Abstract
Four groups of plant homeodomain proteins contain a dimerization motif closely linked to the homeodomain. We here show that two sunflower homeodomain proteins, Hahb-4 and HAHR1, which belong to the Hd-Zip I and GL2/Hd-Zip IV groups, respectively, show different binding preferences at a defined position of a pseudopalindromic DNA-binding site used as a target. HAHR1 shows a preference for the sequence 5′-CATT(A/T)AATG-3′, rather than 5′-CAAT(A/T)ATTG-3′, recognized by Hahb-4. To analyze the molecular basis of this behavior, we have constructed a set of mutants with exchanged residues (Phe→Ile and Ile→Phe) at position 47 of the homeodomain, together with chimeric proteins between HAHR1 and Hahb-4. The results obtained indicate that Phe47, but not Ile47, allows binding to 5′-CATT(A/T)AATG-3′. However, the preference for this sequence is determined, in addition, by amino acids located C-terminal to residue 53 of the HAHR1 homeodomain. A double mutant of Hahb-4 (Ile47→Phe/Ala54→Thr) shows the same binding behavior as HAHR1, suggesting that combinatorial interactions of amino acid residues at positions 47 and 54 of the homeodomain are involved in establishing the affinity and selectivity of plant dimeric homeodomain proteins with different DNA target sequences.
INTRODUCTION
The homeodomain (HD) is a 61 amino acid protein motif found in eukaryotic transcription factors generally involved in the regulation of developmental processes (1–3). It folds into a characteristic three-helix structure that interacts specifically with DNA (3–6). Most contacts are established by amino acids within helix III (the recognition helix), which fits into the major groove of DNA (7–10). The N-terminal portion of the HD (the first nine amino acids) forms a disordered arm that protrudes from helix I and orients towards the DNA, making several specific contacts within the minor groove (7–10). In spite of a structural resemblance between the HD and the helix–turn–helix motif, a striking difference is that HDs usually bind DNA as monomers (11,12). A group of about seven amino acids is conserved in almost all HDs recognized so far. Four of these are hydrophobic amino acids that participate in maintaining the structure of the three-helix core, two are base-contacting amino acids from helix III and the remaining one is Arg5, a DNA-contacting amino acid from the N-terminal arm (3).
HDs are present in almost every eukaryotic organism that has been investigated. In plants, several families of HD proteins have been described (13). One of these families, Hd-Zip, comprises proteins with a typical leucine zipper motif adjacent to the C-terminal end of the HD (14,15). As expected, these proteins bind DNA as dimers (16). Hd-Zip proteins have been grouped into different subfamilies according to sequence conservation (13,17). Hd-Zip I and Hd-Zip II proteins are more related to each other than to other subfamilies and bind the pseudopalindromic sequence 5′-CAAT(N)ATTG-3′ (16,18). Hd-Zip III proteins have four additional amino acids between the HD and the leucine zipper and bind the extended sequence 5′-GTAAT(G/C)ATTAC-3′ (19). HD proteins from a separate group, which are thought to be involved in several aspects of epidermal cell differentiation in plants (20–25), also possess a dimerization motif that resembles a leucine zipper, composed of two short heptad repeats connected by a loop (21,26). This dimerization motif is able to replace the canonical leucine zipper present in the Hd-Zip II protein Athb-2 (21). This has prompted some authors to include this class of proteins within the Hd-Zip family (thus forming the Hd-Zip IV subfamily), while others have grouped these proteins as a separate class, termed GL2, after the first member that was identified (13,17,22). The HD of GL2/Hd-Zip IV proteins differs from those of other Hd-Zip proteins in several positions, some of them important for DNA binding in other HDs (3,7,10). Based on what is known about Hd-Zip I proteins, we have performed studies on the influence of some amino acids on the DNA binding specificity of dimeric HD proteins. We present evidence that the combination of amino acids at positions 47 and 54 of the HD determine the specificity of binding at a single position within the binding site recognized by each monomer. The combination present in GL2/Hd-Zip IV proteins (Phe and Thr, respectively) favors preferential recognition of A instead of T at the fourth position of the sequence 5′-TAAT-3′, recognized by other Hd-Zip proteins and most animal HDs.
MATERIALS AND METHODS
Cloning, expression and purification of recombinant proteins
Hahb-4 coding sequences were amplified and cloned in-frame into the BamHI and EcoRI sites of the expression vector pGEX-3X (27) as described previously (28). Amplifications were performed using oligonucleotides H4F (5′-GGCGGATCCCAACAGAAACAACCACCAGG-3′) and H4R (5′-GGCGAATTCACATTTCTCAGCACCTCC-3′). For construction of the Phe47 mutant of Hahb-4, oligonucleotides 5′-GGCCGATCGCGCGCGTTTGTTCTGGAACCAGAACGCCAC-3′ (introduced mutations underlined) and 5′-GCGCGATCGAAGTCGAG-3′, which generate PvuI restriction sites, were used in PCR reactions in conjunction with H4F and H4R, respectively, to generate fragments representing the N- and C-terminal portions of Hahb-4. These fragments were digested with PvuI and ligated and the resulting product was amplified with H4F and H4R. The amplified fragment was cloned into pGEX-3X as described above.
An NheI/XbaI fragment, containing HAHR1 sequences coding for the entire HD and dimerization motif, was excised from pUC119 with HincII and EcoRI and cloned into the SmaI and EcoRI sites of pGEX-3X as described (28). For the production of a protein with only the first half of the dimerization motif, a KpnI deletion of this clone was used. Both types of proteins showed the same DNA binding preferences. An Ile47 mutant was constructed using primers 5′-CCGAGATCTTCACTTGTCTAGGATG-3′ and 5′-CGCAGATCTGGTTTCAAAATCGCC-3′ (mutations underlined) to amplify the N- and C-terminal portions of HAHR1, respectively, together with vector-specific primers. The products were digested with BglII, ligated, amplified with vector primers and cloned into pGEX-3X.
Fragments encoding chimeric proteins were synthesized using primers containing Hahb-4 sequences fused to HAHR1 sequences between codons 53 and 54 of the HD. Primers 5′-GATTTGAGTGCGTTTGTTCTGGAACCA-3′ and 5′-AACAAACGCACTCAAATCAAGACTATA-3′ (Hahb-4 sequences underlined) were used to amplify hybridizing Hahb-4 N-terminal sequences and HAHR1 C-terminal sequences. The resulting products were mixed in buffer containing 50 mM Tris–HCl, pH 7.2, 10 mM MgSO4 and 0.1 mM DTT, incubated at 95°C for 5 min and annealed by allowing the solution to reach 24°C in ∼1 h. After this, 0.5 mM each dNTP and 5 U Escherchia coli DNA polymerase I Klenow fragment were added and incubation was followed for 1 h at 37°C. A portion of this reaction was directly used to amplify the chimeric fragments essentially as described above. In a similar way, primers 5′-AATCGCCGTGCGCGATCAAAGTCGAGG-3′ and 5′-TGATCGCGCACGGCGATTTTGAAACCA-3′ were used to amplify Hahb-4 C-terminal sequences and HAHR1 N-terminal sequences.
The Phe47/Thr54 double mutant of Hahb-4 was constructed using oligonucleotides 5′-TGATCGCGTGCGTTTGTTCTG-3′ and 5′-AACAAACGCACGCGATCAAAG-3′ (mutations underlined), together with primers H4F and H4R, to amplify partially overlapping N- and C-terminal Phe47-Hahb-4 mutant fragments. The resulting fragments were hybridized and amplified as described above. The Ile47/Ala54 double mutant of HAHR1 was constructed in a similar way using oligonucleotides 5′-TGATTTGAGCACGGCGATT-3′ and 5′-AATCGCCGTGCTCAAATCA-3′, together with vector primers, and Ile47-HAHR1 as template. All constructions were checked by DNA sequence analysis.
For expression, E.coli cells bearing the corresponding plasmids were grown and induced as described previously (28). Purification by affinity chromatography and cleavage of the fusion products were carried out essentially as described by Smith and Johnson (27), with modifications described by Palena et al. (28). Purified proteins (>95% as judged by Coomassie Brilliant Blue staining of denaturing polyacrylamide gels) were used for the assays. Protein amounts were measured as described by Sedmak and Grossberg (29).
DNA binding assays
For electrophoretic mobility shift assays, aliquots of purified proteins were incubated with double-stranded DNA (0.3–0.6 ng, 30000 c.p.m., labeled with [α-32P]dATP by filling in the 3′-ends using the Klenow fragment of DNA polymerase) generated by hybridization of the complementary oligonucleotides 5′-AATTCAGATCTCAATAATTGAGAG-3′ and 5′-GATCCTCTCAATTATTGAGATCTG-3′ (binding site for Hahb-4 underlined) or derivatives with modifications within the binding sequence, as described in the text. Binding reactions (20 µl) containing 20 mM HEPES, pH 7.5, 50 mM KCl, 2 mM MgCl2, 0.5 mM EDTA, 1.0 mM DTT, 0.5% Triton X-100, 1 µg poly(dI·dC) and 10% glycerol were incubated for 20 min at room temperature, supplemented with 2.5% Ficoll and immediately loaded onto a running gel (5% acrylamide, 0.08% bis-acrylamide in 0.5× TBE plus 2.5% glycerol (1× TBE = 90 mM Tris–borate, pH 8.3, 2 mM EDTA). The gel was run in 0.5× TBE at 30 mA for 1.5 h and dried prior to autoradiography. When comparing binding to different oligonucleotides, equal amounts of DNA (as judged by absorbance at 260 nm before labeling and by scintillation counting after labeling) were used. The assays shown are representative of at least three experiments that yielded essentially the same results.
RESULTS
In order to characterize the determinants of DNA binding specificity in plant Hd-Zip proteins, we have compared the relative affinity of the Hd-Zip I protein Hahb-4 and the Hd-Zip IV/GL2 protein HAHR1 (30) for variants of the pseudopalindromic sequence 5′-CAAT(A/T)ATTG-3′ recognized by Hahb-4 (18) and, presumably, all other Hd-Zip I proteins. Changes were introduced at two symmetrical positions in this sequence (i.e. 3 and 7 or 4 and 6) since they presumably interact with similar portions of each monomer. Figure 1 shows that HAHR1 binds preferentially to an oligonucleotide with the sequence 5′-CATT(A/T)AATG-3′, which contains A/T transversions at positions 3 and 7. Notably, a weaker, though significant, binding to 5′-CAAT(A/T)ATTG-3′ was also observed. Conversely, the presence of G:C base pairs at these positions was not tolerated (Fig. 1). Any change at positions 4 and 6 also produced a marked decrease in binding affinity (Fig. 1 and results not shown). In different experiments, proteins bearing the HD and either the complete dimerization motif of HAHR1 (26) or the first half of this motif (helix I) were used. Since both proteins showed similar DNA binding preferences, most experiments were carried out with the shorter form, since it is more stable both in E.coli and after purification.
Figure 1.
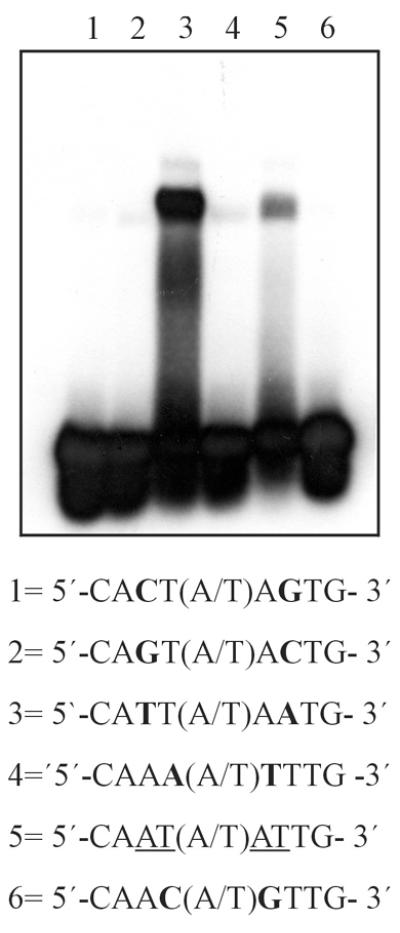
Binding of HAHR1 to oligonucleotides containing different binding sequences. A fragment of HAHR1 composed of the entire HD plus the first half of its dimerization motif (5 pmol) was analyzed by an electrophoretic mobility shift assay for binding to a 24mer synthetic double-stranded oligonucleotide containing variants at two equivalent positions of the sequence CAAT(A/T)ATTG preferred by Hd-Zip I proteins.
The results obtained indicate that HAHR1 displays a different DNA binding specificity to Hahb-4, at least within one of the positions recognized by each monomer. According to current models of the interaction of Hd-Zip proteins with DNA (16), which rely on the crystallographic and NMR structures of monomeric animal HDs (3,10), this position should be contacted by the residue at position 47. This residue is Ile or Val in Hd-Zip I and II proteins (and most animal HDs), while it is Phe in HAHR1 and all other GL2/Hd-Zip IV proteins described up to now. To directly test the role of residue 47 in DNA binding specificity, we have mutated Phe47 in HAHR1 to Ile and Ile47 in Hahb-4 to Phe. Ile47-HAHR1 showed a strong reduction in affinity for 5′-CATT(A/T)AATG-3′ and a slight preference for the 5′-CAAT(A/T)ATTG-3′ sequence, indicating that Ile at this position is a relevant determinant of the specificity of interaction with A and T at positions 3 and 7 (Fig. 2). Although the mutated protein displayed a reduced affinity, indicating that the mutation had disturbed positive interactions, a significant change in specificity was observed.
Figure 2.
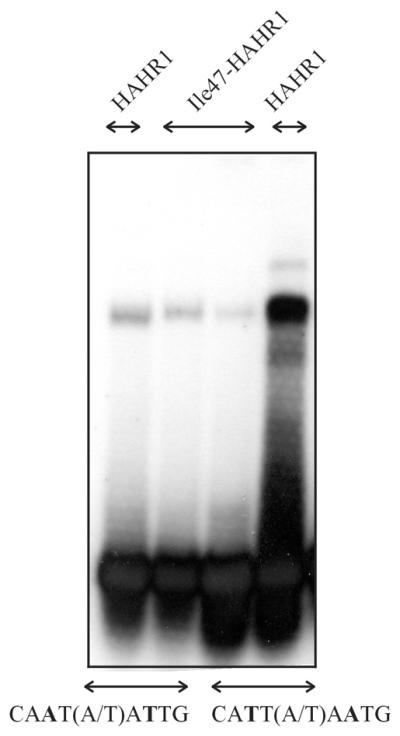
Ile47-HAHR1 shows a preference for 5′-CAAT(A/T)ATTG-3′. Binding of native (Phe47) and Ile47-HAHR1 (5 pmol) to oligonucleotides containing the sequence 5′-CAAT(A/T)ATTG-3′ or 5′-CATT(A/T)AATG-3′ was analyzed as described.
A symmetrical change in Hahb-4 (Ile47→Phe) produced a protein with a significant affinity for 5′-CATT(A/T)AATG-3′, not recognized by wild-type Hahb-4 (Fig. 3), while oligonucleotides with C:G or G:C base pairs at the underlined positions were not bound (not shown). Nevertheless, the mutant protein still showed a preference for 5′-CAAT(A/T)ATTG-3′, although a moderate decrease in binding affinity was evident with respect to wild-type Hahb-4. Again, these results point to an essential role of residue 47 in determining DNA binding specificity at positions 3 and 7. While Ile determines an absolute preference for a T:A base pair, the presence of Phe allows binding to sequences containing either T:A or A:T at these positions within the context of both the Hahb-4 and HAHR1 HDs. The preferences of proteins with Phe at position 47 (HAHR1 and Phe47-Hahb-4), however, indicate that differences in other portions of the HD also influence binding specificity at this single position.
Figure 3.
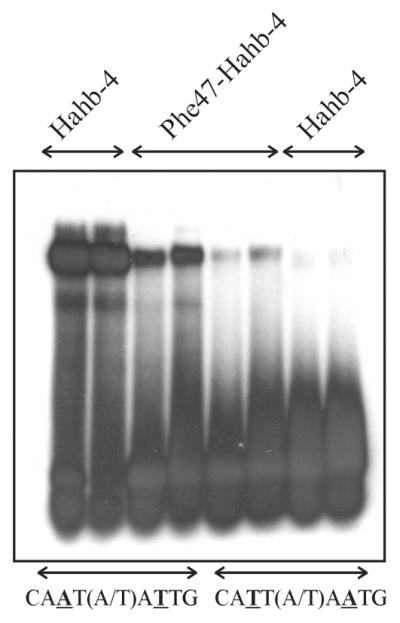

Phe47-Hahb-4 binds to 5′-CAAT(A/T)ATTG-3′ and 5′-CATT(A/T)AATG-3′. Binding of native (Ile47) and Phe47-Hahb-4 to oligonucleotides containing the sequence 5′-CAAT(A/T)ATTG-3′ or 5′-CATT(A/T)AATG-3′ was analyzed as described. For each purified protein, two different concentrations (3 pmol in lanes 1, 3, 5 and 7 and 6 pmol in lanes 2, 4, 6 and 8) were used.
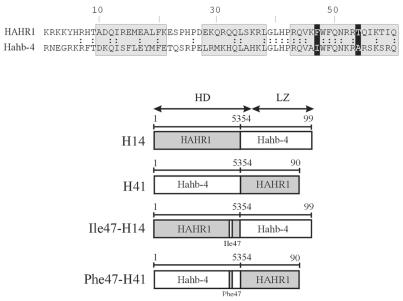
To ascertain which portions of the HD, besides residue 47, are involved in this interaction, we constructed chimeric proteins between HAHR1 and Hahb-4 containing amino acids 1–53 of each protein fused to the rest of the HD and dimerization motif of the other (Fig. 4). It is noteworthy that residues 48–53 are identical in Hahb-4 and HAHR1, with the sole exception of residue 52 (Lys in Hahb-4, Arg in HAHR1), which is not likely to be involved in the determination of specificity (i.e. Hd-Zip I proteins with Arg52 show the same binding specificity as Hahb-4; 16). The chimeric proteins were then used to monitor the influence of sequences N-terminal to residue 47 and C-terminal to residue 53. Protein H41, with Hahb-4 N-terminal sequences followed by HAHR1 C-terminal ones, binds to 5′-CAAT(A/T)ATTG-3′ with low affinity and not at all to 5′-CATT(A/T)AATG-3′, reinforcing the view that Ile47 directs the preference for the first of these sequences (Fig. 5). H14, on the other hand, displayed similar binding to both DNA sequences, with only a slight preference for 5′-CAAT(A/T)ATTG-3′ (Fig. 5). These results suggest that residues C-terminal to amino acid 53 must be responsible for the different binding characteristics of proteins with either Ile or Phe at position 47.
Figure 4.
Chimeric proteins between HAHR1 and Hahb-4. (Top) An amino acid sequence comparison for the HDs of HAHR1 and Hahb-4. Double dots indicate identical amino acids; grey boxes denote helices I, II and III; black boxes indicate the amino acids at positions 47 and 54. (Bottom) A schematic representation of proteins containing portions of HAHR1 and Hahb-4 is shown. Grey and white boxes indicate HAHR1 and Hahb-4 sequences, respectively. Amino acids are numbered from the N-terminus of the HD. Mutations introduced at position 47 are indicated.
Figure 5.
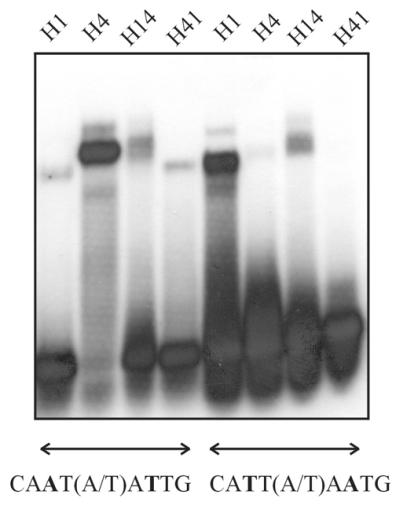
Binding of chimeric proteins between HAHR1 and Hahb-4 to oligonucleotides containing different binding sequences. Proteins (5 pmol each) containing either the first 53 residues of the HAHR1 HD fused to the rest of the HD and dimerization motif of Hahb-4 (H14) or the first 53 residues of the Hahb-4 HD fused to the rest of the HD and dimerization motif of HAHR1 (H41) were analyzed for binding to oligonucleotides containing the sequence 5′-CAAT(A/T)ATTG-3′ or 5′-CATT(A/T)AATG-3′. Native HAHR1 (H1) and Hahb-4 (H4) were included for comparison.
If this assertion is valid, we should be able to recreate the binding site preferences of HAHR1 with a protein containing Phe47 and the HAHR1 C-terminal sequence from amino acid 54 to 90. To test this, we have constructed chimeras with exchanged residues at position 47. As shown in Figure 6, Phe47-H41 exactly matches the binding behavior of native HAHR1, showing a clear preference for 5′-CATT(A/T)AATG-3′. This result suggests that one or more amino acid residues located beyond residue 53 influence binding specificity at the underlined positions of proteins containing Phe47. Ile47-H14, on the other hand, showed a slight preference for 5′-CAAT(A/T)ATTG-3′ and an overall reduced affinity. In this case, residues from the N-terminal part of Hahb-4 may also be required for efficient binding. In this respect, we have recently shown that the N-terminal arm of the Hahb-4 HD is an important determinant of DNA binding affinity (31).
Figure 6.
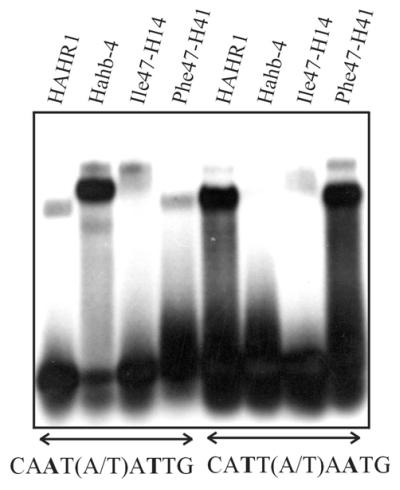
Binding of chimeric proteins with exchanged residues at position 47 to oligonucleotides containing different binding sequences. Proteins (5 pmol each) containing either the first 53 residues of the Ile47-HAHR1 HD fused to the rest of the HD and dimerization motif of Hahb-4 (Ile47-H14) or the first 53 residues of the Phe47-Hahb-4 HD fused to the rest of the HD and dimerization motif of HAHR1 (Phe47-H41) were analyzed for binding to oligonucleotides containing the sequence 5′-CAAT(A/T)ATTG-3′ or 5′-CATT(A/T)AATG-3′. Native HAHR1 (5 pmol) and Hahb-4 (5 pmol) were included for comparison.
A close examination of residues that may participate in DNA binding suggests that Thr54 in HAHR1 may be a good candidate, since it must be located facing the DNA in a similar orientation to residue 47 within an α-helix. Residue 54 is Ala in Hahb-4, and its side chain may be too short to contact DNA, as has been observed for the engrailed–DNA complex (7). To directly test the role of Thr54 in determining the observed DNA binding specificity, we constructed the Hahb-4 double mutant Ile47→Phe/Ala54→Thr. This mutant showed a clear preference for 5′-CATT(A/T)AATG-3′ and also bound 5′-CAAT(A/T)ATTG-3′, although with lower affinity (Fig. 7). This behavior exactly matches that of HAHR1 and Phe47-H41. We conclude that the combination of amino acid residues at positions 47 and 54 of the HD defines the specificity of the interaction of dimeric plant HD proteins with base pairs 3 and 7 of the pseudopalindromic target site used in this study.
Figure 7.
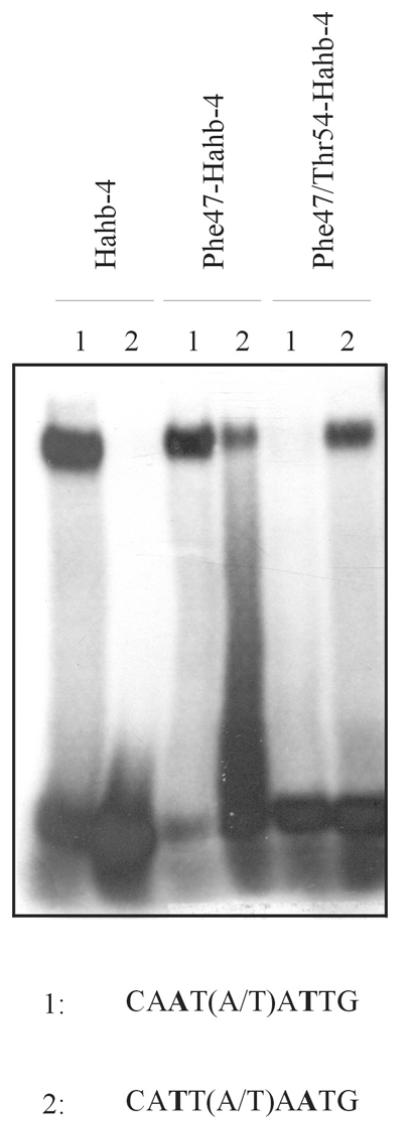
Phe47/Thr54-Hahb-4 binds preferentially to 5′-CATT(A/T)AATG-3′. A double mutant of Hahb-4 (5 pmol) containing Phe and Thr at positions 47 and 54 of the HD, respectively, was analyzed for binding to two different oligonucleotides. Native Hahb-4 (5 pmol) and Phe47-Hahb-4 (5 pmol) were included for comparison.
The opposite change in HAHR1 (Phe47→Ile/Thr54→Ala) produced a protein with an overall reduced affinity for DNA, similar to Ile47-H14 (results not shown). As discussed above, this indicates that other parts of the Hahb-4 HD must be present for efficient binding to DNA of proteins with this combination of amino acids.
DISCUSSION
In the present work, we have investigated the molecular basis of the DNA binding specificity of dimeric plant HD proteins. We have been able to find a correlation between the presence of certain amino acid residues at defined positions and the preference for a given target sequence. Current models on DNA binding by Hd-Zip proteins (16) are based on comparisons of conserved amino acids also present in animal HDs, the structures and DNA binding properties of which have been studied by X-ray crystallography (7,9) and NMR (4,8). These models have been validated by experimental evidence showing that Hd-Zip I and Hd-Zip II proteins bind a 9 bp pseudopalindromic DNA sequence of the type 5′-CAATNATTG-3′ (16), which can be decomposed into two antiparallel partially overlapping 5′-TNATTG-3′ sequences, similar to those bound by most animal HDs. The third and fourth positions within this sequence are recognized by Asn51 (almost invariant) and Ile/Val47 (highly conserved) in animal HDs, while the fifth and sixth positions are thought to interact with amino acids at positions 50 and 54 of the HD (3,10,32). Hd-Zip I and Hd-Zip II proteins contain Asn at position 51, Ile/Val at position 47, Gln (as in Antennapedia and engrailed) at position 50 and Ala (as in engrailed, Met in Antennapedia) at position 54 (33), suggesting a similar mode of interaction. Sessa et al. (16,34) have proposed that position 2 (the central position of the pseudopalindromic sequence) could be in contact with Arg55 in Hd-Zip I and Hd-Zip II proteins.
Hd-Zip IV/GL2 proteins contain a bipartite dimerization motif that resembles a leucine zipper interrupted by a loop (26). The relative location of this motif with respect to the HD is similar to that observed in Hd-Zip I and Hd-Zip II proteins. We reasoned, then, that a comparative analysis of binding of the Hd-Zip I protein Hahb-4 and the Hd-Zip IV/GL2 protein HAHR1 to variants of the sequence 5′-CAAT(A/T)ATTG-3′ might provide clues as to the interactions established by each HD monomer. These studies indicate that HAHR1 shows a preference for 5′-CATT(A/T)AATG-3′. This means that each monomer is able to recognize the sequence 5′-TNAATG-3′ and that Phe47 may be responsible for the different sequence specificities. That Ile47-HAHR1 shows the same specificity as the Hd-Zip I and Hd-Zip II proteins and that Phe47-Hahb-4 is able to bind to 5′-CATT(A/T)AATG-3′ with significant affinity also support this notion. This is the first indication that changes in the amino acid residue present at position 47 produce changes in binding specificity at positions 3 and 7 of the pseudopalindromic binding sequence. This change in specificity most likely reflects the existence of direct amino acid–base pair contacts at these positions. Our data also indicate that Phe at position 47 favors the interaction with 5′-TAAA-3′, instead of the sequence 5′-TAAT-3′ bound by most HDs. Of 750 HD sequences described in The Homeodomain Resource Database (35), we have found only two that contain Phe at this position: one of them is GL2 and the other is PIT-1 from the pig, a POU-HD transcription factor. Thus, GL2/Hd-Zip IV proteins constitute a special group of proteins derived from an ancestor that has acquired a new specificity by incorporating Phe47 into the recognition helix.
Binding site preferences different from 5′-TAAT-3′ have been observed for other HDs, including MATα2 (5′-TTAC-3′; 9), TTF-1 (5′-CAAG-3′; 36) and the two HDs of Zmhox2a (5′-GATC-3′ and 5′-TCCT-3′; 37). TTF-1 has Ile at position 47; however, the presence of Tyr at position 54 determines a preference for G at the fourth position (36). For Zmhox2a it has been proposed that Lys47 and His54 are involved in binding at this position (37), although direct evidence has not been obtained and this does not explain the different binding specificities of the two HDs. Amino acid 47 (Asn) does not participate in DNA binding in MATα2; a C:G base pair at an equivalent position is recognized by Arg54 (9). So far, we have not found reports on changes in DNA binding specificity directly related to changes in amino acid 47 of the HD. The fact that >90% of HDs contain Ile/Val at position 47 (3) suggests that this is not a frequent source of diversity. These examples indicate that single amino acid substitutions at position 47 are not likely to produce new DNA binding specificities, unless they are complemented by changes in other parts of the molecule.
Since the binding site preferences of HAHR1 and Phe47-Hahb-4 are not the same, our results also indicate the existence of additional factors that influence specificity. The construction of chimeric proteins between Hahb-4 and HAHR1, and their respective mutants at residue 47, strongly suggests that amino acids C-terminal to residue 53 are involved. This result helped us to discern the role of Thr54 in DNA binding. Since a mutant of Hahb-4 with Phe47 and Thr54 showed the same binding preferences as HAHR1, we conclude that these two amino acids act in concert to determine DNA binding specificity at the fourth position of the 5′-TAAA-3′ half-sequence, perhaps interacting with different members of the A:T base pair. As mentioned above, residue 54 has been implicated in the interaction with an equivalent position in MATα2 and TTF-1 (9,36), among others, while residue 47 is thought to be important in most HDs (3). We have not found evidence in the literature, however, on a combined effect of residues at the two positions on DNA binding specificity.
An interesting feature of HAHR1 and of the other proteins with Phe at position 47 that we have constructed is that they are able to bind to both 5′-TAAT-3′ and 5′TAAA-3′ half-sequences. This fact can be explained by a model in which alternative amino acid–base pair contacts are established with different DNA target sequences (Fig. 8). The formation of alternative contacts by Arg54 has recently been observed for binding of the bicoid HD to 5′-TAAT-3′ and 5′-TAAG-3′ target sequences (38). Ile at position 47 (as in bicoid) would not allow this flexibility in Hd-Zip proteins, mainly due to steric hindrance with DNA and/or residue 54. In fact, we have observed that the presence of Thr (this study) or Met (as in Antennapedia) reduces the DNA binding affinity of Hd-Zip proteins with Ile47, without changes in DNA binding specificity (results not shown). The presence of Phe47 in GL2/Hd-Zip IV proteins would then be important not only to establish the observed binding specificity, but also to allow binding to different target sites. This fact may eventually be shown to be relevant for the function of this group of proteins within plant cells.
Figure 8.
Alternative modes of interaction of the HD with two different target sequences in plant dimeric HD proteins. The pseudopalindromic DNA-binding sequence 5′-CAATNATTG-3′ can be decomposed into two partially overlapping 5′-TNATTG-3′ sequences, similar to those bound by most animal HDs. The combinatorial interaction of residues 47 and 54 of one monomer of the plant proteins with position 4 of the core sequence 5′-TAA(T/A)-3′ is schematized, according to our observations. Proposed positive, negative and neutral interactions are indicated by +, – and 0, respectively. This model, which implies that different interactions are established with each of the target sequences, explains the selectivity and affinity of the wild-type and mutant proteins under study.
Acknowledgments
ACKNOWLEDGEMENTS
R.L.C. and D.H.G. are members of CONICET; A.E.T. and C.M.P. are fellows of the same Institution. C.W.B. is an undergraduate fellow of Universidad Nacional del Litoral. This work was supported by grants from CONICET, ANPCyT, Universidad Nacional del Litoral and Fundación Antorchas (Argentina).
REFERENCES
- 1.Gehring W.J. (1987) Homeo boxes in the study of development. Science, 236, 1245–1252. [DOI] [PubMed] [Google Scholar]
- 2.Levine M. and Hoey,T. (1988) Homeobox proteins as sequence-specific transcription factors. Cell, 55, 537–540. [DOI] [PubMed] [Google Scholar]
- 3.Gehring W.J., Affolter,M. and Bürglin,T. (1994) Homeodomain proteins. Annu. Rev. Biochem., 63, 487–526. [DOI] [PubMed] [Google Scholar]
- 4.Qian Y.Q., Billeter,M., Otting,G., Müller,M., Gehring,W.J. and Wüthrich,K. (1989) The structure of the Antennapedia homeodomain determined by MNR spectroscopy in solution: comparison with prokaryotic repressors. Cell, 59, 573–580. [DOI] [PubMed] [Google Scholar]
- 5.Qian Y.K., Furukubo-Tokunaga,K., Müller,M., Resendez-Perez,D., Gehring,W.J. and Wüthrich,K. (1994) Nuclear magnetic resonance solution structure of the fushi tarazu homeodomain from Drosophila and comparison with the Antennapedia homeodomain. J. Mol. Biol., 238, 333–345. [DOI] [PubMed] [Google Scholar]
- 6.Tsao D.H.H., Gruschus,J.M., Wang,L., Nirenberg,M. and Ferretti,J.A. (1995) The three dimensional solution structure of the NK-2 homeodomain from Drosophila. J. Mol. Biol., 251, 297–307. [DOI] [PubMed] [Google Scholar]
- 7.Kissinger C.R., Liu,B., Martin-Blanco,E., Kornberg,T.B. and Pabo,C.O. (1990) Crystal structure of an engrailed homeodomain-DNA complex at 2.8 Å resolution: a framework for understanding homeodomain-DNA interactions. Cell, 63, 579–590. [DOI] [PubMed] [Google Scholar]
- 8.Otting G., Qian,Y.Q., Billeter,M., Müller,M., Affolter,M., Gehring,W.J. and Wüthrich,K. (1990) Protein–DNA contacts in the structure of a homeodomain–DNA complex determined by nuclear magnetic resonance spectroscopy in solution. EMBO J., 9, 3085–3092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wolberger C., Vershon,A.K., Liu,B., Johnson,A.D. and Pabo,C.O. (1991) Crystal structure of a MATα2 homeodomain-operator complex suggests a general model for homeodomain-DNA interactions. Cell, 67, 517–528. [DOI] [PubMed] [Google Scholar]
- 10.Gehring W.J., Qian,Y.Q., Billeter,M., Furukubo-Tokunaga,K., Schier,A.F., Resendez-Perez,D., Affolter,M., Otting,G. and Wüthrich,K. (1994) Homeodomain-DNA recognition. Cell, 78, 211–223. [DOI] [PubMed] [Google Scholar]
- 11.Affolter M., Percival-Smith,A., Müller,M., Leupin,W. and Gehring,W.J. (1990) DNA binding properties of the purified Antennapedia homeodomain. Proc. Natl Acad. Sci. USA, 87, 4093–4097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Florence B., Handrow,R. and Laughon,A. (1991) DNA-binding specificity of the fushi tarazu homeodomain. Mol. Cell. Biol., 11, 3613–3623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chan R.L., Gago,G.M., Palena,C.M. and Gonzalez,D.H. (1998) Homeoboxes in plant development. Biochim. Biophys. Acta, 1442, 1–19. [DOI] [PubMed] [Google Scholar]
- 14.Ruberti I., Sessa,G., Lucchetti,S. and Morelli,G. (1991) A novel class of proteins containing a homeodomain with a closely linked leucine zipper motif. EMBO J., 10, 1787–1791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Schena M. and Davis,R.W. (1992) HD-Zip proteins: members of an Arabidopsis homeodomain protein superfamily. Proc. Natl Acad. Sci. USA, 89, 3894–3898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Sessa G., Morelli,G. and Ruberti,I. (1993) The Athb-1 and -2 HD-Zip domains homodimerize forming complexes of different DNA binding specificities. EMBO J., 12, 3507–3517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Sessa G., Carabelli,M., Ruberti,I., Lucchetti,S., Baima,S. and Morelli,G. (1994) Identification of distinct families of HD-Zip proteins in Arabidopsis thaliana. In Puigdomenech,P. and Coruzzi,G. (eds), Molecular-Genetic Analysis of Plant Development and Metabolism. Springer Verlag, Berlin, Germany, pp. 411–426.
- 18.Palena C.M., Gonzalez,D.H. and Chan,R.L. (1999) A monomer-dimer equilibrium modulates the interaction of the sunflower homeodomain leucine-zipper protein Hahb-4 with DNA. Biochem. J., 341, 81–87. [PMC free article] [PubMed] [Google Scholar]
- 19.Sessa G., Steindler,C., Morelli,G. and Ruberti,I. (1998) The Arabidopsis Athb-8, -9 and -14 genes are members of a small gene family coding for highly related HD-ZIP proteins. Plant Mol. Biol., 38, 609–622. [DOI] [PubMed] [Google Scholar]
- 20.Rerie W.G., Feldmann,K.A. and Marks,M.D. (1994) The GLABRA2 gene encodes a homeodomain protein required for normal trichome development in Arabidopsis. Genes Dev., 8, 1388–1399. [DOI] [PubMed] [Google Scholar]
- 21.Di Cristina M., Sessa,G., Dolan,L., Linstead,P., Baima,S., Ruberti,I. and Morelli,G. (1996) The Arabidopsis Athb-10 (GLABRA2) is an HD-Zip protein required for regulation of root hair development. Plant J., 10, 393–402. [DOI] [PubMed] [Google Scholar]
- 22.Lu P., Porat,R., Nadeau,J.A. and O’Neill,S.D. (1996) Identification of a meristem L1 layer-specific gene in Arabidopsis that is expressed during embryonic pattern formation and defines a new class of homeobox genes. Plant Cell, 8, 2155–2168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Masucci J.D., Rerie,W.G., Foreman,D.R., Zhang,M., Galway,M.E., Marks,M.D. and Schiefelbein,J.W. (1996) The homeobox gene GLABRA2 is required for position-dependent cell differentiation in the root epidermis of Arabidopsis thaliana. Development, 122, 1253–1260. [DOI] [PubMed] [Google Scholar]
- 24.Hung C.-Y., Lin,Y., Zhang,M., Pollock,S., Marks,M.D. and Schiefelbein,J. (1998) A common position-dependent mechanism controls cell-type patterning and GLABRA2 regulation in the root and hypocotyl epidermis of Arabidopsis. Plant Physiol., 117, 73–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ingram G.C., Magnard,J.-L., Vergne,P., Dumas,C. and Rogowsky,P.M. (1999) ZmOCL1, an HDGL2 family homeobox gene, is expressed in the outer cell layer throughout maize development. Plant Mol. Biol., 40, 343–354. [DOI] [PubMed] [Google Scholar]
- 26.Palena C.M., Chan,R.L. and Gonzalez,D.H. (1997) A novel type of dimerization motif, related to leucine zippers, is present in plant homeodomain proteins. Biochim. Biophys. Acta, 1352, 203–212. [DOI] [PubMed] [Google Scholar]
- 27.Smith D.B. and Johnson,K.S. (1988) Single-step purification of polypeptides expressed in Escherichia coli as fusions with glutathione S-transferase. Gene, 67, 31–40. [DOI] [PubMed] [Google Scholar]
- 28.Palena C.M., Gonzalez,D.H., Guelman,S. and Chan,R.L. (1998) Expression of sunflower homeodomain containing proteins in Escherichia coli: purification and functional studies. Protein Expr. Purif., 13, 97–103. [DOI] [PubMed] [Google Scholar]
- 29.Sedmak J. and Grossberg,S. (1977) A rapid, sensitive and versatile assay for protein using Coomassie brilliant blue G-250. Anal. Biochem., 79, 544–552. [DOI] [PubMed] [Google Scholar]
- 30.Valle E.M., Gonzalez,D.H., Gago,G. and Chan,R.L. (1997) Isolation and expression pattern of hahr1, a homeobox containing cDNA from Helianthus annuus. Gene, 196, 61–68. [DOI] [PubMed] [Google Scholar]
- 31.Palena C.M., Tron,A.E., Bertoncini,C.W., Gonzalez,D.H. and Chan,R.L. (2001) Positively charged residues at the N-terminal arm of the homeodomain are required for efficient DNA binding by homeodomain-leucine zipper proteins. J. Mol. Biol., 308, 39–47. [DOI] [PubMed] [Google Scholar]
- 32.Laughon A. (1991) DNA binding specificity of homeodomains. Biochemistry, 30, 11357–11367. [DOI] [PubMed] [Google Scholar]
- 33.Gonzalez D.H., Valle,E.M., Gago,G.M. and Chan,R.L. (1997) Interaction between proteins containing homeodomains associated to leucine zippers from sunflower. Biochim. Biophys. Acta, 1351, 137–149. [DOI] [PubMed] [Google Scholar]
- 34.Sessa G., Morelli,G. and Ruberti,I. (1997) DNA-binding specificity on the homeodomain-leucine zipper domain. J. Mol. Biol., 274, 303–309. [DOI] [PubMed] [Google Scholar]
- 35.Banerjee-Basu S., Ryan,J.F. and Baxevanis,A.D. (2000) The Homeodomain Resource: a prototype database for a large protein family. Nucleic Acids Res., 28, 329–330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Damante G., Pellizzari,L., Esposito,G., Fogolari,F., Viglino,P., Fabbro,D., Tell,G., Formisano,S. and Di Lauro,R. (1996) A molecular code dictates sequence-specific DNA recognition by homeodomains. EMBO J., 15, 4992–5000. [PMC free article] [PubMed] [Google Scholar]
- 37.Kirch T., Bitter,S., Kisters-Woike,B. and Werr,W. (1998) The two homeodomains of the Zmhox2a gene from maize originated as an internal gene duplication and have evolved different target site specificities. Nucleic Acids Res., 26, 4714–4720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dave V., Zhao,C., Yang,F., Tung,C.S. and Ma,J. (2000) Reprogrammable recognition codes in bicoid homeodomain-DNA interaction. Mol. Cell. Biol., 20, 7673–7684. [DOI] [PMC free article] [PubMed] [Google Scholar]