Abstract
Amyloid-beta 42 (Aβ42) and phosphorylated tau (pTau) levels in cerebrospinal fluid (CSF) reflect core features of the pathogenesis of Alzheimer’s disease (AD) more directly than clinical diagnosis. Initiated by the European Alzheimer & Dementia Biobank (EADB), the largest collaborative effort on genetics underlying CSF biomarkers was established, including 31 cohorts with a total of 13,116 individuals (discovery n = 8074; replication n = 5042 individuals). Besides the APOE locus, novel associations with two other well-established AD risk loci were observed; CR1 was shown a locus for Aβ42 and BIN1 for pTau. GMNC and C16orf95 were further identified as loci for pTau, of which the latter is novel. Clustering methods exploring the influence of all known AD risk loci on the CSF protein levels, revealed 4 biological categories suggesting multiple Aβ42 and pTau related biological pathways involved in the etiology of AD. In functional follow-up analyses, GMNC and C16orf95 both associated with lateral ventricular volume, implying an overlap in genetic etiology for tau levels and brain ventricular volume.
Supplementary Information
The online version contains supplementary material available at 10.1007/s00401-022-02454-z.
Keywords: GWAS, Alzheimer’s disease, Cerebrospinal fluid, Amyloid-beta, Tau
Introduction
Resolving the genetic background of Alzheimer’s disease (AD) has proven to contribute greatly to our understanding of underlying disease processes, for instance with the discovery of APP [25], PSEN1 [53], and PSEN2 [52] in family-based studies, leading to the amyloid cascade theory [38]. In addition, genome-wide association studies (GWAS) in AD have convincingly highlighted the importance of microglia [33, 56], a finding previously supported by research from other scientific fields [18, 72, 75], and now also widely accepted as a genetic cause rather than a result of AD pathogenesis. Further exploration of genetic risk factors contributing to AD development and pathogenesis might reveal more biological insights, an important step in the quest for AD treatment that will slow down or even halt disease progression.
GWAS of clinically diagnosed AD patients have been successful, and current efforts largely focus on increasing sample size to improve the statistical power to detect genetic variants [6, 70]. An alternative approach is to study effects of genetic variants on pathophysiological features of AD. The strength of such studies is based on the assumption that more objective measurable biological properties are more strongly associated with the underlying AD pathology than the clinical diagnostic classifications (e.g., misclassifications or symptoms not manifested yet), thereby allowing to detect larger effects by reducing heterogeneity [26]. The use of biomarkers further enables to identify genetic effects specific for certain AD-related biological mechanisms. This is an advantage over the conventional GWAS approach for clinical AD diagnosis, where it generally remains unclear through what causal gene or cellular process a locus is associated to AD.
It is possible to measure levels of amyloid-beta-42 (Aβ42) and (phosphorylated) tau (pTau and Tau) in cerebrospinal fluid (CSF), the two major proteins implicated in the AD pathological process. Aβ42 pathology in the brain is negatively correlated with CSF Aβ42 levels, where a decrease in CSF Aβ42 is indicative of AD [49, 58]. CSF (p)Tau is positively correlated with (p)Tau pathology in the brain, and therefore higher CSF (p)Tau levels are observed in patients with AD. CSF pTau is presumed to reflect AD-type tau-tangles more specifically than total tau [49, 58]. Previous studies on CSF amyloid-beta and (p)Tau have identified genetic risk loci, the most recent one including 3,146 individuals [17]. Some of the 8 discovered loci had not been previously associated with AD, emphasizing the potential of endophenotypes to reveal novel genetic risk factors. Our current study aimed to further define the genetic background of AD by studying the genetic effects on CSF Aβ42 and pTau levels in a total of 13,116 individuals.
Materials and methods
Participants
We combined data from 16 European cohorts, encompassing a total of 8074 individuals (Table 1; Online Resource 1—Table 1; Online Resource 2—Fig. 1) with both genotype data and CSF measurements. The majority of these cohorts (82%) are part of the EADB consortium [6], and included the full spectrum of clinical severity potentially leading to AD, from subjective cognitive decline, mild cognitive impairment, to dementia. Written informed consent was obtained from study participants or, for those with substantial cognitive impairment, from a caregiver, legal guardian, or other proxy. Study protocols for all cohorts were reviewed and approved by the appropriate institutional review boards.
Table 1.
Demographic information on cohorts of stage 1 discovery analysis
| Country | Cohort | Age | Gender | Diagnoses | APOE4 carrier | Aβ42 | pTau | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mean years (sd) | % male | AD (%) | MCI (%) | Control (%) | Other dem (%) | % APOE4 + | Mean levels (sd) | n | Mean levels (sd) | n | ||
| Belgium | DEM | 78.3 (7.3) | 40 | 72 | 27 | 1 | 0 | 48 | 562.7 (250.8) | 587 | 71.6 (35.4) | 585 |
| Finland | ADGEN | 70.2 (8.0) | 34 | 89 | 1 | 10 | 0 | 73 | 477.1 (183.2) | 226 | 80.4 (34.8) | 155 |
| France1 | BALTAZAR | 77.0 (6.7) | 45 | 43 | 57 | 0 | 0 | 43 | 825.7 (358.9) | 420 | 73.0 (34.0) | 419 |
| France2 | MEMENTO | 69.2 (8.9) | 47 | 0 | 100 | 0 | 0 | 30 | 1078.6 (406.7) | 389 | 62.6 (29.8) | 386 |
| France3 | CNRMAJ-Lille | 66.0 (8.7) | 47 | 100 | 0 | 0 | 0 | 56 | 664.8 (205.1) | 127 | 101.8 (53.4) | 125 |
| Germany1 | Delcode | 71.7 (5.9) | 52 | 13 | 23 | 64 | 0 | 39 | 965.3 (338.2) | 465 | 63.2 (35.1) | 462 |
| Germany2 | KND | 67.3 (8.7) | 57 | 18 | 82 | 0 | 0 | 49 | 724.0 (347.9) | 309 | 69.4 (36.8) | 305 |
| Germany3 | TUM | 70.2 (9.2) | 48 | 98 | 1 | 0 | 1 | 58 | 536.8 (240.0) | 151 | 88.8 (48.7) | 152 |
| Germany4 | PAGES | 73.4 (7.7) | 40 | 70 | 30 | 0 | 0 | 59 | 528.2 (245.5) | 136 | 90.8 (60.0) | 137 |
| Germany5 | UHB | 70.3 (7.2) | 42 | 69 | 30 | 1 | 0 | 68 | 545.7 (327.8) | 111 | 82.0 (44.2) | 111 |
| Netherlands | ADC & Pearl ND | 64.1 (8.9) | 59 | 42 | 10 | 24 | 23 | 50 | 842.8 (288.9) | 2936 | 65.9 (35.9) | 2931 |
| Spain1 | ACE | 72.7 (8.2) | 43 | 27 | 59 | 8 | 6 | 36 | 779.2 (314.3) | 609 | 68.2 (35.5) | 609 |
| Spain2 | SIGNAL & SPIN | 70.6 (8.0) | 43 | 34 | 45 | 19 | 2 | 39 | 707.2 (368.3) | 394 | 71.1 (41.9) | 370 |
| Spain3 | Valdecilla | 67.0 (9.0) | 39 | 10 | 37 | 45 | 8 | 29 | 887.5 (360.5) | 98 | 56.8 (27.9) | 99 |
| Sweden1 | Birth cohort & Clin. AD | 75.0 (9.4) | 45 | 51 | 0 | 49 | 0 | 52 | 533.5 (271.9) | 856 | 68.9 (35.5) | 694 |
| Sweden2 | Uppsala university | 71.0 (6.3) | 46 | 58 | 37 | 0 | 6 | 64 | 489.0 (238.1) | 260 | 77.6 (37.2) | 259 |
For most cohorts, one of the two CSF levels is missing for a small number of samples. The demographics for age, gender diagnoses and APOE4 carriers status are then displayed for the largest group of samples with at least one CSF measurement. AB42 levels are corrected according to known drift over time for the Dutch ADC and Pearl ND cohorts. All, except the Swedisch Birth cohort and clinical AD samples, are part of EADB
Aβ42 amyloid-beta 42, pTau phosphorylated tau, AD Alzheimer’s disease, MCI mild cognitive impairment, other dem other dementia, n sample size, DEM Antwerp prospective dementia cohort
For replication, 15 cohorts totaling 5042 individuals (Online Resource 1—Table 2) were available to attempt replication of the association signals to Aβ42 and pTau, for the variants with P value < 1e–5 in the discovery analysis. Data from all cohorts, except one (NorCog from University of Oslo, Norway), were obtained through collaboration with the previous largest GWAS on CSF Aβ42 and pTau, mostly including cohorts originating from the United States [17]. Basic demographics are described in Online Resource 1—Table 2, more detailed cohort information is described elsewhere [17].
CSF measurements
Due to the multi-center approach, CSF protein levels were measured with various CSF protein assays (Online Resource 1—Table 2). Aβ42 was measured with ELISA, Lumipulse or V-PLEX, and pTau with ELISA or Lumipulse. For details on specific lab procedures, see the original studies [2–4, 17, 21, 23, 27, 34, 36, 42, 47, 50, 57, 62, 78]. Protein levels were log10 transformed and normalized within cohorts and CSF assay type (if multiple assays were used within a single cohort) to approximate a normal distribution to correct for the application of various CSF assays across different studies. Then, the normalized protein levels were used as continuous phenotypes in the association analyses. The distribution of raw and normalized CSF protein levels are displayed in Online Resource 2—Figs. 2–13, for the cohorts for which we have individual-level data available (indicated in Online Resource 1—Table 1; column D). For the stratified analyses, two subgroups of individuals, amyloid normal and amyloid abnormal, were defined based on their Aβ42 status. Individuals with an untransformed Aβ42 level below a threshold were assigned to the abnormal amyloid level group. The thresholds were defined by the individual research groups as it depends on technical circumstances, and are displayed in Online Resource 1—Table 2.
Genotyping, quality control and imputation
The genetic data for the EADB cohorts have been processed in a homogeneous approach (Online Resource 1—Table 1), in which the Illumina Infinium Global Screening Array (GSA, GSAsharedCUSTOM_24 + v1.0) was predominantly used for data generation. Additional arrays included the Axiom 815 K Spanish biobank array (Thermo Fisher) for ACE (Barcelona, Spain) and Valdecilla (Santander, Spain) cohorts, and the Illumina Neurochip array (Gothenburg, Sweden). Standard quality control (QC) procedures were performed to exclude individuals and variants with low quality, in general followed by imputation with the Trans-Omics for Precision Medicine (TOPMed) reference panel [13, 59]. For the EADB cohorts for which GSA genotype level data were available, the details on QC steps and imputation with the TOPMed reference panel were previously described [6].
For the Spanish ACE and Valdecilla cohorts, QC procedures are described in another study [16], followed by imputation with the TOPMed reference panel. For the Gothenburg H70 Birth Cohort studies and clinical AD samples from Sweden, the QC and imputation procedures were described elsewhere [48]. Post‐imputation QC only included variants with a high imputation quality (RSQ [imputation quality] > 0.8). The UCSC LiftOver program (https://genome-store.ucsc.edu/) and Plink v2.0 (www.cog-genomics.org/plink/2.0/) [10] were used to lift the GRCh37 genomic positions to GRCh38, the genomic build for all other datasets. All genotypes were hard called using the default Plink v2.0 (http://www.cog-genomics.org/plink/2.0/) settings.
Heritability and genetic correlation
For the estimation of the SNP-heritability, two distinct tools were used. With LD score regression (LDSC), it was possible to perform the calculations with the full number of samples as the input for this analysis is the summary statistics. Besides heritability estimates, genetic correlations were also calculated for Aβ42, pTau, tau (to test the similarity in genetic background to pTau), and two previously published AD summary statistics [33, 43]. Precalculated LD scores from the 1,000 Genomes European reference population were obtained online. All estimates were based on HapMap3 SNPs only to ensure high-quality LD score calculations (https://alkesgroup.broadinstitute.org/LDSCORE/). As a rule of thumb, LD Score regression tends to yield very noisy results when applied to datasets with fewer than 5000 individuals (https://github.com/bulik/ldsc/wiki/FAQ) [9]. The summary statistics for the stratified analyses were, therefore, not considered.
For comparison to SNP-heritability estimates of previous studies for Aβ42 and pTau, GCTA v1.9 [74] was applied to the individual-level genotype data of the largest dataset (Netherlands). Other datasets were not considered as the sample size was too low for small standard errors, thereby impossible to draw any meaningful conclusions from the estimates. The restricted maximum likelihood (REML) analysis was performed for the log10-transformed normalized CSF Aβ42 and pTau adjusted for gender, age, and the first 10 principal components. Variance explained could not be calculated for significant loci only as p-values from GWAS results of a large independent sample are unavailable, and calculation in the Dutch sample would be hampered by winners-curse, causing inflation.
Single-marker association
Genome-wide association analysis for each cohort was performed in PLINK v2.0 [10], using linear regression for the continuous phenotypes Aβ42, tau and pTau. Association tests were adjusted for gender, age, assay type (if applicable), and ten ancestry principal components. Only variants with a minor allele frequency threshold above 0.01 were tested. For smaller cohorts (n < 250 individuals) this threshold was set to 0.05 to avoid false positive findings.
Association analyses were repeated for subgroups, stratified according to APOE4 status (based on the high-quality (R2 > 0.8) imputed variants rs429358 and rs7412) or dichotomous Aβ42 status, resulting in the following groups: (1) APOE4 (hetero- and homozygous) carriers; (2) APOE4 non-carriers; (3) individuals with abnormal Aβ42 levels; and (4) individuals with normal Aβ42 levels. After stratification, cohorts with a minimal sample size of 100 individuals were included. Covariates were those described for the main analyses above.
Independent replication
A total of 5042 samples from 15 cohorts were included for the replication analysis. The genetic data for NorCog were generated with 2 different genotyping assays. Extensive QC procedures which are detailed elsewhere [6], allowed for joined genetic analyses of these sub-datasets. Variant association testing was performed according to the association analysis section above. For all other replication cohorts, QC, imputation and association testing procedures are described elsewhere [17]. In short, individual and variant QC standards were met, and imputation was performed using the 1000 Genomes Project Phase 3 reference panel. Each dataset was QCed and imputed independently. The additive linear regression model in PLINK v1.9 [10] was used for single-variant analyses.
Meta-analyses
METAL [71] was used for meta-analyses in stages 1–3 of the per-cohort association results, applying the default approach that utilizes P value and direction of effect, while weighted according to sample size. For stage 1, we used the genome-wide threshold for significance of P < 5 × 10−8, and a suggestive threshold of P < 1 × 10−5 to select variants to study in Stage 2. Stage 2 variants were considered a replication with P < 0.05 and same direction of effect in comparison to stage 1. The genome-wide threshold for significance of P < 5 × 10−8 was used to defined GWAS hits in stage 3.
Colocalization
All variants within 1.5 megabases (Mb) of the lead variant of each genomic risk loci were used in the colocalization analysis. The stage 1 GWAS summary statistics were used for the CSF loci aiming consist sample sizes across the variants. Colocalization comparisons were performed to eQTL data for brain and immune-related tissues and cell-types, which were obtained from the eQTL catalog [39] release 5. The microglia data were obtained from Young et al. [77]. GWAS summary statistics for loci comparison to other GWAS studies were obtained from Kunkle et al. [43] for AD, and from Vojinovic et al. [65] for brain ventricular volume. The GWAS data and eQTL data were trimmed so that all variants overlap. Colocalization was performed with the Coloc R package [24], using the coloc.abf function for the approach assuming a single causal variant, and the runsusie and coloc.susie functions to test for colocalization relaxing this assumption to multiple variants. The latter approach was performed for the comparisons to AD and brain volume loci, and LD matrices were calculated with our own individual-level data for the significant CSF loci using LDstore R package [7]. Default priors were used for prior probability of association with the GWAS data and eQTL data. The prior probability of colocalization was set as 5 × 10−6 as recommended [67]. Nominal P, sample size and MAF were used when beta and variance of beta were not available for the GWAS data or eQTL data. Colocalizations with a posterior probability > 0.8 were considered successful colocalizations. Comparisons were visualized with the R package LocusCompareR (https://github.com/boxiangliu/locuscomparer).
Gene-based analysis
Gene-based and gene-set association tests were performed using MAGMA v1.08 [14], which was implemented by FUMA [69]. The per variant association summary statistics for the main results served as the input, where variants were selected if mapped within 18,870 protein-coding genes (with unique ensembl ID). The mean SNP-wise model was implemented. The Bonferroni-corrected significance threshold was set to P < 2.65 × 10–6, based on the number of tested genes.
Gene mapping
The genome-wide significant loci of the main results were further explored for promising causal AD genes using FUMA [69], after lifting over the results with genomic build GRCh38 to GRCh37 with the UCSC LiftOver Program (https://genome-store.ucsc.edu/). Two gene mapping strategies were used:
Positional mapping maps SNPs to genes based on physical distance (within a 10-kb window) from known protein-coding genes in the human reference assembly (GRCh37/hg19).
eQTL mapping maps SNPs to genes with which they show a significant eQTL association (that is, allelic variation at the SNP is associated with the expression level of that gene). eQTL mapping uses information from 85 brain- and immune-related tissue types in 11 data repositories (BIOSQTL, BloodeQTL, BRAINEAC, CMC, DICE, eQTLcatalogue, eQTLGen, GTEx, PsychENCODE, scRNA_eQTLs, xQTLServer), and is based on cis-eQTLs which can map SNPs to genes up to 1 Mb apart. We used a false discovery rate of 0.05 to define significant eQTL associations.
Phenome-wide association studies (PheWAS)
We conducted phenome-wide association studies (PheWAS) on the top SNPs, rs4844610, rs429358, rs744373, rs9877502, rs4843559. A PheWAS starts out with a single to a few variants of interest that are systematically being tested for association to many phenotypes. We used the ‘phewas’ function of the R-package ‘ieugwasr’ [19, 31]. Using this function, we searched traits that associate with the list of SNPs with P < 1 × 10–7 in all GWAS harmonized summary statistics in the MRC IEU OpenGWAS data infrastructure [31]. In short, this enables us to screen for other traits to which these SNPs are associated. The database (May 2021) includes the GWAS summary statistics of 19,649 traits.
Association with CSF proteomics
We associated the lead variants near GMNC (rs9877502) and in C16orf95 (rs4843559) with CSF proteomics data of two different sources (EMIF-AD MBD and Knight-ADRC). For the EMIF-AD MBD data, a total of 2,136 proteins were quantified centrally using 11-plex tandem mass tag spectrometry in 366 individuals from the EMIF-AD MBD study [61] (subset of Amsterdam Dementia Cohort within EADB). We selected proteins with a maximum of 50% missing values. For related proteins that had identical values due to fragment aspecificity, we randomly selected one protein for analysis (52 proteins were excluded). Out of the 2136 proteins quantified, 1282 (55.4%) proteins respected these criteria and were included in the study.
For the Knight-ADRC data, levels of 1305 proteins were quantified using the SOMAscan assay, a multiplexed, aptamer-based platform CSF (n = 717) [73]. Quality control was performed at the sample and aptamer levels using control aptamers (positive and negative controls) and calibrator samples. As described in detail [73], additional quality control was performed that included limit of detection cut-off, scale factor, coefficient of variation, and outlier variation. Only proteins with a call rate higher than 85% call rate were included. A total of 713 proteins passed quality control. pQTL analyses was performed and reported in previous studies [73].
g:Profiler and Enrichment map [44], a Cytoscape App, were used to perform pathway enrichment analyses on proteins with a certain level of association (EMIF-AD MBD: P < 0.05; Knight-ADRC: P < 0.004, corresponding to proteins with a similar effect size as in EMIF-AD MBD). The results are shown as functionally grouped networks. We used GO biological processes and Reactome as ontology sources. For this explorative analysis, only pathways with P < 0.05 (corrected for multiple testing) are shown.
Effects of AD-associated variants on Aβ42 and pTau
We assessed the most recent GWAS [6] for AD and extracted the top loci of 83 variants (excluding APOE ɛ4 and APOE ɛ2) that showed genome-wide significant association with AD [6]. We extracted Z-scores and P values and plotted them in a heatmap. Rows and columns were clustered using Euclidean distances and average hierarchical clustering. We performed a gene-set enrichment analysis to find molecular pathways enriched within each cluster. The SNP-gene assignment corresponds to the one described in the recent main EADB GWAS [6], including several annotation strategies. When multiple genes were reported to associate with the same SNP (rs12590654 near SLC24A4/RIN3, rs7225151 near SCIMP/RABEP1, rs6846529 near CLNK/HS3ST1, rs7384878 near ZCWPW1/NYAP1 and rs10437655 near CELF1/SPI1), we considered both genes for the gene-set enrichment analysis. In addition, for SNP rs6605556, located in the complex HLA region, we considered HLA-DRB1 gene (eQTL in blood with rs6605556), and for SNPs rs7157106 and rs10131280, both located in the gene-dense IGH region, we considered IGHG2 and IGHV2-70 (eQTLs in blood with rs7157106 and rs10131280, respectively). The gene-set enrichment analysis was performed specifying Biological Processes from Gene Ontology [1, 5] as gene-set and correcting P values with Bonferroni. Biological pathways were considered significant at corrected P < 0.01. To help with the interpretation of each cluster’s function, we plot the most recurring words of the significant terms underlying each cluster using wordclouds. The following R packages were used for these analysis: gprofiler2 [41] and wordcloud2 (https://github.com/lchiffon/wordcloud2).
Results
The overview of the study design is illustrated in Online Resource 2—Fig. 1. The GWA results from 16 studies were combined in stage 1. Variants that reached a suggestive level of significance (P < 1 × 10–5) were subsequently evaluated in an independent sample from 15 studies in stage 2. Finally, the results of stage 1 and stage 2 analyses were combined in stage 3. Detailed information on study participants, CSF acquisition and genotyping is provided in Table 1 and Online Resource 1—Tables 1 and 2. The results for tau and pTau are strongly correlated (rg = 0.94; P = 1.86 × 10–118), and therefore only pTau findings are reported.
Genetic architecture
The fraction of variance in Aβ42 and pTau protein levels that could be explained by the additive effect of the genetic variants tested, was estimated on 0.13 (SE = 0.06) and 0.21 (SE = 0.07) by LDSC, respectively. These SNP-heritabilities are substantially higher than the 0.07 previously estimated with LDSC for the diagnosis AD [80], or similarly reported for AD by this study using the same LDSC method on more recent public GWAS summary statistics of AD [33, 43] (Online Resource 1—Table 3). GCTA estimated the SNP-heritability to be higher for both Aβ42 and pTau, namely 0.27 (SE = 0.13) and 0.34 (SE = 0.12), respectively. Both methods are reporting a higher SNP-heritability for pTau than for Aβ42. Genetic correlation estimates with AD GWAS summary statistics are described in Online Resource 1—Results and Online Resource 1—Table 4.
GWAS variants associated with CSF Aβ42 and pTau
The stage 1 meta-analyses (QQ plots and lambda shown in Online Resource 2—Fig. 14) identified 4 independent significant variant associations, 1 for Aβ42 and 3 for pTau (Table 2). The strongest associations were observed for both Aβ42 and pTau in the APOE locus (Table 2). The variant that determines the APOE ɛ4 allele (rs429358-C) decreased Aβ42 (Z = − 36.29; P = 2.0 × 10–288) and increased pTau (Z = 18.31; P = 6.87 × 10–75) in CSF. In contrast, the variant determining the APOE ɛ2 haplotype (rs7412-T) increased Aβ42 (Z = 11.97; P = 5.09 × 10–33) and decreased pTau (Z = − 6.59; P = 4.49 × 10–11). The APOE ɛ2 association was replicated in stage 2 (Aβ42: Z = 7.27; P = 3.73 × 10–13, and pTau: Z = − 6.43; P = 1.26 × 10–10), and APOE ɛ4 with rs4420638-G for Aβ42 (Z = − 25.51; P = 1.57 × 10–143), and with rs769449-A for pTau (Z = 13.83; P = 1.66 × 10–43), both variants in high linkage disequilibrium with rs429358, as the original APOE ɛ4 variant was not genotyped or imputed in the replication datasets.
Table 2.
Meta-analysis association results for the 3 stages
| Protein | Locus | rsID | Chr | Pos (hg38) | A1 | A2 | Stage 1 | Stage 2 | Stage 3 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Freq A1 | n | Z | P | n | Z | P | n | Z | P | |||||||
| AB42 | 1q32.2 (CR1) | rs4844610 | 1 | 207,629,207 | A | C | 0.20 | 8074 | − 4.87 | 1.13E–06 | 5015 | − 3.54 | 4.08E–04 | 13,089 | − 6.01 | 1.84E–09 |
| APOE | rs429358 | 19 | 44,908,684 | C | T | 0.29 | 8074 | − 36.29 | 2.00E–288 | 4488 | − 25.51 | 1.57E–143 | 12,562 | − 41.68 | 1.00E–379 | |
| pTau | 2q14.3 (BIN1) | rs744373 | 2 | 127,137,039 | G | A | 0.30 | 7798 | 5.39 | 6.99E–08 | 3948 | 3.03 | 2.48E–03 | 11,746 | 6.15 | 7.88E–10 |
| 3q28 (GMNC) | rs9877502 | 3 | 190,951,729 | A | G | 0.39 | 7798 | 9.06 | 1.28E–19 | 4742 | 8.97 | 3.00E–19 | 12,540 | 12.66 | 9.65E–37 | |
| 16q24.2 (C16orf95) | rs4843559 | 16 | 87,191,825 | G | A | 0.38 | 7798 | 6.41 | 1.49E–10 | 3985 | 3.88 | 1.03E–04 | 11,783 | 7.47 | 8.03E–14 | |
| APOE | rs429358 | 19 | 44,908,684 | C | T | 0.28 | 7798 | 18.31 | 6.87E–75 | 4741 | 13.83 | 1.66E–43 | 12,539 | 20.65 | 9.59E–95 | |
Z-statistic summarizes the magnitude and the direction of effect relative to the effect allele. For APOE stage 2 and stage 3, the statistics are reported for the most significant stage 3 variants rs4420638 (G allele) for AB42 and rs769449 (A allele) for pTau, as rs429358 was not genotyped or imputed in the replication datasets
rsID ID number in SNP database, Chr chromosome, Pos genomic base pair position, A1 effect allele, A2 other allele, Freq A1 allele frequency effect allele, n sample size, P p value
In stage 1, no other significant loci were observed for Aβ42. For pTau, we further identified significant associations mapping to two chromosomal regions at 3q28 and 16q24.2 (Table 2). The 3q28-locus (Z = 9.06; P = 1.28 × 10–19) also known as the GMNC locus, was reported previously for its association with pTau [17]. The cohorts from this previous study are the replication cohorts of the current study, thereby logically GMNC was replicated (Z = 8.97; P = 3.00 × 10–19). The 16q24.2 locus (Z = 6.41; P = 1.49 × 10–10), which is novel for pTau, was replicated in the stage 2 meta-analysis (Z = 3.88; P = 1.03 × 10–04).
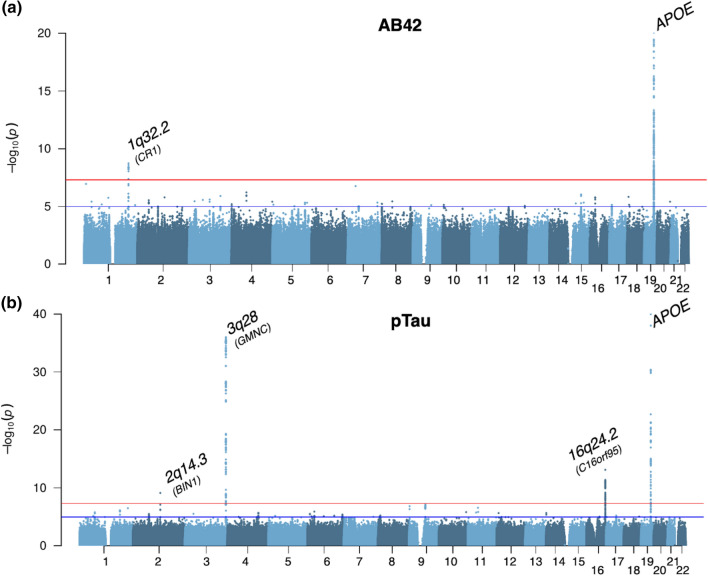
Subsequently, the results from all individual studies were combined in the stage 3 meta-analysis (N = 13,116). In stage 3, two well-known AD loci showed additional genome-wide significant associations (Fig. 1, Table 2) with Aβ42 in chromosomal region 1q32.2 (Z = − 6.01; P = 1.84 × 10–9, CR1; Fig. 1a), and with pTau for the region 2q14.3 (Z = 6.15; P = 7.88 × 10–10, BIN1; Fig. 1b). The per-cohort and zoomed in genomic location details of all significantly associated loci of stage 3 are visualized in Online Resource 2—Figs. 15–20. Colocalization analyses (results detailed in Online Resource 1—Table 5; Online Resource 2—Fig. 21 and 22) showed colocalization of the CR1 (posterior probability = 0.97) and BIN1 (posterior probability = 0.82) loci to these loci in a recent AD GWAS [43]. For the BIN1 locus of the AD GWAS two independent causal signals were observed, of which our BIN1 locus colocalized (posterior probability = 0.84) with the first signal that was tagged by rs6733839, which is the most significant variant of the BIN1 locus of the AD GWAS.
Fig. 1.
Manhattan plots of the stage 3 GWAS results. a Results visualized for CSFAβ42; b Results visualized for CSF pTau. The y-axes are limited to visualize the non-APOE loci. The lowest P values for APOE are 4.07 × 10–355 and 3.74 × 10–94 for Aβ42 and pTau, respectively
Explorative meta-analyses were repeated stratified for APOE (APOE ɛ4 carriers (n = 3240) vs. APOE ɛ4 non-carriers (n = 3201)) and amyloid status (Amyloid normal levels (n = 3182) vs. amyloid abnormal levels (n = 3775)) for stage 1 (QQ plots and lambda shown in Online Resource 2—Figs. 23 and 24), of which the results are visualized in Online Resource 2—Figs. 25 and 26, and detailed in Online Resource 2—Results and Online Resource 1—Table 6. Besides the APOE and GMNC loci, two novel loci are observed that have previously not been linked to any AD phenotype.
Functional interpretation
To interpret the functional effects of the identified variants beyond AD, we performed gene prioritization (based on positional mapping, gene-based association results, and brain and immune eQTL annotations) using FUMA [69], colocalization analyses and PheWAS. The results of the FUMA annotation are detailed in the Online Resource 2—Results and Online Resource 1—Table 7. Ten of our CSF Aβ42 and pTau loci colocalized with one of the brain or immune eQTLs from the 63 tested datasets, which are reported in Online Resource 1—Table 8. The APOE locus for both Aβ42 and pTau colocalized with an eQTL for NKPD1 in a specific immune helper T cell. The CR1 locus for AB42 colocalized with an eQTL for the CR1 gene in 6 brain tissues (hippocampus, caudate, putamen, dorsolateral prefrontal cortex, frontal cortex, cortex). Additionally for the dorsolateral prefrontal cortex, the CR1 locus also colocalized with an eQTL for the AL137789.1 gene. The BIN1 locus colocalized with an eQTL for BIN1 in lymphoblastoid cell lines.
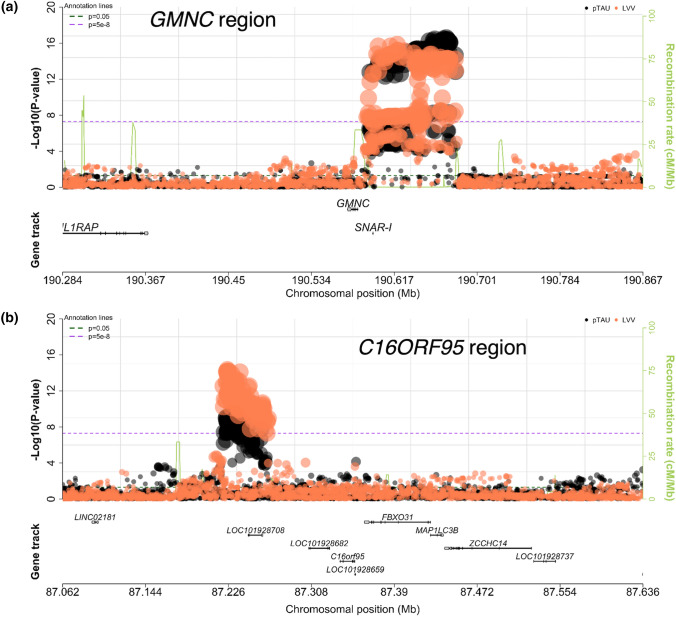
For the PheWAS, using data from publicly available genome-wide association studies (N = 19,649) of the five top variants yielded 529 associations at P < 1 × 10–7 (Online Resource 1—Table 9). The majority is the known wide range of 490 traits associations with the APOE ɛ4 allele. For the other variants 39 associations were reported for 27 unique traits. These traits can be categorized in three groups: traits related to brain ventricular volumes in particular the lateral-ventricle (GMNC and C16orf95), Alzheimer’s disease diagnosis (BIN1 and CR1), and measures of blood cell/lymphocyte counts (CR1). The regional pTau associations of GMNC and C16orf95 overlapped with ventricular volume (Fig. 2). Colocalization results (Online Resource 1—Table 5; Online Resource 2—Figs. 27 and 28) imply the same causal variant for C16orf95 (posterior probability = 0.93). The GMNC locus cannot be convincingly explained by the same causal variant (posterior probability = 0.63). The coloc method combined with Susie reports only 1 causal signal for both GWAS. Of note, the colocalization probability improves to 0.74 when first reducing the GMNC-locus variants to a credible set of variants. Although these results are subthreshold to the predefined posterior probability limit of 0.8, they do imply that the same causal variant is likely underlying the GMNC association signals for pTau and AD.
Fig. 2.
LocusZoom plots showing variant association results for a GMNC and b C16orf95 loci. In black, the pTau association signals of this study; and in orange, the lateral ventricular volume (LVV) association signals observed in other studies
Because of the overlap in effect of GMNC and C16orf95, we hypothesized that these two loci affect the same biological pathways. We explored this hypothesis using CSF proteomics datasets of the EMIF consortium with 1284 quantified proteins and of Knight-ADRC with 696 quantified proteins (of which 42% overlap with the EMIF-AD MBD proteins). For GMNC there were 279 (22%) proteins associated in the EMIF-AD MBD data (Online Resource 1—Table 10) and 255 (36%) proteins in the Knight-ADRC data (Online Resource 1—Table 11). C16orf95 could only be tested in the EMIF-AD MBD data in which 73 (6%) proteins were associated (Online Resource 1—Table 10). Only 2 proteins (CDH9 and DPP6) overlapped between the 2 loci. We studied the overlap in affected pathways between the associated protein lists. For GMNC, consistent functional group networks between the 2 tested datasets were axon guidance and ephrin signaling (Online Resource 2—Fig. 29, Online Resource 1—Tables 12 and 13), while for C16orf95 (only based on the EMIF-AD MBD data) glycosaminoglycan metabolism and ECM organization were overrepresented functional groups (Online Resource 2—Fig. 30, Online Resource 1—Table 14). There was little overlap between the loci in the pathways that emerged from the protein lists.
Relation to AD-associated genetic variants
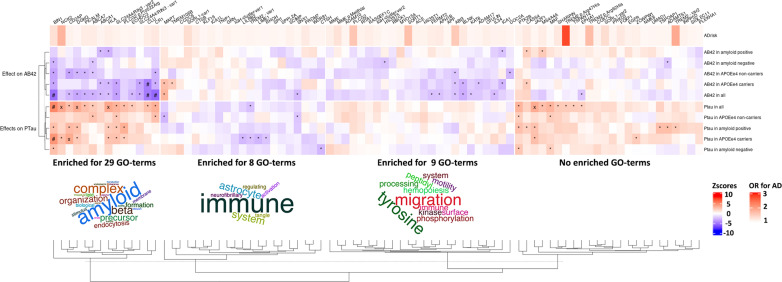
Because of the evident overlap in etiology with clinical AD dementia, we examined the association of all known AD loci (excluding the APOE locus) with CSF Aβ42 and pTau. This analysis has an explorative character as the small contribution of each individual AD loci to CSF amyloid and pTau is in general reflected in moderate effect sizes and association signals. However, patterns of reasonable signal could facilitate the generation of biological hypotheses to test in future experiments. The results are shown in the heatmap of Fig. 3 and Online Resource 1—Tables 15 and 16. The variants could be clustered in 4 groups of AD-associated genes based on their associations with Aβ42 and pTau. The first cluster of 14 variants showed strong association with both decreased levels of Aβ42 and increased levels of pTau in CSF. A pathway enrichment analysis of the genes associated with the variants showed 29 GO terms enriched and ‘amyloid’ is the common denominator in the names of these terms, of which the signal is mostly driven by BIN1, PICALM ABCA7 and CLU. The second cluster contained 21 variants and included genes that have also been related to other dementia types (e.g., GRN, TMEM107B, SNX1, MAPT, CTSB and CTSH). This cluster was associated with decreased pTau levels, an no general effect on Aβ42 levels. Pathway analysis of the genes suggests an enrichment for 8 GO terms of which the names have ‘immune’ as a common denominator that is mostly driven by 12 genes of which TREM2 and GRN are the most frequent contributors. The third cluster consisted of 22 variants, which were related to decreased levels of Aβ42 but not increased levels of pTau. Nine GO terms were enriched and ‘migration’ and ‘tyrosine’ are the words that occur most often in these terms, though each word only based on 2 GO terms each. The last cluster of 20 variants group because they increased pTau, but did not decrease Aβ42 levels. No GO terms are significantly enriched in this gene cluster.
Fig. 3.
The effects of all AD-associated loci. The names of the loci are named according to their linked gene names in Bellinguez et al. (2021). Hierarchical clustering was performed on the rows and columns using Eucledian distances and the method ‘average’ for clustering Pathway enrichment analyses was performed on the four first clusters. The enrichment analyses are in Online Resource 1—Table 9. The upper bar shows the odds ratio for AD, where alleles for variants with protective effects have been flipped to show AD-risk increasing effects for all variants. The increases in Aβ42 and pTau (positive Zscores) are shown in red and decreases in AB42 and PTau (negative Z scores) are shown in blue. * = P value < 0.05, X = 0.05 < P value < 0.001, # = 0.001 < P value < 5 × 10–8, $ = P value < 5 × 10–8
Discussion
We identified 2 loci (CR1 and APOE) for Aβ42, and 4 loci (BIN1, GMNC, C16orf95 and APOE) for pTau in a total of 13,116 individuals (discovery n = 8074; replication n = 5402 individuals). In concordance with previous GWAS studies [12, 17], both proteins showed the strongest association for APOE, where APOE ɛ4 decreased amyloid-beta levels and increased pTau levels, while APOE ɛ2 had the opposite effect. We confirmed GMNC as a risk factor for CSF pTau levels. We identified CR1 as a novel locus for CSF Aβ42 levels, and we observed 2 novel loci (BIN1 and C16orf95) for CSF pTau levels. So other than APOE, no risk loci overlap is observed for Aβ42 and pTau, implying at least partly separate genetic backgrounds for both pathological hallmarks. Amyloid-beta appears to be dominated by the effect of APOE, while pTau is influenced by multiple genetic components. Such a divergence in genetic influences is not in concordance with a genetic etiology where accumulation of Tau tangles is a direct downstream effect of amyloid plaque formation, as proposed by the amyloid cascade theory [29, 30]. Rather, it seems that these pathologies have an independent component in its origin, as already extensively reviewed [64], thereby implying a dual-cascade hypothesis in which sporadic AD pathogenesis is caused by defects in correlated but independent cellular processes. This is highly relevant biological knowledge for the development of potential AD treatments. The limited clinical efficacy of agents that aim to reduce beta-amyloid plaques might potentially be due to this [76].
In line with this implication is the observed difference in genetic subgroups based on CSF protein patterns as proposed by the explorative cluster analysis, suggesting multiple Aβ42 and pTau related biological pathways to be involved in the etiology AD. The first genetic subgroup is defined by the word ‘amyloid’, where BIN1, PICALM, ABCA7 and CLU contribute most to this signal. The involvement of these genes in amyloid-beta pathology is supported by previous studies [15, 46, 63, 79]. ‘Immune’ is labeling the second subgroup, thereby implying that the genes included in this cluster are influencing AD by altering immune responses. The largest genetic drivers of this signal are TREM2 and GRN which have been previously described for their functional role in AD via the immune system [45, 54]. A substantial involvement of the immune system in AD is a generally accepted concept, supported by clinical, functional and genetic research [8]. As mentioned in the Results section, this second cluster includes genes that have also been related to other dementia types (e.g., GRN, TMEM107B, SNX1, MAPT, CTSB and CTSH). This is in line with more recent insights where dysfunction of the immune response is seen as a common cause for multiple neurodegenerative diseases [28]. The third and fourth clusters are less straightforward to interpret as many terms are similarly enriched, or no significant enrichment is observed. The definition of the first 3 subclasses in its current state is the best approximate within reach, though presumably not a perfect reflection of the genuine genetic etiology of AD. Subsequent genetic studies with larger sample sizes are anticipated to facilitate an improved understanding of the biological implications of the underlying genetic subclasses.
The variety in subclasses of genetic contributors for AD etiology could mean that different patient groups might benefit from distinct AD treatment depending on the biological pathway that is affected. Although, our genetic results in its current state are not applicable for clinical trials, we believe by improving the definition of the amyloid-tau clusters (e.g., by increasing sample sizes), it will be possible to assign individuals to one of the groups based on the distribution of their AD risk variants. For example, individuals with relatively more AD risk variants in the first cluster should preferably be included in a clinical trial addressing amyloid formation, while individuals with higher sub PRS scores for the second immune cluster, would have a higher probability benefitting from a clinical trial targeting the immune system. Alternatively, when sample sizes are increasing for future CSF amyloid and tau GWAS, an amyloid or tau PRS could be calculated to identify individuals at high genetic risk for amyloid and tau, thereby defining the individuals that should be included in trials aiming to reduce amyloid plagues or tau neurofibrillary tangles, respectively.
As mentioned above, APOE is the strongest locus for both CSF Aβ42 and pTau. The functional implications of the relation between APOE and the 2 pathological hallmarks has been excessively studied and reviewed in the scientific field of Alzheimer’s disease [20, 32, 60, 64]. In short, studies in human and mice models have shown higher Aβ42 levels and plaque load for APOE4 carriers. Whether this is a result of gain of toxic effect (e.g., enhanced ability of the APOE4 isoform to bind to Aβ42 [40]), or loss of defensive mechanism (e.g., less effective microglial response[20]), or a combination of both, remains to be elucidated. For Tau, studies have shown hyperphosphorylation and faster accumulation for APOE4 carriers, both in mice and human models [32, 64]. Again, the functional route through which APOE is affecting the Tau aggregates is unknown, and multiple cellular processes are being proposed as mediators, including neuronal endocytosis, lipid metabolism and glial function. An important functional observation though is the influence of APOE4 on tau accumulation in absence of Aβ42 pathology [55, 68], which is in agreement with the genetic implications of this current study. An unexpected observation for the APOE locus in this study is the successful colocalization with an eQTL for the NKPD1 in a specific type of T cells. However, due to the convincing functional involvement of APOE in AD pathogenesis via the presence of cysteine or arginine at APOE amino acid residuals 112 and 158, that we anticipate this successful NKPD1 eQTL colocalization result to be a false positive.
The CR1-locus findings are in agreement with the well-known observed association for AD risk, where rs6656401 (R2 = 0.88 with lead SNP of current study) carriers are more susceptible for AD. The similarity in these association signals is strengthened by the convincing colocalization of the CR1 locus between our CSF pTau observation and the CR1 locus of Kunkle et al. [43]. We furthermore observe a successful colocalization with an eQTL for the CR1 gene in multiple brain regions. In concordance with this observation is functional work on the effect of CR1 further which suggests that CR1 is involved in AD pathogenesis by regulating Aβ42 clearance in the brain itself, but also peripherally in blood cells [81]. More recent research in red blood cells of AD patients showed deficient CR1 immunoreactivity, including CR1-mediated capture of circulating amyloid-beta [35]. They observed decreased CR1 protein levels in red blood cells for CR1 SNPs that associate with higher AD risk. The second novel locus in this study is BIN1 for CSF pTau, for which we observe a successful colocalization for an eTQL for the BIN1 gene in lymphoblastoid cell lines, implying BIN1 to be the causal gene for this locus. The BIN1-locus furthermore localizes with the same locus in the AD GWAS, implying that a causal variant of the BIN1-locus for AD contributes to AD pathogenesis via tau pathology. BIN1 has already been linked to Tau pathology in several functional studies, first shown in fruit flies where a decrease in the BIN1 ortholog gene expression suppressed Tau-mediated neurotoxicity [11]. More recently, research in mice showed physical protein interaction between BIN1 and Tau [51], and BIN1 involvement in Tau-dependent hyperexcitability in AD [66]. In human subjects, BIN1-carriers were associated with lower memory performance, mediated by higher tau-PET levels [22]. Our observed BIN1-Tau association contributes valuable knowledge on this topic, by observing the same trend in a substantial larger study (89 vs 13,118 individuals) using different techniques to measure Tau pathology (PET vs. CSF). We provide in vivo confirmation that CR1 is associated with AD via Aβ42, while BIN1 relates to Tau pathology.
The third novel locus in this study is the region on genomic location 16q24.2 for pTau, of which the strongest associated variants are located within intronic regions of C16orf95. This locus has not been linked to (p)Tau pathology or AD in previous research, though it associated to lateral ventricular volume in the CHARGE study, including 23.5 k healthy individuals [65]. Similarly, the 3q28-locus for pTau from our findings colocalizes inconclusively (posterior probability of 0.73) to lateral ventricular volume by the CHARGE study, implying that the same genetic risk factors contribute to both phenotypes, strengthening the notion that neurodegeneration and (p)Tau pathology are highly correlated. 3q28 has been linked to (p)Tau by previous CSF studies in dementia cohorts and was identified as the GMNC locus [12, 17]. In comparison to the latest GWAS of Deming et al. [17], increasing the sample size with a small 10 k individuals in this study strengthens the association of this locus from 3.07 × 10–11 to 1.19 × 10–36, thereby turning it into a well-established locus for pTau pathology (similar for Tau: Deming P = 3.07 × 10−11 (n = 3146); current P = 9.65 × 10–37 (n = 12,540). The C16orf95-locus convincingly colocalized with the same locus in ventricular volume. The formerly reported directions of effect of GMNC and C16orf95 for ventricular volume are counterintuitive. For both loci the allele that associated with an increase in pTau pathology in our dementia cohorts associated with a smaller ventricular volume, implying less neurodegeneration. We explored if these loci work through the same biological pathways using CSF proteomics data. The consistently highlighted functional groups for the GMNC locus (axon guidance and ephrin signaling) were different than for the C16orf95-locus (extracellular matrix components), thereby implying at least partly distinct functional routes via which they influence pTau protein levels in CSF. The power of the proteomics dataset is rather limited. We anticipate that these analyses would benefit and potentially find overlapping biological pathways, when AD proteomics datasets at hand will increase in sample size, or apply the same proteomics assay (rather than using different ones with little overlap in proteins).
We furthermore identified two novel loci in the stratified analyses for Aβ42 levels: 7q11.22 for APOE ɛ4 non-carriers and 12q13.3 in individuals with abnormal amyloid levels. We were unable to test for replication of these loci in independent replication datasets, as such analyses were not performed. Neither loci have been previously linked to AD, or any other trait. The lead SNP for the locus on chromosome 7 is a common intronic variant for the lncRNA LOC105375341, which according to GTEx is only expressed in testis and prostate, and thereby not a promising causal gene for AD or AD-related phenotypes. The locus on chromosome 12 consists of just one rare intronic variant that was only detected in the Dutch cohort with the largest sample size (n = 498). Future studies including more cohorts of large sample sizes are required to study this rare variant in more detail.
Besides APOE and GMNC, no other loci from the latest GWAS of CSF amyloid and tau [17] were replicated by this study. For Aβ42, it was not possible to test the GLIS1-locus as this variant was not observed in our data, which is in concordance with the gnomAD browser [37] reporting extremely low coverage and a MAF of 0.007 for rs185031519, the strongest GLIS1 SNP in Deming et al. For the other loci that we were unable to replicate (SERPINB1 for Aβ42; and GLIS3, PCDH8, CTDP1 for pTau), there were no significant associations (P > 0.05) despite substantial sample sizes (n > 7000). These differences in findings might be due to differences in study design, for example inclusion of cohorts with other diagnoses and/or differences in analysis strategies.
The GCTA-based SNP-heritability estimates of 27% and 34% for Aβ42 and pTau, respectively, are in a similar range to the estimates calculated by Deming et al. [17] (36% for Aβ42 and 25% for pTau). Notable is that the previous study estimated Aβ42 to be most heritable, while we observed the highest estimate for pTau, though the standard errors were about 10% for both studies. Our LDSC-based SNP-heritabilities show a similar trend with higher estimates for pTau. Furthermore, these LDSC-based 13% for Aβ42 and 21% for pTau protein levels are considerably higher than the 7% previously observed for the diagnosis AD. The higher heritability for the tested CSF protein levels strengthens the assumption that more objective measurable biological properties are more strongly associated with AD pathogenesis than the diagnostic classifications.
Due to this assumption, the number of risk loci identified in this study might be lower than anticipated. However, our study design is suboptimal for obtaining the most powerful model for genetic association identification. Meta-analyzing 31 heterogeneous cohorts (differences in genotyping array, imputation references, CSF protein assays, and patient/control ratios) is more challenging rather than analyzing a homogeneous dataset of similar sample size. Alternatively, increasing the sample size by adding more cohorts to increase power is presumably a more feasible approach for the future.
In conclusion, the current findings clearly show that studying the genetic effects of AD-related endophenotypes has the potential to reveal novel associations, and highlight important biological insights. The clear distinction in genetic findings for amyloid-beta and tau emphasizes the (partly) genetic independence of these two biological mechanisms in AD pathogenesis. Moreover, the identification of CR1 and BIN1, which are the second and third strongest associated AD locus after APOE, furthermore implies that by increasing sample size of genetic analysis in CSF biomarkers it will become more apparent through which biological mechanisms certain AD loci have their effect on AD pathogenesis. Even larger collaborative efforts with more homogeneous sample definitions are, therefore, encouraged to be undertaken to enhance our genetic understanding of AD, ultimately leading to improved biological knowledge for the development of drug treatment.
Supplementary Information
Below is the link to the electronic supplementary material.
Acknowledgements
This work was supported by a grant (European Alzheimer DNA BioBank, EADB) from the EU Joint Programme, Neurodegenerative Disease Research (JPND). Amsterdam dementia Cohort (ADC): Research of the Alzheimer center Amsterdam is part of the neurodegeneration research program of Amsterdam Neuroscience. The Alzheimer Center Amsterdam is supported by Stichting Alzheimer Nederland and Stichting VUmc fonds. The clinical database structure was developed with funding from Stichting Dioraphte. Genotyping of the Dutch case-control samples was performed in the context of EADB (European Alzheimer DNA biobank) funded by the JPco-fuND FP-829-029 (ZonMW projectnumber 733051061).Part of the work described in this study was carried out in the context of the Parelsnoer Institute (PSI). PSI was part of and funded by the Dutch Federation of University Medical Centers and has received initial funding from the Dutch Government (from 2007-2011). Since 2020, this work was carried out in the context of Parelsnoer clinical biobanks at Health-RI (https://www.health-ri.nl/initiatives/parelsnoer). Part of the genotyping included in this work was funded by the JPND EADB grant (German Federal Ministry of Education and Research (BMBF) grant: 01ED1619A). Alfredo Ramirez is also supported by the German Research Foundation (DFG) grants Nr: RA 1971/6-1, RA1971/7-1, and RA 1971/8-1. We would like to thank patients and controls who participated in this project. The Genome Research @ Fundació ACE project (GR@ACE) is supported by Grifols SA, Fundación bancaria ‘La Caixa’, Fundació ACE, and CIBERNED. A.R. and M.B. receive support from the European Union/EFPIA Innovative Medicines Initiative Joint undertaking ADAPTED and MOPEAD projects (grant numbers 115975 and 115985, respectively). M.B. and A.R. are also supported by national grants PI13/02434, PI16/01861, PI17/01474, PI19/01240 and PI19/01301. Acción Estratégica en Salud is integrated into the Spanish National R + D + I Plan and funded by ISCIII (Instituto de Salud Carlos III)–Subdirección General de Evaluación and the Fondo Europeo de Desarrollo Regional (FEDER–‘Una manera de hacer Europa’). The position held by I.dR. is funded by grant. FI20/00215. PFIS Contratos Predoctorales de Formación en Investigación en Salud. We would like to thank UCL Genomics, London, UK, for performing the genotyping analyses of the samples within the Gothenburg H70 Birth Cohort Studies and Clinical AD Sweden. The recruitment and clinical characterization of research participants at Washington University were supported by NIH P30AG066444, and P01AG003991. This work was supported by access to equipment made possible by the Hope Center for Neurological Disorders, the Neurogenomics and Informatics Center (NGI: https://neurogenomics.wustl.edu/)and the Departments of Neurology and Psychiatry at Washington University School of Medicine. Research at the Belgian EADB site is funded in part by the Alzheimer Research Foundation (SAO-FRA), The Research Foundation Flanders (FWO), and the University of Antwerp Research Fund. FK is supported by a BOF DOCPRO fellowship of the University of Antwerp Research Fund. The work of Valdecilla was supported by grants from the Instituto de Salud Carlos III (Fondo de Investigación Sanitario, PI08/0139, PI12/02288, PI16/01652, and PI20/01011), the JPND (DEMTEST PI11/03028), the CIBERNED, and the Siemens Healthineers. We thank the Valdecilla Biobank (PT17/0015/0019), integrated into the Spanish Biobank Network, for their support and collaboration in sample collection and management. Our heartfelt thanks to the participants of the Valdecilla Cohort for their generosity. The work of ADGEN was supported by EU Joint Programme—Neurodegenerative Disease Research (301220) and the Academy of Finland (338182). The DELCODE study (Study-ID:BN012 ) was supported and conducted by the German Center for Neurodegenerative Diseases (DZNE). The data samples were provided by the DELCODE study group. Details and participating sites can be found at www.dzne.de/en/research/studies/clinical-studies/delcode. The German Dementia Competence Network (KND) is funded by the German Federal Ministry of Education and Research (BMBF) grants Number: 01G10102, 01GI0420, 01GI0422, 01GI0423, 01GI0429, 01GI0431, 01GI0433, 04GI0434, 01GI0711. WF, SvdL, HHolstege, CT and PhS are recipients of ABOARD, which is a public-private partnership receiving funding from ZonMW (#73305095007) and Health~Holland, Topsector Life Sciences & Health (PPP-allowance; #LSHM20106). More than 30 partners participate in ABOARD (www.aboard-project.nl). ABOARD also receives funding from de Hersenstichting, Edwin Bouw Fonds and Gieskes-Strijbisfonds. IEJ was partially supported by NWO Gravitation program BRAINSCAPES: A Roadmap from Neurogenetics to Neurobiology (NWO: 024.004.012). AZ was supported by the Swedish Alzheimer Foundation (AF-939988, AF-930582, AF-646061, AF-741361), and the Dementia Foundation (2020-04-13, 2021-04-17). ISk was supported by the Swedish state under the agreement between the Swedish government and the county councils, the ALF-agreement (ALF 716681), the Swedish Research Council (no 11267, 825-2012-5041, 2013-8717, 2015-02830, 2017-00639, 2019-01096), Swedish Research Council for Health, Working Life and Welfare (no 2001-2646, 2001-2835, 2001-2849, 2003-0234, 2004-0150, 2005-0762, 2006-0020, 2008-1229, 2008-1210, 2012-1138, 2004-0145, 2006-0596, 2008-1111, 2010-0870, 2013-1202, 2013-2300, 2013-2496), Swedish Brain Power, Hjärnfonden, Sweden (FO2016-0214, FO2018-0214, FO2019- 0163), the Alzheimer's Association Zenith Award (ZEN-01-3151), the Alzheimer's Association Stephanie B. Overstreet Scholars (IIRG-00-2159), the Alzheimer's Association (IIRG-03-6168, IIRG-09-131338) and the Bank of Sweden Tercentenary Foundation. SK was supported by the Swedish state under the agreement between the Swedish government and the county councils, the ALF-agreement (ALFGBG-81392, ALF GBG-771071), the Swedish Alzheimer Foundation (AF-842471, AF-737641, AF-939825), and the Swedish Research Council (2019-02075). MW was supported by the Swedish Research Council 2016-01590. DP was supported by BRAINSCAPES: A Roadmap from Neurogenetics to Neurobiology (grant no. 024.004.012), and a European Research Council advanced grant (Grant No, ERC-2018-AdG GWAS2FUNC 834057. HZ is a Wallenberg Scholar supported by grants from the Swedish Research Council (2018-02532), the European Research Council (681712), Swedish State Support for Clinical Research (ALFGBG-720931), the Alzheimer Drug Discovery Foundation (ADDF), USA (201809-2016862), the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No 860197 (MIRIADE), and the UK Dementia Research Institute at UCL. KB was supported by the Swedish Research Council (#2017-00915), the Alzheimer Drug Discovery Foundation (ADDF), USA (#RDAPB-201809-2016615), the Swedish Alzheimer Foundation (#AF-742881), Hjärnfonden, Sweden (#FO2017-0243), the Swedish state under the agreement between the Swedish government and the County Councils, the ALF-agreement (#ALFGBG-715986), the European Union Joint Program for Neurodegenerative Disorders (JPND2019-466-236), the National Institute of Health (NIH), USA, (grant #1R01AG068398-01), and the Alzheimer’s Association 2021 Zenith Award (ZEN-21-848495). CC receives support from the National Institutes of Health (R01AG044546, R01AG064877, RF1AG053303, R01AG058501, U01AG058922, RF1AG058501, R01AG064614), and the Chuck Zuckerberg Initiative (CZI).
Data availability statement
Genome-wide summary statistics have been deposited to the European Bioinformatics Institute GWAS Catalog (https://www.ebi.ac.uk/gwas/) under accession IDs GCST90129599 and GCST90129600.
Declarations
Conflict of interest
Research programs of Wiesje van der Flier have been funded by ZonMW, NWO, EU-FP7, EU-JPND, Alzheimer Nederland, CardioVascular Onderzoek Nederland, Health ~ Holland, Topsector Life Sciences & Health, stichting Dioraphte, Gieskes-Strijbis fonds, stichting Equilibrio, Pasman stichting, stichting Alzheimer & Neuropsychiatrie Foundation, Biogen MA Inc, Boehringer Ingelheim, Life-MI, AVID, Roche BV, Fujifilm, Combinostics. WF holds the Pasman chair. WF is recipient of ABOARD, which is a public–private partnership receiving funding from ZonMW (#73305095007) and Health ~ Holland, Topsector Life Sciences & Health (PPP-allowance; #LSHM20106). WF has performed contract research for Biogen MA Inc, and Boehringer Ingelheim. WF has been an invited speaker at Boehringer Ingelheim, Biogen MA Inc, Danone, Eisai, WebMD Neurology (Medscape), Springer Healthcare. WF is consultant to Oxford Health Policy Forum CIC, Roche, and Biogen MA Inc. WF participated in advisory boards of Biogen MA Inc and Roche. All funding is paid to her institution. WF was associate editor of Alzheimer, Research & Therapy in 2020/2021. WF is associate editor at Brain. CC receives research support from: Biogen, EISAI, Alector and Parabon. The funders of the study had no role in the collection, analysis, or interpretation of data; in the writing of the report; or in the decision to submit the paper for publication. CC is a member of the advisory board of Vivid genetics, Halia Therapeutics and ADx Healthcare. HZ has served at scientific advisory boards for Denali, Roche Diagnostics, Wave, Samumed, Siemens Healthineers, Pinteon Therapeutics and CogRx, has given lectures in symposia sponsored by Fujirebio, Alzecure and Biogen, and is a co-founder of Brain Biomarker Solutions in Gothenburg AB (BBS), which is a part of the GU Ventures Incubator Program (outside submitted work). KBlennow has served as a consultant, at advisory boards, or at data monitoring committees for Abcam, Axon, Biogen, JOMDD/Shimadzu. Julius Clinical, Lilly, MagQu, Novartis, Roche Diagnostics, and Siemens Healthineers, and is a co-founder of Brain Biomarker Solutions in Gothenburg AB (BBS), which is a part of the GU Ventures Incubator Program. OAA is a consultant to HealthLytix, and received speaker’s honorarium from Lundbeck and Sunovion. SE has served at scientific advisory boards for Biogen, Danone, Eisai, icometrix, Pfizer, Novartis, Nutricia, Roche and has received unrestricted research grants from ADx Neurosciences and Janssen Pharmaceutica. CC receives research support from: Biogen, EISAI, Alector, GSK and Parabon. The funders of the study had no role in the collection, analysis, or interpretation of data; in the writing of the report; or in the decision to submit the paper for publication. CC is a member of the advisory board of Vivid genetics, Halia Therapeutics and ADx Healthcare. HHampel is an employee of Eisai Inc. and serves as Senior Associate Editor for the Journal Alzheimer’s & Dementia; during the past three years he had received lecture fees from Servier, Biogen and Roche, research grants from Pfizer, Avid, and MSD Avenir (paid to the institution), travel funding from Eisai, Functional Neuromodulation, Axovant, Eli Lilly and company, Takeda and Zinfandel, GE-Healthcare and Oryzon Genomics, consultancy fees from Qynapse, Jung Diagnostics, Cytox Ltd., Axovant, Anavex, Takeda and Zinfandel, GE Healthcare, Oryzon Genomics, and Functional Neuromodulation, and participated in scientific advisory boards of Functional Neuromodulation, Axovant, Eisai, Eli Lilly and company, Cytox Ltd., GE Healthcare, Takeda and Zinfandel, Oryzon Genomics and Roche Diagnostics. DA participated in advisory boards from Fujirebio-Europe and Roche Diagnostics and received speaker honoraria from Fujirebio-Europe, Roche Diagnostics, Nutricia, Krka Farmacéutica S.L., Zambon S.A.U. and Esteve Pharmaceuticals S.A. OG reports consulting fees from Eli Lilly, grants to his institution from Actelion, and prescreening activities for Julius Clinical/Toyama. ABK has been a PI in the drug trials Roche BN29553, Boehringer-Ingelheim 1346.0023 and is PI in Novo Nordisk NN6535-4730. AL has received personal fees for advisory board services and/or speaker honoraria from Fujirebio-Europe, Roche Diagnostics, Nutricia,Krka Farmacéutica SL, Biogen and Zambon. PSJ has received personal fees for advisory board from Roche Diagnostics and Zambon. GS participated in one advisory board meeting from Biogen. MI is a paid consultant to BioArctic AB. KS is editor at Acta Neuropathologica and associate editor at Alzheimer’s Research & Therapy. HZ has served at scientific advisory boards for Denali, Roche Diagnostics, Wave, Samumed, Siemens Healthineers, Pinteon Therapeutics and CogRx, has given lectures in symposia sponsored by Fujirebio, Alzecure and Biogen, and is a co-founder of Brain Biomarker Solutions in Gothenburg AB (BBS), which is a part of the GU Ventures Incubator Program (outside submitted work). All other authors declare no financial interests or potential conflicts of interest.
Footnotes
Members of “EADB consortium” and “The GR@ACE study group” are provided in Supplementary File 2.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Iris E. Jansen, Sven J. van der Lee, Duber Gomez-Fonseca, Itziar de Rojas, Maria Carolina Dalmasso and Benjamin Grenier-Boley are co-first authors.
Agustín Ruiz, Alfredo Ramirez, Carlos Cruchaga, Jean-Charles Lambert and Wiesje van der Flier are co-last authors.
Contributor Information
Iris E. Jansen, Email: i.e.jansen@vu.nl
Wiesje van der Flier, Email: wm.vdflier@amsterdamumc.nl.
References
- 1.(2021) The Gene Ontology resource: enriching a GOld mine. Nucleic Acids Res 49:D325–D334. 10.1093/nar/gkaa1113 [DOI] [PMC free article] [PubMed]
- 2.Alcolea D, Martínez-Lage P, Sánchez-Juan P, Olazarán J, Antúnez C, Izagirre A, et al. Amyloid precursor protein metabolism and inflammation markers in preclinical Alzheimer disease. Neurology. 2015;85:626–633. doi: 10.1212/wnl.0000000000001859. [DOI] [PubMed] [Google Scholar]
- 3.Alcolea D, Pegueroles J, Muñoz L, Camacho V, López-Mora D, Fernández-León A, et al. Agreement of amyloid PET and CSF biomarkers for Alzheimer's disease on Lumipulse. Ann Clin Transl Neurol. 2019;6:1815–1824. doi: 10.1002/acn3.50873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Almdahl IS, Lauridsen C, Selnes P, Kalheim LF, Coello C, Gajdzik B, et al. Cerebrospinal fluid levels of amyloid beta 1–43 mirror 1–42 in relation to imaging biomarkers of Alzheimer's disease. Front Aging Neurosci. 2017;9:9. doi: 10.3389/fnagi.2017.00009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–29. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bellenguez C, Küçükali F, Jansen IE, Kleineidam L, Moreno-Grau S, Amin N, et al. New insights into the genetic etiology of Alzheimer’s disease and related dementias. Nat Genet. 2022;54:412–436. doi: 10.1038/s41588-022-01024-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Benner C, Havulinna AS, Järvelin MR, Salomaa V, Ripatti S, Pirinen M. Prospects of fine-mapping trait-associated genomic regions by using summary statistics from genome-wide association studies. Am J Hum Genet. 2017;101:539–551. doi: 10.1016/j.ajhg.2017.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bettcher BM, Tansey MG, Dorothée G, Heneka MT. Peripheral and central immune system crosstalk in Alzheimer disease—a research prospectus. Nat Rev Neurol. 2021;17:689–701. doi: 10.1038/s41582-021-00549-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bulik-Sullivan BK, Loh P-R, Finucane HK, Ripke S, Yang J, Patterson N, Schizophrenia Working Group of the Psychiatric Genomics C et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–295. doi: 10.1038/ng.3211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience. 2015 doi: 10.1186/s13742-015-0047-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chapuis J, Hansmannel F, Gistelinck M, Mounier A, Van Cauwenberghe C, Kolen KV, et al. Increased expression of BIN1 mediates Alzheimer genetic risk by modulating tau pathology. Mol Psychiatry. 2013;18:1225–1234. doi: 10.1038/mp.2013.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cruchaga C, Kauwe JS, Harari O, Jin SC, Cai Y, Karch CM, et al. GWAS of cerebrospinal fluid tau levels identifies risk variants for Alzheimer's disease. Neuron. 2013;78:256–268. doi: 10.1016/j.neuron.2013.02.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Das S, Forer L, Schönherr S, Sidore C, Locke AE, Kwong A, et al. Next-generation genotype imputation service and methods. Nat Genet. 2016;48:1284–1287. doi: 10.1038/ng.3656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11:e1004219. doi: 10.1371/journal.pcbi.1004219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.De Roeck A, Van Broeckhoven C, Sleegers K. The role of ABCA7 in Alzheimer’s disease: evidence from genomics, transcriptomics and methylomics. Acta Neuropathol. 2019;138:201–220. doi: 10.1007/s00401-019-01994-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.de Rojas I, Moreno-Grau S, Tesi N, Grenier-Boley B, Andrade V, Jansen IE, et al. Common variants in Alzheimer’s disease and risk stratification by polygenic risk scores. Nat Commun. 2021;12:3417. doi: 10.1038/s41467-021-22491-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Deming Y, Li Z, Kapoor M, Harari O, Del-Aguila JL, Black K, et al. Genome-wide association study identifies four novel loci associated with Alzheimer's endophenotypes and disease modifiers. Acta Neuropathol. 2017;133:839–856. doi: 10.1007/s00401-017-1685-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.El Khoury J, Toft M, Hickman SE, Means TK, Terada K, Geula C, et al. Ccr2 deficiency impairs microglial accumulation and accelerates progression of Alzheimer-like disease. Nat Med. 2007;13:432–438. doi: 10.1038/nm1555. [DOI] [PubMed] [Google Scholar]
- 19.Elsworth B, Lyon M, Alexander T, Liu Y, Matthews P, Hallett J, et al. The MRC IEU OpenGWAS data infrastructure. bioRxiv. 2020 doi: 10.1101/2020.08.10.244293. [DOI] [Google Scholar]
- 20.Fitz NF, Nam KN, Wolfe CM, Letronne F, Playso BE, Iordanova BE, et al. Phospholipids of APOE lipoproteins activate microglia in an isoform-specific manner in preclinical models of Alzheimer’s disease. Nat Commun. 2021;12:3416. doi: 10.1038/s41467-021-23762-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fladby T, Pålhaugen L, Selnes P, Waterloo K, Bråthen G, Hessen E, et al. Detecting at-risk Alzheimer's disease cases. J Alzheimers Dis. 2017;60:97–105. doi: 10.3233/jad-170231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Franzmeier N, Rubinski A, Neitzel J, Ewers M, Weiner MW, Aisen P, et al. The BIN1 rs744373 SNP is associated with increased tau-PET levels and impaired memory. Nat Commun. 2019;10:1766. doi: 10.1038/s41467-019-09564-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Geijselaers SLC, Aalten P, Ramakers I, De Deyn PP, Heijboer AC, Koek HL, et al. Association of cerebrospinal fluid (CSF) insulin with cognitive performance and CSF biomarkers of Alzheimer's disease. J Alzheimers Dis. 2018;61:309–320. doi: 10.3233/jad-170522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Giambartolomei C, Vukcevic D, Schadt EE, Franke L, Hingorani AD, Wallace C, et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 2014;10:e1004383. doi: 10.1371/journal.pgen.1004383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Goate A, Chartier-Harlin MC, Mullan M, Brown J, Crawford F, Fidani L, et al. Segregation of a missense mutation in the amyloid precursor protein gene with familial Alzheimer's disease. Nature. 1991;349:704–706. doi: 10.1038/349704a0. [DOI] [PubMed] [Google Scholar]
- 26.Gottesman II, Gould TD. The endophenotype concept in psychiatry: etymology and strategic intentions. Am J Psychiatry. 2003;160:636–645. doi: 10.1176/appi.ajp.160.4.636. [DOI] [PubMed] [Google Scholar]
- 27.Grimmer T, Riemenschneider M, Förstl H, Henriksen G, Klunk WE, Mathis CA, et al. Beta amyloid in Alzheimer's disease: increased deposition in brain is reflected in reduced concentration in cerebrospinal fluid. Biol Psychiatry. 2009;65:927–934. doi: 10.1016/j.biopsych.2009.01.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hammond TR, Marsh SE, Stevens B. Immune Signaling in Neurodegeneration. Immunity. 2019;50:955–974. doi: 10.1016/j.immuni.2019.03.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Hardy JA, Higgins GA. Alzheimer's disease: the amyloid cascade hypothesis. Science (New York, NY) 1992;256:184–185. doi: 10.1126/science.1566067. [DOI] [PubMed] [Google Scholar]
- 30.He Z, Guo JL, McBride JD, Narasimhan S, Kim H, Changolkar L, et al. Amyloid-β plaques enhance Alzheimer's brain tau-seeded pathologies by facilitating neuritic plaque tau aggregation. Nat Med. 2018;24:29–38. doi: 10.1038/nm.4443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife. 2018 doi: 10.7554/eLife.34408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Husain MA, Laurent B, Plourde M. APOE and Alzheimer’s disease: from lipid transport to physiopathology and therapeutics. Front Neurosci. 2021 doi: 10.3389/fnins.2021.630502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Jansen IE, Savage JE, Watanabe K, Bryois J, Williams DM, Steinberg S, et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer's disease risk. Nat Genet. 2019;51:404–413. doi: 10.1038/s41588-018-0311-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jessen F, Spottke A, Boecker H, Brosseron F, Buerger K, Catak C, et al. Design and first baseline data of the DZNE multicenter observational study on predementia Alzheimer’s disease (DELCODE) Alzheimer's Res Ther. 2018;10:15. doi: 10.1186/s13195-017-0314-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Johansson JU, Brubaker WD, Javitz H, Bergen AW, Nishita D, Trigunaite A, et al. Peripheral complement interactions with amyloid β peptide in Alzheimer's disease: polymorphisms, structure, and function of complement receptor 1. Alzheimer's Dementia. 2018;14:1438–1449. doi: 10.1016/j.jalz.2018.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kaipainen A, Jääskeläinen O, Liu Y, Haapalinna F, Nykänen N, Vanninen R, et al. Cerebrospinal fluid and MRI biomarkers in neurodegenerative diseases: a retrospective memory clinic-based study. J Alzheimers Dis. 2020;75:751–765. doi: 10.3233/jad-200175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–443. doi: 10.1038/s41586-020-2308-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Karran E, Mercken M, De Strooper B. The amyloid cascade hypothesis for Alzheimer's disease: an appraisal for the development of therapeutics. Nat Rev Drug Discov. 2011;10:698–712. doi: 10.1038/nrd3505. [DOI] [PubMed] [Google Scholar]
- 39.Kerimov N, Hayhurst JD, Peikova K, Manning JR, Walter P, Kolberg L, et al. A compendium of uniformly processed human gene expression and splicing quantitative trait loci. Nat Genet. 2021;53:1290–1299. doi: 10.1038/s41588-021-00924-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kloske CM, Wilcock DM. The important interface between apolipoprotein E and neuroinflammation in Alzheimer’s disease. Front Immunol. 2020 doi: 10.3389/fimmu.2020.00754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kolberg L, Raudvere U, Kuzmin I, Vilo J, Peterson H. gprofiler2–an R package for gene list functional enrichment analysis and namespace conversion toolset g:Profiler. F1000Res. 2020 doi: 10.12688/f1000research.24956.2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kornhuber J, Schmidtke K, Frolich L, Perneczky R, Wolf S, Hampel H, et al. Early and differential diagnosis of dementia and mild cognitive impairment: design and cohort baseline characteristics of the German Dementia Competence Network. Dement Geriatr Cogn Disord. 2009;27:404–417. doi: 10.1159/000210388. [DOI] [PubMed] [Google Scholar]
- 43.Kunkle BW, Grenier-Boley B, Sims R, Bis JC, Damotte V, Naj AC, et al. Genetic meta-analysis of diagnosed Alzheimer's disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat Genet. 2019;51:414–430. doi: 10.1038/s41588-019-0358-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Merico D, Isserlin R, Stueker O, Emili A, Bader GD. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS ONE. 2010;5:e13984. doi: 10.1371/journal.pone.0013984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Minami SS, Min S-W, Krabbe G, Wang C, Zhou Y, Asgarov R, et al. Progranulin protects against amyloid β deposition and toxicity in Alzheimer's disease mouse models. Nat Med. 2014;20:1157–1164. doi: 10.1038/nm.3672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Miyagawa T, Ebinuma I, Morohashi Y, Hori Y, Young Chang M, Hattori H, et al. BIN1 regulates BACE1 intracellular trafficking and amyloid-β production. Hum Mol Genet. 2016;25:2948–2958. doi: 10.1093/hmg/ddw146. [DOI] [PubMed] [Google Scholar]
- 47.Mulder C, Verwey NA, van der Flier WM, Bouwman FH, Kok A, van Elk EJ. Amyloid-beta(1–42), total tau, and phosphorylated tau as cerebrospinal fluid biomarkers for the diagnosis of Alzheimer disease. Clin Chem. 2010;56:248–253. doi: 10.1373/clinchem.2009.130518. [DOI] [PubMed] [Google Scholar]
- 48.Najar J, van der Lee SJ, Joas E, Wetterberg H, Hardy J, Guerreiro R, et al. Polygenic risk scores for Alzheimer's disease are related to dementia risk in APOE ɛ4 negatives. Alzheimer's Dementia: Diagn Assess Dis Monitor. 2021;13:e12142. doi: 10.1002/dad2.12142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Olsson B, Lautner R, Andreasson U, Öhrfelt A, Portelius E, Bjerke M, et al. CSF and blood biomarkers for the diagnosis of Alzheimer's disease: a systematic review and meta-analysis. Lancet Neurol. 2016;15:673–684. doi: 10.1016/s1474-4422(16)00070-3. [DOI] [PubMed] [Google Scholar]
- 50.Ramirez A, van der Flier WM, Herold C, Ramonet D, Heilmann S, Lewczuk P, et al. SUCLG2 identified as both a determinator of CSF Aβ1-42 levels and an attenuator of cognitive decline in Alzheimer's disease. Hum Mol Genet. 2014;23:6644–6658. doi: 10.1093/hmg/ddu372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Sartori M, Mendes T, Desai S, Lasorsa A, Herledan A, Malmanche N, et al. BIN1 recovers tauopathy-induced long-term memory deficits in mice and interacts with Tau through Thr(348) phosphorylation. Acta Neuropathol. 2019;138:631–652. doi: 10.1007/s00401-019-02017-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Sherrington R, Froelich S, Sorbi S, Campion D, Chi H, Rogaeva EA, et al. Alzheimer's disease associated with mutations in presenilin 2 is rare and variably penetrant. Hum Mol Genet. 1996;5:985–988. doi: 10.1093/hmg/5.7.985. [DOI] [PubMed] [Google Scholar]
- 53.Sherrington R, Rogaev EI, Liang Y, Rogaeva EA, Levesque G, Ikeda M, et al. Cloning of a gene bearing missense mutations in early-onset familial Alzheimer's disease. Nature. 1995;375:754–760. doi: 10.1038/375754a0. [DOI] [PubMed] [Google Scholar]
- 54.Shi Y, Holtzman DM. Interplay between innate immunity and Alzheimer disease: APOE and TREM2 in the spotlight. Nat Rev Immunol. 2018;18:759–772. doi: 10.1038/s41577-018-0051-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Shi Y, Yamada K, Liddelow SA, Smith ST, Zhao L, Luo W, et al. ApoE4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature. 2017;549:523–527. doi: 10.1038/nature24016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Sims R, van der Lee SJ, Naj AC, Bellenguez C, Badarinarayan N, Jakobsdottir J, et al. Rare coding variants in PLCG2, ABI3, and TREM2 implicate microglial-mediated innate immunity in Alzheimer's disease. Nat Genet. 2017;49:1373–1384. doi: 10.1038/ng.3916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Somers C, Struyfs H, Goossens J, Niemantsverdriet E, Luyckx J, De Roeck N, et al. A decade of cerebrospinal fluid biomarkers for Alzheimer's disease in Belgium. J Alzheimers Dis. 2016;54:383–395. doi: 10.3233/jad-151097. [DOI] [PubMed] [Google Scholar]
- 58.Sunderland T, Linker G, Mirza N, Putnam KT, Friedman DL, Kimmel LH, et al. Decreased beta-amyloid1-42 and increased tau levels in cerebrospinal fluid of patients with Alzheimer disease. JAMA. 2003;289:2094–2103. doi: 10.1001/jama.289.16.2094. [DOI] [PubMed] [Google Scholar]
- 59.Taliun D, Harris D, Kessler M, Carlson J, Szpiech Z, Torres R, et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. bioRxiv. 2019 doi: 10.1101/563866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Therriault J, Benedet AL, Pascoal TA, Mathotaarachchi S, Chamoun M, Savard M, et al. Association of apolipoprotein E ε4 with medial temporal tau independent of amyloid-β. JAMA Neurol. 2020;77:470–479. doi: 10.1001/jamaneurol.2019.4421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Tijms BM, Gobom J, Reus L, Jansen I, Hong S, Dobricic V, et al. Pathophysiological subtypes of Alzheimer's disease based on cerebrospinal fluid proteomics. Brain. 2020;143:3776–3792. doi: 10.1093/brain/awaa325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Tijms BM, Willemse EAJ, Zwan MD, Mulder SD, Visser PJ, van Berckel BNM, et al. Unbiased approach to counteract upward drift in cerebrospinal fluid amyloid-β 1–42 analysis results. Clin Chem. 2018;64:576–585. doi: 10.1373/clinchem.2017.281055. [DOI] [PubMed] [Google Scholar]
- 63.Uddin MS, Kabir MT, Begum MM, Islam MS, Behl T, Ashraf GM. Exploring the role of CLU in the pathogenesis of Alzheimer's disease. Neurotox Res. 2021;39:2108–2119. doi: 10.1007/s12640-020-00271-4. [DOI] [PubMed] [Google Scholar]
- 64.van der Kant R, Goldstein LSB, Ossenkoppele R. Amyloid-β-independent regulators of tau pathology in Alzheimer disease. Nat Rev Neurosci. 2020;21:21–35. doi: 10.1038/s41583-019-0240-3. [DOI] [PubMed] [Google Scholar]
- 65.Vojinovic D, Adams HH, Jian X, Yang Q, Smith AV, Bis JC, et al. Genome-wide association study of 23,500 individuals identifies 7 loci associated with brain ventricular volume. Nat Commun. 2018;9:3945. doi: 10.1038/s41467-018-06234-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Voskobiynyk Y, Roth JR, Cochran JN, Rush T, Carullo NV, Mesina JS, et al. Alzheimer's disease risk gene BIN1 induces Tau-dependent network hyperexcitability. Elife. 2020 doi: 10.7554/eLife.57354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Wallace C. Eliciting priors and relaxing the single causal variant assumption in colocalisation analyses. PLoS Genet. 2020;16:e1008720. doi: 10.1371/journal.pgen.1008720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wang C, Najm R, Xu Q, Jeong DE, Walker D, Balestra ME, et al. Gain of toxic apolipoprotein E4 effects in human iPSC-derived neurons is ameliorated by a small-molecule structure corrector. Nat Med. 2018;24:647–657. doi: 10.1038/s41591-018-0004-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826. doi: 10.1038/s41467-017-01261-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Wightman DP, Jansen IE, Savage JE, Shadrin AA, Bahrami S, Rongve A, et al. Largest GWAS (N=1,126,563) of Alzheimer’s disease implicates microglia and immune cells. medRxiv. 2020 doi: 10.1101/2020.11.20.20235275. [DOI] [Google Scholar]
- 71.Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics (Oxford, England) 2010;26:2190–2191. doi: 10.1093/bioinformatics/btq340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Wyss-Coray T, Yan F, Lin AH, Lambris JD, Alexander JJ, Quigg RJ, et al. Prominent neurodegeneration and increased plaque formation in complement-inhibited Alzheimer's mice. Proc Natl Acad Sci USA. 2002;99:10837–10842. doi: 10.1073/pnas.162350199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Yang C, Farias FHG, Ibanez L, Suhy A, Sadler B, Fernandez MV, et al. Genomic atlas of the proteome from brain, CSF and plasma prioritizes proteins implicated in neurological disorders. Nat Neurosci. 2021;24:1302–1312. doi: 10.1038/s41593-021-00886-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet. 2011;88:76–82. doi: 10.1016/j.ajhg.2010.11.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Yeh FL, Wang Y, Tom I, Gonzalez LC, Sheng M. TREM2 binds to apolipoproteins, including APOE and CLU/APOJ, and thereby facilitates uptake of amyloid-beta by microglia. Neuron. 2016;91:328–340. doi: 10.1016/j.neuron.2016.06.015. [DOI] [PubMed] [Google Scholar]
- 76.Yiannopoulou KG, Anastasiou AI, Zachariou V, Pelidou SH. Reasons for failed trials of disease-modifying treatments for Alzheimer disease and their contribution in recent research. Biomedicines. 2019 doi: 10.3390/biomedicines7040097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Young AMH, Kumasaka N, Calvert F, Hammond TR, Knights A, Panousis N, et al. A map of transcriptional heterogeneity and regulatory variation in human microglia. Nat Genet. 2021;53:861–868. doi: 10.1038/s41588-021-00875-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Zettergren A, Höglund K, Kern S, Thorvaldsson V, Johan Skoog M, Hansson O, et al. Association of IL1RAP-related genetic variation with cerebrospinal fluid concentration of Alzheimer-associated tau protein. Sci Rep. 2019;9:2460. doi: 10.1038/s41598-018-36650-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Zhao Z, Sagare AP, Ma Q, Halliday MR, Kong P, Kisler K, et al. Central role for PICALM in amyloid-β blood-brain barrier transcytosis and clearance. Nat Neurosci. 2015;18:978–987. doi: 10.1038/nn.4025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Zheng J, Erzurumluoglu AM, Elsworth BL, Kemp JP, Howe L, Haycock PC, et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics (Oxford, England) 2017;33:272–279. doi: 10.1093/bioinformatics/btw613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Zhu X-C, Yu J-T, Jiang T, Wang P, Cao L, Tan L. CR1 in Alzheimer's disease. Mol Neurobiol. 2015;51:753–765. doi: 10.1007/s12035-014-8723-8. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Genome-wide summary statistics have been deposited to the European Bioinformatics Institute GWAS Catalog (https://www.ebi.ac.uk/gwas/) under accession IDs GCST90129599 and GCST90129600.