ABSTRACT
Viral infection activates the type I interferons (IFNs) and cellular antiviral responses. Eukaryotic initiation factor 4A-III (eIF4A3) has been shown to promote influenza A virus (IAV) replication by promoting viral mRNA splicing and spliced mRNA nuclear export. Here, we identified eIF4A3 as a negative regulator of virus-triggered type I IFN induction. Our study found that eIF4A3 promoted multiple RNA viruses’ replication by binding to IFN regulatory factor 3 (IRF3) and impaired the interaction between tank-binding kinase 1 (TBK1) and IRF3, leading to attenuation of the phosphorylation of IRF3 by TBK1, the formation of IRF3 dimer, and the nuclear translocation of IRF3. This impaired its biological functions in the nucleus, which blocked IRF3 binding to interferon-stimulated response element (ISRE) and the interaction of IRF3 and CBP/p300, resulting in inhibiting the transcription of IFN-β and downstream IFN-stimulated genes (ISGs), thereby impairing innate antiviral immune responses against RNA viruses. These findings reveal a previously unknown function of eIF4A3 in host innate immunity and establish a mechanistic link between eIF4A3 and IRF3 activation that expands potential therapeutic strategies for viral infectious diseases.
IMPORTANCE Production of type I IFN is pivotal for the cellular antiviral immunity. Virus infection leads to the activation of transcription factor IRF3 and subsequent production of type I IFN to eliminate viral infection. Thus, the regulation of IRF3 activity is an important way to affect type I IFN production. IRF3 activation requires phosphorylation, dimerization, and nuclear translocation. Here, we first reported that eIF4A3, a member of DEAD box family, served as a negative regulator of antiviral innate immune responses by inhibiting IRF3 activation. Mechanistically, eIF4A3 binds to IRF3 to impair the recruitment of IRF3 by TBK1, which is independent of eIF4A3 ATP binding, ATPase, and RNA helicase activities. Our study delineates a common mechanism of eIF4A3 promoting replication of different RNA viruses and provides important insights into the negative regulation of host antiviral innate immune responses against virus infections.
KEYWORDS: IFN-β, IRF3, eIF4A3, innate immune responses
INTRODUCTION
The innate immune system is the first line of host defense when the virus invades. There are many host germ line-encoded pattern recognition receptors (PRRs) in cells that identify unique pathogen-associated molecular patterns (PAMPs). The PRRs include the Toll-like receptors (TLRs), retinoic acid-inducible gene I (RIG-I)-like receptors (RLRs), NOD-like receptors (NLRs), and DNA receptors (1, 2). Viral nucleic acids are classical PAMPs that are recognized by these receptors and initiate innate immune signaling. Upon viral RNA recognition, the PRR retinoic acid-inducible gene I (RIG-I) directly interacts with the mitochondrial adaptor protein mitochondrial antiviral-signaling protein (MAVS) (also known as VISA, or IPS-1) through homophilic interactions between the CARDs and induces MAVS to form oligomers (3, 4). The oligomerized MAVS further recruits and activates tank-binding kinase 1/inhibitor-κB kinase ε (TBK1/IKKε) complex and inhibitor-κB kinase α/β (IKKα/β) complex to phosphorylate the interferon regulatory factor 3/7 (IRF3/IRF7) and nuclear factor-κB (NF-κB) (5, 6). Then phosphorylated IRF3 and NF-κB enter the nucleus to initiate transcription of downstream genes, including type I IFN and proinflammatory cytokines.
IRF3 serves as the key signal mediator and transcription factor for the type I IFN induction (7). Upon viral infection, the cytoplasmic IRF3 undergoes phosphorylation by inducing a conformational change in IRF3 that allows dimerization, nuclear translocation, and association with the coactivator CBP/p300, ultimately leading to the transcription of type I IFN and the downstream IFN-stimulated genes (ISGs) (8, 9). The phosphorylation of IRF3 is an important step in the induction of type I IFN. There are many negative regulators inhibiting the IRF3 phosphorylation. For instance, protein arginine methyltransferase 6 (PRMT6) inhibits IRF3 activation and type I IFN expression by disrupting the assembly of the TBK1-IRF3 signaling complex (10); AKT3 is an important phosphatase for IRF3, which directly phosphorylates IRF3 Ser385; 7-dehydrocholesterol reductase (DHCR7) strongly decreases IRF3 activation and type I IFN production by modulating AKT3 activation (11); protein phosphatase 2A (PP2A) can dephosphorylate IRF3, and PP2A interacts with RACK1 to restrict type I IFN expression by reducing the IRF3 phosphorylation (12). There are also other IRF3 posttranslational regulation methods to affect the IFN signaling pathway. For instance, the prolyl isomerase Pin1 suppresses IRF3 activity by destabilizing activated IRF3 (13); tripartite motif-containing protein 26 (TRIM26) negatively regulates type I IFN production by promoting IRF3 ubiquitination and proteasomal degradation (14); forkhead box protein O1 (FoxO1) negatively regulates cellular antiviral responses by interacting with IRF3 and promoting K48-linked polyubiquitination (15). For all that, whether and how other molecules are involved in regulating posttranscriptional modifications of IRF3 to affect type I IFN signaling pathway is of great interest.
eIF4A3, the eukaryotic translation initiation factor, is the member of the DEAD box family with the ability of RNA-dependent ATPase and RNA helicase (16). eIF4A3 is a nucleocytoplasmic shuttling protein and is also the core protein of the splicing-dependent multiprotein exon junction complex (EJC), which is involved in splicing, nuclear mRNA export, translation, and nonsense-mediated mRNA decay (NMD) (17, 18). As an important host factor, it has been reported that eIF4A3 promotes avian influenza A virus (AIV) replication by increasing viral mRNA splicing and nuclear export (19). However, the impact of eIF4A3 on other RNA viruses is unknown, and whether and how eIF4A3 involves in antiviral innate immunity remains unclear. In this study, we found that eIF4A3 promoted influenza A virus (IAV), Sendai virus (SeV), and vesicular stomatitis virus (VSV) replication and identified eIF4A3 as a critical negative regulator of virus-triggered type I IFN signaling. eIF4A3 and TBK1 competed for binding to the same region of IRF3 to block TBK1-IRF3 interaction, leading to attenuation of the recruitment of IRF3 to TBK1 and subsequently inhibiting virus-triggered phosphorylation and nuclear translocation of IRF3. This inhibited the IRF3 function in the nucleus, which impaired IRF3 binding to interferon-stimulated response element (ISRE) and the interaction of IRF3 and CBP/p300. We also found that the inhibition of eIF4A3 on antiviral innate immune responses was independent of its ATP binding, ATPase, and RNA helicase activities. Taken together, our studies uncover a novel molecular mechanism of eIF4A3 as a negative regulator of antiviral innate immunity.
RESULTS
eIF4A3 promotes IAV, SeV, and VSV replication.
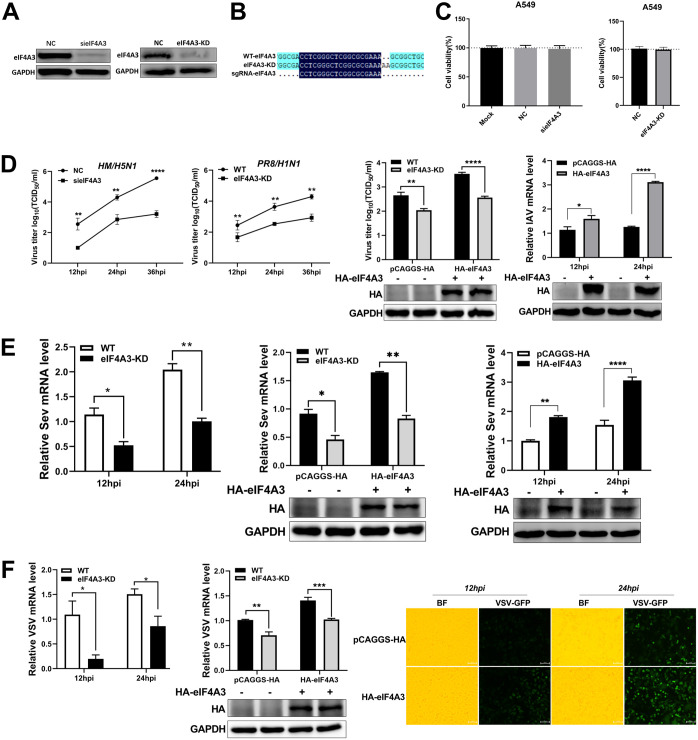
A previous study has reported that eIF4A3 promotes the replication of H9N2 influenza virus (19), but its effects on other RNA viruses remain unknown. To explore the roles of eIF4A3 in other RNA viruses, the eIF4A3 gene was silenced using small interfering RNA (siRNA) or knocked down using the CRISPR/Cas9 technique in A549 cells. The results showed that the expression level of endogenous eIF4A3 protein was decreased in both eIF4A3 silencing cells (sieIF4A3) cells and eIF4A3 knockdown (eIF4A3-KD) cells, and a mutation occurred near the protospacer-adjacent motif (PAM) region in eIF4A3 knockdown cells (Fig. 1A and B). Meanwhile, sieIF4A3 or eIF4A3 knockdown also showed no cytotoxic effect on A549 cells (Fig. 1C). We next determined the impact of eIF4A3 on replication of IAVs, including a highly pathogenic avian A/duck/Hubei/Hangmei01/2006 (HM H5N1) and a PR8 H1N1 influenza virus, which replicate their genomes in the nucleus. Results showed that the viral titer of HM was significantly decreased in eIF4A3 silencing cells, and the replication of PR8 virus was profoundly inhibited in eIF4A3 knockdown cells at all tested time points, including 12, 24, and 36 h postinfection (hpi). Similarly, the complement of eIF4A3 expression restored the replication of PR8 virus in eIF4A3 knockdown cells at 24 hpi, and the proliferation of PR8 virus significantly increased at 12 and 24 hpi in wild-type (WT) cells complemented with eIF4A3 expression (Fig. 1D). In contrast to IAVs, both vesicular stomatitis virus (VSV) and Sendai virus (SeV) replicate their genomes in the cytoplasm. We explored the effects of eIF4A3 on the replication of cytoplasmic RNA viruses using an SeV and a recombinant VSV-green fluorescent protein (GFP) as examples. Results showed that significantly lower levels of viral mRNAs of SeV or VSV-GFP were detected at 12 and 24 hpi in sieIF4A3 cells or eIF4A3 knockdown cells. Moreover, the replication of VSV and SeV restored at 24 hpi in eIF4A3 knockout cells complemented eIF4A3 expression, and the replication of VSV and SeV significantly increased at 12 and 24 hpi in WT cells complemented with eIF4A3 expression (Fig. 1E and F). Together, these data demonstrate that eIF4A3 is a positive regulator of multiple RNA viruses, suggesting that eIF4A3 might play a role in antiviral innate immunity.
FIG 1.
The positive regulation of eIF4A3 on IAV, SeV, and VSV replication. (A to D) eIF4A3 promotes the replication of IAV. (A) The silencing efficiency of eIF4A3 or CRISPR-mediated knockdown efficiency of eIF4A3 was measured by Western blot assay. (B) The Sanger sequencing result indicated that a frameshift occurred in the eIF4A3 knockdown cells (eIF4A3-KD). (C) The cell viability of sieIF4A3 cells or eIF4A3 knockdown cells was measured by CCK-8 assay (mean ± SD of three independent experiments; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test). (D) The effect of eIF4A3 on HM and PR8 replication. A549 cells were transfected with sieIF4A3 or NC for 24 h, and were infected with HM virus (MOI, 0.01). The cell supernatants were harvested at 12 hpi, 24 hpi, and 36 hpi. Virus titers were determined by TCID50 assay on MDCK cells. eIF4A3 knockdown A549 cells or WT A549 cells were infected with PR8 (MOI, 0.1). Cell supernatants were collected at 12 hpi, 24 hpi, and 36 hpi, and virus titers were determined by TCID50 assay on MDCK cells. eIF4A3 knockdown A549 cells and WT A549 cells were transfected with HA-eIF4A3 or vector for 24 h and were infected with PR8 virus (MOI, 0.1). Cell supernatants were collected at 24 hpi, and virus titers were determined by TCID50 assay on MDCK cells. A549 cells were transfected with HA-eIF4A3 or vector for 24 h and were infected with PR8 at an MOI of 0.1. Samples were collected at 12 hpi and 24 hpi. The mRNA level of NP was detected by qRT-PCR. The viral RNAs levels were normalized to the GAPDH level (mean ± SD of three independent experiments *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test). (E) The effect of eIF4A3 on SeV replication. eIF4A3 knockdown A549 cells or WT A549 cells were infected with SeV virus at an MOI of 1.0. Samples were collected at 12 hpi and 24 hpi. The mRNA level of SeV-N was detected by qRT-PCR, and the viral RNAs levels were normalized to the GAPDH level. eIF4A3 knockdown A549 cells and WT A549 cells were transfected with HA-eIF4A3 or vector for 24 h and were infected with SeV virus at an MOI of 1.0. Samples were collected at 24 hpi. The mRNA level of SeV-N was detected by qRT-PCR, and the viral RNAs levels were normalized to the GAPDH level. A549 cells were transfected with HA-eIF4A3 or vector for 24 h and were infected with SeV virus at an MOI of 1.0. Samples were collected at 12 hpi and 24 hpi. The mRNA level of SeV-N was detected by qRT-PCR. The viral RNA levels were normalized to the GAPDH level (mean ± SD of three independent experiments; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test). (F) The effect of eIF4A3 on VSV-GFP replication. eIF4A3 knockdown A549 cells or WT A549 cells were infected with VSV-GFP (MOI, 1.0). Samples were collected at 12 hpi and 24 hpi. The VSV-GFP viral RNA level was determined by qRT-PCR. The viral RNA levels were normalized to the GAPDH level. eIF4A3 knockdown A549 cells or WT A549 cells were transfected with HA-eIF4A3 or vector for 24 h and infected with VSV-GFP (MOI, 1.0). Samples were collected at 24 hpi. The VSV-GFP viral RNA level was determined by qRT-PCR. The viral RNA levels were normalized to the GAPDH level. A549 cells were transfected with HA-eIF4A3 or vector for 24 h and infected with VSV-GFP (MOI, 1.0). The VSV-GFP was visualized by fluorescence microscopy at 12 hpi and 24 hpi (×100 magnification) (mean ± SD of three independent experiments; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test).
eIF4A3 inhibits the antiviral innate immune responses.
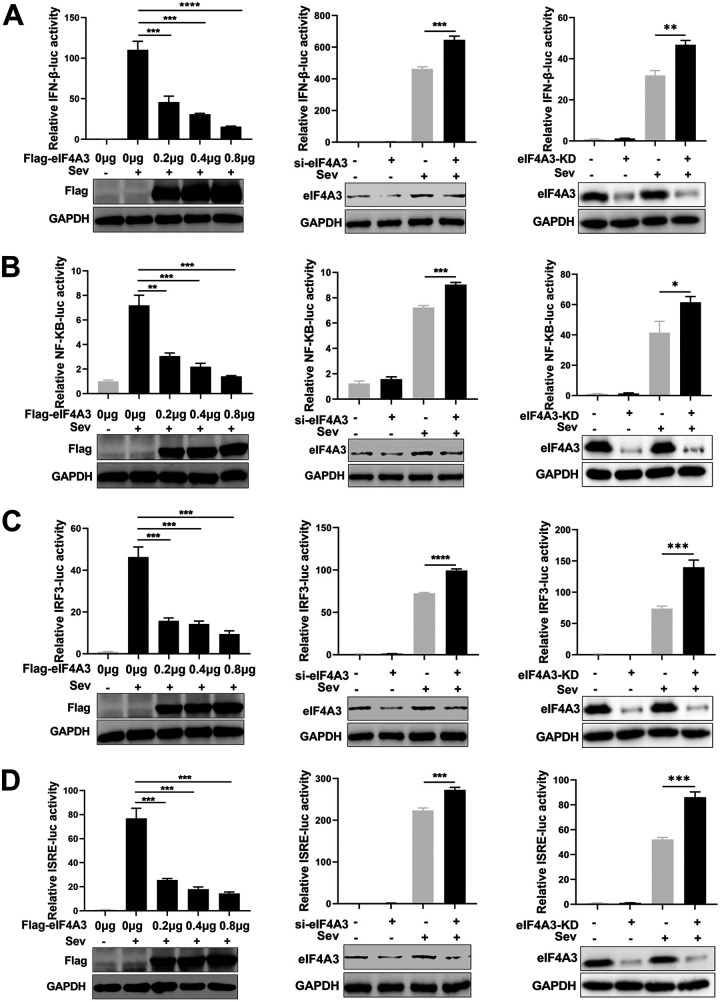
To explore the roles of eIF4A3 in the antiviral innate immune responses, we first examined the effect of eIF4A3 on SeV-induced activation of IFN-β promoter luciferase activity. HEK293T cells were cotransfected with IFN-β-luc, pRL-TK, and sieIF4A3 or Flag-eIF4A3, and the cells were stimulated with SeV for another 10 h after 24 h posttransfection. Results of the dual-luciferase reporter assay showed that overexpression of eIF4A3 reduced the SeV-trigged IFN-β promoter activity in a dose-dependent manner, while sieIF4A3 significantly increased SeV-induced IFN-β promoter activity. The effect of eIF4A3 on the SeV-induced activation of IFN-β promoter luciferase activity was also examined in eIF4A3 knockdown cells and WT cells. Similarly, SeV-induced IFN-β promoter activity could be profoundly enhanced in eIF4A3 knockdown cells (Fig. 2A), indicating that eIF4A3 was an inhibitor of SeV-induced activation of IFN-β promoter. It is well known that IFN-β expression is mainly regulated by NF-κB and IRF3, so we further determined the impacts of eIF4A3 on SeV-triggered activation of NF-κB and IRF3 promoter activity in HEK293T cells and eIF4A3 knockdown cells. Results showed that NF-κB and IRF3 promoter activity gradually decreased along with increasing amounts of eIF4A3 but significantly increased in sieIF4A3 cells and eIF4A3 knockdown cells (Fig. 2B and C), indicating that eIF4A3 was involved in both NF-κB- and IRF3-regulated SeV-triggered type I IFN expression. The effect of eIF4A3 on the SeV-induced activation of interferon-stimulated response element (ISRE) promoter was also examined by the luciferase activity assay. Results showed that overexpressed eIF4A3 obviously reduced the ISRE promoter activity in a dose-dependent manner, while silencing of eIF4A3 or knockdown of eIF4A3 significantly enhanced the ISRE promoter activity (Fig. 2D). These results indicate that eIF4A3 suppresses the SeV-triggered antiviral innate immune responses.
FIG 2.
(A to D) The negative regulation of eIF4A3 on IFN-β, NF-κB, IRF3, and ISRE promoter. HEK293T cells were transfected with increasing amounts of Flag-eIF4A3 or sieIF4A3 and the IFN-β/NF-κB/IRF3/ISRE luciferase reporter. Then, 24 h after transfection, the cells were stimulated by SeV for 10 h and measured by the luciferase assays. eIF4A3 knockdown A549 cells or WT A549 cells were transfected with the IFN-β/NF-κB/IRF3/ISRE luciferase reporter. The cells were stimulated with SeV for another 10 h after 24 h posttransfection and measured by the luciferase assays. For all experiments described above, the data are presented as the mean ± SD of three independent experiments (*, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test).
eIF4A3 suppresses the transcription of IFN-β, ISGs, and cytokines.
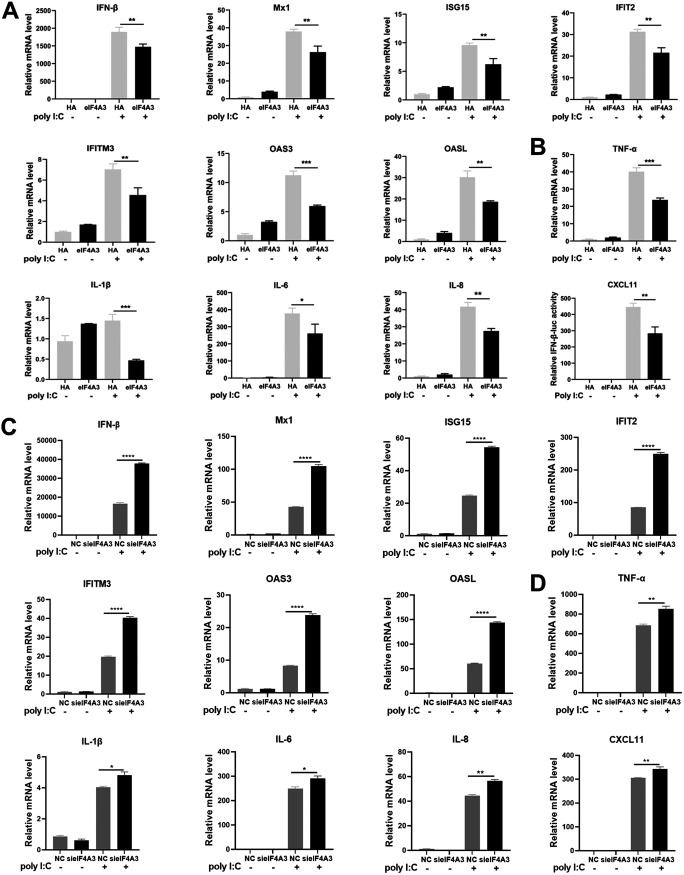
To further investigate the roles of eIF4A3 in the innate immune responses, we detected the effect of eIF4A3 on the expression of IFN-β, ISGs, and proinflammatory cytokines. A549 cells were transfected with HA-eIF4A3 and then were stimulated with poly(I·C) for another 6 h after 20 h posttransfection. A reverse transcription-quantitative PCR (qRT-PCR) assay was performed to detect mRNA levels of each indicated gene. Results showed that eIF4A3 overexpression significantly decreased the mRNA level of IFN-β, the ISGs, including Mx1, ISG15, IFIT2, IFITM3, OAS3, and OASL, and the proinflammatory cytokines, including tumor necrosis factor alpha (TNF-α), interleukin-1β (IL-1β), IL-6, IL-8, and CXCL11 (Fig. 3A and B). To better understand the roles of eIF4A3 in regulating IFN-β transcription, the negative-control A549 cells (NC) and eIF4A3 silencing cells (sieIF4A3) were also induced by poly(I·C) for 6 h and harvested for qRT-PCR assay. Results illustrated that sieIF4A3 promoted the mRNA level of IFN-β, the ISGs (Mx1, ISG15, IFIT2, IFITM3, OAS3, and OASL), and the proinflammatory cytokines (TNF-α, IL-1β, IL-6, IL-8, and CXCL11) (Fig. 3C and D). Collectively, these results indicate that eIF4A3 is likely to inhibit the virus-triggered innate immune signaling pathways.
FIG 3.
The inhibition of eIF4A3 on the transcription of IFN-β and ISGs. (A and B) Overexpression of eIF4A3 inhibited the mRNA of IFN-β, ISGs, and cytokines. A549 cells were transfected with HA-eIF4A3 or control vector (HA), followed by stimulation with or no stimulation with poly(I·C) (100 ng) for 6 h. The mRNA levels of the indicated target genes were measured by qRT-PCR. (C and D) sieIF4A3 promoted the mRNA of IFN-β, ISGs, and cytokines. A549 cells were treated with negative-control RNAs (NC) and siRNAs targeting eIF4A3 (sieIF4A3) for 24 h, and then cells were both induced or uninduced by poly(I·C) (100 ng) for 6 h. The mRNA levels of the indicated target genes were detected by qRT-PCR. All the mRNA levels were normalized to the GAPDH level (mean ± SD of three independent experiments; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test).
eIF4A3 negatively modulates RIG-I-inducing pathway.
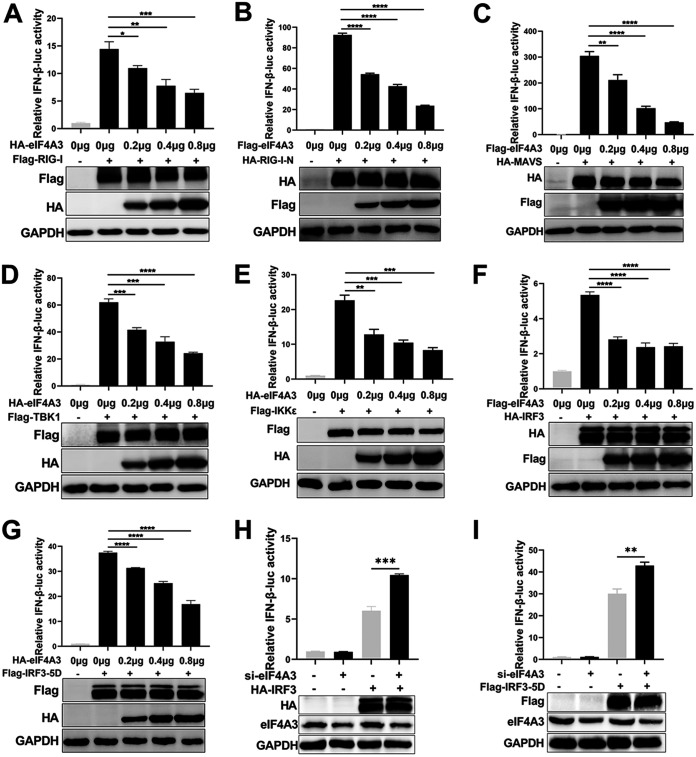
Viral RNAs are classical pathogen-associated molecular patterns that are recognized by TLRs and RIG-I-like receptors (20). Influenza virus can be recognized by RIG-I during infection and activates type I IFN production by signaling transduction from RIG-I, MAVS, and TBK1/IKKε complex to IRF3. To investigate how eIF4A3 regulates IFN-β, we analyzed the impact of eIF4A3 on the RIG-I-inducing signal pathway. HEK293T cells were transfected with IFN-β-luc, pRL-TK, and signaling components along with an increasing amount of eIF4A3 expressing plasmid, and the luciferase activities were examined at 24 h posttransfection. When stimulated with RIG-I, IFN-β production was gradually reduced along with an increasing amount of HA-eIF4A3 (Fig. 4A). The N terminus of RIG-I (RIG-I-N) consists of two N-terminal CARDs responsible for triggering the interaction with its downstream MAVS. Results showed that the activation of the IFN-β promoter induced by HA-RIG-I-N was obviously suppressed by eIF4A3 in a dose-dependent manner (Fig. 4B). Results of a dual-luciferase reporter assay also showed that eIF4A3 dramatically decreased the MAVS-, TBK1-, IKKε-, IRF3-, and IRF3-5D (an active phosphomimetic mutant of IRF3)- mediated activation of the IFN-β in a dose-dependent manner (Fig. 4C to G). In contrast to eIF4A3 overexpression, sieIF4A3 enhanced IRF3- or IRF3-5D-stimulated IFN-β promoter activation (Fig. 4H and I). These data suggest that eIF4A3 may reduce type I IFN signaling by affecting IRF3.
FIG 4.
The reduction of eIF4A3 on the RIG-I inducing pathway. (A to G) The effect of eIF4A3 on the activation of IFN-β promoter induced by RIG-I (A), RIG-I-N (B), MAVS (C), TBK1 (D), IKKε (E), IRF3 (F), and IRF3-5D (G). HEK293T cells were transfected with the IFN-β-luc (0.3 μg) and pRL-TK (0.02 μg), plus the indicated signal molecules (0.3 μg), along with an increasing amount of HA-eIF4A3 (0, 0.2, 0.4, and 0.8 μg). Then, 24 h after transfection, IFN-β promoter luciferase activities were detected by dual-luciferase reporter assay. (H and I) The effect of eIF4A3 on the activation of IFN-β promoter induced by IRF3 (H) and IRF3-5D (I). HEK293T cells were transfected with sieIF4A3 for 6 h and then transfected with IFN-β-luc (0.3 μg), pRL-TK (0.02 μg), and IRF3 or IRF3-5D (1 μg). Then, 24 h after transfection, IFN-β promoter luciferase activities were detected by dual-luciferase reporter assay. For all experiments described above, the data are presented as the mean ± SD of three independent experiments (*, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test).
eIF4A3 inhibits IRF3 phosphorylation and nuclear entry.
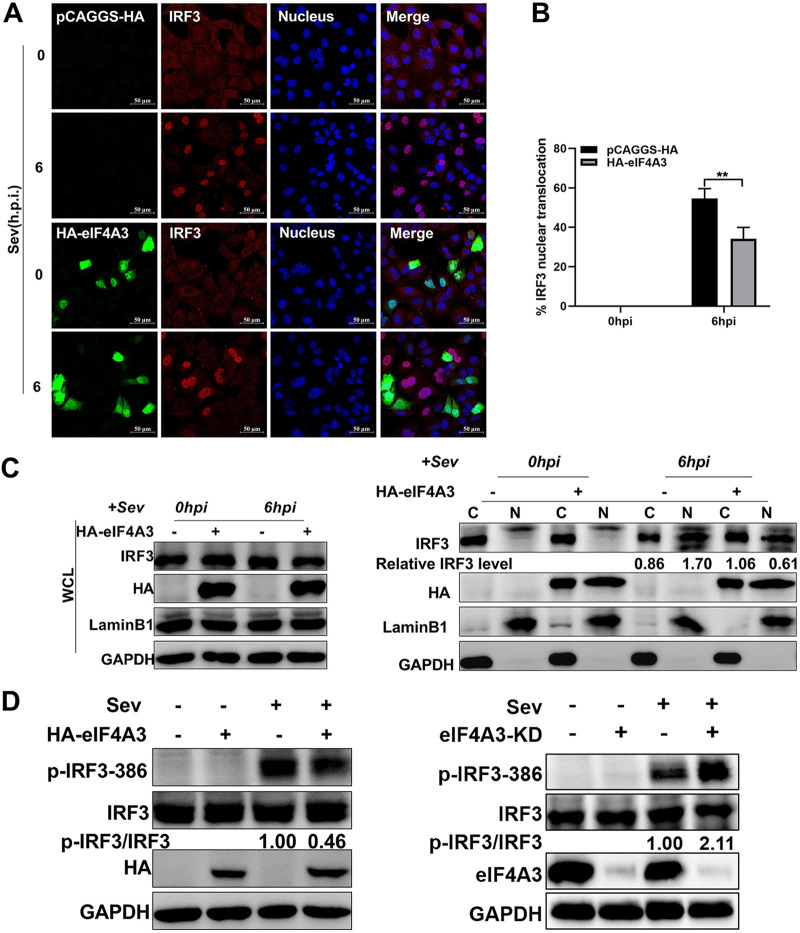
Phosphorylation, dimerization, and nuclear entry of IRF3 are hallmarks for IRF3 activation after viral infection, and activation of IRF3 drives type I IFN production. To determine whether eIF4A3 affected the IRF3 activation, A549 cells transfected with HA-eIF4A3 were stimulated with SeV for 6 h, followed by an indirect immunofluorescence assay. Results of immunofluorescence confocal microscopy showed that overexpression of eIF4A3 substantially impaired nuclear accumulation of IRF3 after SeV infection (Fig. 5A), and approximately 10% of nuclear translocation of IRF3 in eIF4A3-overexpressed cells was decreased compared to control cells (Fig. 5B). To further authenticate the microscopic observation, SeV-infected A549 cells were subjected to cellular fractionation. The nuclear localization of IRF3 was increased upon SeV stimulation, and this nuclear localization was obviously reduced when eIF4A3 was overexpressed, while the expression of IRF3 in the whole-cell fraction was not affected by eIF4A3 in SeV-uninfected cells (Fig. 5C). This result is consistent with that observed in confocal microscopy. The IRF3 phosphorylation levels were also examined by Western blot analysis in eIF4A3-overexpressed cells and eIF4A3 knockdown cells. We observed that the phosphorylation on endogenous IRF3 at Ser386 residue in response to SeV was weaker in eIF4A3-overexpressed cells while stronger in eIF4A3 knockdown cells (Fig. 5D). Together, above-described results demonstrate that eIF4A3 downregulates IFN-β induction by inhibiting the activation of IRF3.
FIG 5.
The inhibition of eIF4A3 on IRF3 activation. (A to C) Effect of eIF4A3 on nuclear translocation of IRF3. A549 cells were transfected with HA-eIF4A3 or vector (HA) and simulated with SeV for 6 h after 24 h of transfection. (A) Confocal microscopy was performed using an anti-IRF3 rabbit antibody (red) and anti-HA mouse antibody (green). DAPI was used to stain for the nucleus (blue). Samples were examined with a confocal microscope (LSM 880; Zeiss). Images are representative of three independent experiments. Scale bar = 50 μM. (B) Quantitative analysis of IRF3 translocation. At least 100 cells in each group were scored (*, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test). (C) Western blot analysis of the distribution of IRF3 in the cytoplasmic and nuclear factions. The indicated cells were subjected to isolation of cytoplasmic and nuclear fractions. Lamin B1 and GAPDH were used as the loading control for nuclear and cytoplasmic fractions, respectively. The band intensities were quantified, and relative cytoplasmic IRF3 levels (IRF3/GAPDH) and relative nuclear IRF3 levels (IRF3/lamin B1) are shown below. (D) Effect of eIF4A3 on the phosphorylation of IRF3 induced by SeV. A549 cells were transfected with HA-eIF4A3 or vector (HA), and simulated with SeV for 6 h after 24 h of transfection or eIF4A3 knockdown A549 cells and WT A549 cells infected with SeV for 6 h. The cell lysates were harvested for the Western blot analysis using anti-IRF3 antibody and anti-Ser386 phosphorylated IRF3 antibody. The band intensities were quantified, and relative IRF3 levels (p-IRF3-386/IRF3) are shown below.
eIF4A3 interacts with IRF3.
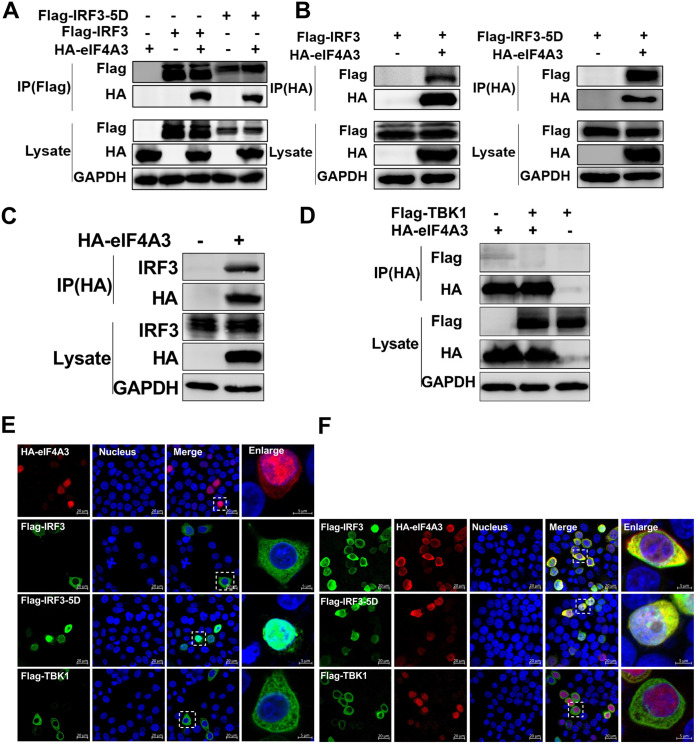
To figure out the relationship between IRF3 and eIF4A3, we investigated the interaction between eIF4A3 and IRF3 by coimmunoprecipitation (co-IP) assay. When cotransfected into HEK293T cells, HA-eIF4A3 coimmunoprecipitated with both Flag-IRF3 and Flag-IRF3-5D (the constitutively active form of IRF3) despite of an anti-Flag or anti-hemagglutinin (HA) antibody used as a bait (Fig. 6A and B). To confirm eIF4A3 targeting IRF3, we next examined the endogenous association between eIF4A3 and IRF3, and immunoblot analysis revealed that eIF4A3 interacted with IRF3 in HEK293T cells (Fig. 6C), suggesting a real interaction between these two proteins. However, HA-eIF4A3 did not coimmunoprecipitate with Flag-TBK1 when overexpressed in HEK293 cells (Fig. 6D). Additionally, the colocalization of eIF4A3 and IRF3, IRF3-5D, or TBK1 was determined using immunofluorescence confocal microscopy. In HeLa cells, eIF4A3 and IRF3-5D predominantly localized to the nucleus, while TBK1 and IRF3 localized to the cytoplasm when expressed alone (Fig. 6E). When eIF4A3 was coexpressed with IRF3, IRF-5D, or TBK1 in HeLa cells, it visibly colocalized with IRF3 and IRF3-5D in the cytoplasm but did not colocalize with TBK1 (Fig. 6F). Overall, these results demonstrate that eIF4A3 targets and physically interacts with IRF3.
FIG 6.
The interaction between eIF4A3 and IRF3. (A and B) Immunoblot analysis of the interactions between IRF3, IRF3-5D, and eIF4A3. HEK293T cells were transfected with HA-eIF4A3, Flag-IRF3, and Flag-IRF3-5D, followed by lysing at 24 h posttransfection. A co-IP assay was carried out using anti-HA immunomagnetic beads or anti-Flag immunomagnetic beads, followed by a Western blot assay. (C) The endogenous interaction between eIF4A3 and IRF3. HEK293T cells were transfected with HA-eIF4A3, followed by lysing at 24 h posttransfection. Immunoblot analysis was performed by using anti-HA immunomagnetic beads. (D) The interaction between eIF4A3 and TBK1. HEK293T cells were transfected with HA-eIF4A3 and Flag-TBK1, followed by lysing at 24 h posttransfection. Immunoblot analysis was performed by using anti-HA immunomagnetic beads. (E) Subcellular location of HA-eIF4A3, Flag-IRF3, Flag-IRF3-5D, or Flag-TBK1. HeLa cells were grown on coverslips and transfected with HA-eIF4A3, Flag-IRF3, Flag-IRF3-5D, or Flag-TBK1. Cells were fixed at 24 h posttransfection and stained for detecting eIF4A3 (red) and signal molecules (green) using the anti-HA rabbit antibodies and anti-Flag mouse antibodies, followed by the Alexa Fluor 594-conjugated AffiniPure goat anti-rabbit secondary antibodies and Alexa Fluor 488-conjugated AffiniPure goat anti-mouse secondary antibodies. DAPI was used to stain the nucleus (blue). Samples were examined with a confocal microscope (LSM 880; Zeiss). Images are representative of three independent experiments. Scale bar = 20 μM. (F) Colocalization of Flag-IRF3, Flag-IRF3-5D, or Flag-TBK1 and HA-eIF4A3. HeLa cells were grown on coverslips and transfected with HA-eIF4A3, Flag-IRF3, Flag-IRF3-5D, and Flag-TBK1. The remaining steps are the same as in panel E.
eIF4A3 inhibits the recruitment of TBK1 to IRF3.
TBK1 is known to directly phosphorylate IRF3 at Ser386 and drive type I IFN production (21). Given that eIF4A3 binds to IRF3 and inhibits the phosphorylation of IRF3, eIF4A3 might inhibit the recruitment of TBK1 to IRF3. To verify our hypothesis, HEK293T cells were transfected with Flag-TBK1 and HA-IRF3, together with increasing amounts of Myc-eIF4A3. Results of the immunoprecipitation assay revealed that TBK1 interacted with IRF3, and the binding of the TBK1-IRF3 module was decreased by eIF4A3 in a dose-dependent manner, whereas sieIF4A3 enhanced TBK1-IRF3 interaction (Fig. 7A). Then, we performed the Western blot assay to determine how eIF4A3 regulates the TBK1 to phosphorylate IRF3. Results showed that coexpression of eIF4A3 and TBK1 significantly reduced phosphorylation of IRF3 at Ser386 in HEK293T cells (Fig. 7B). To further investigate how eIF4A3 blocks TBK1-IRF3 interaction, a truncated IRF3 was constructed to bind TBK1. The results revealed that eIF4A3 and TBK1 competed for binding to the same region of IRF3 (Fig. 7C). The effect of eIF4A3 on the IRF3 dimer formation caused by TBK1 was further determined by a native PAGE assay, in which Flag-TBK1 and HA-IRF3 were cotransfected with increasing amounts of Myc-eIF4A3 in HEK293T cells. The results revealed that TBK1 alone could induce IRF3 dimer formation. Additionally, we noticed that the IRF3 dimerization was reduced when eIF4A3 was overexpressed (Fig. 7D). These results together demonstrate that eIF4A3 attenuates the ability of TBK1 to recruit IRF3 by interacting with IRF3.
FIG 7.
The inhibition of eIF4A3 on TBK1-IRF3 signaling. (A) The effect of eIF4A3 on the interaction between IRF3 and TBK1. Coimmunoprecipitation and immunoblot analysis of HEK293T cells transfected with Flag-TBK1, HA-IRF3, and Myc-eIF4A3 or sieIF4A3 HEK293T cells. The band intensities were quantified, and relative precipitated HA-IRF3/Flag-TBK1 or IRF3/TBK1 ratios are shown below. (B) Effect of eIF4A3 on the phosphorylation of IRF3 induced by TBK1. HEK293T cells were transfected with Flag-TBK1, HA-IRF3, and Myc-eIF4A3 to detect p-IRF3 at Ser 386 by Western blot analysis. The band intensities were quantified, and relative IRF3 levels (p-IRF3/IRF3) are shown below. (C) Identification of a binding site for the IRF3-eIF4A3 interaction. Coimmunoprecipitation and immunoblot analysis of HEK293T cells transfected with HA-eIF4A3, Flag-EGFP, Flag-EGFP-ΔIRF3 and Flag-IRF3 or HA-TBK1, Flag-EGFP, Flag-EGFP-ΔIRF3, and Flag-IRF3. (D) Effect of eIF4A3 on the dimer formation of IRF3 induced by TBK1. HEK293T cells were transfected with Flag-TBK1, HA-IRF3, and Myc-eIF4A3 to detect IRF3 dimer by native PAGE analysis; * represents the indicated protein.
eIF4A3 impairs the DNA binding ability of IRF3 and its association with CBP.
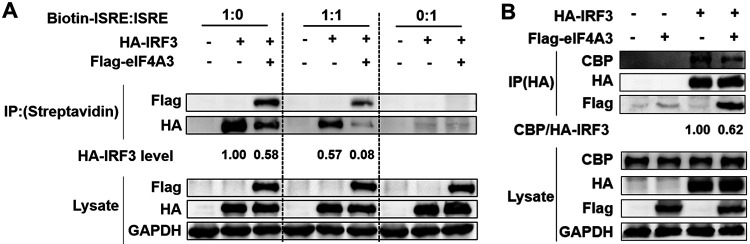
Given that eIF4A3 also impaired IRF3(5D)-induced ISRE activity as depicted in Fig. 2D, eIF4A3 might alter binding efficiency between IRF3 and ISRE, or the formation of the IRF3-CBP/p300 complex. To test this hypothesis, a DNA-binding assay was performed to investigate the effect of eIF4A3 on IRF3 binding. HEK293T cells were cotransfected with HA-IRF3 and Flag-eIF4A3 or vector (Flag) and then infected with SeV at 24 h posttransfection for 12 h. A DNA pulldown assay was carried out to detect bound IRF3 in ISRE oligonucleotide by using a biotinylated ISRE or unbiotinylated ISRE element as the probe. The results showed that the amounts of IRF3 pulled down by the biotin-ISRE were obviously decreased in the presence of eIF4A3, indicating that eIF4A3 reduced IRF3 DNA binding ability (Fig. 8A). As IRF3 forms a complex with CBP/p300 to allow promoter binding to activate the IFN-β gene, we next investigated the impacts of eIFA43 on the interaction between IRF3 and CBP/p300. The results of Western blot analysis showed that the endogenous CBP coprecipitated by IRF3 was decreased after coexpression of eIF4A3 and IRF3, indicating that eIF4A3 inhibited the formation of the CBP/p300-IRF3 complex (Fig. 8B). All results indicate that eIF4A3 impairs IRF3-induced ISRE activity through attenuating the binding between IRF3 and ISRE and the interaction between IRF3 and CBP/p300.
FIG 8.
eIF4A3 impedes IRF3 binding onto ISRE and the interaction of IRF3 and CBP/p300. (A) Effect of eIF4A3 on IRF3 binding to ISRE. HEK293T cells were transfected with HA-IRF3 in the absence or presence of Flag-eIF4A3 and infected with SeV at 24 h posttransfection. At 12 hpi, cells were lysed, the biotinylated or unbiotinylated DNA probes were incubated with streptavidin beads for 2 h at 4°C, and then the probe-coated beads were incubated with cell lysates at 4°C overnight. The beads were washed five times with cold IP lysis buffer, and Western blot analysis was done to detect IRF3 and eIF4A3 by using an anti-HA and anti-Flag antibody. The band intensities were quantified, and the HA-IRF3 levels are shown below. (B) The interaction between IRF3 and CBP/p300 in the absence or presence of Flag-eIF4A3. HEK293T cells were transfected with the HA-IRF3 and Flag- eIF4A3 or its control expression vector (Flag). After 24 h, cells were infected with SeV for 12 h. A co-IP assay was carried out using anti-HA antibody to immunoprecipitate endogenous CBP. The band intensities were quantified, and relative precipitated CBP/HA-IRF3 ratios are shown.
eIF4A3 suppresses the IFN-β independently of ATP binding, RNA-dependent ATPase, and RNA helicase activities.
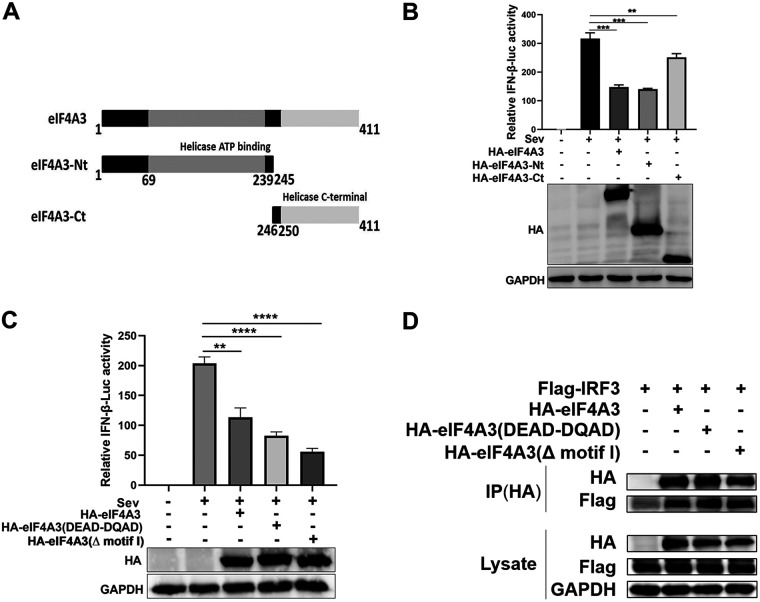
ATP binding, ATPase, and RNA helicase activity are important for many physiological functions of the DEAD box family members. Studies have reported Q motif (38 to 66 amino acids [aa]), motif I (80 to 89 aa), and DEAD box (187to 190 aa) located in the N-terminus of eIF4A3, which are important for ATP binding, RNA-dependent ATPase, and RNA helicase activities (22, 23). Thus, we hypothesized that the functions of eIF4A3 in antiviral immunity might depend on its ATP binding, ATPase, and RNA helicase activities. To test our hypothesis, we generated two truncated eIF4A3, eIF4A3-Nt (1 to 245 aa, N-terminal) and eIF4A3-Ct (246 to 411 aa, C-terminal) as depicted in Fig. 9A, and determined their effects on the type I IFN expression by a luciferase reporter assay. Results showed that the wild type and two truncated eIF4A3s were able to significantly inhibit luciferase reporter expression compared to the control, but eIF4A3-Ct was less efficient than the wild type and eIF4A3-Nt (Fig. 9B), suggesting that eIF4A3-Nt played a dominant role in the inhibition of luciferase reporter expression. Several studies have reported that mutation in the motif I (AxTGoGKT) affect the ATP binding activity of eIF4A3, while mutation of DEAD to DQAD eliminates eIF4A3’s RNA-stimulated ATPase and RNA helicase activities (24, 25). Therefore, we constructed a mutant with deletion of motif I (Δ motif I) of eIF4A3 and a mutant of eIF4A3 (DEAD-DQAD) and then performed a luciferase reporter assay induced by SeV that activates IFN transcription. The results showed that overexpression of eIF4A3, a mutant of eIF4A3 (Δ motif I) or a mutant of eIF4A3 (DEAD-DQAD) inhibited luciferase reporter expression induced by SeV (Fig. 9C). Meanwhile, we found that the interaction between eIF4A3 and IRF3 was not changed when transfected with the plasmid expressing eIF4A3, mutant of eIF4Aa3 (Δ motif I), or mutant of eIF4A3 (DEAD-DQAD) (Fig. 9D). These data indicate that the inhibitory effect of eIF4A3 on type I IFN expression does not depend on its ATP binding, RNA-stimulated ATPase, or RNA helicase activities.
FIG 9.
The inhibition of eIF4A3 on IFN-β does not rely on ATP binding, RNA-dependent ATPase, or RNA helicase activities. (A) Schematic diagram of the eIF4A3 truncated segments. (B) Effect of eIF4A3, eIF4A3-Nt, or eIF4A3-Ct on the IFN-β luciferase activity. HEK293T cells were transfected with the HA-eIF4A3, HA-eIF4A3-Nt, HA-eIF4A3-Ct, or vector (HA) and the IFN-β luciferase reporter, plus the pRL-TK. Then, 24 h after transfection, the cells were stimulated by SeV for 10 h and measured by the luciferase assays (*, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test). (C) Effect of eIF4A3, eIF4A3(DEAD-DQAD), or eIF4A3(Δ motif I) on the IFN-β luciferase activity. HEK293T cells were transfected with the HA-eIF4A3, HA-eIF4A3(DEAD-DQAD), HA-eIF4A3(Δ motif I), or vector (HA) and the IFN-β luciferase reporter, plus the pRL-TK. Then, 24 h after transfection, the cells were stimulated by SeV for 10 h and measured by the luciferase assays (*, P < 0.05; **, P < 0.01; ***, P < 0.001; ****; P < 0.0001; two-tailed Student’s t test). (D) Interactions between Flag-IRF3 and HA-eIF4A3, HA-eIF4A3(DEAD-DQAD), or HA-eIF4A3(Δ motif I). HEK293T cells were transfected with Flag-IRF3 and HA-eIF4A3, HA-eIF4A3(DEAD-DQAD), or HA-eIF4A3(Δ motif I), followed by lysing at 24 h posttransfection. A co-IP assay was carried out using anti-HA immunomagnetic beads, followed by a Western blot assay.
DISCUSSION
In this study, we revealed a novel and critical role of eIF4A3 in host antiviral innate immunity and showed that eIF4A3 overexpression resulted in reduction of the antiviral immune responses by inhibiting the activation of IRF3 and the transcription of IFN-β and ISGs. Mechanistically, eIF4A3 acted as an inhibitor of TBK1-IRF3 signaling to prevent TBK1 to activate IRF3. Therefore, our results provide new insights into the regulation of TBK1-IRF3 signaling and demonstrate a molecule of the DEAD box family as a key regulator of antiviral innate immunity.
Previous studies have reported that eIF4A3 is essential for the replication of IAVs (26, 27). Furthermore, eIF4A3 has been shown to promote AIVs by interacting with PB1, PB2, and NP (19). Our results demonstrate that eIF4A3 not only promotes the replication of avian and human IAVs that replicate in the nucleus, but also enhances the replication of other RNA viruses such as VSV and SeV that replicate in cytoplasm, indicating that eIF4A3 plays an important role in both cytoplasm and nucleus. eIF4A3 belongs to the DEAD box helicase family, eIF4A subfamily, which is involved in various steps in RNA metabolism (16). For example, eIF4A contains the same functional domain as eIF4A3, and its helicase activity is required to unwind extended and highly structured 5′ untranslated regions (UTRs) of several proto-oncogenes (28). eIF4A3 exhibits 65% identity to eIF4A at the amino acid level (29). The eIF4A inhibitor zotatifin is currently in a phase I clinical trial for cancer therapy and also is a potent antiviral molecule to inhibit the novel coronavirus SARS-CoV-2 (30). The natural compound silvestrol as an eIF4A inhibitor can inhibit Ebola virus in human primary macrophages at low nanomolar concentrations (31) and also has a broad-spectrum antiviral activity against corona- and picornaviruses (32). Our findings showed that eIF4A3 not only significantly inhibited the expression of IFN-β, but also extensively suppressed the expression of the ISGs (Mx1, ISG15, OAS3, OASL, IFIT2, IFIT2), which most likely explains why the inhibitors targeting eIF4A3 have a broad-spectrum antiviral activity. Furthermore, our results demonstrate that eIF4A3 inhibits the NF-κB promoter activity and the expression of NF-κB regulated proinflammatory cytokines (IL-6, IL-8, IL-1β, and TNF-α), despite the effect being not as significant as that observed in the IFN-β signaling pathway, suggesting that eIF4A3 may regulate cellular inflammatory responses. To understand the molecular mechanism of eIF4A3 on the NF-κB signaling pathway will be helpful to alleviate the overactive inflammatory responses caused by the use of antiviral drugs in the clinic.
It is acknowledged that infection-induced activation of IRF3 is responsible for the induction of type I IFN production. Activation of IRF3 is usually divided into three steps: phosphorylation, dimerization, and translocation from cytoplasm into nucleus. IRF3 locates in the cytoplasm, while IRF3-5D (the constitutively active form of IRF3) mainly locates in the nucleus. We found that eIF4A3 bound to IRF3 and IRF3-5D to reduce the IRF3- and IRF3-5D-stimulated IFN-β promoter activity. eIF4A3 is a nucleocytoplasmic shuttling protein that can enter the cytoplasm by binding to mRNA (33, 34). Noticeably, our studies revealed that the interaction between eIF4A3 and IRF3 disrupted the formation of the TBK1-IRF3 complex, resulting in the reduction of IRF3 activation. However, eIF4A3 could not interact with TBK1, as shown by the co-IP assay. This may be because the sequence of IRF3 binding to eIF4A3 probably overlaps with that of IRF3 recognized by TBK1, thereby blocking the recruitment of IRF3 to TBK1. The results shown in Fig. 7C confirmed our hypothesis. Upon viral infection and specific phosphorylation, IRF3 accumulates in the nucleus and associates with CREB binding protein/p300 via its DNA binding domain (35, 36). In this study, we also found that eIF4A3 could inhibit IRF3 binding to ISRE and impair the interaction of IRF3 and CBP/p300. This may be an indirect effect due to the fact that the nuclear translocation of IRF3 was blocked, ultimately leading to the biological functions in the nucleus being impaired.
RNA helicases can catalyze the separation of double-stranded nucleic acids in an energy-dependent manner and are encoded by many genomes (37). All helicases contain the common conserved regions: Walker A and B motifs (38). Walker A motif, also known as motif I (AxTGoGKT), is crucial for the ATP binding (39). Walker B motif, also known as motif II (DEAD), is necessary for the ATPase and helicase activities (24). eIF4A3, belongs to the DEAD box family of RNA helicases and possesses all nine conserved motifs (Q, I, Ia, Ib, II, III, IV, V, VI) (25). A previous study revealed that motif I (AxTGoGKT) was essential for ATP binding activity of eIF4A3, while eIF4A3 lost the abilities of the ATPase and helicase and could not participate in the process of ATP hydrolysis when the motif DEAD mutated to DQAD. However, the DQAD mutant was still capable of binding ATP and RNA (24). The Q motif was found to form the binding pocket for the adenylate base of ATP and to establish direct interactions with the binding nucleotides (22, 40). Our results showed that the N terminus of eIF4A3, consisting of motif I, DEAD box and Q motif, played a dominant role in inhibition of the type I IFN expression. Interestingly, the inhibition of eIF4A3 on the type I IFN expression was independent of the abilities of the ATP binding, ATPase, and helicase. Based on our findings, we speculated that the suppression of eIF4A3 on antiviral innate immunity might not be related to ATP.
Our findings reveal the mechanism of eIF4A3 negative regulation of the type I IFN induction pathway. eIF4A3 interacts with IRF3 to impair the formation of the TBK1-IRF3 complex, which impedes the phosphorylation and nuclear translocation of IRF3. This impairs the function of IRF3 in the nucleus and IRF3 binding to ISRE, as well as the interaction of IRF3 and CBP/p300, resulting in the reduction of type I IFN production. In conclusion, our studies provide new understanding of eIF4A3 regulating antiviral innate immune responses, which might be a potential target for drug development against viral infections.
MATERIALS AND METHODS
Cells and viruses.
Human embryonic kidney 293T cells (HEK293T) were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (HyClone, Beijing, China). The human lung epithelial cells (A549) and the human cervix epithelial cells (HeLa) were maintained in Ham’s F-12 medium and RPMI 1640 (HyClone), respectively. All media were supplemented with 10% fetal bovine serum (FBS) (PNA-Biotech, Germany), and all cells were cultured at 37°C in a 5% CO2 humidified atmosphere. All the cell lines listed above were purchased from ATCC (American Type Culture Collection, Manassas, VA, USA).
Influenza A/duck/Hubei/Hangmei01/2006 (HM, H5N1) and influenza A/Puerto Rico/8/1934 (PR8, H1N1) viruses were amplified in 10-day-old specific-pathogen-free (SPF) embryonic chicken eggs (Sparfas, Jinan, China). All experiments with the H5N1 virus were performed in a biosafety level 3 (BSL-3) laboratory. This study was carried out in accordance with the recommendations of the BSL-3 laboratory at Huazhong Agricultural University (HZAU). All procedures were approved by the Institutional Biosafety Committee of HZAU. Sendai virus (SeV) was amplified using 10-day-old embryonic chicken eggs (Sparfas). The recombinant vesicular stomatitis virus carrying green fluorescence protein (VSV-GFP) was a gift from Harbin Veterinary Research Institute (Harbin, China).
Antibodies and reagents.
Rabbit antibodies against human eIF4A3 (17504-1-AP; Proteintech, USA), TBK1 (28397-1-AP; Proteintech), IRF3 (11904; Cell Signaling Technology, USA), Ser386 phosphorylated IRF3 (AP0091; ABclonal Technology, USA), and mouse antibody against HA-tag (PMK013C; PMK Bio, China), Flag-tag (F1804, Sigma, USA), GFP-tag (PMK009S), Myc-tag (AE010; Proteintech), GAPDH (PMK043F; PMK Bio), and lamin B1 (12987-1-AP, Proteintech) were purchased from the indicated manufactures. Alexa Fluor 488-conjugated AffiniPure goat anti-mouse (GM200G-02C; Sungene Biotech) and Alexa Fluor 594-conjugated AffiniPure goat anti-rabbit (GR200G-43C; Sungene Biotech) secondary antibodies were used for the indirect immunofluorescence assay. The small-molecule compounds used in this study were DAPI (4′,6′-diamidino-2-phenylindole dihydrochloride; C1002; Beyotime, China) and poly(I·C) [polyinosinic-poly(C) potassium salt; 31852-29-6; Sigma, USA].
Plasmids and small interfering RNAs.
The eIF4A3 (NM_014740.4) gene was cloned into eukaryotic expression vector pCAGGS-HA at the ClaI and XhaI sites, into the vector p3×FLAG-CMV-14 at the SacI and XboI sites, or into the vector pCMV-Myc at SalI and NotI. Flag-RIG-I, -MAVS, -TBK1, -IKKε, and -IRF3 expression plasmids were kindly provided by Zhengfan Jiang (Peking University, Beijing, China). A luciferase (luc) reporter plasmid for the IFN-β promoter (IFN-β-luc) and Renilla control plasmid pRL-TK were a gift from Ping Qian (Huazhong Agricultural University, Wuhan, China). The Flag-tagged active phosphomimetic mutant of IRF3 (Flag-IRF3-5D) was kindly provided by Yiling Lin (National Defense Medical Center, Taiwan, China). The luciferase (luc) reporter plasmids for the ISRE promoter (ISRE-luc), the IRF3 promoter (IRF3-luc), the NF-κB promoter (NF-κB-luc), HA-RIG-I-N, HA-MAVS, HA-IRF3, HA-eIF4A3(DEAD-DQAD), HA-eIF4A3(Δ motif I), HA-TBK1, Flag-EGFP, and Flag-EGFP-ΔIRF3 were constructed by our laboratory. The PCR primers used in this study are listed in Table 1. Small interfering RNA (siRNA) targeting eIF4A3 (sieIF4A3) (5′-AAGCAGCAGATCAGTGGGATGAG-3′) and a nontarget siRNA (siNC) (5′-UUCUCCGAACGUGUCACGUTT-3′ and 5′-ACGUGACACGUUCGGAGAATT-3′) were synthesized by GenePharma (Shanghai, China).
TABLE 1.
Primers used for PCR
| Primer name | Sequences (5′ to 3′)a |
|---|---|
| Flag-eIF4A3 | F: CGGAATTCAATGGCTACAAACTTCCTAG |
| R: GCTCTAGAGATCTTGTTGAAAGCTGC | |
| Myc-eIF4A3 | F: GCGTCGACCATGGCGACCACGGCCACGAT |
| R: AAGAATGCGGCCGCATAAGATCAGCAACGTTCATCGGCATC | |
| HA-eIF4A3 | F: CCATCGATATGGCGACCACGGCCAC |
| R: CCCTCGAGTTAGATAAGATCAGCAACGTTCATCG | |
| HA-eIF4A3-Nt | F: CCATCGATATGGCGACCACGGCCACGA |
| R: CCCTCGAGTTCATCACGTTTCACCAAGATGCGGA | |
| HA-eIF4A3-Ct | F: CCATCGATTTGACTCTGGAAGGCATCAAGCAAT |
| R: CCCTCGAGGATAAGATCAGCAACGTTCATCGGC | |
| HA-eIF4A3(DEAD-DQAD) | F: TGGATCAAGCTGATGAAATGTT |
| R: AGCTTGATCCAAAACCAACATT | |
| HA-eIF4A3(Δ motif I) | F: CCATCGATATGGCGACCACGGCCAC |
| R: ACTGAGATACTGAAGGTGGCGATGACATCTCTCCCTTTGA | |
| F: TCAAAGGGAGAGATGTCATCGCCACCTTCAGTATCTCAGT | |
| R: CCCTCGAGGATAAGATCAGCAACGTTCATCG | |
| HA-RIG-I-N | F: GGGGTACCATGACCACCGAGCAGCGAC |
| R: CCCTCGAGTTTAAATGGGCTGTACAAGTTTGTATC | |
| HA-IRF3 | F: CGGAATTCATGGGAACCCCAAAGCCACGGATC |
| R: CCCTCGAGGCTCTCCCCAGGGCCCTGGAAAT | |
| HA-TBK1 | F: CCCATCGATATGCAGAGCACTTCTAATCATCTGT |
| R: CCGCTCGAGCTAAAGACAGTCAACGTTGCGAA | |
| Flag-EGFP | F: GGTTCAGGCGGAGGTGGCTCTGGCGGTGGCGGATCCATGGTGAGCAAGGGCGAGGAGCT |
| R: GCTCTAGACTTGTACAGCTCGTCCATGCCGAGAGT | |
| Flag-EGFP-ΔIRF3 | F: CCGGAATTCAATGCGGGTAGGGGGTGCCTCCTCCCT |
| R: ACCGCCAGAGCCACCTCCGCCTGAACCGCCTCCACCGCTCTCCCCAGGGCCCTGGAAAT |
The gene sequences are downloaded from NCBI. F, forward; R, reverse.
Transfection and virus titration.
Transfections were performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. Briefly, plasmids and siRNA transfections were done using Lipofectamine 2000 (Invitrogen) in cells maintained in Opti-MEM. Culture medium was replaced with fresh medium supplemented with 10% FBS after at 6 h posttransfection. For growth dynamics of IAV, A549 cells in 12-well plates were infected with the IAV at a multiplicity of infection (MOI) of 0.01. After virus adsorption for 1 h, cells were washed with phosphate-buffered saline (PBS) and then incubated in Ham’s F-12 medium with or without 0.25 μg/mL TPCK (tosylsulfonyl phenylalanyl chloromethyl ketone; Sigma) at 37°C. Viral supernatants were harvested at the indicated time points and titrated on MDCK cells. Briefly, the serially diluted supernatants were inoculated onto MDCK cell monolayers in 96-well plates for 1 h, and the inoculum was removed and replaced with DMEM containing Trypsin-TPCK (Sigma). The plates were incubated for 72 h at 37°C. Virus titers were determined by calculating log10 50% tissue culture infection dose (TCID50)/milliliter using the Reed-Muench method. A549 cells transfected with the indicated plasmids for 24 h were infected with VSV-GFP (MOI, 0.1) or SeV (MOI, 0.1). Then, 2 h after infection, the cells were washed with PBS twice, and fresh medium with 1% FBS was then added to incubate for 12 h and 24 h. Infected cells were collected for qRT-PCR analysis or used for fluorescence microscopy analysis.
Dual-luciferase reporter assay.
HEK293T cells in 24-well plates were transfected with the IFN-β-luc, IRF3-luc, NF-κB-luc, ISRE-luc (0.3 μg/well), and pRL-TK (10 ng/well), and the cells were harvested 24 h posttransfection and then induced by SeV for 6 h. To stimulate IFN-β promoter luciferase activity, the RIG-I, RIG-I-N, MAVS, TBK1, IKKε, IRF3, or IRF3-5D expression plasmids (0.3 μg/well) were transfected together with luciferase reporter plasmids. Then, 24 h after transfection, the cells were lysed in 100 μL of 1× passive lysis buffer (PLB; Promega) to measure luciferase signals by using a dual-luciferase reporter assay system (Promega) according to the manufacturer’s instructions. The Renilla activity was used as an internal control and to normalize transfection efficiency. All experiments were performed in triplicate and repeated at least three times.
Quantitative RT-PCR assay.
Total RNA was extracted from cells using TRIzol reagent (Invitrogen, USA) following the protocols recommended by the manufacturer. The RNA samples were treated with DNase I (Ambion) at 37°C for 30 min to remove the residual DNA. cDNA was prepared from 2 μg of RNA using oligo(dT) and 1 μL of avian myeloblastosis virus (AMV) reverse transcriptase (TaKaRa, Tokyo) in a total volume of 20 μL at 42°C for 1 h and 72°C for 15 min. Real-time PCR (RT-PCR; ViiA7; ABI, USA) was performed using FastStart Universal SYBR green master (Roche). The relative quantifications of the mRNAs were normalized to GAPDH (glyceraldehyde-3-phosphate dehydrogenase) using the 2–ΔΔCT method. The gene-specific primer sequences for qRT-PCR are shown in Table 2.
TABLE 2.
Primers used for qRT- PCR
| Primer name | Sequences (5′ to 3′)a |
|---|---|
| GAPDH | F: GCTAAGGCTGTGGGCAAGG |
| R: GGAGGAGTGGGTGTCGCTG | |
| SeV-M | F: GTGATTTGGGCGGCATCT |
| R: GATGGCCGGTTGGAACAC | |
| VSV-N | F: ACGGCGTACTTCCAGATGG |
| R: CTCGGTTCAAGATCCAGGT | |
| PR8-NP | F: GCGTCTCAAGGCACCAAAC |
| R: AACCGTCCCTCATAATCAC | |
| IFN-β | F: GCTCCTGTGGCAATTGAATGG |
| R: TTGGCCTTCAGGTAATGCAG | |
| Mx1 | F: CCGAGGGAGACAGGACCAT |
| R: CGTGGCCTTTCCTTCCTCC | |
| ISG15 | F: TGGACAAATGCGACGAACC |
| R: GCCCGCTCACTTGCTGCTT | |
| IFIT2 | F: GGTCTCTTCAGCATTTATTGGTG |
| R: TGCCGTAGGCTGCTCTCCA | |
| IFITM3 | F: GGGACAGGAAGATGGTTGG |
| R: CACTGGGATGACGATGAGC | |
| OAS3 | F: CACAGACCTAAGGGATGGC |
| R: TCAGGAAACTGAAGGCTCA | |
| OASL | F: GCCTTCTCTTCCCAACTCCC |
| R: AGGCATAGATGGTTAGAAGTTCAAGA | |
| TNF-α | F: CAGGCGGTGCTTGTTCC |
| R: AAGAGGACCTGGGAGTAGATGA | |
| IL-1β | F: AGCTACGAATCTCCGACCAC |
| R: CGTTATCCCATGTGTCGAAGAA | |
| IL-6 | F: ACAGCCACTCACCTCTTCAGAAC |
| R: GCTCTGGCTTGTTCCTCACTACTC | |
| IL-8 | F: TGCAGCTCTGTGTGAAGGTG |
| R: CAGCCCTCTTCAAAAACTTCTCC | |
| CXCL11 | F: ATTTGCTGCCTTATCTTTCTGACTCTA |
| R: TGGCCTTCGATTCTGGATTCA |
F, forward; R, reverse.
Indirect immunofluorescence assay.
A549 or HeLa cells were grown on coverslips in 12-well plates and transfected or infected as indicated. At time points postinfection or posttransfection, cells were rinsed twice with PBS and fixed with 4% paraformaldehyde (PFA) for 15 min at room temperature (RT). The cells were subsequently permeabilized with 0.2% Triton X-100 for 15 min and blocked with 2% bovine serum albumin (BSA) in PBS at RT for 30 min. Following incubation with the primary antibody for 2 h, the cells were washed and incubated with the appropriate Alexa Fluor-conjugated secondary antibody for 1 h, and nuclei were stained with DAPI (4′,6-diamidino-2-phenylindole) for 15 min at RT. These samples were observed using a confocal microscope (LSM880; Carl Zeiss, Germany).
Isolation of cytoplasmic and nuclear fractions.
A549 cells were harvested in ice-cold PBS and collected by centrifugation at 1,000 rpm for 5 min. The cell pellet was resuspended in hypotonic lysis buffer (10 mM HEPES-NaOH [pH 7.9], 10 mM KCl, 1.5 mM MgCl2, 0.5 mM beta-mercaptoethanol) supplemented with protease inhibitors and phosphatase inhibitors, vortexed, and then incubated on ice for 15 min, followed by the addition of 3 μL 10% NP-40 for another 5 min. Samples were centrifuged at 16,000 × g for 10 min, the supernatants were used to collect cytoplasmic proteins, and the pellets were used for nuclear proteins extraction. Pellets were washed with cold PBS twice, RIPA buffer (V900854, Sigma, USA) was added as nuclear lysis buffer, and then they mixture was dispersed and incubated on ice for 20 min. The extractions were centrifuged at 16,000 × g for 10 min for nuclear proteins. The proteins were collected and stored at −80°C.
Immunoprecipitation assay and native PAGE.
HEK293T cells seeded on 10-cm dishes were transfected in 10 μg of the indicated expression plasmids by using Lipofectamine 2000 (Invitrogen). Then, 36 h after transfection, cells were lysed in immunoprecipitation (IP) lysis buffer (Beyotime, China) together with a protease inhibitor cocktail (BioTools, USA) on ice, and then the lysis was centrifuged. Supernatants were collected and incubated with anti-HA immunomagnetic beads (B26202; Bimake, USA) or anti-Flag immunomagnetic beads (B26102; Bimake) overnight at 4°C. The beads were washed five times with cold IP lysis buffer and were eluted with 5× loading buffer by boiling of samples for 10 min before SDS-PAGE and immunoblot analysis.
To determine the IRF3 dimerization, native PAGE assays were performed. HEK293T cells transfected with the indicated plasmids were harvested and lysed with ice-cold lysis buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1 mM phenylmethylsulfonyl fluoride [PMSF] and 0.5% NP-40). Next, the cell lysates were diluted with 5× loading buffer. Finally, the samples were loaded onto 12 by 1.5 mm precast 7.5% native gels without SDS and separated at 20 mA for 90 min by electrophoresis on ice, followed by immunoblot analysis.
DNA pulldown assay.
The sequence of triple repeats of IRF3-recognized and bound ISG54 ISRE (5′-GGGAAAGTGAAACTAGGGAAAGTGAAACTAGGGAAAGTGAAACTA-3′) (41, 42) was synthesized by Tsingke Biological Technology (Wuhan, China) and labeled with biotin (biotinylated ISRE) or without biotin (unbiotinylated ISRE). HEK293T cells were transfected with HA-IRF3 in the absence or presence of Flag-eIF4A3 and infected with SeV at 24 h posttransfection. At 12 hpi, cells were lysed, and the BeaverBeads streptavidin beads (22305-1; Beaverbio, China) were washed three times with cold IP lysis buffer. The single-stranded DNA samples were thermally annealed to form double-stranded DNA (dsDNA) and incubated with streptavidin beads for 2 h at 4°C. The beads were washed three times with cold IP lysis buffer, and the lysate was then incubated with probe-bound beads at 4°C overnight. The beads were washed five times with cold IP lysis buffer and were eluted with 5× loading buffer by boiling of samples for 10 min before SDS-PAGE and immunoblot analysis.
Cell viability assay.
Cell viability was assessed using a colorimetric-based CCK-8 assay (Dojindo Molecular Technologies). Briefly, A549 cells in 96-well plates were transfected with si-eIF4A3 or si-NC, and the cell viability was measured at 24 h posttransfection. Then, 10 μL of CCK-8 reagent was added to each well of the plates, and cells were incubated at 37°C for 4 h; the absorbance at 450 nm was measured with a microplate reader.
Statistical analysis.
The data are presented as the means ± standard deviations (SD) from three independent experiments. Statistical analyses were performed using Prism 8 software, version 8 (GraphPad Software, Inc.). Statistical significance was calculated using Student’s two-tailed unpaired t test (*, P < 0.05; **, P < 0.01; ***, P < 0.00; ****, P < 0.0001).
ACKNOWLEDGMENTS
This work was supported by the National Key Research and Development Program (2021YFD1800204), the National Natural Science Foundation of China (32025036 and 31772752), the earmarked fund for CARS-41, and the Natural Science Foundation of Hubei Province (2021CFA016).
We thank Wenjun Ma (University of Missouri, USA) and Xiao Xiao (Huazhong Agricultural University, China) for critically proofreading the manuscript.
H.Z. conceived the project; Q.G., M.J., Y.Z., G.L., and C.R. conducted the experiments; Q.G., M.J., Y.Z., C.Y., H.C., and H.Z. analyzed the data; Q.G., M.J., Y.Z., and H.Z. wrote the paper. All authors reviewed and approved the final manuscript.
Contributor Information
Hongbo Zhou, Email: hbzhou@mail.hzau.edu.cn.
Jae U. Jung, Lerner Research Institute, Cleveland Clinic
REFERENCES
- 1.Akira S, Uematsu S, Takeuchi O. 2006. Pathogen recognition and innate immunity. Cell 124:783–801. 10.1016/j.cell.2006.02.015. [DOI] [PubMed] [Google Scholar]
- 2.Kumar H, Kawai T, Akira S. 2011. Pathogen recognition by the innate immune system. Int Rev Immunol 30:16–34. 10.3109/08830185.2010.529976. [DOI] [PubMed] [Google Scholar]
- 3.Meylan E, Curran J, Hofmann K, Moradpour D, Binder M, Bartenschlager R, Tschopp J. 2005. Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus. Nature 437:1167–1172. 10.1038/nature04193. [DOI] [PubMed] [Google Scholar]
- 4.Xu LG, Wang YY, Han KJ, Li LY, Zhai Z, Shu HB. 2005. VISA is an adapter protein required for virus-triggered IFN-beta signaling. Mol Cell 19:727–740. 10.1016/j.molcel.2005.08.014. [DOI] [PubMed] [Google Scholar]
- 5.Seth RB, Sun L, Ea CK, Chen ZJ. 2005. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-kappaB and IRF 3. Cell 122:669–682. 10.1016/j.cell.2005.08.012. [DOI] [PubMed] [Google Scholar]
- 6.Sharma S, tenOever BR, Grandvaux N, Zhou GP, Lin R, Hiscott J. 2003. Triggering the interferon antiviral response through an IKK-related pathway. Science 300:1148–1151. 10.1126/science.1081315. [DOI] [PubMed] [Google Scholar]
- 7.Honda K, Takaoka A, Taniguchi T. 2006. Type I inteferon gene induction by the interferon regulatory factor family of transcription factors. Immunity 25:349–360. 10.1016/j.immuni.2006.08.009. [DOI] [PubMed] [Google Scholar]
- 8.Lin R, Mamane Y, Hiscott J. 1999. Structural and functional analysis of interferon regulatory factor 3: localization of the transactivation and autoinhibitory domains. Mol Cell Biol 19:2465–2474. 10.1128/MCB.19.4.2465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wathelet MG, Lin CH, Parekh BS, Ronco LV, Howley PM, Maniatis T. 1998. Virus infection induces the assembly of coordinately activated transcription factors on the IFN-beta enhancer in vivo. Mol Cell 1:507–518. 10.1016/S1097-2765(00)80051-9. [DOI] [PubMed] [Google Scholar]
- 10.Zhang H, Han C, Li T, Li N, Cao X. 2019. The methyltransferase PRMT6 attenuates antiviral innate immunity by blocking TBK1-IRF3 signaling. Cell Mol Immunol 16:800–809. 10.1038/s41423-018-0057-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Xiao J, Li W, Zheng X, Qi L, Wang H, Zhang C, Wan X, Zheng Y, Zhong R, Zhou X, Lu Y, Li Z, Qiu Y, Liu C, Zhang F, Zhang Y, Xu X, Yang Z, Chen H, Zhai Q, Wei B, Wang H. 2020. Targeting 7-dehydrocholesterol reductase integrates cholesterol metabolism and IRF3 activation to eliminate infection. Immunity 52:109–122.e6. 10.1016/j.immuni.2019.11.015. [DOI] [PubMed] [Google Scholar]
- 12.Long L, Deng Y, Yao F, Guan D, Feng Y, Jiang H, Li X, Hu P, Lu X, Wang H, Li J, Gao X, Xie D. 2014. Recruitment of phosphatase PP2A by RACK1 adaptor protein deactivates transcription factor IRF3 and limits type I interferon signaling. Immunity 40:515–529. 10.1016/j.immuni.2014.01.015. [DOI] [PubMed] [Google Scholar]
- 13.Saitoh T, Tun-Kyi A, Ryo A, Yamamoto M, Finn G, Fujita T, Akira S, Yamamoto N, Lu KP, Yamaoka S. 2006. Negative regulation of interferon-regulatory factor 3-dependent innate antiviral response by the prolyl isomerase Pin1. Nat Immunol 7:598–605. 10.1038/ni1347. [DOI] [PubMed] [Google Scholar]
- 14.Wang P, Zhao W, Zhao K, Zhang L, Gao C. 2015. TRIM26 negatively regulates interferon-beta production and antiviral response through polyubiquitination and degradation of nuclear IRF3. PLoS Pathog 11:e1004726. 10.1371/journal.ppat.1004726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lei CQ, Zhang Y, Xia T, Jiang LQ, Zhong B, Shu HB. 2013. FoxO1 negatively regulates cellular antiviral response by promoting degradation of IRF3. J Biol Chem 288:12596–12604. 10.1074/jbc.M112.444794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Linder P, Jankowsky E. 2011. From unwinding to clamping: the DEAD box RNA helicase family. Nat Rev Mol Cell Biol 12:505–516. 10.1038/nrm3154. [DOI] [PubMed] [Google Scholar]
- 17.Ryu I, Won YS, Ha H, Kim E, Park Y, Kim MK, Kwon DH, Choe J, Song HK, Jung H, Kim YK. 2019. eIF4A3 phosphorylation by CDKs affects NMD during the cell cycle. Cell Rep 26:2126–2139.e9. 10.1016/j.celrep.2019.01.101. [DOI] [PubMed] [Google Scholar]
- 18.Hauer C, Sieber J, Schwarzl T, Hollerer I, Curk T, Alleaume AM, Hentze MW, Kulozik AE. 2016. Exon junction complexes show a distributional bias toward alternatively spliced mRNAs and against mRNAs coding for ribosomal proteins. Cell Rep 16:1588–1603. 10.1016/j.celrep.2016.06.096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ren X, Yu Y, Li H, Huang J, Zhou A, Liu S, Hu P, Li B, Qi W, Liao M. 2019. Avian influenza A virus polymerase recruits cellular RNA helicase eIF4A3 to promote viral mRNA splicing and spliced mRNA nuclear export. Front Microbiol 10:1625. 10.3389/fmicb.2019.01625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tan X, Sun L, Chen J, Chen ZJ. 2018. Detection of microbial infections through innate immune sensing of nucleic acids. Annu Rev Microbiol 72:447–478. 10.1146/annurev-micro-102215-095605. [DOI] [PubMed] [Google Scholar]
- 21.Shu C, Sankaran B, Chaton CT, Herr AB, Mishra A, Peng J, Li P. 2013. Structural insights into the functions of TBK1 in innate antimicrobial immunity. Structure 21:1137–1148. 10.1016/j.str.2013.04.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cordin O, Tanner NK, Doere M, Linder P, Banroques J. 2004. The newly discovered Q motif of DEAD-box RNA helicases regulates RNA-binding and helicase activity. EMBO J 23:2478–2487. 10.1038/sj.emboj.7600272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cordin O, Banroques J, Tanner NK, Linder P. 2006. The DEAD-box protein family of RNA helicases. Gene 367:17–37. 10.1016/j.gene.2005.10.019. [DOI] [PubMed] [Google Scholar]
- 24.Pause A, Sonenberg N. 1992. Mutational analysis of a DEAD box RNA helicase: the mammalian translation initiation factor eIF-4A. EMBO J 11:2643–2654. 10.1002/j.1460-2075.1992.tb05330.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Shibuya T, Tange TO, Stroupe ME, Moore MJ. 2006. Mutational analysis of human eIF4AIII identifies regions necessary for exon junction complex formation and nonsense-mediated mRNA decay. RNA 12:360–374. 10.1261/rna.2190706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Cao M, Wei C, Zhao L, Wang J, Jia Q, Wang X, Jin Q, Deng T. 2014. DnaJA1/Hsp40 is co-opted by influenza A virus to enhance its viral RNA polymerase activity. J Virol 88:14078–14089. 10.1128/JVI.02475-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Karlas A, Machuy N, Shin Y, Pleissner KP, Artarini A, Heuer D, Becker D, Khalil H, Ogilvie LA, Hess S, Maurer AP, Muller E, Wolff T, Rudel T, Meyer TF. 2010. Genome-wide RNAi screen identifies human host factors crucial for influenza virus replication. Nature 463:818–822. 10.1038/nature08760. [DOI] [PubMed] [Google Scholar]
- 28.Chang JH, Cho YH, Sohn SY, Choi JM, Kim A, Kim YC, Jang SK, Cho Y. 2009. Crystal structure of the eIF4A-PDCD4 complex. Proc Natl Acad Sci USA 106:3148–3153. 10.1073/pnas.0808275106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Li Q, Imataka H, Morino S, Rogers GW, Jr, Richter-Cook NJ, Merrick WC, Sonenberg N. 1999. Eukaryotic translation initiation factor 4AIII (eIF4AIII) is functionally distinct from eIF4AI and eIF4AII. Mol Cell Biol 19:7336–7346. 10.1128/MCB.19.11.7336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O'Meara MJ, Rezelj VV, Guo JZ, Swaney DL, Tummino TA, Huttenhain R, Kaake RM, Richards AL, Tutuncuoglu B, Foussard H, Batra J, Haas K, Modak M, Kim M, Haas P, Polacco BJ, Braberg H, Fabius JM, Eckhardt M, Soucheray M, Bennett MJ, Cakir M, McGregor MJ, Li Q, Meyer B, Roesch F, Vallet T, Mac Kain A, Miorin L, Moreno E, Naing ZZC, Zhou Y, Peng S, Shi Y, Zhang Z, Shen W, Kirby IT, Melnyk JE, Chorba JS, Lou K, Dai SA, Barrio-Hernandez I, Memon D, Hernandez-Armenta C, et al. 2020. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583:459–468. 10.1038/s41586-020-2286-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Biedenkopf N, Lange-Grünweller K, Schulte FW, Weißer A, Müller C, Becker D, Becker S, Hartmann RK, Grünweller A. 2017. The natural compound silvestrol is a potent inhibitor of Ebola virus replication. Antiviral Res 137:76–81. 10.1016/j.antiviral.2016.11.011. [DOI] [PubMed] [Google Scholar]
- 32.Müller C, Schulte FW, Lange-Grünweller K, Obermann W, Madhugiri R, Pleschka S, Ziebuhr J, Hartmann RK, Grüunweller A. 2018. Broad-spectrum antiviral activity of the eIF4A inhibitor silvestrol against corona- and picornaviruses. Antiviral Res 150:123–129. 10.1016/j.antiviral.2017.12.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Michelle L, Cloutier A, Toutant J, Shkreta L, Thibault P, Durand M, Garneau D, Gendron D, Lapointe E, Couture S, Le Hir H, Klinck R, Elela SA, Prinos P, Chabot B. 2012. Proteins associated with the exon junction complex also control the alternative splicing of apoptotic regulators. Mol Cell Biol 32:954–967. 10.1128/MCB.06130-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chan CC, Dostie J, Diem MD, Feng W, Mann M, Rappsilber J, Dreyfuss G. 2004. eIF4A3 is a novel component of the exon junction complex. RNA 10:200–209. 10.1261/rna.5230104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kumar KP, McBride KM, Weaver BK, Dingwall C, Reich NC. 2000. Regulated nuclear-cytoplasmic localization of interferon regulatory factor 3, a subunit of double-stranded RNA-activated factor 1. Mol Cell Biol 20:4159–4168. 10.1128/MCB.20.11.4159-4168.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Honda K, Taniguchi T. 2006. IRFs: master regulators of signalling by Toll-like receptors and cytosolic pattern-recognition receptors. Nat Rev Immunol 6:644–658. 10.1038/nri1900. [DOI] [PubMed] [Google Scholar]
- 37.Silverman E, Edwalds-Gilbert G, Lin RJ. 2003. DExD/H-box proteins and their partners: helping RNA helicases unwind. Gene 312:1–16. 10.1016/s0378-1119(03)00626-7. [DOI] [PubMed] [Google Scholar]
- 38.Walker JE, Saraste M, Runswick MJ, Gay NJ. 1982. Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J 1:945–951. 10.1002/j.1460-2075.1982.tb01276.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rozen F, Pelletier J, Trachsel H, Sonenberg N. 1989. A lysine substitution in the ATP-binding site of eucaryotic initiation factor 4A abrogates nucleotide-binding activity. Mol Cell Biol 9:4061–4063. 10.1128/mcb.9.9.4061-4063.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Tanner NK. 2003. The newly identified Q motif of DEAD box helicases is involved in adenine recognition. Cell Cycle 2:18–19. 10.4161/cc.2.1.296. [DOI] [PubMed] [Google Scholar]
- 41.Nakaya T, Sato M, Hata N, Asagiri M, Suemori H, Noguchi S, Tanaka N, Taniguchi T. 2001. Gene induction pathways mediated by distinct IRFs during viral infection. Biochem Biophys Res Commun 283:1150–1156. 10.1006/bbrc.2001.4913. [DOI] [PubMed] [Google Scholar]
- 42.Meng F, Zhou R, Wu S, Zhang Q, Jin Q, Zhou Y, Plouffe SW, Liu S, Song H, Xia Z, Zhao B, Ye S, Feng XH, Guan KL, Zou J, Xu P. 2016. Mst1 shuts off cytosolic antiviral defense through IRF3 phosphorylation. Genes Dev 30:1086–1100. 10.1101/gad.277533.116. [DOI] [PMC free article] [PubMed] [Google Scholar]