Standfirst
Critical technological advances have enabled the rapid investigations into the immune responses elicited by SARS-CoV-2, the pathogen responsible for the COVID-19 pandemic. In this Comment, we discuss the cutting-edge methods used to deconvolute the B cell responses against this virus, and the significant impact they have had in the ongoing public health crisis.
The COVID-19 pandemic is extracting an enormous toll on human populations worldwide. As overwhelming evidence has accumulated to indicate the participation of inflammatory and autoimmune responses in adverse outcomes, there is also a major need to understand the immunological underpinnings of protective and pathogenic responses to an acute, life-threatening novel virus for which there was little if any preceding immunological memory despite the circulation of other common endemic coronaviruses. The confluence of public health need, scientific opportunity and unparalleled technological and computational tools has provided a unique opportunity to understand the underpinnings and broad heterogeneity of the human immune response in general, and in particular in the context of primary immune responses. The study of B cell responses in this context, whose effective activation is responsible for antibody production in both vaccination and infection, has been a critical point of focus throughout the pandemic in understanding natural immunity development against SARS-CoV-2, vaccine longevity, and memory durability against emergence viral variants.
The rapid development of technology around immunologic investigation generally, and B cell response monitoring specifically, has resulted in a robust experimental toolset capable of extracting significant data down to the single-cell level (Fig. 1). The emergence of these tools, and their application to critical areas of human health such as vaccination1, infection, and autoimmunity2 has allowed for the creation of a framework for B cell response classification and development. Advances in surface phenotyping have led to an increased depth of B cell subset identification and correlated function3. Next-generation sequencing has provided understanding of developmental B cell programs4, with single-cell technology promising to push those efforts even further. Broad antigen-specific screening technologies alongside robust monoclonal antibody (mAb) production pipelines have combined to understand emerging antigen-specific responses and rapidly evaluate potential therapeutics5.
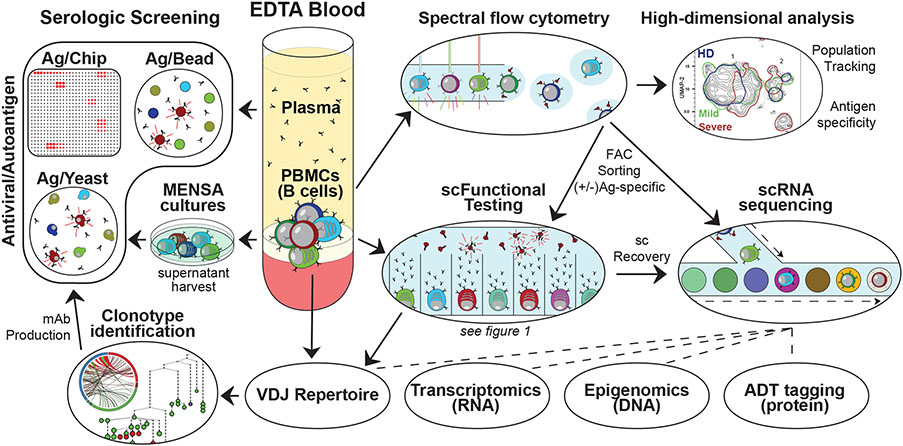
Figure 1. Efficient utilization of patient blood in B cell-focused investigations of COVID-19.
Routine EDTA-tube blood collection from human patients with COVID-19 can be readily processed into plasma and PBMC fractions for downstream investigation. Application of high-dimensional flow cytometry panels to collected PBMCs can reveal alterations in B cell activation pathways, antigen specificity tracking, memory emergence and persistence, and anticipate B cell effector functions. Cell sorting of these analysis platforms allows for the single-cell functional testing of antibody secreting cells to identify clonotype specificity, neutralizing potential, and real-time screening for therapeutic potential. Cells either directly sorted based on fluorescent markers or recovered from single cell functional assays can then be shunted into single cell sequencing applications to investigate the transcriptomic, epigenetic, and repertoire features of interest from selected B cell-derived populations. Using this multi-omics approach, single cells of interest can be identified for mAb production to screen and identify potential binding partners. Resulting mAbs, patient plasma, or ex vivo ASC (MENSA) cultures can then be applied to multiplex antigen screening tools to identify relevant viral and autoantigen reactivities across a multitude of platforms.
With the new technological advances, B cell immunologists are now perfectly poised to rapidly understand mechanisms of viral clearance, disease pathogenesis, and immune protection in both infection and vaccination. Furthermore, some of these novel tools have been successfully deployed to develop mAb therapeutics against SARS-CoV-2 in weeks rather than years5. The result has been an explosion of understanding around humoral immune development in human viral infection. While certainly not a comprehensive list, it is important to document how these technologies have contributed to our collective investigations in dissecting the immune responses surrounding COVID-19.
High-dimensional cytometry
Since its inception, flow cytometry has served as a cornerstone technology in the identification and classification of leukocytes into increasingly refined subpopulations6. As such, its ability to provide increased breadth or depth of cellular characterization is a direct reflection of the number of cellular markers that can be simultaneously and discretely identified with high confidence. In turn, the simultaneous use of multiple markers is critical to study the behavior on an increasing numbers of B cell populations of different functional significance 3. However, the foundational method for detection of those targets -- immunofluorescence -- makes discrete identification of high numbers of targets inherently difficult due to highly overlapping emission spectra of related fluorophores. Attempts to identify highly overlapping fluorophores using traditional technology require careful signal compensation and an unavoidable loss of signal strength.
As a core methodology in immunological labs, several approaches have been tested in bypassing the traditional limitations of fluorescence-based, subtractive compensation-dependent systems. Mass cytometry, where cellular markers are identified through the detection of metal ion-tagged rather than fluorophore-tagged antibodies, have bypassed many of these limitations and introduced the ability to measure in excess of 50 simultaneous markers7. This system is highly sensitive and resistant to the compensation-based errors that can result in erroneous cellular classification, but is lower throughput than its fluorescent-based counterparts and requires intensive reagent quality control. However, due its ability to reliably produce extraordinarily high-dimensional datasets, this approach, continues to find increased applications in immunology, including recently in COVID studies8.
A notable new generation of cytometers, spectral flow cytometers, have replaced traditional compensation methods in favor of spectral unmixing9. Rapid adoption of this technology has made access to antibody panels discerning 30+ cellular markers relatively commonplace within large research institutions. Rapid application of these panels resulted in the broad characterization of lymphocyte responses across COVID-19 disease severity, including the identification of broad patient ‘immunotypes’ with potential therapeutic implications, within months of the start of the pandemic10. Simultaneously, deeper immune profiling of more restricted populations, such as our own study of B lymphocytes, provided the fine characterizations of these populations and their relation to responses identified in other critical areas of human disease11. Regardless of populations of interest, the high-dimensional nature of these data and the development of approachable analysis tools have made the application of unsupervised analysis methods standard –allowing for a deeper probing of cytometric data to uncover unexpected relationships between cellular markers.
Another critical use of flow cytometry is the identification of antigen-specific B cells using antigens labeled with different fluorochromes. When integrated within multi-marker panels, this approach is central for the identification of the specific source of early and late anti-COVID B cell responses, including the memory compartments generated in response to previous infection or vaccination. The enumeration of such cells in different memory compartments with different requirements for induction, maintenance, and re-activation will be critical for our ability to predict, measure, and manipulate the abundance of the more desirable memory responses. A frequently ignored implication of memory studies is the possibility that different vaccines in individual subjects might induce protective B cell responses not reflected in serum antibody titers due to uncoupling of the generation of memory B cells and antibody-producing plasma cells as well as differences in the longevity of the two compartments. This scenario would advocate for the measurement of antigen-specific memory responses in addition to antibody titers to monitor vaccine efficacy and lasting immunity.
Single-cell multi-omics and B cell repertoire
The capabilities and value of single-cell sequencing is now well established across disciplines and was used to great effect in the earliest characterizations of the inflammatory profiles that define severe COVID-1912. In a rapidly developing field, current iterations of single-cell technologies enable multi-omics assessment extending well beyond the transcriptome – making available the simultaneous evaluation of cell-surface phenotypes13, antigen specificity14, chromatin accessibility15, and adaptive immune repertoire16. The vast depth of information available through these combined approaches has propelled single-cell sequencing to the forefront of immunologic investigation into COVID-1916,17.
However, while single-cell sequencing technology has proven its worth in a variety of applications, the unique biological properties of the BCR make it particularly appealing for understanding the developmental pressures associated with the rapidly developing humoral immune response. As in T cell repertoire analysis, by sequencing recombined VDJ sequences at a single-cell level, clonotypes can be readily identified with repeated identification of sequences directly indicating a lineage expansion event18. Using these expanded clonotypes as indicators of selection, v or j-gene biases can be identified and characterized in the context of acute infection18. In addition to these simple assessments, the somatic alterations across the BCR resulting from B cell selection, namely somatic hypermutation and class-switch recombination, provide a wealth of information beyond gene-selection asymmetries that are unavailable in other cell types. These additional parameters, when appropriately utilized, contribute detailed information about the developmental trajectories of individual clonotypes, positive and negative selective pressures acting on the compartment, and even the likely cytokine milieu of the microenvironment. These types of analyses, that have been used to great effect in the identification of disrupted mechanisms of tolerance in human autoimmunity19, have now been wielded similarly in COVID-19 to begin to answer long-standing questions about the emergence of autoreactive antibody responses in primary viral infection20.
The combination of single-cell repertoire alongside traditional transcriptomics and antibody derived tag (ADT)-based sequencing allows for new compiled analysis pipelines21 – providing the potential for extraordinarily detailed developmental trajectories in primary human immunity. While much of this work has focused on blood, the increasing sampling of tissue collection outside of the blood, including fine needle aspirates from human secondary lymphoid tissue, has increasingly allowed probing of developmental B cell responses directly at the differentiation site22. These studies are exciting, feasible, and promise to close many of the gaps in current understanding of both extrafollicular and germinal center B cell origins.
ASC functional assessment and clonotype specificity testing
A commonly identified feature of B cell responses in severe COVID-19 has been the rapid expansion of antibody secreting cells (ASC; ie. plasma cells and plasmablasts) in the blood in the acute phase of infection. Despite their presumed contribution to the early antiviral antibody response, the specificities of these cells are difficult to assess due to their down-regulation of cell-surface B cell receptor rendering traditional cytometry-based antigen-specificity assays ineffective. Intracellular flow cytometry (ICS) methods of antigen-specific Ig staining have been pursued, but the need for cell fixation makes the technique incompatible with downstream antibody discovery. Historically, ELISpot assays have been performed on these cells to identify antigen-specific antibody secretion although these assays are low-throughput, targeted at a single antigen, and require significant numbers of fresh cells for each individual assay. However, a new approach has emerged to help parse the contribution of circulating ASC to the overall antiviral response. Through short-term ASC culture and collection of Media Enriched with Newly Synthesized Antibodies (MENSA), the bulk specificities of circulating ASC enriched following infection can be readily assessed23,24. MENSA collection and testing offers the ability to interrogate multiple antigens simultaneously, as well as assess neutralizing function to provide a snapshot of the antibodies from circulating early-minted ASC in contrast to the serum or plasma which offers the entire immunological/microbial history. Hence, MENSA provides a unique immune snapshot of antibody specificities from only the new active infection compared to historical serum antibody measurements.
While short-term ASC culture is informative, longer-term ex vivo study can be required for functional assessment. Despite the amenability of some lymphocyte populations to long-term culture (including memory B cells), circulating ASC have lagged significantly due to the predisposition of these terminally differentiated B cells to die early in culture25. In vivo, long-lived plasma cells (LLPC) require specialized microniches for maturation and survival. By harnessing factors secreted by the BM mesenchymal stromal cells, and providing survival signals and a hypoxic environment that mimics the natural BM microniche25, we have developed a cell-free plasma cell survival system (PCSS) that allows for the survival and maturation of these temperamental cells for months25 (Fig 2). This system enables interrogation of intrinsic survival programs driving ASC maturity and longevity26,27, and may provide a useful platform for directed genetic editing of ASCs ex vivo for therapeutic use28.
Figure 2. Function-based pre-selection of Ag-specific ASC.
(Left) Development of plasma cell survival system (PCSS). Primary mesenchymal stromal cells (MSC) derived from human bone marrow (BM) are expanded and irradiated. The resulting culture medium is then collected and supplemented with APRIL and hypoxic conditions to promote ASC survival. (Middle) Single cell detection of antigen (Ag)-specific ASC by an in-channel binding assay. A single-assay mixture of Ag-coated beads and fluorescently labeled detection antibodies (Abs) is imported into the channel above the NanoPen chambers. PCSS is perfused through the chip to maintain survival of penned ASC, whose secreted Abs diffuse into the channel and bind the beads. Accumulation of fluorescence on the beads leads to the development of fluorescent halos (“blooms”) in the channel adjacent to the pens that contain Ag-specific ASC. (Right) Single cell functional screening for Ag-specific ASC by an in-pen receptor blocking assay. After positioning Ag-coated beads into pens and saturating the Ag, an assay mixture of fluorescent receptor (ACE2) and fluorescently labeled detection Abs targeting Ag-specific (RBD) binding Abs is imported into the channel and allowed to diffuse into pens. PCSS is perfused through the chip to maintain survival of penned ASC. (Top panel) RBD-binding, ACE2 non-blocking Abs result in accumulation and detection of both fluorophores; (bottom panel) ACE2 blocking Abs exhibit only signals for RBD-binding Abs. ASC secreting blocking Abs are then exported from specific pens for downstream interrogation.
Therapeutic monoclonal antibody identification
Studies of circulating ASC, generally, alongside antigen-specific memory has been critically important in discovery of functional mAb against COVID-19 for use as therapeutics or prophylaxis. Classic approaches to single cell screening and hybridoma creation can require years, and even high-throughput techniques such as emulsion-based single cell methods struggle to differentiate antigen binding from neutralization. Fortunately, a recent technology developed by Berkeley Lights, Inc. (BLI) offer the ability to prescreen ASC for neutralizing potential on a single cell basis to then capture mRNA from these cells. The BLI technology uses OptoElectroPositioning (OEP), which uses light-induced electrokinetics to position and manipulate individual cells in subnanoliter chambers29 (Fig 2). Using PCSS to stabilize these cells, this optofluidic platform accelerated discovery of mAbs for therapeutic potential during the COVID-19 pandemic5,30 and resulted in the highly efficacious and widely-distributed mAb therapy developed for pre-exposure prophylaxis 5,31.
A unique advantage of the BLI system is the feasibility and implementation of ASC functional readouts, ie neutralization potency and receptor blockade5 (Fig. 2), which offers increased efficiency and rapid scalability. Emulsion-based single cell methods do not allow for direct interrogation of cells or a secreted product in supernatants, and while other target-based screening microtools allow for direct single cell manipulation and recovery of living cells (microcapillary arrays32, microfluidic chambers33, micro-engraved/microwell-array systems34), they haven’t been shown to accommodate a neutralization assay. Beyond the identification of therapeutics, these novel approaches will allow for continued probing of ASC biology including their potential for proliferation and secretory dynamics. This platform has already yielded results in COVID-19, allowing for the generation of monoclonal and bi-specific antibodies that are effective against spike mutations associated with emerging viral variants5,30,31.
Whether ASCs are identified and collected through function-based assays or more generic single-cell VDJ sequencing, the paired identification of intact heavy and light chain sequences from quality-controlled samples provides an opportunity to directly evaluate antigen specificities of identified clonotypes of interest. Combined with increasingly mechanized gene synthesis and custom antibody production5, the field inches closer to the long-pursued possibility of deeply screening a donors’ circulating B cell repertoire for specificity within the context of identified cell-surface and transcriptomic features. While still costly, this single-cell-to-antibody pipeline is increasingly used to identify functional antigen specificities within unique B cell populations emerging in COVID-19 and characterized directly through single-cell phenotyping20.
Antibody screening
Taking advantage of these single-cell approaches, and the more generalized need for antigen specificity screening at the serological level, COVID-19 has brought focus on a myriad of high-throughput technologies aimed at providing a detailed understanding of broad antibody targeting. Due to its high throughput and easily customizable antigen sets, Luminex-based fluorescent and Meso Scale platforms have become an invaluable tool for antiviral specificity assessment across comprehensive sets of SARS-CoV-2-based antigens11,35-37. Similarly, targeted systems have been developed through both chip-based and flow-based technologies, with multiplex identification of COVID-19 antigen specificity emerging as a cornerstone approach to serological assessment throughout the pandemic.
In addition to antiviral reactivity, a surprising connection between the B cell responses present in severe COVID-19 and those identified in patients with active autoimmune disorders has driven a need for comprehensive assessments of both antiviral and self-targeted antibodies11. With no clear standard approach for these screening strategies, groups have relied on commercially available protein arrays reflecting the entire human proteome to identify self-targeted antibody responses in COVID-1931, phage-display-based screening of antiviral targeting32, or complex combinations of in-house-validated bead-based platforms to synthesize antiviral and anti-self targeting into a comprehensive strategy33. A notable emerging technology, the REAP system, offers the potential to screen an expansive selection of the exoproteome for autoreactivity and has been used to great effect in the identification of autoreactivity in COVID-1938. Commercial avenues for large-scale autoreactivity testing have also emerged as widely used, albeit expensive technologies requiring a fair amount of individual target validation39. Importantly, while these tools have application in identifying broad autoreactivity against a variety of targets, existing FDA-approved autoantibody testing pipelines routinely employed in healthcare settings should be considered as established methods to identifying clinically relevant autoreactive features of severe infectious disease20.
Conclusions
The questions informed by these various technologies have played a critical role in the extraordinarily public investigation of COVID-19. High-dimensional cytometry paired with single-cell multi-omics and antibody repertoire analysis have helped reveal the intensity, quality, and even pathological potential of the B cell compartment in severe COVID-19. Multiplex antibody screening has yielded critical information about the duration of responses resulting from both infection and vaccination, and their durability in the face of emerging SARS-CoV-2 variants. Single cell functional studies have allowed for the rapid identification of therapeutic mAbs deployed to great effect throughout the pandemic and may reveal therapeutic targets to intervene in misdirected humoral immune targeting. Together, these technologies have allowed the B cell field to reliably address some of the most rapidly evolving and important questions emerging throughout the pandemic – informing not only scientific understanding, but also human health and public policy.
However, in addition to technological advance, the rapid pace of discovery since November 2019 is a direct reflection of the willingness of the immunological community to alter research trajectories towards unfamiliar and often dangerous work in support of the common good. It is also a testament to the value of continuing technological development aimed at basic science discovery in human immunology. Though targeted funding opportunities and the creation of critical national research consortia (SeroNet, PRISM, and RECOVER, among others), laboratories at the forefront of the investigation of B cell biology were able to deploy new technology on an emerging global threat at an unprecedented pace. Now firmly embedded in the COVID-19 literature, these technologies, in our opinion will serve as a benchmark for future investigations and applications – driving critical understanding of developing humoral immunity and B cell effector responses even beyond the realm of infectious disease.
Despite the unprecedented pace of developments enabled by the confluence of technological and scientific prowess, there remains a need for additional advances in our ability to interrogate the B cell and antibody immune response to SARS-CoV2 infection and vaccination. Areas of particular need include extensive antigen platforms for the analysis of autoreactivity of serum and monoclonal antibodies. While multiple such platforms currently exist, there is a need for robust and affordable screening systems which in turn should be subjected to cross-validation. Similarly, it will be important to continue to develop methods for high-throughput identification of anti-viral and autoreactive antibody-secreting cells. These capabilities would enable investigators to readily determine the protective and pathogenic potential of serum antibodies generated in response to vaccination, and of monoclonal antibodies considered as candidates for passive prevention and treatment of COVID-19 infection. In all, a final goal should be the generation of B cell and antibody signatures associated with favorable responses to infection and vaccination, and conversely, capable of predicting acute and long-term autoimmune complications.
Acknowledgments
This work was supported by National Institutes of Health grants: UL TR000424 (Emory Library IT), U54-CA260563-01 Emory SeroNet (I.S., F.E.L.), U19-AI110483 Emory Autoimmunity Center of Excellence (I.S.), P01-AI125180-01 (I.S., F.E.L.), R37-AI049660 (I.S.), 1R01AI12125 (F.E.L.), 1U01AI141993 (F.E.L). Bill and Melinda Gates Foundation: INV-002351 (F.E.L.)
Footnotes
Competing Interest Statement
Dr. Lee is the founder of MicroB-plex, Inc and have patents with the MENSA technology, and has grants with the Gates foundation and Genentech. Drs. Lee, Nguyen, and Sanz have patents with the PCSS. The authors have collaborated with Exagen, Inc. to provide clinical autoreactivity testing in several COVID-19-related studies.
References
- 1.Halliley JL et al. Long-Lived Plasma Cells Are Contained within the CD19(−)CD38(hi)CD138(+) Subset in Human Bone Marrow. Immunity 43, 132–145, doi: 10.1016/j.immuni.2015.06.016 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Jenks SA et al. Distinct Effector B Cells Induced by Unregulated Toll-like Receptor 7 Contribute to Pathogenic Responses in Systemic Lupus Erythematosus. Immunity 49, 725–739 e726, doi: 10.1016/j.immuni.2018.08.015 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sanz I et al. Challenges and opportunities for consistent classification of human B cell and plasma cell populations. Frontiers in immunology 10, 2458 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Scharer CD et al. Epigenetic programming underpins B cell dysfunction in human SLE. Nat Immunol 20, 1071–1082, doi: 10.1038/s41590-019-0419-9 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zost SJ et al. Rapid isolation and profiling of a diverse panel of human monoclonal antibodies targeting the SARS-CoV-2 spike protein. Nat Med 26, 1422–1427, doi: 10.1038/s41591-020-0998-x (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Herzenberg LA et al. The history and future of the fluorescence activated cell sorter and flow cytometry: a view from Stanford. Clin Chem 48, 1819–1827 (2002). [PubMed] [Google Scholar]
- 7.Palit S, Heuser C, de Almeida GP, Theis FJ & Zielinski CE Meeting the Challenges of High-Dimensional Single-Cell Data Analysis in Immunology. Front Immunol 10, 1515, doi: 10.3389/fimmu.2019.01515 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Wang W et al. High-dimensional immune profiling by mass cytometry revealed immunosuppression and dysfunction of immunity in COVID-19 patients. Cell Mol Immunol 17, 650–652, doi: 10.1038/s41423-020-0447-2 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ferrer-Font L et al. Panel Optimization for High-Dimensional Immunophenotyping Assays Using Full-Spectrum Flow Cytometry. Curr Protoc 1, e222, doi: 10.1002/cpz1.222 (2021). [DOI] [PubMed] [Google Scholar]
- 10.Mathew D et al. Deep immune profiling of COVID-19 patients reveals distinct immunotypes with therapeutic implications. Science 369, doi: 10.1126/science.abc8511 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Woodruff MC et al. Extrafollicular B cell responses correlate with neutralizing antibodies and morbidity in COVID-19. Nat Immunol 21, 1506–1516, doi: 10.1038/s41590-020-00814-z (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Arunachalam PS et al. Systems biological assessment of immunity to mild versus severe COVID-19 infection in humans. Science 369, 1210–1220, doi: 10.1126/science.abc6261 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Su Y et al. Multi-Omics Resolves a Sharp Disease-State Shift between Mild and Moderate COVID-19. Cell 183, 1479–1495 e1420, doi: 10.1016/j.cell.2020.10.037 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dugan HL et al. Profiling B cell immunodominance after SARS-CoV-2 infection reveals antibody evolution to non-neutralizing viral targets. Immunity 54, 1290–1303 e1297, doi: 10.1016/j.immuni.2021.05.001 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.You M et al. Single-cell epigenomic landscape of peripheral immune cells reveals establishment of trained immunity in individuals convalescing from COVID-19. Nat Cell Biol 23, 620–630, doi: 10.1038/s41556-021-00690-1 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang JY et al. Single-cell landscape of immunological responses in patients with COVID-19. Nat Immunol 21, 1107–1118, doi: 10.1038/s41590-020-0762-x (2020). [DOI] [PubMed] [Google Scholar]
- 17.Unterman A et al. Single-cell multi-omics reveals dyssynchrony of the innate and adaptive immune system in progressive COVID-19. Nat Commun 13, 440, doi: 10.1038/s41467-021-27716-4 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Nielsen SCA et al. Human B Cell Clonal Expansion and Convergent Antibody Responses to SARS-CoV-2. Cell Host Microbe 28, 516–525 e515, doi: 10.1016/j.chom.2020.09.002 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Tipton CM et al. Diversity, cellular origin and autoreactivity of antibody-secreting cell population expansions in acute systemic lupus erythematosus. Nat Immunol 16, 755–765, doi: 10.1038/ni.3175 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Woodruff MC et al. Relaxed peripheral tolerance drives broad de novo autoreactivity in severe COVID-19. medRxiv, doi: 10.1101/2020.10.21.20216192 (2021). [DOI] [Google Scholar]
- 21.Babcock Benjamin R., K. A, Yang Junkai, White Mackenzie L., Ghosn Eliver E. B.. Data Matrix Normalization and Merging Strategies Minimize Batch-specific Systemic Variation in scRNA-Seq Data. bioRxiv, doi: 10.1101/2021.08.18.456898 (2021). [DOI] [Google Scholar]
- 22.Turner JS et al. SARS-CoV-2 mRNA vaccines induce persistent human germinal centre responses. Nature 596, 109–113, doi: 10.1038/s41586-021-03738-2 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kyu S et al. Diagnosis of Streptococcus pneumoniae infection using circulating antibody secreting cells. PLoS One 16, e0259644, doi: 10.1371/journal.pone.0259644 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Haddad NS et al. Detection of Newly Secreted Antibodies Predicts Non-recurrence in Primary Clostridioides difficile Infection. J Clin Microbiol, jcm0220121, doi: 10.1128/jcm.02201-21 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Nguyen DC et al. Factors of the bone marrow microniche that support human plasma cell survival and immunoglobulin secretion. Nat Commun 9, 3698, doi: 10.1038/s41467-018-05853-7 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Garimalla S et al. Differential transcriptome and development of human peripheral plasma cell subsets. JCI Insight 4, doi: 10.1172/jci.insight.126732 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Joyner CJ et al. Generation of human long-lived plasma cells by developmentally regulated epigenetic imprinting. Life Sci Alliance 5, doi: 10.26508/lsa.202101285 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Faiq MA B-cell engineering: A promising approach towards vaccine development for COVID-19. Med Hypotheses 144, 109948, doi: 10.1016/j.mehy.2020.109948 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Jorgolli M et al. Nanoscale integration of single cell biologics discovery processes using optofluidic manipulation and monitoring. Biotechnol Bioeng 116, 2393–2411, doi: 10.1002/bit.27024 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Cho H et al. Bispecific antibodies targeting distinct regions of the spike protein potently neutralize SARS-CoV-2 variants of concern. Sci Transl Med 13, eabj5413, doi: 10.1126/scitranslmed.abj5413 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.VanBlargan LA et al. An infectious SARS-CoV-2 B.1.1.529 Omicron virus escapes neutralization by therapeutic monoclonal antibodies. Nat Med, doi: 10.1038/s41591-021-01678-y (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Fitzgerald V et al. Exploiting highly ordered subnanoliter volume microcapillaries as microtools for the analysis of antibody producing cells. Anal Chem 87, 997–1003, doi: 10.1021/ac503547j (2015). [DOI] [PubMed] [Google Scholar]
- 33.Burman L et al. Isolation of monoclonal antibodies from anti-synthetase syndrome patients and affinity maturation by recombination of independent somatic variants. MAbs 12, 1836718, doi: 10.1080/19420862.2020.1836718 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jin A et al. A rapid and efficient single-cell manipulation method for screening antigen-specific antibody-secreting cells from human peripheral blood. Nat Med 15, 1088–1092, doi: 10.1038/nm.1966 (2009). [DOI] [PubMed] [Google Scholar]
- 35.Haddad NS et al. One-Stop Serum Assay Identifies COVID-19 Disease Severity and Vaccination Responses. Immunohorizons 5, 322–335, doi: 10.4049/immunohorizons.2100011 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Runnstrom MC et al. Reduced COVID-19 Vaccine Response in Patients Treated with Biologic Therapies for Asthma. Am J Respir Crit Care Med, doi: 10.1164/rccm.202111-2496LE (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Cillo AR et al. People critically ill with COVID-19 exhibit peripheral immune profiles predictive of mortality and reflective of SARS-CoV-2 lung viral burden. Cell Rep Med 2, 100476, doi: 10.1016/j.xcrm.2021.100476 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wang EY et al. Diverse functional autoantibodies in patients with COVID-19. Nature, doi: 10.1038/s41586-021-03631-y (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wong AKH et al. Broad auto-reactive IgM responses are common in critically ill patients, including those with COVID-19. Cell Rep Med 2, 100321, doi: 10.1016/j.xcrm.2021.100321 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]