Abstract
Monotopic phosphoglycosyl transferase enzymes (MonoPGTs) initiate the assembly of prokaryotic glycoconjugates essential for bacterial survival and proliferation. MonoPGTs belong to an expansive superfamily with a diverse and richly annotated sequence space; however, the biochemical roles of most monoPGTs in glycoconjugate biosynthesis pathways remain elusive. To better understand these critical enzymes, we have implemented activity-based protein profiling (ABPP) probes as protein-centric, membrane-protein compatible tools that lay the groundwork for understanding the activity and regulation of the monoPGT superfamily from a cellular proteome. With straightforward gel-based read outs we demonstrate robust, covalent labeling at the active site of various representative monoPGTs from unfractionated cell membrane fractions using 3-phenyl-2H-azirine probes.
Graphical Abstract

INTRODUCTION
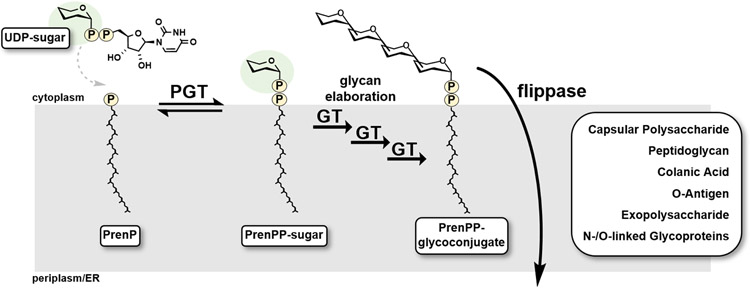
Bacterial glycoconjugate biosynthesis is initiated at the cytoplasmic face of the cell by phosphoglycosyl transferases (PGTs), which catalyze the transfer of a phospho-sugar from a soluble nucleoside diphosphate (NDP)-activated donor to a polyprenol phosphate (Pren-P) acceptor (Figure 1).1 The membrane-bound PGT product then undergoes sequential glycosylations by the action of glycosyltransferases (GTs), and is ultimately flipped to the periplasm to afford the substrate for mature glycoconjugate biosynthesis. These glycoconjugates contribute to the structural integrity of cells and feature macromolecular landscapes that interact with human hosts, and include the O-antigen of lipopolysaccharide (LPS),2, 3 capsular polysaccharide (CPS),4 and peptidoglycan (PG).5
Figure 1.
Phosphoglycosyl transferases (PGTs) catalyze the initial membrane-committed step in the en bloc mechanism of glycoconjugate biosynthesis. The PrenPP-sugar product of the PGT reaction is further elaborated via sequential modification by glycosyltransferases (GTs) to afford PrenPP-glycoconjugates, that are translocated to the periplasmic space by a flippase. These membrane-bound glycoconjugates then continue on support essential functions related to bacterial survival and proliferation.
PGTs are represented by two broad enzyme superfamilies that are distinct in both structure and mechanism of action.6 The polytopic PGTs (polyPGTs) are exemplified by the mammalian GPT and bacterial WecA and have been shown to invoke a mechanism involving a ternary complex utilizing a conserved Asp-Asp-X-X-Asp motif.7-9 In contrast the monotopic PGTs (monoPGTs), exemplified by the Campylobacter PglC, invoke covalent catalysis exploiting an active site aspartic acid carboxyl group. This nucleophilic aspartic acid activity is unique and a hallmark of the mechanism of monoPGTs, which supports probe development with specificity towards the monoPGT superfamily.10
The monoPGTs are named for the distinct membrane topology of the minimal functional unit in which a re-entrant membrane helix penetrates a single leaflet of the bilayer and anchors the enzyme to the cytoplasmic face of the membrane. The monoPGT superfamily is exclusively prokaryotic and members share a signature functional core that can be elaborated at the N- or C- terminus with various modalities.11 A recent sequence similarity network (SSN) analysis has classified over 38,000 non-redundant sequences in the monoPGT superfamily; the SSN has illuminated a wide range of functional domains, including sugar-modifying enzymes, glycosyl transferases or regulatory domains that may be appended to the monoPGT functional core.12 The diversification of a common basic scaffold across the superfamily could allow for differential regulation of pathways with shared membrane-embedded substrates. In particular, one cluster in the SSN contained PGTs fused to regulatory domains, suggesting that PGTs may play a central role in the regulation of glycoconjugate biosynthesis pathways. Additional recent work showed that Cap5M, a monoPGT in the pathway for CPS production in Staphylococcus aureus, is activated upon Tyr phosphorylation by the kinase CapB,13 further corroborating the idea that regulation of bacterial glycosylation may occur at the PGT step.
To understand these critical regulatory processes, the gap in knowledge between gene sequence and protein activity must be addressed. Implementation of protein-forward techniques and analyses can serve to complement the invaluable genetic information uncovered by the SSN. However, protein-forward analyses typically rely on purified proteins, and membrane protein isolation and handling is rife with technical challenges: membrane proteins, such as the monoPGTs, are prone to low expression levels and solubilization from lipidic environments typically relies on detergents that can perturb structure and function, which can further promote aggregation and precipitation from solution.14 As such, the expression and activity of monoPGTs is best studied in the context of the native complex cellular milieu.
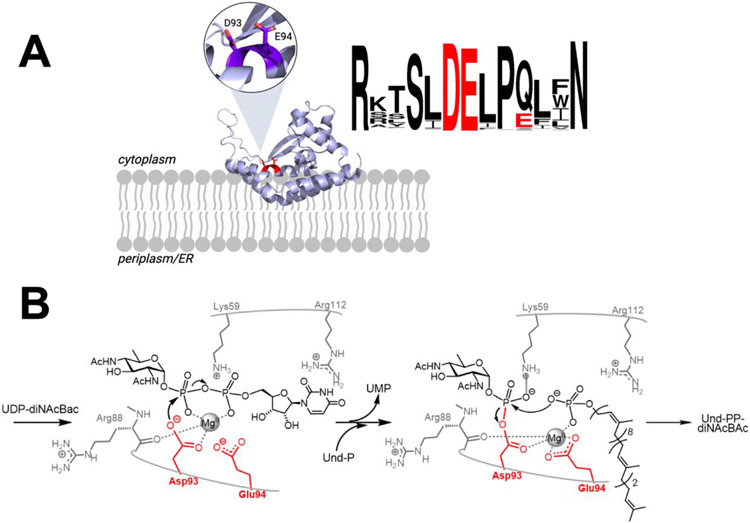
Activity-based protein profiling (ABPP) is a powerful proteomics technique used to selectively interrogate the functional proteome by covalently targeting the active sites of classes of proteins with shared mechanistic signatures.15, 16 ABPP is uniquely suited for probing complex proteomes, and is suited to membrane protein analysis in lysates or in cell envelope fractions (CEFs). By extension, ABPP is potentially ideal as a proteomics approach for probing the expansive monoPGT superfamily. Biochemical studies and sequence analysis have identified a highly-conserved active site in the monoPGT superfamily, including an Asp-Glu dyad catalytic motif (Figure 2A). This motif contributes to monoPGT catalysis, through a Bi-Bi ping pong mechanism.10 In this mechanism, the first reaction step involves formation of a covalent phosphoglycosyl derivative of the carboxyl group of the active site aspartic acid (Figure 2B). Within glycoconjugate biosynthesis pathways, this mechanism is exclusive to monoPGTs and affords the potential for selective intervention as the nucleophilic Asp is primed for probing with active-site targeted electrophilic agents. In contrast, the polyPGTs include a conserved active site Asp-Asp-Xaa-Xaa-Asp motif that plays a role in substrate orientation and catalysis, but not in nucleophilic catalysis.
Figure 2.
A) Ribbon diagram of monoPGT PglC (PDB 5W7L), inset and red coloring shows close-up view of active site highlighting the PglC Asp-Glu catalytic dyad. Sequence logo showing conservation in the catalytic residues among PglC orthologs. Logo generated using https://weblogo.berkeley.edu/logo.cgi. B) Bi-Bi ping-pong mechanism between a UDP-N,N'-diacetylbacillosamine (UDP-diNAcBac) as the UDP-sugar donor, and undecaprenol phosphate (Und-P) as the Pren-P acceptor. Proposed roles for binding are shown and catalytic residues are highlighted in red (numbering from Campylobacter concisus PglC).
Carboxylic acid-modifying ABPP probes in general have shown notoriously poor amino acid specificity; amino acids that are more nucleophilic, such as cysteine or serine, can out-compete carboxylic acids in reactions with electrophiles.17 Typical moieties used for carboxylic acid labeling include carbodiimides, epoxides, diazo compounds, phenylsulfonate esters, α-chloroacetamides, and α-β unsaturated ketones.17-19 We explored many of these classical labeling options for monoPGTs. Compounds containing carbodiimides and phenylsulfonate esters resulted in nonspecific labeling, and compounds comprising epoxides and α-β unsaturated ketones led to no observable monoPGT labeling. Alternatively, specificity can be controlled employing a warhead approach by synthesizing known protein ligands strategically equipped with electrophiles. However, the warhead approach requires prior knowledge of either protein structure or protein ligands and many important enzyme classes do not yet have this information available. Notably, amino acids with carboxylic acid side chains have been shown to be efficiently labeled in glycosidases exhibiting the Koshland double displacement mechanism.20 These probes employed deoxyfluorosugar substrate mimics21 as well as aziridine-based cyclophellitol inhibitor scaffolds.22
Recently, Ma et al. have described a 2H-azirine (2H-Az) electrophilic moiety for carboxylic acid labeling,23 and the 3-phenyl-2H-Az compounds were shown to be ideal for ABPP; the reactive group is small, stable and shows carboxylate-selective reactivity. Utility was demonstrated in live cells, and derivatives can be prepared using straightforward chemistry, enabling integration of reactive handles such as azides or alkynes for protein labeling and detection as well as mass spectrometry (MS)-based proteomic analysis. Herein, we present ABPP of monoPGTs with 3-phenyl-2H-Az probes that target the highly-conserved nucleophilic Asp of the Asp-Glu dyad. This work lays the foundation for the identification of glycoconjugate biosynthesis pathways in diverse prokaryotes and provides tools for studying regulatory mechanisms in the multistep processes. We also provide a straightforward synthetic route that will allow for tunable 3-phenyl-2H-Az derivatives.
METHODS
Preparation of Cell Envelope Fractions (CEFs)
The CEF preparation was carried out at 4 °C. Cell pellets from 500 mL culture were re-suspended in 50 mL buffer A (50 mM PBS, pH 7.5) containing lysozyme (50 mg) (Research Products International, cat L38100), protease inhibitor cocktail (50 μL) (Roche, cat 11836170001) and DNAse I (100 μL) (New England BioLabs cat M0303S). Cells were sonicated (50 % amplitude, 1 sec ON/2 sec OFF, 2 x 2 min). Cells were kept on ice during sonication. The resultant lysate was centrifuged at 9,000 g for 45 min at 4 °C. The cloudy supernatant was collected in a new centrifuge tube and further centrifuged at 140,000 g for 65 min at 4 °C. A membrane pellet (cell envelope fraction, CEF) was produced. The CEF was re-suspended using a Dounce homogenizer in 10 mL buffer A,
Probe labeling and click conditions
CEF was diluted in Buffer A to a total protein concentration of 10 mg/mL, and was subsequently treated with DMSO-dissolved probe at the appropriate concentrations. The probe-treated CEF was allowed to react with the probe for either 18 h at 4 °C, or 2 h at RT. Click with the azide-conjugate (N3-biotin or N3-fluorophore) was carried out following previous literature reports.4 Azide-conjugate was added to the eppendorf containing the probe reaction to a final concentration of 1 mM. This was followed by the addition of 100 μM TBTA, 1 mM sodium ascorbate, and 100 μM CuSO4. The reaction was agitated for 2 h at RT, then quenched with loading dye comprising BME.
Western blot analysis
Labeled CEF samples were separated by gel electrophoresis on Biorad 4-20% gradient gels. 7 uL of sample was loaded per lane for Western blot and SDS-PAGE analysis. Western blot analyses were run at 100 V for 1 h at 4 °C. For the detection of PGTs, membranes were incubated with a 1:5000 dilution of mouse anti-His antibody (LifeTein) in TBS-T 3% BSA followed by a 1:5000 dilution of IRDye 800CW secondary goat anti-mouse antibody in Odyssey blocking buffer (LI-COR Biosciences). The blots were imaged using a LICOR imager. For subsequent detection of labeled protein, the Western blots were stripped by incubation at 37 °C in stripping buffer (62.5 mM Tris, pH 6.8, 1% SDS, 0.35% β-mercaptoethanol (v/v)) for 1 hour. After thorough washing in TBS-T, the membrane was blocked in Odyssey blocking buffer for 1 hour at RT, and incubated with 0.1 mg/mL Streptavidin-647 in TBS-T 3% BSA overnight at 4 °C. The blots were washed three times in TBS prior to imaging. Labeled proteins were visualized by scanning fluorescence on a Typhoon FLA 9000 (GE Healthcare).
RESULTS AND DISCUSSION
Representative enzymes from the monoPGT superfamily were selected for activity profiling: PglC from Campylobacter jejuni (Cj), PglC from Helicobacter pullorum (Hp), and PglB from Clostridium botulinum (Cb). C. jejuni is a human food borne enteropathogen that is a significant cause of gastroenteritis worldwide.24 H. pullorum is a zoonotic pathogen known to colonize the gastrointestinal tract of poultry. C. botulinum is a food borne pathogen that can produce botulinum toxin. PglC from C. jejuni and H. pullorum catalyze the first membrane-committed step in the biosynthesis of N-linked glycoproteins, and PglC (Hp) displays 68% sequence identity to PglC (Cj). C. botulinum PglB catalyzes this same reaction in an unknown glycoconjugate biosynthesis pathway and shares 36% sequence identity with PglC (Cj).
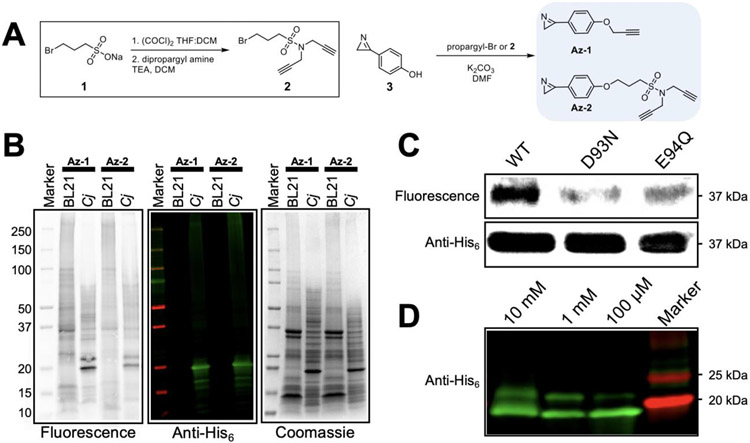
Following the established protocol, we first resynthesized the principal 3-phenyl-2H-Az probe described by Ma, et al. The final product matched the literature characterization including NMR analysis and mass spectrometry. For a broader range of ABPP applications we established a revised route that would enable diversification of the probe structures without impacting the carefully balanced electrophilic reactivity. With an activated C=N double bond and a lone pair of electrons on the nitrogen, the 2H-Az is a key precursor for a wide variety of transformations due to the strong reactivity towards a wide range of electrophiles and nucleophiles.17 This makes developing synthetic routes for ABPP challenging because many standard reaction conditions to incorporate biologically useful pharmacophores or functional groups used in copper catalyzed azide-alkyne cycloaddition (CuAAC) click chemistry may not be orthogonal to both the synthesis and the preservation of the 2H-Az moiety. To avoid introducing multiple synthetic routes that are only orthogonal to specific moieties, we instead employed a late-stage diversification strategy that uses modular attachments to bring together the p-phenyl-2H-azirine and select building blocks at the final synthetic step. An alkyl halide sulfonyl chloride linker is key to this strategy; it has two distinct electrophilic groups capable of selective step-wise nucleophilic addition. We exemplified this route by first chlorinating sodium sulfonate (1) with oxalyl-chloride, then selectively reacting the sulfonyl chloride with dipropargyl amine to introduce the essential terminal alkyne for click chemistry (Figure 3A). The resulting compound (2) is then used to alkylate the base-activated p-phenol-2H-azirine (3), yielding Az-2.
Figure 3.
A) Synthesis the sulfonamide linker, and synthesis of probes Az-1 and Az-2. B) Western blot analysis of CEF containing overexpressed PglC-His6 (Cj) in BL21, compared to CEF without any overexpressed proteins. The left most Western blot shows the fluorescence read out from the streptavidin fluorophore conjugate, and the middle shows the same Western blot imaged with Anti-His6 fluorophore conjugate. The right most is a gel with the total protein stained by Coomassie. C) Probe labeling of His6-SUMO-PglC (Cc) active site variants with Az-1 (100 μM). D) Probe labeling with Az-1 in a concentration dependent manner, followed by a click reaction with monodispersed N3-PEG36, demonstrating a single covalent modification visualized with a single mass shift.
Probes Az-1 and Az-2 (100 μM) were first applied to the cell envelope fraction (CEF) of Escherichia coli BL21 overexpressing PglC-His6 (Cj).25 Although we are heterologously overexpressing PglC, expression yields are modest using this method; membrane protein overexpression is considerably lower compared to that of soluble proteins. Each probe was incubated with CEF for 18 h at 4 °C and subsequently conjugated by CuAAC click chemistry with N3-biotin. As a negative control, these probes were also tested against BL21 CEF not overexpressing a monoPGT. Labeling was analyzed by Western blot with Alexa Fluor™ 647 streptavidin. The presence of PglC-His6 was confirmed by anti-His antibody detection and total protein was visualized by Coomassie staining (Figure 3B). Analysis of the resulting labeled proteome using fluorescence imaging revealed robust covalent labeling of PglC (Cj) by Az-1 and Az-2, with the most prominent band occurring at the expected molecular weight (23 kDa). Additional fluorescence labeling of background proteins was minimal when directly compared to the prominence of the PglC (Cj) band. Notably, Az-2 maintains reactivity towards activated protein carboxylic acids, demonstrating that the balanced reactivity of the 2H-azirine is not impacted by the additional chemical moieties. This provides a viable strategy for introducing features such as known protein ligands to boost probe affinity and selectivity. The fluorescent labeling for both Az-1 and Az-2 occurred with an intensity that is dependent on the final concentration of the probe (Figure S1). This concentration dependent experiment allowed us to select the probe concentration for our experiments to afford the best visual for our gel-based read outs.
Active site specificity was confirmed by probe labeling Asp-Glu active site mutants. These experiments were carried out using PglC from C. concisus (Cc), because active site mutations are not well tolerated in PglC (Cj). Furthermore, we used a SUMO construct, His6-SUMO-PglC, to express active site variants; the N-terminal SUMO tag moderately mitigates the low expression yields and poor stability these mutants display. Asp93, which has been biochemically determined to form the covalent intermediate,10 was mutated to Asn (D93N). Separately, Glu94, which is proposed to coordinate with the Mg2+ cofactor, was mutated to Gln (E94Q). The same labeling methodology was applied to the variants. Labeling was analyzed by Western blot with Alexa Fluor™ 647 streptavidin, and the presence of PglC was confirmed with anti-His6 antibody detection (Figure 3C). By fluorescence, each active site mutant displays attenuated labeling relative to WT protein. Labeling is eliminated with the D93N mutant, capturing the essentiality of the catalytic Asp93 residue. In contrast, the E94Q variant also shows reduced labeling, but not to the extent of the catalytically essential D93N. We propose that the reactivity of the Asp is perturbed by this local mutation change. Interestingly, the Glu94 appears to impact the nucleophilicity of the Asp93, but it is clearly not the main contributor. The observed labeling corroborates the two-step ping-pong mechanism of the monoPGTs, involving a catalytic Asp-Glu (DE) dyad, wherein the Asp carboxyl group attacks at the β-phosphate of the NDP-sugar substrate forming a covalent phospho-sugar intermediate.10
To further test the specificity of Az-1 for catalytic residues, we carried out a concentration-dependent gel-shift assay on CEF containing PglC (Cj). Labeling was carried out in the same manner as previously described, and the probe was clicked with monodisperse N3-PEG36. A direct fluorescence read-out visualized by anti-His antibody detection correlates the number of new bands to the number of probe additions to a monoPGT, where each new band represents a mass shift of approximately 1.5 kDa. Probe concentrations ranged from 100 μM to 10 mM (Figure 3D). Importantly, only a single mass shift was visualized with each concentration, even with treatment up to the 10 mM probe; this clearly demonstrates site-specific probe modification to PglC (Cj). The efficiency of labeling is concentration dependent. This single labeling is presumed to occur predominantly at the Asp93 residue, based on the previously demonstrated enhanced chemical nucleophilicity and supporting biochemical analysis.10
To validate the specificity of labeling for active site residues, we measured changes in activity of Cj PglC upon labeling with Az-1. CEF containing overexpressed Cj PglC was treated with 1 mM and 10 mM Az-1 for 18 h at 4 °C. We selected these probe concentrations based on densitometry from the previous gel-shift assay. At these concentrations, labeling is observed at an efficiency that would sufficiently attenuate activity if active site residues are in fact targeted by Az-1. We applied a radioactivity-based assay which directly measures Und-PP-sugar product by monitoring the transfer of a radiolabeled-phosphosugar from the UDP-[3H]-diNAcBac to the unlabeled Und-P acceptor; substrate conversion is quantified by scintillation counting after extraction.26 A time course of activity was carried out under linear initial rate conditions, and the sample treated with 10 mM Az-1 had activity diminished to 53% compared to an unlabeled CEF sample (WT unlabeled CEF maintains complete activity under these conditions). The sample treated with 1 mM Az-1 had activity that was reduced to 84% compared to unlabeled CEF (Figure S2). This concentration dependent decrease in activity is related to active-site labeling, inhibiting catalysis and product formation.
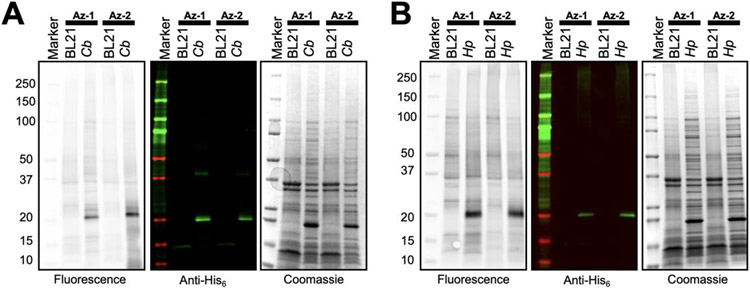
After characterizing probe labeling of PglC (Cj), we employed the activity probes to test labeling of additional superfamily members, including PglC (Hp) and PglB (Cb) (Figure 4). These select monoPGTs comprise the minimal functional catalytic core. Following the same labeling methodology, probes Az-1 and Az-2 showed very robust and targeted labeling of monoPGTs from the complex protein-rich CEF mixture. Strong fluorescent bands are seen at the molecular weight corresponding to each monoPGT, indicating high levels of labeling, and importantly, visualization also showed very low levels of non-specific protein labeling. These labeling results exemplify how effective Az-1 and Az-2 are at targeting the highly-conserved monoPGT catalytic dyad, and how these probes label other members of this expansive superfamily. Based on several biological replicates, we did not observe a significant difference in reactivity of Az-1 and Az-2 towards monoPGTs. Native expression levels of monoPGTs, such as PglC, are expected to be too low to visualize labeling by gels and Western Blot. Labeling of CEF from Campylobacter jejuni NCTC 11168 was carried out, however no labeled PglC was observed, likely due to low endogenous expression levels. Labeling of CEF from Campylobacter jejuni NCTC 11168 was carried out, however no labeled PglC was observed, likely due to low endogenous expression levels (data not shown). We assessed the sensitivity of the current probe and have determined that monoPGTs at concentrations down to 300 nM can be observed by fluorescence in gels of CEF by Az-1 labeling (Figure S3).
Figure 4.
Western blot analysis of CEF containing overexpressed PglC-His6 in BL21, compared to CEF without any overexpressed proteins. The left most Western blot shows the fluorescence read out from the streptavidin fluorophore conjugate, and the middle shows the same Western blot imaged with Anti-His6 fluorophore conjugate. The right most is a gel with the total protein stained by Coomassie. Analysis with A) Cb and B) Hp.
In summary, we have demonstrated that we can covalently target the active site Asp in the catalytic Asp-Glu by employing 2H-Az probes. We have established active site-specific covalent labeling of monoPGTs from complex cellular fractions based on simple efficient gel-based read outs, and carried this out with representative monoPGTs, including PglC from C. concisus, PglC from C. jejuni, PglC from H. pullorum, and PglB from C. botulinum.
This work provides a strong foundation for employing activity-based protein profiling (ABPP) as a tool for understanding the activity and regulation of the monoPGT superfamily. Future work will focus on increasing the specificity of the probes to support ABPP of monoPGTs at the low native expression levels. To achieve this goal, we will use the presented synthetic route to tailor a small library of probes to monoPGTs. Specifically, we will utilize previously described monoPGTs inhibitors27 as templates and will incorporate small structural components of the best inhibitors via addition to the sulfonyl chloride linker. We will then identify the best structural features that enhance binding affinity to carry out more targeted biological profiling. With targeted ABPP probes in hand, characterization of unknown glycoconjugate biosynthesis pathways in pathogenic bacteria can begin, with the ultimate goal of identification of new mono PGTs as targets for mechanistic analysis and screening for inhibitors in cells and cell membrane fractions.
ABBP has been invaluable to proteomics providing the ultimate springboard for enzyme discovery, and has greatly contributed to our understanding of myriad of diverse enzyme classes including metalloproteins,28 oxidoreductases,29 and glutathione S-transferases.30 With these comprehensive ABPP studies we are now poised to accelerate monoPGT research in an analogous path.
Supplementary Material
ACKNOWLEDGMENT
The authors thank S. McNamara (Haverford College) for providing edits to the manuscript.
Funding Sources
Financial support from the National Institute of Health (GM131627 and GM039334 to BI) and the MIT RSC Fund is gratefully acknowledged.
ABBREVIATIONS
- ABPP
activity based protein profiling
- PGT
phosphoglycosyl transferase
- mono
monotopic
- NDP
nucleoside diphosphate
- GT
glycosyltransferase
- LPS
lipopolysaccharide
- CPS
capsular polysaccharide
- PG
peptidoglycan
- CEF
cell envelope fraction
- SSN
sequence similarity network
- UDP
uridine diphosphate
- UMP
uridine monophosphate
- Az
azirine
- Cj
Campylobacter jejuni
- Cc
Campylobacter concisus
- Hp
Helicobacter pullorum
- Cb
Clostridium botulinum
- Pren-P
polyprenol phosphate
- diNAcBac
di-N-acetylbacillosamine
- Pgl
protein glycosylation
- SUMO
Small Ubiquitin-like Modifier
Footnotes
Supporting Information. The Supporting Information is available free of charge on the ACS Publications website http://pubs.acs.org.
Supporting experimental methods include protein expression and CEF preparation, protocols for gels and Western blots, detailed synthetic procedures and full structural characterization of new compounds.
The authors declare no competing financial interest.
REFERENCES
- 1.Price NP; Momany FA, Modeling bacterial UDP-HexNAc: polyprenol-P HexNAc-1-P transferases. Glycobiology 2005, 15 (9), 29R–42R. [DOI] [PubMed] [Google Scholar]
- 2.Tytgat Hanne LP; Lebeer S, The Sweet Tooth of Bacteria: Common Themes in Bacterial Glycoconjugates. Microbiol. Mol. Biol. Rev 2014, 78 (3), 372–417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Whitfield C; Trent MS, Biosynthesis and export of bacterial lipopolysaccharides. Annu. Rev. Biochem 2014, 83, 99–128. [DOI] [PubMed] [Google Scholar]
- 4.Whitfield C, Biosynthesis and Assembly of Capsular Polysaccharides in Escherichia coli. Annu. Rev. Biochem 2006, 75 (1), 39–68. [DOI] [PubMed] [Google Scholar]
- 5.Bouhss A; Trunkfield AE; Bugg TDH; Mengin-Lecreulx D, The biosynthesis of peptidoglycan lipid-linked intermediates. FEMS Microbiol. Rev 2008, 32 (2), 208–233. [DOI] [PubMed] [Google Scholar]
- 6.Hartley MD; Imperiali B, At the membrane frontier: a prospectus on the remarkable evolutionary conservation of polyprenols and polyprenyl-phosphates. Arch. Biochem. Biophys 2012, 517 (2), 83–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lehrer J; Vigeant Karen A; Tatar Laura D; Valvano Miguel A, Functional Characterization and Membrane Topology of Escherichia coli WecA, a Sugar-Phosphate Transferase Initiating the Biosynthesis of Enterobacterial Common Antigen and O-Antigen Lipopolysaccharide. J. Bacteriol. Res 2007, 189 (7), 2618–2628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chung BC; Zhao J; Gillespie RA; Kwon D-Y; Guan Z; Hong J; Zhou P; Lee S-Y, Crystal Structure of MraY, an Essential Membrane Enzyme for Bacterial Cell Wall Synthesis. Science 2013, 341 (6149), 1012–1016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Brandish PE; Kimura KI; Inukai M; Southgate R; Lonsdale JT; Bugg TD, Modes of action of tunicamycin, liposidomycin B, and mureidomycin A: inhibition of phospho-N-acetylmuramyl-pentapeptide translocase from Escherichia coli. Antimicrob. Agents Chemother 1996, 40 (7), 1640–1644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Das D; Kuzmic P; Imperiali B, Analysis of a dual domain phosphoglycosyl transferase reveals a ping-pong mechanism with a covalent enzyme intermediate. Proc. Natl. Acad. Sci 2017, 114 (27), 7019–7024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.O'Toole KH; Bernstein HM; Allen KN; Imperiali B, The surprising structural and mechanistic dichotomy of membrane-associated phosphoglycosyl transferases. Biochem. Soc. Trans 2021, 49 (3), 1189–1203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.O’Toole KH; Imperiali B; Allen KN, Glycoconjugate pathway connections revealed by sequence similarity network analysis of the monotopic phosphoglycosyl transferases. Proc. Natl. Acad. Sci 2021, 118 (4), e2018289118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Rausch M; Deisinger JP; Ulm H; Müller A; Li W; Hardt P; Wang X; Li X; Sylvester M; Engeser M; Vollmer W; Müller CE; Sahl HG; Lee JC; Schneider T, Coordination of capsule assembly and cell wall biosynthesis in Staphylococcus aureus. Nat. Commun 2019, 10 (1), 1404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Seddon AM; Curnow P; Booth PJ, Membrane proteins, lipids and detergents: not just a soap opera. Biochim. Biophys. Acta. Biomembr 2004, 1666 (1), 105–117. [DOI] [PubMed] [Google Scholar]
- 15.Liu Y; Patricelli MP; Cravatt BF, Activity-based protein profiling: The serine hydrolases. Proc. Natl. Acad. Sci 1999, 96 (26), 14694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cravatt BF; Wright AT; Kozarich JW, Activity-Based Protein Profiling: From Enzyme Chemistry to Proteomic Chemistry. Annu. Rev. Biochem 2008, 77 (1), 383–414. [DOI] [PubMed] [Google Scholar]
- 17.Weerapana E; Simon GM; Cravatt BF, Disparate proteome reactivity profiles of carbon electrophiles. Nat. Chem. Biol 2008, 4 (7), 405–407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shin N-Y; Liu Q; Stamer SL; Liebler DC, Protein Targets of Reactive Electrophiles in Human Liver Microsomes. Chem. Res. Toxicol 2007, 20 (6), 859–867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang H; Wen J; Huang RYC; Blankenship RE; Gross ML, Mass spectrometry-based carboxyl footprinting of proteins: Method evaluation. Int. J. Mass Spectrom 2012, 312, 78–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wu L; Armstrong Z; Schröder SP; de Boer C; Artola M; Aerts JMFG; Overkleeft HS; Davies GJ, An overview of activity-based probes for glycosidases. Current Opinion in Chemical Biology 2019, 53, 25–36. [DOI] [PubMed] [Google Scholar]
- 21.Wicki J; Williams SJ; Withers SG, Transition-State Mimicry by Glycosidase Inhibitors: A Critical Kinetic Analysis. J. Am. Chem. Soc 2007, 129 (15), 4530–4531. [DOI] [PubMed] [Google Scholar]
- 22.Armstrong Z; Kuo C-L; Lahav D; Liu B; Johnson R; Beenakker TJM; de Boer C; Wong C-S; van Rijssel ER; Debets MF; Florea BI; Hissink C; Boot RG; Geurink PP; Ovaa H; van der Stelt M; van der Marel GM; Codée JDC; Aerts JMFG; Wu L; Overkleeft HS; Davies GJ, Manno-epi-cyclophellitols Enable Activity-Based Protein Profiling of Human α-Mannosidases and Discovery of New Golgi Mannosidase II Inhibitors. J. Am. Chem. Soc 2020, 142 (30), 13021–13029. [DOI] [PubMed] [Google Scholar]
- 23.Ma N; Hu J; Zhang Z-M; Liu W; Huang M; Fan Y; Yin X; Wang J; Ding K; Ye W; Li Z, 2H-Azirine-Based Reagents for Chemoselective Bioconjugation at Carboxyl Residues Inside Live Cells. J. Am. Chem. Soc 2020, 142 (13), 6051–6059. [DOI] [PubMed] [Google Scholar]
- 24.Acheson D; Allos BM, Campylobacter jejuni Infections: Update on Emerging Issues and Trends. Clin. Infect. Dis 2001, 32 (8), 1201–1206. [DOI] [PubMed] [Google Scholar]
- 25.Swiecicki JM; Santana JT; Imperiali B, A Strategic Approach for Fluorescence Imaging of Membrane Proteins in a Native-like Environment Resource A Strategic Approach for Fluorescence Imaging of Membrane Proteins in a Native-like Environment. Cell Chem. Biol 2020, 27, 1–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Walvoort MTC; Lukose V; Imperiali B, A Modular Approach to Phosphoglycosyltransferase Inhibitors Inspired by Nucleoside Antibiotics. Eur. J. Chem 2016, 22 (11), 3856–3864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Madec AGE; Schocker NS; Sanchini S; Myratgeldiyev G; Das D; Imperiali B, Facile Solid-Phase Synthesis and Assessment of Nucleoside Analogs as Inhibitors of Bacterial UDP-Sugar Processing Enzymes. ACS Chem. Biol 2018, 13 (9), 2542–2550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Saghatelian A; Jessani N; Joseph A; Humphrey M; Cravatt BF, Activity-based probes for the proteomic profiling of metalloproteases. Proc. Natl. Acad. Sci 2004, 101 (27), 10000–10005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Fuerst R; Breinbauer R, Activity-Based Protein Profiling (ABPP) of Oxidoreductases. ChemBioChem 2021, 22 (4), 630–638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Stoddard EG; Killinger BJ; Nair RN; Sadler NC; Volk RF; Purvine SO; Shukla AK; Smith JN; Wright AT, Activity-Based Probes for Isoenzyme- and Site-Specific Functional Characterization of Glutathione S-Transferases. J. Am. Chem. Soc 2017, 139 (45), 16032–16035. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.