Summary
Chromodomain helicase DNA-binding domain 2 (CHD2) is a chromatin remodeling factor involved in many developmental processes. However, its role in male germ cell development has not been elucidated. Here, we confirm that CHD2 expression is enriched in the male germline. In a heterozygous knockout mouse model of Chd2 (Chd2+/−), we demonstrated that Chd2 haploinsufficiency resulted in testicular developmental delay, an increased rate of abnormal sperm, and impaired fertility in mice. In vitro experiments in mouse spermatogonia showed that CHD2 knockdown inhibits spermatogonial self-renewal. Mechanistically, CHD2 maintains the enrichment of H3K4me3 in the Ccnb1 and Ccnd2 promotors, consequently promoting the transcription of Ccnb1 and Ccnd2. In addition, CHD2 interacts with the cleavage stimulation factor CSTF3 and upregulates the expression of OCT4 and PLZF by improving mRNA stability. This is the first study to reveal the role and mechanism of CHD2 in maintaining spermatogonial self-renewal.
Subject areas: Biological sciences, molecular mechanism of gene regulation, cell biology, transcriptomics, alternate contact
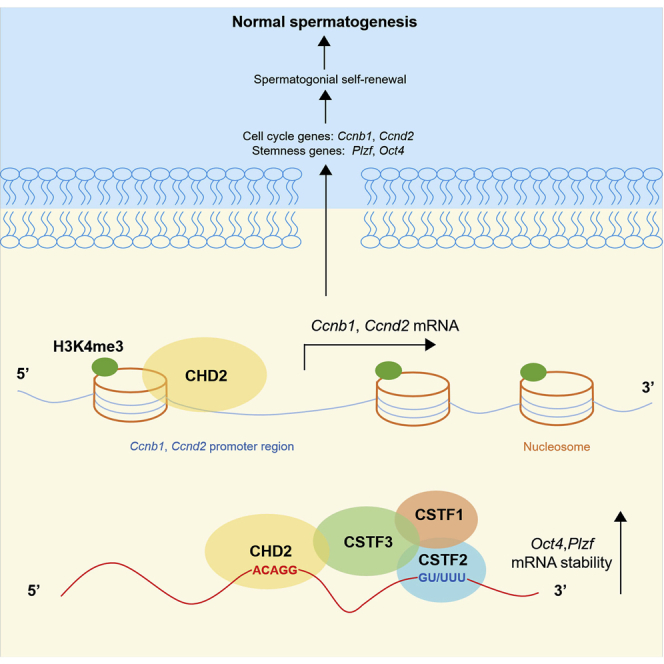
Graphical abstract
Highlights
-
•
CHD2 maintains spermatogonial self-renewal in spermatogenesis
-
•
CHD2 maintains H3K4me3 enrichment in the promotor of Ccnb1 and Ccnd2
-
•
CHD2 interacts with CSTF3 to enhance the mRNA stability of Oct4 and Plzf
Biological sciences; Molecular mechanism of gene regulation; Cell biology; Transcriptomics; Alternate contact.
Introduction
Spermatogenesis is a complex and precise process that begins with spermatogonial proliferation, spermatogonial differentiation into spermatocytes, spermatocyte meiosis into round spermatozoa, chromatin concentration and morphological change, and release of mature spermatozoa into the convoluted tubules.1,2 This process, accompanied by changes in the morphology and function of germ cells, is finely regulated by a complicated network composed of transcription factors, epigenetic regulators and signaling pathways. In the early stage of spermatogenesis, spermatogonia are faced with the choice of self-renewal or differentiation.3 Multiple genes have been confirmed to determine germ cell fate. OCT4, NANOG, PLZF, ID4, GFRα1, and LIN28A are essential for spermatogonial stem cell (SSC) self-renewal,4,5,6,7,8,9,10 while C-KIT, STRA8, and SPO11 are hallmarks of differentiating spermatogonia.11,12,13 Disrupted expression of these genes can lead to the failure of spermatogonial self-renewal or differentiation, which are important causes of male infertility.
Mammalian ATP-dependent chromatin remodeling complexes are chromatin structure regulators that are highly conserved in evolution and play vital roles in regulating genomic architecture through the energy produced by the hydrolysis of ATP. Due to the modular organization and pathways of assembly of these chromatin regulators, the chromatin remodeling status of different gene loci in various tissues will differ, which determines the specificity of gene expression and cell fate.14,15 The chromatin remodeling complexes can be classified into the switching defective/sucrose nonfermenting (SWI/SNF), imitation switch (ISWI), inositol-requiring 80 (INO80) and chromodomain helicase DNA-binding protein (CHD) families.16 Increasing studies have confirmed the close relationship between chromatin remodeling defects and male sterility. For example, knockout of Brg1, the catalytic subunit of the SWI/SNF chromatin remodeling complex, causes impaired synapsis and compromised DNA repair in mouse spermatocytes and results in meiotic arrest.17 Deletion of INO80 leads to a failure to repair DNA damage during meiosis recombination and arrest.18,19 Zinc finger HIT-type containing 1 (Znhit1), a subunit of the SRCAP chromatin remodeling complex, controls meiotic initiation in male germ cells.20 CHD4 and CHD5 are both members of the CHD family. CHD4 is essential for SSC maintenance, and loss of CHD4 impairs the regenerative capacity of SSCs upon transplantation back into the mouse testis.21 CHD5 has been shown to be involved in chromatin concentration during spermiogenesis, and its deficiency leads to chromatin compression defects and male sterility in mice.22,23 The above research shows that chromatin remodeling factors are involved in the regulation of gene expression, DNA damage repair and replacement of classic histones and protamine in spermatogenesis, and their loss causes spermatogenesis arrest. Therefore, exploring the function of chromatin remodeling factors in spermatogenesis may help explain the causes of male sterility.
As a member of the CHD family, CHD2 is involved in embryonic development, embryonic neurogenesis, neural circuit development, myelopoiesis, DNA damage repair and other biological processes.24,25,26,27,28 In mice, homozygous knockout of Chd2 results in overall developmental delay and perinatal lethality, and heterozygous mutation of Chd2 causes histological abnormalities of the kidney, spleen, heart and liver.29 Chd2 heterozygous mutant mice also showed neuronal proliferation defects and neuronal excitability changes, including divergent changes in excitatory and inhibitory synaptic function.26 During mouse embryonic neurogenesis in vivo, CHD2 is essential for the development of the neocortex and the proliferation of progenitor cells by promoting the expression of REST.25 In mouse embryonic stem cells, CHD2 interacts with OCT3/4 to promote the deposition of histone variant H3.3 on developmental genes and inhibit the formation of heterochromatin, thereby promoting developmental gene expression during the differentiation of embryonic stem cells.24 Similar functions of CHD2 are also found in myelopoiesis.27 During these processes, CHD2 plays an important role in global transcriptional regulation in stem cells. In humans, CHD2 haploinsufficiency is associated with various catastrophic childhood epilepsies, intellectual disabilities, autism spectrum disorders, and photosensitivity.30 It has been reported that among 611 patients with epilepsy carrying pathogenic mutations, 17 (2.8%) carried CHD2 mutations.31 In the research field of reproduction, a study on systems biology methods combining GWAS and RNA-seq technology was performed to explore the molecular basis of sperm quality in boars, and 12 quantitative trait loci (QTLSs) associated with abnormal sperm quality were identified, among which Chd2 was one of the candidate gene loci.32 Interestingly, Rs1406714 in CHD2 has been reported to be associated with nonobstructive azoospermia (NOA) in Han Chinese men.33 To date, the role of CHD2 during spermatogenesis has not been explored, and a previous study suggested the possibility of CHD2 involvement in sperm quality and azoospermia. Thus, it is important to determine whether and how CHD2 mutations or abnormal expression affect male fertility in humans, which may be helpful to diagnose male sterility.
CHD2 is highly conserved in the mammalian genome, and the homology of the CHD2 amino acid sequence between humans and mice is 94%, which makes it possible to study the function of human CHD2 using a mouse model. The aim of the present study was to clarify the functional role and regulatory mechanism of CHD2 in spermatogenesis in vitro and in vivo. We confirmed that CHD2 promotes spermatogonia self-renewal by maintaining H3K4me3 enrichment and increasing the mRNA stability of target genes. Our study may provide a potential target for male reproductive disorders.
Results
Chromodomain helicase DNA-binding domain 2 expression is enriched in the male germline
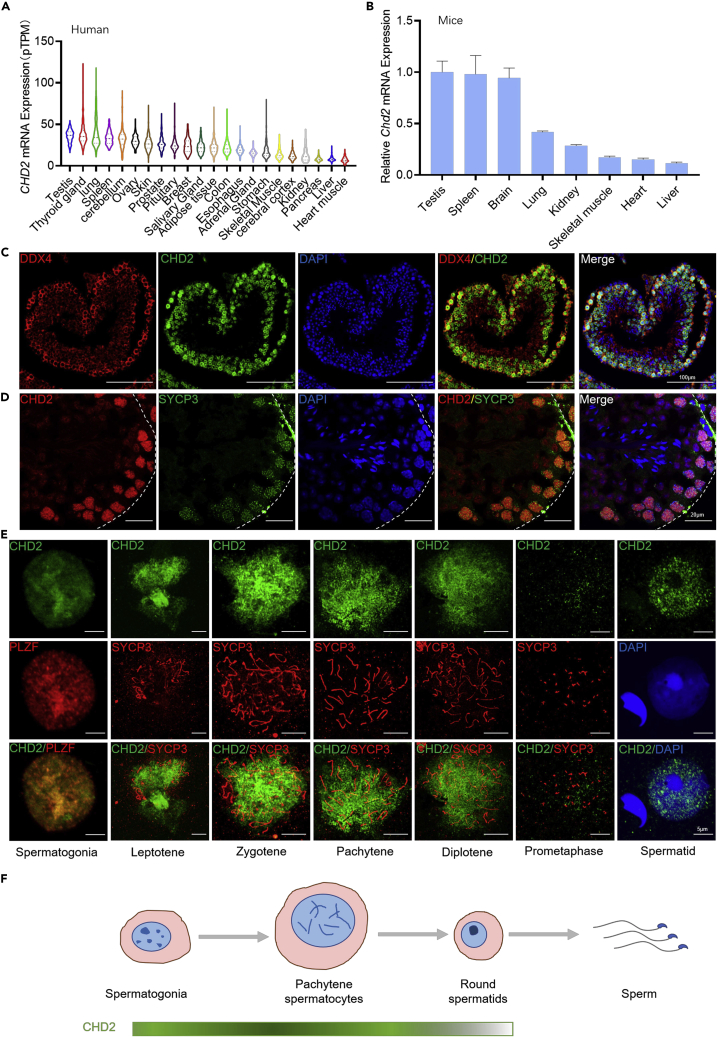
The expression profile of the CHD2 gene in the Human Protein Atlas (HPA, https://www.proteinatlas.org/) indicates that CHD2 is more highly expressed in human testes than in other tissues (Figure 1A). In mice, RT‒qPCR results of multiple organs showed that Chd2 mRNA was also more highly expressed in mouse testes (Figure 1B). The HPA and GEO databases (GSE4193) show that Chd2 mRNA is expressed in both human and mouse type A spermatogonia, type B spermatogonia, pachytene spermatocytes and round spermatids (Figures S1A and S1B). Using DDX4 and SYCP3 as markers of spermatogenic cells and meiosis I spermatocytes, the immunofluorescence analysis of frozen sections of mouse adult testis showed that CHD2 is abundantly expressed throughout the germ cells, with the highest expression in the spermatocytes of meiosis I, while no fluorescence signal was detected in sperm (Figures 1C–1E). Using SOX9 as a Sertoli cell marker and CYP11A1 as a Leydig cell marker, immunofluorescence staining indicated that CHD2 is also expressed in Sertoli cells and Leydig cells (Figures S1C and S1D). The same phenomenon was observed by immunohistochemical analysis in mouse testis (Figure S1E). Negative controls of the immunofluorescence and immunohistochemical experiments are presented in Figure S2. These data suggest that CHD2 may have physiological functions in the testis.
Figure 1.
CHD2 is enriched throughout the testicular germ cells, with the highest expression in the spermatocytes of meiosis I
(A) Expression analysis of CHD2 mRNA in human tissues based on the HPA database.
(B) Expression analysis of Chd2 mRNA in mouse tissues by RT‒qPCR detection relative to 18S rRNA. Data are shown as the mean ± SD.
(C and D) The expression and location of CHD2 in adult mouse seminiferous tubule frozen sections by immunofluorescence analysis. Scale bars, 100 μm and 20 μm, respectively.
(E) Immunofluorescence analysis of CHD2 expression in spermatogenic cells using mouse testicular cell smears. Scale bar, 5 μm.
(F) Model of CHD2 expression during spermatogenesis in mice.
Chromodomain helicase DNA-binding domain 2 haploinsufficiency causes compromised testicular germ cell proliferation and impaired fertility
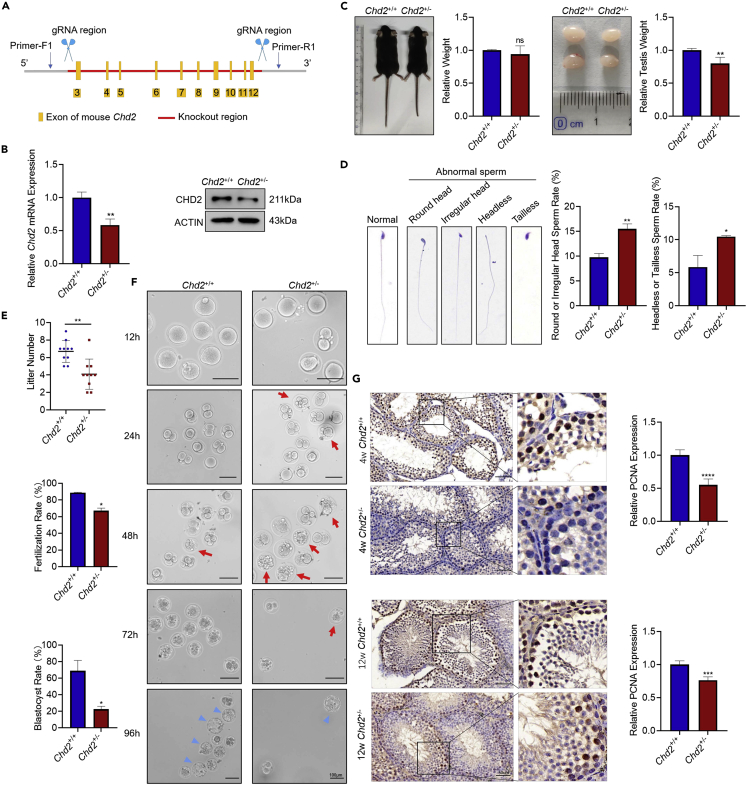
Previous studies have shown that homozygous knockout of Chd2 leads to perinatal lethality in mice. Even heterozygous knockout of Chd2 in mice still causes histological abnormalities in the kidney, spleen, heart, and liver, suggesting that CHD2 has important physiological functions. Clinically, CHD2 haploinsufficiency was observed in some patients with neurological diseases, implying that we could use heterozygous knockout of Chd2 (Chd2+/−) in mice to simulate the disease status related to CHD2 in humans. Here, using the CRISPR‒Cas9 system, male Chd2+/− mice were generated to observe the effect of Chd2 haploinsufficiency on spermatogenesis. Two fragments of gRNA were targeted to intron 2 and intron 13 of the Chd2 gene to knock out exons 3-12, which led to the frameshift mutation of Chd2, thus generating Chd2 haploinsufficiency mice (Figures 2A, S3A, and S3B). RT‒qPCR and western blot results showed an obvious decrease in CHD2 in the Chd2+/−mice (Figure 2B). Phenotypic analysis showed that the testicular weight of 4- and 12-week-old Chd2+/− mice was significantly lower than that of wild-type mice, although the body weights of these mice were similar (Figures 2C and S3C). In addition, compared with wild-type mice, the quantity of sperm was decreased, and the rates of abnormal sperm were increased in the epididymis of 12-week-old Chd2+/− mice (Figure S4A). There were four main types of abnormal sperm in the Chd2+/− mice, including round head, irregular head, headless and tailless sperm, and their rates were increased (Figure 2D). By counting the litter numbers and performing in vitro fertilization (IVF) experiments, male Chd2+/− mice showed smaller litters, a decreased fertilization rate and more degenerated embryos, which led to a reduced blastocyst rate (Figures 2E and 2F). This evidence suggests impaired fertility in the Chd2+/− mice.
Figure 2.
Chd2 haploinsufficiency causes compromised testicular germ cell proliferation and impaired fertility
(A) Diagram illustrating the CRISPR/Cas9 targeting strategy to generate Chd2+/− mice, including gRNA sites and PCR primer sites.
(B) The expression of CHD2 in testes of Chd2+/− mice by RT‒qPCR and western blot analysis. Data are shown as the mean ± SD. Student’s t-test, ∗∗p < 0.01.
(C) Relative body weight and testes volume analysis of 4w Chd2+/+ and Chd2+/− mice (n = 4). Data are shown as the mean ± SD, Student’s t-test; ∗∗p < 0.01; ns, no significance.
(D) Morphology analysis and rate of abnormal sperm in Chd2+/+ and Chd2+/− mice. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗p < 0.01. Scale bar, 50 μm.
(E) Litter number of male Chd2+/+ and Chd2+/− mice. Student’s t-test, ∗∗p < 0.01.
(F) The development of zygotes after in vitro fertilization experiments with Chd2+/+ and Chd2+/− mouse sperm. Images were taken 12, 24, 48, 72, and 96 h after fertilization. The red arrows represent degenerated embryos, and the blue triangles represent blastocysts. The fertilization rate and blastocyst rate are shown as the mean ± SD. Student’s t-test, ∗p < 0.05. Scale bar, 100 μm.
(G) Immunohistochemical detection of PCNA expression in testicular sections of 4- and 12-week-old Chd2+/+ and Chd2+/− mice. Data are shown as the mean ± SD. Student’s t-test, ∗∗∗p < 0.001, ∗∗∗∗p < 0.0001. Scale bar, 50 μm.
Hematoxylin and eosin staining of 4- and 12-week-old Chd2+/− and Chd2+/+ mouse testes showed no significant change in morphology (Figure S4B), and flow cytometry assays showed that the composition of different germ cells in Chd2+/− mice was not significantly changed. Only a slight decrease in diploid cells was found in the Chd2+/− mice (Figure S4C). To explore why the testes are smaller in the Chd2+/− mice, we performed the immunohistochemical analysis of PCNA protein to detect cell proliferation in the Chd2+/− and Chd2+/+ mouse testes. PCNA expression was downregulated in the Chd2+/− mice (Figure 2G). However, TdT-mediated dUTP nick-end labeling (TUNEL) staining showed no significant change in cell apoptosis (Figure S4D), which indicates that cell proliferation rather than apoptosis caused the smaller testes in the Chd2+/− mice.
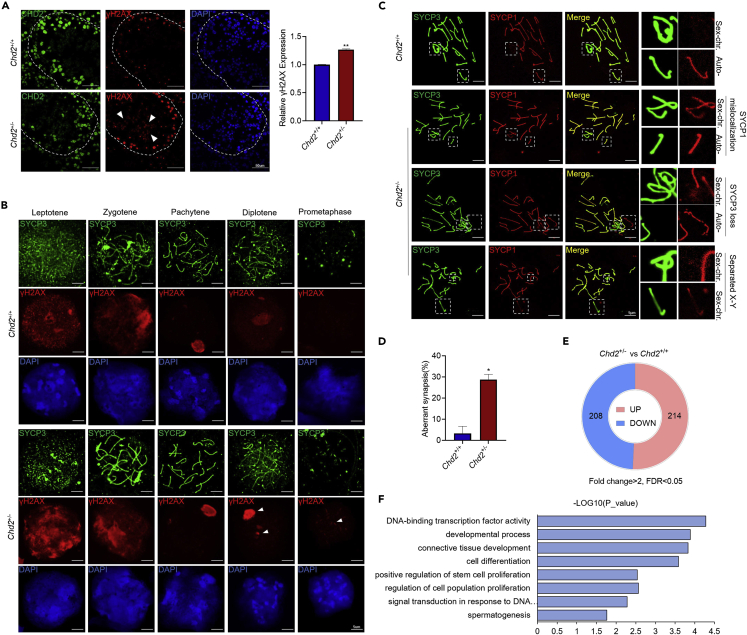
During meiosis, homologous chromosome pairing, recombination, and separation occur in meiosis I, which undergoes a large number of DNA double-strand breaks (DSBs) and repairs, followed by cell division in meiosis II.34 It has been reported that from mid-pachytene to diplotene, DSB repair is mainly mediated by nonhomologous end joining (NHEJ) and comprises Ku70 and XRCC4 to repair DSB on the autosomes and 53BP1 to repair DSB on the XY chromosomes.35,36 In meiosis I spermatocytes, the phosphorylated form of histone variant H2AX (γH2AX) marks sites of DNA damage throughout the nucleus, and as DNA is repaired, it remains on the sex chromosomes.37 Because CHD2 is highly expressed in meiosis I spermatocytes and previous studies have shown that CHD2 is involved in NHEJ repair to promote DSB repair,28 we wanted to know whether Chd2 haploinsufficiency would cause impaired DNA damage repair during mouse spermatogenesis. Immunofluorescence staining showed increased γH2AX expression in the testes of 12-week-old Chd2+/− mice compared with Chd2+/+ mice. In addition, more residual γH2AX in spermatids was observed in the Chd2+/− mice (Figure 3A). In testicular cell smears, abnormal γH2AX residues were also observed in diplotene and prometaphase spermatocytes (Figure 3B). The western blot results confirmed the overexpression of γH2AX in Chd2+/− mouse testes (Figure S4E). Along with impaired DNA damage repair, the synapsis of meiosis may be affected. SYCP1 and SYCP3 are key components of the synaptic complex and are often used as synapsis markers. Further immunofluorescence staining of SYCP1 and SYCP3 in testicular cell smears showed more abnormal chromosomal synapsis in pachytene spermatocytes of Chd2+/− mice, including the mislocalization of SYCP1 to unsynapsed sex chromosomes and loss of SYCP3 on autosomal and separated X-Y chromosomes (Figures 3C and 3D).38 Such defective synapses may affect homologous recombination in spermatogenesis. Interestingly, Chd1, a member of the Chd1-Chd2 subfamily, has been found to reduce histone occupancy near the DSB ends and promotes DSB repair by homologous recombination.39,40 The protein domains of CHD2 and CHD1 are similar. We hypothesized that CHD2, like CHD1, may promote homologous recombination repair by increasing chromatin accessibility. The mechanism of how CHD2 affects homologous recombination during meiosis needs further study.
Figure 3.
Impaired DNA damage repair, abnormal synapses, and multigene expression changes in Chd2+/− mouse testes
(A) Expression analysis of γH2AX in frozen sections of mouse testes by immunofluorescence assay. Triangles represent the abnormal expression of γH2AX. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05. Scale bar, 50 μm.
(B) Expression analysis of γH2AX in meiosis by immunofluorescence assay. Scale bar, 5 μm.
(C) Co-immunofluorescence staining of SYCP3 and SYCP1 in spermatocytes at pachytene from Chd2+/+ and Chd2+/− male mice. Scale bar, 5 μm.
(D) The aberrant synapsis rate of Chd2+/+ and Chd2+/− male mice. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05.
(E) Global gene expression alterations with fold changes of ≥2 or ≤0.5 (p ≤ 0.05) in Chd2+/− versus Chd2+/+ testes at 12 w.
(F) Gene ontology analysis of differentially expressed genes identified by RNA-seq in Chd2+/− testes.
Subsequently, differentially expressed genes in testicular tissues of Chd2+/+ and Chd2+/− mice were detected by transcriptome sequencing. Compared with Chd2+/+ mice, the mRNA levels of 422 genes were significantly changed in Chd2+/− mice, among which 208 genes were downregulated and 214 genes were upregulated (Figure 3E). Gene Ontology (GO) analysis revealed that differentially expressed genes are associated with DNA-binding transcription factor activity, development process, stem cell proliferation, and other functions (Figure 3F). We found that self-renewal-associated genes of spermatogonia, such as Plzf (Zbtb16) and Sall4, and progenitor-associated gene Sox3 were downregulated (Figure S5). In summary, these results show that Chd2 haploinsufficiency causes impaired DNA damage repair during meiosis and an increased abnormal sperm rate. Downregulation of self-renewal-associated and progenitor-associated genes may contribute to the inhibition of spermatogonial self-renewal in testes. All of these phenotypes led to compromised testicular germ cell proliferation and impaired fertility in the Chd2+/− mice.
Chromodomain helicase DNA-binding domain 2 knockdown inhibits spermatogonial self-renewal
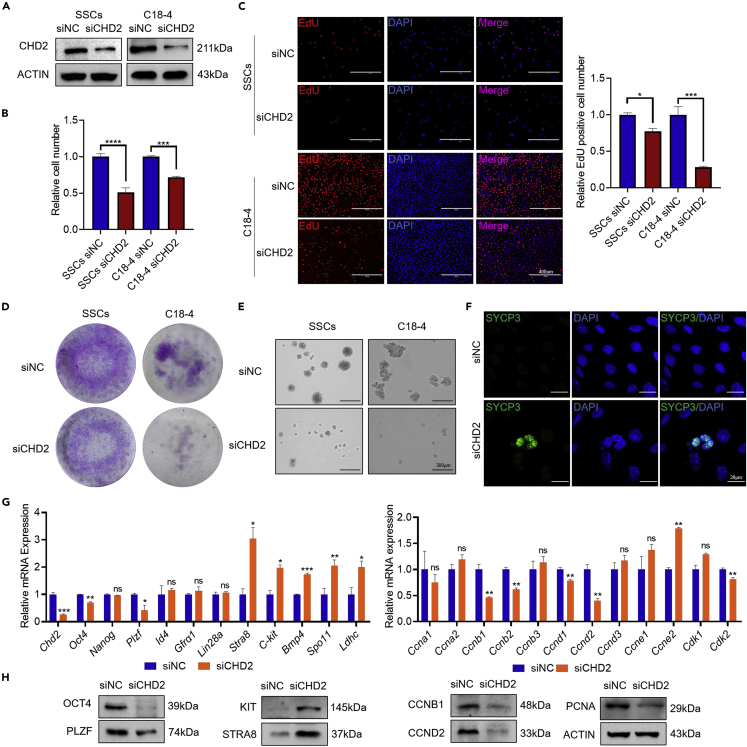
Spermatogonia in testes not only differentiate and undergo whole spermatogenesis to form mature sperm but also maintain the spermatogonia number through continuous self-renewal. As shown in Figures 2C and 2G, Chd2+/− mice exhibited smaller testes and downregulated PCNA expression, suggesting that this abnormal reproductive phenotype may be due to the inhibition of spermatogonial proliferation. Here, in vitro experiments in SSCs and a spermatogonial cell line (C18-4)41 were performed to investigate the mechanism of CHD2 in spermatogonial self-renewal. SSCs were identified by immunofluorescence staining of PLZF and cultured on laminin-coated plates (Figure S6A). The western blot results showed that CHD2 was highly expressed in both C18-4 and GC-1 cells (mouse type B spermatogonial cells), whereas it was weakly expressed in GC-2 spermatocytal cells (Figure S6B). SSCs, C18-4, and GC-1 were chosen for subsequent experiments. After the knockdown of CHD2 by siRNA in SSCs, C18-4 cells (Figure 4A), and GC-1 cells (Figure S6C), the results of the Cell Counting Kit-8 (CCK8) assay (Figures 4B and S6D) and 5-ethynyl-2′-deoxyuridine (EdU) assay (Figures 4C and S6E) showed that the proliferation of germ cells was inhibited and the number of EdU-positive cells decreased. Knockdown of CHD2 significantly reduced the colony-formation ability of SSCs and C18-4 (Figure 4D) and GC-1 cells (Figure S6F). Then, we detected the stemness of SSCs, C18-4 cells (Figure 4E), and GC-1 cells (Figure S6G) by sphere-formation assay. Some SSCs differentiated into SYCP3-positive cells with CHD2 knockdown (Figure 4F). These results suggest that CHD2 knockdown inhibits the self-renewal ability of spermatogonia. Subsequently, six stemness-associated genes (Oct4, Nanog, Plzf, Id4, Gfrα1, Lin28a), five differentiation-related genes (Stra8, C-kit, Bmp4, Spo11, Ldhc), and 12 cell cycle-related genes that are closely associated with spermatogonial self-renewal were detected by RT‒qPCR and western blotting. CHD2 knockdown upregulated the expression of the Stra8 and C-kit genes and inhibited the expression of the Oct4, Plzf, Ccnb1, and Ccnd2 genes (Figures 4G and 4H). These data indicate that CHD2 may regulate the expression of target genes to maintain spermatogonial self-renewal.
Figure 4.
Knockdown of CHD2 inhibits spermatogonial self-renewal
(A) Detection of RNA interference efficiency for siCHD2 by western blot in SSCs and C18-4 cells.
(B–D) Cellular proliferation in SSCs and C18-4 cells with CHD2 knockdown analyzed by CCK8, EdU, and colony-formation assays. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗∗p < 0.001. Scale bar, 400 μm.
(E) The impacts of CHD2 knockdown on the sphere-formation ability of SSCs and C18-4 cells. Scale bar, 300 μm.
(F) Immunofluorescence staining of SYCP3 after CHD2 knockdown in SSCs. Scale bar, 20 μm.
(G and H) Expression changes of stem-associated genes, differentiation genes, and cell cycle genes in C18-4 cells with CHD2 knockdown determined by RT‒qPCR and western blot assays. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001; ns, no significance.
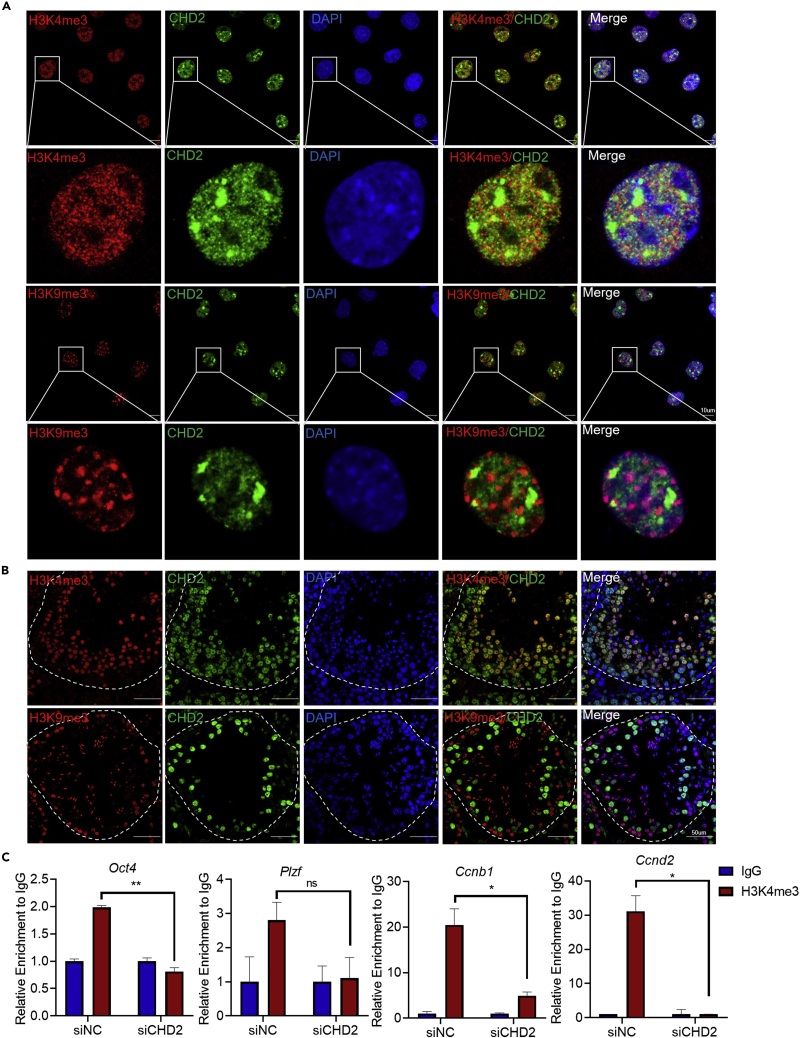
Chromodomain helicase DNA-binding domain 2 promotes the enrichment of H3K4me3 in the promotor region of target genes
Altering chromatin accessibility to affect gene expression is one of the regulatory modes of chromatin remodeling complexes, including CHD2. Using H3K4me3 as the marker of active chromatin and H3K9me3 as the marker of heterochromatin,42 we conducted immunofluorescence experiments in C18-4 cells and found that CHD2 was colocalized with H3K4me3 rather than H3K9me3 (Figure 5A). In mouse testicular sections, most CHD2 colocalized with H3K4me3 but separated from H3K9me3 (Figure 5B), indicating that in germ cells, CHD2 may be associated with an active chromatin status to promote the transcriptional activation of target genes. Furthermore, the chromatin immunoprecipitation (ChIP) and qPCR results showed that after CHD2 knockdown in C18-4 cells, H3K4me3 enrichment decreased in the promotor regions of the Oct4, Ccnb1, and Ccnd2 genes but not Plzf, which might be regulated by CHD2 in another way (Figure 5C). In Chd2+/− mouse testes, the downregulation of the Oct4, Plzf, Ccnb1, and Ccnd2 genes was also detected by RT‒qPCR and western blotting (Figures S7A and S7B). In addition, both the immunofluorescence staining of testes sections of Chd2+/− mice (Figure S7C) and western blot analysis in C18-4 cells (Figure S7D) showed that CHD2 knockdown also inhibited the active chromatin state, which indicates that CHD2 may regulate the whole landscape of chromatin, not only the selected genes. Therefore, we speculate that CHD2 can maintain the active chromatin state of the Ccnb1 and Ccnd2 promotor regions to promote their expression.
Figure 5.
CHD2 maintains an active chromatin state in the promotor of target genes
(A and B) The colocalization analysis of CHD2 and H3K4me3 or H3K9me3 in C18-4 cells (A) and sections of testes (B) by immunofluorescence assay. Scale bars, 10 μm and 50 μm, respectively.
(C) ChIP‒qPCR assay to determine the relative enrichment of H3K4me3 or H3K9me3 in the promotor regions of the Oct4, Plzf, Ccnb1, and Ccnd2 genes in C18-4 cells with CHD2 knockdown. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗p < 0.01; ns, no significance.
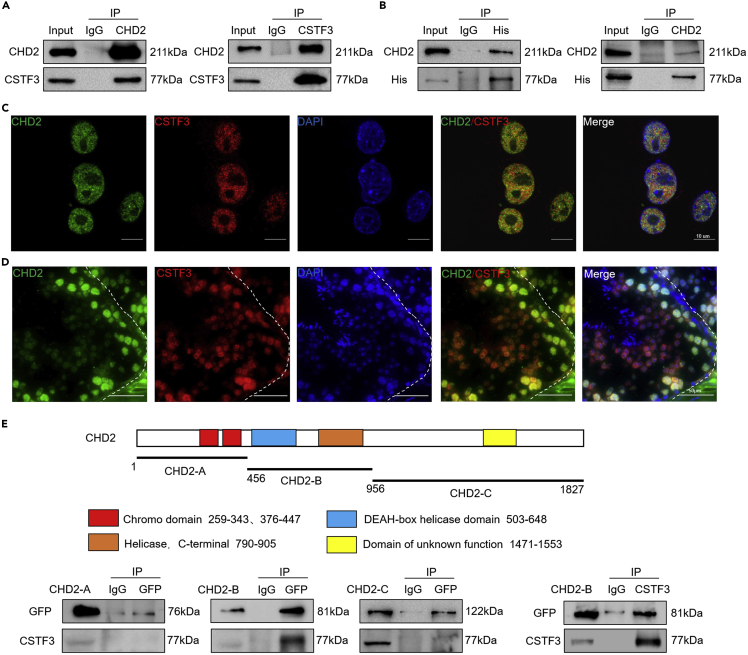
Chromodomain helicase DNA-binding domain 2 interacts with cleavage stimulation factor subunit 3
Immunoprecipitation combined with LC‒MS mass spectrometry was performed to identify the interacting proteins of CHD2 in GC-1 cells (Table S1). Among these proteins, cleavage stimulation factor subunit 3 (CSTF3) was ranked first. Together, CSTF3, CSTF1, and CSTF2 constitute the cleavage stimulation factor (CSTF) complex.43 The CSTF complex is involved in cleavage and polyadenylation (C/P) of nascent mRNA, and cleavage and polyadenylation of nascent transcripts are essential for the maturation of the 3′ ends of most eukaryotic mRNAs.44 CSTF3 acts as a scaffold linking CSTF2 and CSTF1, and CSTF3 is also necessary for the transport of the CSTF complex into the nucleus.45 It has been reported that CSTF3 is important for the expression of cell cycle genes, and downregulation of CSTF3 may help halt proliferation and launch differentiation in C2C12 cells.46 Therefore, we hypothesized that CSTF3 has a similar function in spermatogonial stem cells. In the present study, the binding between CHD2 and CSTF3 was verified by immunoprecipitation experiments in SSCs and C18-4 cells (Figure 6A and S8A). Furthermore, the CDS of the Cstf3 gene linked with the His tag sequence was cloned into the pCD513B-1 plasmid. After overexpression of CSTF3 protein with a His tag (CSTF3-HIS) in 293T cells, we proved the mutual binding between CHD2 and CSTF3 by immunoprecipitation (Figure 6B). Immunofluorescence analysis in C18-4 cells (Figure 6C) and testicular sections (Figure 6D) confirmed the colocalization of CHD2 and CSTF3 in the nucleus. Because CSTF3 usually interacts with CSTF1 and CSTF2 to form the cleavage stimulation factor complex and is involved in the polyadenylation and 3′ end cleavage of pre-mRNAs,47 the binding between CHD2 and CSTF1 and CSTF2 in C18-4 cells was also detected. It was shown that the interaction between CHD2 and CSTF2 was detectable, but that between CHD2 and CSTF1 was not (Figure S8B). These results suggest the function of CHD2 in regulating the mRNA process in spermatogonial cells.
Figure 6.
CHD2 interacts with CSTF3
(A) Immunoprecipitation assay in SSCs to detect the interaction between CHD2 and CSTF3.
(B) Immunoprecipitation detection of the interaction between CHD2 and CSTF3-HIS after overexpression of CSTF3-HIS in 293T cells.
(C and D) Immunofluorescence detection of the colocalization of CHD2 and CSTF3 in C18-4 cells (C) and sections of mouse testes (D). Scale bars, 10 μm and 50 μm, respectively.
(E) Schema diagram of the CHD2 protein domains. Immunoprecipitation assay with GFP antibody to detect the interaction between CHD2-A, CHD2-B, CHD2-C, and CSTF3 (left and middle). Immunoprecipitation assay with CSTF3 antibody to detect the interaction between CSTF3 and CHD2-B (right).
CHD2 contains N-terminal tandem chromodomains, SNF2-like ATPase domains located in the central region, and DNA-binding domains in the C-terminal region.16 To identify which domain of CHD2 binds to CSTF3, three segments, CHD2-A (CHD21−455), CHD2-B (CHD2456−955) and CHD2-C (CHD2955−1827), were cloned and inserted into the pEFGP-C2 plasmid to express fusion proteins with a GFP tag. After being overexpressed in 293T cells, immunoprecipitation analysis performed with GFP and CSTF3 antibodies showed that CSTF3 interacts with CHD2-B, which contains the SNF2-like ATPase domain of CHD2 (Figure 6E). These results suggest that CHD2 interacts with CSTF3 via the SNF2-like ATPase domain.
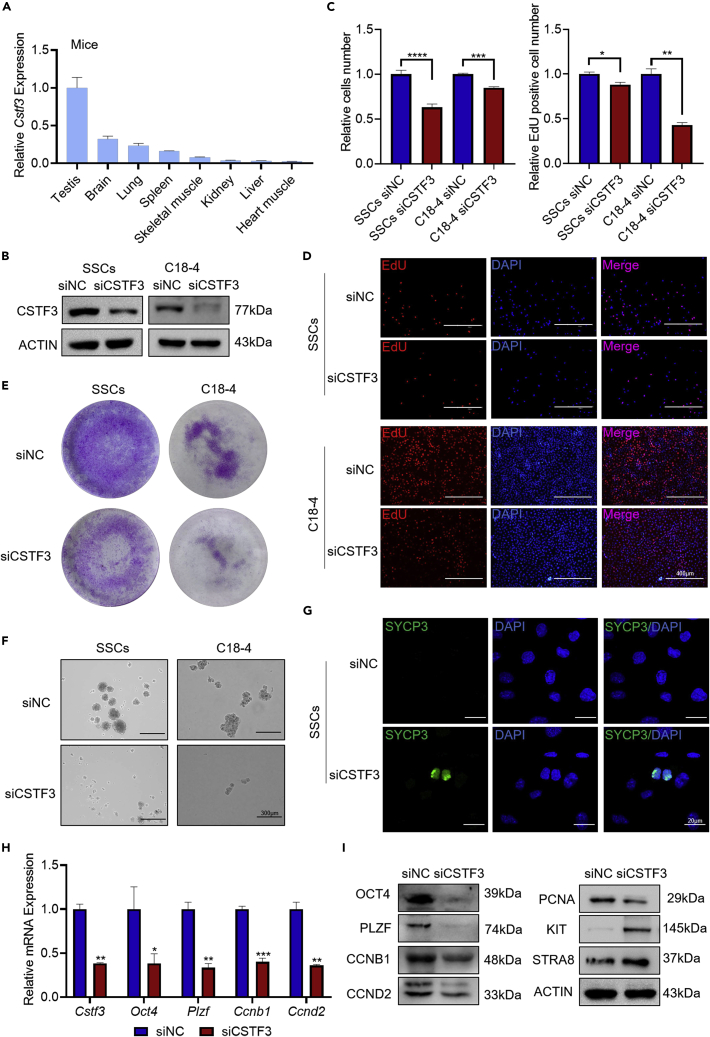
Knockdown of cleavage stimulation factor subunit 3 inhibits spermatogonial self-renewal
Considering the interaction between CSTF3 and CHD2, we hypothesized that CSTF3 might play a similar role in spermatogonia. The HPA database shows that CSTF3 is also highly expressed in testes in humans (Figure S8C), and the highest expression of Cstf3 mRNA was observed in mouse testes (Figure 7A). CSTF3 was more highly expressed in C18-4 and GC-1 cells than in GC-2 cells (Figure S8D). In SSCs, C18-4 (Figure 7B) and GC-1 cells (Figure S8E) treated with siCSTF3, the CCK8 (Figures 7C and S8F) and EdU assay data (Figures 7D and S8G) showed that cellular proliferation was inhibited and the number of EdU-positive cells decreased. Knockdown of CSTF3 also reduced the colony-formation ability of SSCs, C18-4 cells (Figure 7E) and GC-1 cells (Figure S8H). Then, we detected the stemness of SSCs, C18-4 cells (Figure 7F), and GC-1 cells (Figure S8I) by sphere-formation assay. Some SSCs differentiated into SYCP3-positive cells with CSTF3 knockdown (Figure 7G). In addition, the RT‒qPCR and western blot results confirmed that CSTF3 knockdown inhibited the expression of the Oct4, Plzf, Ccnb1, Ccnd2, and Chd2 genes (Figures 7H, 7I, and S8J). These data indicate that both CHD2 and CSTF3 are involved in maintaining spermatogonial self-renewal.
Figure 7.
Knockdown of CSTF3 inhibits spermatogonial self-renewal
(A) Cstf3 mRNA expression in mouse tissues by RT‒qPCR. Data are shown as the mean ± SD.
(B) CSTF3 expression analysis by western blot after CSTF3 knockdown in SSCs and C18-4 cells.
(C–E) Cell proliferation analysis of SSCs and C18-4 cells with CSTF3 knockdown determined by CCK8, EdU, and colony-formation assays. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, ∗∗∗∗p < 0.0001. Scale bar, 400 μm.
(F) The impacts of CSTF3 knockdown on the sphere-formation ability of SSCs and C18-4 cells. Scale bar, 300 μm.
(G) Immunofluorescence staining of SYCP3 after CSTF3 knockdown in SSCs. Scale bar, 20 μm.
(H and I) RT‒qPCR and western blot assays of the expression changes in the Oct4, Plzf, Ccnb1, and Ccnd2 genes in C18-4 cells. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001.
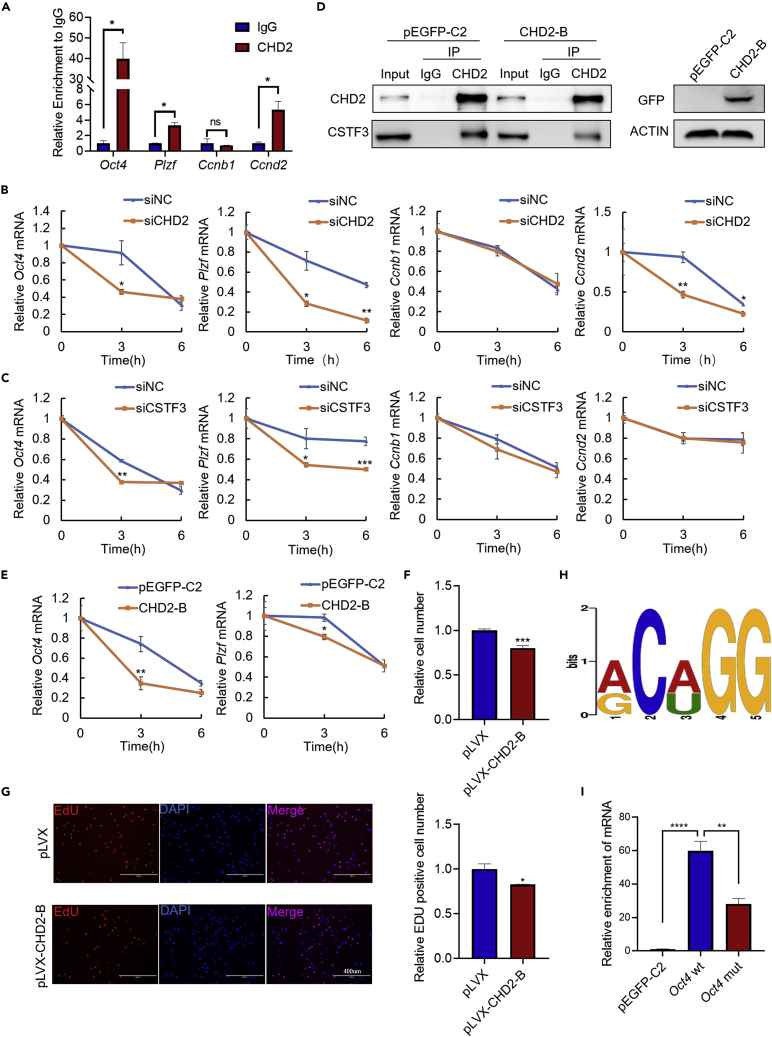
Chromodomain helicase DNA-binding domain 2 interacts with cleavage stimulation factor subunit 3 to promote the mRNA stability of target genes
In mouse embryonic fibroblasts, a recent study performed using an RNA immunoprecipitation (RIP) assay with CHD2 antibody followed by rRNA-depleted RNA sequencing reported that CHD2 may bind mRNA.48 Given that CSTF3, the interacting protein of CHD2, can affect the mRNA stability of target genes,49 we wondered whether CHD2 was involved in the regulation of mRNA stability.
Here, RIP-qPCR analysis in mouse testes confirmed the binding of CHD2 to Oct4, Plzf, and Ccnd2 mRNA but not Ccnb1 (Figure 8A). In addition, in C18-4 cells with CHD2 knockdown, the mRNA stability of Oct4, Plzf, and Ccnd2 but not Ccnb1 decreased after treatment with actinomycin D for 0, 3, and 6 h (Figure 8B). After the knockdown of CSTF3, the mRNA stability of Oct4 and Plzf also decreased (Figure 8C). Moreover, in an immunoprecipitation assay of NIH3T3 cells overexpressing CHD2-B, we confirmed that exogenous CHD2-B could compete with endogenous CHD2 to bind CSTF3 (Figure 8D). Moreover, the RT‒qPCR results showed that the mRNA stability of Oct4 and Plzf also decreased in NIH3T3 cells transfected with CHD2-B, indicating that the interaction between CHD2 and CSTF3 could affect the mRNA stability of Oct4 and Plzf (Figure 8E). Then, we cloned the CHD2-B (CHD2456−955) fragment into the pLVX-IRES-Puro-GFP lentivirus plasmid to generate pLVX-CHD2-B recombinants. In lentivirus pLVX-CHD2-B-infected C18-4 cells, the CCK8 (Figure 8F) and EdU assay (Figure 8G) results confirmed that the overexpression of CHD2-B inhibited C18-4 cell proliferation.
Figure 8.
The interaction between CHD2 and CSTF3 promotes the mRNA stability of target genes
(A) RIP-qPCR detection of the mRNA binding between CHD2 and the Oct4, Plzf, Ccnb1, and Ccnd2 genes in mouse testes. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05; ns, no significance.
(B and C) Effects of CHD2 or CSTF3 knockdown on the mRNA stability of Oct4, Plzf, Ccnb1, and Ccnd2 in C18-4 cells. C18-4 cells were treated with 10 μM actinomycin D for 0, 3, and 6 h. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001.
(D) Interaction analysis between CHD2 and CSTF3 after CHD2-B overexpression in NIH3T3 cells.
(E) Effect of CHD2-B overexpression on the mRNA stability of Oct4 and Plzf in NIH3T3 cells. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗p < 0.01.
(F and G) CCK8 and EdU assays of the proliferation capacity of C18-4 cells with pLVX-CHD2-B overexpression. pLVX represents the pLVX-IRES-Puro-GFP plasmid. Scale bar, 400 μm. Data are shown as the mean ± SD. Student’s t-test, ∗p < 0.05, ∗∗∗p < 0.001.
(H) The mRNA binding motif of CHD2.
(I) RIP-qPCR detection of the mRNA binding between CHD2 and pEGFP-C2, Oct4 wt, Oct4 mut. Data are shown as the mean ± SD. Student’s t-test, ∗∗p < 0.01, ∗∗∗∗p < 0.0001.
Finally, by using the RIP-seq data of CHD2 in the GEO datasets (GSE124375) and analyzing the possible binding motif of CHD2 with MEME Suite (https://meme-suite.org/meme/), the binding motif with the smallest p value was predicted to be ACAGG (Figure 8H). To verify whether CHD2 regulates target mRNAs via ACAGG sites, we searched the mRNA sequences of Oct4 and Plzf and found two possible sites for CHD2 binding on Oct4 mRNA and fifteen on Plzf mRNA. Then, the Oct4 gene was selected for further validation. We constructed a part of the Oct4 sequence containing the two possible binding sites (Oct4 wt) or mutations of the two sites (Oct4 mut) in the pEFGP-C2 plasmid. RIP-qPCR assays were performed after the transfection of pEGFP-C2, Oct4 wt or Oct4 mut plasmids into 293T cells. The results showed that the binding of CHD2 to Oct4 mut was weaker than that of CHD2 to Oct4 wt but stronger than CHD2 to pEGFP-C2 (Figure 8I). This means that the mutation of the binding site can indeed weaken the binding of CHD2 to mRNA. After the transfection of Oct4 wt or Oct4 mut plasmid into NIH3T3 cells, we found that CHD2 knockdown weakened the mRNA stability of Oct4 wt. Mutation of the binding site caused less CHD2 binding, and CHD2 knockdown led to even less CHD2 binding. Thus, the mRNA stability of Oct4 mutation worsened with CHD2 knockdown (Figure S8K).
Discussion
In the present study, we showed that Chd2 haploinsufficiency causes impaired fertility, testicular growth retardation, and impaired DNA damage repair in Chd2+/− mice. In vitro experiments in SSCs and C18-4 and GC-1 cells confirmed that the knockdown of CHD2 inhibits the self-renewal ability of spermatogonia. By detecting the expression of stemness-associated genes and cell cycle-related genes, the downregulated expression of OCT4, PLZF, CCNB1, and CCND2 was observed. Mechanistically, CHD2 promotes the expression of CCNB1 and CCND2 by maintaining H3K4me3 enrichment in the promotor regions of the Ccnb1 and Ccnd2 genes. Moreover, CSTF3 was identified as an interacting protein of CHD2, and by interacting with CSTF3, CHD2 upregulates the expression of OCT4 and PLZF by binding Oct4 and Plzf mRNA and improving the mRNA stability. Our findings verify the important role of CHD2 in maintaining spermatogonial self-renewal in spermatogenesis (graphical abstract).
In the testis, transcription of the genome is more widespread than in other organs, and male germ cells have the largest number of tissue-specific RNA binding proteins.50 RNA binding proteins sequester, modify, translate and degrade individual mRNAs to ensure smooth transitions from one stage of spermatogenesis to the next.51,52 A study in mouse embryonic fibroblasts based on RIP-seq with CHD2 antibody suggested that CHD2 might play the role of an RNA-binding protein,48 but how CHD2 affects mRNA during spermatogenesis was not clear. In the present study, CSTF3 was identified as an interacting protein of CHD2 in spermatogonia. As a subunit of the cleavage stimulation factor (CSTF) complex, CSTF3 has an N-terminal HAT domain and a proline-rich segment, through which CSTF3 interacts with CSTF1 and CSTF2 to form a heterotrimeric.53 The function of the CSTF complex is to define polyadenylation by recognizing G/U-rich sequences located downstream of the 3′ end cleavage site in pre-mRNA.54 To date, only CSTF2T, a paralog gene of CSTF2, in the CSTF complex has been reported to be involved in spermatogenesis. Cstf2t knockout mice showed abnormal sperm morphology and reduced sperm motility, which led to male infertility.55,56 Our study showed that CSTF3 is highly expressed in human and mouse testes. Similar to CHD2, CSTF3 is related to spermatogonial proliferation and differentiation, which expands our understanding of the CSTF complex in spermatogenesis. More interestingly, the interaction between CHD2 and CSTF3 suggests a novel role of CHD2 in regulating RNA processing, such as mRNA stability. As a chromodomain helicase DNA binding protein, the function of CHD2 in chromatin remodeling and its interaction with transcription factors such as OCT4 and histone and histone variants have been discussed in previous studies.24 Here, we hypothesized that the role of CHD2 in PLZF expression may be the regulation of Plzf mRNA stability. Through an mRNA stability assay, we confirmed that CHD2 can promote the mRNA stability of Oct4 and Plzf, which may contribute to the role of CHD2 in maintaining spermatogonial self-renewal in spermatogenesis. In addition, CHD2-B, which contains the SNF2-like ATPase domain of CHD2, was identified to interact with CSTF3. The SNF2-like ATPase domain contains a conserved set of amino acid motifs that have been found in proteins involved in a myriad of cellular processes, including chromatin assembly, DNA replication, DNA repair, transcription regulation, development, and differentiation.57,58 Whether the interaction between CHD2 and CSTF3 is involved in other cellular processes requires further investigation in future studies.
We also found that Chd2 haploinsufficiency in mice resulted in increased DNA damage in spermatogenesis. Increased γH2AX deposition was observed in diplotene and prometaphase spermatocytes and round sperm cells in Chd2+/− mice, and the enhanced γH2AX signal in leptotene and zygotene may explain the increase in γH2AX. CHD2 has been shown to be involved in NHEJ repair of DSBs28; thus, more DNA damage in Chd2+/− pachytene and subsequent spermatogenesis may be caused by defects in NHEJ repair. This may also be the reason for increased abnormal sperm in the Chd2+/− mice.
In conclusion, our study is the first to reveal the important role of CHD2 in spermatogenesis. We confirmed that CHD2 promotes spermatogonial self-renewal by regulating chromatin accessibility and mRNA stability. This study provides a new perspective to understand the role of chromatin remodeling and cleavage stimulation factor complexes in spermatogenesis and provides a potential target and molecular basis for the diagnosis and treatment of male infertility.
Limitations of the study
Our study found that CHD2 maintains spermatogonial self-renewal by promoting chromatin accessibility and mRNA stability. Several issues remain; for example, conditional knockout of the Chd2 gene in male germ cells would be a better model to investigate the function of CHD2 in germ cells. Given that CHD2 is also expressed in Sertoli cells and Leydig cells and that the testis microenvironment is important for spermatogenesis, further work is needed to investigate whether CHD2 regulates the testis microenvironment to affect spermatogenesis. Whether the nongonadal phenotypes of Chd2 haploinsufficiency affect spermatogenesis requires further investigation, which may help find connections between spermatogenesis and other organs.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Rat monoclonal anti-CHD2 | Sigma-Aldrich | Cat# MABE873 Lot: 3407335 RRID:AB_2737345 |
| Rabbit polyclonal anti-CHD2 | ABclonal | Cat# A5895 Lot: 0203250201 RRID: AB_2766643 |
| Mouse monoclonal anti-DDX4 | Proteintech | Cat# 67147-2-Ig Lot:10009967 RRID:AB_2882446 |
| Rabbit polyclonal anti-SYCP3 | Proteintech | Cat# 23024-1-AP Lot:00091736 RRID:AB_11232426 |
| Mouse monoclonal anti-SYCP3 | Santa cruz | Cat# sc-74569 Lot: G1521 RRID:AB_2197353 |
| Rabbit monoclonal anti-SYCP1 | Abcam | Cat# ab175191 Lot: GR135907-7 RRID: N/A |
| Rabbit polyclonal anti-β-ACTIN | Bioworld | Cat# AP0060 Lot: AA1217J1199ZJ1 RRID:AB_2797445 |
| Mouse monoclonal anti-PLZF | Proteintech | Cat# 66672-1-Ig Lot:10006043 RRID:AB_2882026 |
| Rabbit polyclonal anti-PCNA | Cohesion Biosciences | Cat# CPA1859 Lot: CV0298A RRID: N/A |
| Mouse monoclonal anti-γH2AX | ZEN-BIOSCIENCE | Cat# 201082-7G9 RRID:AB_2722720 |
| Rabbit polyclonal anti-OCT4 | Cohesion Biosciences | Cat# CPA1915 Lot: CV0283A RRID: N/A |
| Rabbit polyclonal anti-CCNB1 | Proteintech | Cat# 28603-1-AP Lot:00073891 RRID:AB_2881179 |
| Rabbit polyclonal anti-CCND2 | Proteintech | Cat# 10934-1-AP RRID:AB_2275319 |
| Rabbit polyclonal anti-KIT | Wanleibio | Cat# WL00125 Lot:H10100125 RRID: N/A |
| Rabbit polyclonal anti-STRA8 | CUSABIO | Cat# CSB-PA767212LA01HU Lot: G1020A RRID: N/A |
| Rabbit polyclonal anti-H3K4me3 | Beyotime | Cat# AF5704 RRID: N/A |
| Rabbit polyclonal anti-H3K9me3 | Beyotime | Cat# AF5707 RRID: N/A |
| Mouse monoclonal anti-CSTF3 | Santa cruz | Cat# sc-376553 Lot: B1914 RRID:AB_11149910 |
| Rabbit polyclonal anti-CSTF3 | Proteintech | Cat# 24290-1-AP Lot:00020373 RRID:AB_2879475 |
| Mouse monoclonal anti-His | Sangon Biotech | Cat# D191001 Lot: D191001-0025 RRID: N/A |
| Rabbit Monoclonal anti-GFP | Beyotime | Cat# AF1483 RRID: N/A |
| Rabbit polyclonal anti-CSTF1 | Proteintech | Cat# 10064-2-AP Lot:00000083 RRID:AB_2261145 |
| Rabbit polyclonal anti-CSTF2 | Proteintech | Cat# 26825-1-AP Lot:00047950 RRID:AB_2880649 |
| Mouse monoclonal anti-SOX9 | Thermo Fisher Scientific | Cat# 14-9765-80 Lot:2171014 RRID:AB_2573005 |
| Rabbit polyclonal anti-CYP11A1 | Beyotime | Cat# AF6636 RRID: N/A |
| Mouse IgG | Beyotime | Cat# A7028 RRID:AB_2909433 |
| Rabbit IgG | Beyotime | Cat# A7016 RRID:AB_2905533 |
| Rat IgG | Beyotime | Cat# A7031 RRID: N/A |
| HRP-labeled Goat Anti-Mouse IgG(H + L) | Beyotime | Cat# A0216 RRID:AB_2860575 |
| HRP-labeled Goat Anti-Rabbit IgG(H + L) | Beyotime | Cat# A0208 RRID:AB_2892644 |
| CoraLite594 – conjugated Goat Anti-Mouse IgG(H + L) | Proteintech | Cat# SA00013-3 Lot:2000200 RRID:AB_2797133 |
| CoraLite488-conjugated Goat Anti-Rabbit IgG(H + L) | Proteintech | Cat# SA00013-2 Lot:20000236 RRID:AB_2797132 |
| CoraLite594 – conjugated Goat Anti-Rabbit IgG(H + L) | Proteintech | Cat# SA00013-4 Lot:20000239 RRID:AB_2810984 |
| Alexa Fluor 488-conjugated Goat anti-rat IgG | Sangon Biotech | Cat# D110268 RRID: N/A |
| Cy3-labeled Goat Anti-Rabbit IgG (H + L) | Beyotime | Cat# A0516 Lot:021622220711 RRID:AB_2893015 |
| Alexa Fluor 488-labeled Goat Anti-Mouse IgG(H + L) | Beyotime | Cat# A0428 Lot:102221220614 RRID:AB_2893435 |
| Cy3-labeled Goat Anti-Rat IgG(H + L) | Beyotime | Cat# A0507 Lot:102021220607 RRID: N/A |
| Biological samples | ||
| C57BL/6J-Chd2em1Cya | Cyagen | Cat# S-KO-07492 |
| Chemicals, peptides, and recombinant proteins | ||
| Gelatin | Sangon Biotech | Cat# A609764 |
| mouse spermatogonial stem cell medium | Procell Life Science&Technology | Cat# CM-M160 |
| pregnant mare serum gonadotropin | Nanjing Aibei Biotechnology | Cat# M2620 |
| human chorionic gonadotropin | Nanjing Aibei Biotechnology | Cat# M2520 |
| HTF solution | Nanjing Aibei Biotechnology | Cat# M1130 |
| KSOM embryo culture medium | Nanjing Aibei Biotechnology | Cat# M1430 |
| Laminin Mouse Protein, Natural | Thermo Fisher Scientific | Cat# 23017015 |
| Actinomycin D | AbMole | Cat# M4881 |
| Critical commercial assays | ||
| Cell Counting Kit-8 | TOPSCIENCE | Cat# C0005 |
| BeyoClick™ EdU Cell Proliferation Kit with Alexa Fluor 555 | Beyotime | Cat# C0075S |
| Hoechst 33342/PI Double Stain Kit | Solarbio Life Science | Cat# CA1120 |
| One Step TUNEL Apoptosis Assay Kit | Beyotime | Cat# C1089 |
| Hematoxylin-Eosin/HE Staining Kit | Solarbio Life Science | Cat# G1120 |
| Deposited data | ||
| RNA-seq of Chd2+/+ and Chd2+/− | This paper |
GSE198581https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE198581 Enter token: ovmvaweozrmbjsd |
| LC-MS mass spectrometry result of CHD2 | This paper | ProteomeXchange: PXD032277 Username:reviewer_pxd032277@ebi.ac.uk Password: YlPMRuEC |
| Experimental models: Cell lines | ||
| GC-1 spg | ATCC | CRL-2053 |
| GC-2 spd | ATCC | CRL-2196 |
| NIH3T3 | ATCC | CRL-1658 |
| HEK293T cells | ATCC | ACS-4500 |
| C18-4 cells | Marie-Claude Hofmann et al., 2005 | N/A |
| Oligonucleotides | ||
| siCHD2 GAUGGCAAGUCUUCAAGUA | Dharmacon | Cat# D-060656-03 |
| siCSTF3 CGGAGCAGGCAGCGGAAUA | Dharmacon | Cat# D-051652-17 |
| Primers of this paper: Table S2 | This paper | N/A |
| Recombinant DNA | ||
| pEGFP-C2 | MiaoLing Plasmid Platform | Cat# P0135 |
| pLVX-IRES-Puro-GFP | A gift from PhD. Zhang Yunsheng | N/A |
| pCD513B-1 (pCDH-CMV-MCS-EF1-CopGFP-T2A-Puro) | MiaoLing Plasmid Platform | Cat# P0268 |
| Software and algorithms | ||
| FlowJo | https://www.flowjo.com/solutions/flowjo | |
| ImageJ | https://imagej.nih.gov/ij/ | |
| GraphPad Prism 8 | https://www.graphpad.com/ | |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Dan Li (sw_ld@hnu.edu.cn).
Materials availability
This study did not generate new unique reagents.
Experimental model and subject details
Animals
All mice used in this study were C57BL/6J background. 4w, 8w and 12w male Chd2+/+ and Chd2+/− mice were used for immunofluorescence and immunohistochemical experiments. 12w male Chd2+/+, Chd2+/− and 4w female Chd2+/+ mice were used for in vitro fertilization. 12w male Chd2+/+, Chd2+/− and 12w female Chd2+/+ mice were used for breeding. Mice were housed in an environment controlled for light (12 h on/off) at ambient temperature (21–23°C) with ad libitum access to water and food (normal mouse chow). All animal housing and handling were approved by the Experimental Animal Ethics Committee of the College of Biology, Hunan University.
Cell lines
GC-1 spg (ATCC, CRL-2053), GC-2 spd (ATCC, CRL-2196), NIH3T3 (ATCC, CRL-1658) and HEK293T (ATCC, ACS-4500) cells were cultured in DMEM (Gibco) supplemented with 10% fetal bovine serum (Gibco), 100 units/mL penicillin and 100 μg/mL streptomycin at 37°C with 5% CO2. C18-4 cells (provided by Professor Runsheng Li of Fudan University and Professor Zuping He of Hunan Normal University) were cultured in DMEM/F12 (1:1) (Biosharp) supplemented with 10% fetal bovine serum (Gibco), 100 units/mL penicillin and 100 μg/mL streptomycin at 37°C with 5% CO2. SSCs were cultured on 20 μg/mL laminin-coated plates and in mouse SSC medium (Procell Life Science & Technology) at 37°C with 5% CO2. Both SSCs, C18-4, GC-1 and GC-2 cells are male germ cells.
Method details
Generation of Chd2+/− mouse models
The Chd2 knockout (KO) heterozygous mouse model (C57BL/6J) was established using CRISPR/Cas9-mediated genome engineering by Cyagen Biosciences Inc. In brief, the designed gRNA was targeted to exons 3∼12 of the mouse Chd2 gene (NM_001081345.2; Ensembl: ENSMUSG00000078671), and the effective KO region size was ∼20467 bp. The constructed gRNA vector and Cas9 vector were transcribed in vitro, and then the gRNA and Cas9 mRNA were coinjected into the zygote. The microinjected fertilized egg was then returned to the fallopian tube of the surrogate mouse. PCR and DNA sequencing were used to identify mouse genotypes. The primers and gRNA sequences are presented in Table S2.
Spermatogonial stem cells isolation and culture
For primary culture of spermatogonial stem cells (SSCs), 6-day-old C57BL/6J mouse pups were used. The isolation and culture were performed according to the literature with slight changes.59 In brief, testicular cells were digested with 2 mg/mL collagenase IV (Biofroxx) and 200 U/mL DNase I (Biofroxx) for 10 min at 37°C. Then, the cells were digested with 0.25% trypsin-EDTA (BI) for 5 min. After centrifugation, the cells were plated on 0.1% gelatin-coated plates and incubated overnight. Germ cells were enriched by collecting nonattached cells. Cells were then transferred to laminin-coated plates and cultured in mouse SSC medium (Procell Life Science & Technology) at 37°C with 5% CO2.
SiRNA, vectors and cell transfection
A final concentration of 100–120 nmol/L siRNA was used, and siRNAs for siCHD2 (D-060656-03) and siCSTF3 (D-051652-17) were purchased from Dharmacon. The cDNA sequence of CSTF3 with a His tag sequence was cloned into the pCD513B-1 plasmid to generate the CSTF3-His plasmid. CHD21−455, CHD2456−955 or CHD2955−1827 was inserted into the pEGFP-C2 plasmid to generate CHD2-A, CHD2-B and CHD2-C plasmids. The CHD2456−955 fragment was cloned into the pLVX-IRES-Puro-GFP lentivirus plasmid to generate the pLVX-CHD2-B plasmid. The +933 to +1244 part of the Oct4 (NM_013633.3) cDNA sequence was cloned into the pEGFP-C2 plasmid to generate Oct4 wt and Oct4 mut. The primers used in the above experiments are presented in Table S2. Transient transfection was performed with TurboFect Transfection Reagent (Thermo Fisher Scientific, Inc. R0531) according to the manufacturer’s protocol. HEK293T cells were transfected with pLVX-CHD2-B, psPAX2 and pMD2.G plasmids at a 4:3:1 ratio to package lentivirus pLVX-CHD2-B.
Immunofluorescence assay
Immunofluorescence of mouse testes was performed with frozen sections. The preparation of frozen sections was performed according to the literature with slight changes.60 Briefly, mouse testes were fixed in 4% paraformaldehyde for 24 h at 4°C, washed in PBS and incubated in 30% w/v sucrose solution for 24 h at 4°C. Testes samples were placed in Optimum Cutting Temperature compound (OCT) (Sakura Finetek), frozen at −80°C and then sectioned using a freezing microtome. For immunofluorescence staining of frozen sections, 5 μm thick testis cross-sections were washed in PBS and blocked for 1 h at room temperature with 5% v/v FBS in PBS (block solution I). The sections were incubated overnight at 4°C with primary antibodies diluted in block solution I. The sections were then washed in PBS and incubated for 2 h at room temperature with appropriate fluorescent secondary antibodies.
Immunofluorescence of germ cells was performed with mouse testicular cell smears.61 Fresh testis removed from the albuginea was sheared into pieces in PBS and filtered with a 200-mesh sieve. The cells were washed with cold PBS 3 times and collected by centrifugation at 1500 rpm for 5 min at 4°C. Then, the cells were treated with hypotonic for 15 min, fixed with 4% paraformaldehyde for 10 min, smeared on lysine-coated glass slides and air-dried. Finally, the smears were blocked for 1 h at room temperature with 5% v/v FBS and 0.3% Triton X-100 in PBS (block solution II). After incubation overnight at 4°C with primary antibodies diluted in block solution II, the sections were then washed in PBS and incubated for 2 h at room temperature with appropriate fluorescent secondary antibodies.
For cellular immunofluorescence, SSCs and C18-4 cells were fixed on glass-bottom dishes with 4% paraformaldehyde for 10 min. Then, the cells were blocked for 1 h at room temperature with 5% v/v FBS and 0.3% Triton X-100 in PBS (block solution II). After incubation overnight at 4°C with primary antibodies diluted in block solution II, the cells were washed in PBS and incubated for 2 h at room temperature with appropriate fluorescent secondary antibodies.
The primary antibodies, fluorescent secondary antibodies and dilution rates are listed in key resources table.
In vitro fertilization experiment
The in vitro fertilization experiments were performed according to the literature with slight changes.62 Six-week-old C57BL/6J female mice were injected with 10 IU of pregnant mare serum gonadotropin (PMSG) (Nanjing Aibei Biotechnology) via intraperitoneal injection. After 48 h, they were injected with 10 IU of human chorionic gonadotropin (HCG) (Nanjing Aibei Biotechnology) via intraperitoneal injection. Thirteen hours after the HCG injection, the oocytes were harvested from the oviduct ampulla and transferred to KSOM embryo culture medium (Nanjing Aibei Biotechnology) under paraffin oil at 37°C. Sperm were harvested from the epididymis of Chd2+/+ and Chd2+/− mice and capacitated in HTF solution (Nanjing Aibei Biotechnology) for 1 h. Subsequently, the oocytes were incubated with capacitated sperm at a final concentration of 2×106 sperm/mL in KSOM embryo culture medium for 6 h. Then, the unbound sperm cells were removed by washing with KSOM medium. Finally, the fertilization rate was calculated using the percentage of two-cell embryos. Photographs were taken 12, 24, 48, 72 and 96 h after fertilization, and the blastocyst rate was calculated at 96 h. All animal studies were approved by the Experimental Animal Ethics Committee of the College of Biology, Hunan University.
Flow cytometry analysis of testicular cell populations
The flow cytometry analysis of testicular cell populations was performed according to the literature with slight changes.63,64 Testes were collected from 12-week-old Chd2+/+ and Chd2+/− male mice, and the albuginea was removed. The testes cells were washed with DPBS three times and treated with 2 mg/mL collagenase IV and 10 μg/mL DNase I at 37°C for 10 min. Then, the cells were digested with 4 mg/mL collagenase IV, 2.5 mg/mL hyaluronidase (Biofroxx), 0.25% trypsin-EDTA (BI), and 1 μg/μL DNase I (Biofroxx). After centrifugation, the cells were plated on 0.1% gelatin-coated plates for 2 h to remove most of the Sertoli cells. Suspended cells were collected and stained with Hoechst 33342 and propidium iodide according to the instructions of the Hoechst 33342/PI Double Stain Kit (Solarbio Life Sciences). The DNA content of germ cells was analyzed with flow cytometry (Beckman CytoFLEX).
Coimmunoprecipitation (IP), mass spectrometry and western blot analyses
For co-IP, total proteins were extracted with western blot and IP lysis buffer (Beyotime Biotech) from SSCs and C18-4, 293T or NIH3T3 cells. After preclearing with 30 μL protein A/G magnetic beads (Selleck) at 4°C, the proteins were incubated with 40 μL magnetic beads and primary antibodies or control IgG at 4°C overnight. Then, the beads were washed with PBS three times for 10 min each. Finally, the beads were resuspended in 50 μL of western blot and IP lysis buffer with SDS loading buffer (Beyotime) and boiled at 100°C for 5 min. Mass spectrometry analysis was conducted by OE Biotech Co. Ltd.
Total proteins of cultured cells were extracted with western blot and IP lysis buffer (Beyotime) and boiled with SDS loading buffer at 100°C for 5 min. Then, western blotting was performed with standard procedures. For testicular protein extraction, fresh testis removed from the albuginea was sheared into pieces in PBS and washed with PBS 3 times. Then, total protein was extracted from the pieces with western blotting and IP lysis buffer. Finally, the total proteins were used for western blotting. The antibodies used in the co-IP and western blots are listed in key resources table.
RNA extraction and RT‒qPCR
Total RNA was extracted using RNAiso Plus (Takara) according to the manufacturer’s protocol. cDNA was synthetized using PrimeScript™ RT reagent Kit with gDNA Eraser (Takara). RT‒qPCR was performed using SYBR Green qPCR Master Mix (Bimake) and a CFX96 Touch Deep Well real-time PCR detection system (Bio-Rad). The primers used in RT‒qPCR are presented in Table S2. 2−ΔΔCt was the ratio of gene expression of the experimental group and the control group.
Cell counting kit-8 (CCK-8) and 5-ethynyl-2′-deoxyuridine (EdU) staining
For the CCK8 assay, SSCs, C18-4 cells or GC-1 cells were seeded in 96-well plates at a density of 2000 cells/well. According to the manufacturer’s instructions, the absorbance was measured at 450 nm with a microplate reader to detect cell numbers. For EdU staining, SSCs, C18-4 cells or GC-1 cells were seeded in 6-well plates, and EdU staining was performed with a BeyoClickTM EdU Cell Proliferation Kit (C0075S, Beyotime, China) according to the manufacturer’s protocols and analyzed by fluorescence microscopy.
TdT-mediated dUTP nick-end labeling (TUNEL) staining
Paraffin sections of mouse testes were detected by a One Step TUNEL Apoptosis Assay Kit (Beyotime) according to the manufacturer’s protocol. Then, the sections were counterstained with DAPI and analyzed by fluorescence microscopy.
Colony-formation and sphere-formation assays
Digested SSCs, C18-4 cells or GC-1 cells were placed in 12-well plates at a density of 500 cells/well. After 10 days, the cells were fixed with 4% paraformaldehyde and stained with crystal violet for the colony-formation assay. Digested SSCs, C18-4 cells or GC-1 cells were placed in low-adsorption 6-well plates at a density of 1000 cells/well. After 10 days, the formation of microspheres was detected by microscopy.
Immunohistochemistry analysis and hematoxylin and eosin staining
Paraffin sections of mouse testes were used for immunohistochemistry according to standard procedures. PCNA or CHD2 antibody was used to detect cell proliferation in testes and the expression of CHD2 in mouse testes. Paraffin sections of mouse testes and sperm in the epididymis were used for hematoxylin and eosin staining with a HE Staining Kit (Solarbio) according to the manufacturer’s protocols.
Chromatin immunoprecipitation (ChIP)
The ChIP experiments were performed according to the literature.65 After transfection with siNC or siCHD2 for 48 h, C18-4 cells were cross-linked with 1% formaldehyde for 10 min at 37°C, terminated by 0.125 mol/L glycine at room temperature and then lysed by SDS lysis. Samples were sonicated at 30% power with a 5 s pulse and 5 s intervals at 4°C for 8 min using a Bioruptor Pico (Diagenode s.a., Seraing, Belgium) to obtain 200∼1000 bp fragments. After centrifugation, the supernatant was diluted with CHIP dilution buffer and incubated with 40 μL protein A/G magnetic beads and 4 μg CHD2 antibody or control anti-IgG (Beyotime) overnight at 4°C. The magnetic beads were washed with low-salt buffer, high-salt buffer, LiCl buffer, and twice with TE buffer. The complexes were eluted and decrosslinked at 65°C overnight. After treatment with RNase at 37°C for 1 h and proteinase K at 56°C for 1 h, DNA was purified with a DNA purification kit (TIANGEN) and amplified by qPCR as described above. Primer sequences for CHIP-qPCR are presented in Table S2.
RNA sequencing
RNA was extracted from Chd2+/+ and Chd2+/− mouse testes. RNA-seq library preparation and sequencing were performed by Shanghai Origin-gene Biological Company. Genes with a false discovery rate (FDR)<0.05 were considered to be differentially expressed. GO enrichment analysis provided the GO terms that were significantly enriched in the differentially expressed genes (DEGs).
mRNA stability assays
The mRNA stability assays were performed according to the literature with slight changes.60 C18-4 or NIH3T3 cells were treated with 10 μM actinomycin D (AbMole) for up to six hours. Cell samples were collected at 0, 3, and 6 h posttreatment, and total RNA was extracted using RNAiso Plus. cDNA synthesis and RT‒qPCR analysis was performed as described above (see RNA extraction and RT‒qPCR).
RNA immunoprecipitation (RIP)
The RIP experiments were performed according to the literature with slight changes.66 Fresh mouse testis removed from the albuginea was sheared into pieces and washed with PBS 3 times. Then, testis cells were fixed with 1% formaldehyde for 10 min at 37°C, terminated by 0.125 mol/L glycine at room temperature, and lysed by SDS lysis with RNase Inhibitor. After centrifugation, the supernatant was diluted with RIP dilution buffer and incubated with 40 μL protein A/G magnetic beads and 4 μg CHD2 antibody or control anti-IgG (Beyotime) overnight at 4°C. The samples were washed with washing buffer 3 times and eluted with RIP elution buffer at 65°C. After treatment with proteinase K at 56°C for 1 h, RNA was extracted using RNAiso Plus. RNase inhibitor was added in all steps above. cDNA synthesis and RT‒qPCR analysis was performed as described above (see RNA extraction and RT‒qPCR). The primer sequences used are shown in Table S2.
Quantification and statistical analysis
All experiments were performed in triplicate, and the results are presented as the mean ± standard deviation. Statistical significance was assessed by two-tailed unpaired t tests. A value of p < 0.05 (∗) was considered significant, and p < 0.01 (∗∗), p < 0.001 (∗∗∗), p < 0.0001 (∗∗∗∗) were considered prominently significant.
Acknowledgments
This work was supported by the National Key Research and Development Program of China [2021YFA0910103], the National Natural Science Foundation of China [81571494], and Central Government Funds for Guiding Local Scientific and Technological Development of Shenzhen City in China [2021Szvup034].
Author contributions
ZM performed the experiments and wrote the article. HX and XL were responsible for the IHC and IF experiment. JW and ZF were responsible for cell culture and protein extraction. XR and JF were responsible for reagents and materials. LY revised the article. DL designed the research and wrote the article. All authors contributed to the article and approved the submitted version.
Declaration of interests
The authors declare no competing interests.
Published: December 22, 2022
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2022.105552.
Supplemental information
Data and code availability
Data reported in this paper will be shared by the lead contact upon request. The accession codes for mass spectrometry proteomics and RNA-seq are provided in the key resources table.
This paper does not report original code.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
References
- 1.Neto F.T.L., Bach P.V., Najari B.B., Li P.S., Goldstein M. Spermatogenesis in humans and its affecting factors. Semin. Cell Dev. Biol. 2016;59:10–26. doi: 10.1016/j.semcdb.2016.04.009. [DOI] [PubMed] [Google Scholar]
- 2.Xu C., Shah M.A., Mipam T., Wu S., Yi C., Luo H., Yuan M., Chai Z., Zhao W., Cai X. Bovid microRNAs involved in the process of spermatogonia differentiation into spermatocytes. Int. J. Biol. Sci. 2020;16:239–250. doi: 10.7150/ijbs.38232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kanatsu-Shinohara M., Shinohara T. Spermatogonial stem cell self-renewal and development. Annu. Rev. Cell Dev. Biol. 2013;29:163–187. doi: 10.1146/annurev-cellbio-101512-122353. [DOI] [PubMed] [Google Scholar]
- 4.Dann C.T., Alvarado A.L., Molyneux L.A., Denard B.S., Garbers D.L., Porteus M.H. Spermatogonial stem cell self-renewal requires OCT4, a factor downregulated during retinoic acid-induced differentiation. Stem Cell. 2008;26:2928–2937. doi: 10.1634/stemcells.2008-0134. [DOI] [PubMed] [Google Scholar]
- 5.Izadyar F., Wong J., Maki C., Pacchiarotti J., Ramos T., Howerton K., Yuen C., Greilach S., Zhao H.H., Chow M., et al. Identification and characterization of repopulating spermatogonial stem cells from the adult human testis. Hum. Reprod. 2011;26:1296–1306. doi: 10.1093/humrep/der026. [DOI] [PubMed] [Google Scholar]
- 6.Hobbs R.M., Seandel M., Falciatori I., Rafii S., Pandolfi P.P. Plzf regulates germline progenitor self-renewal by opposing mTORC1. Cell. 2010;142:468–479. doi: 10.1016/j.cell.2010.06.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hammoud S.S., Low D.H.P., Yi C., Lee C.L., Oatley J.M., Payne C.J., Carrell D.T., Guccione E., Cairns B.R. Transcription and imprinting dynamics in developing postnatal male germline stem cells. Genes Dev. 2015;29:2312–2324. doi: 10.1101/gad.261925.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Carrieri C., Comazzetto S., Grover A., Morgan M., Buness A., Nerlov C., O'Carroll D. A transit-amplifying population underpins the efficient regenerative capacity of the testis. J. Exp. Med. 2017;214:1631–1641. doi: 10.1084/jem.20161371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Chakraborty P., Buaas F.W., Sharma M., Snyder E., de Rooij D.G., Braun R.E. LIN28A marks the spermatogonial progenitor population and regulates its cyclic expansion. Stem Cell. 2014;32:860–873. doi: 10.1002/stem.1584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nakagawa T., Jörg D.J., Watanabe H., Mizuno S., Han S., Ikeda T., Omatsu Y., Nishimura K., Fujita M., Takahashi S., et al. A multistate stem cell dynamics maintains homeostasis in mouse spermatogenesis. Cell Rep. 2021;37:109875. doi: 10.1016/j.celrep.2021.109875. [DOI] [PubMed] [Google Scholar]
- 11.Correia S., Alves M.R., Cavaco J.E., Oliveira P.F., Socorro S. Estrogenic regulation of testicular expression of stem cell factor and c-kit: implications in germ cell survival and male fertility. Fertil. Steril. 2014;102:299–306. doi: 10.1016/j.fertnstert.2014.04.009. [DOI] [PubMed] [Google Scholar]
- 12.Endo T., Romer K.A., Anderson E.L., Baltus A.E., de Rooij D.G., Page D.C. Periodic retinoic acid-STRA8 signaling intersects with periodic germ-cell competencies to regulate spermatogenesis. Proc. Natl. Acad. Sci. USA. 2015;112:E2347–E2356. doi: 10.1073/pnas.1505683112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hou H., Kyriacou E., Thadani R., Klutstein M., Chapman J.H., Cooper J.P. Centromeres are dismantled by foundational meiotic proteins Spo11 and Rec8. Nature. 2021;591:671–676. doi: 10.1038/s41586-021-03279-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Mashtalir N., D'Avino A.R., Michel B.C., Luo J., Pan J., Otto J.E., Zullow H.J., McKenzie Z.M., Kubiak R.L., St Pierre R., et al. Modular organization and assembly of SWI/SNF family chromatin remodeling complexes. Cell. 2018;175:1272–1288.e20. doi: 10.1016/j.cell.2018.09.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kadoch C., Crabtree G.R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv. 2015;1:e1500447. doi: 10.1126/sciadv.1500447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Clapier C.R., Iwasa J., Cairns B.R., Peterson C.L. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat. Rev. Mol. Cell Biol. 2017;18:407–422. doi: 10.1038/nrm.2017.26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kim Y., Fedoriw A.M., Magnuson T. An essential role for a mammalian SWI/SNF chromatin-remodeling complex during male meiosis. Development. 2012;139:1133–1140. doi: 10.1242/dev.073478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Serber D.W., Runge J.S., Menon D.U., Magnuson T. The mouse INO80 chromatin-remodeling complex is an essential meiotic factor for spermatogenesis. Biol. Reprod. 2016;94:8. doi: 10.1095/biolreprod.115.135533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chakraborty P., Magnuson T. INO80 requires a polycomb subunit to regulate the establishment of poised chromatin in murine spermatocytes. Development. 2022;149 doi: 10.1242/dev.200089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sun S., Jiang Y., Zhang Q., Pan H., Li X., Yang L., Huang M., Wei W., Wang X., Qiu M., et al. Znhit1 controls meiotic initiation in male germ cells by coordinating with Stra8 to activate meiotic gene expression. Dev. Cell. 2022;57:901–913.e4. doi: 10.1016/j.devcel.2022.03.006. [DOI] [PubMed] [Google Scholar]
- 21.Cafe S.L., Skerrett-Byrne D.A., De Oliveira C.S., Nixon B., Oatley M.J., Oatley J.M., Lord T. A regulatory role for CHD4 in maintenance of the spermatogonial stem cell pool. Stem Cell Rep. 2021;16:1555–1567. doi: 10.1016/j.stemcr.2021.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li W., Wu J., Kim S.Y., Zhao M., Hearn S.A., Zhang M.Q., Meistrich M.L., Mills A.A. Chd5 orchestrates chromatin remodelling during sperm development. Nat. Commun. 2014;5:3812. doi: 10.1038/ncomms4812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zhuang T., Hess R.A., Kolla V., Higashi M., Raabe T.D., Brodeur G.M. CHD5 is required for spermiogenesis and chromatin condensation. Mech. Dev. 2014;131:35–46. doi: 10.1016/j.mod.2013.10.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Semba Y., Harada A., Maehara K., Oki S., Meno C., Ueda J., Yamagata K., Suzuki A., Onimaru M., Nogami J., et al. Chd2 regulates chromatin for proper gene expression toward differentiation in mouse embryonic stem cells. Nucleic Acids Res. 2017;45:8758–8772. doi: 10.1093/nar/gkx475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Shen T., Ji F., Yuan Z., Jiao J. CHD2 is required for embryonic neurogenesis in the developing cerebral cortex. Stem Cell. 2015;33:1794–1806. doi: 10.1002/stem.2001. [DOI] [PubMed] [Google Scholar]
- 26.Kim Y.J., Khoshkhoo S., Frankowski J.C., Zhu B., Abbasi S., Lee S., Wu Y.E., Hunt R.F. Chd2 is necessary for neural circuit development and long-term memory. Neuron. 2018;100:1180–1193.e6. doi: 10.1016/j.neuron.2018.09.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Shahin Varnoosfaderani F., Palau A., Dong W., Persson J., Durand-Dubief M., Svensson J.P., Lennartsson A. A regulatory role for CHD2 in myelopoiesis. Epigenetics. 2020;15:702–714. doi: 10.1080/15592294.2019.1710913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Luijsterburg M.S., de Krijger I., Wiegant W.W., Shah R.G., Smeenk G., de Groot A.J.L., Pines A., Vertegaal A.C.O., Jacobs J.J.L., Shah G.M., van Attikum H. PARP1 links CHD2-mediated chromatin expansion and H3.3 deposition to DNA repair by non-homologous end-joining. Mol. Cell. 2016;61:547–562. doi: 10.1016/j.molcel.2016.01.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Marfella C.G.A., Ohkawa Y., Coles A.H., Garlick D.S., Jones S.N., Imbalzano A.N. Mutation of the SNF2 family member Chd2 affects mouse development and survival. J. Cell. Physiol. 2006;209:162–171. doi: 10.1002/jcp.20718. [DOI] [PubMed] [Google Scholar]
- 30.Chénier S., Yoon G., Argiropoulos B., Lauzon J., Laframboise R., Ahn J.W., Ogilvie C.M., Lionel A.C., Marshall C.R., Vaags A.K., et al. CHD2 haploinsufficiency is associated with developmental delay, intellectual disability, epilepsy and neurobehavioural problems. J. Neurodev. Disord. 2014;6:9. doi: 10.1186/1866-1955-6-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Chen J., Zhang J., Liu A., Zhang L., Li H., Zeng Q., Yang Z., Yang X., Wu X., Zhang Y. CHD2-related epilepsy: novel mutations and new phenotypes. Dev. Med. Child Neurol. 2020;62:647–653. doi: 10.1111/dmcn.14367. [DOI] [PubMed] [Google Scholar]
- 32.Gòdia M., Reverter A., González-Prendes R., Ramayo-Caldas Y., Castelló A., Rodríguez-Gil J.E., Sánchez A., Clop A. A systems biology framework integrating GWAS and RNA-seq to shed light on the molecular basis of sperm quality in swine. Genet. Sel. Evol. 2020;52:72. doi: 10.1186/s12711-020-00592-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Qin Y., Ji J., Du G., Wu W., Dai J., Hu Z., Sha J., Hang B., Lu C., Xia Y., Wang X. Comprehensive pathway-based analysis identifies associations of BCL2, GNAO1 and CHD2 with non-obstructive azoospermia risk. Hum. Reprod. 2014;29:860–866. doi: 10.1093/humrep/deu013. [DOI] [PubMed] [Google Scholar]
- 34.Handel M.A., Schimenti J.C. Genetics of mammalian meiosis: regulation, dynamics and impact on fertility. Nat. Rev. Genet. 2010;11:124–136. doi: 10.1038/nrg2723. [DOI] [PubMed] [Google Scholar]
- 35.Enguita-Marruedo A., Martín-Ruiz M., García E., Gil-Fernández A., Parra M.T., Viera A., Rufas J.S., Page J. Transition from a meiotic to a somatic-like DNA damage response during the pachytene stage in mouse meiosis. PLoS Genet. 2019;15:e1007439. doi: 10.1371/journal.pgen.1007439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Dai X.X., Jiang Y., Gu J.H., Jiang Z.Y., Wu Y.W., Yu C., Yin H., Zhang J., Shi Q.H., Shen L., et al. The CNOT4 subunit of the CCR4-NOT complex is involved in mRNA degradation, efficient DNA damage repair, and XY chromosome crossover during male germ cell meiosis. Adv. Sci. 2021;8:2003636. doi: 10.1002/advs.202003636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Turner J.M.A., Aprelikova O., Xu X., Wang R., Kim S., Chandramouli G.V.R., Barrett J.C., Burgoyne P.S., Deng C.X. BRCA1, histone H2AX phosphorylation, and male meiotic sex chromosome inactivation. Curr. Biol. 2004;14:2135–2142. doi: 10.1016/j.cub.2004.11.032. [DOI] [PubMed] [Google Scholar]
- 38.Feng S., Li J., Wen H., Liu K., Gui Y., Wen Y., Wang X., Yuan S. hnRNPH1 recruits PTBP2 and SRSF3 to modulate alternative splicing in germ cells. Nat. Commun. 2022;13:3588. doi: 10.1038/s41467-022-31364-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gnugnoli M., Casari E., Longhese M.P. The chromatin remodeler Chd1 supports MRX and Exo1 functions in resection of DNA double-strand breaks. PLoS Genet. 2021;17:e1009807. doi: 10.1371/journal.pgen.1009807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wild P., Susperregui A., Piazza I., Dörig C., Oke A., Arter M., Yamaguchi M., Hilditch A.T., Vuina K., Chan K.C., et al. Network rewiring of homologous recombination enzymes during mitotic proliferation and meiosis. Mol. Cell. 2019;75:859–874.e4. doi: 10.1016/j.molcel.2019.06.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hofmann M.C., Braydich-Stolle L., Dettin L., Johnson E., Dym M. Immortalization of mouse germ line stem cells. Stem Cell. 2005;23:200–210. doi: 10.1634/stemcells.2003-0036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Igolkina A.A., Zinkevich A., Karandasheva K.O., Popov A.A., Selifanova M.V., Nikolaeva D., Tkachev V., Penzar D., Nikitin D.M., Buzdin A. H3K4me3, H3K9ac, H3K27ac, H3K27me3 and H3K9me3 histone tags suggest distinct regulatory evolution of open and condensed chromatin landmarks. Cells. 2019;8 doi: 10.3390/cells8091034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Grozdanov P.N., Masoumzadeh E., Latham M.P., MacDonald C.C. The structural basis of CstF-77 modulation of cleavage and polyadenylation through stimulation of CstF-64 activity. Nucleic Acids Res. 2018;46:12022–12039. doi: 10.1093/nar/gky862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Tian B., Manley J.L. Alternative cleavage and polyadenylation: the long and short of it. Trends Biochem. Sci. 2013;38:312–320. doi: 10.1016/j.tibs.2013.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Hockert J.A., Yeh H.J., MacDonald C.C. The hinge domain of the cleavage stimulation factor protein CstF-64 is essential for CstF-77 interaction, nuclear localization, and polyadenylation. J. Biol. Chem. 2010;285:695–704. doi: 10.1074/jbc.M109.061705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Luo W., Ji Z., Pan Z., You B., Hoque M., Li W., Gunderson S.I., Tian B. The conserved intronic cleavage and polyadenylation site of CstF-77 gene imparts control of 3' end processing activity through feedback autoregulation and by U1 snRNP. PLoS Genet. 2013;9:e1003613. doi: 10.1371/journal.pgen.1003613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Yang W., Hsu P.L., Yang F., Song J.E., Varani G. Reconstitution of the CstF complex unveils a regulatory role for CstF-50 in recognition of 3'-end processing signals. Nucleic Acids Res. 2018;46:493–503. doi: 10.1093/nar/gkx1177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Rom A., Melamed L., Gil N., Goldrich M.J., Kadir R., Golan M., Biton I., Perry R.B.T., Ulitsky I. Regulation of CHD2 expression by the Chaserr long noncoding RNA gene is essential for viability. Nat. Commun. 2019;10:5092. doi: 10.1038/s41467-019-13075-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Miles W.O., Lembo A., Volorio A., Brachtel E., Tian B., Sgroi D., Provero P., Dyson N. Alternative polyadenylation in triple-negative breast tumors allows NRAS and c-JUN to bypass PUMILIO posttranscriptional regulation. Cancer Res. 2016;76:7231–7241. doi: 10.1158/0008-5472.CAN-16-0844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Soumillon M., Necsulea A., Weier M., Brawand D., Zhang X., Gu H., Barthès P., Kokkinaki M., Nef S., Gnirke A., et al. Cellular source and mechanisms of high transcriptome complexity in the mammalian testis. Cell Rep. 2013;3:2179–2190. doi: 10.1016/j.celrep.2013.05.031. [DOI] [PubMed] [Google Scholar]
- 51.Morgan M., Kumar L., Li Y., Baptissart M. Post-transcriptional regulation in spermatogenesis: all RNA pathways lead to healthy sperm. Cell. Mol. Life Sci. 2021;78:8049–8071. doi: 10.1007/s00018-021-04012-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Dai X., Zhang Q., Yu Z., Sun W., Wang R., Miao D. Bmi1 deficient mice exhibit male infertility. Int. J. Biol. Sci. 2018;14:358–368. doi: 10.7150/ijbs.23325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Legrand P., Pinaud N., Minvielle-Sébastia L., Fribourg S. The structure of the CstF-77 homodimer provides insights into CstF assembly. Nucleic Acids Res. 2007;35:4515–4522. doi: 10.1093/nar/gkm458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Bai Y., Auperin T.C., Chou C.Y., Chang G.G., Manley J.L., Tong L. Crystal structure of murine CstF-77: dimeric association and implications for polyadenylation of mRNA precursors. Mol. Cell. 2007;25:863–875. doi: 10.1016/j.molcel.2007.01.034. [DOI] [PubMed] [Google Scholar]
- 55.Dass B., Tardif S., Park J.Y., Tian B., Weitlauf H.M., Hess R.A., Carnes K., Griswold M.D., Small C.L., Macdonald C.C. Loss of polyadenylation protein tauCstF-64 causes spermatogenic defects and male infertility. Proc. Natl. Acad. Sci. USA. 2007;104:20374–20379. doi: 10.1073/pnas.0707589104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Tardif S., Akrofi A.S., Dass B., Hardy D.M., MacDonald C.C. Infertility with impaired zona pellucida adhesion of spermatozoa from mice lacking TauCstF-64. Biol. Reprod. 2010;83:464–472. doi: 10.1095/biolreprod.109.083238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Tsukiyama T. The in vivo functions of ATP-dependent chromatin-remodelling factors. Nat. Rev. Mol. Cell Biol. 2002;3:422–429. doi: 10.1038/nrm828. [DOI] [PubMed] [Google Scholar]
- 58.de la Serna I.L., Ohkawa Y., Imbalzano A.N. Chromatin remodelling in mammalian differentiation: lessons from ATP-dependent remodellers. Nat. Rev. Genet. 2006;7:461–473. doi: 10.1038/nrg1882. [DOI] [PubMed] [Google Scholar]
- 59.Kanatsu-Shinohara M., Ogonuki N., Matoba S., Morimoto H., Ogura A., Shinohara T. Improved serum- and feeder-free culture of mouse germline stem cells. Biol. Reprod. 2014;91:88. doi: 10.1095/biolreprod.114.122317. [DOI] [PubMed] [Google Scholar]
- 60.Legrand J.M.D., Chan A.L., La H.M., Rossello F.J., Änkö M.L., Fuller-Pace F.V., Hobbs R.M. DDX5 plays essential transcriptional and post-transcriptional roles in the maintenance and function of spermatogonia. Nat. Commun. 2019;10:2278. doi: 10.1038/s41467-019-09972-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Li X., Wu Y., Huang L., Yang L., Xing X. SPAG4L/SPAG4Lbeta interacts with Nesprin2 to participate in the meiosis of spermatogenesis. Acta Biochim. Biophys. Sin. 2019;51:669–676. doi: 10.1093/abbs/gmz051. [DOI] [PubMed] [Google Scholar]
- 62.Yan Q., Huang H., Lu S., Ou B., Feng J., Shan W., Li H., Wang Z., Hong A., Ma Y. PACAP ameliorates fertility in obese male mice via PKA/CREB pathway-dependent Sirt1 activation and p53 deacetylation. J. Cell. Physiol. 2020;235:7465–7483. doi: 10.1002/jcp.29651. [DOI] [PubMed] [Google Scholar]
- 63.Bastos H., Lassalle B., Chicheportiche A., Riou L., Testart J., Allemand I., Fouchet P. Flow cytometric characterization of viable meiotic and postmeiotic cells by Hoechst 33342 in mouse spermatogenesis. Cytometry A. 2005;65:40–49. doi: 10.1002/cyto.a.20129. [DOI] [PubMed] [Google Scholar]
- 64.Gaysinskaya V., Bortvin A. Flow cytometry of murine spermatocytes. Curr. Protoc. Cytom. 2015;72:7.44.1–7.44.24. doi: 10.1002/0471142956.cy0744s72. [DOI] [PubMed] [Google Scholar]
- 65.Liu X., Fan Z., Li Y., Li Z., Zhou Z., Yu X., Wan J., Min Z., Yang L., Li D. microRNA-196a-5p inhibits testicular germ cell tumor progression via NR6A1/E-cadherin axis. Cancer Med. 2020;9:9107–9122. doi: 10.1002/cam4.3498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Li K., Zhong S., Luo Y., Zou D., Li M., Li Y., Lu Y., Miao S., Wang L., Song W. A long noncoding RNA binding to QKI-5 regulates germ cell apoptosis via p38 MAPK signaling pathway. Cell Death Dis. 2019;10:699. doi: 10.1038/s41419-019-1941-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data reported in this paper will be shared by the lead contact upon request. The accession codes for mass spectrometry proteomics and RNA-seq are provided in the key resources table.
This paper does not report original code.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.