Abstract
Intrinsic and acquired antibiotic resistance of the nosocomial pathogen Pseudomonas aeruginosa is mediated mainly by the expression of several efflux pumps of broad substrate specificity. Here we report that nfxC type mutants, overexpressing the MexEF-OprN efflux system, produce lower levels of extracellular virulence factors than the susceptible wild type. These include pyocyanin, elastase, and rhamnolipids, three factors controlled by the las and rhl quorum-sensing systems of P. aeruginosa. In agreement with these observations are the decreased transcription of the elastase gene lasB and the rhamnosyltransferase genes rhlAB measured in nfxC type mutants. Expression of the lasR and rhlR regulator genes was not affected in the nfxC type mutant. In contrast, transcription of the C4-homoserine lactone (C4-HSL) autoinducer synthase gene rhlI was reduced by 50% in the nfxC type mutant relative to that in the wild type. This correlates with a similar decrease in C4-HSL levels detected in supernatants of the nfxC type mutant. Transcription of an rhlAB-lacZ fusion could be partially restored by the addition of synthetic C4-HSL and Pseudomonas quinolone signal (PQS). It is proposed that the MexEF-OprN efflux pump affects intracellular PQS levels.
Pseudomonas aeruginosa is an opportunistic pathogen which may cause pneumonia and bacteremia in immunocompromised hosts and is responsible for chronic destructive lung disease in patients suffering from cystic fibrosis. The pathogenicity of P. aeruginosa is attributable to an arsenal of virulence factors, some of which are cell associated (pili, nonpilus adhesins, lipopolysaccharide, and alginate) while others are secreted (proteases, rhamnolipids, exotoxin A, exoenzyme S, and pyocyanin). The production of many of these extracellular virulence factors is controlled by two cell-to-cell signaling systems, called las and rhl, which are both composed of a transcriptional regulator (LasR and RhlR, respectively) and an autoinducer synthase (LasI and RhlI, respectively). LasI and RhlI catalyze the last step in the synthesis of the cell-to-cell signaling molecules 3-oxo-C12-homoserine lactone (3-oxo-C12-HSL) and C4-HSL, respectively; each of these molecules binds to, and activates, its corresponding transcriptional regulator. The systems are connected via a hierarchical cascade (19) and allow coordinated production of extracellular virulence factors, which occurs only when the bacterial cell density has reached a threshold (quorum). Recently, a novel signaling molecule, called PQS, for Pseudomonas quinolone signal (39), has been identified. Furthermore, the published genome sequence of PAO1 (53) has revealed a new modulator of cell-to-cell signaling, termed QscR (4). This protein is homologous to both LasR and RhlR and seems to prevent premature transcription of quorum-sensing regulated genes.
Besides its pathogenic capabilities, P. aeruginosa is well known for its intrinsic resistance to a wide range of antimicrobial agents and its ability to develop multidrug resistance following antibiotic therapy. Recent investigations in several laboratories have demonstrated that both intrinsic and acquired resistance is caused mainly by active efflux systems which efficiently expel antimicrobial compounds without any apparent structural similarity. So far, four genetically distinct efflux systems have been characterized for P. aeruginosa. They are similar in genetic and structural organization but differ in substrate specificity and regulation. The MexAB-OprM system (22, 42) has the broadest substrate spectrum of all bacterial efflux pumps described so far, including quinolones, tetracycline, chloramphenicol (20), trimethoprim (17), β-lactam antibiotics (21), β-lactamase inhibitors (24), and detergents and solvent molecules (23). The transcriptional repressor MexR (43) keeps expression of the mexAB-oprM operon at a low constitutive level, but one sufficient to contribute significantly to the elevated intrinsic antibiotic resistance of this organism. A second efflux system, MexCD-OprJ (41), is responsible for efflux of quinolones, erythromycin (29), and cephalosporins (12, 27). Its expression is totally repressed by the transcriptional regulator NfxB (35, 49). The third efflux pump, MexEF-OprN, transports chloramphenicol as well as quinolones, is overexpressed in nfxC type mutants (18), and is positively regulated by the transcriptional activator MexT (16). Recently, a fourth efflux system of P. aeruginosa, called MexXY, has been cloned into Escherichia coli, on which it conferred resistance to quinolones and erythromycin (31). This efflux system was subsequently shown to be involved in the intrinsic resistance of P. aeruginosa to aminoglycosides and erythromycin (45).
nfxC type mutants were originally isolated from P. aeruginosa strain PAO4009 after exposure to the quinolone norfloxacin (9). These mutants displayed cross-resistance to other quinolones but also to nonquinolone antibiotics such as imipenem and chloramphenicol. The nfxC locus was mapped to 46 min on the PAO1 chromosome (9), near the catA gene, which is located within 15 kb of the mexEF-oprN operon (18). Mutations which lead to overexpression of the MexEF-OprN pump have recently been shown to result from variations in the transcriptional activator gene mexT (26). NfxC is therefore to be considered a phenotype, since overexpression of the MexEF-OprN pump might result from mutations which are not necessarily linked to mexT (26) (T. Köhler and J. L. Dumas, unpublished data).
We previously showed that an nfxC type mutant which overexpressed the MexEF-OprN efflux operon produces about 20 times less pyocyanin than the isogenic wild-type strain (18). Since pyocyanin is a typical secondary metabolite whose production is controlled by the rhl cell-to-cell signaling system (2), we decided to investigate the production of other virulence factors in nfxC type mutants. Our results show that overexpression of the MexEF-OprN efflux pump is correlated with a decrease in production of extracellular virulence factors, particularly those controlled by the rhl system. Evidence is presented that the PQS (39) is involved in this response.
MATERIALS AND METHODS
Bacteria, media, and growth conditions.
Bacterial strains and plasmids are listed in Table 1. E. coli and P. aeruginosa were routinely grown in Luria-Bertani (LB) broth supplemented when necessary with antibiotics at the following concentrations, in milligrams per liter: gentamicin, 15; ampicillin (for E. coli), 100; carbenicillin, 250 (for P. aeruginosa); tetracycline, 10 (for E. coli) or 50 (for P. aeruginosa); mercury chloride, 12.5. For analysis of exoproducts and autoinducers in culture supernatants and for lacZ fusion experiments, P. aeruginosa strains were grown as follows. Strains to be tested were streaked from −80°C glycerol stocks on selective LB agar plates. Single colonies were inoculated into 5 ml of PB (2% Bacto Peptone [select peptone 140; Gibco-BRL], 1.4 g of MgCl2/liter, 10 g of K2SO4/liter) (6) supplemented with antibiotics where appropriate. Cultures were grown overnight at 37°C with agitation in 50-ml flasks. One milliliter of this overnight culture was centrifuged and resuspended in 1 ml of fresh PB. From this suspension, 25 ml of prewarmed PB without antibiotics was inoculated 1:100 and grown in 250-ml flasks with agitation. Defined media were based on M9 salts (25) supplemented with 2 mM MgSO4 and 0.4% glucose. For phage transductions, donor strains were grown in LB broth and recipient strains were resuspended in TNM medium (10 mM Tris-HCl [pH 7.4], 150 mM NaCl, 10 mM MgSO4).
TABLE 1.
Bacterial strains, plasmids, and bacteriophage used in this study
| Strain, plasmid, or bacteriophage | Relevant characteristicsa | Source or reference |
|---|---|---|
| Strains | ||
| E. coli | ||
| MC1061 | F−araD139 Δ(ara-leu)7696 galE15 galK16 Δ(lac)X74 rpsL thi | Laboratory collection |
| S17-1 | thi pro hsdR recA chr::RP4-2 | 51 |
| DH5α | endA1 hsdR17(rk−mk+) glnV44 thi-1 recA1 gyrA relA1 Δ(lacIZYA-argF)U169 deoR [φ80dlacΔ(lacZ)M15] | Laboratory collection |
| MG4λI14 | λ lysogen carrying a chromosomal lasI::lacZ fusion | 48 |
| P. aeruginosa | ||
| PAO-BI | PAO1 wild type | 10 |
| PAO-R1 | PAO-BIΔlasR Tcr | 10 |
| JP2 | PDO100ΔlasI Tcr Hgr | 37 |
| JP3 | PDO111ΔlasR Tcr Hgr | 37 |
| PT5 | PAO1 wild type | Laboratory collection |
| PT121 | PT5mexE::ΩHg (formerly PAOmexE) Hgr | 18 |
| PT149 | PT5nfxC (formerly PAO-7H); overproduces MexEF-OprN | 18 |
| PT466 | PT5ΔlasI Tcr | 15 |
| PT498 | PT5ΔlasR Tcr | 15 |
| PT454 | PT5ΔrhlI::Tn501 Hgr | 15 |
| PT462 | PT5rhlR::Tn501 Hgr | 15 |
| PT531 | PT5rhlR::Tn501 ΔlasRI Tcr Hgr | This study |
| PT469 | PT149ΔlasI; transduced from PAO-JP1 Tcr | This study |
| PT456 | PT149ΔrhlI::Tn501; transduced from PDO100 Hgr | This study |
| PT464 | PT149rhlR::Tn501; transduced from PAO-JP3 Hgr | This study |
| PT500 | PT149rhlR::Tn501; ΔlasR Tcr Hgr | This study |
| PT509 | PT149ΔlasR; transduced from PAO-JP3 Tcr | This study |
| PT637 | PT149mexE::ΩHg; transduced from PT121 Hgr | This study |
| Plasmids | ||
| pTS400 | lasB::lacZ on pSW205; Apr | 36 |
| pECP60 | rhlAB::lacZ on pSW205; Apr | 40 |
| pPCS1001 | lasR::lacZ on pLP170; Apr | 40 |
| pPCS223 | lasI::lacZ on pLP170; Apr | 56 |
| pPCS1002 | rhlR::lacZ on pLP170; Apr | 40 |
| pMAL-I | rhlI::lacZ on pMP220; Tcr | 19 |
| pMP220 | Promoterless lacZ fusion vector; Tcr | 52 |
| pECP61.5 | rhlAB::lacZ, ptac::rhlR; Apr | 37 |
| pLP170 | Promoterless lacZ fusion vector; Apr | 44 |
| pSW205 | Promoterless lacZ fusion vector; Apr | 11 |
| pEZ5 | mexE::lacZ on pSW205; Apr | This study |
| Bacteriophage E79tv2 | Transducing phage | 32 |
Resistance phenotypes: Hg, mercury; Tc, tetracycline; Ap, ampicillin.
Strain and plasmid constructions.
Mutations in the cell-to-cell signaling regulator genes were transferred into the PAO1 wild-type strain PT5 and the nfxC mutant PT149 using the transducing phage E79tv2 (32). The genotype of the strains was verified by Southern hybridization as described previously (15). The mexE gene was inactivated in strain PT149 by using bacteriophage E79tv2 (32) to transduce the mexE::ΩHg mutation from strain PT121 (called PAOmexE in reference 18). In all of the 12 transductants analyzed, wild-type antibiotic susceptibility to the efflux pump substrates was restored. However, 4 of the 12 transductants remained imipenem resistant. One of these strains, called PT637, was shown by sequencing to contain a full-length mexT open reading frame (ORF) (see Results). The mexE-lacZ fusion pEZ5 was constructed by ligating a 1.8-kbp BglII-EcoRV fragment from plasmid pNFZ4 (16) into BamHI-EcoRV-cleaved pSW205 (11).
DNA-manipulations.
Plasmids were introduced into P. aeruginosa by electroporation or by triparental mating using pRK2013 as a helper plasmid (8). Genomic DNA was isolated as described previously (1). PCR amplification was performed by using 100 ng of genomic DNA as a template. PCR mixtures contained primers at 0.1 μmol, 2.5 mM deoxynucleoside triphosphates, and 2 U of Taq polymerase (Appligene, Illkirch, France) in a total volume of 50 μl. Reaction mixtures were subjected to an initial 1-min denaturation step at 95°C, followed by 25 cycles of 30 s at 95°C, 30 s at 55°C, and 2 min at 72°C, with a final 5-min elongation at 72°C. Amplification of rhlI with primers RhlI-P30 (5′-CCATCATCCTGAGCATCTCCAGAGAGC-3′) and RhlI-M6 (5′-GGAATGACTTCGGCATGGCGACTCC-3′) yielded a 1,074-bp fragment, and amplification of rhlR with primers RhlR-P4 (5′-CGGCGTTTCAATGGAATTGTCACAACC-3′) and RhlR-M5 (5′-GGCGGCATCCCTACCCTGATACTCCC-3′) yielded a 1,109-bp fragment. PCR products were run on Tris-acetate-EDTA gels (1.2% agarose) and then purified using a Qiagen gel extraction kit. The mexT DNA region was amplified using primers nfxC-P1 (5′-TCTCGCACGCAAGGCTTGACG-3′) and nfxC-M2 (5′-TCCCACTCGTTCAGCGGTTGTTC-3′). PCR conditions were as follows: 1 min at 95°C, followed by 25 cycles of 30 s at 95°C, 20 s at 52°C, and 2 min at 72°C, with a final 5-min elongation at 72°C. DNA sequences were determined from double-stranded templates according to the dideoxy chain termination method (47) using an automatic sequencer (model 377A; Applied Biosystems).
Qualitative plate assays.
Rhamnolipid production was estimated by inoculating strains on M9-based agar plates supplemented with 0.2% glucose (vol/vol), 2 mM MgSO4, trace elements, 0.05% (vol/vol) glutamate (unless otherwise stated) instead of NH4Cl as an N source, 0.0005% (vol/vol) methylene blue, and 0.02% (vol/vol) cetyltrimethylammonium bromide (50). Plates were incubated first at 37°C for 24 h and then for at least 48 h at room temperature until a blue halo appeared around the colony. Swarm plates were prepared and inoculated as described previously (15). Incubation was carried out for 18 h at 37°C.
Quantitative exoproduct assays on culture supernatants.
Samples of 0.5 ml were taken at various time points during growth in PB, centrifuged (at 8,000 × g for 5 min), and filtered (pore size, 0.22 μm). Filtrates were immediately frozen and kept at −80°C. Elastolytic activity was determined by the elastin Congo red (ECR) method (56). Five milligrams of ECR (Elastin Products Company, Owensville, Mo.) was used per assay. Triplicate samples were analyzed for each time point. Pyocyanin was determined in culture supernatants as described previously (6).
β-Galactosidase assays.
Cultures were grown at 37°C with agitation as described above. Triplicate 100-μl samples were taken to determine the optical density at 600 nm (OD600) and β-galactosidase activity (30). For complementation assays with autoinducers, strains were grown overnight in PB medium. Cells were resuspended in M9-based medium supplemented with 0.2% glucose (vol/vol), 2 mM MgSO4, trace elements, and 0.05% (vol/vol) tryptophan instead of NH4Cl as an N source. C4-HSL was added from a dimethyl sulfoxide stock solution. PQS was synthesized as described previously (39)and dissolved in dimethyl sulfoxide.
Autoinducer bioassays.
Aliquots (3 ml) were taken at different time points during growth in PB medium and centrifuged, and the supernatants were filtered (pore size, 0.22 μm). Aliquots (2 ml) were extracted twice with 2 ml of ethyl acetate (containing 0.01% acetic acid). The extracts were kept at −20°C. Aliquots of the ethyl acetate extract were evaporated, and the dried residue was resuspended directly in 1 ml of the bioassay strain culture. The E. coli bioassay strain was grown in M9 glucose medium supplemented with 0.001% thiamine, 1% LB medium, 50 μg of ampicillin/ml, and 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) when required. The P. aeruginosa bioassay strain JP2 (pECP61.5) was grown in LB medium. Incubation and β-galactosidase determinations were performed as described above.
RESULTS
Production of extracellular virulence factors is affected in nfxC type mutants.
We previously observed that the nfxC type mutant PT149 produced 20-fold less pyocyanin than the wild-type strain PT5 (18). We therefore tested the production of other extracellular virulence factors in these strains and compared their levels to those of isogenic lasR, lasI, rhlR, and rhlI mutants. As expected, the nfxC type mutant and the two rhl mutants showed drastically reduced production of pyocyanin, while the lasR and lasI mutants still secreted substantial amounts of pyocyanin (Table 2). Elastase activity was reduced by more than 50% in the nfxC mutant, while the rhlR and lasR mutants showed only marginal activity in culture supernatants after 10 h of growth in PB medium (Table 2).
TABLE 2.
Exoproduct assays and resistance profiles in P. aeruginosa multidrug efflux and quorum-sensing mutants
| Strain | Genotype | Exoproduct activitya
|
MIC of:
|
|||
|---|---|---|---|---|---|---|
| Pyocyanin | Elastase | Ciprofloxacin | Chloramphenicol | Imipenem | ||
| PT5 | wtb | 100 | 100 | 0.125 | 32 | 1 |
| PT149 | nfxC | 5 | 42 | 1 | 1,024 | 4 |
| PT454 | rhlI | 3 | ND | 0.125 | 32 | 1 |
| PT462 | rhlR | <1 | 4 | 0.125 | 32 | 1 |
| PT466 | lasI | 80 | ND | 0.125 | 64 | 1 |
| PT498 | lasR | 45 | 6 | 0.125 | 32 | 1 |
| PT531 | rhlR lasR | <1 | 4 | 0.125 | 32 | 1 |
| PT637 | nfxC mexE | ND | 88 | 0.125 | 32 | 4 |
| PAO-BI | nfxC | ND | 45 | 1 | 1,024 | 4 |
| PAO-RI | nfxC lasR | ND | <1 | 1 | 1,024 | 4 |
Expressed as a percentage of the activity in the wild-type strain PT5. Pyocyanin activities were determined in supernatants of cultures grown for 18 h in PB medium. ECR assays were performed on supernatants of cultures grown for 10 h in PB medium. At this point the cultures were in stationary phase, and the OD600s of the cultures were 5 ± 0.5. ND, not determined.
wt, wild type.
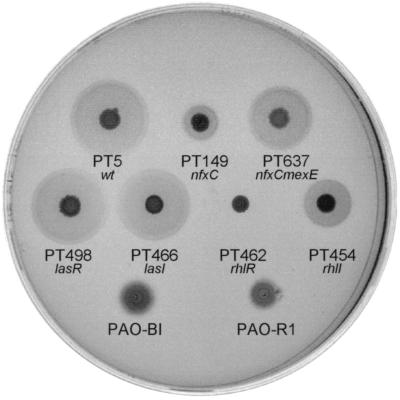
We then compared the production of rhamnolipids using a standard plate assay. As a reference, we included the PAO1 wild-type strain (PAO-BI) (10) and its lasR derivative, PAO-R1 (10). The nfxC mutant PT149 showed strongly reduced rhamnolipid production (Fig. 1). Surprisingly the wild-type strain PAO-BI also showed reduced rhamnolipid production. Unlike PAO-R1, the lasR mutant PT498 constructed in the PT5 background was still able to produce rhamnolipids. As expected, the rhlR mutant PT462 (Fig. 1) was completely deficient in rhamnolipid production as shown previously in other wild-type backgrounds (2, 33). We therefore verified the resistance profiles of PAO-BI and PAO-R1. Indeed, both strains were resistant to chloramphenicol, ciprofloxacin, and imipenem, a phenotype reminiscent of our nfxC mutant PT149 (Table 2). Furthermore β-galactosidase levels expressed from a mexE-lacZ fusion were similar in PAO-BI and the nfxC mutant PT149 (data available upon request). This strongly suggests that strains PAO-BI and PAO-R1 are nfxC mutants (see also below).
FIG. 1.
Rhamnolipid plate assay. Strains to be tested were grown for 8 h in LB medium, and 2 μl was spotted on the plate. Tyrosine at a final concentration of 0.05% was used as the nitrogen source. Incubation was carried out for 24 h at 37°C and then for 72 h at room temperature. The presence of a dark halo around the colony indicates production of rhamnolipids.
As expected, the rhlR lasR double mutant PT531 was completely deficient in production of all exoproducts tested (Table 2).
To rule out the possibility that a particular mutation in the nfxC mutant strain PT149, which had been selected previously on ciprofloxacin (strain PAO-7H in reference 18), was responsible for the decrease in virulence factor production, new nfxC type mutants were selected by plating the wild-type strain PT5 on LB agar plates containing chloramphenicol at 600 μg/ml, a condition which exclusively selects nfxC type mutants (T. Köhler, unpublished data). Fifty spontaneous independent Cmr colonies were analyzed. All of them were cross-resistant to quinolones and imipenem, as expected for nfxC type mutants. All 50 colonies showed drastically decreased production of rhamnolipids in the plate assay, demonstrating the link between the nfxC phenotype and exoproduct synthesis.
nfxC mutants are deficient in swarming.
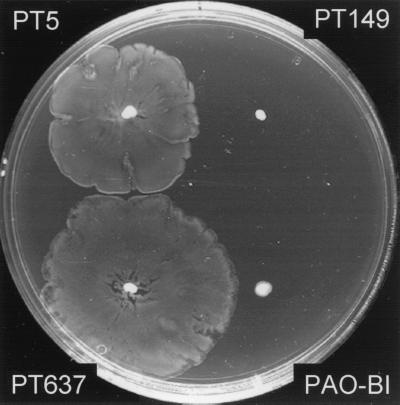
We (15) and others (46) recently demonstrated the swarming motility of P. aeruginosa on semisolid agar plates. Swarming was shown to depend on rhamnolipids as biosurfactants (15). We therefore tested the nfxC mutant PT149 on swarm plates. While the wild type showed normal swarming behavior, both the nfxC mutant and strain PAO-BI were unable to swarm (Fig. 2). This is in agreement with the finding that rhamnolipid production was decreased in these strains.
FIG. 2.
Swarming was tested on M8-based minimal medium (M9 medium without NH4Cl), supplemented with 0.2% glucose and 0.05% glutamate as the sole nitrogen source and solidified with agar to a final concentration of 0.6%. Strains were inoculated by using a toothpick, and plates were incubated at 37°C for 18 h.
Efflux pump overexpression is responsible for decreased virulence factor production.
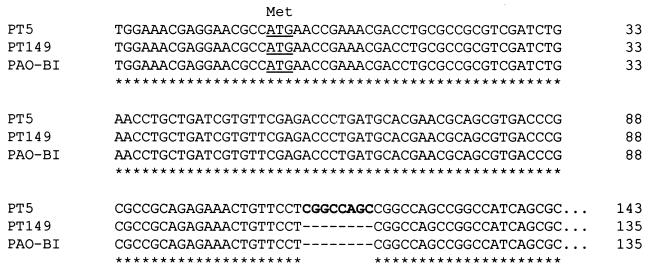
MexEF-OprN is positively regulated by the transcriptional activator MexT (16), located upstream of the efflux operon. In agreement with the findings of Maseda et al. (26), we recently found that the nfxC mutant PT149 contains a full-length mexT ORF, while in the wild-type strain PT5, the mexT ORF is interrupted by an 8-bp insert (CGGCCAGC), resulting in a truncated MexT protein (Fig. 3). This means that wild-type strains which do not express the MexEF-OprN efflux pump may encode an inactive mexT gene, while nfxC type mutants express a functional mexT gene. As expected from the phenotype, we found that strain PAO-BI encodes a functional mexT gene, whose complete sequence is identical to that of the mexT gene in our nfxC mutant PT149 (data available on request).
FIG. 3.
Alignment of a partial DNA sequence of mexT from the wild-type strain PT5, the nfxC mutant PT149, and strain PAO-BI. The 8-bp insert inactivating the mexT ORF is boldfaced. The entire mexT gene was sequenced for all strains, and no other mutation was found.
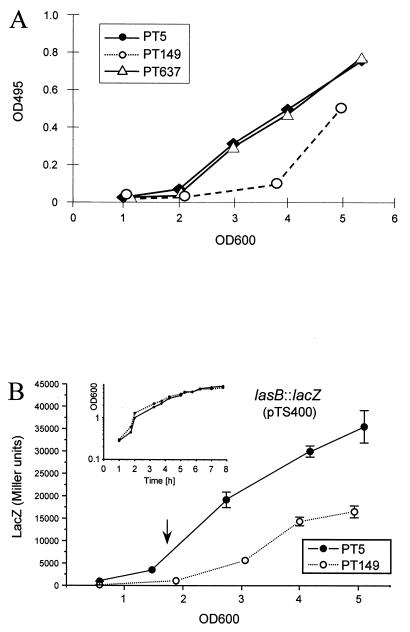
To determine whether possible pleiotropic effects of the functional MexT protein or overexpression of the MexEF-OprN efflux pump per se was responsible for the decrease in virulence factor production, the MexEF-OprN pump was inactivated in the nfxC type mutant PT149 by insertion of an ΩHg cassette into the mexE gene. The resulting strain, called PT637, expressed a full-length mexT ORF and was susceptible to the pump substrates chloramphenicol and ciprofloxacin but remained resistant to imipenem. This is in agreement with previous observations demonstrating that imipenem resistance is independent of MexEF-OprN overexpression (see the last row of Table 1 in reference 18) but results from decreased expression of the porin OprD. Strain PT637 was used in subsequent experiments as a means of distinguishing between phenotypes related to MexEF-OprN pump overexpression and those related to other MexT-mediated effects. Indeed, LasB activities, as measured by elastase production (Fig. 4A), were restored to wild-type levels in strain PT637, and both rhamnolipid production and swarming ability were comparable to those of the wild type (Fig. 1 and 2). We therefore concluded that overexpression of the MexEF-OprN efflux pump is solely responsible for the decrease in virulence factor production in the nfxC type mutant PT149.
FIG. 4.
(A) ECR assay. Elastase production was determined on filtered culture supernatants of strains grown in PB. Determinations were performed on three different occasions. Results from one typical experiment are shown. (B) Expression of the lasB gene was monitored during growth in PB using a lasB::lacZ fusion carried on plasmid pTS400. Growth, expressed as theOD600, was monitored (inset).
Mutations in the cell-to-cell signaling regulators do not affect expression of the MexEF-OprN efflux system in the nfxC type mutant.
Since virulence factor production is affected in nfxC mutants, we asked whether the cell-to-cell signaling regulators were required for expression of the MexEF-OprN efflux system in the nfxC mutant PT149. For this purpose lasI, lasR, rhlI, and rhlR knockout mutations were transduced into PT149. The resistance profiles of the resultant mutants PT469 (nfxC lasI), PT509 (nfxC lasR), PT456 (nfxC rhlI), PT464 (nfxC rhlR), and PT500 (nfxC rhlR lasR) were compared with those of PT5 and PT149 on antibiotic gradient plates. All of the cell-to-cell signaling mutants showed the same susceptibilities to pefloxacin, chloramphenicol, and imipenem as the parental strain PT149 (data not shown). We further introduced plasmid pEZ5, carrying a mexE-lacZ fusion, into the nfxC type mutants PT149, PT464, PT500, and PT509 and measured β-galactosidase activities during exponential growth. In all four strains similar β-galactosidase activities (170 ± 25 Miller units) were obtained. This clearly establishes the quorum-sensing-independent regulation of the mexEF-oprN operon.
Expression of the elastase (lasB) and rhamnosyltransferase (rhlAB) genes is affected in nfxC type mutants.
To further analyze the mechanism of extracellular virulence factor production of the nfxC type mutant PT149, LasB elastase activity was recorded over time and compared to the expression of a plasmid-encoded lasB-lacZ fusion. In both the wild type and the nfxC type mutant, LasB activity and lasB expression started to appear at an OD600 of 2. However, LasB activity and lasB expression continued to increase at a lower rate in the nfxC type mutant, although growth was comparable to that of the wild type (Fig. 4). Hence, the reduced elastase production in strain PT149 results from decreased expression of the lasB gene.
Expression of the rhlAB operon encoding rhamnosyltransferase was assayed using the translational rhlAB-lacZ fusion carried by plasmid pECP60. rhlAB expression was determined after 18 h of incubation in M8 medium supplemented with 0.2% glucose and 0.05% glutamate as the sole nitrogen source. While the wild-type PT5 yielded 1,976 ± 10 Miller units, the nfxC type mutant PT149 and the rhlR mutant PT462 yielded 269 ± 11 and 59 ± 2 Miller units, respectively. These results suggest that the drastically reduced rhamnolipid production in strain PT149 (Fig. 1) is caused by a strong reduction in rhlAB transcription.
Expression of cell-to-cell signaling regulator genes in the nfxC mutant.
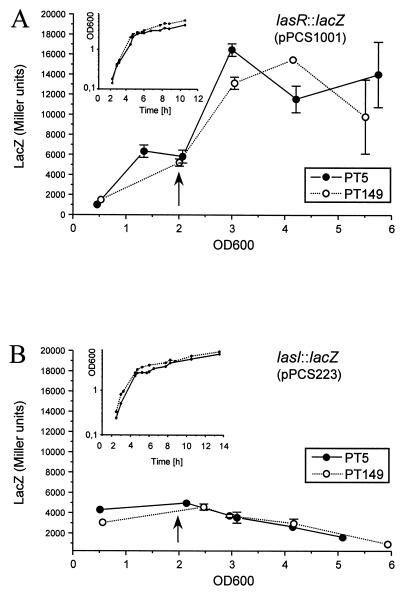
Both lasB and rhlAB are controlled by the las and rhl cell-to-cell signaling systems. We therefore introduced lacZ fusions carried on plasmids to the lasR, lasI, rhlR, and rhlI genes into PT5 and PT149 in order to determine whether their expression was altered, which could account for the decreased elastase and rhamnolipid production in the nfxC mutant. Both lasR(pPCS1001) and lasI(pPCS223) expression reached similar levels in the wild type and the nfxC mutant (Fig. 5). The expression of lasR increased in both strains during early stationary phase, as previously reported with the pPCS1001 lasR::lacZ fusion in strain PAO1 (40). In contrast, the expression of lasI was constant and even decreased slightly in both strains when stationary phase was reached (Fig. 5B). This surprising expression profile is very likely due to the absence on pPCS223 of the rsaL gene, encoding the recently described inhibitor of lasI expression (5). In the absence of multiple copies of the RsaL repressor, expression of lasI is already at a maximum during the exponential-growth phase and therefore does not display an induction profile typical of other genes regulated by the cell-to-cell signaling system. Importantly, the nfxC mutation did not affect the expression of lasI compared to that in the wild type strain.
FIG. 5.
β-Galactosidase activities expressed from lasR::lacZ (A) and lasI::lacZ (B) fusions were determined during growth in PB. Growth, expressed as the OD600, was monitored (inset). Experiments were repeated on three different occasions. Error bars represent standard deviations of triplicate LacZ determinations for one typical experiment. Where error bars are not shown, the standard deviation was within the size of the symbol. Arrows indicate the end of exponential growth. The antibiotic phenotypes of the strains at the end of the experiment were determined on pefloxacin-containing gradient plates. The LacZ activities of control plasmids pSW205 and pLP170 were approximately 3 and 400 Miller units, respectively, and remained fairly constant during growth.
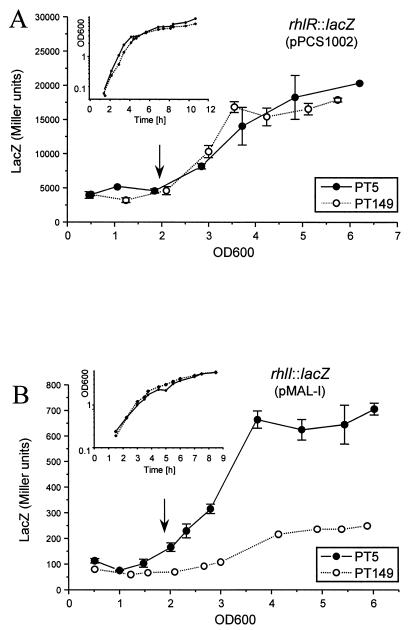
Expression of the regulator gene rhlR(pPCS1002) also increased after the exponential-growth phase in both the wild type and the nfxC mutant and remained comparable even during stationary phase (Fig. 6A). Surprisingly, a significant difference was found when expression of rhlI(pMAL-I) was determined. Indeed, in the nfxC type mutant PT149, rhlI transcription was drastically decreased and reached only 35% of wild-type levels in stationary phase (Fig. 6B). We subsequently sequenced the rhlR-rhlI region in strains PT5 and PT149. However, no differences were found between the two strains, suggesting that the observed effect on rhlI expression in the nfxC type mutant PT149 does not result from mutations in the rhlR-rhlI regulatory region.
FIG. 6.
β-Galactosidase activities expressed from rhlR::lacZ (A) and rhlI::lacZ (B) fusions were determined during growth in PB. Growth, expressed as the OD600, was monitored (inset). Error bars represent standard deviations of triplicate LacZ determinations for one typical experiment. Where error bars are not shown, the standard deviation was within the size of the symbol. Arrows indicate the end of exponential growth.
Autoinducer production in culture supernatants.
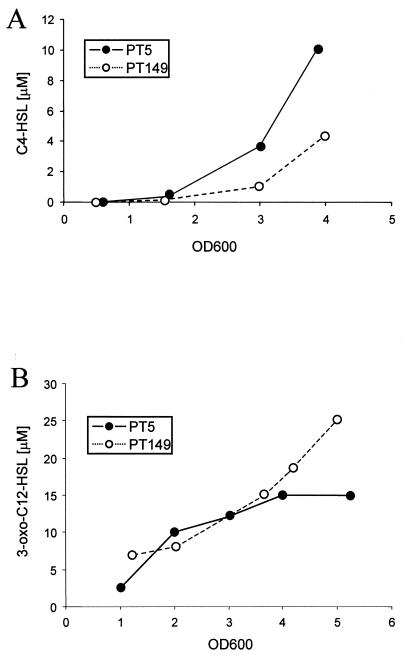
Since the transcription of the autoinducer synthase gene rhlI was affected by the mexEF-oprN expression level, production of the C4-HSL autoinducer was determined in culture supernatants of PT5 and PT149 and compared to that in the rhlI mutant PT454. While the wild-type supernatants reached a C4-HSL concentration of 10 ± 0.6 μM, the PT149 supernatants contained 3.5 ± 0.7 μM C4-HSL (Fig. 7A). As expected, no C4-HSL was detectable in the rhlI mutant PT454 (data not shown). C4-HSL levels determined in supernatants of strain PT637 (nfxC mexE) were comparable to those for the wild type (data available on request), suggesting again that MexEF-OprN overexpression is solely responsible for the decreased amounts of C4-HSL in the nfxC type mutants. When the concentrations of the 3-oxo-C12-HSL autoinducer were determined, we found increased concentrations in supernatants of strain PT149 at ODs above 4 (Fig. 7B). This suggests that MexEF-OprN may contribute to the secretion of the hydrophobic 3-oxo-C12-HSL molecule, as was previously shown for the MexAB-OprM pump (7, 38).
FIG. 7.
Autoinducer concentrations determined in culture supernatants. C4-HSL (A) and 3-oxo-C12-HSL (B) were determined using the bioassay strains JP2(pECP61.5) and MG4λI14, respectively.
Complementation of the nfxC mutant with exogenous autoinducers.
The results described above show an effect of MexEF-OprN overexpression on rhlI transcription and hence on the levels of C4-HSL produced. For full induction, the rhl system requires the 3-oxo-C12-HSL autoinducer and a recently identified novel regulator molecule, PQS (39). We therefore tested the effects of 3-oxo-C12-HSL, C4-HSL, and PQS (kindly synthesized by the group of U. Burger, Faculty of Chemistry, University of Geneva, Geneva, Switzerland) on rhamnolipid production using the plate assay. Addition of 3-oxo-C12-HSL alone or in combination with either PQS or C4-HSL did not affect rhamnolipid production. In contrast, addition of either C4-HSL alone or C4-HSL and PQS increased rhamnolipid production (data not shown). Hence, we tested the effects of the three signaling molecules on expression of the rhlAB-lacZ fusion in strain PT5, in the nfxC mutant PT149, and in the rhl nfxC double mutant PT464. PQS alone at a final concentration of 50 μM had no significant effect on the transcription of the rhlAB fusion in any of the three strains (Table 3). In contrast, the presence of 10 μM C4-HSL alone increased the transcription of rhlAB about sixfold, but only in the nfxC mutant. Surprisingly, when both molecules were present at these concentrations, the transcription increased 15-fold in the nfxC mutant. When 3-oxo-C12-HSL (final concentration, 5 μM) was added to the two other signaling molecules, the expression of rhlAB did not increase further. These results strongly suggest that the levels of both C4-HSL and PQS are affected by mexEF-oprN overexpression. Since rhlAB transcription in the rhlR nfxC mutant was not influenced by the addition of autoinducers, it can be concluded that the partial complementation observed in the nfxC mutant requires the presence of the RhlR regulator.
TABLE 3.
Complementation of the rhlAB::lacZ fusion with C4-HSL and PQS
| Signal added | β-Gal activitiesa (Miller units)
|
||
|---|---|---|---|
| PT5 (pECP60) | PT149 (pECP60) | PT464 (pECP60) | |
| None | 23,298 ± 4,318 | 268 ± 10 | 168 ± 45 |
| C4-HSL (10 μM) | 21,512 ± 274 | 1,716 ± 484 | 133 ± 24 |
| PQS (50 μM) | 16,258 ± 3,453 | 249 ± 4 | 103 ± 11 |
| C4-HSL (10 μM) + PQS (50 μM) | 21,058 ± 4,674 | 4,073 ± 146 | 129 ± 24 |
Cultures were grown for 3 h in the absence of signal; after signal was added, cultures were incubated for 18 h and β-galactosidase (β-Gal) activity was determined. Values are means from two independent experiments done in triplicate.
In conclusion, our work provides evidence that overexpression of the MexEF-OprN multidrug efflux pump reduces the production of virulence factors controlled mainly by the rhl cell-to-cell signaling system (pyocyanin and rhamnolipids) and that this reduction results from decreased rhlI transcription and decreased C4-HSL autoinducer production.
DISCUSSION
P. aeruginosa is known for its ability to develop resistance to a number of structurally unrelated antibiotics, a phenomenon which can now be attributed predominantly to chromosomal mutations leading to overexpression of multidrug efflux systems. P. aeruginosa also produces a series of exoproducts, several of which, such as elastase, alkaline protease, exotoxins, and pyocyanin, have been shown to be virulence factors (3, 54, 55). In this study, we show a link between the active efflux system MexEF-OprN and the production of virulence factors regulated by the las (10, 11, 19) and rhl (2, 33) cell-to-cell signaling systems. This important finding suggests that P. aeruginosa strains becoming resistant to multiple antibiotics by overexpression of MexEF-OprN are likely to be less virulent. Indeed, we recently found that nfxC mutants exhibit significantly reduced virulence both in a nonmammalian system and in a rat model of acute pneumonia (P. Cosson et al., submitted for publication).
The connection between multidrug resistance and virulence factor production was previously suggested in a study comparing 18 multidrug-resistant P. aeruginosa clinical samples collected in a Japanese hospital. All multidrug-resistant strains were deficient in production of pyoverdine, pyocyanin, elastase, hemolysin, and casein protease, while at least 8 out of 13 antibiotic-susceptible strains from the same ward were positive for these virulence factors (34).
Furthermore, two recent reports established a link between the expression of efflux pumps and the quorum-sensing system in P. aeruginosa. Evans et al. (7) showed that strains overexpressing the MexAB-OprM system secrete less 3-oxo-C12-HSL. Furthermore, these investigators found reduced production of pyocyanin, elastase, and casein protease compared to that in the wild type. However, several strains used by Evans et al. were derived from PAO-BI, which we show here to be an nfxC mutant. We sequenced the mexT gene in these strains and confirmed that all express a functional mexT gene and are therefore nfxC mutants. Thus, it is not clear whether the effects on virulence factor production observed by Evans et al. are due to indirect effects of MexAB-OprM overexpression on the expression of MexEF-OprN. In obvious contradiction of these results, Pearson et al. (38) found that in a non-nfxC background, overexpression of the MexAB-OprM pump increased secretion of 3-oxo-C12-HSL, while deletion of mexAB-oprM resulted in decreased release of this autoinducer, suggesting that this hydrophobic molecule is actively secreted by the MexAB-OprM efflux pump.
Our finding that strain PAO-BI is an nfxC type mutant also allows us to explain discrepancies between several laboratories working in the field of quorum sensing. While it was observed that the lasR mutant PAO-R1 was deficient in rhamnolipid production (37), several investigators showed substantial rhamnolipid production in lasR mutants constructed in other strain backgrounds (2, 33). Our findings suggest that the las system has only a marginal effect on rhamnolipid production. Therefore, the strongly reduced rhamnolipid production in strain PAO-R1 is mainly due to its NfxC phenotype.
How can we explain the effect of MexEF-OprN overexpression on the rhl quorum system? One possibility is that autoinducers are substrates of the MexEF-OprN efflux pump, in which case the overexpression of this pump could lead to decreased intracellular autoinducer concentrations and hence diminished production of virulence factors. In the nfxC type mutant, we observed increased amounts of 3-oxo-C12-HSL at OD600 values above 4, suggesting that 3-oxo-C12-HSL could also be a substrate for the MexEF-OprN efflux pump, as suggested for MexAB-OprM (7, 38). On the other hand, supernatants of the nfxC type mutant contained about 60% less of the second autoinducer, C4-HSL, than those of the wild-type strain. Short-chain autoinducers like 3-oxo-C6-HSL of Photobacterium fischeri (14) and C4-HSL of P. aeruginosa apparently (38) diffuse freely across the bacterial cell membrane. It is therefore unlikely that an efflux system, such as the MexEF-OprN pump, is involved in active export of C4-HSL. Our data support the conclusion that the reduced amounts of C4-HSL produced by the nfxC type mutant are the result of altered rhlI expression. Indeed, rhlI transcription levels in PT149 were reduced to 50% of those in the wild type, while rhlR transcription levels were unaffected. Since sequencing of the rhlR-rhlI DNA region obtained from the nfxC type mutant did not reveal any mutation, it seems likely that altered expression or activity of another regulatory element required for rhlI expression might be involved in the nfxC type mutant. The existence of such a regulator of the rhl operon has already been suggested (19). Furthermore, a novel signaling molecule, called PQS (39), has been identified and shown to positively regulate the transcription of lasB and also of rhlI (28). We propose that the MexEF-OprN pump decreases intracellular PQS levels, which could result either from the transport of PQS by the pump or from efflux of a precursor required for PQS biosynthesis, like, for example, tryptophan (13).This would explain the observed decrease in rhlI transcription and the concomitant decrease in C4-HSL levels. The combined decrease in PQS and C4-HSL levels could therefore be responsible for the diminished exoproduct synthesis in the nfxC type mutant. In agreement with this hypothesis is the observation that PQS in combination with C4-HSL is able to partially restore rhlAB transcription in the nfxC type mutant. PQS has a quinolone structure to which a 7-carbon-atom acyl side chain is attached. This confers a hydrophobic character on the molecule which probably prevents diffusion through the membrane, as in the case of 3-oxo-C12-HSL. The results presented here, together with our observation on the reduced virulence of the nfxC mutant PT149 (Cosson et al., submitted), demonstrate that antibiotic resistance can have dramatic effects on the virulence properties of a strain without necessarily affecting its overall fitness.
ACKNOWLEDGMENTS
We are grateful to J. P. Pearson, B. Iglewski, D. Haas, and M. Foglino for providing strains, phages, and plasmids. We thank R. Comte for excellent technical assistance and C. Rossier for performing the sequencing. Many thanks to the members of the group of U. Burger, Faculty of Chemistry, University of Geneva, for the synthesis of C4-HSL, 3-oxo-C12-HSL, and PQS.
This work was supported by grants 31-55961.98 (to T.K.) and 3231-051940.97 and 3200-052189.97 (to C.V.D.) from the Swiss National Science Foundation.
REFERENCES
- 1.Ausubel F M, Brent R, Kingston R E, Moore D D, Smith J A, Seidman J G, Struhl K. Current protocols in molecular biology. New York, N. Y: John Wiley & Sons, Inc.; 1987. [Google Scholar]
- 2.Brint J M, Ohman D E. Synthesis of multiple exoproducts in Pseudomonas aeruginosa is under the control of RhlR-RhlI, another set of regulators in strain PAO1 with homology to the autoinducer-responsive LuxR-LuxI family. J Bacteriol. 1995;177:7155–7163. doi: 10.1128/jb.177.24.7155-7163.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Britigan B E, Roeder T L, Rasmussen G T, Shasby D M, McCormick M L, Cox C D. Interaction of the Pseudomonas aeruginosa secretory products pyocyanin and pyochelin generates hydroxyl radical and causes synergistic damage to endothelial cells. Implications for Pseudomonas-associated tissue injury. J Clin Investig. 1992;90:2187–2196. doi: 10.1172/JCI116104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Chugani S A, Whiteley M, Lee K M, D'Argenio D, Manoil C, Greenberg E P. QscR, a modulator of quorum-sensing signal synthesis and virulence in Pseudomonas aeruginosa. Proc Natl Acad Sci USA. 2001;98:2752–2757. doi: 10.1073/pnas.051624298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.de Kievit T, Seed P C, Nezezon J, Passador L, Iglewski B H. RsaL: a novel repressor of virulence gene expression in Pseudomonas aeruginosa. J Bacteriol. 1999;181:2175–2184. doi: 10.1128/jb.181.7.2175-2184.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Essar D W, Eberly L, Hadero A, Crawford I P. Identification and characterization of genes for a second anthranilate synthetase in Pseudomonas aeruginosa: interchangeability of the two anthranilate synthases and evolutionary implications. J Bacteriol. 1990;172:884–900. doi: 10.1128/jb.172.2.884-900.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Evans K, Passador L, Srikumar R, Tsang E, Nezezon J, Poole K. Influence of the MexAB-OprM multidrug efflux system on quorum sensing in Pseudomonas aeruginosa. J Bacteriol. 1998;180:5443–5447. doi: 10.1128/jb.180.20.5443-5447.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Figurski D H, Helinski D R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci USA. 1979;76:1648–1652. doi: 10.1073/pnas.76.4.1648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Fukuda H, Hosaka M, Hirai K, Iyobe S. New norfloxacin resistance gene in Pseudomonas aeruginosa PAO. Antimicrob Agents Chemother. 1990;34:1757–1761. doi: 10.1128/aac.34.9.1757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gambello M J, Iglewski B H. Cloning and characterization of the Pseudomonas aeruginosa lasR gene, a transcriptional activator of elastase expression. J Bacteriol. 1991;173:3000–3009. doi: 10.1128/jb.173.9.3000-3009.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gambello M J, Kaye S, Iglewski B H. LasR of Pseudomonas aeruginosa is a transcriptional activator of the alkaline protease gene (apr) and an enhancer of exotoxin A expression. Infect Immun. 1993;61:1180–1184. doi: 10.1128/iai.61.4.1180-1184.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gotoh N, Tsujimoto H, Tsuda M, Okamoto K, Nomura A, Wada T, Nakahashi M, Nishino T. Characterization of the MexC-MexD-OprJ multidrug efflux system in ΔmexA-mexB-oprM mutants of Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1998;42:1938–1943. doi: 10.1128/aac.42.8.1938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Holden I, Swift I, Williams I. New signal molecules on the quorum-sensing block. Trends Microbiol. 2000;8:101–104. doi: 10.1016/s0966-842x(00)01718-2. [DOI] [PubMed] [Google Scholar]
- 14.Kaplan H B, Greenberg E P. Diffusion of autoinducer is involved in regulation of the Vibrio fischeri luminescence system. J Bacteriol. 1985;163:1210–1214. doi: 10.1128/jb.163.3.1210-1214.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Köhler T, Curty L K, Barja F, van Delden C, Pechère J C. Swarming of Pseudomonas aeruginosa is dependent on cell-to-cell signaling and requires flagella and pili. J Bacteriol. 2000;182:5990–5996. doi: 10.1128/jb.182.21.5990-5996.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Köhler T, Epp S F, Kocjancic-Curty L, Pechère J C. Characterization of MexT, the transcriptional activator of the MexEF-OprN multidrug efflux system of Pseudomonas aeruginosa. J Bacteriol. 1999;181:6300–6305. doi: 10.1128/jb.181.20.6300-6305.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Köhler T, Kok M, Michea-Hamzehpour M, Plesiat P, Gotoh N, Nishino T, Curty L K, Pechère J C. Multidrug efflux in intrinsic resistance to trimethoprim and sulfamethoxazole in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1996;40:2288–2290. doi: 10.1128/aac.40.10.2288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Köhler T, Michea-Hamzehpour M, Henze U, Gotoh N, Curty L K, Pechère J C. Characterization of MexE-MexF-OprN, a positively regulated multidrug efflux system of Pseudomonas aeruginosa. Mol Microbiol. 1997;23:345–354. doi: 10.1046/j.1365-2958.1997.2281594.x. [DOI] [PubMed] [Google Scholar]
- 19.Latifi A, Foglino M, Tanaka K, Williams P, Lazdunski A. A hierarchical quorum-sensing cascade in Pseudomonas aeruginosa links the transcriptional activators LasR and RhIR (VsmR) to expression of the stationary-phase sigma factor RpoS. Mol Microbiol. 1996;21:1137–1146. doi: 10.1046/j.1365-2958.1996.00063.x. [DOI] [PubMed] [Google Scholar]
- 20.Li X-Z, Livermore D M, Nikaido H. Role of efflux pump(s) in intrinsic resistance of Pseudomonas aeruginosa: resistance to tetracycline, chloramphenicol, and norfloxacin. Antimicrob Agents Chemother. 1994;38:1732–1741. doi: 10.1128/aac.38.8.1732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li X-Z, Ma D, Livermore D M, Nikaido H. Role of efflux pump(s) in intrinsic resistance of Pseudomonas aeruginosa: active efflux as a contributing factor to β-lactam resistance. Antimicrob Agents Chemother. 1994;38:1742–1752. doi: 10.1128/aac.38.8.1742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li X-Z, Nikaido H, Poole K. Role of MexA-MexB-OprM in antibiotic efflux in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1995;39:1948–1953. doi: 10.1128/aac.39.9.1948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Li X Z, Zhang L, Poole K. Role of the multidrug efflux systems of Pseudomonas aeruginosa in organic solvent tolerance. J Bacteriol. 1998;180:2987–2991. doi: 10.1128/jb.180.11.2987-2991.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Li X Z, Zhang L, Srikumar R, Poole K. Beta-lactamase inhibitors are substrates for the multidrug efflux pumps of Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1998;42:399–403. doi: 10.1128/aac.42.2.399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Maniatis T, Fritsch E F, Sambrook J. Molecular cloning: a laboratory manual. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1982. [Google Scholar]
- 26.Maseda H, Saito K, Nakajima A, Nakae T. Variation of the mexT gene, a regulator of the MexEF-oprN efflux pump expression in wild-type strains of Pseudomonas aeruginosa. FEMS Microbiol Lett. 2000;192:107–112. doi: 10.1111/j.1574-6968.2000.tb09367.x. [DOI] [PubMed] [Google Scholar]
- 27.Masuda N, Gotoh N, Ohya S, Nishino T. Quantitative correlation between susceptibility and OprJ production in NfxB mutants of Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1996;40:909–913. doi: 10.1128/aac.40.4.909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.McKnight S L, Iglewski B H, Pesci E C. The Pseudomonas quinolone signal regulates rhl quorum sensing in Pseudomonas aeruginosa. J Bacteriol. 2000;182:2702–2708. doi: 10.1128/jb.182.10.2702-2708.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Michéa-Hamzehpour M, Pechère J-C, Plésiat P, Köhler T. OprK and OprM define two genetically distinct multidrug efflux systems in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1995;39:2392–2396. doi: 10.1128/aac.39.11.2392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Miller J H. Experiments in molecular genetics. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1972. pp. 352–355. [Google Scholar]
- 31.Mine T, Morita Y, Kataoka A, Mizushima T, Tsuchiya T. Expression in Escherichia coli of a new multidrug efflux pump, MexXY, from Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1999;43:415–417. doi: 10.1128/aac.43.2.415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Morgan A F. Transduction of Pseudomonas aeruginosa with a mutant of bacteriophage E79. J Bacteriol. 1979;139:137–140. doi: 10.1128/jb.139.1.137-140.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ochsner U A, Koch A K, Fiechter A, Reiser J. Isolation and characterization of a regulatory gene affecting rhamnolipid biosurfactant synthesis in Pseudomonas aeruginosa. J Bacteriol. 1994;176:2044–2054. doi: 10.1128/jb.176.7.2044-2054.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Okazaki M, Onogawa T, Araki K, Egami T, Furuya N, Endo N, Uchimura H. Isolation frequency and biological characteristics of the multiple-antibiotic resistant Pseudomonas aeruginosa isolated from clinical specimens. Kansenshogaku Zasshi. 1997;71:1181–1186. doi: 10.11150/kansenshogakuzasshi1970.71.1181. [DOI] [PubMed] [Google Scholar]
- 35.Okazaki T, Hirai K. Cloning and nucleotide sequence of the Pseudomonas aeruginosa nfxB gene, conferring resistance to new quinolones. FEMS Microbiol Lett. 1992;97:197–202. doi: 10.1016/0378-1097(92)90386-3. [DOI] [PubMed] [Google Scholar]
- 36.Passador L, Cook J M, Gambello M J, Rust L, Iglewski B H. Expression of Pseudomonas aeruginosa virulence genes requires cell-to-cell communication. Science. 1993;260:1127–1130. doi: 10.1126/science.8493556. [DOI] [PubMed] [Google Scholar]
- 37.Pearson J P, Pesci E C, Iglewski B H. Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of elastase and rhamnolipid biosynthesis genes. J Bacteriol. 1997;179:5756–5767. doi: 10.1128/jb.179.18.5756-5767.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Pearson J P, van Delden C, Iglewski B H. Active efflux and diffusion are involved in transport of Pseudomonas aeruginosa cell-to-cell signals. J Bacteriol. 1999;181:1203–1210. doi: 10.1128/jb.181.4.1203-1210.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Pesci E C, Milbank J B, Pearson J P, McKnight S, Kende A S, Greenberg E P, Iglewski B H. Quinolone signaling in the cell-to-cell communication system of Pseudomonas aeruginosa. Proc Natl Acad Sci USA. 1999;96:11229–11234. doi: 10.1073/pnas.96.20.11229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Pesci E C, Pearson J P, Seed P C, Iglewski B H. Regulation of las and rhl quorum sensing in Pseudomonas aeruginosa. J Bacteriol. 1997;179:3127–3132. doi: 10.1128/jb.179.10.3127-3132.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Poole K, Gotoh N, Tsujimoto H, Zhao Q, Wada A, Yamasaki T, Neshat S, Yamagishi J, Li X Z, Nishino T. Overexpression of the mexC-mexD-oprJ efflux operon in nfxB-type multidrug-resistant strains of Pseudomonas aeruginosa. Mol Microbiol. 1996;21:713–724. doi: 10.1046/j.1365-2958.1996.281397.x. [DOI] [PubMed] [Google Scholar]
- 42.Poole K, Krebes K, McNally C, Neshat S. Multiple antibiotic resistance in Pseudomonas aeruginosa: evidence for involvement of an efflux operon. J Bacteriol. 1993;175:7363–7372. doi: 10.1128/jb.175.22.7363-7372.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Poole K, Tetro K, Zhao Q, Neshat S, Heinrichs D E, Bianco N. Expression of the multidrug resistance operon mexA-mexB-oprM in Pseudomonas aeruginosa: mexR encodes a regulator of operon expression. Antimicrob Agents Chemother. 1996;40:2021–2028. doi: 10.1128/aac.40.9.2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Preston M J, Seed P C, Toder D S, Iglewski B H, Ohman D E, Gustin J K, Goldberg J B, Pier G B. Contribution of proteases and LasR to the virulence of Pseudomonas aeruginosa during corneal infections. Infect Immun. 1997;65:3086–3090. doi: 10.1128/iai.65.8.3086-3090.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ramos-Aires J, Köhler T, Nikaido H, Plésiat P. Involvement of an active efflux system in the natural resistance of Pseudomonas aeruginosa to aminoglycosides. Antimicrob Agents Chemother. 1999;43:2624–2628. doi: 10.1128/aac.43.11.2624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Rashid M H, Kornberg A. Inorganic polyphosphate is needed for swimming, swarming, and twitching motilities of Pseudomonas aeruginosa. Proc Natl Acad Sci USA. 2000;97:4885–4890. doi: 10.1073/pnas.060030097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sanger F, Nicklen S, Coulson A R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA. 1977;74:5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Seed P C, Passador L, Iglewski B H. Activation of the Pseudomonas aeruginosa lasI gene by LasR and the Pseudomonas autoinducer PAI: an autoinduction regulatory hierarchy. J Bacteriol. 1995;177:654–659. doi: 10.1128/jb.177.3.654-659.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Shiba T, Ishiguro K, Takemoto N, Koibuchi H, Sugimoto K. Purification and characterization of the Pseudomonas aeruginosa NfxB protein, the negative regulator of the nfxB gene. J Bacteriol. 1995;177:5872–5877. doi: 10.1128/jb.177.20.5872-5877.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Siegmund I, Wagner F. New method for detecting rhamnolipids excreted by Pseudomonas species during growth in mineral agar. BioTechniques. 1991;5:265–268. [Google Scholar]
- 51.Simon R, Priefer U, Pühler A. A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in Gram-negative bacteria. Bio/Technology. 1983;1:784–790. [Google Scholar]
- 52.Spaink H P, Okker R J H, Wijffelman C A, Pees E, Lugtenberg B J J. Promoters in the nodulation region of the Rhizobium leguminosarum Sym plasmid pRL1JI. Plant Mol Biol. 1987;9:27–39. doi: 10.1007/BF00017984. [DOI] [PubMed] [Google Scholar]
- 53.Stover C K, Pham X Q, Erwin A L, Mizoguchi S D, Warrener P, Hickey M J, Brinkman F S, Hufnagle W O, Kowalik D J, Lagrou M, Garber R L, Goltry L, Tolentino E, Westbrock-Wadman S, Yuan Y, Brody L L, Coulter S N, Folger K R, Kas A, Larbig K, Lim R, Smith K, Spencer D, Wong G K, Wu Z, Paulsen I T. Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature. 2000;406:959–964. doi: 10.1038/35023079. [DOI] [PubMed] [Google Scholar]
- 54.Tamura Y, Suzuki S, Sawada T. Role of elastase as a virulence factor in experimental Pseudomonas aeruginosa infection in mice. Microb Pathog. 1992;12:237–244. doi: 10.1016/0882-4010(92)90058-v. [DOI] [PubMed] [Google Scholar]
- 55.Twining S S, Kirschner S E, Mahnke L A, Frank D W. Effect of Pseudomonas aeruginosa elastase, alkaline protease, and exotoxin A on corneal proteinases and proteins. Investig Ophthalmol Vis Sci. 1993;34:2699–2712. [PubMed] [Google Scholar]
- 56.van Delden C, Pesci E C, Pearson J P, Iglewski B H. Starvation selection restores elastase and rhamnolipid production in a Pseudomonas aeruginosa quorum-sensing mutant. Infect Immun. 1998;66:4499–4502. doi: 10.1128/iai.66.9.4499-4502.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]