Abstract
In this world, there are several acute viral infections. One of them is influenza, a respiratory disease caused by the influenza virus. Stochastic modelling of infectious diseases is now a popular topic in the current century. Several stochastic epidemiological models have been constructed in the research papers. In the present article, we offer a stochastic two-strain influenza epidemic model that includes both resistant and non-resistance strains. We demonstrate both the existence and uniqueness of the global positive solution using the stochastic Lyapunov function theory. The extinction of our research sickness results from favourable circumstances. Additionally, the infection’s persistence in the mean is demonstrated. Finally, to demonstrate how well our theoretical analysis performs, various noise disturbances are simulated numerically.
Introduction
The respiratory system, which includes the nose, throat, and lungs, is affected by viruses that cause influenza, sometimes known as the flu [1]. Flu is frequently characterized by acute symptoms and potentially fatal consequences. Viruses with the names influenza A, B, C, and D are four different varieties [2]. Seasonal diseases brought on by influenza types A and B occur nearly every winter. The disease brought on by type C influenza is often quite mild and frequently symptomless. Cattle are affected by type D influenza viruses, which are not known to cause any illnesses in people. All subtype of type A influenza viruses is split into strains, and each strain is additionally categorized into subcategories. Just viruses of type A have sparked pandemic. The various types of proteins found on the outside of the influenza virus envelope are designated by the letters H and N. the different influenza subtypes Hemagglutinin, also known as the HA protein, and neuraminidase, sometimes known as the NA protein, are two types of proteins that attach to the surface of viruses. The immune system of the body may produce antibodies that can identify these particular viral proteins (antigens) and hence can combat this particular influenza virus.
Scholars have identified 18 distinct HA protein forms and 11 distinct NA protein types that may co-occur in a wide range of combinations in influenza viruses that infect birds. According to reports, each of these mixtures represents a unique strain of influenza virus with a specific number of H(number) and N(number) proteins, such as , , , etc [3, 4]. Although they might be classified as strains, type B influenza viruses are not classified into sub-types. Rarely does vaccination offer protection against novel influenza viruses. This was evident during the 2009 influenza pandemic. Antiviral medication is thus necessary to prevent the spread of the flu epidemic [5]. Resistance to the influenza virus is increasingly a problem. As an illustration, consider the and viruses’ resistance to aminoadamantanes and oseltamivir, respectively [8–10]. Future pandemics might be brought on through resistance, which is lethal. In comparison to the original strain, a new strain’s force of transmission is typically thought to be quite weak. According to references [10–12], mutation reduces the viral strength, which is connected to this event.
In epidemiology, mathematical modelling is crucial for a deeper understanding of the numerous facets of many illnesses. Because there are several diseases in which more than one pathogen strain is noted due to the process of viral mutation, for example, influenza [32], human immunodeficiency virus [33], tuberculosis [34], and COVID-19 [35], multi-strain epidemics models have attracted the focus of many researchers. Recently, in [26–28], the research of the two-strain epidemic model by fractional differential equation was also established, because the fractional-order differential equations can be helpful in modelling biological systems [29–31]. In actuality, some unknown environmental perturbations invariably affect population dynamics and epidemic systems. As we all know, real life is filled with randomness and unpredictability. Stochastic models can better conform to the actual situation, because most epidemic models are influenced by environmental factors, such as percipitation, temperature, relative humidity. Thus, the variability of epidemic growth and spread is random due to the different infectious periods. It has equally been shown that stochastic models can provide additional degree of realism as compared with their deterministic study. Furthermore, several writers have extensively examined certain stochastic epidemics models, including [36–40]. An epidemic model with a twofold hypothesis that combines two transmission mechanisms, SIS and SIR, with two distinct saturation incidence rates is addressed in [36] Boukanjime et al. Although there might be two epidemic illnesses in the current world, one brought on by virus A and the other by virus B, the authors of [37] explored an SIS model with the twin epidemic theory. With two distinct saturation incidence rates, Chang et al. [38] constructed a stochastic SIRS model and determined the thresholds that determine whether the disease will remain or go away. The existence of an ergodic stationary distribution of the nonnegative solutions to a stochastic SIS epidemic model with double illnesses and the Beddington-DeAngelis incidence was demonstrated by Liu and Jiang, who used [39] as their source. In [40], it was looked at how two different infectious diseases might spread vertically under a stochastic epidemic model.
In our case we will study two strains of an influenza epidemic model, after analyze the situation in which the two strains can coexist and the difference in their mode of transmission, we employ the use of mathematical modeling. Principal element in mathematical modeling is the incidence rate. Its significance in epidemiology can’t be over emphasized.
Recently, Baba et al. [41] constructed and studied a resistance and non-resistance strains of influenza.
| 1.1 |
Here S(t) is the susceptibles, is the infective resistant individuals, is the infective non-resistant individuals and R(t) is the removed ones. The parameters in the model (1.1) are positive constants where : is a recruitment into susceptible. is natural mortality rate, The rate of infection by resistant strain is represented by , the rate of infection by non-resistant strain is denoted by , removal of individuals carrying the resistant strain from the population is , removal of individuals carrying the non-resistant strain is , and the mutation effect on the resistant strain is . Both illnesses are spread by interaction between people in the susceptible compartment and those in the and compartments, which have, respectively, bilinear and saturation incidence rates. We anticipate that the populations that reside in environments where random accidents are prevalent are mostly impacted by the contact rate, which will primarily present itself as changes in the saturated response rate, so that turn into and turn into where and are standard Brownian motion with intensities and . Now, the corresponding stochastic model of the system (1.1) is as follows:
| 1.2 |
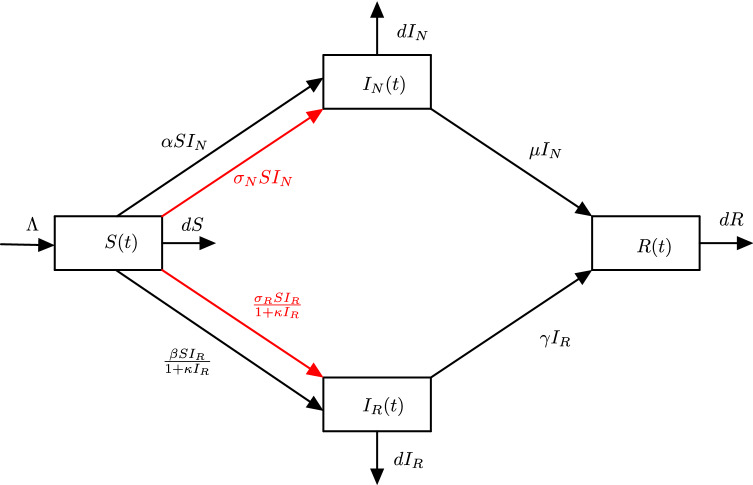
Fig. 1.
The detailed flowchart of system (1.2)
The remaining of this paper is arranged as: In the Sect. 2, we show the positivity and the boundedness of solutions of the stochastic system (1.2). The extinction of the non-resistance and resistance infectious diseases will be discussed in Sect. 3. In Sect. 4 we study the persistence in mean of the epidemic. In Sect. 5, the numerical simulations are carried out to confirm our theoretical results. Lastly, a brief discussion is given in the end to conclude this paper.
Existence and uniqueness of the global nonnegative solution
The notations, definitions, and lemmas we utilised to examine our primary outcomes are provided in this part.
Consider a filtration with a complete probability space that fulfills the usual conditions with increasing and right continuous while is the set of -null sets. The function B(t) denotes a scalar Brownian motion which is defined on .
We introduce the following notations:
In general, consider the d-dimensional stochastic differential equation
with initial value . B(t) denotes a d-dimensional standard Brownian motion defined on the complete probability space . Denote by the family of all nonnegative functions V(X) defined on such that they are continuously twice differentiable in X. The differential operator L of Eq. (1.5) is defined by [42]
If L acts on a function , then
where .
In view of Itô’s formula [42], if , then
For arbitrary integrable function h on , define .
Let and
Definition 1
The diseases and are said to go extinction if and .
The diseases and will be persist in mean if and such that and
Remark 2
Let the set
The total population in systems (1.1) and (1.2) verifies, the equation
which gives by integration
If , then almost surely. Thus, the set is almost surely positively invariant by the systems (1.1) and (1.2) respectively, throughout the rest, we assume that
Lemma 3
For the initial condition , the model (1.2) has at most one solution and will belong to with probability one almost surely.
Proof
As all the coefficients of the proposed stochastic model (1.2) are locally Lipschitz continuous, then for each initial condition , exclusive local solution on , where denotes the explosion time.
It is obligatory to verify that the solution is global, one need only to prove that almost surely.
For this, let us take sufficiently large to get that , integer . Next, we express the stopping time by:
| 2.1 |
where one can set . Thus, increases as m tends to .
Let , and almost surely. When almost surely is true, then almost surely and almost surely . To put it another way, we just need to demonstrate that almost surely. Otherwise, there will be constants and with
| 2.2 |
So, with
| 2.3 |
Let us take a -function as
| 2.4 |
where ,
Applying the Itô’s method on , one get
| 2.5 |
where is defined by
| 2.6 |
| 2.7 |
Thus
| 2.8 |
Integrating (2.8) from 0 to and then using the notion of expectations, we have
| 2.9 |
Let for . Using (2.3), one can acquire Notice that , or or or equals either m or
Therefore,
Then we attain
| 2.10 |
where is the indicator function of . For , one reach
| 2.11 |
is a contradiction. Hence, .
Lemma 4
[42] Let satisfies model (1.2) with . Then
| 2.12 |
Extinction
Here, we create the conditions that result in extinction of the non-resistance and resistance infectious strains motioned in the system (1.2).
Proposition 5
If
| 3.1 |
then the non-resistance strain of (1.2) go to the extinction almost surely.
Proof
Let satisfies the model(1.2) with . Using the Itô’s method, one get
| 3.2 |
| 3.3 |
Integrating (3.3) from 0 to t and doing some manipulation, we obtain
| 3.4 |
where is the local continuous martingale satisfying , and by the Lemma 4, we obtain
| 3.5 |
Since . Applying superior limit of 3.4, we conclude
| 3.6 |
which means that almost surely. Hence the theorem.
Proposition 6
If
| 3.7 |
then the resistance strain of (1.2) go to the extinction almost surely.
Proof
Let satisfies the model (1.2) with . Implementing the Itô’s technique on model (1.2) results in
| 3.8 |
| 3.9 |
Integrating Eq. (3.9), we reach
| 3.10 |
where is the local continuous martingale satisfying , and by the Lemma 4, one may reach
| 3.11 |
Since . Applying superior limit to (3.10), we conclude
| 3.12 |
which implies that almost surely.
Remark 7
Proposition 5 and Proposition 6 shows that when and the non-resistance strain and resistance strain of system (1.2) die out almost surely, respectively. In other words, large white noise stochastic disturbance yield the two strains extinct. Hence, we presume that the white noise disturbance is not large in the rest of this manuscript.
Let
Theorem 8
Let satisfies the model (1.2) with .
Proof
Firstly, taking integral of both sides of (3.2) and doing some manipulations gives
| 3.13 |
| 3.14 |
where is the local continuous martingale satisfying , and by the Lemma 4, one have
| 3.15 |
Using superior limit on Eq. (3.14), one get
| 3.16 |
Consequently, , almost surely.
Secondly, for both sides of (3.8), integrating from 0 to t first and doing some manipulations gives
| 3.17 |
| 3.18 |
where is the local continuous martingale satisfying , and by the Lemma 4, one have
| 3.19 |
We achieve the following result by using superior limit
| 3.20 |
Consequently, , almost surely.
Lastly, without loss of generality, one can suppose that and , from the first class of the model (1.2), one obtain
| 3.21 |
As and , thus
| 3.22 |
Also,
| 3.23 |
Let and , one attain
| 3.24 |
From (3.24) and (3.22), one get
| 3.25 |
Next, we prove the last conclusion. Using the third equation of (1.2), we obtain
| 3.26 |
Its clear by comparison theorem we deduce
| 3.27 |
Extending and to 0, we have
| 3.28 |
Remark 9
From Theorem 8 we show that the non-resistant and the resistant strains will die out if the white noise disturbances are large than certain values or and , and the white noise disturbances are not so large.
Persistence in mean
In this section, our main concern to determine sufficient conditions for the persistence of the infectious disease.
Theorem 10
Let satisfies the model (1.2) with ,
-
If , and , then the resistance strain will go to extinct and the strain will persist, furthermore, satisfies
-
If , and , then the non-resistance strain go to extinct and the strain will persist, furthermore, satisfies
-
If , , then the two strains and are persistent in mean, furthermore, and satisfy
where
Proof
-
Let the function define by Then the first three equation of model (1.2), implies
Since and one can see from Proposition 6 that, almost surely. Then we can choose for all t large enough small enough, such that therefore,4.1
Using the Itô’s formula to model (1.2), we obtain4.2
Hence,4.3
Integration of (4.4) gives4.4
So, we obtain4.5
where is the local continuous martingale satisfying , and using Lemma 4, the result is:4.6
Since for all t large enough we can choose small enough, such that4.7
Let yields
Therefore, -
Hence, we have4.10
where is the local continuous martingale satisfying , and using Lemma 4, one have4.11
Since for all t large enough we can choose small enough, such that4.12
Let yields
Therefore -
Notice that
Define4.13
With the help of Itô’s formula, we reach:4.14
Therefore4.15
Integration of (4.16) gives4.16
Remark 11
Proposition 5 and Proposition 6 shows that the non-resistance and the resistance infections diseases can be extinct if the white noise disturbances are larger than certain values. Theorem 8 and 10 show that the non-resistant (resistant) infection diseases can prevail if the white noise disturbances are small enough such that ( ) respectively. This implies that the stochastic disturbance may cause epidemic diseases to die out.
Graphical analysis
In this section, we implement the Milstein procedure which is given in [43] to test numerically the persistence and the extinction of the disease. The discretization of system 1.2 is given by
| 5.1 |
where are the Guassian random variables which Obey Gaussian distribution .
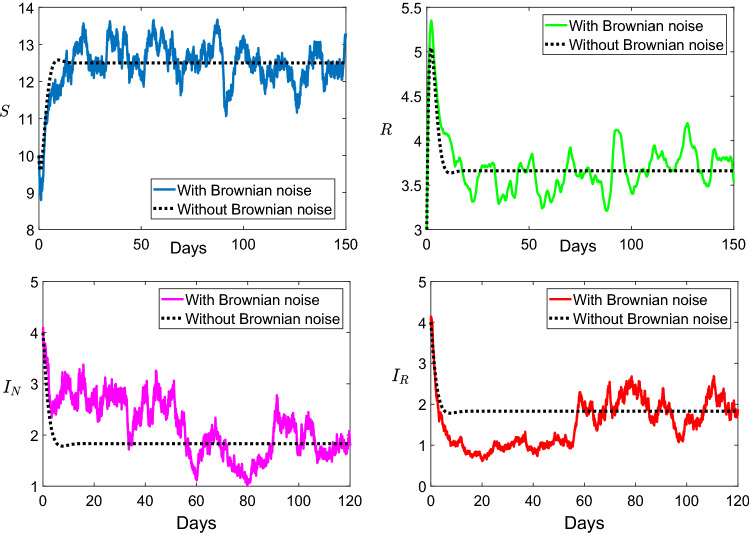
Indeed, Fig. 2 shows the dynamics of the non-resistance and resistance strains of influenza for the chosen values of the parameters , , , , , , , and . We clearly see that the all the model variables stay at a strictly positive level. Within this parameters, we have and , then the two infectious diseases and will persist. This result is consistent with the theoretical result given in Theorem 10.
Fig. 2.
Dynamic describing the persistence of the non-resistance and resistance diseases
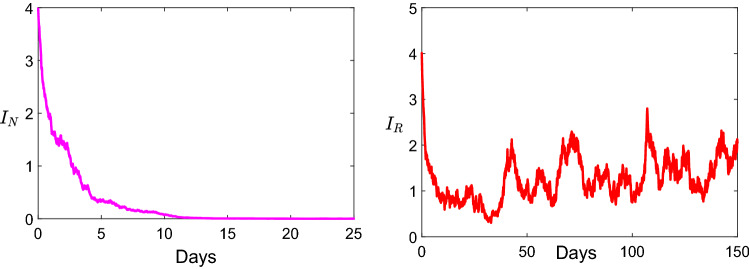
Next, we take the parameters values for the stochastic model 1.2 as: , , , , , , and . Within this parameters we get and , . Thus, non-resistance strain goes to the extinction, and resistance strain will persist (see Fig. 3). This result is consistent with the theoretical result given in Theorem 10.
Fig. 3.
Dynamic describing the persistence of the non-resistance strain and the extinction of resistance strain
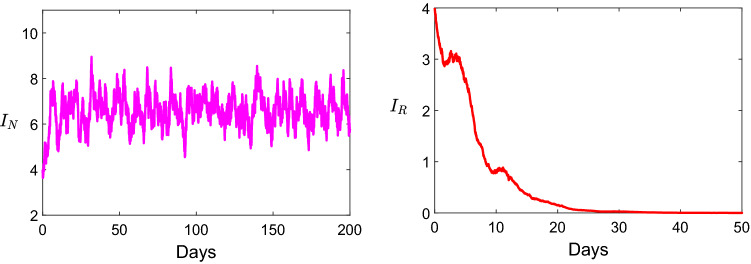
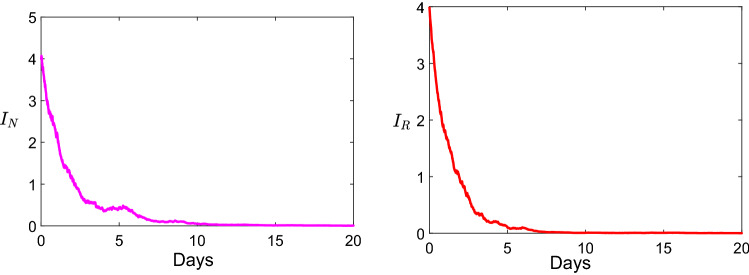
In Fig. 4, we take the parameters values for the model 1.2 as: , , , , ,, , and . Within this parameters we get , and . Thus, non-resistance strain is persistent, and resistance strain go to extinction.
Fig. 4.
Dynamics of the infection describing the persistence of the non-resistance strain and the extinction of resistance strain
In Fig. 5, we take the parameters values for the model 1.2 as: , , , , , , , and . Within this parameters we get , and both conditions and . Thus, both of them go to the extinction which is consistent with the theoretical result given in Sect. 3.
Fig. 5.
Dynamics of the infection describing the persistence of the non-resistance strain and the extinction of resistance strain
Conclusion and discussion
The novelty of this study is that we analyzed the dynamics of two-strain SIR epidemic model including non-resistance and resistance sub-strain of influenza, by considering different incidence rates for these strains. This is due to the fact that the mutated strain will have a minimal effect. We have assumed saturated and bilinear incidence rates for the resistant and non-resistant strains respectively. Saturated incidence rate grasps the negotiating alteration and swarming impact of the infected people and hinders the unboundedness of the interconnection rate by fitting parameters, which was reused in several epidemic issue . Indeed, a stochastic two-strain epidemic model describing resistance and non-resistance strains of influenza was suggested and studied. The existence and uniqueness of the positive solution to the stochastic model (1.2) are proved. The extinction of our studied disease was derived with sufficient conditions. The persistence in the mean of the infection was also established. Different numerical simulations for different noises disturbance were performed to illustrate the efficiency of our theoretical study.
Some interesting topics deserve further investigation. On the one hand, one may propose some more realistic models, such as considering the effects of impulsive perturbations on system (1.2). On the other hand, it is interesting to introduce the telegraph noise, such as continuous-time Markov chain, into system (1.2). Also it is interesting to consider more complex influenza virus models, for example, multi-group model. These problems will be the subject of future work.
Data Availability
No data associated in the manuscript.
Contributor Information
Shabir Ahmad, Email: shabirahmad2232@gmail.com.
Kamsing Nonlaopon, Email: nkamsi@kku.ac.th.
References
- 1.Mohler L, Flockerzi D, Sann H, Reichl U. Mathematical model of influenza A virus production in large scale micro carrier culture. Biotechnol. Bioeng. 2005;90:46–58. doi: 10.1002/bit.20363. [DOI] [PubMed] [Google Scholar]
- 2.Tamura S, Tanimoto T, Kurata T. Mechanism of cross-protection provided by influenza virus infection and their application to vaccines. Jpn. J. Infect. Dis. 2005;58:195–207. [PubMed] [Google Scholar]
- 3.Webster RG, Peiris M, Chen H, Guan Y. H5N1 outbreaks and enzootic influenza. Emerg. Inf. Dis. 2016;12:3–8. doi: 10.3201/eid1201.051024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Martcheva M, Iannelli M, Li XZ. Sub threshold coexistence of strains: the impact of vaccination and mutation. Math. Biosci. Eng. 2007;4:287–317. doi: 10.3934/mbe.2007.4.287. [DOI] [PubMed] [Google Scholar]
- 5.Ward P, Small I, Smith J, Suter P, Dutkowski R. Oseltamivir (Tamiflu) and its potential for use in the event of an influenza pandemic. J. Antimicrob. Chemother. 2005;55:i5–i21. doi: 10.1093/jac/dki018. [DOI] [PubMed] [Google Scholar]
- 6.Schünemann HJ, Hill SR, Kakad M, et al. WHO Rapid Advice Guidelines for pharmacological management of sporadic human infection with avian influenza A (H5N1) virus. Lancet Infect. Dis. 2007;7:21–31. doi: 10.1016/S1473-3099(06)70684-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Fiore AE, Fry A, Shay D, Gubareva L, Bresee JS, Uyeki TM. Antiviral agents for the treatment and chemoprophylaxis of influenza-recommendations of the Advisory Committee on Immunization Practices (ACIP) MMWR Recomm. Rep. 2011;60:1–24. [PubMed] [Google Scholar]
- 8.Baranovich T, Saito R, Suzuki Y, et al. Emergence of H274Y oseltamivir-resistant A(H1N1) influenza viruses in Japan during the 2008–2009 season. J. ClinVirol. 2010;47:23–28. doi: 10.1016/j.jcv.2009.11.003. [DOI] [PubMed] [Google Scholar]
- 9.Monto AS, McKimm-Breschkin JL, Macken C, et al. Detection of influenza viruses resistant to neuraminidase inhibitors in global surveillance during the first 3 years of their use. Antimicrob. Agents Chemother. 2006;50:2395–2402. doi: 10.1128/AAC.01339-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Carr J, Ives J, Kelly L, et al. Influenza virus carrying neuraminidase with reduced sensitivity to oseltamivir carboxylate has altered properties in vitro and is compromised for infectivity and replicative ability in vivo. Antivir. Res. 2002;54:79–88. doi: 10.1016/S0166-3542(01)00215-7. [DOI] [PubMed] [Google Scholar]
- 11.Herlocher ML, Carr J, Ives J, et al. Influenza virus carrying an R292K mutation in the neuraminidase gene is not transmitted in ferrets. Antivir. Res. 2002;54:99–111. doi: 10.1016/S0166-3542(01)00214-5. [DOI] [PubMed] [Google Scholar]
- 12.Bouvier NM, Lowen AC, Palese P. Oseltamivir-resistant influenza A viruses are transmitted efficiently among guinea pigs by direct contact but not by aerosol. J. Virol. 2008;82:10052–10058. doi: 10.1128/JVI.01226-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ives JA, Carr JA, Mendel DB, et al. The H274Y mutation in the influenza A/H1N1 neuraminidase active site following oseltamivir phosphate treatment leave virus severely compromised both in vitro and in vivo. Antivir. Res. 2002;55:307–317. doi: 10.1016/S0166-3542(02)00053-0. [DOI] [PubMed] [Google Scholar]
- 14.Herlocher ML, Truscon R, Elias S, et al. Influenza viruses resistant to the antiviral drug oseltamivir: transmission studies in ferrets. J. Infect. Dis. 2004;190:1627–1630. doi: 10.1086/424572. [DOI] [PubMed] [Google Scholar]
- 15.Abed Y, Goyette N, Bovin G. A reverse genetics study of resistance to neuraminidase inhibitors in an influenza A/H1N1 virus. AntivirTher. 2004;9:577–581. [PubMed] [Google Scholar]
- 16.M.A. Rameix-Welti, V. Enouf, F. Cuvelier, P. Jeannin, S. van der Werf. Enzymatic properties of the neuraminidase of seasonal H1N1 influenza viruses provide insights for the emergence of natural resistance to oseltamivir. PLoS Pathog. 4. Article e1000103 (2008) [DOI] [PMC free article] [PubMed]
- 17.Baz M, Abed Y, Simon P, Hamelin ME, Boivin G. Effect of the neuraminidase mutation H274Y conferring resistance to oseltamivir on the replicative capacity and virulence of old and recent human influenza A(H1N1) viruses. J. Infect. Dis. 2010;201:740–745. doi: 10.1086/650464. [DOI] [PubMed] [Google Scholar]
- 18.Matsuzaki Y, Mizuta K, Aoki Y, et al. A two-year survey of the oseltamivir-resistant influenza A(H1N1) virus in Yamagata, Japan and the clinical effectiveness of oseltamivir and zanamivir. Virol. J. 2010;7:53. doi: 10.1186/1743-422X-7-53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Bloom JD, Gong LI, Baltimore D. Permissive secondary mutations enable the evolution of influenza oseltamivir resistance. Science. 2010;328:1272–1275. doi: 10.1126/science.1187816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Baba IA, Hincal E. Global stability analysis of two-strain epidemic model with bilinear and non-monotone incidence rates. Eur. Phys. J. Plus. 2017;132:208. doi: 10.1140/epjp/i2017-11476-x. [DOI] [Google Scholar]
- 21.Bentaleb D, Amine S. Lyapunov function and global stability for a two-strain SEIR model with bilinear and non-monotone. Int. J. Biomath. 2019;12(2):1950021. doi: 10.1142/S1793524519500219. [DOI] [Google Scholar]
- 22.Meskaf A, Khyar O, Danane J, Allali K. Global stability analysis of a two-strain epidemic model with non-monotone incidence rates. Chaos Solitons Fractals. 2020;133:109647. doi: 10.1016/j.chaos.2020.109647. [DOI] [Google Scholar]
- 23.Farah EM, Amine S, Allali K. Dynamics of a time-delayed two-strain epidemic model with general incidence rates. Chaos Solitons Fractals. 2021;153:111527. doi: 10.1016/j.chaos.2021.111527. [DOI] [Google Scholar]
- 24.Martcheva M. A non-autonomous multi-strain SIS epidemic model. J. Biol. Dyn. 2016;3:235–251. doi: 10.1080/17513750802638712. [DOI] [PubMed] [Google Scholar]
- 25.Naji RK, Hussien RM. The dynamics of epidemic model with two types of infectious diseases and vertical transmission. J. Appl. Math. 2016;2016:1–16. doi: 10.1155/2016/4907964. [DOI] [Google Scholar]
- 26.A. Khan, K. Shah, T. Abdeljawad, M.A. Alqudah, Existence of results and computational analysis of a fractional order two strain epidemic model. Results Phys., 105649 (2022)
- 27.A. Allali, S. Amine, Stability analysis of a fractional-order two-strain epidemic model with general incidence rates. Commun. Math. Biol. Neurosci. Article-ID 43 (2022)
- 28.S. Rezapour, J.K.K. Asamoah, A. Hussain, H. Ahmad, R. Banerjee, S. Etemad, T. Botmart, A theoretical and numerical analysis of a fractal-fractional two-strain model of meningitis. Results Phys. 105775 (2022)
- 29.Shah K, Arfan M, Ullah A, Al-Mdallal Q, Ansari KJ, Abdeljawad T. Computational study on the dynamics of fractional order differential equations with applications. Chaos Solitons Fractals. 2022;157:111955. doi: 10.1016/j.chaos.2022.111955. [DOI] [Google Scholar]
- 30.Arfan M, Mahariq I, Shah K, Abdeljawad T, Laouini G, Mohammed PO. Numerical computations and theoretical investigations of a dynamical system with fractional order derivative. Alex. Eng. J. 2022;61(3):1982–1994. doi: 10.1016/j.aej.2021.07.014. [DOI] [Google Scholar]
- 31.Sinan M, Shah K, Kumam P, Mahariq I, Ansari KJ, Ahmad Z, Shah Z. Fractional order mathematical modeling of typhoid fever disease. Results Phys. 2022;32:105044. doi: 10.1016/j.rinp.2021.105044. [DOI] [Google Scholar]
- 32.Baba IA, Hincal E. A model for influenza with vaccination and awareness. Chaos Solitons Fractals. 2018;106:49–55. doi: 10.1016/j.chaos.2017.11.003. [DOI] [Google Scholar]
- 33.S. Amine, E.M. Farah, Global stability of HIV-1 and HIV-2 model with drug resistance compartment. Commun. Math. Biol. Neurosci. Article-ID 38 (2021)
- 34.Bhunu CP, Garira W. A two strain tuberculosis transmission model with therapy and quarantine. Math. Model. Anal. 2009;14(3):291–312. doi: 10.3846/1392-6292.2009.14.291-312. [DOI] [Google Scholar]
- 35.Khyar O, Allali K. Global dynamics of a multi-strain SEIR epidemic model with general incidence rates: application to COVID-19 pandemic. Nonlinear Dyn. 2020;102(1):489–509. doi: 10.1007/s11071-020-05929-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Boukanjime B, El Fatini M, Laaribi A, Taki R. Analysis of a deterministic and a stochastic epidemic model with two distinct epidemics hypothesis. Physica A. 2019;534:122321. doi: 10.1016/j.physa.2019.122321. [DOI] [Google Scholar]
- 37.Meng X, et al. Dynamics of a novel nonlinear stochastic SIS epidemic model with double epidemic hypothesis. J. Math. Anal. Appl. 2016;433(1):227–242. doi: 10.1016/j.jmaa.2015.07.056. [DOI] [Google Scholar]
- 38.Chang Z, Meng X, Xiao L. Analysis of a novel stochastic SIRS epidemic model with two different saturated incidence rates. Physica A. 2017;472:103–116. doi: 10.1016/j.physa.2017.01.015. [DOI] [Google Scholar]
- 39.Liu Q, Jiang D. Stationary distribution of a stochastic SIS epidemic model with double diseases and the Beddington-DeAngelis incidence. Chaos: Interdiscip. J. Nonlinear Sci. 2017;27(8):083126. doi: 10.1063/1.4986838. [DOI] [PubMed] [Google Scholar]
- 40.Wang X, Huang C, Hao Y, Shi Q. A stochastic mathematical model of two different infectious epidemic under vertical transmission. Math. Biosci. Eng. 2022;19(3):2179–2192. doi: 10.3934/mbe.2022101. [DOI] [PubMed] [Google Scholar]
- 41.Baba IA, Ahmad H, Alsulami MD, Abualnaja KM, Altanji M. A mathematical model to study resistance and non-resistance strains of influenza. Results Phys. 2021;26:104390. doi: 10.1016/j.rinp.2021.104390. [DOI] [Google Scholar]
- 42.Mao X. Stochastic Differential Equations and Applications. Amsterdam: Elsevier; 2007. [Google Scholar]
- 43.Higham DJ. An algorithmic introduction to numerical simulation of stochastic differential equations. SIAM Rev. 2001;43:525–546. doi: 10.1137/S0036144500378302. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
No data associated in the manuscript.