Abstract
Chromosome 17q12-q21 is the most replicated genetic locus for childhood-onset asthma. Polymorphisms in this locus containing ∼10 genes interact with a variety of environmental exposures in the home and outdoors to modify asthma risk. However, the functional basis for these associations and their linkages to the environment have remained enigmatic. Within this extended region, regulation of GSDMB (gasdermin B) expression in airway epithelial cells has emerged as the primary mechanism underlying the 17q12-q21 genome-wide association study signal. Asthma-associated SNPs influence the abundance of GSDMB transcripts as well as the functional properties of GSDMB protein in airway epithelial cells. GSDMB is a member of the gasdermin family of proteins, which regulate pyroptosis and inflammatory responses to microbial infections. The aims of this review are to synthesize recent studies on the relationship of 17q12-q21 SNPs to childhood asthma and the evidence pointing to GSDMB gene expression or protein function as the underlying mechanism and to explore the potential functions of GSDMB that may influence the risk of developing asthma during childhood.
Keywords: asthma, 17q12-q21, gene-by-environment interaction, GSDMB, pyroptosis
Clinical Relevance
Although the chromosomal 17q12-q21 locus has been robustly associated with asthma, particularly childhood-onset asthma, which gene(s) at this locus and what cellular mechanisms contribute to this risk have not been definitively determined. This Perspective article highlights recent discoveries at this locus, gene-by-environment interactions, and the emerging evidence supporting the gasdermin B gene and cellular process of pyroptosis as mediators of asthma risk.
Genome-wide association studies (GWASs) have identified more than 150 risk loci for asthma and led to novel insights into the pathogenesis of asthma in children and adults (1–5). These discoveries have highlighted the importance of both immune cells and the airway epithelium in childhood-onset asthma, as well as differences between childhood-onset and adult-onset asthma with respect to the number of associated risk loci and tissue enrichments of the genes expressed at these loci (1). Since its discovery in 2007 (6), a region on chromosome 17q12-q21 has remained the most significant and most replicated locus with childhood-onset asthma. Like GWASs more broadly, most genetic studies of 17q12-q21 and asthma have been conducted in populations with predominantly European ancestry. Although the association between 17q12-q21 SNPs and asthma has been robustly observed in studies of different ethnic groups, including African American (2, 3, 7), Asian (8, 9), American Indian (10), and multiethnic (2) cohorts, most of these have been underpowered compared with studies in individuals of European ancestry. However, the extensive linkage disequilibrium at this locus in European-ancestry populations has made it challenging to identify the variant(s) and gene(s) in this region that confer risk for childhood-onset asthma.
Using complementary strategies, three recent studies have converged on the conclusion that risk for childhood-onset asthma at the 17q12-q21 locus is due to genetic variation that alters degrees of expression of GSDMB (gasdermin B) or affects GSDMB protein function in airway epithelial cells. Using the significant breakdown of linkage disequilibrium in African-ancestry populations, we integrated genetic association and gene expression studies in African Americans to fine map this locus (7). Our results suggested that the risk for childhood-onset asthma conferred by variation at the 17q12-q21 locus was attributable to SNPs regulating GSDMB expression or GSDMB function in airway epithelial cells.
The missense SNP highlighted in our study, rs2305480 (c.892G→A, p.Pro298Ser), is in near perfect linkage disequilibrium with an intronic SNP, rs11078928 (c.661 + 742T→C), that alters splicing of the GSDMB transcript and protein abundance in cells (11, 12). In all populations studied, the p.298Pro allele and the rs11079829-T allele are associated with childhood-onset asthma. The localization of this association to SNPs in the GSDMB gene was then confirmed by Gui and colleagues (13) using whole-genome sequencing in African Americans that further suggested that the causal SNP at this locus is the splice variant rs11078928. These and previous studies indicate that the rs11078928-T and rs2305480-G asthma-risk alleles are associated with increased expression of the full-length transcript (11, 13, 14) and with changes in protein conformation and surface charge (15), respectively.
Li and colleagues (16) recently demonstrated that GSDMB expression in airway epithelial cells was also correlated with a greater number of exacerbations, reduced asthma control, and more emergency room visits, further implicating GSDMB in asthma severity in adults. Despite this convergence on GSDMB as the “risk gene” at the 17q12-q21 locus, the mechanisms through which GSDMB affects asthma pathogenesis are incompletely understood.
We performed a comprehensive literature review of the 17q12-q21 locus and asthma in general and of GSDMB in particular. We reviewed clinical and environmental correlates with 17q12-q21 and childhood asthma, the expression patterns of GSDMB in different cell types and its regulation at the molecular level, and the functional properties of GSDMB and related proteins at the cellular level. This paper provides an update to our previous review of the 17q12-q21 locus (17) and focuses on functional studies linking genetic regulation of GSDMB to childhood asthma development.
Clinical and Environmental Correlates at the 17q12-q21 Locus and Childhood Asthma
Associations at the 17q12-q21 locus with asthma are driven nearly exclusively by individuals with asthma diagnosed in childhood (1, 4, 6, 17). Despite a shared genetic architecture between asthma and allergic phenotypes (4, 18), SNPs at the 17q12-q21 locus are not associated with allergic sensitization (19), total IgE concentrations (20) or atopic dermatitis (21). However, some studies have shown associations with specific allergic diseases, such as hay fever and allergic rhinitis (22–24). The latter potentially reflects a role for this locus in epithelial barrier function, which is a shared feature of childhood-onset asthma, atopic dermatitis, and allergic rhinitis (1, 18).
Associations of SNPs at this locus have been strongest among children with early-life respiratory illnesses (25, 26), exacerbations and hospitalizations (27), exposure to environmental tobacco smoke (28–30), wheezing-associated illnesses (29), and wheezing phenotypes (31). Moreover, genotypes at this locus moderate the effects of many environmental risk and protective factors for childhood-onset asthma, such as prenatal and postnatal tobacco smoke exposure (28–30, 32), rhinovirus-associated wheezing illness (25) and respiratory illnesses in general (26), older siblings (33), farm animals (33), house pets (30, 34, 35), allergens in the home (26), mold exposure (36), breastfeeding (37), and prenatal vitamin D supplementation (38). These relationships are listed in Table 1 and illustrated in an online interactive diagram (https://crew.biostat.wisc.edu). It is notable that nearly all the epidemiological risk factors involved in interactions involve inhaled irritants or microbial exposures, which can act directly or indirectly on the epithelium, further supporting a role for airway epithelial cell barrier function as a primary mechanism for 17q12-q21–associated risks.
Table 1.
Gene-Environment Interactions at the 17q12-21 Locus
| SNP(s)* | Effect Allele or Genotype | Nearest Gene | Interaction Effect (95% CI) (Ratio) | Asthma Outcome | Study Population (Number of Asthma Cases/Control Subjects) | Reference | |
|---|---|---|---|---|---|---|---|
| Interactions associated with increased risk | |||||||
| Environmental tobacco smoke exposure in early life | rs2305480 | G | GSDMB | 1.98 | Early-onset asthma | European family cohort (651/776) | 29 |
| Maternal smoking during pregnancy and tobacco exposure at age 2 yr | rs2305480 | G | GSDMB | NR | Asthma-like symptoms during first 4 yr of life | Two European birth cohorts (315/2,350 and 572/2,350) | 28 |
| Current smoking status† | rs12936231 | C | ZPBP2 | NR | Current asthma | European birth cohorts (312/983) | 32 |
| rs2290400 | C | GSDMB | |||||
| rs8079416 | C | ORMDL3 | |||||
| Rhinovirus wheezing illness in the first 3 yr | rs7216389 | TT | GSDMB | 26.1 (5.1–133.0) (OR) | Asthma at age 6 yr | European and European American birth cohorts (122/375) | 25 |
| Early life exposure to tobacco | rs4795405 | TT | ORMDL3 | NR | Asthma symptoms in previous 12 mo | Cross-sectional cohort (423/414) | 30 |
| Number of colds in the first year of life | rs8069202 | A | GSDMA | 3.71 (1.67–8.25) (OR) | Asthma at age 7 yr | Black children in U.S. birth cohort (262) | 26 |
| rs3859192 | T | GSDMA | 4.36 (1.87–10.17) | ||||
| Visible mold exposure | rs7216389 | T | GSDMA | 1.32 (1.11–1.57) (aOR) | Childhood asthma | Cross-sectional cohort (608/839) | 36 |
| Older siblings | rs8076131 | GA/AA | ORMDL3 | 1.53 (1.16–2.01) (OR) | Wheeze in the first year of life | European birth cohort | 33 |
| rs2290400 | TC/TT | GSDMB | 1.38 (1.04–1.83) (OR) | ||||
| rs7216389 | CT/TT | GSDMB | 1.41 (1.06–1.87) (OR) | ||||
| Interactions associated with decreased risk | |||||||
| Exposure to furred pets by 18 mo of age | rs7216389 | TT | GSDMB | 0.76 (0.50–1.15)‡ | Four or more wheezing episodes by 18 mo | European birth cohort (1,111/734) | 34 |
| Current cat and/or dog ownership | rs921651 | AA | GSDMA | NR | Asthma symptoms in previous 12 mo | Cross-sectional cohort (423/414) | 30 |
| Exposure to animal sheds in the first year | rs8076131 | GA/AA | ORMDL3 | 0.44 (0.33–0.60) (OR) | Wheeze in the first year of life | European birth cohort§ | 33 |
| rs2290400 | TC/TT | GSDMB | 0.42 (0.31–0.58) (OR) | ||||
| rs7216389 | CT/TT | GSDMB | 0.44 (0.32–0.60) (OR) | ||||
| Cat and/or dog living in the home at birth | rs7216389 | TT | GSDMB | 0.16 (HR) | Asthma at 12 yr of age | European birth cohort (85/292) | 35 |
| Breastfeeding | rs7216389 | T | GSDMB | 0.82 (RR) | Respiratory symptoms during the weeks of breastfeeding | European birth cohorts (252/116)‖ | 37 |
| rs4795405 | C | ORMDL3 | 0.85 (RR) | ||||
| Vitamin D during pregnancy | rs12936231 | GG | ZPBP2 | 0.49 (0.26–0.94) (HR) | Asthma/recurrent wheezing between 0 and 3 yr of age | European and European American prenatal cohorts (260/875) | 38 |
| Indoor allergen concentration | rs7216389 | TT | GSDMB | 0.46 (0.02–0.87) (OR) | Asthma at age 7 yr | Black children in U.S. birth cohort (262) | 26 |
Definition of abbreviations: aOR = adjusted odds ratio; CI = confidence interval; GASDMB = gasdermin B; HR = hazard ratio; NR = not reported; OR = odds ratio; ORMDL3 = ORM1-like protein 3; RR = relative risk; ZPBP2 = zona pellucida binding protein 2.
SNPs are those reported in the studies to have significant interactions.
Initial analysis of “smoking pack-years” was not significant; “smoking status” was significant.
Reported P value was for the difference in the effect of the genotype (CC).
Numbers of cases and control subjects were not reported.
Number of subjects in the discovery cohort.
GWASs of autoimmune diseases (inflammatory bowel disease, rheumatoid arthritis, and type 1 diabetes) have also identified GSDMB SNPs as mediators of risk, although the direction of effect is opposite; for example, the rs2305480-G (c.298Pro) in asthma-risk allele is associated with protection against inflammatory bowel disease and type 1 diabetes, for which rs2305480-T (c.298Ser) is associated with risk (39). The latter is consistent with previous reports of opposite-effect alleles in asthma and autoimmune disease (40, 41), suggesting that GSDMB is a crucial regulator of epithelial function in the airways and other tissues and may be an essential clue to understanding shared pathogenic mechanisms for these diseases.
Expression Patterns of GSDMB in Different Cell Types and Its Regulation at the Molecular Level
A recent transcriptome-wide association study in lung and blood identified GSDMB as the gene most significantly associated with asthma (42). GSDMB is broadly expressed across tissues and cell types in the Genotype-Tissue Expression (GTEx) database (see Figure E1 in the data supplement) (43), with intermediate degrees of GSDMB expression in lung, skin, spleen, and whole blood, tissues that are also enriched for expression of genes at asthma GWAS loci (1). These tissues include mixed cell types, which likely have variable degrees of GSDMB expression. Single-cell RNA sequencing in lung tissue, including that from GTEx, indicate that GSDMB is expressed in multiple cell types, including immune, epithelial, and endothelial cell subsets, but only a small fraction of cells within each of these cell types express GSDMB (see Figure E2). Therefore, additional studies of GSDMB expression in specific lung cells (e.g., epithelial cell subsets and airway smooth muscle) and immune cells (e.g., CD4 tissue-resident T cells and lung-resident macrophages) are needed, particularly in the context of environmental exposures, such as rhinovirus infection, airway microbes, and tobacco smoke.
Effects of asthma-associated genetic variants on GSDMB also vary among tissues. For example, the effect of the asthma-risk rs11078928-T allele on GSDMB gene expression is largest in whole blood, followed by spleen, small intestine, and lung, among all tissues in the GTEx database (43). The asthma-risk allele is associated with increased GSDMB expression in all these tissues (see Figure E2). Notably, however, this SNP is not an expression quantitative trait locus for GSDMB in other tissues (e.g., heart, esophagus, brain) and has opposite effects on GSDMB expression in yet others (e.g., breast, adipose, liver), highlighting the tissue specificity of gene regulation at this locus (see Figure E2).
The splicing precision of GSDMB transcripts and the abundance of functional GSDMB protein and asthma risk have been linked to rs11078928 (11, 13, 44). There are many alternatively spliced forms of GSDMB transcripts; the two most common transcripts are shown in Figure 1. The rs11078928-C nonrisk allele leads to skipping of exon 6 and reduced expression of the full-length transcript and of GSDMB protein (11, 13, 44), suggesting that overexpression of GSDMB protein in carriers of the rs11078928-T allele mediates the asthma risk associated with this locus.
Figure 1.

GSDMB exon expression in lung from Genotype-Tissue Expression Analysis Release V8 (43). Full-length GSDMB transcript (top) and the alternatively spliced variant with approximate positions of rs2305480 and rs11078928 are depicted. Boxes indicate exons, and lines indicate introns. GSDMB = gasdermin B.
Functional Properties of GSDMB and Related Proteins at the Cellular Level
The gasdermin family of proteins includes GSDMA (gasdermin A), GSDMB, GSDMC (gasdermin C), GSDMD (gasdermin D), GSDME (gasdermin E), and PJVK (pejvakin); these proteins share 23.9–49.4% sequence similarity (45). GSDMB (416 amino acids, 47 kD) is the most divergent member of the gasdermin family and is found exclusively in primates and ruminants (46). Herein, we discuss the known mechanisms by which GSDMB affects airway cell function, including pyroptotic cell death, airway remodeling, and cellular lipid transport. These GSDMB functions are also depicted in an interactive online diagram (https://crew.biostat.wisc.edu).
Pyroptosis
Gasdermins mediate pyroptosis, a form of programmed cell death initiated by either activation of caspase-1 (canonical pathway) or caspase-4/5 (caspase-11 in mice; noncanonical pathway) (47). Both pathways lead to similar cellular damage characterized by gasdermin protein–mediated pore formation in the cell membrane, K+ efflux/Na+ influx, cell swelling, and eventual rupture (Figure 2). Gasdermins consist of a repressor C terminus and a cytotoxic N terminus joined by a flexible linker domain (Figure 2). Cleavage of the linker domain releases the N terminus from autoinhibition. Once released, the N terminus forms multimers that bind to the cell membrane, subsequently forming pores. These pores in the membrane are large enough to allow leakage of cytosolic contents, including IL-1β, IL-18, and damage-associated molecular patterns, but not bacteria and organelles. The damaged cells, known as pore-induced intracellular traps, can be efferocytosed by neutrophils in a second wave of killing (48).
Figure 2.
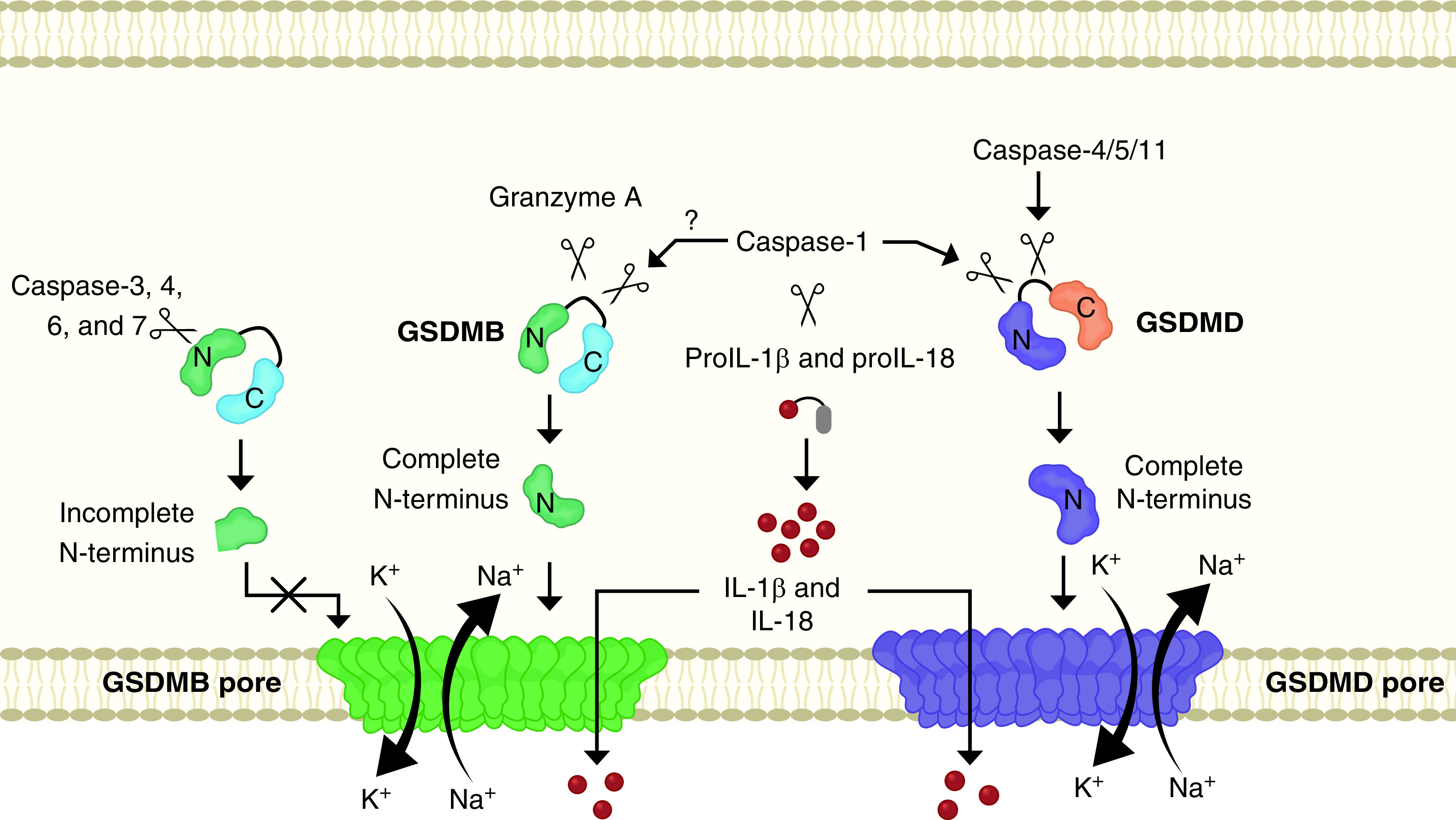
Mechanisms of GSDMB-induced pyroptosis. In macrophages, caspase-1, 4, 5, and 11 cleave the linker domain of GSDMD and release the N terminus, which then oligomerizes to form pores in the cell membrane. This results in leakage of proinflammatory cytokines IL-1β and IL-18. The pore also allows Na+ efflux and K+ influx, leading to cell swelling and rupture. In epithelial cells, the linker domain of GSDMB is cleaved by granzyme A (from natural killer cells or cytotoxic T lymphocytes) or possibly caspase-1, leading to similar consequences. Conversely, cleavage of GSDMB by caspase-3, 4, 6, or 7 releases an incomplete N terminus fragment that cannot oligomerize to form a pore in the cell membrane. GSDMD = gasdermin D.
GSDMD has been well studied and is the dominant executor of pyroptosis in macrophages (47). In response to inflammasome assembly, procaspase-1 is cleaved to release the active caspase-1 that cleaves GSDMD and pro–IL-1β and pro–IL-18 into their mature forms. In contrast, cytoplasmic LPS directly activates caspase-4/5/11 through the N terminus CARD (caspase activation and recruitment domain), leading to oligomerization of the N terminus cleavage products (49).
There are conflicting data as to whether caspase-1 can cleave GSDMB to initiate pyroptosis. Panganiban and colleagues (44) reported that GSDMB in airway epithelial cells could be specifically cleaved by caspase-1 at the D236 position to release the complete cytotoxic N terminus (Figure 3). They further demonstrated that the asthma nonrisk splice variant rs11078928-C abolishes pyroptotic activity by preventing the splicing of GSDMB exon 6 within the N terminus, while the missense variant rs2305480 (p.Pro298Ser) did not affect pyroptosis. In contrast, Chen and colleagues (50) reported that caspase-1 could cleave GSDMD but not GSDMB. In their model, caspases-1, 3, 4, 6, and 7 cleaved GSDMB to form an incomplete N terminus (1–91 amino acids) that could not oligomerize and execute pyroptosis. Instead, GSDMB indirectly promoted pyroptosis by binding to the CARD of caspase-4 and increasing its ability to cleave GSDMD (Figure 3) (50). Regardless of the precise mechanisms involved, both models implicate GSDMB in pyroptosis.
Figure 3.
GSDMB protein cleavage sites. Granzyme A cleaves the C-terminal domain at position 244, releasing a complete N terminus that can form a membrane pore. Caspase-1 may also cleave the linker domain to release the complete N terminus that can form the pore. In contrast, caspase-3, 6, 7, and 4 cleave GSDMB at position 91 on the N terminus, and this results in a fragment that cannot form a pore. aa = amino acids.
More recently, Zhou and colleagues (51) demonstrated that granzyme A, derived from cytotoxic lymphocytes, can directly cleave GSDMB at Lys244 to release the N terminus. They further showed that IFN-γ increased the expression of GSDMB, consistent with the findings of Li and colleagues (16) that implicated IFN gene pathways in GSDMB-associated asthma exacerbations. These combined findings suggest that pathogens that induce IFN and stimulate natural killer cells or CD8+ T cells to secrete granzyme A could initiate GSDMB-mediated pyroptosis in infected airway epithelial cells. Viral and bacterial infections could damage the respiratory epithelium through this mechanism, suggesting that SNPs that increase GSDMB expression or functional activity may increase epithelial damage during infections.
Hansen and colleagues (52) confirmed the role of granzyme A in cleaving GSDMB and also demonstrated that the cleaved N terminus binds to intracellular gram-negative bacterial membrane proteins, particularly cardiolipin, to cause pore-mediated lytic death of bacteria. As the GSDMB N terminus did not bind to phospholipids commonly found in the mammalian plasma membrane, the authors speculated that GSDMB-mediated lytic death of host cells could be triggered by a disarrangement of membrane lipids between inner mitochondrial and plasma membranes during infection. The latter finding also suggests that overexpression of GSDMB could further damage infected epithelium. Interestingly, the IpaH7.8 protein of Shigella flexneri (an enteroinvasive gram-negative pathogen) counters GSDMB by ubiquitinating the cleaved N terminus and marking it for proteasomal degradation. It will be interesting to investigate whether respiratory pathogens associated with asthma exacerbations use similar evasion mechanisms that aid survival and growth, resulting in increased epithelial damage during infections.
Airway Hyperresponsiveness and Remodeling
Using immunohistochemistry, Das and colleagues (53) showed that the GSDMB protein is expressed specifically on human bronchial epithelial cells and not on other lung cell populations. The numbers of GSDMB+ bronchial epithelial cells were higher in asthmatics (n = 14) compared with control subjects (n = 7) and were highest in subjects with severe asthma (n = 7). However, in contrast to the results of genetic studies in which the allele associated with the full-length transcript is associated with asthma, Das and colleagues reported nearly exclusive expression of the truncated transcript in these cells (transcript ENST00000394179.5 in Figure 1). Following up on this finding, they further showed that in vitro overexpression of the truncated transcript upregulated airway remodeling genes, such as TGF-β1 (transforming growth factor-β 1) and 5-lipoxygenase, in bronchial epithelial cells. Although GSDMB is not natively expressed in the mouse, they used a transgenic mouse model with the human GSDMB gene. Mice expressing the transgene demonstrated the expected increased GSDMB expression, which was associated with airway hyperresponsiveness and remodeling in the absence of inflammation. These studies suggest that GSDMB can promote airway hyperresponsiveness and remodeling even in the absence of allergen exposure in mice, although further studies are needed to determine whether similar findings are observed in humans.
Cellular Lipid Transport
Full-length GSDMB can bind lipids, including sulfatide, a 3-O-sulfogalactosylceramide and a component of the apical membrane of epithelial cells with roles in immune regulation and the antimicrobial response. GSDMB is unique in binding lipids, as the C termini of other GSDM proteins inhibit this function, allowing only free N termini to bind lipids (15). Chao and colleagues (15) demonstrated that the GSDMB protein polymorphisms Arg299:Ser306, corresponding to SNPs rs2305479 (Gly299Arg) and rs2305480 (Pro306Ser; referred to above as Pro298Ser), showed changes in conformation and surface charge distributions. Although the cellular transporters of sulfatide are unknown, the authors speculated that if GSDMB is involved in this process, overexpression of GSDMB could lead to higher sulfatide concentrations on the apical surface of airway epithelial cells. In this context, it is worth noting that several clinically significant bacterial strains, including Moraxella catarrhalis and Haemophilus influenzae, bind to mucosal surfaces through sulfatide (54, 55) and can be increased in airway secretions during exacerbations of asthma in children (56). GSDMB could therefore indirectly affect bacterial adhesion by altering the expression of binding proteins on the epithelium’s apical surface. Further studies are needed to confirm this role for GSDMB in sulfatide transport and its impact on epithelial barrier integrity and microbial interactions, particularly during respiratory tract infections.
Conclusions and Future Directions
The etiology of childhood asthma has long been associated with type 2 inflammation, and it was therefore not surprising that variation at loci including the IL13, TSLP (thymic stromal lymphopoietin), IL33, and IL1R1 (IL-1 receptor type 1) (encoding the IL-33 receptor) genes were significantly associated with asthma in GWAS. In contrast, connections between genes at the 17q12-q21 locus and childhood-onset asthma were not obvious but promised to bring to light novel mechanistic insights into the pathogenesis of the most common chronic disease of childhood. Recent studies suggest that SNPs regulating GSDMB expression and protein function in airway epithelium may mediate the association between 17q12-q21 and childhood asthma. Together with experimental information linking GSDMB to microbial responses and pyroptosis, these findings provide plausible mechanisms linking respiratory pathogens and irritants to epithelial cell survival and function, airway inflammation, and asthma. These collective findings represent a significant step toward understanding the most important GWAS locus for childhood-onset asthma.
Several questions related to the context specificity of this locus remain, including those on mechanisms of gene–environment interactions. First, with respect to molecular mechanisms, what nongenetic factors regulate the expression of GSDMB in airway cells? Defining the molecular biology of this molecule and the GSDMB interactome should lead to a greater understanding of how the airway epithelium is involved in childhood-onset asthma and how GSDMB-mediated pathways affect the long-term health of the airway epithelium.
Second, with respect to gene–environment interactions, which variants and which genes at this locus interact with environmental factors, and what causal pathways underlie these interactions? Understanding these interactions is essential for developing personalized interventions that modify exposures to environmental factors in early life to reduce asthma risk. Although we present evidence that the main effect of this locus on asthma risk is due to perturbations of GSDMB expression in airway epithelial cells, we acknowledge the complexity of this locus and the possibility that other genes, such as ORMDL3 (ORM1-like protein 3), GSDMA, or PGAP3 (post-GPI attachment to proteins phospholipase 3), may be involved in context-specific risks, such as those associated with rhinovirus-associated wheezing illnesses and exposures to large farm animals or house pets in early life. Elucidating these additional intricacies should be a priority area of research over the next few years.
A final set of questions relate to the natural history of asthma. Although it is clear that the 17q12-q21 locus is associated exclusively with asthma that starts in childhood, is this endotype of asthma transient or persistent? The recent study of Li and colleagues (16) suggests that GSDMB expression is associated with the risk of exacerbation and disease severity in adults, and that of Hallmark and colleagues (31) suggests that variation at this locus is associated with the risk of transient, late-onset, and persistent wheeze in young children. These combined data suggest that the effects of this locus on wheezing illness in early life (31) and exacerbations in childhood (27) persist into adulthood (16). Further studies are required to determine whether this locus affects exacerbation risk into adulthood only among individuals with childhood-onset asthma, or it is an independent risk factor for exacerbations in adults regardless of the age of asthma onset. Related to this is the important question as to whether 17q12-q21–associated childhood-onset asthma persists into adulthood.
Footnotes
Supported by National Institutes of Health (NIH) grant K08 HL153955 (N.S.) and NIH grants T15 LM007359 and UG3/UH3 OD023282 (J.S.Y.) and the Children’s Respiratory and Environmental Workgroup consortium, NIH Office of the Director (award UG3 OD023282). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author Contributions: N.S., E.D., M.W.C., E.M.S., R.F.L., C.O., and J.E.G. made contributions to the conception or design of the study. N.S. and E.D. made the figures and table. M.W.C., J.S.Y., and J.E. designed the interactive online diagram with input from all authors. N.S. and E.D. wrote the initial draft of the manuscript. N.S., E.D., E.S., R.F.L., C.O., and J.E.G. read and edited early drafts of the manuscript. All authors reviewed and/or critically revised the manuscript for important intellectual content and provided final approval.
This article has a data supplement, which is accessible from this issue’s table of contents at www.atsjournals.org.
Originally Published in Press as DOI: 10.1165/rcmb.2022-0043PS on May 17, 2022
Author disclosures are available with the text of this article at www.atsjournals.org.
References
- 1. Pividori M, Schoettler N, Nicolae DL, Ober C, Im HK. Shared and distinct genetic risk factors for childhood-onset and adult-onset asthma: genome-wide and transcriptome-wide studies. Lancet Respir Med . 2019;7:509–522. doi: 10.1016/S2213-2600(19)30055-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Daya M, Rafaels N, Brunetti TM, Chavan S, Levin AM, Shetty A, et al. CAAPA Association study in African-admixed populations across the Americas recapitulates asthma risk loci in non-African populations. Nat Commun . 2019;10:880. doi: 10.1038/s41467-019-08469-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Demenais F, Margaritte-Jeannin P, Barnes KC, Cookson WOC, Altmüller J, Ang W, et al. Australian Asthma Genetics Consortium (AAGC) Collaborators Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nat Genet . 2018;50:42–53. doi: 10.1038/s41588-017-0014-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Ferreira MAR, Mathur R, Vonk JM, Szwajda A, Brumpton B, Granell R, et al. 23andMe Research Team; eQTLGen Consortium; BIOS Consortium Genetic architectures of childhood- and adult-onset asthma are partly distinct. Am J Hum Genet . 2019;104:665–684. doi: 10.1016/j.ajhg.2019.02.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Olafsdottir TA, Theodors F, Bjarnadottir K, Bjornsdottir US, Agustsdottir AB, Stefansson OA, et al. Eighty-eight variants highlight the role of T cell regulation and airway remodeling in asthma pathogenesis. Nat Commun . 2020;11:393. doi: 10.1038/s41467-019-14144-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Moffatt MF, Kabesch M, Liang L, Dixon AL, Strachan D, Heath S, et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature . 2007;448:470–473. doi: 10.1038/nature06014. [DOI] [PubMed] [Google Scholar]
- 7. Ober C, McKennan CG, Magnaye KM, Altman MC, Washington C, III, Stanhope C, et al. Environmental Influences on Child Health Outcomes-Children’s Respiratory Research Workgroup Expression quantitative trait locus fine mapping of the 17q12-21 asthma locus in African American children: a genetic association and gene expression study. Lancet Respir Med . 2020;8:482–492. doi: 10.1016/S2213-2600(20)30011-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Anantharaman R, Andiappan AK, Nilkanth PP, Suri BK, Wang Y, Chew FT. Genome-wide association study identifies PERLD1 as asthma candidate gene. BMC Med Genet . 2011;12:170. doi: 10.1186/1471-2350-12-170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Kim KW, Kim DY, Yoon D, Kim KK, Jang H, Schoettler N, et al. Genome-wide association study identifies TNFSF15 associated with childhood asthma. Allergy . 2022;77:218–229. doi: 10.1111/all.14952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Best LG, Azure C, Segarra A, Enright KJ, Hamley S, Jerome D, et al. Genetic variants and risk of asthma in an American Indian population. Ann Allergy Asthma Immunol . 2017;119:31–36.e1. doi: 10.1016/j.anai.2017.05.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Morrison FS, Locke JM, Wood AR, Tuke M, Pasko D, Murray A, et al. The splice site variant rs11078928 may be associated with a genotype-dependent alteration in expression of GSDMB transcripts. BMC Genomics . 2013;14:627. doi: 10.1186/1471-2164-14-627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Cardamone G, Paraboschi EM, Rimoldi V, Duga S, Soldà G, Asselta R. The characterization of GSDMB splicing and backsplicing profiles identifies novel isoforms and a circular RNA that are dysregulated in multiple sclerosis. Int J Mol Sci . 2017;18:576. doi: 10.3390/ijms18030576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Gui H, Levin AM, Hu D, Sleiman P, Xiao S, Mak ACY, et al. Mapping the 17q12-21.1 locus for variants associated with early-onset asthma in African Americans. Am J Respir Crit Care Med . 2021;203:424–436. doi: 10.1164/rccm.202006-2623OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Pagnamenta AT, Murakami Y, Taylor JM, Anzilotti C, Howard MF, Miller V, et al. DDD Study Analysis of exome data for 4293 trios suggests GPI-anchor biogenesis defects are a rare cause of developmental disorders. Eur J Hum Genet . 2017;25:669–679. doi: 10.1038/ejhg.2017.32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Chao KL, Kulakova L, Herzberg O. Gene polymorphism linked to increased asthma and IBD risk alters gasdermin-B structure, a sulfatide and phosphoinositide binding protein. Proc Natl Acad Sci U S A . 2017;114:E1128–E1137. doi: 10.1073/pnas.1616783114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Li X, Christenson SA, Modena B, Li H, Busse WW, Castro M, et al. NHLBI Severe Asthma Research Program (SARP) Genetic analyses identify GSDMB associated with asthma severity, exacerbations, and antiviral pathways. J Allergy Clin Immunol . 2021;147:894–909. doi: 10.1016/j.jaci.2020.07.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Stein MM, Thompson EE, Schoettler N, Helling BA, Magnaye KM, Stanhope C, et al. A decade of research on the 17q12-21 asthma locus: piecing together the puzzle. J Allergy Clin Immunol . 2018;142:749–764.e3. doi: 10.1016/j.jaci.2017.12.974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Zhu Z, Lee PH, Chaffin MD, Chung W, Loh PR, Lu Q, et al. A genome-wide cross-trait analysis from UK Biobank highlights the shared genetic architecture of asthma and allergic diseases. Nat Genet . 2018;50:857–864. doi: 10.1038/s41588-018-0121-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Bønnelykke K, Matheson MC, Pers TH, Granell R, Strachan DP, Alves AC, et al. AAGC Meta-analysis of genome-wide association studies identifies ten loci influencing allergic sensitization. Nat Genet . 2013;45:902–906. doi: 10.1038/ng.2694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Levin AM, Mathias RA, Huang L, Roth LA, Daley D, Myers RA, et al. A meta-analysis of genome-wide association studies for serum total IgE in diverse study populations. J Allergy Clin Immunol . 2013;131:1176–1184. doi: 10.1016/j.jaci.2012.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Sliz E, Huilaja L, Pasanen A, Laisk T, Reimann E, Magi R, et al. Uniting biobank resources reveals novel genetic pathways modulating susceptibility for atopic dermatitis. J Allergy Clin Immunol . 2022;149:1105–1112.e9. doi: 10.1016/j.jaci.2021.07.043. [DOI] [PubMed] [Google Scholar]
- 22. Karunas AS, Fedorova YY, Gimalova GF, Etkina EI, Khusnutdinova EK. Association of gasdermin B gene GSDMB polymorphisms with risk of allergic diseases. Biochem Genet . 2021;59:1527–1543. doi: 10.1007/s10528-021-10073-8. [DOI] [PubMed] [Google Scholar]
- 23. Fuertes E, Söderhäll C, Acevedo N, Becker A, Brauer M, Chan-Yeung M, et al. Associations between the 17q21 region and allergic rhinitis in 5 birth cohorts. J Allergy Clin Immunol . 2015;135:573–576. doi: 10.1016/j.jaci.2014.08.016. [DOI] [PubMed] [Google Scholar]
- 24. Tomita K, Sakashita M, Hirota T, Tanaka S, Masuyama K, Yamada T, et al. Variants in the 17q21 asthma susceptibility locus are associated with allergic rhinitis in the Japanese population. Allergy . 2013;68:92–100. doi: 10.1111/all.12066. [DOI] [PubMed] [Google Scholar]
- 25. Calışkan M, Bochkov YA, Kreiner-Møller E, Bønnelykke K, Stein MM, Du G, et al. Rhinovirus wheezing illness and genetic risk of childhood-onset asthma. N Engl J Med . 2013;368:1398–1407. doi: 10.1056/NEJMoa1211592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Gereige JD, Morin A, Calatroni A, Visness CM, Wood RA, Kattan M, et al. 17q12-q21 variants interact with early-life exposures to modify asthma risk in Black children. Clin Exp Allergy . 2022;52:565–568. doi: 10.1111/cea.14074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Bønnelykke K, Sleiman P, Nielsen K, Kreiner-Møller E, Mercader JM, Belgrave D, et al. A genome-wide association study identifies CDHR3 as a susceptibility locus for early childhood asthma with severe exacerbations. Nat Genet . 2014;46:51–55. doi: 10.1038/ng.2830. [DOI] [PubMed] [Google Scholar]
- 28. van der Valk RJ, Duijts L, Kerkhof M, Willemsen SP, Hofman A, Moll HA, et al. Interaction of a 17q12 variant with both fetal and infant smoke exposure in the development of childhood asthma-like symptoms. Allergy . 2012;67:767–774. doi: 10.1111/j.1398-9995.2012.02819.x. [DOI] [PubMed] [Google Scholar]
- 29. Bouzigon E, Corda E, Aschard H, Dizier MH, Boland A, Bousquet J, et al. Effect of 17q21 variants and smoking exposure in early-onset asthma. N Engl J Med . 2008;359:1985–1994. doi: 10.1056/NEJMoa0806604. [DOI] [PubMed] [Google Scholar]
- 30. Blekic M, Kljaic Bukvic B, Aberle N, Marinho S, Hankinson J, Custovic A, et al. 17q12-21 and asthma: interactions with early-life environmental exposures. Ann Allergy Asthma Immunol . 2013;110:347–353.e2. doi: 10.1016/j.anai.2013.01.021. [DOI] [PubMed] [Google Scholar]
- 31. Hallmark B, Wegienka G, Havstad S, Billheimer D, Ownby D, Mendonca EA, et al. Chromosome 17q12-21 variants are associated with multiple wheezing phenotypes in childhood. Am J Respir Crit Care Med . 2021;203:864–870. doi: 10.1164/rccm.202003-0820OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Marinho S, Custovic A, Marsden P, Smith JA, Simpson A. 17q12-21 variants are associated with asthma and interact with active smoking in an adult population from the UK. Ann Allergy Asthma Immunol . 2012;108:402–411.e9. doi: 10.1016/j.anai.2012.03.002. [DOI] [PubMed] [Google Scholar]
- 33. Loss GJ, Depner M, Hose AJ, Genuneit J, Karvonen AM, Hyvärinen A, et al. PASTURE (Protection against Allergy Study in Rural Environments) Study Group The early development of wheeze: environmental determinants and genetic susceptibility at 17q21. Am J Respir Crit Care Med . 2016;193:889–897. doi: 10.1164/rccm.201507-1493OC. [DOI] [PubMed] [Google Scholar]
- 34. Bräuner EV, Loft S, Raaschou-Nielsen O, Vogel U, Andersen PS, Sørensen M. Effects of a 17q21 chromosome gene variant, tobacco smoke and furred pets on infant wheeze. Genes Immun . 2012;13:94–97. doi: 10.1038/gene.2011.51. [DOI] [PubMed] [Google Scholar]
- 35. Stokholm J, Chawes BL, Vissing N, Bønnelykke K, Bisgaard H. Cat exposure in early life decreases asthma risk from the 17q21 high-risk variant. J Allergy Clin Immunol . 2018;141:1598–1606. doi: 10.1016/j.jaci.2017.07.044. [DOI] [PubMed] [Google Scholar]
- 36. Zhang Y, Hua L, Liu QH, Chu SY, Gan YX, Wu M, et al. Household mold exposure interacts with inflammation-related genetic variants on childhood asthma: a case-control study. BMC Pulm Med . 2021;21:114. doi: 10.1186/s12890-021-01484-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Gorlanova O, Illi S, Toncheva AA, Usemann J, Latzin P, Kabesch M, et al. BILD and PASTURE study groups Protective effects of breastfeeding on respiratory symptoms in infants with 17q21 asthma risk variants. Allergy . 2018;73:2388–2392. doi: 10.1111/all.13568. [DOI] [PubMed] [Google Scholar]
- 38. Kelly RS, Chawes BL, Guo F, Zhang L, Blighe K, Litonjua AA, et al. The role of the 17q21 genotype in the prevention of early childhood asthma and recurrent wheeze by vitamin D. Eur Respir J . 2019;54:1900761. doi: 10.1183/13993003.00761-2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Tang L, Lu C, Zheng G, Burgering BM. Emerging insights on the role of gasdermins in infection and inflammatory diseases. Clin Transl Immunology . 2020;9:e1186. doi: 10.1002/cti2.1186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Li X, Ampleford EJ, Howard TD, Moore WC, Torgerson DG, Li H, et al. Genome-wide association studies of asthma indicate opposite immunopathogenesis direction from autoimmune diseases. J Allergy Clin Immunol . 2012;130:861–868.e7. doi: 10.1016/j.jaci.2012.04.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Kreiner E, Waage J, Standl M, Brix S, Pers TH, Couto Alves A, et al. Shared genetic variants suggest common pathways in allergy and autoimmune diseases. J Allergy Clin Immunol . 2017;140:771–781. doi: 10.1016/j.jaci.2016.10.055. [DOI] [PubMed] [Google Scholar]
- 42. Valette K, Li Z, Bon-Baret V, Chignon A, Bérubé JC, Eslami A, et al. Prioritization of candidate causal genes for asthma in susceptibility loci derived from UK Biobank. Commun Biol . 2021;4:700. doi: 10.1038/s42003-021-02227-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science . 2020;369:1318–1330. doi: 10.1126/science.aaz1776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Panganiban RA, Sun M, Dahlin A, Park HR, Kan M, Himes BE, et al. A functional splice variant associated with decreased asthma risk abolishes the ability of gasdermin B to induce epithelial cell pyroptosis. J Allergy Clin Immunol . 2018;142:1469–1478.e2. doi: 10.1016/j.jaci.2017.11.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Broz P, Pelegrín P, Shao F. The gasdermins, a protein family executing cell death and inflammation. Nat Rev Immunol . 2020;20:143–157. doi: 10.1038/s41577-019-0228-2. [DOI] [PubMed] [Google Scholar]
- 46. Tamura M, Tanaka S, Fujii T, Aoki A, Komiyama H, Ezawa K, et al. Members of a novel gene family, Gsdm, are expressed exclusively in the epithelium of the skin and gastrointestinal tract in a highly tissue-specific manner. Genomics . 2007;89:618–629. doi: 10.1016/j.ygeno.2007.01.003. [DOI] [PubMed] [Google Scholar]
- 47. Shi J, Zhao Y, Wang K, Shi X, Wang Y, Huang H, et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature . 2015;526:660–665. doi: 10.1038/nature15514. [DOI] [PubMed] [Google Scholar]
- 48. Jorgensen I, Zhang Y, Krantz BA, Miao EA. Pyroptosis triggers pore-induced intracellular traps (PITs) that capture bacteria and lead to their clearance by efferocytosis. J Exp Med . 2016;213:2113–2128. doi: 10.1084/jem.20151613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Shi J, Zhao Y, Wang Y, Gao W, Ding J, Li P, et al. Inflammatory caspases are innate immune receptors for intracellular LPS. Nature . 2014;514:187–192. doi: 10.1038/nature13683. [DOI] [PubMed] [Google Scholar]
- 50. Chen Q, Shi P, Wang Y, Zou D, Wu X, Wang D, et al. GSDMB promotes non-canonical pyroptosis by enhancing caspase-4 activity. J Mol Cell Biol . 2019;11:496–508. doi: 10.1093/jmcb/mjy056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Zhou Z, He H, Wang K, Shi X, Wang Y, Su Y, et al. Granzyme A from cytotoxic lymphocytes cleaves GSDMB to trigger pyroptosis in target cells. Science . 2020;368:eaaz7548. doi: 10.1126/science.aaz7548. [DOI] [PubMed] [Google Scholar]
- 52. Hansen JM, de Jong MF, Wu Q, Zhang LS, Heisler DB, Alto LT, et al. Pathogenic ubiquitination of GSDMB inhibits NK cell bactericidal functions. Cell . 2021;184:3178–3191.e18. doi: 10.1016/j.cell.2021.04.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Das S, Miller M, Beppu AK, Mueller J, McGeough MD, Vuong C, et al. GSDMB induces an asthma phenotype characterized by increased airway responsiveness and remodeling without lung inflammation. Proc Natl Acad Sci U S A . 2016;113:13132–13137. doi: 10.1073/pnas.1610433113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Ozcelik P, Bezirci FB, Suzuki Y, Uzawa H, Nishida Y, Kobayashi K, et al. Sulfatide and its synthetic analogues recognition by Moraxella catarrhalis. Microbiol Immunol . 2006;50:967–970. doi: 10.1111/j.1348-0421.2006.tb03873.x. [DOI] [PubMed] [Google Scholar]
- 55. Hartmann E, Lingwood CA, Reidl J. Heat-inducible surface stress protein (Hsp70) mediates sulfatide recognition of the respiratory pathogen Haemophilus influenzae. Infect Immun . 2001;69:3438–3441. doi: 10.1128/IAI.69.5.3438-3441.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Bashir H, Grindle K, Vrtis R, Vang F, Kang T, Salazar L, et al. Association of rhinovirus species with common cold and asthma symptoms and bacterial pathogens. J Allergy Clin Immunol . 2018;141:822–824.e9. doi: 10.1016/j.jaci.2017.09.027. [DOI] [PMC free article] [PubMed] [Google Scholar]



