Abstract
G-typing of rotavirus strains enables the study of molecular epidemiology and gathering of information to promote disease prevention and control. Rotavirus strains in fecal specimens from neonatal calves in Swedish cattle herds were therefore characterized by using G1 to -4-, G6-, G8-, and G10-specific primers in reverse transcription (RT)-PCR. Fecal samples were collected from one dairy herd (herd A) for 4 consecutive years and from 41 beef and dairy herds (herd B) experiencing calf diarrhea outbreaks. Altogether, 1,700 samples were analyzed by group A rotavirus enzyme-linked immunosorbent assay, and 98 rotavirus-positive specimens were selected for G-typing by RT-PCR. The effect of herd type, time, geographic region, and clinical symptoms on the G-type distribution was evaluated. Altogether (herds A and B), G10 was found in 59 (60.2%) fecal specimens, G6 was found in 30 (30.6%) specimens, G3 was found in 1 (1.0%) specimen, and G8 was found in 1 (1.0%) specimen. Seven (7.1%) fecal specimens were not typeable. Herd type specificity in the G-type distribution was demonstrated in the herds in herd B. In the 6 beef suckler herds, only G6 was detected, while rotavirus strains from the 35 dairy herds were predominantly (54%) G10. The G-type distribution was restricted in herds A and B. Twenty-nine of 30 strains from herd A were characterized as G10. In the vast majority of herds in herd B, a single G-type was identified. The serotype G10 and the electropherotype persisted over time in herd A. No characteristic G-type variation in the geographic distribution of cattle herds in herd B was obvious. There was no difference in the G-type distributions between the strains from clinically and subclinically rotavirus-infected calves in dairy herd A. The results from this study strongly indicate a pronounced stability in the rotavirus G-type distribution in Swedish cattle herds, which emphasizes the importance of continuous preventive measures for control of neonatal calf diarrhea. A future bovine rotavirus vaccine in Sweden should contain G10 and G6 strains.
Neonatal enteritis caused by bovine group A rotavirus (BRV) is a common and costly disease in cattle populations worldwide. In Sweden, a clear association between infections with BRV and calf diarrhea (6a) and reduction of weight gain in diarrheic calves has been demonstrated (6b). Control of neonatal diarrhea caused by BRV is based on prophylaxis. Useful information for vaccine development and other prevention strategies may be gained by insight into the BRV molecular epidemiology (e.g., the serotype distribution). BRV strains are classified into G- and P-types according to the two type-specific, outer capsid proteins VP7 and VP4 (7). Serotype-specific classification is based on neutralization tests, but other assays are also available for typing, including monoclonal antibodies in enzyme-linked immunosorbent assays (ELISAs) nucleic acid hybridization, reverse transcription-PCR (RT-PCR), and nucleotide sequence analysis (for review, see reference 11).
Significant information about the distribution of rotavirus serotypes in different species has been obtained, especially about the G-types. Out of 14 identified rotavirus G-types, at least 4 epidemiologically important BRV G-types (G1, G6, G8, and G10) have been described in cattle populations (2, 22). The G6 and G10 types are regarded as the main types, while G1 and G8 are less common. The occurrence of G2, G3, G7, and G11 BRV strains has also been reported (4, 12, 13). The distribution of BRV serotypes has been suggested to be associated with herd type, region, management conditions (1, 17, 18), clinical symptoms (14), and calf age (6).
In Sweden, the cattle herd structure comprises approximately 31,000 herds, equally divided between dairy and suckled beef herds. Herds are comparatively small; the average size of dairy herds is 28 cows, and that of beef cattle herds is 10 cows. The cattle population in the northern regions of Sweden has a sparse density (100 cows/km2 of grazed land), while the density in the south is three times as high (Statistics Sweden 1998).
The aim of this study was to evaluate for the first time the effects of time, herd type, geographic region, and clinical symptoms on the BRV G-type distribution in Swedish cattle herds, in order to gather epidemiological information to be used in disease prevention and control.
MATERIALS AND METHODS
Field samples.
Fecal samples were collected from (i) a single dairy herd on a long-term basis and (ii) 41 different herds experiencing calf diarrhea outbreaks.
Herd A.
Fecal sampling was performed for 4 consecutive years (1993 to 1996) in a large, closed dairy herd, as previously described (6b). In brief, at 4, 14, and 28 days of age, calves were sampled on a regular basis by farm personnel. In addition, fecal samples were also collected on the day of onset from all diarrheic calves, up to 31 days of age. All 1,400 fecal samples were submitted to the National Veterinary Institute (SVA) by mail. From each sample, a 10% fecal slurry was prepared in 0.9% NaCl and centrifuged at 1,000 × g for 5 min, after which the supernatant was collected. The clarified fecal specimens were stored at −20°C before analysis or examined directly by a group A rotavirus ELISA (23, 24). A total of 104 fecal specimens had an absorbance value (optical density) equal to or more than 0.10 and were thus considered BRV positive. Out of these 104 fecal specimens, 30 were selected for G-typing. The selection of fecal specimens for G-typing was performed to comprise symptomatic as well as asymptomatic infections (15 specimens originating from diarrheic calves and 15 from nondiarrheic calves) and also to cover the entire sampling time period.
Herd B.
Roughly 300 fecal samples from diarrheic calves were collected by veterinary practitioners in 1992 to 1997 and submitted to SVA for enteropathogenic agent analysis. The samples were delivered directly to the laboratory or submitted by mail. Preparation of fecal slurries and examination and storage of the fecal specimens for BRV analysis were performed as for herd A. A total of 68 fecal specimens, originating from calves in 35 dairy herds and 6 suckled beef herds, were tested positive by group A rotavirus ELISA (23, 24). Each herd contributed either 1 (24 herds), 2 (10 herds), 3 (4 herds), or 4 (3 herds) BRV-positive fecal samples. All 68 BRV-positive fecal specimens were analyzed by G-type RT-PCR.
Reference strains.
The BRV reference strains UK (G6), 678 (G8), and B223 (G10) and the rotavirus reference strains Wa (G1), DS-1 (G2), and RRV (G3) were used as controls in the RT-PCR assays. The BRV reference strain NCDV (G6) was used in the RNA polyacrylamide gel electrophoresis (RNA-PAGE). All reference strains had been cultured in MA 104 cells.
Extraction of dsRNA.
Extraction of double-stranded RNA (dsRNA) from the fecal specimens and the reference strains was performed by a modification of the previously published guanidinium thiocyanate and silica extraction method (3). Briefly, 50 μl of 10% clarified fecal suspension was mixed and incubated for 15 min with 10 μl of silica and 500 μl of a buffer consisting of a mixture of 120 g of guanidinium thiocyanate (GTC), 100 ml of 0.1 M Tris-HCl (pH 6.4), 22 ml of 0.2 M EDTA (pH 8.0), and 2.6 g of Triton X-100 (KEBO Lab, Stockholm, Sweden). The silica was pelleted by centrifugation and washed twice with a buffer consisting of 120 g of GTC and 100 ml of 0.1 M Tris-HCl (pH 6.4), twice with 70% ethanol, and once with acetone. The pellet was dried at 56°C for 15 min. The dsRNA was then eluted with 25 μl of distilled water (dH2O) and incubated at 56°C for 15 min, which was followed by centrifugation of the pellet and collection of the supernatant. The dsRNA was used directly in the RT-PCR assay or stored at −20°C.
G-typing by RT-PCR.
G-typing was performed by a modification of the previously published RT-PCR method (8–10). The nucleotide sequences of the primers are shown in Table 1. Typing with G1 to -4 primers and G6, G8, and G10 primers was performed in two separate assays. Briefly, 0.8 μl of 100 mM methylmercuric hydroxide (LabKemi, Stockholm, Sweden) was mixed in a PCR tube with either 3 μl of End 9 and End 9 UK (33 μM each), (G1 to -4) or 9 μl of 10 μM primer sBeg9 (G6-G8-G10), followed by addition of 10 μl of dsRNA and subsequent denaturation for 5 min. Denaturation was stopped by addition of 0.8 μl of 700 mM β-mercaptoethanol and incubation of this mixture for 5 min at room temperature. The denaturated dsRNA was mixed to a final volume of 30 μl with an RT reaction mixture consisting of 3 μl of 10× PCR buffer (Perkin-Elmer), 1.8 μl of 25 mM MgCl2 (Perkin-Elmer), 3 μl of 2 mM deoxynucleoside triphosphates (dNTPs) (Pharmacia Biotech), 0.5 μl of Superscript (200 U/μl) (Gibco BRL), 0.5 μl of RNasin (40 U/μl) (Promega), and dH2O. The dsRNA was reverse transcribed by incubation in a Perkin-Elmer M2400 PCR machine for 60 min at 42°C. The cDNA produced was amplified in the presence of 70 μl of PCR mixture, which consisted of 10 μl of 10× PCR buffer, 6 μl of 25 mM MgCl2, 10 μl of 2 mM dNTPs, a serotype-specific primer mix (either 4 μl of 33 μM aBT1-aCT2-aET3-aDT4 or 12 μl of 10 μM sBeg9-DT6-HT8-ET10), 0.5 μl of Taq polymerase (5 U/μl) (Perkin-Elmer), and dH2O. The PCR was conducted for 30 cycles at 94°C for 2 min, 55°C for 1 min, and 72°C for 1 min, followed by 72°C for 5 min. The G-type-specific PCR product was visualized by electrophoresis for 30 min at 100 V on a 2% agarose–ethidium bromide gel in Tris borate buffer and photographed under UV light on Polaroid film.
TABLE 1.
Oligonucleotide primers for PCR amplification
| Primer (reference) | Nucleotide sequence (5′→3′) | Position | G-type |
|---|---|---|---|
| sBeg9 (10) | GGC TTT AAA AGA GAG AAT TTC | 1–21 | |
| End 9 (8) | GGT CAC ATC ATA CAA TTC TAA TCT AAG | 1062–1036 | |
| End 9 UK (9) | GGT CAC ATC ATA CAA CTC TAA TCT | 1062–1039 | |
| aBT1 (8) | CAA GTA CTC AAA TCA ATG ATG G | 314–335 | 1 |
| aCT2 (8) | CAA TGA TAT TAA CAC ATT TTC TGT G | 411–435 | 2 |
| aET3 (8) | CGT TTG AAG AAG TTG CAA CAG | 689–709 | 3 |
| aDT4 (8) | CGT TTC TGG TGA GGA GTT G | 480–498 | 4 |
| DT6 (10) | CTA GTT CCT GTG TAG AAT C | 499–481 | 6 |
| HT8 (10) | CGG TTC CGG ATT AGA CAC | 273–256 | 8 |
| ET10 (10) | TTC AGC CGT TGC GAC TTC | 714–697 | 10 |
RT-PCR of full-length VP7.
For the BRV-positive fecal specimens that were not typeable by G-type RT-PCR, amplification of the full-length VP7 gene was performed. The VP7 RT-PCR procedure was identical to the G6-G8-G10-typing procedure, with the exception that a VP7 primer mix (33 μM sBeg 9-End 9-End 9 UK) was used instead of the type-specific primer mix in the PCR mixture.
Demonstration of RNA electrophoretic migration pattern and detection of BRV by RNA-PAGE.
The RNA electrophoretic patterns of BRV strains from dairy herd A were examined by RNA-PAGE (24). The RNA segment bands were visualized by silver staining with the Silver Stain Plus kit (Bio-Rad), according to the manufacturer’s instructions.
RESULTS
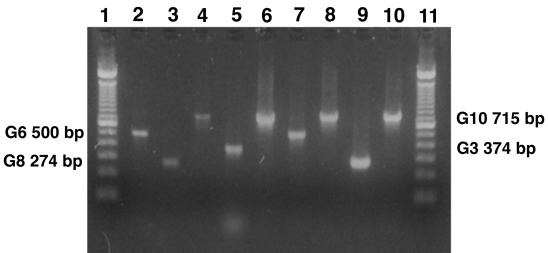
A total of 91 of 98 BRV strains from dairy herd A and the herds in herd B were successfully G-typed. Types G10 and G6 predominated; G10 was detected in 59 (60.2%) and G6 was detected in 30 (30.6%) of the BRV-positive fecal specimens. Both G3 and G8 were each demonstrated in 1 (1.0%) of the BRV-positive fecal specimens. Figure 1 illustrates PCR results. No fecal specimen with double G-types (i.e., more than one amplified G-type-specific PCR product visualized) was identified.
FIG. 1.
G-typing of bovine field strains by RT-PCR. Lanes: 6 to 10, field strains; 2 to 5, rotavirus reference strains G6 (UK), G8 (678), G10 (B223), and G3 (RRV), respectively; 1 and 11, molecular weight marker (100-bp DNA ladder).
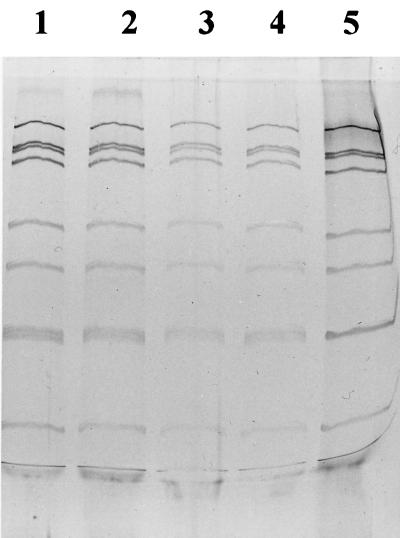
The G-type distribution in dairy herd A from 1993 to 1996 is summarized in Table 2 and shows that, with one exception, G10 strains were the only ones circulated. Furthermore, it was interesting to observe that all identified RNA patterns in the herd showed great similarity (four RNA patterns are shown in Fig. 2), suggesting that a single G-type strain circulated in herd A. No effect of time or clinical symptoms on the G-type distribution was apparent.
TABLE 2.
Rotavirus G-types in 30 fecal specimens from diarrheic and nondiarrheic calves in dairy herd A
| Yr/quarter | Distribution in
G-typea:
|
|||
|---|---|---|---|---|
| G1–4 | G6 | G8 | G10 | |
| 1993/1 | NN | |||
| 1994/4 | DNDDDNN | |||
| 1995/1 | DDNN | |||
| 1995/2 | D | |||
| 1995/3 | NDDD | |||
| 1995/4 | NDNNND | |||
| 1996/1 | N | DDN | ||
| 1996/3 | ND | |||
D, diarrheic; N, nondiarrheic.
FIG. 2.
Illustration of rotavirus RNA electrophoretic pattern in bovine rotavirus field strains by PAGE and silver staining. Lanes 1 to 4 are field strains originating from calves in dairy herd A. Fecal samples were collected in 1993 (lane 1), 1994 (lane 2), 1995 (lane 3), and 1996 (lane 4). Lane 5 contained the bovine rotavirus reference strain NCDV. (Photo taken by Bengt Ekberg.)
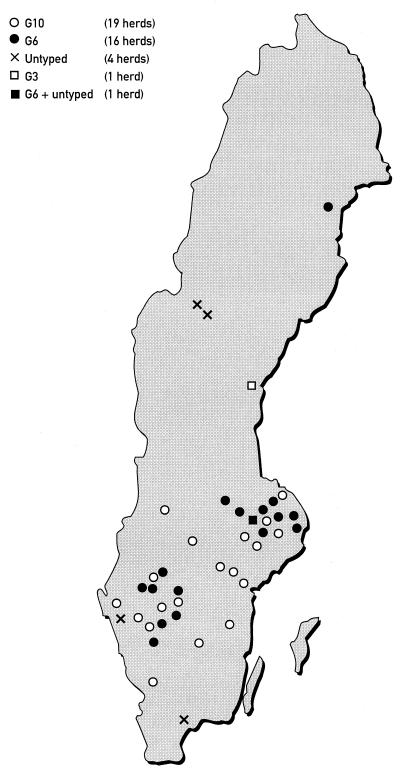
The typing results from the B herds are shown in Table 3. Most interesting was to find that in 15 out of 17 herds where multiple samples were collected, a single G-type was identified. In the remaining two herds, two fecal specimens from one herd were not typeable, and two G6 plus two untyped strains were identified in the other herd. All strains from the beef suckler herds in herd B were characterized as G6, whereas in the dairy herds, G10 was the predominating G-type (54%) and G6 was less frequently detected (32%). The geographic distribution of the herds in study B is illustrated in Fig. 3.
TABLE 3.
Rotavirus G-types in 68 fecal specimens from diarrheic calves in study B
| Herd type | No.
of:
|
||||||
|---|---|---|---|---|---|---|---|
| Herds | Rotavirus strainsa | G6 strains | G10 strains | G3 strains | G1, G2, G4, and G8 strains | Untyped fecal specimens | |
| Beef suckler | 6 | 12 | 12 | 0 | 0 | 0 | 0 |
| Dairy | 35 | 56 | 18 | 30 | 1 | 0 | 7 |
| Total | 41 | 68 | 30 | 30 | 1 | 0 | 7 |
Each herd contributed one to four rotavirus-positive fecal samples.
FIG. 3.
Geographic distribution of rotavirus G-types in 41 Swedish cattle herds with calf diarrhea outbreaks.
Altogether, seven BRV-positive fecal specimens (7.1%), originating from five distinct dairy herds in different parts of Sweden, were not typeable with the G1- to -4-, G6-, G8-, and G10-specific primers (Table 3). Amplification of full-length copies of the VP7 gene obtained from five of the seven untyped fecal specimens indicated that they are of group A rotavirus. The remaining two fecal specimens, originating from two diarrheic calves in the same herd, were examined by RNA-PAGE without detection of dsRNA bands. Their optical densities in ELISA were close to the 0.10 cutoff level (0.12 and 0.24).
DISCUSSION
Several different BRV G-types (G3, G6, G8, and G10) were detected in Swedish cattle herds A and B. G10 clearly dominated in dairy herd A and was also an important G-type in herd B. This differs from previously reported studies, in which G6 was shown to be the predominating G-type (1, 14, 17, 20, 22). The results from this study strongly suggest G-type restriction in the herds. From a vast majority (15 of 18) of the herds from which multiple samples were collected, only a single G-type was detected. Most interestingly, all strains from the suckled beef herds were G6, while G10 predominated or was the only G-type in the dairy herds. A large proportion of the BRV-positive fecal specimens originated from dairy herds, a circumstance that naturally reflects the G-type prevalence. The strong association between herd type and G-type distribution found in this study is supported by others (1, 17, 18). In the Swedish cattle population, exchange of animals between dairy herds and suckled beef herds is limited, which probably enhances the connection between herd type and BRV G-type. If cattle trading is restricted and herds are closed, BRV can still persist in a herd from one year to the next and serve as a source of infection to neonates. Long-term persistence of one G-type was apparent in the closed dairy herd A, which is consistent with a previous report (16). The RNA electrophoretic pattern of the BRV strains from herd A showed remarkable congruity. This suggests that the same BRV strain persisted in the herd for several years, possibly due to heavy contamination of BRV in the calf barn. No characteristic G-type variation was obvious in the geographic distribution. From the northern part of Sweden, two BRV strains were identified as G3 and G6, and two were untyped, which might indicate a different G-type entity compared with the rest of the BRV strains. However, the number of fecal specimens from the north, where the cattle population is sparse, was very low. In southern Sweden, G10 and G6 predominated. The untyped fecal specimens originated from different parts of the country. The results from this study support the opinion (22) that BRV forms a distinct epidemiological entity, which is globally spread due to cattle trading. In previously reported studies, G3 and G8 have occasionally been detected in calf feces (13, 20). In this study, two G3 and G8 strains were identified.
There was no difference in the G-type distribution between the BRV strains from calves with symptomatic infections and those with asymptomatic infections in dairy herd A (Table 2), suggesting that G-type specificity is not associated with virulence. Virulence variation may be associated with other differences in rotavirus genes (5).
The group A rotavirus ELISA and RT-PCR assays used in this study were previously described in several publications (6a, 8–10, 19, 21, 23, 24). The RT-PCR method for G-typing of BRV strains used in this study proved to be a reliable typing method, which is consistent with previous observations (8–10, 15). RT-PCR offered a rapid and sensitive screening method for G-typing directly from feces. Out of 98 BRV-positive fecal specimens in the study, only 7 (7.1%) remained untyped by the G1- to -4-, G6-, G8-, and G10-specific primers. This number is low in comparison with those from other reports (14, 15, 17, 20). The successful G-typing was likely an effect of the silica extraction, the seminested PCR, and the use of methylmercuric hydroxide for denaturation of dsRNA. The untypeability of some BRV-positive fecal specimens may have resulted from the occurrence of G-types other than G1- to -4, G6, G8, and G10 or antigenic variation within the G1- to -4, G6, G8, and G10 serotypes. Further studies, including nucleotide sequence determination, might reveal the G-type and origin of these strains. Two untyped fecal specimens proved BRV positive by ELISA, but no full-length copies of the VP7 gene were amplified and no dsRNA bands were detected by RNA-PAGE, suggesting that these were false positive by ELISA. The fact that these specimens had optical density values close to the cutoff supports this conclusion.
In conclusion, the results from this study strongly indicate a pronounced stability in the BRV G-type distribution in Swedish cattle herds, which probably reflects the resistant character of rotavirus. The resistance and persistence of BRV in a cattle herd are highly significant epidemiological factors and stress the importance of continuous preventive measures in control of the disease. Preventive measures for a herd should aim at reducing BRV contamination in the calf barn and elevating the calves’ resistance to disease through hygiene, management, and vaccination programs. In Sweden, no BRV vaccines are currently registered. An efficient vaccine would clearly be a useful agent for cattle herds experiencing problems related to calf diarrhea. A future vaccine in Sweden should contain both the G6 and G10 BRV strains.
ACKNOWLEDGMENTS
This work was supported by grant 114/93 from the Swedish Farmers’ Foundation for Agricultural Research.
We are deeply grateful to D. R. Snodgrass for kindly providing the BRV reference strains, Qiao Hai-ping for generous help in running the RT-PCR, Helena B. Reineck for excellently performed ELISA and RNA-PAGE, and Stefan Alenius for helpful discussions. Our thanks also go to the veterinary practitioners and to Örjan Hansson and the staff at NötCenter Viken for excellently performed sampling.
REFERENCES
- 1.Bellinzoni R C, Blackhall J O, Mattion N M, Estes M K, Snodgrass D R, LaTorre J L, Scodeller E A. Serological characterization of bovine rotaviruses isolated from dairy and beef herds in Argentina. J Clin Microbiol. 1989;27:2619–2623. doi: 10.1128/jcm.27.11.2619-2623.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Blackhall J, Bellinzoni R, Mattion N, Estes M, La Torre J L, Magnusson G. A bovine rotavirus serotype 1: serologic characterization of the virus and nucleotide sequence determination of the structural glycoprotein VP7 gene. Virology. 1992;189:833–837. doi: 10.1016/0042-6822(92)90617-x. [DOI] [PubMed] [Google Scholar]
- 3.Boom R, Sol C J A, Salimans M M M, Jansen C L, Wertheim van Dillen P M E, van der Noordaa J. Rapid and simple method for purification of nucleic acids. J Clin Microbiol. 1990;28:495–503. doi: 10.1128/jcm.28.3.495-503.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Brüssow H, Nakagomi O, Gerna G, Eichhorn W. Isolation of an avianlike group A rotavirus from a calf with diarrhea. J Clin Microbiol. 1992;30:67–73. doi: 10.1128/jcm.30.1.67-73.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Burke B, Bridger J C, Desselberger U. Temporal correlation between a single amino acid change in the VP4 of a porcine rotavirus and a marked change in pathogenicity. Virology. 1994;202:754–759. doi: 10.1006/viro.1994.1397. [DOI] [PubMed] [Google Scholar]
- 6.Clark K J, Tamborello T J, Xu Z, Mann F E, Bonnot C E, Woode G N. An unusual group-A rotavirus associated with an epidemic of diarrhea among three-month-old calves. J Am Vet Med Assoc. 1996;208:552–554. [PubMed] [Google Scholar]
- 6a.de Verdier Klingenberg K, Svensson L. Group A rotavirus as a cause of neonatal calf enteritis in Sweden. Acta Vet Scand. 1998;39:195–199. doi: 10.1186/BF03547792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6b.de Verdier Klingenberg, K., I. Vågsholm, and S. Alenius. Reduction of calf diarrhea after strict closure and eradication of bovine viral diarrhea virus in a dairy herd. Submitted for publication. [PubMed]
- 7.Estes M K, Cohen J. Rotavirus gene structure and function. Microbiol Rev. 1989;53:410–449. doi: 10.1128/mr.53.4.410-449.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gouvea V, Glass R I, Woods P, Taniguchi K, Clark H F, Forrester B, Fang Z-Y. Polymerase chain reaction amplification and typing of rotavirus nucleic acid from stool specimens. J Clin Microbiol. 1990;28:276–282. doi: 10.1128/jcm.28.2.276-282.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gouvea V, Ramirez C, Li B, Santos N, Saif L, Clark H F, Hoshino Y. Restriction endonuclease analysis of the vp7 genes of human and animal rotaviruses. J Clin Microbiol. 1993;31:917–923. doi: 10.1128/jcm.31.4.917-923.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gouvea V, Santos N, d. Carmo Timenetsky M. Identification of bovine and porcine rotavirus G types by PCR. J Clin Microbiol. 1994;32:1338–1340. doi: 10.1128/jcm.32.5.1338-1340.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hoshino Y, Kapikian A Z. Rotavirus antigens. Curr Top Microbiol Immunol. 1994;185:179–227. doi: 10.1007/978-3-642-78256-5_7. [DOI] [PubMed] [Google Scholar]
- 12.Hurtado O, Parwani A V, Tineo A, Lucchelli A, Saif L J. Serotypic analysis of group A bovine rotavirus field specimens using G1-, G2-, and G3-specific monoclonal antibodies in ELISA. J Vet Diagn Investig. 1995;7:251–254. doi: 10.1177/104063879500700215. [DOI] [PubMed] [Google Scholar]
- 13.Hussein H A, Parwani A V, Rosen B I, Lucchelli A, Saif L J. Detection of rotavirus serotypes G1, G2, G3, and G11 in feces of diarrheic calves by using polymerase chain reaction-derived cDNA probes. J Clin Microbiol. 1993;31:2491–2496. doi: 10.1128/jcm.31.9.2491-2496.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hussein H A, Cornaglia E, Saber M S, El-Azhary Y. Prevalence of serotypes G6 and G10 group A rotaviruses in dairy calves in Quebec. Can J Vet Res. 1995;59:235–237. [PMC free article] [PubMed] [Google Scholar]
- 15.Hussein H A, Frost E, Talbot B, Shalaby M, Cornaglia E, El-Azhary Y. Comparison of polymerase chain reaction and monoclonal antibodies for G-typing of group A bovine rotavirus directly from fecal material. Vet Microbiol. 1996;51:11–17. doi: 10.1016/0378-1135(95)00202-2. [DOI] [PubMed] [Google Scholar]
- 16.Ishizaki H, Ohta C, Shirahata T, Goto H, Taniguchi K, Urasawa T, Urasawa S. Persistence of a single electropherotype and serotype (G6P5) of bovine rotavirus in calves on a closed dairy farm from 1990 to 1993. Am J Vet Res. 1995;56:1019–1024. [PubMed] [Google Scholar]
- 17.Lucchelli A, Kang S Y, Jayasekera M K, Parwani A V, Zeman D H, Saif L J. A survey of G6 and G10 serotypes of group A bovine rotaviruses from diarrheic beef and dairy calves using monoclonal antibodies in ELISA. J Vet Diagn Investig. 1994;6:175–181. doi: 10.1177/104063879400600207. [DOI] [PubMed] [Google Scholar]
- 18.Muñoz M, Parwani A V, Lucchelli A, Saif L J. Abstracts of the 74th Annual Conference of Research Workers in Animal Diseases. 1993. G typing of group A bovine rotaviruses in diarrheic beef and dairy calves by using polymerase chain reaction derived cDNA probes, abstr. 56. [Google Scholar]
- 19.Nakagomi O, Oyamada H, Nakagomi T. Experience with serotyping rotavirus strains by reverse transcription and two-step polymerase chain reaction with generic and type-specific primers. Mol Cell Probes. 1991;5:285–289. doi: 10.1016/0890-8508(91)90051-k. [DOI] [PubMed] [Google Scholar]
- 20.Parwani A V, Hussein H A, Rosen B I, Lucchelli A, Navarro L, Saif L J. Characterization of field strains of group A bovine rotaviruses by using polymerase chain reaction-generated G and P type-specific cDNA probes. J Clin Microbiol. 1993;31:2010–2015. doi: 10.1128/jcm.31.8.2010-2015.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Santos N, Lima R C C, Pereira C F A, Gouvea V. Detection of rotavirus types G8 and G10 among Brazilian children with diarrhea. J Clin Microbiol. 1998;36:2727–2729. doi: 10.1128/jcm.36.9.2727-2729.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Snodgrass D R, Fitzgerald T, Campbell I, Scott F M M, Browning G F, Miller D L, Herring A J, Greenberg H B. Rotavirus serotypes 6 and 10 predominate in cattle. J Clin Microbiol. 1990;28:504–507. doi: 10.1128/jcm.28.3.504-507.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Svensson L, Grandien M, Pettersson C-A. Comparison of solid-phase immune electron microscopy by use of protein A with direct electron microscopy and enzyme-linked immunosorbent assay for direct detection of rotavirus in stool. J Clin Microbiol. 1983;18:1244–1249. doi: 10.1128/jcm.18.5.1244-1249.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Svensson L, Uhnoo I, Grandien M, Wadell G. Molecular epidemiology of rotavirus infections in Uppsala, Sweden, 1981. J Med Virol. 1986;18:101–111. doi: 10.1002/jmv.1890180202. [DOI] [PubMed] [Google Scholar]