Abstract
Background
Modern laboratory methods such as next generation sequencing and MALDI-TOF allow identification of novel bacterial species. This can affect surveillance of infections and antimicrobial resistance. From 2017, increasing numbers of medical microbiology laboratories in Switzerland differentiated Klebsiella variicola from Klebsiella pneumoniae complex using updated MALDI-TOF databases, whereas many laboratories still report them as K. pneumoniae or K. pneumoniae complex.
Aim
Our study explored whether separate reporting of K. variicola and the Klebsiella pneumoniae complex affected the ANRESIS surveillance database.
Methods
We analysed antibiotic susceptibility rates and specimen types of K. variicola and non-K. variicola-K. pneumoniae complex isolates reported by Swiss medical laboratories to the ANRESIS database (Swiss Centre for Antibiotic Resistance) from January 2017 to June 2022.
Results
Analysis of Swiss antimicrobial resistance data revealed increased susceptibility rates of K. variicola compared with species of the K. pneumoniae complex other than K. variicola in all six antibiotic classes tested. This can lead to underestimated resistance rates of K. pneumoniae complex in laboratories that do not specifically identify K. variicola. Furthermore, K. variicola strains were significantly more often reported from blood and primarily sterile specimens than isolates of the K. pneumoniae complex other than K. variicola, indicating increased invasiveness of K. variicola.
Conclusion
Our data suggest that refined differentiation of the K. pneumoniae complex can improve our understanding of its taxonomy, susceptibility, epidemiology and clinical significance, thus providing more precise information to clinicians and epidemiologists.
Keywords: Klebsiella pneumoniae complex, Klebsiella variicola, ANRESIS, antimicrobial resistance, surveillance
Key public health message.
What did you want to address in this study?
Our previously known microorganism species classification is increasingly refined by novel technological methods. In this study, we investigated how this technological leap affects antibiotic resistance surveillance data using Klebsiella pneumoniae as an example.
What have we learnt from this study?
The introduction of further differentiation of the Klebsiella pneumoniae species complex into individual species was inconsistent across laboratories. This affects antibiotic resistance monitoring: resistance rates of the species complex as a whole appear lower in those laboratories where Klebsiella variicola, one of the less resistant species in the complex, is still included.
What are the implications of your findings for public health?
Harmonisation of resistance monitoring requires early identification of such inhomogeneous data and agreement on a common approach, otherwise resistance data may differ across countries (and even within countries) for methodological reasons.
Introduction
DNA sequencing methods refine phylogenetic population structures and allow identification of novel species. Novel species may show distinct resistance rates and clinical significance, revising our understanding of epidemiology, ecology, pathogenesis and antimicrobial susceptibility [1,2]. Examples include Staphylococcus argenteus, Gemella taiwanensis, Actinotignum spp [1], Macrococcus spp [2], which have been identified as novel sources of infections. The taxonomies of Corynebacteriae and Enterobacterales have been reclassified considerably, with new species and subspecies [2]. Furthermore, colistin-resistant Citrobacter europaeus strains and multi-resistant Enterobacter bugandensis isolates with the potential of increased virulence have been described [2]. In addition to molecular tools, matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS) can reliably identify a growing number of species in routine diagnostics, which allows treatment adjustment and increased understanding of clinical consequences in practical healthcare [3]. A recent study has shown the potential of MALDI-TOF to identify subgroups within a bacterial species for epidemiological assessments [4].
The Klebsiella pneumoniae complex is highly diverse: as a member of Enterobacterales, this complex of closely related species occupies various ecological niches. Several virulence and resistance factors have been described in strains causing a variety of infections in humans and animals [5]. To date, phylogenetic research on the population structure of the K. pneumoniae complex has identified four species with a total of seven subspecies [6]. Recently, an additional species has been proposed [7]: ‘K. quasivariicola’.
Klebsiella pneumoniae sensu stricto has been isolated from asymptomatic humans, but is also a frequent cause of serious invasive infections associated with strains carrying large varieties of virulence or resistance accessory genes [5]. Other Klebsiella species are considered less virulent and resistant, although recent studies applying refined species discrimination methods revealed virulence and antibiotic resistance genes also in these species [7]. For example, K. variicola has been recovered from a wide range of environments and hosts [8]. It is an opportunistic human pathogen causing e.g. bloodstream, respiratory and urinary tract infections [9]. Moreover, studies report genetic potential of virulence traits [10], increased clinical virulence [11] and antibiotic resistance [12].
Increasing antimicrobial resistance rates are a concern for public health [13]. Many countries, including Switzerland, address this issue by antibiotic stewardship guidelines and surveillance databases to control resistance rates. However, data quality and comparability are very important to provide meaningful surveys on a national and international level, while appropriately integrating novel findings such as the identification of novel species, emerging resistance genes or virulence factors. Since 2017, commercial MALDI-TOF databases have enabled differentiation of K. variicola from other species of the K. pneumoniae complex. An increasing number of laboratories has started reporting K. variicola separately and report the remaining species of the K. pneumoniae complex as K. pneumoniae or K. pneumoniae complex. Thus, K. pneumoniae or K. pneumoniae complex can include different sets of species depending on the reporting policies of individual laboratories. For antimicrobial resistance surveillance databases, inconsistent reporting practices may affect trend analyses of resistance rates if novel species show differential antimicrobial susceptibilities.
The Swiss Centre for Antibiotic Resistance (ANRESIS) has hosted the Swiss national antimicrobial resistance surveillance database since 2004 and collects antibiotic resistance data of all bacterial species isolated and reported from various invasive and non-invasive specimens (primary sterile specimens, swabs, blood cultures, urine, sputum, stool) of up to 37 medical diagnostic laboratories (in 2020). Reported isolates were recovered from patients treated in different facilities, ranging from tertiary hospitals to general practices. The contributing laboratories are widely distributed across Switzerland, representing 89% of all hospitalised patients [14]. With the emergence of reported K. variicola isolates, the aim of this study was to investigate whether differentiation of K. variicola from other species of the K. pneumoniae complex (we refer to those as non-K. variicola-K. pneumoniae-complex (non-kv-kpc) in this article) may affect the ANRESIS surveillance database.
Methods
We extracted susceptibility rates and specimen types of K. variicola and non-kv-kpc isolates from the ANRESIS database for all laboratories differentiating K. variicola from K. pneumoniae complex, starting from 1 January 2017 when K. variicola became available in the MALDI-TOF database for routine diagnostics, or as soon as individual laboratories began K. variicola differentiation, until 13 June 2022. Species identification and antimicrobial susceptibility tests (AST) were performed at local laboratories following the guidelines from European Committee on Antimicrobial Susceptibility Testing (EUCAST) (36 laboratories) or the Clinical and Laboratory Standards Institute (CLSI) (one laboratory until 2020). The individual laboratories used various AST methods, such as automated minimum inhibitory concentration (MIC) tests or disc diffusion tests measuring inhibition zones manually or through an automated process. All reporting laboratories are accredited by the Swiss Accreditation Service (SAS).
Resistance categories were defined resistant if categorised R (resistant) by reporting laboratories, and non-resistant if categorised S (susceptible) or I (susceptible, increased exposure), according to EUCAST's revised susceptibility categorisation that has been introduced gradually in Swiss laboratories since 2019 [15]. For third- and fourth-generation cephalosporins, we considered isolates resistant if at least one substance of this class was categorised R. Our database included the following third- and fourth-generation cephalosporins: cefepime, cefixime, cefotaxime, cefpodoxime, ceftazidime, ceftibuten and ceftriaxone. For carbapenems, we only considered meropenem and imipenem, and rated them resistant if they were categorised as R, because many laboratories did not include ertapenem in their AST panels. For all remaining antibiotic classes, we included the most commonly reported substance in the analysis: amoxicillin-clavulanic acid, ciprofloxacin, gentamicin and trimethoprim-sulfamethoxazole, resulting in a total of six antibiotic classes tested.
Invasive strains were defined as isolates obtained from blood or primarily sterile specimen types, such as cerebrospinal fluid (CSF) and tissue specimens. We compared susceptibility rates and proportions of invasive strains of K. variicola and non-kv-kpc using Fisher's exact test. Repeated testing was accounted for by applying sequential Bonferroni corrections. All statistical tests were conducted in R v4.2.1 [16].
Results
Laboratories routinely identifying K. variicola increased from four of 30 laboratories reporting antibiotic susceptibility data to ANRESIS in 2017 to 18 of 35 laboratories in June 2022. Klebsiella variicola was less frequently isolated than non-kv-kpc isolates: from 1 January 2017 to 13 June 2022, 14.1% (n = 9,899) of the 70,114 isolates analysed were reported as K. variicola. All six antibiotic substances or antibiotic classes tested showed significantly higher susceptibility rates in K. variicola than in non-kv-kpc isolates. These differences remained after applying sequential Bonferroni corrections (Table and Figure) and after considering EUCAST's previous definition of ‘non-susceptibility’ (isolates tested I (intermediate) or R (resistant)). See Supplementary Table S1 for the detailed results of Fisher’s exact tests after applying EUCAST’s previous definition of ‘non-susceptibility’.
Table. Resistance rates against six antibiotics of Klebsiella variicola and non-kv-kpc isolates collected by ANRESIS, Switzerland, 1 January 2017–13 June 2022 (n = 70,114).
| Antibiotic | Non-kv-kpc | Klebsiella variicola | OR | 95% CI | p value | ||
|---|---|---|---|---|---|---|---|
| Resistant (%) | n (total) | Resistant (%) | n (total) | ||||
| Amoxicillin-clavulanic acid | 9.1 | 60,114 | 4.7 | 9,889 | 2.0 | 1.8–2.2 | < 0.001 |
| Carbapenem | 0.4 | 45,460 | 0.1 | 7,832 | 7.7 | 3.0–28.5 | < 0.001 |
| Cephalosporin 3rd/4th generation | 6.6 | 54,530 | 2.9 | 8,883 | 2.4 | 2.1–2.7 | < 0.001 |
| Ciprofloxacin | 6.9 | 59,743 | 2.1 | 9,856 | 3.4 | 2.9–3.9 | < 0.001 |
| Gentamicin | 2.8 | 37,806 | 1.7 | 6,447 | 1.6 | 1.3–2.0 | < 0.001 |
| Trimethoprim-sulfamethoxazole | 9.5 | 58,489 | 3.9 | 9,542 | 2.6 | 2.3–2.9 | < 0.001 |
ANRESIS: Swiss Centre for Antibiotic Resistance; CI: confidence interval; non-kv-kpc: non-Klebsiella variicola-Klebsiella pneumoniae-complex; OR: odds ratio.
Fisher’s exact tests were used and p values adjusted using sequential Bonferroni corrections. Non-kv-kpc refers to all species of the K. pneumoniae complex except K. variicola. Carbapenem includes imipenem and meropenem, cephalosporin 3rd/4th generation includes all third- and fourth-generation cephalosporins available in the ANRESIS database (i.e. cefepime, cefixime, cefotaxime, cefpodoxime, ceftazidime, ceftibuten and ceftriaxone). Resistant isolates consist of all isolates categorised resistant (R) by reporting laboratories.
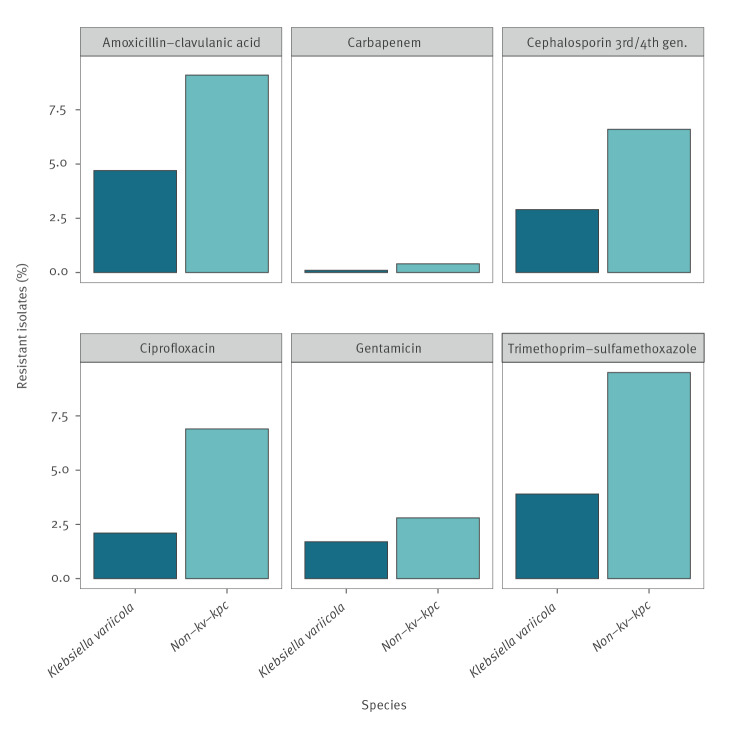
Figure.
Antibiotic resistance rates in Klebsiella variicola isolates (n = 9,899) vs non-kv-kpc isolates (n = 60,215) collected by ANRESIS, Switzerland, 1 January 2017–13 June 2022
ANRESIS: Swiss Centre for Antibiotic Resistance; gen.: generation; non-kv-kpc: non-Klebsiella variicola-Klebsiella pneumoniae-complex.
Non-kv-kpc refers to all species of the K. pneumoniae complex except K. variicola. Carbapenem includes imipenem and meropenem, cephalosporin 3rd/4th gen. includes all third- and fourth-generation cephalosporins available in the ANRESIS database (i.e. cefepime, cefixime, cefotaxime, cefpodoxime, ceftazidime, ceftibuten and ceftriaxone).
Non-kv-kpc and K. variicola isolates showed, respectively, the highest resistance rates against trimethoprim-sulfamethoxazole (9.5% and 3.9%) and against β-lactams (maximum rate for amoxicillin-clavulanic acid: 9.1 and 4.7%), followed by ciprofloxacin (6.9 and 2.1%). Resistance rates against gentamicin, however, only showed a small difference with 2.8% for non-kv-kpc and 1.7% for K. variicola isolates, and carbapenem resistance rates were generally low (respectively 0.4 and 0.1%), resulting in broader confidence intervals.
In addition to different resistance rates, K. variicola isolates were significantly more often reported from blood and primarily sterile specimens than non-kv-kpc isolates, namely 8.7% K. variicola isolates (n = 9,899) and 6.1% of non-kv-kpc isolates (n = 60,215) (Fisher’s exact test: odds ratio = 0.68, 95% confidence interval: 0.63–0.73; p < 0.001).
Discussion
Our data show that K. variicola isolates of the ANRESIS database were significantly more susceptible to six antibiotic classes tested than non-kv-kpc isolates. This is consistent with Cuénod et al., a collaborative international study of several institutions published in 2021 [17]. That study used new generation sequencing data and their own MALDI-TOF spectra to identify eight species of Klebsiella within the K. pneumoniae and K. oxytoca complex and genomic data on the abundance of antimicrobial resistance genes. The authors suggested that higher susceptibility rates in K. variicola may be linked to lower abundance of plasmids.
Differential susceptibility rates may affect surveillance data, as laboratories not differentiating K. variicola (in our study accounting for 14.1% of all K. pneumoniae complex isolates) tend to underestimate resistance rates of non-kv-kpc isolates. Similar concerns have been raised by a recent review focusing on genomic data of the K. oxytoca complex. The authors describe several resistance and virulence genes with the potential to create multidrug-resistant pathogens [18]. Based on data from SENTRY, a global antimicrobial surveillance programme collecting in vitro antimicrobial susceptibility data, the authors observed increasing non-susceptibility rates to carbapenems and cephalosporins over 7 years and highly variable non-susceptibility rates across regions, with higher rates in Asia and Europe than in North America [18]. However, confounding biases as described in our study could not be fully considered due to lack of data. In future large-scale studies comparing different regions and countries, these issues should be addressed to improve our understanding of epidemiological contexts. Similar methodological issues arise in a study by Holt et al. in 2015 [5], showing that the abundance of resistance genes of K. pneumoniae isolates from six countries (Australia, Indonesia, Laos, Singapore, the United States and Vietnam) varied between countries. Even within Europe, resistance rates vary considerably, increasing from north to south and from west to east, with Switzerland among the countries with lowest resistance rates [19]. These differences are consistent over time, and even though the confounding bias described in our study may be negligible in some countries, data homogenisation is still needed to increase our understanding of the epidemiological trends in these different species.
Higher resistance rates come with higher abundance of transferable resistance genes [5,17], potentially increasing opportunities for gene transfers between species [20], which may decrease differences in resistance rates between species. This may be especially true in environments where closely related species occur in close proximity [21]. Thus, the impact of an estimation bias of epidemiology and resistance rates may be less severe in countries with higher resistance rates than in countries with lower resistance rates such as Switzerland. Future studies including data from different countries will have to clarify potential differential bias effects between countries.
Studies estimating global health burdens based on statistical modelling or machine learning rely on empirical data of mortality, resistance and virulence genes. These data are obtained from literature reviews, hospitals and surveillance initiatives. Up-to-date knowledge of taxonomy, virulence and antimicrobial resistance is thus crucial to avoid misleading conclusions about the clinical significance or resistance rates of some pathogens [22,23]. Lower susceptibility rates of K. variicola, for example, may lead to an underestimation of resistance rates of other members of the K. pneumoniae complex and thus to less appropriate treatments of infections. This may affect possible positive effects of appropriately tailored antimicrobial treatments [24].
Our data also indicate that K. variicola may be more invasive than non-kv-kpc. However, this association has to be confirmed by clinical and functional studies addressing underlying causes. A recent study by Cuénod et al. comparing different clinical outcomes of patients infected with different Klebsiella species did not confirm our data [17], whereas a review of several earlier studies reported associations with blood stream infections in animals and humans [9].
A growing body of data, including our study, suggests that discrimination of species within K. pneumoniae complex will increase our knowledge of species-specific susceptibilities and clinical significance, providing valuable information to clinicians and epidemiologists [5,6,9,17]. Correct differentiation of K. variicola from the K. pneumoniae complex is possible not only by molecular methods such as sequencing of rpoB, fusA, gapA, gyrA, leuS genes [12,25], but also by updated MALDI-TOF databases used in medical diagnostic laboratories. Since 2017, MALDI-TOF profiles of K. variicola have been included in commercial databases allowing identification of K. variicola. However, differentiation of K. variicola from the K. pneumoniae complex by MALDI-TOF is not yet accepted by all microbiological laboratories because the close relatedness of Klebsiella spp. prevents verification by 16S rDNA sequencing, a species identification technique used as gold standard by many laboratories. Rodrigues et al. and Cuénod et al. both independently developed MALDI-TOF databases which allow differentiation of all currently known species of the K. pneumoniae complex, but discriminatory precision of these two MALDI-TOF databases have not yet been validated using large datasets [17,25]. Continued optimisation of identification methods will be necessary to improve discriminatory power between species of the K. pneumoniae complex.
Conclusion
Higher susceptibility and invasiveness rates of K. variicola isolates in our ANRESIS antimicrobial resistance database suggest that refined differentiation of K. pneumoniae complex can improve our understanding of its ecological niches, epidemiology, antibiotic susceptibility and clinical significance, thus providing more accurate information to clinicians and epidemiologists. In addition, refined species information has the potential to improve antibiotic prescribing by tailoring antibiotic treatments more appropriately.
Ethical statement
Data extracted from the ANRESIS database have been anonymised and provided by contributing laboratories from bacterial strains isolated during clinical routine diagnostic procedures. Thus, no ethical approval or informed consent was required by the Swiss human research act to conduct this data analysis.
Funding statement
ANRESIS is funded by the Swiss Federal Office of Public Health and the Institute for Infectious Diseases of the University Bern, Switzerland. There was no specific funding of this project.
Acknowledgements
We thank all laboratories who provided their antimicrobial resistance data to the ANRESIS database and to our anonymous reviewers for their invaluable comments which improved our manuscript significantly.
Conflict of interest
Authors’ contributions
References
Supplementary Data
None declared.
IKV analysed the data and wrote the manuscript incorporating input from all authors, CL initiated the research question and contributed to the critical manuscript review and data interpretation, MG supported and proofread data analysis, AK supervised this work including data interpretation and critical review of the manuscript. All authors read and approved the final manuscript.
References
- 1. Munson E, Carroll KC. An update on the novel genera and species and revised taxonomic status of bacterial organisms described in 2016 and 2017. J Clin Microbiol. 2019;57(2):e01181-18. 10.1128/JCM.01181-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Munson E, Carroll KC. Summary of novel bacterial isolates derived from human clinical specimens and nomenclature revisions published in 2018 and 2019. J Clin Microbiol. 2021;59(2):e01309-20. 10.1128/JCM.01309-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Torres-Sangiao E, Leal Rodriguez C, García-Riestra C. Application and perspectives of MALDI-TOF mass spectrometry in clinical microbiology laboratories. Microorganisms. 2021;9(7):1539. 10.3390/microorganisms9071539 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Giraud-Gatineau A, Texier G, Fournier PE, Raoult D, Chaudet H. Using MALDI-TOF spectra in epidemiological surveillance for the detection of bacterial subgroups with a possible epidemic potential. BMC Infect Dis. 2021;21(1):1109. 10.1186/s12879-021-06803-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Holt KE, Wertheim H, Zadoks RN, Baker S, Whitehouse CA, Dance D, et al. Genomic analysis of diversity, population structure, virulence, and antimicrobial resistance in Klebsiella pneumoniae, an urgent threat to public health. Proc Natl Acad Sci USA. 2015;112(27):E3574-81. 10.1073/pnas.1501049112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Wyres KL, Lam MMC, Holt KE. Population genomics of Klebsiella pneumoniae. Nat Rev Microbiol. 2020;18(6):344-59. 10.1038/s41579-019-0315-1 [DOI] [PubMed] [Google Scholar]
- 7. Long SW, Linson SE, Ojeda Saavedra M, Cantu C, Davis JJ, Brettin T, et al. Whole-genome sequencing of a human clinical isolate of the novel species Klebsiella quasivariicola sp. nov. Genome Announc. 2017;5(42):e01057-17. 10.1128/genomeA.01057-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Potter RF, Lainhart W, Twentyman J, Wallace MA, Wang B, Burnham CA, et al. Population structure, antibiotic resistance, and uropathogenicity of Klebsiella variicola. MBio. 2018;9(6):e02481-18. 10.1128/mBio.02481-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Rodríguez-Medina N, Barrios-Camacho H, Duran-Bedolla J, Garza-Ramos U. Klebsiella variicola: an emerging pathogen in humans. Emerg Microbes Infect. 2019;8(1):973-88. 10.1080/22221751.2019.1634981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Andrade BGN, de Veiga Ramos N, Marin MF, Fonseca EL, Vicente AC. The genome of a clinical Klebsiella variicola strain reveals virulence-associated traits and a pl9-like plasmid. FEMS Microbiol Lett. 2014;360(1):13-6. 10.1111/1574-6968.12583 [DOI] [PubMed] [Google Scholar]
- 11. Maatallah M, Vading M, Kabir MH, Bakhrouf A, Kalin M, Nauclér P, et al. Klebsiella variicola is a frequent cause of bloodstream infection in the Stockholm area, and associated with higher mortality compared to K. pneumoniae. PLoS One. 2014;9(11):e113539. 10.1371/journal.pone.0113539 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Barrios-Camacho H, Aguilar-Vera A, Beltran-Rojel M, Aguilar-Vera E, Duran-Bedolla J, Rodriguez-Medina N, et al. Molecular epidemiology of Klebsiella variicola obtained from different sources. Sci Rep. 2019;9(1):10610. 10.1038/s41598-019-46998-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.World Health Organisation (WHO). Antimicrobial resistance. Geneva: WHO; 2021. Available from: https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance
- 14. Barnsteiner S, Baty F, Albrich WC, Babouee Flury B, Gasser M, Plüss-Suard C, et al. Antimicrobial resistance and antibiotic consumption in intensive care units, Switzerland, 2009 to 2018. Euro Surveill. 2021;26(46):2001537. 10.2807/1560-7917.ES.2021.26.46.2001537 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.European Committee on Antimicrobial Susceptibility Testing (EUCAST). New definitions of S, I and R from 2019. Växjö: EUCAST. [Accessed: 7 Jul 2022]; Available from: https://www.eucast.org/newsiandr
- 16.R Core Team. R: A language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2021. Available from: https://www.R-project.org
- 17. Cuénod A, Wüthrich D, Seth-Smith HMB, Ott C, Gehringer C, Foucault F, et al. Whole-genome sequence-informed MALDI-TOF MS diagnostics reveal importance of Klebsiella oxytoca group in invasive infections: a retrospective clinical study. Genome Med. 2021;13(1):150. 10.1186/s13073-021-00960-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Yang J, Long H, Hu Y, Feng Y, McNally A, Zong Z. Klebsiella oxytoca complex: update on taxonomy, antimicrobial resistance, and virulence. Clin Microbiol Rev. 2022;35(1):e0000621. 10.1128/CMR.00006-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.World Health Organization Regional Office for Europe (WHO/Europe)/European Centre for Disease Prevention and Control. Antimicrobial resistance surveillance in Europe 2022 – 2020 data. 2022, Copenhagen: WHO/Europe; 2022. Available from: https://www.ecdc.europa.eu/en/publications-data/antimicrobial-resistance-surveillance-europe-2022-2020-data
- 20. Liao H, Friman VP, Geisen S, Zhao Q, Cui P, Lu X, et al. Horizontal gene transfer and shifts in linked bacterial community composition are associated with maintenance of antibiotic resistance genes during food waste composting. Sci Total Environ. 2019;660:841-50. 10.1016/j.scitotenv.2018.12.353 [DOI] [PubMed] [Google Scholar]
- 21. Broaders E, Gahan CG, Marchesi JR. Mobile genetic elements of the human gastrointestinal tract: potential for spread of antibiotic resistance genes. Gut Microbes. 2013;4(4):271-80. 10.4161/gmic.24627 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Murray CJL, Ikuta KS, Sharara F, Swetschinski L, Robles Aguilar G, Gray A, et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet. 2022;399(10325):629-55. 10.1016/S0140-6736(21)02724-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kim JI, Maguire F, Tsang KK, Gouliouris T, Peacock SJ, McAllister TA, et al. Machine learning for antimicrobial resistance prediction: current practice, limitations, and clinical perspective. Clin Microbiol Rev. 2022;e0017921. [DOI] [PMC free article] [PubMed]
- 24. Nathwani D, Varghese D, Stephens J, Ansari W, Martin S, Charbonneau C. Value of hospital antimicrobial stewardship programs [ASPs]: a systematic review. Antimicrob Resist Infect Control. 2019;8(1):35. 10.1186/s13756-019-0471-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Rodrigues C, Passet V, Rakotondrasoa A, Brisse S. Identification of Klebsiella pneumoniae, Klebsiella quasipneumoniae, Klebsiella variicola and related phylogroups by MALDI-TOF mass spectrometry. Front Microbiol. 2018;9:3000. 10.3389/fmicb.2018.03000 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.