Abstract
Ancestral sequence reconstruction (ASR) infers predicted ancestral states for sites within sequences and can constrain the functions and properties of ancestors of extant protein families. Here, we compare the likely sequences of inferred nitrogenase ancestors to extant nitrogenase sequence diversity. We show that the most-likely combinations of ancestral states for key substrate channel residues are not represented in extant sequence space, and rarely found within a more broadly defined physiochemical space—supporting that the earliest ancestors of extant nitrogenases likely had alternative substrate channel composition. These differences may indicate differing environmental selection pressures acting on nitrogenase substrate specificity in ancient environments. These results highlight ASR's potential as an in silico tool for developing hypotheses about ancestral enzyme functions, as well as improving hypothesis testing through more targeted in vitro and in vivo experiments.
Keywords: ancestral sequence reconstruction, nitrogenase, Nif, early life
Introduction
Ancestral sequence reconstruction (ASR) is widely used for modeling sequence properties and functions of ancient enzyme variants. Maximum-likelihood reconstructions combine substitution models with aligned sequence data and phylogenetic information to calculate the residue likelihood of aligned sites for all internal nodes in a tree (Merkl and Sterner 2016; Selberg Gaucher and Liberles 2021). This approach enables exploration of targeted hypotheses about ancestral proteins in silico and is also an essential starting point for enzyme resurrection and in vitro or in vivo activity assays (Garcia and Kaçar 2019; Liberles et al. 2020). Because most of Earth's earliest organisms and metabolisms have no preserved record, deep-time evolution must be inferred using modern genomic and biochemical data (Barnabas et al.1982). ASR models protein “fossils,” providing biological data about ancestral enzymes and organisms to consider in context with environmental data, such as geochemical or atmospheric modeling (Thornton 2004; Hochberg and Thornton 2017). This process enables quantitative assessment of otherwise-elusive evolutionary hypotheses, making ASR essential for studying ancient, geochemically relevant enzyme families (Garcia and Kaçar 2019).
Nitrogen-fixing nitrogenases are the lynchpin of Earth's biological nitrogen cycle, and have been extensively studied for their role in global biogeochemistry (Raymond et al. 2004; Canfield Glazer and Falkowski 2010; Sickerman et al. 2019). Modern nitrogenases are categorized by their metal cofactors (Sickerman et al. 2019; Raymond et al. 2004; Garcia et al. 2020; North et al. 2020). All diazotrophs express molybdenum nitrogenase (Nif), with its iron-molybdenum cofactor (FeMoCo); some nitrogen fixers also contain alternative vanadium nitrogenase (Vnf) or iron-only nitrogenase (Anf; Boyd et al. 2011; Sickerman et al. 2019; Garcia et al. 2020). Phylogenetic analyses, molecular clock data, and cofactor-type-specific nitrogen isotope fractionations support the parsimonious inference that Nif is the oldest, ancestral form of nitrogenase. Recent studies suggest that the ancestor of this enzyme may date to the early Archaean eon (Boyd et al. 2011; Stüeken et al. 2015; Garcia et al. 2020; Parsons et al. 2021).
In addition to their unique capacity for dinitrogen (N2) fixation, modern nitrogenases can promiscuously reduce other linear, triple-bond-containing molecules at the FeMoCo active site (Fani et al. 2000; Seefeldt et al. 2013). It is possible that these off-target substrates could have played a larger role in early Earth systems. Several alternative substrates for nitrogenase enzymes have been identified, including carbon dioxide (CO2), carbon monoxide (CO), acetylene (C2H2), and hydrogen cyanide (HCN; Li et al. 1982; Conradson et al. 1989; Seefeldt et al. 2013; Stripp et al. 2022). It has been proposed that HCN promiscuity reflects “relic” chemistry of nitrogenases from highly reduced early environments, with HCN serving as a substrate or a target for detoxification (Silver and Postgate 1973; Fani et al. 2000; Pickett et al. 2004). HCN was produced and maintained on early Earth (Tian et al. 2011; Ferus et al. 2017; Todd and Öberg 2020), especially during the Archaean eon (Zahnle 1986; Tian et al. 2011), and is a prominent candidate feedstock molecule for abiogenesis, as it can be abiotically converted into several key nucleic and amino acids (Oró 1961; Oró and Kamat 1961; Ferris and Orgel 1966; Ferris et al. 1978; Miyakawa et al. 2002; LaRowe and Regnier 2008; Pearce and Pudritz 2015; Sutherland 2016). Given the age of the nitrogenase ancestor, no direct biological or biochemical data exist to support the presence or significance of HCN in early ecosystems. But ASR may be able to constrain such hypotheses for ancient biochemistry (Garcia and Kaçar 2019).
A higher affinity for alternative substrates in ancestral nitrogenases would likely require different amino acid residues in the substrate-binding part of the enzyme (the substrate channel), while preserving catalytic activity. As ASRs are inherently probabilistic, individual sites may be inferred to have several possible ancestral states with non-negligible probabilities. Variation across sampled extant diversity allows the combinations within likely ancestor sequences to explode, producing a far greater number of plausible sequence ancestors than can be experimentally tested. Here, we use ASR to reconstruct the amino acid identities of alignment sites directly interacting with the substrate binding channel in the common ancestor of the active site-containing nitrogenase subunit (nif/vnf/anfD). A library of plausible sequence ancestors for this collection of sites was generated, and their relative probabilities calculated. We applied a simple hypothesis-testing strategy: Are likely ancestral active site sequence states found within sampled extant nitrogenase diversity? We find that this is not always the case, permitting the possibility that early nitrogenase ancestors had substrate binding channels with properties distinct from modern enzymes.
Results
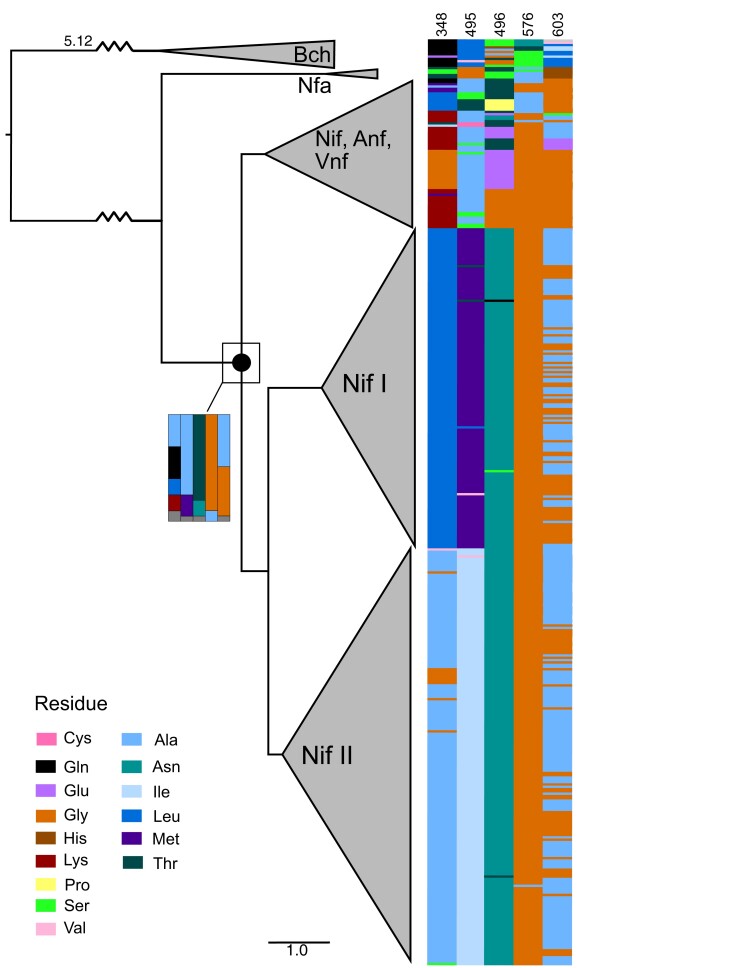
Of 24 sites known to play a role in substrate docking or coordination in modern Azotobacter vinelandii Nif (Seefeldt et al. 2013; Smith et al. 2014), five residues showed variability (>1 ancestral state with P > 0.10) in the predicted nitrogenase ancestor (table 1). For comparison with ASR results, sequence diversity for these five variable sites was evaluated in extant nitrogenases (fig. 1). Extant variation at these sites roughly maps onto the major nitrogenase groups in the gene tree. Sites 398 and 495 in the alignment highlight the distinction between Group I Nif (comprising well-studied proteobacterial and cyanobacterial nifD sequences, including that of A. vinelandii) and Group II Nif (comprising sequences from Clostridia, Bacteroidetes, and methanogenic archaea, among others; Raymond et al. 2004). Group I Nif conserves leucine (Leu) at site 395, whereas Group II conserves primarily alanine (Ala) (with fewer cases of glycine [Gly], and individual substitutions of serine [Ser] and cysteine [Cys]). Group I Nif contains primarily methionine (Met) at site 495, whereas Group II Nif primarily displays isoleucine (Ile). Variation at site 496, meanwhile, highlights the split between Nif I/II nitrogenases and the clade including alternative (Vnf, Anf) nitrogenases (which also contains several divergent Nif sequences): With very few exceptions, Group I and II nitrogenases conserve asparagine (Asn) at this site, whereas alternative clade sequences are poorly constrained, most frequently expressing glutamate (Glu), threonine (Thr), and Gly.
Table 1.
ASR-Predicted Residue for key Substrate Channel Sites
| Conserved residue in alignment (Av) | ASR Nif/Anf/Vnf ancestor residue, p(residue) | Other residues P > 0.10 | Putative role in modern Nif active site | Corresponding residue in Av nifD (Seefeldt et al. 2013; Smith et al. 2014) |
|---|---|---|---|---|
| Val-182 | Val, 0.99998 | No | Modulates substrate access to catalytic core/active site | α-70Val |
| His-350 | His, 0.99999 | No | H-bond with FeMoCo, flexible (S2A or S2B) | α-195His |
| Tyr-497 | Tyr, 0.99997 | No | Substrate channel formation; gating residue | α-281Tyr |
| Arg-493 | Arg, 0.99991 | No | H-bond with FeMoCo in substrate channel (surface “flap” for substrate access) | α-277Arg |
| His-604 | His, 1 | No | Substrate channel formation; gating residue | α-383His |
| Cys-491 | Cys, 1 | No | Coordinates FeMoCo | α-275Cys |
| His-745 | His, 0.99998 | No | Coordinates FeMoCo | α-442His |
| Arg-224 | Arg, 0.99912 | No | Coordinates FeMoCo | α-96Arg |
| Gln-346 | Gln, 0.99995 | No | Coordinates FeMoCo | α-191Gln |
| Gly-181 | Gly, 0.99934 | No | Switch control for α-70Val side chain | α-69Gly |
| Ser-347 | Ser, 0.99888 | No | N2 interaction in substrate channel | α-192Ser |
| Asn-155 | Asn, 0.99972 | No | Lines substrate channel | α-49Asn |
| Gly-178 | Gly, 0.99994 | No | Lines substrate channel | α-66Gly |
| Val-183 | Val, 0.99824 | No | Lines substrate channel | α-71Val |
| Ser-345 | Ser, 0.99918 | No | Lines substrate channel | α-190Ser |
| Leu-348 | Ala, 0.29253 | Gln, 0.27519; Leu, 0.16779; Lys, 0.16716 | Lines substrate channel | α-193Leu |
| His-351 | His, 0.99992 | No | Lines substrate channel | α-196His |
| Asn-354 | Asn, 0.99946 | No | Lines substrate channel | α-199Asn |
| Ser-494 | Ser, 0.99993 | No | Lines substrate channel | α-278Ser |
| Met-495 | Ala, 0.74242 | Met, 0.17555 | Lines substrate channel | α-279Met |
| Asn-496 | Thr, 0.79331 | Asn, 0.14693 | Lines substrate channel | α-280Asn |
| Gly-576 | Gly, 0.87529 | Ala, 0.12413 | Lines substrate channel | α-357Gly |
| Phe-602 | Phe, 1 | No | Lines substrate channel | α-381Phe |
| Ala-603 | Ala, 0.47988 | Gly, 0.46717 | Lines substrate channel | α-382Ala |
Fig. 1.
Extant nitrogenase sequence diversity. A maximum-likelihood gene tree for the D subunit of extant nitrogenases and outgroup sequences (Nfa, Group IV nitrogenase-like enzymes; Bch, bacteriochlorophyll/protochlorophyllide oxidoreductases), displaying the residue state at each of five variable sites in the Azotobacter vinelandii nifD substrate channel. The last common ancestor evaluated for nitrogenases is marked with a black node; reconstructed site likelihoods for all residues P > 0.10 are shown by residue color in the inset bar chart.
Predicted combinations of residue states were evaluated in the ASR for the ancestor of Nif, Anf, and Vnf (fig. 1). The 64 evaluated ancestral sequence combinations of five variant sites had predicted joint likelihood ratios >0.00025% (fig. 2). Only 5/64 (7.8%) combinations were found in extant sequences (fig. 2). Of these five combinations, only one—the second-highest likelihood sequence, AATGG—was in the top 20 most-likely predicted ancestral combinations. Four of the five sequences were “rare,” observed in no more than two extant sequences; the only non-rare sequence (LMNGG) had the lowest likelihood of these five. Several sequence patterns inferred as high-likelihood reconstructed ancestors were not observed within extant nitrogenases. For example, in sampled extant sequences, Gln-348 does not co-occur with Gly-576; Ala-348 does not co-occur with Met-495; and Leu-348 does not co-occur with Ala-495. Ala-348 and Ala-495 only co-occur in one extant sequence.
Fig. 2.
Ancestral sequence and state likelihoods. Likelihood ratios, amino acid sequence, and residue physicochemical states are shown for all combinations of the five variable sites (alignment sites 348, 495, 496, 576, and 603) predicted in the nitrogenase substrate channel ASR. “Rare” states occur in no more than two extant sequences.
It is possible that observed differences between extant sequences and inferred ancestors involve conservative amino acid substitutions that are unlikely to substantially affect substrate binding or phenotype. To test this, amino acids were recoded by general physiochemical properties, to identify radical changes of amino acid type. This showed that 29/64 (45%) of reconstructed ancestors contained substitutions yielding a combination not represented by extant sequences or physicochemical types (fig. 2). Of the 35 ancestors represented among extant physicochemical types, 28/35 (80%) were rare. The 32 most-likely ancestors all displayed physicochemical types that were rare or nonexistent within extant sequences. The top ten most-likely ancestors sampled rare physicochemical types only found within the divergent Nif/Anf/Vnf clade, not within Nif I/II.
The effects of substitution models and topology on predicted ancestor sequence and type were also evaluated by repeating tree construction and ASR under WAG + R10 (the highest likelihood non-LG model) and BLOSUM62 + R10 (a lower likelihood, simple model). Substitution models slightly impacted recovered tree topologies within recently diverged lineages but did not alter phylogenetic relationships between major nitrogenase types and their outgroups. Relative likelihoods for specific amino acids are generally conserved at sites 495, 496, and 603, and sample the same plurality at site 576 (Gly, Ala) and site 348 (Ala, Glu, Gln, Lys) (fig. 3). Moreover, residue type combinations are robust to model selection.
Fig. 3.
Substitution model effect on nitrogenase ASR. Proportional likelihoods are shown for residue states and physicochemical types at the five variable sites (alignment sites 348, 495, 496, 576, and 603) predicted in the nitrogenase ancestor for three different substitution models: LG + R9; WAG + R10; and BLOSUM62 + R10. Relative amino acid likelihoods for the nitrogenase ancestor were generally conserved across models at sites 495, 496, and 603. At site 576, LG and BLOSUM favor Gly over Ala (88% vs. 12% for LG and 81% vs. 19% for BLOSUM, respectively), whereas WAG favors Ala over Gly (88% vs. 12%, respectively). At site 348, all models recover a plurality of Ala, Glu, Gln, and Lys, with relative probabilities varying between 17.5% and 1.9%. Residue type combinations are robust to model selection across sites.
Discussion
These results suggest that the inferred ancestor of Nif, Vnf, and Anf nitrogenases most likely contained a substrate channel sequence distinct from that found in extant sequence space. Twenty-two of the 23 highest likelihood sequence combinations of the five variant substrate channel ancestral sites are not observed within known modern nitrogenases; the only represented sequence pattern occurs in only one observed extant enzyme, that of Chloroflexales bacterium ZM16-3. The top ten most-likely ASR residue combination physicochemical types were represented by very few extant sequences, and only within the divergent group of Nif that also contains the alternative nitrogenases. Divergent homologs within this clade may sample ancestral sequence patterns to a greater extent than type I or II nitrogenases. If so, characterizing these extant genes could provide useful analogs for ancestral phenotypes.
The conservative substitutions observed in the highest likelihood predicted ancestors are unlikely to radically influence substrate affinity or binding, at least when compared with the extant enzymes in the Nif/Anf/Vnf group. In the case of HCN, for example, because both modern Vnf (Seefeldt et al. 2013) and Nif can reduce this alternative substrate, there is no reason to believe that the ASR-predicted ancestors could not as well. Nevertheless, the observed departures from extant sequence space in many likely ancestor sequences open the possibility of greater phenotypic variation of substrate binding in nitrogenase ancestors. Recent work has indicated that even a small number of substitutions, even outside the active site, can elicit different ancestral substrate affinities (Tatsaki et al. 2021). This justifies experimental investigation of alternative substrate binding in reconstructed ancestor candidates, especially those displaying the novel residue type combinations sampled above.
Extant nitrogenases that remain unsampled and excluded from the tree could contain currently missing high-likelihood sequence combinations in the ASR. Additionally, likelihood ratios calculated from only a handful of sites do not represent overall sequence likelihoods, as site likelihoods are calculated independently. Thus, the most-likely ancestral site combinations in the substrate channel are unlikely to identify the sequence in the most-likely overall ancestor. However, the likelihood calculations for these substrate binding sites may be more informative for inferring ancestral phenotype changes than the overall ancestral sequence of the enzyme. Indeed, physicochemical type combinations predicted by the ASR are more robust to changes in substitution model and tree topology than sequence-level variation (fig. 3); this underscores the importance of considering physicochemical type for predicted substitutions. These results could be further contextualized by evaluating more ancient ancestors—for example, the last common ancestor of nitrogenases and maturases (Garcia et al. 2022)—and less-likely ancestral candidates with methods such as Bayesian sampling and laboratory reconstruction (Merkl and Sterner 2016; Garcia and Kaçar 2019; Selberg et al. 2021). But this likelihood-based, nitrogenase-exclusive approach provides the most conservative evaluation of functional hypotheses based on modern enzyme variants.
The results highlight the value of ASR as a first step in evaluating enzyme evolution hypotheses. In some cases, ASR itself may prove sufficient to refute certain evolutionary hypotheses; for example predicted ancestral states may be sterically incompatible with hypothesized substrate interactions or protein folding. Such results would preclude additional experimental work. It is also possible that when ASR fails to identify combinations of key ancestral residues absent from extant diversity, the null hypothesis of uniform phenotype across the reconstructed history should be preferred. However, even in this case, ASR will efficiently identify key sites to target for mutagenesis for in vitro or in vivo analyses—such as the novel sequence and type combinations sampled in this work.
Materials and methods
An initial data set of extant nitrogenase D-subunit homologs was constructed from hits gathered using BLASTp to search NCBI's non-redundant protein database with a query sequence from A. vinelandii (WP_012698832.1). Hits were included for molybdenum (NifD), vanadium (VnfD), and iron-only (AnfD) nitrogenases and manually curated to remove partial sequences, prune oversampled clades, and ensure the presence of conserved sequence features known to be critical for N2 reduction (e.g., Cys-275, His-442). The curated initial data set included 385 nitrogenase sequences and 385 outgroup sequences from light-independent protochlorophyllide oxidoreductases (Boyd et al. 2011; Ghebreamlak and Mansoorabadi 2020). These initial homologs were aligned using MAFFT under auto-parameterization (selecting L-INS-i; Nakamura et al. 2018). Outgroup sequences were further subsampled to retain one sequence per phylum for major monophyletic groups (12 sequences). A second nested outgroup (five sequences) of Group IV-A nitrogenase-like homologs (nfaD; Raymond et al. 2004; North et al. 2020) was profile-aligned to the existing alignment with MAFFT, providing an additional ancestral node required to reconstruct the nif/anf/vnf ancestor as an internal node in the tree. The tree was rooted along the split between nitrogenases and the other oxidoreductases. Maximum-likelihood tree construction and subsequent ASR were performed in IQ-Tree (Nguyen et al. 2015), under the best-fit model identified by the Bayesian Information Criterion, LG + R9. Tree construction and ASR were also repeated using a similar, lower likelihood model, WAG + R10; and a simpler, lower likelihood model, BLOSUM62 + R10. Joint likelihoods were calculated as the product of individual residue likelihoods reported for sites of interest in the ASR state file. Amino acid states were grouped with the following physicochemical categories for residues: Acidic (Glu, Asp); Aliphatic (Ile, Leu, Met, Val); Amine (Gln, Asn); Aromatic (Trp, Tyr, Phe); Basic (Lys, Arg); Small Hydroxyl (Ser, Thr); Tiny (Ala, Gly).
Acknowledgments
This work was supported by the Simons Foundation Collaboration on the Origins of Life (grant #339603 to G.P.F.); a National Defense Science and Engineering Graduate Fellowship (S.L.S.); the National Aeronautics and Space Administration (Postdoctoral Program Fellowship to A.K.G.); the NASA Interdisciplinary Consortium for Astrobiology Research: Metal Utilization and Selection Across Eons, MUSE (19- ICAR19-2-0007; B.K.) a member of the LIFE from Early Cells to Multicellularity Research Coordination Network sponsored by the NASA Science Mission Directorate. The authors thank the Jackson Lab at University of Wisconsin-Madison and Martin Krzywinski of the Michael Smith Genome Sciences Centre for information on the 15-level colorblind-friendly palette data utilized in these figures.
Contributor Information
Sarah L Schwartz, Microbiology Graduate Program, Massachusetts Institute of Technology, Cambridge, MA; Department of Earth, Atmospheric, and Planetary Sciences, Massachusetts Institute of Technology, Cambridge, MA.
Amanda K Garcia, Department of Bacteriology, University of Wisconsin-Madison, Madison, WI.
Betül Kaçar, Department of Bacteriology, University of Wisconsin-Madison, Madison, WI.
Gregory P Fournier, Department of Earth, Atmospheric, and Planetary Sciences, Massachusetts Institute of Technology, Cambridge, MA.
Data Availability
Data files used for analyses in this work are freely available at https://doi.org/10.6084/m9.figshare.c.6039527.
References
- Barnabas J, Schwartz RM, Dayhoff MO. 1982. Evolution of major metabolic innovations in the Precambrian. Orig Life. 12(1):81–91. [DOI] [PubMed] [Google Scholar]
- Boyd ES, Anbar AD, Miller S, Hamilton TL, Lavin M, Peters JW. 2011. A late methanogen origin for molybdenum-dependent nitrogenase. Geobiology 9(3):221–232. [DOI] [PubMed] [Google Scholar]
- Canfield DE, Glazer AN, Falkowski PG. 2010. The evolution and future of Earth's nitrogen cycle. Science 330(6001):192–196. [DOI] [PubMed] [Google Scholar]
- Conradson SD, Burgess BK, Vaughn SA, Roe AL, Hedman B, Hodgson KO, Holm RH. 1989. Cyanide and methylisocyanide binding to the isolated iron-molybdenum cofactor of nitrogenase. J Biol Chem. 264(27):15967–15974. [PubMed] [Google Scholar]
- Fani R, Gallo R, Liò P. 2000. Molecular evolution of nitrogen fixation: the evolutionary history of the nifD, nifK, nifE, and nifN genes. J Mol Evol. 51(1):1–11. [DOI] [PubMed] [Google Scholar]
- Ferris JP, Joshi PC, Edelson EH, Lawless JG. 1978. HCN: a plausible source of purines, pyrimidines and amino acids on the primitive earth. J Mol Evol. 11(4):293–311. [DOI] [PubMed] [Google Scholar]
- Ferris JP, Orgel LE. 1966. An unusual photochemical rearrangement in the synthesis of adenine from hydrogen cyanide1. J Am Chem Soc. 88(5):1074. [Google Scholar]
- Ferus M, Kubelík P, Knížek A, Pastorek A, Sutherland J, Civiš S. 2017. High energy radical chemistry formation of HCN-rich atmospheres on early earth. Sci Rep. 7(1):6275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcia AK, McShea H, Kolaczkowski B, Kaçar B. 2020. Reconstructing the evolutionary history of nitrogenases: evidence for ancestral molybdenum-cofactor utilization. Geobiology 18(3):394–411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcia AK, Kaçar B. 2019. How to resurrect ancestral proteins as proxies for ancient biogeochemistry’, Free Radical Biol Med. 140:260–269. [DOI] [PubMed] [Google Scholar]
- Garcia AK, Kolaczkowski B, Kaçar B. 2022. Reconstruction of nitrogenase predecessors suggests origin from maturase-like proteins. Genome Biol Evol. 14:3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghebreamlak SM, Mansoorabadi SO. 2020. Divergent members of the nitrogenase superfamily: tetrapyrrole biosynthesis and beyond. ChemBioChem. 21(12):1723–1728. [DOI] [PubMed] [Google Scholar]
- Hochberg GKA, Thornton JW. 2017. Reconstructing ancient proteins to understand the causes of structure and function. Ann Rev Biophy. 46(1):247–269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LaRowe DE, Regnier P. 2008. Thermodynamic potential for the abiotic synthesis of adenine, cytosine, guanine, thymine, uracil, ribose, and deoxyribose in hydrothermal systems. Orig Life Evol Biosphy. 38(5):383. [DOI] [PubMed] [Google Scholar]
- Li J, Burgess BK, Corbin JL. 1982. Nitrogenase reactivity: cyanide as substrate and inhibitor. Biochemistry. 21(18):4393–4402. [DOI] [PubMed] [Google Scholar]
- Liberles DA, Chang B, Geiler-Samerotte K, Goldman A, Hey J, Kaçar B, Meyer M, Murphy W, Posada D, Storfer A. 2020. Emerging frontiers in the study of molecular evolution. J Mol Evol. 88:211–226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merkl R, Sterner R. 2016. Ancestral protein reconstruction: techniques and applications. Biol Chem. 397(1):1–21. [DOI] [PubMed] [Google Scholar]
- Miyakawa S, Cleaves H, Miller SL. 2002. The cold origin of life: hydrogen cyanide and formamide. Orig Life Evol Biosph. 32:195–208. [DOI] [PubMed] [Google Scholar]
- Nakamura T, Yamada KD, Tomii K, Katoh K. 2018. Parallelization of MAFFT for large-scale multiple sequence alignments. Bioinformatics 34(14):2490–2492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ. 2015. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol. 32(1):268–274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- North JA, Narrowe AB, Xiong W, Byerly KM, Zhao G, Young SJ, Murali S, Wildenthal JA, Cannon WR, Wrighton KC, et al. 2020. A nitrogenase-like enzyme system catalyzes methionine, ethylene, and methane biogenesis. Science 369(6507):1094–1098. [DOI] [PubMed] [Google Scholar]
- Oró J. 1961. Mechanism of synthesis of adenine from HCN under possible primitive earth conditions. Nature 191(4794):1193–1194. [DOI] [PubMed] [Google Scholar]
- Oró J, Kamat SS. 1961. Amino-acid synthesis from hydrogen cyanide under possible primitive earth conditions. Nature 190:442–443. [DOI] [PubMed] [Google Scholar]
- Parsons C, Stüeken EE, Rosen CJ, Mateos K, Anderson RE. 2021. Radiation of nitrogen-metabolizing enzymes across the tree of life tracks environmental transitions in earth history. Geobiology 19(1):18–34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pearce BKD, Pudritz RE. 2015. Seeding the pregenetic earth: meteoritic abundances of nucleobases and potential reaction pathways. Astrophy J. 807(1):85. [Google Scholar]
- Pickett CJ, Vincent KA, Ibrahim SK, Gormal CA, Smith BE, Fairhurst SA, Best SP. 2004. Synergic binding of carbon monoxide and cyanide to the FeMo cofactor of nitrogenase: relic chemistry of an ancient enzyme? Chemistry 10(19):4770–4776. [DOI] [PubMed] [Google Scholar]
- Raymond J, Siefert JL, Staples CR, Blankenship RE. 2004. The natural history of nitrogen fixation. Mol Biol Evol. 21(3):541–554. [DOI] [PubMed] [Google Scholar]
- Seefeldt LC, Yang Z, Duval S, Dean DR. 2013. Nitrogenase reduction of carbon-containing compounds. Biochim Biophys Acta. 1827(8):1102–1111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Selberg AGA, Gaucher EA, Liberles DA. 2021. Ancestral sequence reconstruction: from chemical paleogenetics to maximum likelihood algorithms and beyond. J Mol Evol. 89(3):157–164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sickerman NS, Hu Y, Ribbe MW. 2019. Nitrogenases. Methods Mol Biol. 1876:3–24. [DOI] [PubMed] [Google Scholar]
- Silver WS, Postgate JR. 1973. Evolution of asymbiotic nitrogen fixation. J Theoretical Biol. 40(1):1–10. [DOI] [PubMed] [Google Scholar]
- Smith D, Danyal K, Raugei S, Seefeldt LC. 2014. Substrate channel in nitrogenase revealed by a molecular dynamics approach. Biochemistry 53(14):2278–2285. [DOI] [PubMed] [Google Scholar]
- Stripp ST, Duffus BR, Fourmond V, Léger C, Leimkühler S, Hirota S, Hu Y, Jasniewski A, Ogata H, Ribbe MW. 2022. Second and outer coordination sphere effects in nitrogenase, hydrogenase, formate dehydrogenase, and CO dehydrogenase. Chem Rev. 122(14):11900–11973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stüeken EE, Buick R, Guy BM, Koehler MC. 2015. Isotopic evidence for biological nitrogen fixation by molybdenum-nitrogenase from 3.2 gyr. Nature 520(7549):666–669. [DOI] [PubMed] [Google Scholar]
- Sutherland JD. 2016. The origin of life - out of the blue. Angewandte Chemie. 55(1):104–121. [DOI] [PubMed] [Google Scholar]
- Tatsaki E, Anagnostopoulou E, Zantza I, Lazou P, Mikros E, Frillingos S. 2021. Identification of new specificity determinants in bacterial purine nucleobase transporters based on an ancestral sequence reconstruction approach. J Mol Biol. 433(24):167329. [DOI] [PubMed] [Google Scholar]
- Thornton JW. 2004. Resurrecting ancient genes: experimental analysis of extinct molecules. Nat Rev Genet. 5(5):366–375. [DOI] [PubMed] [Google Scholar]
- Tian F, Kasting JF, Zahnle K. 2011. Revisiting HCN formation in Earth's early atmosphere. Earth Planet Sci Lett. 308(3):417–423. [Google Scholar]
- Todd ZR, Öberg KI. 2020. Cometary delivery of hydrogen cyanide to the early earth. Astrobiology 20(9):1109–1120. [DOI] [PubMed] [Google Scholar]
- Zahnle KJ. 1986. Photochemistry of methane and the formation of hydrocyanic acid (HCN) in the Earth's early atmosphere. J Geophys Res. 91(D2):2819–2834. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Data files used for analyses in this work are freely available at https://doi.org/10.6084/m9.figshare.c.6039527.