and death in lung adenocarcinoma cancer patients, Cancer Biomark, 33 (2022) 369–377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Hareendran S, Albraidy B, Yang X, Liu A, Breggia A, Chen CC, Loh YP, Exosomal Carboxypeptidase E (CPE) and CPE-shRNA-Loaded Exosomes Regulate Metastatic Phenotype of Tumor Cells, Int J Mol Sci, 23 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- [23].Hareendran S, Yang X, Lou H, Xiao L, Loh YP, Carboxypeptidase E-N Promotes Proliferation and Invasion of Pancreatic Cancer Cells via Upregulation of CXCR2 Gene Expression, Int J Mol Sci, 20 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Sun J, Meng D, Yu T, Li F, Zhang G, Tian X, Zhao N, Li G, Li L, Wang H, Liu Y, Meng Y, Ma Y, Wan Z, Bao J, Piao H, N-terminal truncated carboxypeptidase E represses E-cadherin expression in lung cancer by stabilizing the Snail-HDAC complex, Am J Cancer Res, 10 (2020) 925–938. [PMC free article] [PubMed] [Google Scholar]
- [25].Gomaa A, Waked I, Management of advanced hepatocellular carcinoma: review of current and potential therapies, Hepatoma Res, 3 (2017) 112–122. [Google Scholar]
- [26].Huang SF, Wu HD, Chen YT, Murthy SR, Chiu YT, Chang Y, Chang IC, Yang X, Loh YP, Carboxypeptidase E is a prediction marker for tumor recurrence in early-stage hepatocellular carcinoma, Tumour Biol, 37 (2016) 9745–9753. [DOI] [PubMed] [Google Scholar]
- [27].Murthy SRK, Dupart E, Al-Sweel N, Chen A, Cawley NX, Loh YP, Carboxypeptidase E promotes cancer cell survival, but inhibits migration and invasion, Cancer Lett, 341 (2013) 204–213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [28].Xia Y, Shen S, Verma IM, NF-kappaB, an active player in human cancers, Cancer Immunol Res, 2 (2014) 823–830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Chung EJ, Hwang SG, Nguyen P, Lee S, Kim JS, Kim JW, Henkart PA, Bottaro DP, Soon L, Bonvini P, Lee SJ, Karp JE, Oh HJ, Rubin JS, Trepel JB, Regulation of leukemic cell adhesion, proliferation, and survival by beta-catenin, Blood, 100 (2002) 982–990. [DOI] [PubMed] [Google Scholar]
- [30].Bae DH, Jansson PJ, Huang ML, Kovacevic Z, Kalinowski D, Lee CS, Sahni S, Richardson DR, The role of NDRG1 in the pathology and potential treatment of human cancers, J Clin Pathol, 66 (2013) 911–917. [DOI] [PubMed] [Google Scholar]
- [31].Ho KH, Chen PH, Chou CM, Shih CM, Lee YT, Cheng CH, Chen KC, A Key Role of DNA Damage-Inducible Transcript 4 (DDIT4) Connects Autophagy and GLUT3-Mediated Stemness To Desensitize Temozolomide Efficacy in Glioblastomas, Neurotherapeutics, 17 (2020) 1212–1227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [32].Bonazzi VF, Irwin D, Hayward NK, Identification of candidate tumor suppressor genes inactivated by promoter methylation in melanoma, Genes Chromosomes Cancer, 48 (2009) 10–21. [DOI] [PubMed] [Google Scholar]
- [33].Kamerkar S, LeBleu VS, Sugimoto H, Yang S, Ruivo CF, Melo SA, Lee JJ, Kalluri R, Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer, Nature, 546 (2017) 498–503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [34].Horing E, Harter PN, Seznec J, Schittenhelm J, Buhring HJ, Bhattacharyya S, von Hattingen E, Zachskorn C, Mittelbronn M, Naumann U, The “go or grow” potential of gliomas is linked to the neuropeptide processing enzyme carboxypeptidase E and mediated by metabolic stress, Acta Neuropathol, 124 (2012) 83–97. [DOI] [PubMed] [Google Scholar]
- [35].Ilina EI, Armento A, Sanchez LG, Reichlmeir M, Braun Y, Penski C, Capper D, Sahm F, Jennewein L, Harter PN, Zukunft S, Fleming I, Schulte D, Le Guerroue F, Behrends C, Ronellenfitsch MW, Naumann U, Mittelbronn M, Effects of soluble CPE on glioma cell migration are associated with mTOR activation and enhanced glucose flux, Oncotarget, 8 (2017) 67567–67591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Armento A, Ilina EI, Kaoma T, Muller A, Vallar L, Niclou SP, Kruger MA, Mittelbronn M, Naumann U, Carboxypeptidase E transmits its anti-migratory function in glioma cells via transcriptional regulation of cell architecture and motility regulating factors, Int J Oncol, 51 (2017) 702–714. [DOI] [PubMed] [Google Scholar]
- [37].Liu A, Shao C, Jin G, Liu R, Hao J, Shao Z, Liu Q, Hu X, Downregulation of CPE regulates cell proliferation and chemosensitivity in pancreatic cancer, Tumour Biol, 35 (2014) 12459–12465. [DOI] [PubMed] [Google Scholar]
- [38].Lou H, Loh YP, Silencing of Carboxypeptidase E expression inhibits proliferation and invasion of Panc-1 pancreatic cancer cells, F1000Res, 10 (2021) 489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [39].Bai Z, Feng M, Du Y, Cong L, Cheng Y, Carboxypeptidase E down-regulation regulates transcriptional and epigenetic profiles in pancreatic cancer cell line: A network analysis, Cancer Biomark, 29 (2020) 79–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Sun J, Meng D, Li L, Tian X, Jia Y, Wang H, Yu H, Sun T, Qu A, Shen H, Bao J, Zhang G, N-terminal truncated carboxypeptidase E expression is associated with poor prognosis of lung adenocarcinoma, Oncology letters, 12 (2016) 4659–4664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Liang XH, Li LL, Wu GG, Xie YC, Zhang GX, Chen W, Yang HF, Liu QL, Li WH, He WG, Huang YN, Zeng XC, Upregulation of CPE promotes cell proliferation and tumorigenicity in colorectal cancer, BMC Cancer, 13 (2013) 412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Clevers H, Wnt/beta-catenin signaling in development and disease, Cell, 127 (2006) 469–480. [DOI] [PubMed] [Google Scholar]
- [43].Skalka N, Caspi M, Lahav-Ariel L, Loh YP, Hirschberg K, Rosin-Arbesfeld R, Carboxypeptidase E (CPE) inhibits the secretion and activity of Wnt3a, Oncogene, 35 (2016) 6416–6428. [DOI] [PubMed] [Google Scholar]
- [44].Fan S, Li X, Li L, Wang L, Du Z, Yang Y, Zhao J, Li Y, Silencing of carboxypeptidase E inhibits cell proliferation, tumorigenicity, and metastasis of osteosarcoma cells, OncoTargets and therapy, 9 (2016) 2795–2803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [45].Fan S, Gao X, Chen P, Li X, Carboxypeptidase E-DeltaN promotes migration, invasiveness, and epithelial-mesenchymal transition of human osteosarcoma cells via the Wnt-beta-catenin pathway, Biochem Cell Biol, 97 (2019) 446–453. [DOI] [PubMed] [Google Scholar]
- [46].Shen HW, Tan JF, Shang JH, Hou MZ, Liu J, He L, Yao SZ, He SY, CPE overexpression is correlated with pelvic lymph node metastasis and poor prognosis in patients with early-stage cervical cancer, Archives of gynecology and obstetrics, 294 (2016) 333–342. [DOI] [PubMed] [Google Scholar]
- [47].Sharma VK, Yang X, Kim SK, Mafi A, Saiz-Sanchez D, Villanueva-Anguita P, Xiao L, Inoue A, Goddard WA 3rd, Loh YP, Novel interaction between neurotrophic factor-alpha1/carboxypeptidase E and serotonin receptor, 5-HTR1E, protects human neurons against oxidative/neuroexcitotoxic stress via beta-arrestin/ERK signaling, Cell Mol Life Sci, 79 (2021) 24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [48].Stelzl U, Worm U, Lalowski M, Haenig C, Brembeck FH, Goehler H, Stroedicke M, Zenkner M, Schoenherr A, Koeppen S, Timm J, Mintzlaff S, Abraham C, Bock N, Kietzmann S, Goedde A, Toksoz E, Droege A, Krobitsch S, Korn B, Birchmeier W, Lehrach H, Wanker EE, A human protein-protein interaction network: a resource for annotating the proteome, Cell, 122 (2005) 957–968. [DOI] [PubMed] [Google Scholar]
- [49].Xiao L, Hareendran S, Loh YP, Function of exosomes in neurological disorders and brain tumors, Extracell Vesicles Circ Nucl Acids, 2 (2021) 55–79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [50].Milane L, Singh A, Mattheolabakis G, Suresh M, Amiji MM, Exosome mediated communication within the tumor microenvironment, J Control Release, 219 (2015) 278–294. [DOI] [PubMed] [Google Scholar]
- [51].Xu R, Rai A, Chen M, Suwakulsiri W, Greening DW, Simpson RJ, Extracellular vesicles in cancer - implications for future improvements in cancer care, Nat Rev Clin Oncol, 15 (2018) 617–638. [DOI] [PubMed] [Google Scholar]
- [52].Han M, Pang B, Zhou C, Li X, Wang Q, Jiang J, Li Y, Liquid biopsy of extracellular vesicle biomarkers for prostate cancer personalized treatment decision, Extracell Vesicles Circ Nucl Acids, 3 (2022) 3–9. [Google Scholar]
- [53].Liu C, He D, Cen H, Chen H, Li L, Nie G, Zhong Z, He Q, Yang X, Guo S, Wang L, Fan Z, Nucleic acid functionalized extracellular vesicles as promising therapeutic systems for nanomedicine, Extracell Vesicles Circ Nucl Acids, 3 (2022) 14–30. [Google Scholar]
- [54].Pedraz-Valdunciel C, Rosell R, Defining the landscape of circRNAs in non-small cell lung cancer and their potential as liquid biopsy biomarkers: a complete review including current methods, Extracell Vesicles Circ Nucl Acids, 2 (2021) 179–201. [Google Scholar]
- [55].Tamura T, Yoshioka Y, Sakamoto S, Ichikawa T, Ochiya T, Extracellular vesicles as a promising biomarker resource in liquid biopsy for cancer, Extracell Vesicles Circ Nucleic Acids, 2 (2021) 148–174. [Google Scholar]


