Abstract
Flavonoid secondary metabolites can treat and prevent many diseases, but systematic studies on regulation of the biosynthesis of such metabolites in aboveground parts of Salvia miltiorrhiza are lacking. In this study, metabonomic and transcriptomic analyses of different S. miltiorrhiza phenotypes were conducted to explore pathways of synthesis, catalysis, accumulation, and transport of the main flavonoid secondary metabolites regulating pigment accumulation. Tissue localization and quantitative analysis of flavonoid secondary metabolites were conducted by laser scanning confocal microscopy (LSCM). A total 3090 differentially expressed genes were obtained from 114,431 full-length unigenes in purple and green phenotypes, and 108 functional genes were involved in flavonoid biosynthesis. Five key phenylpropane structural genes (PAL, 4CL, ANS, 3AT, HCT) were highly differentially expressed, and four transcription factor genes (MYB, WRKY, bHLH, bZiP) were identified. In addition, six GST genes, nine ABC transporters, 22 MATE genes, and three SNARE genes were detected with key roles in flavonoid transport. According to LSCM, flavonoids were mainly distributed in epidermis, cortex, and collenchyma. Thus, comprehensive and systematic analyses were used to determine biosynthesis, accumulation, and transport of flavonoids in stems and leaves of different S. miltiorrhiza phenotypes. The findings will provide a reference for flavonoid production and cultivar selection.
Subject terms: Molecular biology, Plant sciences
Introduction
Salvia miltiorrhiza Bunge is an important herb in Chinese medicine that was first recorded in Shennong’s herbal classics and is currently widely used to prevent and treat cardiovascular and cerebrovascular diseases1,2. Tanshinones and phenolic acids are two major groups with significant pharmacological constituents obtained from S. miltiorrhiza, which have important roles in traditional Chinese clinical practice and Western medicine3,4. Flavonoids are another active ingredient that are widespread in aboveground parts of S. miltiorrhiza5–7. Because aerial parts of S. miltiorrhiza, are nonmedicinal, aboveground parts are always discarded during harvesting, resulting in huge waste of resources and increased environmental pressure. However, owing to the high flavonoid content and associated antioxidant characteristics, aerial parts of S. miltiorrhiza can be developed into natural antioxidants and used in foods.
Flavonoids are a group of bioactive compounds found extensively in foodstuffs of plant origin, and regular consumption of flavonoids is associated with reduced risk of several chronic diseases9. Flavonoids are important secondary metabolites involved in plant growth and development but have also been studied in many plants because of excellent antioxidant and anticancer effects5,7. Flavonoids primarily include anthocyanins (red to purple pigments), flavonols (colorless to light yellow pigments), flavanols (colorless pigments that turn brown after oxidation), and procyanidins (PAS)8. In addition, flavonoid metabolite distribution and accumulation vary with plant species, tissues, organs, development stages, and growth conditions10,11.
The growing evidence of the versatile health benefits of flavonoids including anti-inflammatory, antioxidant, antiproliferative and anticancer activity8,11. Diets that include flavonoid containing products are currently recommended12, further studies are necessary for confirmation of the beneficial effects, such as determine their mechanisms of action at the molecular level, to lay a foundation for the rational promotion of diet rich in flavonoids. Biosynthesis of flavonoids shares the same upstream pathway as lignin and phenolic acid biosynthesis, which is derived from the phenylpropane pathway13. In addition, biosynthesis and accumulation of flavonoids in S. miltiorrhiza are influenced by a series of structural genes, including PAL, C4H, 4CL, CHS, CHI, FLS, FNS, DFR, F3H, F3′H, F3′5′H, ANS, 3GT, and 3AT, and regulatory transcription factors, including MYB, bHLH, bZiP, WD40, and WRKY, or transcriptional complexes (MYB-bHLH-WD) in the flavonoid metabolic pathway. Genes (e.g., HCT) from other pathways can also be involved. Although expression of those genes affects the formation of color in plant organs14,15, biological and abiotic factors, especially hormones and environmental factors, also regulate flavonoid biosynthesis16.
Color phenotypes of plants organs are associated with the biosynthesis and accumulation of flavonoids8,17,18. For example, a decrease in flavonoids/anthocyanins was the primary reason leaf color changed in purple-leaf tea plant19,20. In addition, the purple inner leaves of purple ornamental cabbage result from a high level of anthocyanin biosynthesis, a high level of chlorophyll degradation, and an extremely low level of chlorophyll biosynthesis18. In Suaeda salsa, two anthocyanins are responsible for red in leaves19, and in Ziziphus jujuba, flavonoids are the major differential metabolites determining leaf color20.
To better understand the mechanism of color change and the dynamic changes in related metabolites and genes expression levels, metabolic and transcriptional profiles of stems and leaves of S. miltiorrhiza purple and green phenotypes were investigated in this work. Differentially accumulated metabolites (DAMs), including flavonoids and phenolic acids, and differentially expressed genes (DEGs) in flavonoid biosynthesis pathways were determined. To further explore the biosynthesis, accumulation, and transport of flavonoids in aboveground tissues of S. miltiorrhiza, laser scanning confocal microscopy (LSCM) with 2-Aminoethyl diphenylborinate (NA) taining-induced fluorescence was performed. On the basis of the analyses, the biosynthesis, accumulation, and transport of flavonoids in stems and leaves of different phenotypes were determined. The work revealed dynamic changes in metabolites and molecular mechanisms regulating underlying color changes in S. miltiorrhiza stem and leaf.
Results
Comparison of morphological phenotypes
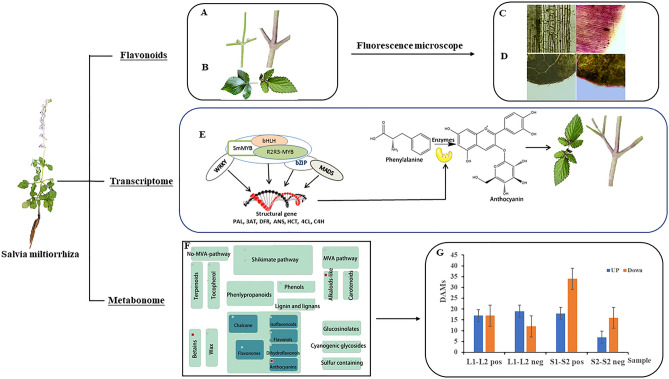
Morphological phenotypes of green and purple stems in S. miltiorrhiza were significantly different. Purple plants had a better agronomic phenotype than that of green plants, with tall plants, thick stems, and purple stem skin with more villi. In addition, the color of buds and leaves on purple plants was different from that of old leaves (Fig. 1A). The edges of young buds and leaves were purple, and the edge of old leaves was also purple. In the comparison of leaves with purple edges and completely green leaves, leaves with purple edges were yellowish green, sharply serrated edges were obvious, and some of the stems were purple. In addition, there was more fluff on the back (Fig. 1B). Microscopic observation indicated the pigment of purple stems primarily accumulated in several layers of cells below the epidermis (Fig. 1C). Therefore, stem skin tissues were observed by hand section, and edge tissues of green and purple leaves were compared. Purple pigment was primarily in the epidermal cells at leaf edges (Fig. 1D). Thus, the purple and green aboveground parts were taken as the materials, we analyzed them using metabonoics (Fig. 1F) and transcriptomics (Fig. 1E), and differentially expressed genes and metabolites were identified (Fig. 1G). In order to establish a dynamic accumulation model of biosynthesis pathways of S. miltiorrhiza using transcriptome and metabolome, and the biological transport pathway leading to accumulation and transport of purple pigment was investigated (Fig. 1).
Figure 1.
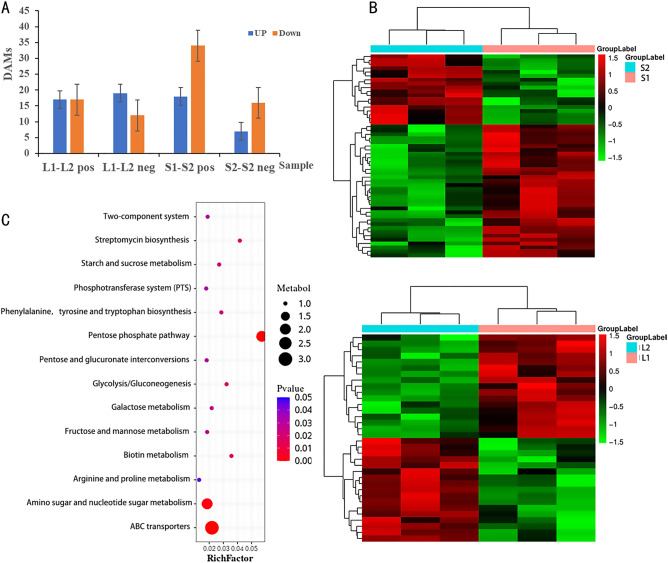
Response of S. miltiorrhiza Phenotypes to flavonoids. (A) Surface view of green and purple stem internode. (B) Surface view of green leaf and the leaves with purple edges. (C) Periderm tissue of green and purple stems under fluorescence microscope. (D) Periderm and mesophyll tissue of green leaves and the leaves with purple edges under fluorescence microscope. (E) A model to explain the internal mechanism of purple pigmentation in stems and leaves of S. miltiorrhiza. (F) The main secondary metabolic pathway in S. miltiorrhiza. (G) DAMs in different phenotypes of S. miltiorrhiza stems and leaves (L1: green leaf, L2: the leaves with purple edges; S1: green stem, S2: purple stem, pos indicated positive ion mode; neg indicated negative ion mode).
Genome sequencing, assembly, and annotation
A total of 76.95 GB of data were measured using a DNBSEQ sequencing platform to sequence the test materials (S1, S2, L1, L2). The clean data of each sample reached 6.3 GB; the Q20 value of obtained sequences was greater than 96%; the percentage of Q30 bases was ≥ 90%; and the GC content was more than 43% (Supplementary Table S2). A total of 40,077 simple sequence repeats (SSR) loci were obtained. The 19,216 unigene sequences included 93 repeat units, of which dinucleotide repeat was the dominant repeat type (55.27%), followed by mononucleotide repeat (15.5%) and trinucleotide repeat (23.1%). The dominant units of mononucleotide and dinucleotide repeats were A/T and AG/TC, respectively (Table 1). In the comparison with the reference database, 114,431 unigenes were successfully annotated, and 92,326 (80.68%) genes were annotated in the Non-Redundant Protein Sequence Database (NR), of which 60,704 (65.75%) genes were consistent with the characteristics of Salvia splendens and 3566 genes were unique to S. miltiorrhiza. The results indicated that the quality of sequencing data was high, and therefore, the data met the requirements of transcriptome analysis.
Table 1.
Transcriptome SSR repeat type and number.
| Repeats motifs | Duplicate times | Total | Ratio (%) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 4 | 5 | 6 | 7 | 8 | 9 | 10 | > 10 | |||
| Mono-nucleotide | – | – | – | – | – | – | 1445 | 6198 | 6198 | 15.5 |
| Di-nucleotide | – | – | 6791 | 4410 | 3540 | 2261 | 1468 | 3679 | 22,149 | 55.27 |
| Tri-nucleotide | – | 4980 | 2171 | 1013 | 607 | 141 | 93 | 214 | 9219 | 23.1 |
| Quad-nucleotide | – | 205 | 53 | 8 | 5 | 12 | 3 | 6 | 292 | 0.73 |
| Penta-nucleotide | 558 | 172 | 30 | 8 | 7 | 2 | 6 | 1 | 784 | 1.96 |
| Hexa-nucleotide | 1088 | 196 | 95 | 39 | 10 | 3 | 2 | 2 | 1435 | 3.58 |
| Total | 1676 | 5553 | 9140 | 5478 | 4169 | 2419 | 1572 | 10,100 | 40,077 | – |
| Ratio (%) | 4.18 | 13.9 | 22.8 | 13.7 | 10.4 | 6.04 | 3.92 | 25.2 | – | – |
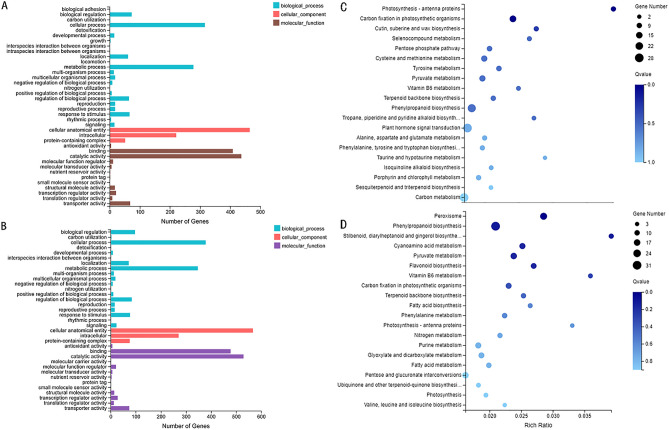
According to the criteria q-value ≤ 0.05 and log2 (fold-change) ≥ 1, 1299 DEGs were identified in S1–S2, including 666 up-regulated genes and 633 down-regulated genes, and 1091 DEGs were identified in L1–L2, including 393 up-regulated genes and 698 down-regulated genes (Fig. S1). Gene Ontology analysis (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed on DEGs. In the GO analysis of the S1–S2 group, 481, 572, and 805 DEGs were assigned to the categories biological processes, molecular functions, and cellular components, respectively, with DEGs further classified into 35 functional subcategories (Fig. 2A). In the L1–L2 group, 402, 469, and 676 DEGs were assigned to the categories biological processes, cellular components, and molecular functions, respectively, with DEGs further classified into 38 functional subcategories (Fig. 2B).
Figure 2.
(A) GO functional classification of differentially expressed genes in L1 (green leaf), and L2 (the leaves with purple edges) in S. miltiorrhiza. (B) GO functional classification of differentially expressed genes in S1 (green stem), and S2 (purple stem). (C) Scatter plots of the KEGG pathway enrichment statistics in L1–L2. (D) Scatter plots of the KEGG pathway enrichment statistics in S1–S2.
To understand biological functions and gene interactions, DEGs were annotated by the KEGG database (www.kegg.jp/kegg/kegg1.html) 21. In the S1–S2 group (Fig. 2C), 473 of 1299 DEGs were in 122 KEGG pathways, including cellular processes, environmental information processing, genetic information processing, metabolism, and organismal systems. In the L1–L2 group (Fig. 2D), 438 of 1091 DEGs were in 113 KEGG pathways. In the S1-S2 group, the main enriched metabolic process was phenylpropanoid biosynthesis, and twelve DEGs were involved in flavonoid biosynthesis. In further analysis of the DEGs, six Glutathione S-transferase (GST) genes, nine ABC transporter genes, 22 MATE genes, and three SNARE (soluble NSF attachment protein receptor) genes were also detected, which may have important roles in transporting anthocyanin to plant vacuoles7. In the L1–L2 group, the main enriched metabolic process was phenylpropanoid biosynthesis and carbon metabolism. Further analysis revealed eight GST genes and seven ABC transporters.
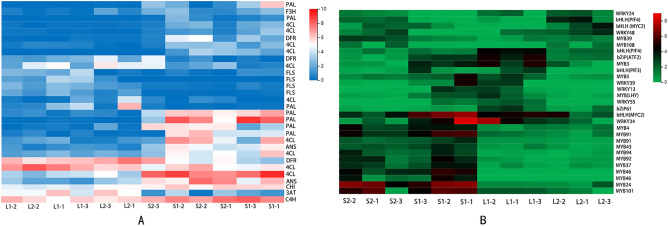
Anthocyanins and flavonoid secondary metabolites are the main components that affect the color of petals, stems, leaves, and fruits6,7. In the enrichment analysis, PAL, C4H/CYP73A, 4CL, DFR, ANS, FLS, CHI, F3H, and 3AT were significantly different in the flavonoid biosynthesis pathway. Hierarchical Cluster Analysis (HCA) was performed on DEGs with a pheatmap package (Fig. 3A, Supplementary Table S3).
Figure 3.
(A) Expression patterns of flavonoids biosynthetic genes in stems and leaves samples of S. miltiorrhiza. (B) Expression of flavonoids biosynthetic transcription factors in nine samples of S. miltiorrhiza. The horizontal axis represents log2 (FPKM + 1) of the sample and the vertical axis represents genes. L1(green leaf), L2(the leaves with purple edges), S1(green stem), S2 (purple stem).
Flavonoid contents in plants are strictly regulated by complex regulatory networks. Transcription factors (TFs), including members of MYB, bHLH, MADS, WRKY, bZiP, and WD40 families, have important roles in regulating plant flavonoid biosynthesis 22. In this study, differential expression levels of MYB, bHLH, bZiP, and WRKY were detected. It is worth mentioning that the MYB family had the largest number and the greatest differences among all TF genes detected. The differential expression of TF genes was most obvious in stem tissue (Supplementary Table S4, Fig. 3B). Transcription factors MYC2 and WRKY34 were expressed at higher levels in green stems than in purple stems. The findings indicated that the TFs were closely associated with phenotypic differences in color in different tissues.
Metabolomic analysis
A representative sample (S1, S2, L1, L2) was selected from each group (three biological replicates) of samples to display the base peak ion Base Peak Chromatogram (BPC) chromatogram. The total ion flow chromatogram of different phenotypes of stems and leaves of S. miltiorrhiza is shown in Fig. S2. In the BPC diagram, the peak results of the tested samples were good, the peak shape was good, and the peak area was large. However, there were some differences among the overall contours of the four samples. To reveal the component differences between the different phenotypes of S. miltiorrhiza, metabolite separation and detection were conducted. After data preprocessing, in leaves, 10,636 metabolites were identified in positive ion mode, and 5231 metabolites were identified in negative ion mode. In stems, 6264 metabolites were identified in positive ion mode, and 3503 metabolites were identified in negative ion mode (Table 2).
Table 2.
Statistical table of compound detection.
| Mode | Number of metabolites | Number of metabolites with identification information | |
|---|---|---|---|
| Stem | Pos | 6264 | 816 |
| Neg | 3503 | 466 | |
| Leaf | Pos | 10,636 | 944 |
| Neg | 5231 | 524 |
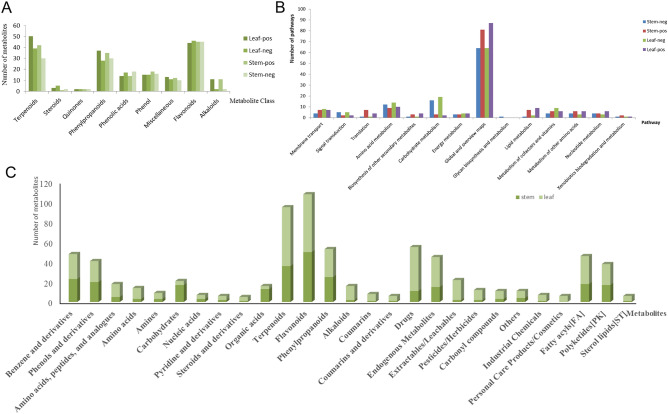
Metabolites identified in S. miltiorrhiza leaves were classified in the Beijing Genomics Institute (BGI) library. Of the metabolites, 44 were flavonoids, 50 were terpenoids, 37 were phenylpropanoids, 14 were phenolic acids, 15 were phenols, 13 were classified as miscellaneous, and 3 were steroids. Eleven metabolites were alkaloids (alkaloids and glycosides), and two were quinones (quinones and glycosides). Metabolites identified in S. miltiorrhiza stems were also classified in the BGI library. Of the metabolites, 45 were flavonoids and glycosides, 42 were terpenoids and glycosides, 35 were phenylpropanoids and glycosides, 18 were phenolic acids and glycosides, 14 were phenols and glycosides, 12 were in other classes, and one was in the steroids and glycosides group. Eleven metabolites were alkaloids and glycosides, and two metabolites were quinones and glycosides. Thus, flavonoids and glycosides, terpenoids and glycosides, and phenylpropanoids and glycosides were the main metabolites identified in all samples of aboveground parts of S. miltiorrhiza (Fig. 4A).
Figure 4.
(A) Metabolite classification bar chart in BGI library. (B) Bar chart of metabolite KEGG function annotation. (C) Bar chart of metabolite classification.
Metabolites were classified and annotated in Human Metabolome Database (HMDB and KEGG databases23. All metabolites were classified in three main pathways in the KEGG database, which were environmental information processing and metabolism and genetic information processing. In leaf tissue, environmental information processing had two branches: membrane transport (77.78%) and signal transduction (22.22%). Genetic information processing also had two branches: translation (80%) and folding, sorting, and degradation (20%). All metabolites accounted for the largest proportion (65.93%) in global and overview maps. In stem tissue of S. miltiorrhiza, two branches under environmental information processing included membrane transport (77.78%) and signal transduction (22.22%). Under genetic information processing, there was a branch of translation. All metabolites accounted for the largest proportion (65.32%) in global and overview maps (Fig. 4B). In leaves, all metabolites were divided into four categories: compounds with biological roles (31.42%), phytochemical compounds (41.09%), others (11.18%), and lipids (16.31%). Flavonoids and terpenoids accounted for large proportions of phytochemical compounds, with 37.5% and 36.03%, respectively. In stems, terpenoids (32.8%), flavonoids (32.22%), and phenylpropanoids (15.56%) were the three most abundant phytochemical compounds (Fig. 4C).
The DAMs between green and purple cultivars were determined according to the criteria of fold-change ≥ 1.2 or ≤ 0.83 and P-value < 0.05. In green stems and purple stems, 23 DAMs were screened, of which seven were up-regulated and 16 were down-regulated (Supplementary Table S5). In green leaves and leaves with purple edges, 34 DAMs were screened, of which 17 were up-regulated and 17 were down-regulated (Supplementary Table S6, Fig. 5A, Fig. S3). Then, an HCA was performed on the accumulation pattern of metabolites among different samples24. The HCA revealed two clusters of metabolites, with one the metabolites before the color change (S1, L1) and the other the metabolites after the color change (S2, L2) (Fig. 5B).
Figure 5.
(A) Statistical map of differential metabolites (DAMs). (B) The Hierarchical cluster analysis of differential accumulated metabolites (DAMs) in L1–L2, S1–S2 of S. miltiorrhiza. (C) Bubble Diagram of metabolic pathway enrichment analysis in L1–L2.
To better identify which pathways are enriched between different DAMs, the metabolic pathways of all DAMs were enriched and analyzed based on the KEGG database. Forty-eight metabolites were enriched in 46 metabolic pathways, and 36 metabolites were enriched in the biosynthesis of secondary metabolites. The pathways with significant enrichment (P < 0.05) were selected. In stems, seven DAMs were annotated in the metabolic pathway (guanosine, aflatoxin G2, estriol, arachidonic acid, 5'-S-methyl-5ʹ-thioadenosine, 4-coumaric acid, and sinapic acid), and four DAMs were annotated in the secondary metabolite biosynthesis pathway (coumarin, 4-coumaric acid, sinapic acid, and phlorizin) (Table 3). For leaves, bubble plots show significant enrichment of DAMS (P < 0.05) in 14 pathways. The largest proportion of DAMs was annotated to the pentose phosphate pathway. The largest number of DAMs was annotated to ABC transporters, accounting for 17.65%. The metabolites (d-(+)-glucose, biotin, d-(+)-xylose), d-(+)-glucose, and d-(+)-xylose were involved in the pentose phosphate pathway and antibiotic biosynthesis. The differential metabolite d-(−)-quinic acid was involved in the metabolic pathway of phenylalanine, tyrosine and tryptophan biosynthesis, and d-(+)-glucose was involved in almost all other metabolic pathways (Fig. 5C).
Table 3.
Metabolic pathway enrichment with Pvalue < 0.05 in S1–S2.
| Pathway | Count | P value | DAMs names |
|---|---|---|---|
| Metabolic pathways | 7 | 0.0018 | Guanosine, aflatoxin G2, estriol, arachidonic acid, 5ʹ-S-methyl-5ʹ-thioadenosine, 4-coumaric acid, sinapic acid |
| Biosynthesis of secondary metabolites | 4 | 0.0113 | Coumarin, 4-coumaric acid, sinapic acid, phlorizin |
| Cysteine and methionine metabolism | 1 | 0.0463 | 5ʹ-S-methyl-5ʹ-thioadenosine |
| Arachidonic acid metabolism | 1 | 0.0549 | Arachidonic acid |
| Purine metabolism | 1 | 0.069 | Guanosine |
| ABC transporters | 1 | 0.098 | Guanosine |
| Degradation of aromatic compounds | 1 | 0.1804 | 4-Coumaric acid |
| Biosynthesis of antibiotics | 1 | 0.5483 | Aflatoxin G2 |
Network of differentially accumulated metabolites regulated by differentially expressed genes
To further correlate gene expression patterns with metabolite accumulation, coexpression analysis was applied to metabolome and transcriptome data from S. miltiorrhiza purple and green stems and leaves. Expression levels of PAL, 3AT, 4CL, ANS, HCT, MYB43, MYB101, MYB108, MYB94, WRKY24, and WRKY48 were higher in purple samples, whereas expression levels of DFR and CYP73A were higher in green samples (Supplementary Tables S3, S4). It was hypothesized that those key genes and transcription factors were involved in the regulation of flavonoid and phenolic acid biosynthesis, thereby affecting the color of S. miltiorrhiza stem and leaf. The anthocyanin accumulation in purple characters was closely related to the high expression of the synthases involved in anthocyanin biology.
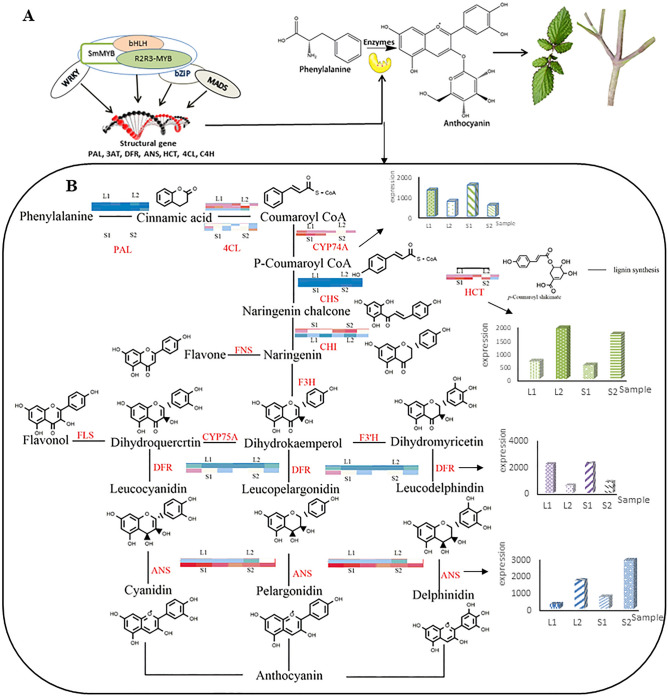
In the main phenylpropanoid and flavone biosynthesis pathway, strong activation and expression of all structural genes eventually led to the production of many anthocyanins. In the flavonoid biosynthesis pathway, under the catalysis of MYB, bHLH, MADS, WRKY, and bZiP families, yields of flavones and anthocyanins also increased on a large scale. Anthocyanins are a group of water-soluble pigments that help to form various colors of plants. Flavonoids, especially those in the anthocyanin biosynthesis pathway, led to the accumulation of purple pigment (Fig. 6).
Figure 6.
Molecular regulation of color phenotype characters of S. miltiorrhiza. (A) A working model to explain the internal mechanism of purple pigmentation in the aboveground part of S. miltiorrhiza. (B) Schematic diagram of flavonoids biosynthesis and regulation in S. miltiorrhiza. The redder the color of the color block, the higher the expression, and the bluer the color, the lower the expression.
Reverse-transcription quantitative PCR of transcriptomic data
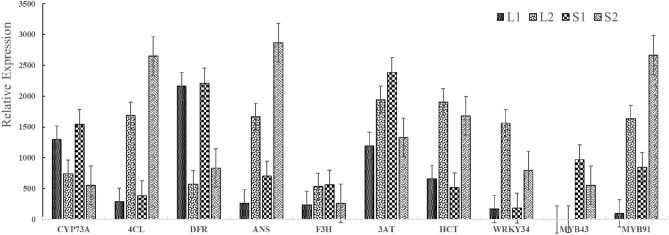
To validate transcriptome information, 10 DEGs were selected to validate the sequencing results. The comparative CT method (2−ΔΔCTmethod) was used to quantify gene expression. Reverse-transcription quantitative PCR (RT-qPCR) indicated that six genes had higher expression levels and four had lower expression levels in the purple phenotype than in the green phenotype (Fig. 7). The RT-qPCR results were consistent with those obtained with RNA-seq.
Figure 7.
Expression levels of 10 genes involved in the flavonoids biosynthetic pathway by qRT-PCR analysis.
Distribution dynamics of flavonoids by laser scanning confocal microscopy
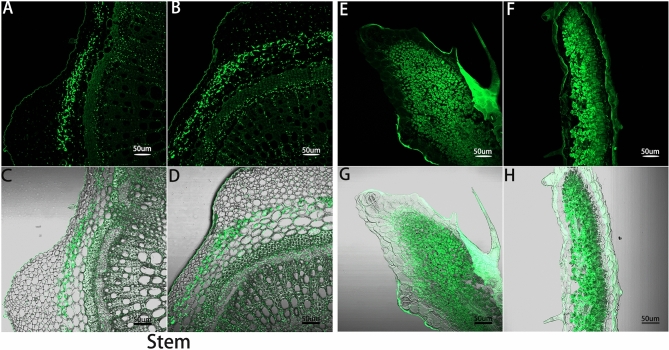
To investigate locations of flavonoids in purple and green characters from S. miltiorrhiza, frozen sections were prepared and observed. At the excitation wavelength of 488 nm, samples showed chloroplast autofluorescence before staining. The interference was eliminated by subtracting the background signal. After staining with NA solution, epidermis, glandular hairs, and thick horn tissue cells of stems in S. miltiorrhiza showed strong fluorescence. In addition, a small amount of fluorescence also appeared in the phloem. The observations suggested that flavonoids were biosynthesized and accumulated in epidermis, villi, collenchyma, phloem, and fascicle of S. miltiorrhiza (Fig. 8A–D). Flavonoids in S. miltiorrhiza leaves were primarily distributed in external tissues such as glandular hair and epidermis (Fig. 8E–H), possibly to prevent microbial invasion and UV radiation damage, and also accumulated in palisade and spongy tissues 25. Because methanol is volatile and strongly solubilizes flavonoids, it acts as a solvent of color developing agent. Therefore, flavonoids are dissolved and diffused in the dyeing process, and a small amount of fluorescence can appear in some parts. The content of flavonoids in the green phenotype was significantly higher than that in purple phenotype, which might be because the lower expression level of ANS resulted in a decrease in anthocyanin biosynthesis and accumulation in green samples but an increase in flavonoid biosynthesis and accumulation.
Figure 8.
Leaves and stems were stained with NA solution and observed with CLSM (50 µm). The figure showed the distribution of flavonoids in the cross section of S. miltiorrhiza after staining. (A,B,E,F) indicated the distribution of flavonoids in the AF488 channel. (C,D,G,H) Indicated the distribution of flavonoids in superimposed bright field of AF488 channel. (A,C) Indicated the purple stem. (B,D) Indicated the green stem; (E,G) indicated the leaf with purple edge. (F,H) Indicated the green leaf.
Discussion
The color of plant tissues and organs is determined primarily by the accumulation of flavonoids. Flavonoids, phenolic acids, and lignin are all in downstream branches of the phenylpropane metabolic pathway20,26. Expression levels of the main structural genes or TFs in the phenylpropane metabolic pathway can determine final flavonoid biosynthesis and then affect formation of color phenotypes. In this study, five key phenylpropane structural genes (PAL, 4CL, ANS, 3AT, HCT) had higher expression in purple than in green samples, and two key anthocyanin structural genes (DFR, CYP73A/C4H) had higher expression in green than in purple samples. As key upstream enzyme genes of the phenylpropane pathway, PAL, 4CL, and CYP73A affect proportions in the metabolic flow of flavonoids, phenolic acids, and lignin1. The key enzymes in anthocyanins biosynthesis are DFR, ANS, and 3AT6. Overexpression of SmANS also increases anthocyanin accumulation and alters phenolic acids content in S. miltiorrhiza17. The genes ANS, DFR, and F3′5′H also have key roles in regulating the biosynthesis of anthocyanins in purple stems of Astragalus membranaceus13. The 3AT from Perilla frutescens can be used as a new molecular tool to modify anthocyanin structure and further increase flower color by recombinant DNA technology27. The gene HCT mainly controls the metabolic flow between lignin and the flavonoid branches. In Arabidopsis, HCT can inhibit the synthesis of lignin, which redirects metabolic flux to flavonoids via chalcone synthase activity28. In S. miltiorrhiza leaves in this study, expression level of HCT and content of flavonoids (rhusflavanone, herbacetin, and ipriflavone) were significantly higher in the purple phenotype than in the green phenotype. Therefore, HCT might be a positive regulator of flavonoid biosynthesis and thereby affect the color of organs and tissues in S. miltiorrhiza.
The structural genes that catalyze the formation of flavonoids in S. miltiorrhiza, are regulated by single or multiple TFs as well as complexes formed between TFs14. Transcription factors MYB, AP2/ERFs, WRKY, bHLH, MADS, WD40, zinc finger, NACs, bZiP, and LBD may be involved in the formation of white and purple flowers in S. miltiorrhiza7. In this work, MYB, WRKY, bHLH, and bZiP also had important roles in pigment formation of S. miltiorrhiza stems and leaves. The regulatory network of MYB in S. miltiorrhiza is important and complex and therefore can provide useful information to improve the growth and defense ability of S. miltiorrhiza as well as the production of active compounds. Transcription factors MYB58 and MYB63 can activate PAL, C4H, 4CL, and other genes involved in lignin biosynthesis in Arabidopsis29. In S. miltiorrhiza, MYB37 is in the same subgroup as MYB58 and MYB63. Therefore, it was inferred that MYB37 had a similar role in lignin biosynthesis30. Arabidopsis thaliana MYB4 can interfere with the transcriptional activity of MBW complexes or inhibit biosynthesis of phenylalanine by repressing expression of the gene encoding arogenate dehydratase 631, which catalyzes the last step in phenylalanine biosynthesis that produces the precursor of flavonoid biosynthesis. In the current study, the green phenotype had high content of MYB4e, indicating that the gene had an obvious inhibitory effect on phenylalanine and thereby affected biosynthesis of flavonoids. The color variation in sweet gum young and senescent leaves is attributed to the composition of anthocyanidins produced by transcriptional regulation of F3′H and F3′5′H by MYB5, MYB113, and MYB12332. Therefore, because the MYB3 and MYB5 in S. miltiorrhiza are in the same group, the two TFs are also likely involved in anthocyanin biosynthesis28. The TFs MYB5-like and bHLH have roles in the first step of flavonoid production in flowers and immature fruits, and expression patterns of those TFs may be the key to determine differences in flavonoids in pomegranate flowers and fruits33. In addition, MYB37 has an important and positive role in plant response to abscisic acid (ABA) and drought stress34. The interaction of TFs MYB21 and MYB24 affects the regulation of jasmonic acid35, and SmMYB39 participates in regulation of the rosemary acid pathway by inhibiting the transcription of key enzyme factors36.
The WRKY family is one of the largest TF families in plants, and it regulates the growth and development of plants37. The TF WRKY55 positively regulates leaf senescence and defense by regulating the transcription of reactive oxygen species (ROS) and salicylic acid (SA) biosynthesis-related genes in Arabidopsis38. The TF WRKY34 interacts with the VQ20 protein to regulate pollen development and function39. The bHLH family TF PIF4 is necessary for plants to adapt to light and high ambient temperature, and interaction between PIF4 and PIF4 interacting proteins establishes a strong network for plants to respond to changes in the environment40. In addition, phytochrome interaction factors PIF4 and PIF5 negatively regulate anthocyanin synthesis in Arabidopsis seedlings under red light41. In plant response to jasmonic acid, MYC2 is the core TF, and overexpression of MYC2 can promote the production of phenolic acids in S. miltiorrhiza42. In this study, the differential expression of those genes in S. miltiorrhiza purple and green phenotypes was also significant. However, the complexity and integrity of biological processes, the complex regulatory network of plant anthocyanin metabolism, and various genetic and environmental factors affect the whole biosynthesis process.
Among the different metabolites, multiple carbohydrates were highly enriched, including d-glucose, l-rhamnose, d-galactose, d-xylose, and l-arabinose. Glycosylated forms of flavonoids and anthocyanins in flavonoids appear most often and are highly soluble in water. Therefore, hydrolysis may occur and other substances may form, resulting in a decrease in anthocyanin content. Moreover, the high expression of the glycosyltransferase 3AT could explain how glucose and flavonoids form anthocyanins through glycosylation, which eventually led to the difference in anthocyanin components between the two S. miltiorrhiza phenotypes. The DEGs and DAMs were more abundant in stems than in leaves, and expression in purple stems was relatively high. Flavonoids can be selectively transported from one organ to another in plants and from sites of synthesis to other parts43. Flavonoid distribution may be mediated by the action of MRP/ABC transporters rather than passive diffusion44. In this study, GST, ABC transporter, MATE, and SNARE genes were detected, which may have important roles in transporting anthocyanins to plant vacuoles. Therefore, some flavonoids and anthocyanins were likely transported to leaves by ABC transporters and plant hormone signal transduction, resulting in purple characters on the edges of some leaves. However, some anthocyanins might have been hydrolyzed during transport, which could also explain the accumulation of many of differential metabolites in the ABC transporter pathway and the variety of differential carbohydrate metabolites.
Materials and methods
Plant materials
Salvia miltiorrhiza seedlings were provided by Shaanxi Tasly Plant Medicine Co., Ltd. (Shangluo, China) and were identified by Prof. Yuan Chen (Gansu Agricultural University, Lanzhou City, Gansu Province). Seedlings were planted in the herbal garden of Gansu Agricultural University (Longitude: 103.700153, latitude: 36.090657). The collection of plant materials complied with relevant institutional, national, and international guidelines and legislation. Completely green stems (S1), purple stems (S2), completely green leaves (L1), and leaves with purple edges (L2) were collected in a full-blossom period, frozen with liquid nitrogen, and stored at − 80 °C. Samples from six individual plants were pooled as one biological replicate, The voucher specimens are deposited in the herbarium of medicinal plants of Gansu Agricultural University (Nos. GAUAB-SM-20210622-S1, GAUAB-SM-20210622-S2, GAUAB-SM-20210622-L1, and GAUAB-SM-20210622-L2). Three biological replicates were used in transcriptome and metabolome analyses conducted at the Shenzhen Genomics Institute (BGI) Co., Ltd.
Transcriptomic sequencing
Because original sequence data obtained by sequencing could not be directly used for subsequent information analysis, the filtering software SOA Pnuke45 was used to filter out low-quality, joint pollution, and reads with high content of unknown base N (> 5%). The filtered data were considered “clean reads”46. The clean reads were assembled to obtain unigenes, and the unigenes were analyzed for function annotation, predicted coding area, and SSRs. The expression amount of each sample was calculated on the basis of all unigenes. The DEGs were screened according to q-value (adjusted P-value) ≤ 0.05. Based on the principle of negative binomial distribution, DEseq2 software was used for differential analysis47. The screening threshold was q-value ≤ 0.05 and log2 |(fold-change)|≥ 148.
Metabolomics analysis
Untargeted metabolomics analysis was performed using LC–MS/MS. The data obtained were processed by Compound Discoverer 3.1 (Thermo Fisher Scientific, USA)49. The metabolomics software package metaX and metabolomic information analysis processing were used for data preprocessing, statistical analysis, metabolite classification, and functional annotation50. This work used VIP values of the first two principal components in a multivariate Partial Least Squares Discriminant Analysis (PLS-DA) model51, combined with fold-change and t-test of univariate analysis to identify DAMs according to fold-change ≥ 1.2 or ≤ 0.83 and P < 0.0552.
Combined analysis to reveal biosynthesis of flavonoids
Each metabolite may be regulated by multiple genes, or multiple metabolites may be regulated by one gene. The differential genes obtained by sequencing the transcriptome were targeted at the utilization site(http://www.plabipd.de/portal/mercator-sequence-annotation). The provided online annotation tool performed functional annotation classification, obtained a mapping file with similar structure, imported log2 fold-change data of DEGs, and then used MAPMAN software for joint analysis of transcriptional and metabolomic data53. Last, a working model was proposed to explain the internal mechanism of purple pigmentation in aboveground parts of S. miltiorrhiza, and a schematic diagram of flavonoid biosynthesis and regulation in S. miltiorrhiza was constructed.
Reverse-transcription quantitative PCR expression analysis
Total RNA of S. miltiorrhiza stem and leaf was extracted according to a DP441 RNA Prep Prue Polysaccharide Polyphenol Plant Total RNA Extraction Kit (Tiangen Biotech, Beijing, China) and reverse-transcribed according to a PrimeScriptTM RT Reagent Kit with gDNA Eraser (Code No. RR047A; Takara Biotech, Beijing, China). The cDNA was used as the template to measure gene expression. The specific primers for genes involved in flavonoid biosynthesis and the S. miltiorrhiza actin gene (internal control) are listed in Supplementary Table S1. Reverse-transcription quantitative polymerase chain reaction (RT-qPCR) was conducted by a real-time PCR system (QuantStudio5) using TB Green® Premix ExT aq™ II (Code No. RR820A; Takara Biotech). Three biological replicates were used for qRT-PCR of each genes. The comparative CT method (2−ΔΔCTmethod) was used to quantify gene expression, firstly, for all test samples and calibration samples, CT values of actin gene are used to normalize CT values of target genes: ΔCT(test) = CT(target, test) − CT(ref, test), ΔCT(calibrator) = CT(target, calibrator) − CT(actin, calibrator). Secondly, the ΔCT value is the same as that of the test sample ΔCT value: ΔΔCT = ΔCT(test) − ΔCT(calibrator). Finally, the comparative CT method (2−ΔΔCT method) was used to quantify gene expression. Statistical analysis was performed using Excel 2020 software (Microsoft Office, USA).
Laser scanning confocal microscopy
Microstructure and distribution of flavonoids in aboveground parts of different S. miltiorrhiza phenotypes were evaluated by an LSCM method54. Flavonoids react with NA reagent (0.1% 2-aminoethyl diphenylborinate (w/v) in 80% methanol) and produce fluorescence55. The middle part of a stem or leave was cut into small pieces (width of 3 mm), which were then cut into 30-μm sections with a freezing microtome (CM1950; Leica, Germany). Sections required treatment to induce fluorescence. A drop of self-made phosphoric acid buffer solution (with the addition of 1% NaCl (w/v), pH 6.5) was added to the sections, followed by a drop of NA solution. After incubation for 5 min, sections were analyzed with LSCM (LSCM 800; Carl Zeiss, Germany).
Conclusions
Contents of flavonoids and anthocyanins in the purple phenotype depended on the up-regulation of genes associated with biosynthesis of flavonoids and anthocyanins. The difference in color between purple and green phenotypes was the result of differential anthocyanin accumulation and differential gene expression. Those results and differences in morphological phenotypes suggested that regulation of structural genes in the flavonoid biosynthesis pathway promoted anthocyanin accumulation and contributed to the formation of the purple phenotype. The results clarify the important role of flavonoids in developmental processes and provide a basis for further understanding of organ coloring and the networks regulating flavonoid metabolism. Overall, the work revealed the dynamic changes in metabolites and the molecular regulation mechanisms underlying the color change in stem and leaf in S. miltiorrhiza and thus can also provide valuable theoretical support for further exploration of new medicines and health foods with flavonoids.
Supplementary Information
Acknowledgements
This work was supported by Scientific Research Start-up Funds for Openly recruited Doctors of Gansu Agricultural University (GAU-KYQD-2019-21).
Author contributions
H.L. conceived and designed the manuscript. H.Y. wrote the manuscript. H.Y. edited the manuscript. H.L., Q.L. and H.Y. performed the language editing and gave final approval of manuscript. All authors read and approved the final manuscript.
Data availability
The RNA-seq data sets for the transcriptome analysis were available at the Genome Sequence Archive in NCBI, under Biological Project ID: PRJNA796114. that are publicly accessible at https://www.ncbi.nlm.nih.gov/bioproject/PRJNA796114. The raw values obtained from the metabolomic data sets were available in Supplemental Data. If any data sets are unavailable through the links stated above, they can be obtained from the corresponding author on request.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Hongyan Li, Email: lihongyan2019@126.com.
Qian Li, Email: liqian1984@gsau.edu.cn.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-022-21517-5.
References
- 1.Ma P, Liu J, Zhang C, Liang Z. Regulation of water-soluble phenolic acid biosynthesis in S. miltiorrhiza Bunge. Appl. Biochem. Biotechnol. 2013;170:1253–1262. doi: 10.1007/s12010-013-0265-4. [DOI] [PubMed] [Google Scholar]
- 2.Wu S, et al. Protocatechuic aldehyde from Salvia miltiorrhiza exhibits an anti-inflammatory effect through inhibiting MAPK signalling pathway. BMC Complement. Med. Ther. 2020;20:1–9. doi: 10.1186/s12906-020-03090-4. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 3.Wu D, Huo M, Chen X, Zhang Y, Qiao Y. Mechanism of tanshinones and phenolic acids from Danshen in the treatment of coronary heart disease based on co-expression network. BMC Complement. Med. Ther. 2020;20:28. doi: 10.1186/s12906-019-2712-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yu H, et al. Systematic analysis of kelch repeat f-box (KFB) protein gene family and identification of phenolic acid regulation members in S. miltiorrhiza Bunge. Genes. 2020;11:557. doi: 10.3390/genes11050557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ravipati AS, et al. Antioxidant and anti-inflammatory activities of selected Chinese medicinal plants and their relation with antioxidant content. BMC Complement. Altern. Med. 2012;12:173. doi: 10.1186/1472-6882-12-173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Deng Y, Li C, Li H, Lu S. Identification and characterization of flavonoid biosynthetic enzyme genes in Salvia miltiorrhiza (Lamiaceae) Molecules. 2018;23:1467. doi: 10.3390/molecules23061467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Jiang T, et al. Integrated metabolomic and transcriptomic analysis of the anthocyanin regulatory networks in S. miltiorrhiza Bge. flowers. BMC Plant Biol. 2020;20:349. doi: 10.1186/s12870-020-02553-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Shen J, et al. Metabolic analyses reveal different mechanisms of leaf color change in two purple-leaf tea plant (Camellia sinensis L.) cultivars. Hortic. Res. 2018;5:7. doi: 10.1038/s41438-017-0010-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhang Z, Zhang YA. Transcriptional network makes normal tomato fruit not purple. Mol. Plant. 2020;13:11–13. doi: 10.1016/j.molp.2019.12.012. [DOI] [PubMed] [Google Scholar]
- 10.Liu Y, et al. Integrative analysis of metabolome and transcriptome reveals the mechanism of color formation in pepper fruit (Capsicum annuum L.) Food Chem. 2020;306:125629. doi: 10.1016/j.foodchem.2019. [DOI] [PubMed] [Google Scholar]
- 11.Petrussa E, et al. Plant flavonoids—bioSynthesis, transport and involvement in stress responses. Int. J. Mol. Sci. 2013;14:14950–14973. doi: 10.3390/ijms140714950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Tutel'ian VA, Lashneva NV. Biologically active substances of plant origin. Flavonols and flavones: Prevalence, dietary sourses and consumption. Vopr. Pitan. 2013;82:4–22. [PubMed] [Google Scholar]
- 13.Dong N, Lin H. Contribution of phenylpropanoid metabolism to plant development and plant-environment interactions. J. Integr. Plant Biol. 2021;63:180–209. doi: 10.1111/jipb.13054. [DOI] [PubMed] [Google Scholar]
- 14.Li H, et al. Overexpression of SmANS enhances anthocyanin accumulation and alters phenolic acids content in Salvia miltiorrhiza and Salvia miltiorrhiza Bge f. alba plantlets. J. Mol. Sci. 2019;20:2225. doi: 10.3390/ijms20092225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Albert NW, et al. A conserved network of transcriptional activators and repressors regulates anthocyanin pigmentation in eudicots. Plant Cell. 2014;26:962–980. doi: 10.1105/tpc.113.122069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mintz-Oron S, et al. Gene expression and metabolism in tomato fruit surface tissues. Plant Physiol. 2014;147:823–851. doi: 10.1104/pp.108.116004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cho K, et al. Network analysis of the metabolome and transcriptome reveals novel regulation of potato pigmentation. J. Exp. Bot. 2016;67:1519–1533. doi: 10.1093/jxb/erv549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jin S, et al. Transcriptome profiling of two contrasting ornamental cabbage (Brassica oleracea var. acephala) lines provides insights into purple and white inner leaf pigmentation. BMC Genomics. 2018;19:797. doi: 10.1186/s12864-018-5199-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wang X, Bai J, Wang W, Zhang G. Leaf metabolites profiling between red and green phenotypes of Suaeda salsa by widely targeted metabolomics. Funct. Plant Biol. 2019;46:845–856. doi: 10.1071/FP18182. [DOI] [PubMed] [Google Scholar]
- 20.Li S, et al. Metabolic and transcriptomic analyses reveal different metabolite biosynthesis profiles between leaf buds and mature leaves in Ziziphus jujuba Mill. Food Chem. 2021;347:129005. doi: 10.1016/j.foodchem.2021.129005. [DOI] [PubMed] [Google Scholar]
- 21.Kanehisa M, et al. KEGG: Integrating viruses and cellular organisms. Nucleic Acids Res. 2021;49:D545–D551. doi: 10.1093/nar/gkaa970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Luo F, et al. Transcriptome profiling reveals the roles of pigment mechanisms in postharvest broccoli yellowing. Hortic. Res. 2019;6:74. doi: 10.1038/s41438-019-0155-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wishart DS, et al. HMDB 4.0: The human metabolome database for 2018. Nucleic Acids Res. 2018;46:D608–D617. doi: 10.1093/nar/gkx1089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Caesar L, Kvalheim O, Cech N. Hierarchical cluster analysis of technical replicates to identify interferents in untargeted mass spectrometry metabolomics. Anal. Chim. Acta. 2018;1021:69–77. doi: 10.1016/j.aca.2018.03.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ferreyra M, Serra P, Casati P. Recent advances on the roles of flavonoids as plant protective molecules after UV and high light exposure. Physiol. Plant. 2021;173:736–749. doi: 10.1111/ppl.13543. [DOI] [PubMed] [Google Scholar]
- 26.Li H, et al. Molecular characterization and expression analysis of dihydroflavonol 4-reductase (DFR) gene in Saussurea medusa. Mol. Biol. Rep. 2012;39:2991–2999. doi: 10.1007/s11033-011-1061-2. [DOI] [PubMed] [Google Scholar]
- 27.Yonekura-Sakakibara K, et al. Molecular and biochemical characterization of a novel hydroxycinnamoyl-CoA: Anthocyanin 3-O-glucoside-6"-O-acyltransferase from Perilla frutescens. Plant Cell Physiol. 2000;41:495–502. doi: 10.1093/pcp/41.4.495. [DOI] [PubMed] [Google Scholar]
- 28.Besseau S, et al. Flavonoid accumulation in Arabidopsis repressed in lignin synthesis affects auxin transport and plant growth. Plant Cell. 2007;19:148–162. doi: 10.1105/tpc.106.044495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zhou J, Lee C, Zhong R, Ye ZH. MYB58 and MYB63 are transcriptional activators of the lignin biosynthetic pathway during secondary cell wall formation in Arabidopsis. Plant Cell. 2009;21:248–266. doi: 10.1105/tpc.108.063321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Li C, Lu S. Genome-wide characterization and comparative analysis of R2R3-MYB transcription factors shows the complexity of MYB-associated regulatory networks in Salvia miltiorrhiza. BMC Genomics. 2014;15:277. doi: 10.1186/1471-2164-15-277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wang X, et al. Arabidopsis MYB4 plays dual roles in flavonoid biosynthesis. Plant J. 2020;101:637–652. doi: 10.1111/tpj.14570. [DOI] [PubMed] [Google Scholar]
- 32.Wen C, Tsao N, Wang S, Chu F. Color variation in young and senescent leaves of Formosan sweet gum (Liquidambar formosana) by the gene regulation of anthocyanidin biosynthesis. Physiol. Plant. 2021;172:1750–1763. doi: 10.1111/ppl.13385. [DOI] [PubMed] [Google Scholar]
- 33.Arlotta C, et al. MYB5-like and bHLH influence flavonoid composition in pomegranate. Plant Sci. 2020;298:110563. doi: 10.1016/j.plantsci.2020.110563. [DOI] [PubMed] [Google Scholar]
- 34.Yu YT, et al. Overexpression of the MYB37 transcription factor enhances abscisic acid sensitivity, and improves both drought tolerance and seed productivity in Arabidopsis thaliana. Plant Mol. Biol. 2016;90:267–279. doi: 10.1007/s11103-015-0411-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Song S, et al. The Jasmonate-ZIM domain proteins interact with the R2R3-MYB transcription factors MYB21 and MYB24 to affect Jasmonate-regulated stamen development in Arabidopsis. Plant Cell. 2011;23:1000–1013. doi: 10.1105/tpc.111.083089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhang S, et al. Cloning and characterization of a putative R2R3 MYB transcriptional repressor of the rosmarinic acid biosynthetic pathway from Salvia miltiorrhiza. PLoS ONE. 2013;8:e73259. doi: 10.1371/journal.pone. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Niu Y, et al. Analysis of drought and salt-alkali tolerance in tobacco by overexpressing WRKY39 gene from Populus trichocarpa. Plant Signal Behav. 2021;16:1918885. doi: 10.1080/15592324.2021.1918885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wang Y, et al. WRKY55 transcription factor positively regulates leaf senescence and the defense response by modulating the transcription of genes implicated in the biosynthesis of reactive oxygen species and salicylic acid in Arabidopsis. Development. 2020;147:189647. doi: 10.1242/dev.189647. [DOI] [PubMed] [Google Scholar]
- 39.Lei R, et al. Arabidopsis WRKY2 and WRKY34 transcription factors interact with VQ20 protein to modulate pollen development and function. Plant J. 2017;91:962–976. doi: 10.1111/tpj.13619. [DOI] [PubMed] [Google Scholar]
- 40.Xu Y, Zhu Z. PIF4 and PIF4-interacting proteins: At the nexus of plant light, temperature and hormone signal integrations. Int. J. Mol. Sci. 2021;22:10304. doi: 10.3390/ijms221910304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Liu Z, et al. Phytochrome-interacting factors PIF4 and PIF5 negatively regulate anthocyanin biosynthesis under red light in Arabidopsis seedlings. Plant Sci. 2015;238:64–72. doi: 10.1016/j.plantsci.2015.06.001. [DOI] [PubMed] [Google Scholar]
- 42.Yang N, et al. Overexpression of SmMYC2 increases the production of phenolic acids in Salvia miltiorrhiza. Front. Plant Sci. 2017;8:1804. doi: 10.3389/fpls.2017.01804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Brown DE, et al. Flavonoids act as negative regulators of auxin transport in vivo in arabidopsis. Plant Physiol. 2001;126:524–535. doi: 10.1104/pp.126.2.524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lewis D, Wu G, Ljung K, Spalding E. Auxin transport into cotyledons and cotyledon growth depend similarly on the ABCB19 multidrug resistance-like transporter. Plant J. 2009;60:91–101. doi: 10.1111/j.1365-313X.2009.03941.x. [DOI] [PubMed] [Google Scholar]
- 45.Chen Y, et al. SOAPnuke: A MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience. 2018;7:1–6. doi: 10.1093/gigascience/gix120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Pichler M, et al. A 16S rRNA gene sequencing and analysis protocol for the Illumina MiniSeq platform. Microbiol. Open. 2018;7:e00611. doi: 10.1002/mbo3.611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Love M, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chen X, et al. Comparison of metabolome and transcriptome of flavonoid biosynthesis pathway in a purple-leaf tea germplasm Jinmingzao and a green-leaf tea germplasm Huangdan reveals their relationship with genetic mechanisms of color formation. Int. J. Mol. Sci. 2020;21:4167. doi: 10.3390/ijms21114167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Züllig T, et al. A metabolomics workflow for analyzing complex biological samples using a combined method of untargeted and target-list based approaches. Metabolites. 2020;10:342. doi: 10.3390/metabo10090342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wen B, Mei Z, Zeng C, Liu S. metaX: A flexible and comprehensive software for processing metabolomics data. BMC Bioinform. 2017;18:183. doi: 10.1186/s12859-017-1579-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Saccenti E, Timmerman ME. Approaches to sample size determination for multivariate data: Applications to PCA and PLS-DA of omics data. J. Proteome Res. 2016;15:2379–2393. doi: 10.1021/acs.jproteome.5b01029. [DOI] [PubMed] [Google Scholar]
- 52.Ye LX, et al. Comparative analysis of the transcriptome, methylome, and metabolome during pollen abortion of a seedless citrus mutant. Plant Mol. Biol. 2020;104:151–171. doi: 10.1007/s11103-020-01034-7. [DOI] [PubMed] [Google Scholar]
- 53.Thimm O, et al. MAPMAN: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004;37:914–939. doi: 10.1111/j.1365-313x.2004.02016.x. [DOI] [PubMed] [Google Scholar]
- 54.He Y, et al. Metabolomic and confocal laser scanning microscopy (CLSM) analyses reveal the important function of flavonoids in Amygdalus pedunculata pall leaves with temporal changes. Front Plant Sci. 2021;12:648277. doi: 10.3389/fpls.2021.648277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Córdoba A, et al. Flavonoid-modified surfaces: Multifunctional bioactive biomaterials with osteopromotive, anti-inflammatory, and anti-fibrotic potential. Adv. Healthc. Mater. 2015;4:540–549. doi: 10.1002/adhm.201400587. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The RNA-seq data sets for the transcriptome analysis were available at the Genome Sequence Archive in NCBI, under Biological Project ID: PRJNA796114. that are publicly accessible at https://www.ncbi.nlm.nih.gov/bioproject/PRJNA796114. The raw values obtained from the metabolomic data sets were available in Supplemental Data. If any data sets are unavailable through the links stated above, they can be obtained from the corresponding author on request.