Abstract
Niche-derived factors regulate tissue stem cells, but apart from the mechanosensory pathways, the effect of niche geometry is not well understood. We used organoids and bioengineered tissue culture platforms to demonstrate that the conical shape of Lgr5+ small intestinal stem cells (ISCs) facilitate their self-renewal and function. Inhibition of non-muscle myosin II (NM II)–driven apical constriction altered ISC shape and reduced niche curvature and stem cell capacity. Niche curvature is decreased in aged mice, suggesting that suboptimal interactions between old ISCs and their niche develop with age. We show that activation of NM IIC or physical restriction to young topology improves in vitro regeneration by old epithelium. We propose that the increase in lateral surface area of ISCs induced by apical constriction promotes interactions between neighboring cells, and the curved topology of the intestinal niche has evolved to maximize signaling between ISCs and neighboring cells.
Stem cells create a shape to listen to their surroundings.
INTRODUCTION
Cellular scale architecture is required for proper embryonic development and supports the functionality of adult tissues (1–3). However, how much the physical cell shape affects stem cell behavior is not well understood. We exploited the intestine as a model to investigate whether the geometry of cells and tissues plays a role in stem cell maintenance. Intestinal stem cells (ISCs) reside in a highly curved environment and constantly renew the epithelium (4), and their function is regulated by paracrine factors from the neighboring epithelial cells (5–7), stipulating effective cell-cell interactions.
To explore how tissue geometry affects ISC function, we used in vitro organoid cultures (8) and bioengineered culture scaffolds. ISCs in organoids, such as in tissue, are surrounded by neighboring Paneth cells, which produce ligands that maintain ISCs by activating crucial pathways such as Wnt and Notch (7). The epithelium containing ISCs and Paneth cells can self-organize and generate in vivo–like crypt domains with curvature during organoid culture (8, 9). Crypt morphogenesis occurs in vitro despite the missing stroma that would provide additional physical constraint and spatial cues in the form of locally produced factors, such as R-spondins and bone morphogenetic protein (BMP) inhibitors (10, 11). These factors are provided uniformly via medium in the organoid culture. Therefore, this setup allows examination of the intrinsic relationship between epithelial contour and stem cell function. Using organoids together with bioengineered scaffolds, we found that cell shape is critical for ISC function.
RESULTS
ISCs actively acquire a conical shape
The Lgr5-EGFP-IRES-CreERT2 mouse model (12) allows identification of the bona fide ISCs (Lgr5hi) and differentiating transit amplifying (TA; Lgr5lo) cells based on their enhanced green fluorescent protein (EGFP) intensity. ISCs located at the crypt bottom have a conical shape to accommodate the positive curvature, while TA cells are pushed upward to less curved sides of the crypt (Fig. 1A). When differentiating cells emerge from the crypt, they encounter negative curvature at the exit (13) and, lastly, migrate upward toward the villus tip where they experience extreme curvature opposite in concavity to that in the bottom of the crypts and are extruded into the gut lumen (9). Epithelial organoids form in vivo–like crypt domains and exhibit a similar gradual transition from positive to negative curvature along the crypt to villus domain axis (Fig. 1B).
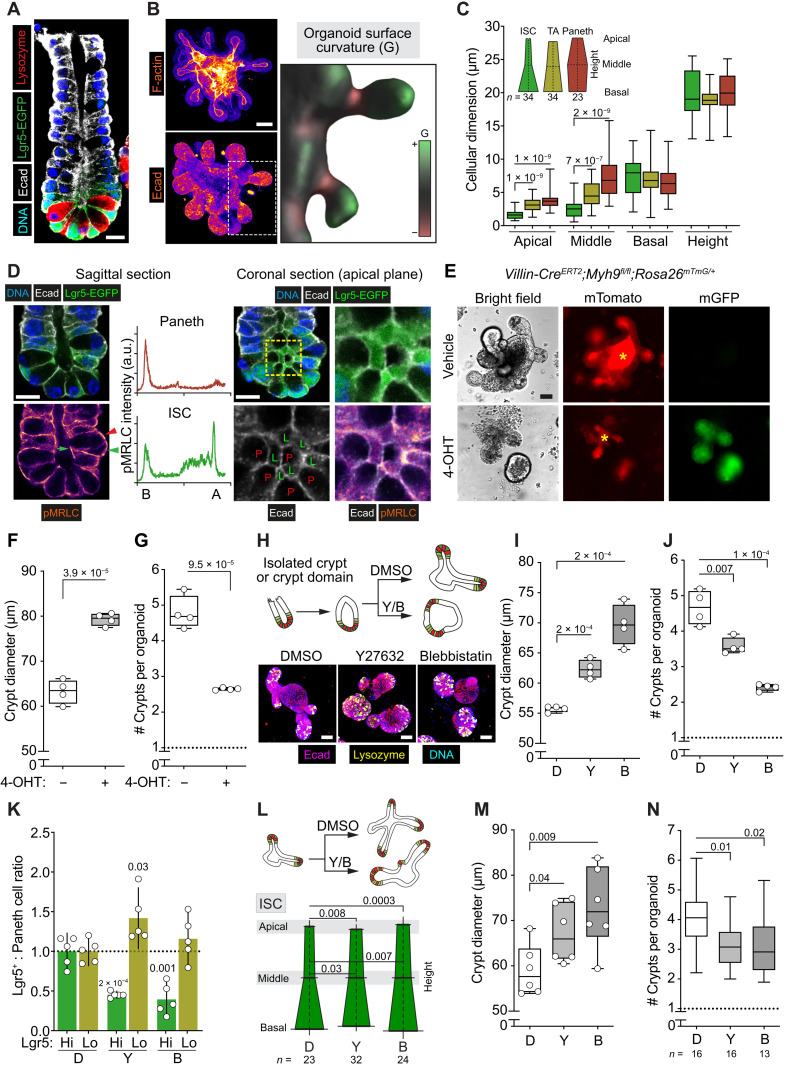
Fig. 1. Apical constriction required for ISC function.
(A) Immunostaining of intestinal crypt. Lgr5-EGFP (green), lysozyme (red), DNA (blue), and E-cadherin (white). (B) Intestinal organoids stained with phalloidin (F-actin) (top) and E-cadherin (bottom). Inset: Surface plot of the organoid crypt domains. Positive (green), zero (gray), and negative (red) Gaussian curvature (G). (C) Quantification of ISC, TA, and Paneth cell morphology in organoids. (D) Left: Sagittal plane of isolated crypt. Lgr5-EGFP (green), E-cadherin (white), DNA (blue), and phosphorylated myosin regulatory light chain (pMRLC; fire). ISCs (green arrowhead) show higher pMRLC intensity apically (green arrow) compared with Paneth cells (red arrowhead). Distribution of pMRLC in Paneth and ISCs: B, basal and A, apical. Right: Coronal plane visualizing presence and lack of apical pMRLC in ISCs (L) and Paneth cells (P). (E) Myh9 knockout induced with 4-hydroxytamoxifen (4-OHT) 2 days after passage in Villin-CreERT2;Myh9fl/fl;Rosa26mTmG/+ mouse–derived organoids. Images 6 days after passage. Note that membrane-localized EGFP (mGFP) is absent in vehicle-treated organoids. Asterisk, luminal background. (F and G) Crypt width and crypt number in organoids from (E) (n = 4). (H) Schematics and images from organoids treated 48 hours with dimethyl sulfoxide (DMSO) (D), Y-27632 (Y), or Blebbistatin (B), E-cadherin (magenta), lysozyme (yellow), and DNA (blue). (I and J) Crypt width and number after treatment (n = 4 replicate wells). (K) ISC and TA cell frequencies from Lgr5-EGFP-IRES-CreERT2 mouse–derived organoids relative to Paneth cell number (n = 5). (L) Organoid treatment schematics and ISC morphology after 48 hours of exposure to inhibitors. (M) Crypt diameter after 48 hours of exposure to inhibitors (n = 6). (N) Regenerative capacity of subcultured crypt domains in the absence of inhibitors. n values represent independent experiments. Unless otherwise indicated, in box plots, the line represents median, the box shows interquartile range, and whiskers indicate the range. All other data are represented as means ± SD. P values are shown in corresponding panels. Scale bars, 10 μm (A and D) and 50 μm (B, E, and H). a.u., arbitrary units.
We first addressed the contribution of various cell types to the overall curvature of the intestinal niche in the domains at the bottom of the crypts of self-organized organoids by analyzing their cellular dimensions. Within the crypt domain, ISCs exhibit the smallest apical surface (Fig. 1C and fig. S1A). Moreover, ISCs in freshly isolated crypts demonstrated apical non-muscle myosin II (NM II) activity based on the phosphorylation status of NM II regulatory light chain (pMRLC; Fig. 1D and fig. S1B). NM II activity is the driver of apical constriction, promoting a conical cellular shape (14), suggesting that active constriction happens in vivo like previously reported for growing organoids (15). We confirmed the apical localization of pMRLC in ISCs of in vitro organoids (fig. S1C). Paneth cells, the cell type dominating the crypt bottom together with the ISCs, lacked apical pMRLC and had larger apical surfaces (Fig. 1, C and D, and fig. S1, B and C), suggesting that crypt curvature is mainly driven by ISCs. In support of this, when organoids were cultured under conditions promoting a stem cell identity (16), they preferably formed narrow tubular crypt domains in contrast to organoids where differentiation was directed to the Paneth cell lineage (fig. S1, D and E). Together, these data demonstrate that ISCs actively acquire a conical shape without mechanical help from the stroma or Paneth cells and that they are a driver of epithelial curvature in the intestinal niche.
Prevention of apical constriction alters ISC shape and reduces capacity to regenerate
To test whether ISCs require tissue-like crypt topology for their function, we targeted crypt formation in organoids with a conditional knockout of Myh9, one of the two epithelial isoforms of NM II (NM IIA) (13). The removal of NM IIA from the epithelium enlarged the size of forming crypts, demonstrating a loss of apical constriction, and reduced their number, suggesting attenuated ISC function (Fig. 1, E to G). To have more precise control on the level and timing of NM II activity, we titrated two NM II inhibitors Y-27632 and Blebbistatin (17, 18) to find a dose that prevents apical constriction to a similar level (fig. S1, F to H). The chosen concentrations of NM II inhibitors effectively prevented crypt formation in organoids (Fig. 1H) without affecting cell proliferation marked by 5-ethynyl-2′-deoxyuridine incorporation (fig. S1I). Similar to the NM IIA knockout, we observed increased width and reduced number of crypt domains, indicating that NM II activity and not actomyosin structure maintains crypt shape via apical constriction (Fig. 1, H to J, and fig. S1, F to H), corroborating previous reports (19). We next analyzed cellular frequencies using flow cytometry to address the number of ISCs when curvature is altered. Treatment with Y-27632 and Blebbistatin decreased the portion of Lgr5hi ISCs supported by the Paneth cells, while frequency of Lgr5lo TA cells was increased or unaltered (Fig. 1K), suggesting that stem cell maintenance is compromised in a less curved niche.
To functionally probe whether ISCs in the wider niche have altered stem cell capacity, we allowed organoids to first grow crypt domains before applying the inhibitors (Fig. 1L). After 48 hours of treatment with Y-27632 and Blebbistatin, the apical and middle diameter of the ISCs was significantly increased, whereas the shapes of Paneth and TA cells were not significantly altered (Fig. 1L and fig. S1J). Consequently, the observed increase in crypt diameter (Fig. 1M) was induced specifically by changes in the shape of ISCs. When the crypt domains of treated organoids were isolated and recultured in the absence of inhibitors, the regenerative output of wider crypts was significantly decreased (Fig. 1N). This indicates that NM II–mediated apical constriction maintains typical ISC shape, which is required for the maintenance of the regenerative capacity of intact crypt units. Yes-associated protein (YAP) is a key mediator of the Hippo pathway and involved in mechanosensing of the surrounding environment (20). In addition, YAP is activated when tension is generated within the cell (21). Therefore, YAP might mediate effects of altered ISC shape.
YAP activity does not mediate the effect of altered ISC shape but regulates Paneth cell differentiation
In the small intestine, YAP has been shown both to maintain stemness and to limit overgrowth of the tissue (22, 23). As inhibition of NM II contractility could alter the mechanosensing properties of intestinal epithelium, we analyzed the nuclear localization of YAP from inhibitor-treated organoids as a proxy for their YAP activity (20, 24).
In control organoids and in vivo, YAP accumulation in the nucleus was higher in ISCs than in the neighboring Paneth cells (fig. S2, A and B), which is consistent with previous reports (23). Treatment of organoids with Y-27632 or Blebbistatin decreased nuclear YAP (fig. S2C). A similar reductive effect was observed in treatments with low doses of a YAP inhibitor, verteporfin (fig. S2C) (25). Verteporfin at 10 nM did not affect organoid growth or change the crypt shape (fig. S2D), suggesting that NM II inhibitors’ effects on regeneration (Fig. 1, H, J, and N) are driven by mechanisms independent of YAP. Moreover, reduced YAP activity has been reported to drive Paneth cell formation (23). Correspondingly, we observed increased Paneth cell number in the NM II and YAP-inhibited crypts (fig. S2E). Together, these data indicate that the reduction in regenerative growth by Y-27632 and Blebbistatin was not driven by YAP and occurred despite the increase in number of supporting niche cells.
Mimicking the native niche curvature maintains ISC function
To directly address whether the reduction in stemness caused by NM II inhibition was induced by altered cell shape and decreased niche curvature, we designed and bioengineered collagen scaffolds that allow the culture of epithelium on custom topographies (Fig. 2A). The median crypt diameter in the mouse gastrointestinal tract ranges between 45 and 60 μm (fig. S3A), and the average crypt diameter in cultured organoids is approximately 60 ± 5 μm (Fig. 1, F, I, and M, and fig. S1D). Therefore, we engineered scaffolds with a range of pit sizes (50, 75, 100, and 125 μm in diameters; Fig. 2A) that would accommodate epithelial curvatures from typical to supraphysiological ranges.
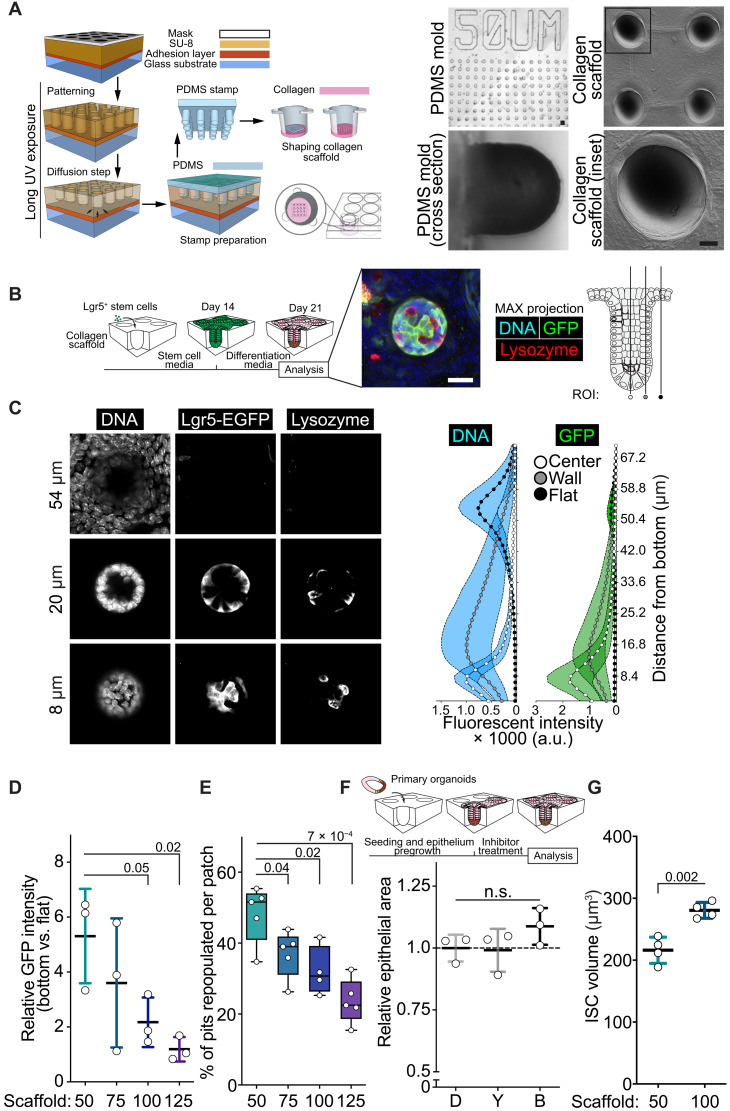
Fig. 2. Bioengineered scaffold mimicking native crypt curvature supports ISC function.
(A) Schematics of the photolithography technique used to prepare rounded-bottom collagen scaffolds. Representative images of the 50-μm PDMS (bright field; scale bar, 50 μm) or collagen [scanning electron microscopy (SEM); scale bar, 10 μm] scaffolds prepared using the technique. (B) Timeline for the established epithelial cultured originating from isolated Lgr5hi ISCs. Maximum projection image of a formed crypt. DNA (blue), Lgr5-EGFP (green), and lysozyme (red). Schematic showing location of the regions of interest (ROIs) used in analysis. Scale bar, 25 μm. (C) Left: Single plane images of bottom, middle, and top layer of the immunostained crypt. Right: Quantification of crypt domains formed on collagen scaffolds. n = 3. (D) Quantification of 4′,6-diamidino-2-phenylindole (DAPI)–normalized intensity of GFP from Lgr5-EGFP-IRES-CreERT2 mouse–derived epithelium grown on various-sized (50 to 125 μm) collagen scaffolds. Data are represented in relation to the flat surface. n = 3 independent experiments. (E) Quantification of the capacity to maintain crypts populated after prolonged culture on various crypt sizes. n = 5 independent experiments. (F) Growth capacity of epithelium cultured on 50-μm collagen scaffold of 48 hours with 20 μM Y-27632 (Y), 10 μM Blebbistatin (B), or vehicle (DMSO) (D). n = 3 independent experiments. (G) Quantification of ISC volume from epithelium grown on 50- and 100-μm scaffolds. n = 4 independent experiments. Unless otherwise indicated, in box plots, the line represents median, the box shows interquartile range and whiskers show the range. All other data are represented as means ± SD. P values are shown in corresponding panels. n.s., not significant.
Isolated Lgr5hi ISCs were then cultured on scaffolds under conditions that maintain ISCs in an undifferentiated state (“stem cell media”; see Materials and Methods; Fig. 2B). After 14 days of culture, a continuous monolayer of Lgr5+ cells covered most of the scaffold and followed its topology. At this point, the medium was switched to regular organoid culture medium (“differentiation media”; see Materials and Methods for ENR) allowing differentiation (Fig. 2B).
After differentiation, the highest Lgr5-EGFP expression was restricted to the bottom of the curved pits in the 50-μm scaffolds, similar to that shown in similar hydrogel devices (26, 27). Lgr5-EGFP expression gradually decreased in cells oriented toward the upper part of the crypt and was lowest in cells outside the crypt (Fig. 2, B and C, and fig. S3, B and C). Cells in pits with wider diameters expressed Lgr5-EGFP at a lower level (Fig. 2D and fig. S3, C and D), suggesting that decreases in the physical curvature of the monolayer reduces stem cell maintenance, similar to the impact of NM II inhibition in organoids (Fig. 1, I to N). Moreover, on scaffolds with wide crypts, a lower percentage of crypts remained populated (Fig. 2E). As these wide crypts contained a higher number of potentially ISC-supporting Paneth cells (fig. S3E), these results suggest that an increase in crypt width alters niche interactions that are critical for stem cell maintenance. Cells grown on the widest crypts (125 μm) often formed multiple crypt domains within a single scaffold pit. Each such crypt had a diameter of ~50 μm (fig. S3F), implying a strong preference for the native crypt width even when the environment supports a lower degree of curvature.
The bioengineered scaffolds allowed us to test whether mechanical restriction of crypt size to 50 μm would be enough to blunt the effects of NM II inhibitors on organoids. Y-27632 or Blebbistatin did not reduce the growth kinetics of epithelium grown on 50-μm scaffolds (Fig. 2F), contrary to the inhibitors’ effects on traditional Matrigel-embedded organoids (Fig. 1I). Furthermore, crypt cells on the wider 100-μm scaffolds accumulated YAP protein in the nucleus more than cells on 50-μm scaffolds (fig. S4, A to C), unlike ISCs in organoids treated with Y-27632 or Blebbistatin (fig. S2C). This dichotomy in YAP signaling between crypt cells cultured on wider scaffolds and wide organoid crypts induced by NM II inhibitors suggests that additional mechanisms influence the decline in regenerative capacity of the intestinal epithelium in a lower-curvature environment.
To probe for alternative mechanisms, we examined the effect of wider scaffold features on the physical appearance of cells. Theoretically, cells at the bottom of 100-μm scaffolds experience four times lower curvature than on 50-μm scaffolds (Gaussian curvature = 0.0004 versus 0.0016, respectively). Cellular density was not altered between epithelium grown within 100- and 50-μm features, suggesting that independent of the physical restraint induced by the epithelial curvature, cells are able to maintain equal density (fig. S4D). This indicates that changes in the individual cell parameters, such as shape or volume, have to compensate for the change in curvature to accommodate the same number of cells evenly. We found that the volume of individual ISCs was larger when cultured on 100-μm scaffolds (Fig. 2G and fig. S4E).
Together, these data suggest that NM II–mediated apical constriction of ISCs maintains cell shape and optimal niche curvature required for stem cell maintenance and function. In line with this notion, both pharmacological and physical reductions of the niche curvature decrease the regenerative potential of the epithelium. The effect of NM II inhibition is blunted by physically preventing crypt relaxation, suggesting that the mechanosensing pathways, including the YAP pathway, have no major contribution to the phenotype. ISCs on wider 100-μm scaffolds exhibit larger cell volumes, suggesting that altered physical features might mediate ISC function.
High surface-to-volume ratio improves signal reception in ISCs
To mechanistically understand how the drop in the regenerative capacity of ISCs manifests when the apical surface, and consequently the shape of the niche, is altered, we reviewed physical properties of other adult stem and progenitor cells. A common feature among undifferentiated cells is their smaller size (28). Many adult stem cells, such as hematopoietic and basal keratinocytes, are smaller than their differentiated progeny (29, 30). Consistently, in the intestinal crypt, ISCs are smaller than their progeny (fig. S5A). We reasoned that, like with a smaller size, the changes in cell morphology could increase the surface-to-volume ratio, allowing, for example, more effective sensing of extracellular factors. In the context of adult stem cells, this could improve the capacity to receive signals from the surrounding niche, such as neighboring Paneth cells. Similarly, the conical shape of ISCs further increases the lateral surface-to-volume (LSV) ratio and could therefore provide a larger signaling interface with neighboring Paneth cells (Fig. 3A). Moreover, reducing apical constriction with NM II inhibition led to increased volumes and smaller LSV ratios (Fig. 3, A and B), suggesting that stem cells’ ability to receive niche signals is attenuated under these conditions.
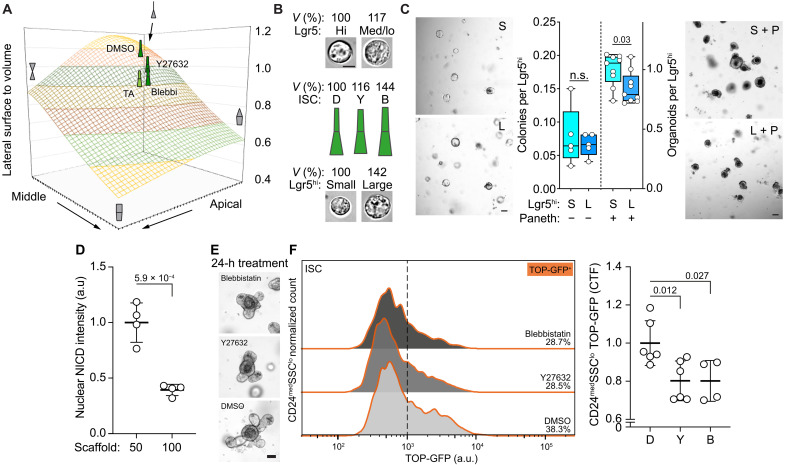
Fig. 3. Increased surface-to-volume ratio promotes signaling capacity between Paneth and ISCs.
(A) Relation between the apical constriction and lateral surface area to volume ratio. Basal width and cell height reflects the mean observed values from cells in organoid crypt domains. Diameter of apical and middle position varies from 0 to 8 μm. (B) Volumes (V %) of the corresponding ISCs and TA cells presented in relation to the smallest within respective groups. Scale bar, 5 μm (C) Left: Colony forming capacity of isolated S-ISCs and L-ISCs in the absence of Paneth cells or external Wnt supplementation. Colonies quantified and representative images taken on day 5. n = 5 animals. Right: Organoid forming capacity of S-ISCs and L-ISCs when cocultured together with the Paneth cell niche. Organoids quantified and representative images taken on day 6. n = 9 animals. (D) Quantification of nuclear NICD in Lgr5+ ISCs cultured on 50- or 100-μm scaffolds. Data represented in relation to average ISC on 50-μm scaffolds. n = 4 crypts quantified. (E) Representative images of TOP-GFP organoids culture for 24 hours with NM II inhibitors. (F) Left: Representative TOP-GFP histogram from flow analysis of ISC containing population CD24medSSClo (see fig. S5L for gating strategy). Right: Quantification of the flow analysis. Data are represented in relation to the average of DMSO control. n = 6 (for DMSO and Y27632) or 4 (Blebbistatin) replicate cultures from two independent experiments. Unless otherwise indicated, in box plots, the line represents median, the box shows interquartile range, and whiskers show the range. All other data are represented as means ± SD. P values are shown in corresponding panels. Unless otherwise noted, scale bars are 50 μm.
To experimentally test whether ISCs benefit from a large LSV ratio, we sorted ISCs on the basis of their size (Fig. 3B and fig. S5C), which did not select for cells with differential expression of stemness-related genes (fig. S5D). On average, the large ISCs (L-ISC) were 42% (±36%) larger in volume than the small ISCs (S-ISC; fig. S5C), corresponding to the increase in size and reduction in LSV ratio observed in Blebbistatin-treated ISCs (Fig. 3, A and B). When cultured alone in niche-free conditions where ISCs can form spheroids, indicative of their intrinsic clonogenicity, different-sized ISCs form colonies at the same frequency, indicating that both populations had equal intrinsic stem cell potency regardless of the difference in their cell cycle profile (Fig. 3C and fig. S5, D to F). However, the population of S-ISCs initiated organoids with a higher frequency than larger cells when cocultured together with their Paneth cell niche (Fig. 3C). This supports the idea that niche-derived signals are more efficiently received and integrated into ISC function when LSV ratio of stem cells is higher. S- and L-ISCs retained equal levels of mammalian target of rapamycin complex 1 (mTORC1) activity (fig. S5, G and H), indicating that enlarged cell size is not driven by hyperactive protein synthesis, known to reduce stem cell function (31). To compare Paneth-to-ISC signaling with ISCs of different LSV ratio, we used the bioengineered scaffolds, where ISC volumes increase, but LSV ratio decreases with increase in crypt size (Fig. 2G). We first focused on the nuclear levels of Notch intracellular domain (NICD) that is cleaved from Notch receptor and translocates to the nucleus upon Notch receptor activity (32). Consistent with ISCs responding to Paneth cell–produced delta ligands in an attenuated fashion, nuclear NCID of Lgr5+ cells was reduced on wider scaffolds (Fig. 3D and fig. S5I). To probe for alterations in other intercellular signaling pathways that are important for ISCs, we generated reporter organoids sensitive to Wnt3 signals from Paneth cells (see the “Generation and analysis of TOP-GFP organoids” section in Materials and Methods; fig. S5J). Twenty-four–hour treatment with NM II inhibitors reduced Wnt response in the population containing ISCs (Fig. 3, E and F, and fig. S5, K and L). Jointly, these findings indicate that a high LSV ratio in ISCs may facilitate effective reception and utilization of stemness supporting niche factors.
Niche curvature is altered during aging and reduces stem cell function
To investigate whether alterations in LSV ratio affect ISC function under physiological conditions, we set out to study the tissue architecture of aged intestine. The regenerative potential of intestinal epithelium declines with age because of intrinsic and extrinsic factors of ISCs (5, 33). We found that crypts in old mice are larger than in young mice, and as a result, overall crypt density is lower in older mice (Fig. 4A and fig. S6A). These factors contribute to a lower-curvature environment that ISCs in old individuals experience in comparison with that in young individuals. We and others have recently observed increased numbers of Paneth cells in aged tissue, while ISC number is either unchanged or decreased (5, 34). Increase of crypt width either by NM II inhibition or by scaffold diameter resulted in similar effects (Figs. 1, I and K and, 2, D and E, and fig. S3E). Furthermore, cell volume of ISCs was increased in old animals, while sizes of progenitors and Paneth cells were unchanged (Fig. 4B and fig. S6B). We therefore hypothesized that reduced curvature of old crypts results from changes in cellular shape, and by decreasing the LSV ratio of ISCs, these changes may contribute to the reduced ISC maintenance in the old intestine.
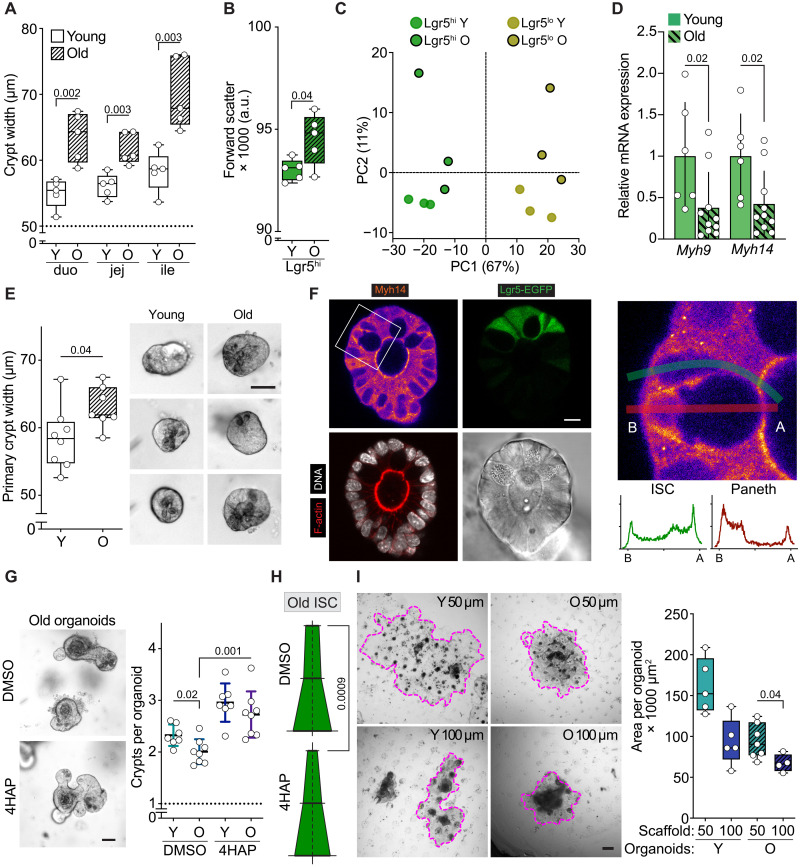
Fig. 4. Young topology enhances regenerative capacity of old ISCs.
(A) Quantification of crypt width from resected intestinal sections of young (Y) and old (O) animals. n = 5 animals per group. (B) Analysis of the ISC size by flow cytometry. n = 5 animals per group. (C) Two-dimensional (2D) principal components analysis of mRNA-profiled young and old Lgr5hi (ISCs) and Lgr5lo (TA) cells. n = 3 animals per group. (D) Expression of NM II heavy chain coding genes Myh9 and Myh14 in young and old ISCs. n = 6 young and 10 old. Fold chain compared to young ISCs. Means ± SD. (E) Quantification and representative images of 2-day-old organoids derived from young and old mice and cultured in the absence of Y-27632. Scale bar, 50 mm. (F) Left: Immunofluorescent staining of organoid crypt domain demonstrating apical localization of Myh14 (fire) in ISCs (Lgr5-EGFP, green). DNA (DAPI, white) and phalloidin (red) were used to identify cell borders. Scale bar, 10 μm. Right: Myh14 staining and quantification of basal (B) to apical (A) distribution along the ISC (green line) and Paneth cell (red line). n = 3 cell pairs analyzed with similar outcome. (G) Left: Representative images of old organoids grown for 4 days in the presence of 500 μM 4HAP or vehicle. Scale bar, 50 μm. Right: Regenerative growth of young and old organoids supplemented with 500 μM 4HAP or vehicle. (H) Quantification of old ISC shape cultured 4 days in the presence of 4HAP or vehicle. n = 31 cells. (I) Representative images and quantification of epithelial area of young or old organoids grown on 50- or 100-μm scaffolds. Scale bar, 200 μm. n = 4 animals per group. Unless otherwise indicated, in box plots, the line represents median, the box shows interquartile range, and whiskers show the range. All other data are represented as means ± SD. P values are shown in corresponding panels.
To discover potential mechanisms for reshaping of the old ISCs, we expanded our previously published transcriptome analysis of ISCs (Lgr5hi) (5) to include more differentiated TA cells (Lgr5lo) from the same animals. This allowed us to characterize the machinery involved in the maintenance of apical constriction in ISCs (see Materials and Methods). Principal components analysis demonstrated that the four groups analyzed are separated by axes of differentiation state and age (x and y, respectively, in Fig. 4C). Comparison of young ISCs and their progeny revealed high enrichment of actin cytoskeleton and cellular morphology–related genes in ISCs (fig. S6C and table S1), highlighting that cellular shape is actively maintained in stem cells.
Among the genes altered with age, we identified that myosin heavy chain 14 (Myh14), a defining component of the NM IIC isoform, had reduced expression in old ISCs (table S2). In intestinal crypts, the main NM II constituent is Myh9 (NM IIA isoform), Myh14 (NM IIC) contributes but to a lesser extent, and Myh10 (NM IIB) is not expressed (13). Quantitative polymerase chain reaction (qPCR) analysis of ISCs confirmed age-associated reduction in both NM II paralogs, suggesting reduced capacity for apical constriction in old ISCs (Fig. 4D). In support of this, the morphological analysis of isolated crypts cultured ex vivo for 48 hours indicated that even in the absence of stroma, old crypt domains are wider, possibly because of reduced NM II activity in stem cells (Fig. 4E). These data, together with observations that NM II activity contributes to cellular stiffness (35), support the idea of aged ISCs having reduced capacity to control their shape, a feature that could contribute to the increased crypt width in old tissue.
Regeneration of old epithelium is improved by NM IIC activation or youthful topology
Our data suggest that the age-associated decrease in crypt curvature (Fig. 4, A and B) may result from reduced apical constriction in stem cells, in turn, reducing their LSV ratio, and might thereby dilute the impact of niche-derived factors. This could further aggravate the known reduction in Wnt signaling activity of old ISCs (5, 34), which was recently demonstrated to attenuate regenerative capacity during aging. Therefore, we were inspired to test whether NM II activation could increase crypt curvature and thus enhance the performance of old ISCs.
Immunofluorescence staining of the NM II paralogs revealed both apical and basolateral localization of Myh9/NM IIA in all cell types composing the organoids, whereas Myh14/NM IIC was mostly localized to the apical surface (Fig. 4F and fig. S6D). Within the crypt, Paneth cells show opposite localization of NM IIC, possibly reflecting their function as secretory cells with capacity to rapidly release antimicrobial granules to the gut lumen (Fig. 4F) (36). Given these differences in subcellular localization, we sought to induce apical constriction specifically in ISCs through targeted activation of NM IIC.
4′-Hydroxyacetophenone (4HAP) is a natural compound that has been shown to activate both NM IIB and NM IIC but not NM IIA (35, 37). As NM IIB is not expressed by the intestinal epithelium (13), 4HAP acts as a specific NM IIC activator in the organoid culture model. Consistently, treating organoid cultures with a tolerable dose of 4HAP (fig. S6E) induced higher crypt curvature and promoted crypt formation without causing ectopic expression of Myh10 or altering expression levels of Myh9 and Myh14 (fig. S6, F to I).
To investigate whether NM IIC activity could improve signaling in the old intestine, freshly isolated crypts from young and old animals were treated with 4HAP. Four-day treatment increased ISC numbers and regenerative growth of young and old epithelium (Fig. 4G and fig. S6J). Moreover, the apical size of old ISCs together with the organoid crypt width was reduced upon 4HAP treatment (Fig. 4H and fig. S6K), implying enhanced activity of apical NM IIC in ISCs. Crypts isolated from 4HAP-treated organoids maintained improved growth in the secondary culture free of 4HAP, likely because of the increased ISC numbers after treatment (fig. S6, J and L). In further support of the hypothesis that the growth-promoting effects of 4HAP occur via changes in cell and crypt shape, 4HAP treatment had no effect on organoid growth in 50-μm scaffolds where crypt curvature was already physically enforced through scaffold shape (fig. S6M).
Last, to investigate whether simple physical restriction of crypt topology could enhance the function of old ISCs and their niche, we cultured epithelium-derived ISCs from young and old animals on scaffolds with 50- and 100-μm-wide pits (Fig. 4H). Old organoids produced new epithelium faster on 50 μm when compared to growth on 100 μm (Fig. 4H), suggesting improved signaling between the Paneth and ISCs. However, the growth was not completely rescued to a youthful level (Fig. 4H), underlining that other factors intrinsic to the aged epithelium (5, 33) are still limiting ISC function in this setting. However, the positive response of old epithelium to an environment with increased curvature indicates the importance of proper niche topology in maintaining an optimal physiological context and suggests that apical constriction could be a promising target for proregenerative therapies.
DISCUSSION
Geometry plays an important role in tissue function. On a cellular level, specialized cell types come in various sizes and shapes, which are tailored to facilitate their function (38). Cellular shape has been shown to affect stem cell function through experiments where cells are studied outside their native environment (39–42). These studies have indicated that changes in cell shape induced by substrate stiffness or mechanical spreading can influence cell fate decisions, demonstrating that cellular shape is an important feature of stem cells (39, 43). Moreover, the columnar shape of intestinal cells has been suggested to facilitate the typical patterning of their progeny (44). However, whether stem cell shape affects stem cell function in the context of their natural surroundings is not well understood. On a molecular level, focus has been around the YAP pathway, a key sensor of the physical environment around the cells (21). YAP has been shown to participate in stem cell regulation in multiple tissues, including the intestine (22, 23). Here, we report that cellular shape affects ISC function by regulating YAP-independent signaling platforms between stem cells and their niche (Fig 1H and fig. S2D).
In the small intestine, conical ISCs continuously renew the epithelium and rely on tightly controlled levels of niche-produced factors (7–9). We found that the stereotypical conical shape of ISCs is an actively acquired trait (Fig. 1, A to D) and required for regenerative function (Fig. 1, H and N). NM II–mediated apical constriction maintains the typical ISC shape and simultaneously bends the surrounding epithelium (Fig. 1, A, B, and D). NM II inhibition relaxed the apical surface of ISCs and reduced their regenerative capacity (Fig. 1, H, L and N), suggesting that the physical shape is a major regulator of stem cell function. While ISC-intrinsic NM II activity creates near-optimal niche curvature in vitro, it does not fully reach the in vivo–type uniform crypt diameter (Fig. 1M and fig. S3A). This suggests that both epithelial and stromal mechanics contribute to the final tissue architecture and supports the use of bioengineered culture platforms with accurate control of epithelial topology in vitro (Fig. 2, A to C). Our data also highlight that tissue morphology is not just a passive outcome of developmental programs but a critical factor for tissue function, and it can change under physiological perturbations, such as aging.
What are the physiological constraints that have set the evolutionary limits on ISC geometry? Previous reports have revealed that smaller cell volumes can increase the intracellular crowding, facilitating signaling pathway activity (40, 45). Moreover, an excessively large cytoplasmic volume can reduce cellular function by inducing senescence (46), underlining the importance of correct cell size. Further reduction of the ISC volume is likely limited by their need to maintain apical and basal connections, which are fundamental to their epithelial nature. Polarity defects are known to compromise ISC function (47). Simultaneously, a large contact area with ISC lateral neighbors, Paneth cells, could increase signal intensity, as is shown for the Notch pathway (48), thus establishing another parameter to be balanced in cell size optimization. An apically constricted conical shape allows cells to have a relatively large lateral surface area while maintaining a minimal volume. Whether the typical geometry of individual ISCs is frustum-like or scutoid shaped, allowing additional neighbor contacts (49), remains an open question.
Here, we show that ISC LSV ratio increases with apical constriction (Fig. 3A) and hypothesize that their conical shape provides a means to amplify signals from their neighboring Paneth cells. In support, lower ISC volume and improved stem cell function correlated only when niche factor–producing Paneth cells were present (Fig. 3C). Our data demonstrating that signaling activity of two pathways critical for ISC self-renewal, Notch and Wnt, is reduced when LSV ratio decreases suggest that mechanisms to maintain minimal cellular volume and maximal lateral surface have evolved to keep tissue-resident stem cells receptive to their surrounding niche signals (29, 30). This also reveals a new mode of self-regulation by ISCs, where the self-promoted tissue geometry improves niche to stem cell signaling.
We found that the size of ISCs is increased in older mice, coinciding with reduced crypt curvature (Fig. 4, A and B). Our data suggests that the apical contractility is impaired in old ISCs, but by activating apically localized NM IIC, the growth potential of old epithelium can be improved (Fig 4G and fig S6, F to L). Whether similar reduction in stem cell mechanics underline the age-associated morphological changes in other tissues, such as the skin (50), will be an interesting avenue for future research. Furthermore, these results demonstrate that improving cell contractility can improve function in old stem cells. To our knowledge, this has gone previously unexplored.
Measuring individual cell behavior in tightly packed epithelium is technically challenging. Moreover, separating the effects of cellular shape from mechanosensing with the current tools is practically impossible. In the current study, we used several approaches to investigate how cellular shape affects ISC function. Often, these experiments rely on a setup where the whole epithelial curvature was altered, rather than precisely affecting only ISCs. While we show that ISC shape was altered under these conditions, we cannot fully rule out the effect being indirect via other crypt cells or the surrounding stroma in vivo. Simultaneously, we measured the activity of a key mechanosensing pathway YAP to ensure that the effects were not due to altered mechanics in the epithelia. To minimize the confounding effects caused by modulation of the whole intact epithelium and to probe the role of individual cells, we also performed experiments where only ISCs and Paneth cells were coupled together. The technological advances in high-resolution live cell imaging of epithelial organoids coupled with sensitive signaling reporters and optogenetic modulation of individual cells will help precisely tackle these issues in the future.
In conclusion, active contraction creates an ISC shape that maximizes the lateral signaling area in relation to cellular volume. Physiological aging results in increased ISC size, which can reduce the lateral surface in relation to cell volume and potentially reduces the ISCs’ ability to receive niche signals and manifesting as impaired stem cell capacity (5). We show that the function of old crypt units, containing ISCs and Paneth cells, can be improved by promoting NM IIC contractility or forcing the epithelial contour to youthful dimensions (Fig. 4, G to I). These results provide evidence that cellular shape is an important characteristic of the adult ISC niche in creating optimal signaling platforms. Our findings may explain why ready tissue contour improves intestinal patterning (51) and is consistent with recent reports on cell shape in animal development (3). Future research will be needed to identify which pathways are most dependent on apical constriction of ISCs. Our data on niche-derived signaling indicates that at least Notch and Wnt signaling in ISCs is sensitive to niche contour, and because of their role in supporting the stem cell identity, they may also promote ISC shape with a positive feedback and their own signal reception, as seen in other epithelial systems (52, 53). Whether changes in the cell shape and contact area influence clonal dynamics of ISCs (54), like in other models of cell competition (55), requires further investigation. Moreover, our study suggests a possible role for tumor architecture in interactions between cancer cells and their microenvironment (56, 57). The work here highlights how cell-scale architecture has evolved to maximize cellular functions and underlines tissue and cell geometry as important components of the stem cell niche.
MATERIALS AND METHODS
Animal housing
Lgr5-EGFP-IRES-CreERT2 (12), Vil-CreERT2 (58), Myh9(fl/fl) (59), Rosa26(mTmG/+) (60), or wild-type mice were maintained with a C57BL/6J background and housed in individually ventilated cages (IVC) cages at consistent temperature (19° to 23°C) and humidity (55 ± 10%) under a 12-hour light/12-hour dark cycle. Standard chow and water were accessible ad libitum. Experiments were done with animals aged 3 to 9 months (referred as “young”) and 22 months or older (referred as “old”), and both sexes were used. Animal housing and experiments were performed in accordance with the Finnish National Animal Experimentation Board and Laboratory Animal Resource Center at the University of California, San Francisco.
Organoid culture
Crypts and single cells were isolated from mouse small intestines as described elsewhere (5). Isolated crypts were cultured in 60% Matrigel (Corning) overlaid with Advanced Dulbecco’s modified Eagle’s medium (DMEM)/F12 (Life Technologies) supplemented with 1× GlutaMAX, 1× N2 (Life Technologies), 1× B27 (Life Technologies), 10 mM Hepes, 1× penicillin-streptomycin (Sigma-Aldrich), 1 μM N-acetylcysteine (Sigma-Aldrich), epidermal growth factor (EGF; 50 ng/ml; R&D Systems), noggin (100 ng/ml; PeproTech), and R-spondin1 (500 ng/ml; R&D Systems), referred as ENR. When indicated, 20 μM Y-27632 (Sigma-Aldrich), 10 μM Blebbistatin (Sigma-Aldrich), 10 to 50 nM verteporfin (Tocris Bioscience), 3 μM Chir99021 (Sigma-Aldrich), 1 mM valproic acid (Cayman Chemical), 500 μM 4HAP (Sigma-Aldrich), or corresponding amount of vehicle [dimethyl sulfoxide (DMSO)] was introduced into the ENR medium. Myh9 knockout was induced with 1 μM 4-hydroxytamoxifen (Sigma-Aldrich), and an equal volume of vehicle (ethanol) was used.
Flow cytometry analysis
Organoids were harvested by pipetting, mechanically broken, and pelleted by 30-s centrifugation at 200g. Matrigel was washed with 1 ml of ice-cold Advanced DMEM/F12; cells were pelleted again and incubated for 5 min at 37°C in 100 μl of TrypLE Express (Life Technologies). Suspension was pipetted 10 times to release single cells and washed with 1 ml of ice-cold Advanced DMEM/F12. Cells were centrifuged for 3 min at 300g, followed by antibody staining with 1:500 dilution of anti–CD24–allophycocyanin (APC) or anti–CD24-PacificBlue (both BioLegend, M1/69), anti–Epcam-BV786 (BD, G8.8), or anti–Epcam-APC (eBioscience, G8.8). After washing away unbound antibody, cells were incubated for 10 min in 1:500 solution of SYTOX Blue or 7-aminoactinomycin D (7-AAD) (both Life Technologies) before analysis with FACSAria Fusion or FACSAria II (both BD). Live EGFPhi and EGFPlo were considered as ISCs and TA cells, respectively. Paneth cells were gated as live CD24hiSideScatterhi.
For cell cycle analysis, Lgr5hi cells were isolated as describe elsewhere (5). Lin (CD31, CD45, Ter119)−CD24med/loEGFPhi cells were subdivided on the basis of their forward scatter width to FSClo and FSChi. Sorted cells were fixed with 4% paraformaldehyde (PFA) for 15 min at room temperature, washed with 1 ml of 1% (w/v) bovine serum albumin (BSA)–phosphate-buffered saline (PBS) twice, and resuspended to 4′,6-diamidino-2-phenylindole (DAPI; 10 μg/ml) in 0.1% (v/v) Triton X-100–PBS. Cells were incubated in the dark for 30 min before flow analysis. EGFP+ singlets were gated, and the DNA content was analyzed on the basis of DAPI intensity.
Coculture of fluorescence-activated cell sorting–isolated single cells
Equal number of ISCs Lin−CD24med/loEGFPhi and Paneth cells Lin−EGFPnegCD24hiSSChi cells were mixed and cultured in 60% Matrigel overlaid with ENR media supplemented with extra R-spondin1 (500 ng/ml), Wnt3A (100 ng/ml; R&D Systems), and 1 μM Jagged-1 (AnaSpec). Media contained 10 μM Y-27632 for the first 2 days. Alternatively, for colony formation assay, single ISCs alone were cultured in ENR media supplemented with 10 μM Chir99021 and 10 μM Y-27632. Medium was changed every 2 days, cocultures were quantified on day 6, and colony cultures were quantified on day 5.
Generation and analysis of TOP-GFP organoids
To generate Wnt-reporter intestinal organoid lines (called “TOP-GFP” throughout the text), organoids were transduced with a lentivirus prepared from a pXL010-Wnt dual (GFP-Fire) (Addgene, no. 40588) as previously described in (5). After puromycin selection, organoids were expanded in ENR media containing 3 μM Chir99021 and 1 mM valproic acid. TOP-GFP+ cells were sorted and cultured in ENR + C + VPA until organoids emerged and were manually picked to initiate clonal cultures. Clonal cultures were maintained in ENR + C + VPA and before experiments were passaged and cultured in ENR.
TOP-GFP organoids grown 2 days in ENR were exposed to NM II inhibitors of vehicle (DMSO). After 24 hours, single cells were dissociated and stained with anti–Epcam-BV786, anti–CD24-APC, and SYTOX Blue like above. Wnt response in ISCs was determined as a corrected total fluorescence (CTF) in CD24medSSClo population. CTF was calculated as frequency of TOP-GFP+CD24medSSClo cells multiplied by the mean GFP fluorescence (measured with fluorescein isothiocyanate–A detector) in the positive population. The distinction between negative and positive TOP-GFP signals was determined by comparing signal to wild-type organoids. See fig. S5 for example gating. pXL010-Wnt dual (GFP-Fire) reporter was a gift from S. Palecek (Addgene, plasmid no. 40588; http://n2t.net/addgene:40588; RRID:Addgene_40588) (61).
Immunofluorescent staining
For immunofluorescent staining, organoids were grown on glass-bottom dishes (MatTek Corporation) before fixation with 4% PFA and subsequent staining. Fixed organoids were permeabilized with 0.5% (v/v) Triton X-100 in PBS for 30 min, followed by 30-min blocking in 5% (v/v) normal goat serum–0.2% (w/v) BSA–0.25% (v/v) Triton X-100-PBS. Subsequently, organoids were incubated at 4°C overnight in the blocking buffer containing primary antibodies, rb–anti-lysozyme (DAKO), mo–anti–E-cadherin (BD), rb–anti–phospho-myosin light chain 2 (Ser19; CST), rb–anti-Myh14 (Proteintech), rb–anti-Myh9 (BioLegend), and rb–anti-YAP (D8HX1; CST). The following day, organoids were washed with 0.2% (w/v) BSA in PBS for 2 hours and incubated in the blocking buffer with diluted secondary antibodies (anti–rb–Alexa Fluor 488, anti–rb–Alexa Fluor 546, anti–rb–Alexa Fluor 647, and anti–mo–Alexa Fluor 647) all 1:500 for 2 hours at room temperature or 4°C overnight. Costaining of F-actin and the nuclei was performed through 45-min incubation in phalloidin–Alexa Fluor 647 (1:50; Sigma-Aldrich) or phalloidin-Atto555 (1:500; Sigma-Aldrich) and DAPI (1 μg/ml; Life Technologies) diluted in 0.2% (w/v) BSA-PBS.
Image analysis
Crypt width in live organoids was quantified from images using ImageJ software. Width was measured at the widest point of the crypt bottom where granulated Paneth cells were visible. In stem cell–rich organoids, cultured in ENR supplemented with Chir99021 and valproic acid (CVPA), width was assessed from the crypt bottom, as Paneth cells were absent. In vivo crypt width and density were quantified by drawing a polygon around the crypt from bottom-view images of resected mouse intestine. Feret diameter was used as a crypt width and in calculation of crypt area. Cellular morphology in vitro was measured from confocal sections of organoid crypt domains. Lysozyme-positive cells were considered as Paneth cells. Cells with direct contact with Paneth cells were considered ISCs, and other cells in the crypt domain were designated TA cells. Cell dimensions were measured from apical (the innermost) and basal (the outermost) surface. A line was drawn from the center of the basal side to the center of the apical side and labeled as cell height. At the midpoint of the height, the middle width was measured perpendicular to the lateral membrans. Lateral surface area and volume were calculated by using Eqs. 1 and 2. Ex vivo cell size was analyzed from bright-field images taken from sorted cells that were allowed to settle on the MatTek glass bottom dish. Volume was calculated by using Eq. 3. Gaussian curvature (G) (62) analysis was done on z-stacks of organoids and isolated crypts stained with E-cadherin antibody using the LimeSeg macro for ImageJ (63). The area and Gaussian curvature of a hemisphere were calculated by using Eqs. 4 and 5.
Mathematical equations
Mathematical equations used were as follows
| (1) |
| (2) |
| (3) |
| (4) |
| (5) |
RNA isolation and qPCR
Sorted cells were lysed in TRIzol (Life Technologies), and RNA was isolated according to the manufacturer’s instructions. Isolated RNA was treated with deoxyribonuclease I (Sigma-Aldrich) and then transcribed to cDNA using the RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific) and oligo dT primers according to the manufacturer’s instructions. qPCR was conducted using a Power SYBR Green master mix (Thermo Fisher Scientific) according to the manufacturer’s instructions. Gene expression was evaluated on the basis of the ΔCt method against the housekeeping gene, beta-Actin. The following primers (Sigma-Aldrich) were used: Actin (forward: CCTCTATGCCAACACAGTGC; reverse: CCTGCTTGCTGATCCACATC), Myh9 (forward: AGAAGTTGGTATGGGTGCCTT; reverse: CCCTGAGTAGTATCGCTCCTTG); Myh10 (forward: GGAATCCTTTGGAAATGCGAAGA; reverse: GCCCCAACAATATAGCCAGTTAC), Myh14 (forward: CAGTGACCATGTCCGTGTCTG; reverse: CGTAGAGGAACGATTGGGCTG); Hes1 (forward: CTACCCCAGCCAGTGTCAAC; reverse: ATGCCGGGAGCTATCTTTCT), Olfm4 (forward: ACCACACCTCCAACATCACC; reverse: TAAGCGCTCCACTCTGTCAG), Axin2 (forward: AGTGCAAACTCTCACCCACC; reverse: TCGCTGGATAACTCGCTGTC), cMyc (forward: CAAATCCTGTACCTCGTCCGATTC; reverse: CTTCTTGCTCTTCTTCAGAGTCGC), Sox9 (forward: GAGCCGGATCTGAAGAGGGA; reverse: GCTTGACGTGTGGCTTGTTC); Lgr5 (forward: ACCCGCCAGTCTCCTACATC; reverse: GCATCTAGGCGCAGGGATTG); Mex3a (forward: CAGGCAAGGCTGCAAGATTA; reverse: TCACCATGAACACTGGCTCC), Bmi1 (forward: CCTGGAGAAGAAATGGCCCA; reverse: GGCAAACAGGAAGAGGTGGA), and Lrig1 (forward: TTGAGGACTTGACGAATCTGC; reverse: CTTGTTGTGCTGCAAAAAGAGAG).
Scanning electron microscopy
The collagen scaffold was fixed by immersion in 2.5% glutaraldehyde (Sigma-Aldrich) in 0.1 M phosphate buffer (pH 7.4). The fixed scaffold was then rinsed in 0.1 M phosphate buffer and Milli-Q water before stepwise ethanol dehydration. The scaffold was adhered onto a phosphoenolpyruvate filter support and subjected to critical point drying (Balzer, CPD 010; Liechtenstein). Last, the specimens were mounted on an aluminum pin stub with adhesive carbon tape and sputter-coated with a thin layer of platinum (Quorum, Q150T ES). SE2 images were acquired using an Ultra 55 field-emission scanning electron microscope (Zeiss, Oberkochen, Germany) at 3 kV.
Fabrication of a polydimethylsiloxane substrate for an intestinal crypt model
Polymer-based intestine crypt models were fabricated by standard soft lithography and replica molding processes (64, 65). For the molding process, glass slides (75 mm by 50 mm; Sigma-Aldrich) were used as a substrate. Since SU-8 photoresist adheres poorly to glass, a thin layer of a negative photoresist, SU-8 2 (thickness, ~10 μm; MicroChem) was spin-coated as an adhesive layer first at 1000 rpm for 30 s (66). The photoresist was soft-baked on a hot plate at 65°C for 1 min and at 95°C for 2 min before ultraviolet (UV) light exposure for 2 min. The photoresist was then postbaked on a hot plate at 65°C for 1 min and 95°C for 2 min. SU-8 2100 (MicroChem) was spin-coated onto the adhesive layer at 1500 rpm for 30 s, yielding a layer with a thickness of approximately 200 μm. The SU-8 layer was soft-baked on a hot plate at 65°C for 3 min and 100°C for 15 min. A photomask with one of four potential circle patterns—diameter of 50, 75, 100, and 125 μm—and 100-μm edge-to-edge spacing (designated D50, D75, D100, and D125, respectively) was placed on top of the SU-8 layer. Then, the SU-8 layer was exposed to UV light through the photomask at 10 mW cm−2 for 20 s (for D50 and D75) and 40 s (for D100 and D125). We intentionally applied long exposure times to allow for the diffusion of photoactive components and obtain rounded edges after development. In addition, after exposure, the SU-8 substrate was incubated for 15 min at room temperature to allow for the diffusion of photoacid (67), further assisting the formation of curved edges. Next, the patterned SU-8 layer was postbaked on a hot plate at 65°C for 5 min and 100°C for 20 min. After cooling to room temperature, the SU-8 layer was developed in propylene glycol monomethyl ether acetate (Sigma-Aldrich) for 7 min (for D50 and D75) or 9 min (for D100 and D125), rinsed with isopropanol alcohol (IPA; Sigma-Aldrich), and dried with compressed nitrogen. The resulting SU-8 master pattern was treated with oxygen plasma for 30 min followed by silanization, a surface treatment with trimethylchlorosilane (Sigma-Aldrich) in a vacuum chamber overnight for casting. This SU-8 master pattern was then cast with a mixture of polydimethylsiloxane (PDMS; SYLGARD 184, Dow Corning) and curing agent at 10:1 weight ratio. The mixture was degassed in a vacuum chamber for 10 min and cured overnight at room temperature. After curing, the patterned PDMS was peeled off from the SU-8 master pattern (PDMS stamp). The patterned PDMS was treated with the same silanization process as the SU-8 master pattern, and a new PDMS sample was cast using the first PDMS sample as a primary mold. Last, PDMS intestinal crypts were peeled off from the PDMS primary and cleaned with ethanol, IPA, and Milli-Q water in a sonication bath.
Preparation of collagen scaffold
To improve the collagen integrity, collagen was cross-linked with 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide hydrochloride (EDC) and N-hydroxysuccinimide (NHS) at pH 5, as previously described (68). Collagen solution (in 0.02 M acetic acid) was lyophilized for 72 hours to remove water and acetic acid. The lyophilized collagen was dissolved in MES buffer (0.1 M; pH 5) at a concentration of 5 mg/ml. A collagen scaffold was micromolded and cross-linked on the surface of a porous polytetrafluoroethylene (PTFE) membrane (0.4-μm pore size) held in a 24-well insert (Merck Millipore). A mixture of collagen (4 mg/ml), 60 mM EDC, and 15 mM NHS in MES buffer (0.1 M; pH 5) was prepared, and bubbles were removed by centrifugation at 3000g for 1 min. The collagen mixture (60 μl) was added to the center of the insert, and the PDMS stamp was placed on top of the mixture for 2 hours. The PDMS stamp was then removed from the collagen scaffold, and the scaffold was then soaked in PBS for 24 hours to remove unincorporated EDC and NHS. The following day, the collagen scaffold was coated overnight with laminin-521 (20 μg/ml), laminin-111 (BioLamina), fibronectin (50 μg/ml; Thermo Fisher Scientific), and 10% (v/v) Matrigel.
Culture of isolated cells or organoids on collagen scaffolds
Ten thousand sorted Lgr5hi cells were cultured on each scaffold for 1 week in ENR media supplemented with 10 μM Y-27632, R-spondin1 (500 ng/ml), Wnt3A (100 ng/ml), 3 μM Chi99021, 1 mM valproic acid, 1 μM Jagged-1 (tebu-bio; stem cell media) followed by a release period in ENR + 10 μM Y-27632 for 2 days and in plain ENR for 2 weeks (differentiation media). When seeding organoids, isolated crypts were cultured for 2 days in three-dimensional (3D) Matrigel (see the “Organoid culture” section), followed by washing and plating on collagen scaffolds. Areas of the colonies were quantified as indicated in the figure legends.
RNA sequencing and analysis
RNA was isolated and sequenced from sorted Lgr5lo simultaneously with Lgr5hi cells, as described by Pentinmikko et al. (5). The fastq files from Lgr5lo samples were combined with Lgr5hi from the work of Pentinmikko et al., and adapter and low-quality sequences were trimmed from the data using Trimmomatic (69) and checked for quality using FastQC (70). The trimmed reads were subsequently aligned to the Mus musculus transcriptome version 6.24 and counted using Salmon version 0.14 (71). The counted transcripts were then transformed into gene counts using the tximportData package for R version 3.6 (72). To correct for potential batch effects, because of differences in mouse origin and other experimental variation, a batch correction analysis and data preparation for DESeq analysis were performed using the R package sva (73). This yielded improved clustering within sample groups based on age and cell type as visualized through principal components analysis in R version 3.6. Differential gene expression after batch correction between the Lgr5hi old versus young cells was estimated using the DESeq2 package for R (74). Differentially expressed genes that were identified to be statistically significant (false discovery rate < 0.05) were used for gene set enrichment analysis on the GenePattern platform (75, 76) based on Gene Ontology and Reactome identified gene sets (77, 78).
Statistical methods
A two-tailed Student’s t test was used to compare groups when indicated. P value is represented in the corresponding figure panels. P < 0.05 was considered significant. A paired t test was used when indicated in the figure legend to analyze effects of treatments between independent biological replicates. For statistical methods of RNA sequencing, see the “RNA sequencing and analysis” section. Microsoft Excel v16 and GraphPad Prism 8 were used for statistical analyses.
Acknowledgments
We thank M. Simula and J. Bärlund for technical assistance and the whole Katajisto Laboratory for commenting on the manuscript. Imaging was performed at the Light Microscopy Unit of Institute of Biotechnology and at the Live Cell Imaging Unit/Nikon Center of Excellence at the Karolinska Institute.
Funding: This work was supported by the Academy of Finland Centre of Excellence MetaStem (266869, 304591, and 320185 to P.K., N.P., R.L., S.S., J.I.E., S.A., A.S.C., and M.B.), ERC grants 677809, 101045009 (to P.K., N.P., R.L., S.S., J.I.E., S.A., A.S.C., and M.B.), Swedish Research Council 2018-03078 (to P.K., N.P., R.L., S.S., J.I.E., S.A., A.S.C., and M.B.), the EU/EFPIA/OICR/McGill/KTH/Diamond Innovative Medicines Initiative 2 Joint Undertaking (EUbOPEN grant no. 875510 to M.S. and P.K.), the EU/EFPIA Innovative Medicines Initiative Joint Undertaking (ULTRA-DD grant no. 115766 to M.S. and P.K.), the Wenner-Gren Foundation (UPD2017-0254, UPD2018-0269 to M.M.S. and A.T.S.), Vetenskapsrådet (VR 2015-02904 to M.M.S. and A.T.S.), and NIH R35-DE026602 (to O.D.K.).
Author contributions: N.P. and P.K. designed the study. N.P., R.L., S.S., J.I.E., S.A., D.C.-A., A.G., and A.S.C. performed the experiments. M.B. analyzed the RNA sequencing data. R.L. engineered the scaffolds with the help of K.-Y.S., A.T.S., M.S., M.M.S., N.A., and A.T. O.D.K. provided reagents and commented on the manuscript. N.P., R.L., and P.K. interpreted the data and wrote the manuscript.
Competing interests: D.C.-A. is an employee of Genentech Inc. and owns shares of Roche. N.A. owns shares of Altis Biosystems Inc. All other authors declare that they have no competing interests.
Data and materials availability: All data needed to evaluate the conclusions in this paper are present in the paper and/or the Supplementary Materials. RNA sequencing data are publicly available at ArrayExpress (E-MTAB-12122).
Supplementary Materials
This PDF file includes:
Figs. S1 to S6
Other Supplementary Material for this manuscript includes the following:
Tables S1 and S2
REFERENCES AND NOTES
- 1.Chacon-Martinez C. A., Koester J., Wickstrom S. A., Signaling in the stem cell niche: Regulating cell fate, function and plasticity. Development 145, dev165399 (2018). [DOI] [PubMed] [Google Scholar]
- 2.Nelson C. M., Bissell M. J., Of extracellular matrix, scaffolds, and signaling: Tissue architecture regulates development, homeostasis, and cancer. Annu. Rev. Cell Dev. Biol. 22, 287–309 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Guignard L., Fiúza U.-M., Leggio B., Laussu J., Faure E., Michelin G., Biasuz K., Hufnagel L., Malandain G., Godin C., Lemaire P., Contact area-dependent cell communication and the morphological invariance of ascidian embryogenesis. Science 369, eaar5663 (2020). [DOI] [PubMed] [Google Scholar]
- 4.Barker N., Adult intestinal stem cells: Critical drivers of epithelial homeostasis and regeneration. Nat. Rev. Mol. Cell Biol. 15, 19–33 (2014). [DOI] [PubMed] [Google Scholar]
- 5.Pentinmikko N., Iqbal S., Mana M., Andersson S., Cognetta A. B. III, Suciu R. M., Roper J., Luopajärvi K., Markelin E., Gopalakrishnan S., Smolander O.-P., Naranjo S., Saarinen T., Juuti A., Pietiläinen K., Auvinen P., Ristimäki A., Gupta N., Tammela T., Jacks T., Sabatini D. M., Cravatt B. F., Yilmaz Ö. H., Katajisto P., Notum produced by Paneth cells attenuates regeneration of aged intestinal epithelium. Nature 571, 398–402 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Yilmaz O. H., Katajisto P., Lamming D. W., Gültekin Y., Bauer-Rowe K. E., Sengupta S., Birsoy K., Dursun A., Yilmaz V. O., Selig M., Nielsen G. P., Mino-Kenudson M., Zukerberg L. R., Bhan A. K., Deshpande V., Sabatini D. M., mTORC1 in the Paneth cell niche couples intestinal stem-cell function to calorie intake. Nature 486, 490–495 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Sato T., van Es J. H., Snippert H. J., Stange D. E., Vries R. G., van den Born M., Barker N., Shroyer N. F., van de Wetering M., Clevers H., Paneth cells constitute the niche for Lgr5 stem cells in intestinal crypts. Nature 469, 415–418 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sato T., Vries R. G., Snippert H. J., van de Wetering M., Barker N., Stange D. E., van Es J. H., Abo A., Kujala P., Peters P. J., Clevers H., Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009). [DOI] [PubMed] [Google Scholar]
- 9.Clevers H., The intestinal crypt, a prototype stem cell compartment. Cell 154, 274–284 (2013). [DOI] [PubMed] [Google Scholar]
- 10.Shoshkes-Carmel M., Wang Y. J., Wangensteen K. J., Tóth B., Kondo A., Massasa E. E., Itzkovitz S., Kaestner K. H., Subepithelial telocytes are an important source of Wnts that supports intestinal crypts. Nature 557, 242–246 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.McCarthy N., Manieri E., Storm E. E., Saadatpour A., Luoma A. M., Kapoor V. N., Madha S., Gaynor L. T., Cox C., Keerthivasan S., Wucherpfennig K., Yuan G.-C., de Sauvage F. J., Turley S. J., Shivdasani R. A., Distinct mesenchymal cell populations generate the essential intestinal BMP signaling gradient. Cell Stem Cell 26, 391–402.e5 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Barker N., van Es J. H., Kuipers J., Kujala P., van den Born M., Cozijnsen M., Haegebarth A., Korving J., Begthel H., Peters P. J., Clevers H., Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007). [DOI] [PubMed] [Google Scholar]
- 13.Sumigray K. D., Terwilliger M., Lechler T., Morphogenesis and compartmentalization of the intestinal crypt. Dev. Cell 45, 183–197.e5 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Vicente-Manzanares M., Ma X., Adelstein R. S., Horwitz A. R., Non-muscle myosin II takes centre stage in cell adhesion and migration. Nat. Rev. Mol. Cell Biol. 10, 778–790 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Yang Q., Xue S.-L., Chan C. J., Rempfler M., Vischi D., Maurer-Gutierrez F., Hiiragi T., Hannezo E., Liberali P., Cell fate coordinates mechano-osmotic forces in intestinal crypt formation. Nat. Cell Biol. 23, 733–744 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Yin X., Farin H. F., van Es J. H., Clevers H., Langer R., Karp J. M., Niche-independent high-purity cultures of Lgr5+ intestinal stem cells and their progeny. Nat. Methods 11, 106–112 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Straight A. F., Cheung A., Limouze J., Chen I., Westwood N. J., Sellers J. R., Mitchison T. J., Dissecting temporal and spatial control of cytokinesis with a myosin II Inhibitor. Science 299, 1743–1747 (2003). [DOI] [PubMed] [Google Scholar]
- 18.Uehata M., Ishizaki T., Satoh H., Ono T., Kawahara T., Morishita T., Tamakawa H., Yamagami K., Inui J., Maekawa M., Narumiya S., Calcium sensitization of smooth muscle mediated by a Rho-associated protein kinase in hypertension. Nature 389, 990–994 (1997). [DOI] [PubMed] [Google Scholar]
- 19.Perez-Gonzalez C., Ceada G., Greco F., Matejčić M., Gómez-González M., Castro N., Menendez A., Kale S., Krndija D., Clark A. G., Gannavarapu V. R., Álvarez-Varela A., Roca-Cusachs P., Batlle E., Vignjevic D. M., Arroyo M., Trepat X., Mechanical compartmentalization of the intestinal organoid enables crypt folding and collective cell migration. Nat. Cell Biol. 23, 745–757 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hansen C. G., Moroishi T., Guan K. L., YAP and TAZ: A nexus for Hippo signaling and beyond. Trends Cell Biol. 25, 499–513 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Totaro A., Panciera T., Piccolo S., YAP/TAZ upstream signals and downstream responses. Nat. Cell Biol. 20, 888–899 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Barry E. R., Morikawa T., Butler B. L., Shrestha K., de la Rosa R., Yan K. S., Fuchs C. S., Magness S. T., Smits R., Ogino S., Kuo C. J., Camargo F. D., Restriction of intestinal stem cell expansion and the regenerative response by YAP. Nature 493, 106–110 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Gregorieff A., Liu Y., Inanlou M. R., Khomchuk Y., Wrana J. L., Yap-dependent reprogramming of Lgr5+ stem cells drives intestinal regeneration and cancer. Nature 526, 715–718 (2015). [DOI] [PubMed] [Google Scholar]
- 24.Serra D., Mayr U., Boni A., Lukonin I., Rempfler M., Challet Meylan L., Stadler M. B., Strnad P., Papasaikas P., Vischi D., Waldt A., Roma G., Liberali P., Self-organization and symmetry breaking in intestinal organoid development. Nature 569, 66–72 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wang C., Zhu X., Feng W., Yu Y., Jeong K., Guo W., Lu Y., Mills G. B., Verteporfin inhibits YAP function through up-regulating 14-3-3σ sequestering YAP in the cytoplasm. Am. J. Cancer Res. 6, 27–37 (2016). [PMC free article] [PubMed] [Google Scholar]
- 26.Nikolaev M., Mitrofanova O., Broguiere N., Geraldo S., Dutta D., Tabata Y., Elci B., Brandenberg N., Kolotuev I., Gjorevski N., Clevers H., Lutolf M. P., Homeostatic mini-intestines through scaffold-guided organoid morphogenesis. Nature 585, 574–578 (2020). [DOI] [PubMed] [Google Scholar]
- 27.Verhulsel M., Simon A., Bernheim-Dennery M., Gannavarapu V. R., Gérémie L., Ferraro D., Krndija D., Talini L., Viovy J.-L., Vignjevic D. M., Descroix S., Developing an advanced gut on chip model enabling the study of epithelial cell/fibroblast interactions. Lab Chip 21, 365–377 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li Q., Rycaj K., Chen X., Tang D. G., Cancer stem cells and cell size: A causal link? Semin. Cancer Biol. 35, 191–199 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Berardi A. C., Wang A., Levine J. D., Lopez P., Scadden D. T., Functional isolation and characterization of human hematopoietic stem cells. Science 267, 104–108 (1995). [DOI] [PubMed] [Google Scholar]
- 30.Watt F. M., Green H., Involucrin synthesis is correlated with cell size in human epidermal cultures. J. Cell Biol. 90, 738–742 (1981). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lengefeld J., Cheng C.-W., Maretich P., Blair M., Hagen H., McReynolds M. R., Sullivan E., Majors K., Roberts C., Kang J. H., Steiner J. D., Miettinen T. P., Manalis S. R., Antebi A., Morrison S. J., Lees J. A., Boyer L. A., Yilmaz Ö. H., Amon A., Cell size is a determinant of stem cell potential during aging. Sci. Adv. 7, eabk0271 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mumm J. S., Kopan R., Notch signaling: From the outside in. Dev. Biol. 228, 151–165 (2000). [DOI] [PubMed] [Google Scholar]
- 33.Mihaylova M. M., Cheng C.-W., Cao A. Q., Tripathi S., Mana M. D., Bauer-Rowe K. E., Abu-Remaileh M., Clavain L., Erdemir A., Lewis C. A., Freinkman E., Dickey A. S., La Spada A. R., Huang Y., Bell G. W., Deshpande V., Carmeliet P., Katajisto P., Sabatini D. M., Yilmaz Ö. H., Fasting activates fatty acid oxidation to enhance intestinal stem cell function during homeostasis and aging. Cell Stem Cell 22, 769–778.e4 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nalapareddy K., Nattamai K. J., Kumar R. S., Karns R., Wikenheiser-Brokamp K. A., Sampson L. L., Mahe M. M., Sundaram N., Yacyshyn M.-B., Yacyshyn B., Helmrath M. A., Zheng Y., Geiger H., Canonical Wnt signaling ameliorates aging of intestinal stem cells. Cell Rep. 18, 2608–2621 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Surcel A., Schiffhauer E. S., Thomas D. G., Zhu Q., DiNapoli K. T., Herbig M., Otto O., West-Foyle H., Jacobi A., Kräter M., Plak K., Guck J., Jaffee E. M., Iglesias P. A., Anders R. A., Robinson D. N., Targeting mechanoresponsive proteins in pancreatic cancer: 4-hydroxyacetophenone blocks dissemination and invasion by activating MYH14. Cancer Res. 79, 4665–4678 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Farin H. F., Karthaus W. R., Kujala P., Rakhshandehroo M., Schwank G., Vries R. G. J., Kalkhoven E., Nieuwenhuis E. E. S., Clevers H., Paneth cell extrusion and release of antimicrobial products is directly controlled by immune cell-derived IFN-γ. J. Exp. Med. 211, 1393–1405 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Surcel A., Ng W. P., West-Foyle H., Zhu Q., Ren Y., Avery L. B., Krenc A. K., Meyers D. J., Rock R. S., Anders R. A., Freel Meyers C. L., Robinson D. N., Pharmacological activation of myosin II paralogs to correct cell mechanics defects. Proc. Natl. Acad. Sci. U.S.A. 112, 1428–1433 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Amodeo A. A., Skotheim J. M., Cell-size control. Cold Spring Harb. Perspect. Biol. 8, a019083 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.McBeath R., Pirone D. M., Nelson C. M., Bhadriraju K., Chen C. S., Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev. Cell 6, 483–495 (2004). [DOI] [PubMed] [Google Scholar]
- 40.Bao M., Xie J., Piruska A., Huck W. T. S., 3D microniches reveal the importance of cell size and shape. Nat. Commun. 8, 1962 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Vasilevich A. S., Vermeulen S., Kamphuis M., Roumans N., Eroumé S., Hebels D. G. A. J., van de Peppel J., Reihs R., Beijer N. R. M., Carlier A., Carpenter A. E., Singh S., de Boer J., On the correlation between material-induced cell shape and phenotypical response of human mesenchymal stem cells. Sci. Rep. 10, 18988 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kilian K. A., Bugarija B., Lahn B. T., Mrksich M., Geometric cues for directing the differentiation of mesenchymal stem cells. Proc. Natl. Acad. Sci. U.S.A. 107, 4872–4877 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Engler A. J., Sen S., Sweeney H. L., Discher D. E., Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006). [DOI] [PubMed] [Google Scholar]
- 44.McKinley K. L., Stuurman N., Royer L. A., Schartner C., Castillo-Azofeifa D., Delling M., Klein O. D., Vale R. D., Cellular aspect ratio and cell division mechanics underlie the patterning of cell progeny in diverse mammalian epithelia. eLife 7, (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Li Y., Chen M., Hu J., Sheng R., Lin Q., He X., Guo M., Volumetric compression induces intracellular crowding to control intestinal organoid growth via Wnt/β-catenin signaling. Cell Stem Cell 28, 63–78.e7 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Neurohr G. E., Terry R. L., Lengefeld J., Bonney M., Brittingham G. P., Moretto F., Miettinen T. P., Vaites L. P., Soares L. M., Paulo J. A., Harper J. W., Buratowski S., Manalis S., van Werven F. J., Holt L. J., Amon A., Excessive cell growth causes cytoplasm dilution and contributes to senescence. Cell 176, 1083–1097.e18 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sakamori R., das S., Yu S., Feng S., Stypulkowski E., Guan Y., Douard V., Tang W., Ferraris R. P., Harada A., Brakebusch C., Guo W., Gao N., Cdc42 and Rab8a are critical for intestinal stem cell division, survival, and differentiation in mice. J. Clin. Invest. 122, 1052–1065 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Shaya O., Binshtok U., Hersch M., Rivkin D., Weinreb S., Amir-Zilberstein L., Khamaisi B., Oppenheim O., Desai R. A., Goodyear R. J., Richardson G. P., Chen C. S., Sprinzak D., Cell-cell contact area affects notch signaling and notch-dependent patterning. Dev. Cell 40, 505–511.e6 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Gomez-Galvez P., Vicente-Munuera P., Tagua A., Forja C., Castro A. M., Letrán M., Valencia-Expósito A., Grima C., Bermúdez-Gallardo M., Serrano-Pérez-Higueras Ó., Cavodeassi F., Sotillos S., Martín-Bermudo M. D., Márquez A., Buceta J., Escudero L. M., Scutoids are a geometrical solution to three-dimensional packing of epithelia. Nat. Commun. 9, 2960 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Giangreco A., Goldie S. J., Failla V., Saintigny G., Watt F. M., Human skin aging is associated with reduced expression of the stem cell markers beta1 integrin and MCSP. J. Invest. Dermatol. 130, 604–608 (2010). [DOI] [PubMed] [Google Scholar]
- 51.Gjorevski N., Nikolaev M., Brown T. E., Mitrofanova O., Brandenberg N., Del Rio F. W., Yavitt F. M., Liberali P., Anseth K. S., Lutolf M. P., Tissue geometry drives deterministic organoid patterning. Science 375, eaaw9021 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kozlovskaja-Gumbriene A., Yi R., Alexander R., Aman A., Jiskra R., Nagelberg D., Knaut H., Clain M. M., Piotrowski T., Proliferation-independent regulation of organ size by Fgf/Notch signaling. eLife 6, e21049 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Fumoto K., Takigawa-Imamura H., Sumiyama K., Kaneiwa T., Kikuchi A., Modulation of apical constriction by Wnt signaling is required for lung epithelial shape transition. Development 144, 151–162 (2017). [DOI] [PubMed] [Google Scholar]
- 54.Snippert H. J., van der Flier L. G., Sato T., van Es J. H., van den Born M., Kroon-Veenboer C., Barker N., Klein A. M., van Rheenen J., Simons B. D., Clevers H., Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010). [DOI] [PubMed] [Google Scholar]
- 55.Levayer R., Hauert B., Moreno E., Cell mixing induced by myc is required for competitive tissue invasion and destruction. Nature 524, 476–480 (2015). [DOI] [PubMed] [Google Scholar]
- 56.Messal H. A., Alt S., Ferreira R. M. M., Gribben C., Wang V. M.-Y., Cotoi C. G., Salbreux G., Behrens A., Tissue curvature and apicobasal mechanical tension imbalance instruct cancer morphogenesis. Nature 566, 126–130 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Fiore V. F., Krajnc M., Quiroz F. G., Levorse J., Pasolli H. A., Shvartsman S. Y., Fuchs E., Mechanics of a multilayer epithelium instruct tumour architecture and function. Nature 585, 433–439 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.el Marjou F., Janssen K.-P., Hung-Junn Chang B., Li M., Hindie V., Chan L., Louvard D., Chambon P., Metzger D., Robine S., Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004). [DOI] [PubMed] [Google Scholar]
- 59.Jacobelli J., Friedman R. S., Conti M. A., LennonDumenil A.-M., Piel M., Sorensen C. M., Adelstein R. S., Krummel M. F., Confinement-optimized three-dimensional T cell amoeboid motility is modulated via myosin IIA-regulated adhesions. Nat. Immunol. 11, 953–961 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Muzumdar M. D., Tasic B., Miyamichi K., Li L., Luo L., A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007). [DOI] [PubMed] [Google Scholar]
- 61.Azarin S. M., Lian X., Larson E. A., Popelka H. M., de Pablo J. J., Palecek S. P., Modulation of Wnt/β-catenin signaling in human embryonic stem cells using a 3-D microwell array. Biomaterials 33, 2041–2049 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.E. Kreyszig, Differential Geometry (Dover Publications, 1991). [Google Scholar]
- 63.Machado S., Mercier V., Chiaruttini N., LimeSeg: A coarse-grained lipid membrane simulation for 3D image segmentation. BMC Bioinformatics 20, 2 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Xia Y., Whitesides G. M., Soft lithography. Angew. Chem. Int. Ed. Engl. 37, 550–575 (1998). [DOI] [PubMed] [Google Scholar]
- 65.Qin D., Xia Y., Whitesides G. M., Soft lithography for micro- and nanoscale patterning. Nat. Protoc. 5, 491–502 (2010). [DOI] [PubMed] [Google Scholar]
- 66.Liu J. S., Song D., Zong G., Yin P., Zhang X., Xu Z., Du L., Liu C., Wang L., Fabrication of SU-8 moulds on glass substrates by using a common thin negative photoresist as an adhesive layer. J. Micromech. Microeng. 24, 035009 (2014). [Google Scholar]
- 67.Chang S. I., Yoon J. B., Shape-controlled, high fill-factor microlens arrays fabricated by a 3D diffuser lithography and plastic replication method. Opt. Express 12, 6366–6371 (2004). [DOI] [PubMed] [Google Scholar]
- 68.Wang Y., Gunasekara D. B., Reed M. I., DiSalvo M., Bultman S. J., Sims C. E., Magness S. T., Allbritton N. L., A microengineered collagen scaffold for generating a polarized crypt-villus architecture of human small intestinal epithelium. Biomaterials 128, 44–55 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Bolger A. M., Lohse M., Usadel B., Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.S. Andrews, (2010). FastQC: A Quality Control Tool for High Throughput Sequence Data [Online]. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
- 71.Patro R., Duggal G., Love M. I., Irizarry R. A., Kingsford C., Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Soneson C., Love M. I., Robinson M. D., Differential analyses for RNA-seq: Transcript-level estimates improve gene-level inferences. F1000Res 4, 1521 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.J. T. Leek, W. E. Johnson, H. S. Parker, E. J. Fertig, A. E. Jaffe, J. D. Storey, Y. Zhang, L. C. Torres, vol. R package version 3.34.0 (2017).
- 74.Love M. I., Huber W., Anders S., Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Reich M., Liefeld T., Gould J., Lerner J., Tamayo P., Mesirov J. P., GenePattern 2.0. Nat. Genet. 38, 500–501 (2006). [DOI] [PubMed] [Google Scholar]
- 76.Subramanian A., Tamayo P., Mootha V. K., Mukherjee S., Ebert B. L., Gillette M. A., Paulovich A., Pomeroy S. L., Golub T. R., Lander E. S., Mesirov J. P., Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U.S.A. 102, 15545–15550 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Croft D., O’Kelly G., Wu G., Haw R., Gillespie M., Matthews L., Caudy M., Garapati P., Gopinath G., Jassal B., Jupe S., Kalatskaya I., Mahajan S., May B., Ndegwa N., Schmidt E., Shamovsky V., Yung C., Birney E., Hermjakob H., D’Eustachio P., Stein L., Reactome: A database of reactions, pathways and biological processes. Nucleic Acids Res. 39, D691–D697 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Harris M. A., Clark J., Ireland A., Lomax J., Ashburner M., Foulger R., Eilbeck K., Lewis S., Marshall B., Mungall C., Richter J., Rubin G. M., Blake J. A., Bult C., Dolan M., Drabkin H., Eppig J. T., Hill D. P., Ni L., Ringwald M., Balakrishnan R., Cherry J. M., Christie K. R., Costanzo M. C., Dwight S. S., Engel S., Fisk D. G., Hirschman J. E., Hong E. L., Nash R. S., Sethuraman A., Theesfeld C. L., Botstein D., Dolinski K., Feierbach B., Berardini T., Mundodi S., Rhee S. Y., Apweiler R., Barrell D., Camon E., Dimmer E., Lee V., Chisholm R., Gaudet P., Kibbe W., Kishore R., Schwarz E. M., Sternberg P., Gwinn M., Hannick L., Wortman J., Berriman M., Wood V., de la Cruz N., Tonellato P., Jaiswal P., Seigfried T., White R.; Gene Ontology Consortium , The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 32, D258–D261 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figs. S1 to S6
Tables S1 and S2